Abstract
Streptococcus pyogenes isolates obtained in 1981 to 1997 from patients and healthy subjects were characterized by pulsed-field gel electrophoresis (PFGE) patterns, biotyping, and the presence of spe genes encoding streptococcal pyrogenic exotoxins. Changes in the profiles were shown in the serotype M1/T1 isolates from pharyngitis over this period, but not in serotype M3/T3 isolates. The characteristics of isolates from patients with toxic shock-like syndrome (TSLS) were comparable to those of the other isolates, including those from healthy subjects. This finding suggests that further phenotypic and molecular characterization, such as investigating the genomic difference represented by the pathogenicity island, of isolates with apparently the same profiles would be necessary to determine the etiology of diseases caused by S. pyogenes, including TSLS.
Streptococcus pyogenes (group A Streptococcus) is a major etiological agent causing a variety of human diseases ranging from pharyngitis to severe invasive disease, such as toxic shock-like syndrome (TSLS) (13). In the United States, the predominant serotypes of S. pyogenes causing TSLS were reported to be M1 and M3 (4, 14, 16). Molecular profiling of S. pyogenes serotypes M1 and M3 by pulsed-field gel electrophoresis (PFGE) has demonstrated that the majority of episodes of invasive disease in the United States have been caused by clonal spread (3, 7). Identical PFGE patterns had been observed in isolates from patients with both invasive and noninvasive diseases in Japan (8). In order to determine the role of molecular typing in pathogenicity of S. pyogenes, this report compared strains isolated from patients and healthy carriers in Kanagawa Prefecture, Japan, during 1981 to 1997.
A total of 73 clinical isolates of S. pyogenes were obtained from pediatric patients (3 to 9 years old) with pharyngitis living in Kanagawa Prefecture, Japan, who attended sentinel clinics of the National Epidemiological Surveillance of Infectious Diseases between 1981 and 1997. A total of 35 S. pyogenes strains were isolated from asymptomatic healthy children (5 to 10 years old) who were attending 20 different schools in different geographical regions of Kanagawa Prefecture between 1981 and 1985. The strains were examined for Lancefield serogroups and T-protein serotypes by using slide agglutination with commercial rabbit antisera (Denka Seiken Co., Ltd., Tokyo, Japan) (5). The strains with serotypes T1 and T3 were tested for M-protein serotypes by microdiffusion with acid extracts and rabbit antisera as described elsewhere (12). In addition, a total of 20 strains derived from TSLS patients (8 to 83 years old) collected in Japan during 1992 to 1995 had known M and T serotypes and streptococcal pyrogenic exotoxin A (speA), speB, and speC genes (5). It was previously reported (2) that S. pyogenes strains isolated from patients with pharyngitis or superficial skin infection showed changes in their restriction profiles between 1988 and 1989. In this study, we compared S. pyogenes strains obtained from patients with pharyngitis to those obtained from patients who were asymptomatic over a 7- to 8-year interval. Strain comparisons were performed by using PFGE with SmaI-digested (15) and SfiI-digested (7) chromosomal DNA; PCR-detected speA, speB, or speC genes (11); and biotyping with a commercial biochemical test kit (Rapid ID 32 Strep; bioMeriuex, Marcy-l'Etoile, France) (1). Numerical analysis of SmaI-digested patterns was performed by identifying the proportion of fragments shared by pairs of isolates by the method of Nei and Li (9) (also known as the Dice coefficient of similarity) and calculated as F = 2nxy/(nx + ny), where nx and ny are the total numbers of fragments from isolates x and y, respectively, and nxy is the number of fragments identical in the two isolates. An F value for two PFGE patterns of ≥0.80 represents closely related strains (10).
