Whipps first described the microbiome as “the theater of activity of micro-organisms,”1 which referred to the numerous micro-organisms that thrive in many human systems. The word “theater,” used by Whipps, indicates a specific medium and environment in which micro-organisms interplay with their surroundings and contribute to the development of various diseases in different organs.1 Nowadays, it is well known that the microbiome acts as an incremental component for the intact functioning of the human body and our well-being. The term infectome also relates to micro-organisms that contribute to the development and progression of diseases, particularly those of autoimmune nature.2 The gut microbiome, for instance, has been of tremendous interest to researchers for years, because the gastrointestinal tract constitutes a vibrant medium of micro-organisms known to play a role in the pathogenesis of numerous diseases of the gastrointestinal system. Nonetheless, many extraintestinal disorders are also closely influenced by gut microbiome alterations.2
Regarding the immune system, its interaction with the normal microbial flora of human organisms, that is, the microbiome, helped shed light on the immune system and unclear mechanisms responsible for the pathogenesis of autoimmune diseases. The microbiome contributes to the homeostasis of the immune system. In contrast, alterations of the microbiome, a state that is known mainly as dysbiosis, have been implicated in the pathogenesis of autoimmune diseases, such as systemic lupus erythematosus and rheumatoid arthritis.3
Early in the coronavirus disease 2019 (COVID-19) pandemic, questions were raised concerning whether the lung microbiome has an active role in the pathogenesis and manifestations of COVID-19, mainly due to the lung involvement in disease’s progression, as well as the extrapulmonary immune-mediated manifestations of the disease and its heterogeneous clinical severity.4 Although the COVID-19 pandemic is still emerging, affecting millions, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) represents an ideal paradigm of the close interplay between the immune system and exacerbated autoimmune manifestations.4 At least 15 distinct autoantibodies and more than 10 different autoimmune diseases appear to develop as a result of the SARS-CoV-2 infection, some of which were shown to be significantly associated.4 The propensity of de novo autoimmunity in patients with COVID-19 primarily seen, but not limited to, in severely ill patients supports the notion that the virus is instrumental for inducing autoimmunity, irrespective of disease status.4 Here, we aim to address the implication of microbiome alterations in COVID-19 with its various aspects in pathogenesis, clinical manifestations, and possibly treatment.
Alterations in the normal microbial populations, including the proliferation of a new organism, may give rise to disease development.2 Some of the pathogenic microbial populations and transformations of microbiome alterations disrupt the immune system’s ability to distinguish between self and nonself, leading to autoimmune manifestations. Thus, many hypothesized that some preexisting lung and gut microbiome compositions could develop autoimmune presentations in SARS-CoV-2–infected patients through direct and indirect mechanisms.5 In addition, the wide diversity of lung microbiota across the public may explain some of the heterogeneity noted in COVID-19–induced autoimmunity.
We had introduced the terms infectome and autoinfectome as the exposure to infectious agents throughout an individual’s lifetime and their contribution to autoimmunity.6 It was proposed that identifying such agents would be necessary for establishing an instrumental link between infection and autoimmunity. Much like genetic imprints, a unique genome of an individual, each individual has a distinctive microbiota.6 According to the concept of autoinfectome, the microbiome variability could contribute to the polymorphism of autoimmunity across the general public.2 Thus, the development of autoimmunity in only some patients with COVID-19 could be a consequence of specific microbial compositions and their complex alterations, which provides a state prone to autoimmunity. Because of genetic predisposition, some individuals are prone to develop autoimmunity, whereas others will not. Similarly, individuals prone to microbiome changes, which present specific alterations of microbiome pattern compositions, will also be prone to develop autoimmune disorders, whereas others, with no such alterations, will not.
The lung microbiome was shown to be correlated with the clinical severity of respiratory syncytial virus infection in infants.7 According to the same study, the severe disease was associated with the presence of H influenza in the airways of patients infected with respiratory syncytial virus. In another study, alterations in the lung microbiome were found to play an essential role in the clinical course and pathogenesis of chronic obstructive pulmonary disease and its exacerbations.8 In relevance to COVID-19, Merenstein et al9 investigated lung microbiome in patients with COVID-19 and its correlation with the severity of the disease, immunologic characteristics, and outcomes by collecting samples from the respiratory tract of 83 patients with COVID-19 and healthy individuals without COVID-19. Moreover, using 16S rRNA sequencing, the authors found a substantial change in lung microbiome in patients with COVID-19 compared with patients without COVID-19. Thus, patients with severe COVID-19 showed massive alterations in the lung microbiome, including the prominence of Staphylococcus and a higher colonization rate of Anelloviridae and Redondoviridae viruses. The authors conclude that the disrupted lung microbiome was correlated to immune response and clinical severity of COVID-19. Nevertheless, it should be noted that the lung microbiome had not been checked before SARS-CoV-2 infection in the affected individuals; thus, additional research is necessary to draw an unambiguous conclusion.
