Abstract
This is the first published report of a PCR assay for detecting porcine cytomegalovirus (PCMV), the causative agent of inclusion body rhinitis in pigs. The DNA to be tested was extracted directly from lungs and nasal scrapings of pigs with various clinical syndromes. Fifty-nine percent (74 of 126) of tested pigs with various clinical syndromes were found to be PCR positive for PCMV. It is hoped that veterinary diagnostic laboratories will benefit by using this PCR assay for routine testing and surveillance of PCMV in pigs.
Porcine cytomegalovirus (PCMV) causes inclusion body rhinitis and abortion or neonatal piglet losses in pigs (3, 9). Cytomegalovirus causes large intranuclear inclusion bodies in cytomegalic cells (9). In pigs, a major site of infection tends to be the turbinates and the rest of the upper respiratory tract. Inclusion body rhinitis is often confused with atrophic rhinitis, an upper respiratory tract disease of multiple etiologies (3, 9). In susceptible herds, infection with PCMV can lead to fetal and piglet death, runting, rhinitis, pneumonia, and poor weight gain. In herds where management conditions tend to be good or exceptional, the virus may be endemic without causing any apparent clinical disease or economic loss (3). Inclusion body rhinitis is characterized by intranuclear inclusions in macrophages in the lungs and in tubuloalveolar gland cells in the nasal mucosa and by small intranuclear inclusions in reticuloendothelial cells (3). Antibodies to this virus have been found in a high percentage of swine herds worldwide (2, 12, 13). Based on these previously reported serology studies, our laboratory was inspired to develop the first reported PCR assay for PCMV, which is described in the present study.
The methodologies for extracting DNA, designing and synthesizing PCR oligonucleotide primers, performing the PCR, restriction enzyme digestion, and gel electrophoresis, and photographing samples are described in detail in other studies reported by our laboratory (7, 8). The primer sequences used for the PCMV PCR assay were 5′-CCCTGATCTTAAATGACGAGGACGTGAC-3′ and 5′-ACCGTCTGAGAGACTGAACTTCTCTGACAC-3′, corresponding to nucleotide sequence positions 37 to 64 and 449 to 420, respectively, of the PCMV polymerase gene, represented by GenBank accession number AJ222640. These PCR primer sequences were found by BLAST computer analysis (1) to possess similarity with the PCMV polymerase gene and not with any other sequences found in the GenBank database (data not shown).
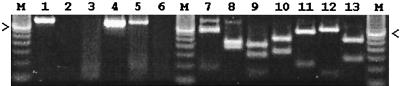
Case submissions consisted of pigs with various clinical syndromes and were brought directly to our facility from farms in Manitoba and other Canadian provinces. The DNA tested by this PCR assay was extracted directly from the lungs of randomly chosen pigs or from lungs and nasal scrapings of pigs suspected to have inclusion body rhinitis. Figure 1 shows some examples of the PCR assay for PCMV, including restriction enzyme analysis. All PCR-positive amplification products were the expected size of 413 bp.
FIG. 1.
Detection of PCMV by PCR. Samples were as follows: positive-control PCMV from cell culture (lane 1), PCMV-negative control with water instead of DNA (lane 2), PCMV-negative porcine lung (lane 3), PCMV-positive porcine lung (lane 4), PCMV-positive porcine nasal scraping (lane 5), PCMV-negative porcine nasal scraping (lane 6). PCR amplification products from PCMV-positive tissue were identified by restriction enzyme analysis by digestion with the following: AciI (lane 7), AluI (lane 8), DdeI (lane 9), HinPI (lane 10), HinfI (lane 11), RsaI (lane 12), and XbaI (lane 13). The 350-bp fragment of a 50-bp DNA ladder (lanes M) (Bethesda Research Laboratories, Inc.) is indicated by the arrows in the right and left margins.
The predicted positions and sizes of fragments from PCMV cleavage by several restriction endonucleases are listed in Table 1. The amplification products from 22 randomly chosen PCR-positive cases were treated with restriction endonucleases, and all yielded the cleavage fragments of the expected size (data not shown).
TABLE 1.
List of restriction enzyme digests of the 413-bp PCR product of PCMV
| Enzymea | Digest position(s)b | Fragment sizes (bp) |
|---|---|---|
| AciI (CCGC) | 301, 324 | 300, 90, 23 |
| AluI (AGCT) | 222 | 221, 192 |
| BstUI (CGCG) | 168 | 246, 167 |
| DdeI (CTNAG) | 142, 334 | 192, 141, 71, 9 |
| HinPI (GCGC) | 166, 168 | 246, 165 |
| HinfI (GANTC) | 109 | 305, 108 |
| RsaI (GTAC) | 82 | 332, 81 |
| XbaI (TCTAGA) | 133, 373 | 240, 132, 41 |
The name of each restriction enzyme is given with the nucleotide sequence of its recognition site in parentheses.
Digest positions correspond to nucleotide positions in the 413-bp PCR amplification product represented in Fig. 1.
In total, 59% (74 of 126) of pigs, 65% (47 of 62) of cases, and 67% (46 of 58) of different farms tested positive by PCR for the presence of PCMV. Case histories and postmortem examinations revealed that only 59% (44 of 74) of PCR-positive pigs (35% [44 of 126] of all tested pigs) had clinical signs and lesions consistent with inclusion body rhinitis. This should be of little surprise, considering that the virus may be endemic without causing any apparent clinical disease or economic loss in well-managed herds (3). Indeed, most of the pigs tested were from well-managed, “high health, high biosecurity” farms.