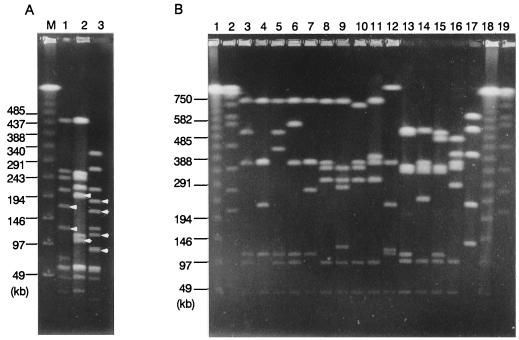
By following the established criteria for bacterial strain typing by PFGE methodology (20), S. pyogenes isolates were classified into types I to III by PFGE analysis with SmaI (Fig. 1A). PFGE patterns with DNA bands of 160 and 120 kb and those with DNA bands of 200 and 100 kb were arbitrarily designated as types I and II, respectively. Those isolates with four DNA bands (180, 155, 110, and 90 kb) were designated as type III. Each of the above PFGE patterns was further classified. Subtypes were determined when the patterns varied by two to four bands compared with the predominant pattern (i.e., Ia, IIa, and IIIa) of each PFGE type. The patterns from isolates within type I were distinguished from a predominant pattern (Ia) by two to four bands and a total of five patterns (Ia, Ib, Ic, Id, and Ie) were recognized (Fig. 2A). Similarly, four to five PFGE patterns were observed in isolates within each of the types II and III (Fig. 2B and C). Since the numbers of differences of bands between the predominant patterns and the others in each of the types ranged from two to four, which can be explained by one to two genetic events (20), the isolates within each PFGE type are considered to be genetically closely related (F values ranging from 0.80 to 0.91). The PFGE pattern of a serotype M3/T3 isolate was untypeable (UT) (Table 1 [PFGE data not shown]). As shown in Fig. 1B, SfiI-digested PFGE patterns were classified into six types (A through F) in a similar manner. The patterns of isolates with pattern types A, B, and C were distinguished by two or three bands from each of the predominant patterns, named A1, B1, and C1, and a total of five, four, and three types respectively, were recognized.
FIG. 1.
(A) Representative PFGE patterns of SmaI-digested chromosomal DNA of S. pyogenes isolates. Lanes: 1, type Ia; 2, type IIa; 3, type IIIa; M, lambda ladders. Arrows indicate the fragments which are common within each of the PFGE types (see text). (B) SfiI-digested patterns of S. pyogenes isolates. Lanes: 1 and 18, lambda DNA ladder; 2 and 19, Saccharomyces cerevisiae DNA; 3, type A1; 4, type A2; 5, type A3; 6, type A4; 7, type A5; 8, type B1; 9, type B2; 10, type B3; 11, type B4; 12, type E; 13, type C1; 14, type C2; 15, type C3; 16, type D; 17, type F. The sizes of the markers (in kilobase pairs) are indicated to the left of each panel.
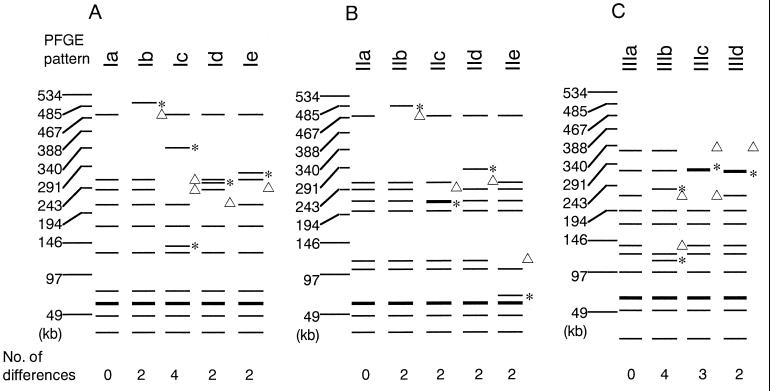
FIG. 2.
Schematic diagram showing the differences between the PFGE patterns and the predominant patterns in each of the serotypes—types Ia (A), IIa (B), and IIIa (C). Triangles indicate fragments present in the predominant patterns and missing from those patterns after a possible genetic event(s); asterisks indicate the fragment is absent from the predominant patterns, but present in the patterns after a possible genetic event(s).
TABLE 1.