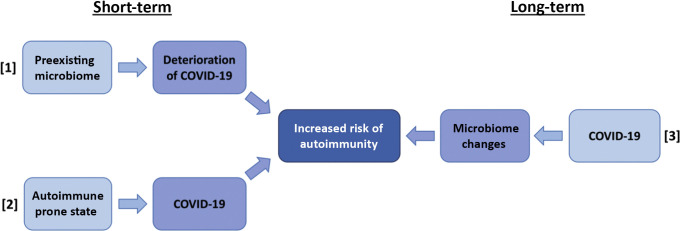
The interaction of the novel SARS-CoV-2 and specific microbiome populations may de novo induce or exacerbate subclinical autoimmunity5 by various mechanisms (Fig 1 ): (1) Specific profiles of preexisting lung microbiota that enable excessive viral proliferation may deteriorate severity of illness and raise the chance of autoimmunity. (2) COVID-19 might act as a final trigger leading to new onset of autoimmunity in genetically susceptible individuals with an autoimmune-prone preexisting microbial composition. (3) Long-term implications of COVID-19 on the immune response could initiate long-lasting transformational changes in microbial populations into microbiota prone to be engaged in foreign self-immunologic alterations, which induce autoimmunity.
Fig 1.
Mechanisms in which interaction of the novel SARS-CoV-2 and specific microbiome populations may increase autoimmunity: (1) Specific profiles of preexisting lung microbiota that enable excessive viral proliferation may increase the severity of illness and the rising chance of viral-induced autoimmunity. (2) COVID-19 might act as a final trigger leading to new onset of autoimmunity in individuals with an autoimmune-prone preexisting microbial composition, that is, an autoimmunity-prone microbiome “signature.” (3) Long-term implications of COVID-19 on the immune responses could initiate long-lasting transformational changes in microbial populations into microbiota, altering microbiota-self interactions and enhancing autoimmunity.
Interestingly, apart from the lung microbiome, the severity of COVID-19 and its outcomes were associated with the gut microbiome. By analyzing the plasma for gut microbiome and gut permeability markers (FABP-2, PGNN, and LPS) in 30 patients with COVID-19 and 16 healthy controls, Prasad et al10 demonstrated, by using 16S rRNA sequencing, higher levels of gut permeability markers as well as higher rates of the gut microbiome in the bloodstream of patients with COVID-19. The authors concluded their results as the reason behind bacteremia and worse outcomes in patients with COVID-19, which is yet to be seen in more extensive studies.
In conclusion, the microbiome, particularly the lung and gut microbiome, could have a crucial role in the pathogenesis, clinical severity, outcomes, and treatment of COVID-19; nevertheless, further research is required to improve our understanding. Studies that deal with the definition and isolation of microbiome should be conducted to provide reliable data that can assist in targeted treatment to patients with COVID-19 and other conditions.
Footnotes
Disclosure of potential conflict of interest: The authors declare that they have no relevant conflicts of interest.
References
- 1.Berg G., Rybakova D., Fischer D., Cernava T., Vergès M.C., Charles T., et al. Microbiome definition re-visited: old concepts and new challenges. Microbiome. 2020;8:103. doi: 10.1186/s40168-020-00875-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Li B., Selmi C., Tang R., Gershwin M.E., Ma X. The microbiome and autoimmunity: a paradigm from the gut–liver axis. Cell Mol Immunol. 2018;15:595–609. doi: 10.1038/cmi.2018.7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.De Luca F., Shoenfeld Y. The microbiome in autoimmune diseases. Clin Exp Immunol. 2019;195:74–85. doi: 10.1111/cei.13158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dotan A., Muller S., Kanduc D., David P., Halpert G., Shoenfeld Y. The SARS-CoV-2 as an instrumental trigger of autoimmunity. Autoimmun Rev. 2021;20 doi: 10.1016/j.autrev.2021.102792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ferreira C., Viana S.D., Reis F. Is gut microbiota dysbiosis a predictor of increased susceptibility to poor outcome of COVID-19 patients? An update. Microorganisms. 2021;9:53. doi: 10.3390/microorganisms9010053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bogdanos D.P., Smyk D.S., Rigopoulou E.I., Sakkas L.I., Shoenfeld Y. Infectomics and autoinfectomics: a tool to study infectious-induced autoimmunity. Lupus. 2015;24:364–373. doi: 10.1177/0961203314559088. [DOI] [PubMed] [Google Scholar]
- 7.Chu C.Y., Qiu X., McCall M.N., Wang L., Corbett A., Holden-Wiltse J., et al. Airway gene expression correlates of RSV disease severity and microbiome composition in infants. J Infect Dis. 2021;223:1639–1649. doi: 10.1093/infdis/jiaa576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Santoro A., Tomino C., Prinzi G., Cardaci V., Fini M., Macera L., et al. Microbiome in chronic obstructive pulmonary disease: role of natural products against microbial pathogens. Curr Med Chem. 2020;27:2931–2948. doi: 10.2174/0929867327666191213110551. [DOI] [PubMed] [Google Scholar]
- 9.Merenstein C., Liang G., Whiteside S.A., Cobián-Güemes A.G., Merlino M.S., Taylor L.J., et al. Signatures of COVID-19 severity and immune response in the respiratory tract microbiome. medRxiv. 2021 doi: 10.1128/mBio.01777-21. 2021.04.02.21254514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Prasad R., Patton M.J., Floyd J.L., Vieira C.P., Fortmann S.D., DuPont M., et al. Plasma microbiome in COVID-19 subjects: an indicator of gut barrier defects and dysbiosis. bioRxiv. 2021 doi: 10.3390/ijms23169141. 2021.04.06.438634. [DOI] [PMC free article] [PubMed] [Google Scholar]