The PCR assay was tested for specificity by assaying DNA extracted from tissues that had previously tested positive for the following organisms: bovine viral diarrhea virus, equine rhinopneumonitis virus, infectious bovine rhinotracheitis virus, infectious laryngotracheitis virus, Lawsonia intracellularis, Marek's disease herpesvirus 1, Mycoplasma hyopneumoniae, type-2 porcine circovirus, and swine influenza virus. None of these specimens yielded any detectable amplification products or artifacts (data not shown). Likewise, DNA extracted directly from lungs, tonsils, or lymph nodes of PCMV-negative pigs failed to produce any detectable amplification products or artifacts (Fig. 1).
PCMV appears to be present in a high percentage (59%) of the diseased pigs that were tested in our facility. Although this was a preliminary survey of a small number of diseased pigs, PCMV appears to be common in many Canadian swine herds. The PCR assay detected PCMV in all 44 pigs which were found by postmortem examination to have clinical signs and lesions consistent with inclusion body rhinitis. PCMV was detected by PCR in an additional 30 pigs, each with various clinical signs and lesions which were not easily assigned to any single disease pathogen. This appears to be a common observation for a few clinical conditions of pigs. Type 2 porcine circovirus and porcine reproductive and respiratory syndrome virus cause a wide range of similar clinical signs and lesions in pigs, and many of these symptoms are difficult to differentiate from one another (4). Indeed, the guidelines for defining a causal relationship between a microbe and a disease (Koch's postulates) need to be rewritten in light of recent applications of new diagnostic technologies such as the PCR (5).
Based on our laboratory's extensive experience with using PCR to detect other infectious disease agents, we believe that this technology will prove to be a powerful diagnostic tool (7, 8, 10, 11). In our laboratory, PCR assays are reliable, fast, sensitive, cost-effective, and versatile. PCR versatility can be demonstrated by using restriction enzymes to digest PCR amplification products in order to characterize and identify them and to perform molecular epidemiology studies (6). Hopefully, the PCR assay for PCMV described in this report will prove useful to veterinary diagnostic laboratories for the routine testing and surveillance of PCMV infection in pigs.
Acknowledgments
We thank the veterinary pathologists, technologists, and clerical staff of the Veterinary Services Branch Diagnostic Laboratory for their contributions to diagnostic reports.
REFERENCES
- 1.Altschul S F, Gish W, Miller W, Myers E W, Lipman D J. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 2.Assaf R, Bouillant A M, Di Franco E. Enzyme linked immunosorbant assay (ELISA) for the detection of antibodies to porcine cytomegalovirus. Can J Comp Med. 1982;46:183–185. [PMC free article] [PubMed] [Google Scholar]
- 3.Edington N. Cytomegalovirus. In: Lemen A D, Straw B, Glock R D, Mengeling W L, Penny R H C, Scholl E, editors. Diseases of swine. 6th ed. Ames, Iowa: Iowa State University Press; 1986. pp. 330–336. [Google Scholar]
- 4.Ellis J, Krakowka S, Allan G, Clark E, Kennedy S. The clinical scope of porcine reproductive and respiratory syndrome virus infection has expanded since 1987: an alternative perspective. Vet Pathol. 1999;36:262–265. doi: 10.1354/vp.36-3-262. [DOI] [PubMed] [Google Scholar]
- 5.Fredericks D N, Relman D A. Sequence-based identification of microbial pathogens: a reconsideration of Koch's postulates. Clin Microbiol Rev. 1996;9:18–33. doi: 10.1128/cmr.9.1.18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Gagnon C A, Dea S. Differentiation between porcine reproductive and respiratory syndrome virus isolates by restriction fragment length polymorphism of their ORF 6 and 7 genes. Can J Vet Res. 1998;62:110–116. [PMC free article] [PubMed] [Google Scholar]
- 7.Hamel A L, Wasylyshen M D, Nayar G P S. Rapid detection of bovine viral diarrhea virus by using RNA extracted directly from assorted specimens and a one-tube reverse transcription PCR assay. J Clin Microbiol. 1995;33:287–291. doi: 10.1128/jcm.33.2.287-291.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hamel, A. L., L. Lin, C. Sachvie, E. Grudeski, and G. P. S. Nayar. Unpublished data.
- 9.Narita M, Kawamura H, Shirai J, Haritani M. Morphologic study of inclusions in tissues from pigs inoculated with cytomegalovirus. Am J Vet Res. 1987;48:1398–1402. [PubMed] [Google Scholar]
- 10.Nayar G P S, Hamel A, Lin L. Detection and characterization of porcine circovirus associated with postweaning multisystemic wasting syndrome in pigs. Can Vet J. 1997;38:385–386. [PMC free article] [PubMed] [Google Scholar]
- 11.Nayar G P S, Hamel A L, Lin L, Sachvie C, Grudeski E, Spearman G. Evidence of circovirus in cattle with respiratory disease and in aborted bovine fetuses. Can Vet J. 1999;40:277–278. [PMC free article] [PubMed] [Google Scholar]
- 12.Rondhuis P R, de Jong M F, Schep J. Indirect fluorescence antibody studies of porcine cytomegalovirus infection in the Netherlands. Tijdschr Diergeneeskd. 1980;105:56–68. [PubMed] [Google Scholar]
- 13.Tajima T, Hironao T, Kajikawa T, Kawamura H. Application of enzyme-linked immunosorbent assay for the seroepizootiological survey of antibodies against porcine cytomegalovirus. J Vet Med Sci. 1993;55:421–424. doi: 10.1292/jvms.55.421. [DOI] [PubMed] [Google Scholar]