Results of typing of S. pyogenes isolates
| PFGE pattern
|
Serotype | spe genotype | Biotype | No. of subjects
|
||||
|---|---|---|---|---|---|---|---|---|
| 1981–1988
|
1989–1997
|
|||||||
| SmaI | SfiI | Healthy | Pharyngitis | Pharyngitis | TSLS | |||
| Ia | A1 | M1/T1 | B | 1 | 1 | 4 | ||
| Ia | A1 | M1/T1 | B | 3 | 9 | 10 | 1 | |
| Ia | A1 | M1/T1 | B | 4 | 2 | 1 | ||
| Ia | A1 | M1/T1 | B | 8 | 1 | |||
| Ia | A1 | M1/T1 | B | UTa | 1 | |||
| Ia | A2 | M1/T1 | B | 1 | 2 | |||
| Ia | A2 | M1/T1 | B | 3 | 2 | |||
| Ib | A3 | M1/T1 | B | 1 | 1 | |||
| Ic | A1 | M1/T1 | B | 3 | 1 | |||
| Id | A4 | M1/T1 | BC | 1 | 1 | |||
| Id | A5 | M1/T1 | BC | 3 | 1 | 1 | ||
| Id | A5 | M1/T1 | BC | 8 | 1 | |||
| Ie | E | M1/T1 | B | UT | 1 | |||
| IIa | B1 | M1/T1 | AB | 1 | 18 | 6 | ||
| IIa | B1 | M1/T1 | AB | UT | 1 | |||
| IIb | B2 | M1/T1 | ABC | 1 | 1 | 1 | ||
| IIc | B3 | M1/T1 | AB | 1 | 1 | |||
| IId | B4 | M1/T1 | ABC | 1 | 3 | |||
| IIe | B3 | M1/T1 | B | 1 | 1 | |||
| IIIa | C1 | M3/T3 | AB | 3 | 15 | 7 | 7 | 5 |
| IIIa | C1 | M3/T3 | AB | 4 | 2 | |||
| IIIa | C2 | M3/T3 | AB | 4 | 3 | 4 | ||
| IIIa | C1 | M3/T3 | AB | 8 | 1 | |||
| IIIa | C1 | M3/T3 | AB | UT | 1 | |||
| IIIb | C1 | M3/T3 | AB | 3 | 2 | 1 | ||
| IIIc | C3 | M3/T3 | AB | 3 | 1 | |||
| IIId | D | M3/T3 | AB | 3 | 1 | |||
| IIId | D | M3/T3 | AB | UT | 2 | 2 | ||
| UT | F | M3/T3 | BC | 1 | 1 | |||
UT, untypeable.
The results of the typing of S. pyogenes isolates in the present study are summarized in Table 1. Isolates from patients with pharyngitis in Japan that belonged to serotype M1/T1 showed appreciable shifts in their phenotypic and genotypic characteristics over the 17-year period between 1981 and 1997. Twenty-four of the 25 serotype M1/T1 isolates in the first 8-year period (1981 to 1988) were classified as SmaI-digested PFGE type I and SfiI-digested type A (SmaI-SfiI:A) and did not carry the speA gene, although they belonged to various biotypes. In contrast, 24 of the 26 isolates in the next 9-year period (1989 to 1997) belonged to PFGE type SmaI:II-SfiI:B and were biotype 1. All of the type SmaI:II strains except one carried the speA gene, which seems to have coincided with the first report of the isolation of the speA-positive strain of serotype M1 in North Carolina in 1989 (6). A chi-square analysis indicated that the number of type SmaI:I-SfiI:A isolates significantly decreased (P < 0.001) and the number of type SmaI:II-SfiI:B isolates increased (P < 0.001) during this period (1981 to 1997). A similar shift was observed within S. pyogenes isolates from patients with pharyngitis in a different part of Japan (Toyama Prefecture) in which SmaI-digested PFGE patterns and biotypes of isolates obtained in 1987 to 1988 were different from those in 1991 to 1993 (19). These data imply that the type SmaI:II-SfiI:B strains in this study may have an etiological advantage, because of either ineffectiveness of previous acquired immunity or inadequate immunity against these strains in the overall population. The PFGE patterns observed in 19 of the 26 M1/T1 serotype isolates from patients with pharyngitis in 1989 to 1997 were identical to those of TSLS-associated isolates. As for serotype M3/T3, all isolates from both patients and healthy subjects except one were classified as PFGE type SmaI:III-SfiI:C or SmaI:III-SfiI:D and carried both speA and speB genes.
Musser et al. (7) analyzed strains expressing M1 protein from 13 countries on five continents (Japan not included) by comparing PFGE patterns of SfiI-digested chromosomal DNA. They concluded that most invasive disease episodes were caused by a distinct subclone that spread among the countries. The results of the present study confirm their findings. Although we have not examined isolates from their study, the pattern B1 that was shown in SfiI-digested chromosomal DNA of M1/T1 serotype strains obtained from patients with TSLS in this study appeared to be identical to the published PFGE pattern of the subclone revealed by Musser et al. (7). However, the patterns observed in serotype M1/T1 isolates from patients with pharyngitis in 1989 to 1997 were identical to those of TSLS-associated isolates. Interestingly, the SmaI-digested pattern (IIa) was comparable to the published PFGE patterns of the isolates from patients with pharyngotonsillitis and TSLS in Europe and the United States (3, 6, 10, 15). In the present study, the association of both genotypic and phenotypic characteristics of the M3 isolates with their pathogenicities seems to be obscure, since the profiles observed in healthy subjects were identical to those of the TSLS-associated isolates. Two possibilities can be raised in this context. One possibility is that the occurrence of TSLS is related to the susceptibility of individuals to a particular strain. Stevens (17) considered that the preexisting immunity against virulence factors of S. pyogenes, such as M proteins and streptococcal pyrogenic exotoxins, participates in the clinical syndrome and outcome of infection. Kiska et al. (6) suggested that cases of pharyngeal infections might have served as a reservoir for virulent strains in persons susceptible to invasive infection. Alternatively, novel virulence factors not found in the strains from the healthy subjects could be possessed by those TSLS-associated strains. Recently, Stockbauer et al. (18) revealed that considerable variations in a particular gene encoding a streptococcal inhibitor of complement, which would aid the bacterium to avoid the host's immune response in infection, existed among serotype M1 isolates that apparently belonged to the same clone. Further phenotypic and molecular characterization of isolates with apparently the “same” genotypic and phenotypic profiles would be necessary in order to elucidate the etiology of TSLS.
Acknowledgments
We are grateful to R. A. Whiley of the Department of Oral Microbiology, St. Bartholomew's and Royal London School of Medicine and Dentistry.
This work was carried out under the Research on Emerging and Reemerging Infectious Diseases, Health Sciences Research Grants, provided by the Ministry of Health and Welfare (Japan).
REFERENCES
- 1.Bouvet A, Geslin P, Kriz-Kuzemenska P, Blanc V, Devine C, Grimont F. Restricted association between biotypes and serotypes within group A streptococci. J Clin Microbiol. 1994;32:1312–1317. doi: 10.1128/jcm.32.5.1312-1317.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cleary P P, Kaplan E L, Handley J P, Wlazlo A, Kim M H, Hauser A R, Schlievert P M. Clonal basis for resurgence of serious Streptococcus pyogenes disease in the 1980s. Lancet. 1992;339:518–521. doi: 10.1016/0140-6736(92)90339-5. [DOI] [PubMed] [Google Scholar]
- 3.Cockerill F R, III, Thomson R L, Musser J M, Schlievert P M, Talbot J, Holley K E, Harmsen W S, Ilstrup D M, Kohner P C, Kim M H, Frankfort B, Manahan J M, Steckelberg J M, Roberson F, Wilson W R the Southeastern Minnesota Streptococcal Working Group. Molecular, serological, and clinical features of 16 consecutive cases of invasive streptococcal disease. Clin Infect Dis. 1998;26:1448–1458. doi: 10.1086/516376. [DOI] [PubMed] [Google Scholar]
- 4.Hauser A R, Stevens D L, Kaplan E L, Schlievert P M. Molecular analysis of pyrogenic exotoxins from Streptococcus pyogenes isolates associated with toxic shock-like syndrome. J Clin Microbiol. 1991;29:1562–1567. doi: 10.1128/jcm.29.8.1562-1567.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Inagaki Y, Konda T, Murayama S, Yamai S, Matsushima A, Gyobu Y, Tanaka D, Tamaru A, Katsukawa C, Katayama A, Tomita M, Fuchi Y, Hoashi K, Watanabe H the Working Group for Group A Streptococci in Japan. Serotyping of Streptococcus pyogenes isolated from common and severe invasive infections in Japan, 1990–5: implication of the T3 serotype strain-expansion in TSLS. Epidemiol Infect. 1997;119:41–48. doi: 10.1017/s0950268897007644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kiska D L, Thiede B, Caracciolo J, Jordan M, Johnson D, Kaplan E L, Gruninger R P, Lohr J A, Gilligan P H, Denny F W., Jr Invasive group A streptococcal infections in North Carolina: epidemiology, clinical features, and genetic and serotype analysis of causative organisms. J Infect Dis. 1997;176:992–1000. doi: 10.1086/516540. [DOI] [PubMed] [Google Scholar]
- 7.Musser J M, Kapur V, Szeto J, Pan X, Swanson D S, Martin D R. Genetic diversity and relationships among Streptococcus pyogenes strains expressing serotype M1 protein: recent intercontinental spread of a subclone causing episodes of invasive disease. Infect Immun. 1995;63:994–1003. doi: 10.1128/iai.63.3.994-1003.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Nakashima K, Ichiyama S, Iimura Y, Hasegawa Y, Ohta M, Ooe K, Shimizu Y, Igarashi H, Murai T, Shimokata K. A clinical and bacteriologic investigation of invasive streptococcal infections in Japan on the basis of serotypes, toxin production, and genomic DNA fingerprints. Clin Infect Dis. 1997;25:260–266. doi: 10.1086/514543. [DOI] [PubMed] [Google Scholar]
- 9.Nei M, Li W H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA. 1979;76:5269–5273. doi: 10.1073/pnas.76.10.5269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nguyen L, Levy D, Ferroni A, Gehanno P, Berche P. Molecular epidemiology of Streptococcus pyogenes in an area where acute pharyngotonsillitis is endemic. J Clin Microbiol. 1997;35:2111–2114. doi: 10.1128/jcm.35.8.2111-2114.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Norgren M, Norrby A, Holm S E. Genetic diversity in T1M1 group A streptococci in relation to clinical outcome of infection. J Infect Dis. 1992;166:1014–1020. doi: 10.1093/infdis/166.5.1014. [DOI] [PubMed] [Google Scholar]
- 12.Rotta J, Facklam R R. Manual of microbiological diagnostic methods for streptococcal infections and their sequelae. Bull W H O. 1980;1:1–50. [Google Scholar]
- 13.Ruoff . Streptococcal diseases. In: Collier L, Balows A, Sussman M, editors. Topley and Wilson's microbiology and microbial infections. 9th ed. Vol. 3. London, United Kingdom: Arnold; 1998. pp. 257–275. [Google Scholar]
- 14.Schwartz B, Facklam R R, Breiman R F. Changing epidemiology of group A streptococcal infection in the USA. Lancet. 1990;336:1167–1171. doi: 10.1016/0140-6736(90)92777-f. [DOI] [PubMed] [Google Scholar]
- 15.Stanley J, Linton D, Desai M, Efstratiou A, George R. Molecular subtyping of prevalent M serotypes of Streptococcus pyogenes causing invasive disease. J Clin Microbiol. 1995;33:2850–2855. doi: 10.1128/jcm.33.11.2850-2855.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Stevens D L. Streptococcal toxic-shock syndrome: spectrum of disease, pathogenesis, and new concepts in treatment. Emerg Infect Dis. 1995;1:69–78. doi: 10.3201/eid0103.950301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Stevens D L. Invasive group streptococcus infections. Clin Infect Dis. 1992;14:2–13. doi: 10.1093/clinids/14.1.2. [DOI] [PubMed] [Google Scholar]
- 18.Stockbauer K E, Grigsby D, Pan X, Fu Y-X, Perea Mejia L M, Cravioto A, Musser J M. Hypervariability generated by natural selection in an extracellular complement-inhibiting protein of serotype M1 strains of group A Streptococcus. Proc Natl Acad Sci USA. 1998;95:3128–3133. doi: 10.1073/pnas.95.6.3128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Tanaka D, Gyobu Y, Kodama H. Typing of group A streptococci by pulsed field gel electrophoresis. Kansenshogaku Zasshi. 1996;70:283–286. doi: 10.11150/kansenshogakuzasshi1970.70.283. [DOI] [PubMed] [Google Scholar]
- 20.Tenover F C, Arbeit R D, Goering R V, Mickelsen P A, Murray B E, Persing D H, Swaminathan B. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33:2233–2239. doi: 10.1128/jcm.33.9.2233-2239.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]