Abstract
Inhibitors of bromodomain and extra-terminal motif (BET) proteins are emerging epigenetic therapeutics that suppress gene expressions that drive cancer and inflammation. The present study examined anti-inflammatory effects of a quinazoline-based BET inhibitor, CN210, in a murine ileitis model. CN210 was given orally 30 min before and 24 h after a subcutaneous administration of indomethacin. Macroscopic and histological evidences of ileitis, mucosal myeloperoxidase (MPO) activity and cytokine expressions were evaluated 48 h after the indomethacin administration. To further characterize the anti-inflammatory pathways modulated by CN210, its effects on RAW264 cells treated with lipopolysaccharide (LPS) were investigated. Competitive ligand binding and docking studies of CN210 to CREB-binding protein (CBP) and p300 were also performed. Oral administration of CN210 significantly reduced the severity of ileitis, normalized both proinflammatory MPO activity and concomitant cytokine expressions induced by indomethacin administration. Furthermore, CN210 attenuated the expression of cytokines and reversed the activation of nuclear factor κB (NF-κB) and mitogen-activated protein kinases (MAPK) induced by LPS. Competitive ligand binding assays showed that CN210 bound to the bromodomains of two paralogous histone acetyltransferases, CBP and p300, in addition to the bromodomains of BET proteins. Docking studies of CN210 to the bromodomains of CBP and p300 showed a similarity to the binding mode of SGC-CBP30, a specific CBP/p300 inhibitor. CN210 ameliorates indomethacin-induced ileitis by inhibiting the expression of inflammatory cytokines through the attenuation of NF-κB and MAPK pathways. CN210 thus represents a new mode of therapy for non-steroidal anti-inflammatory drug-induced ileitis and inflammatory bowel disease.
Keywords: bromodomain and extra-terminal motif (BET) proteins, CREB-binding protein (CBP) and p300, ileitis
1 |. INTRODUCTION
Bromodomain and extra-terminal motif (BET) proteins—BRD2, BRD3, BRD4 and BRDT—are epigenetic “reader” proteins that bind to acetylated lysine residues on proteins (e.g., histones) by means of their bromodomains (Filippakopoulos et al., 2012; Wu & Chiang, 2007). Of the four BET proteins, BRD4 associates with the general transcription cofactor complex, Mediator, and is a key structural component of extensive transcription factor complexes formed at genomic regions known as super-enhancers (Loven et al., 2013; Whyte et al., 2013; Wu & Chiang, 2007). By inhibiting the binding of BRD4 to acetylated histones, small-molecule inhibitors of BET proteins (BET inhibitors) have been shown to suppress the expression of the MYC oncogene and the genes that encode proinflammatory cytokines, such as interleukin (IL)-6, interferon (INF)-β, IL-1β, IL-12α, chemokine (C-X-C motif) ligand 9 and chemokine (C-C motif) ligand 12 in mice (Delmore et al., 2011; Mertz et al., 2011; Nicodeme et al., 2010). Accordingly, BET inhibitors are now considered as promising drug candidates for both cancer and inflammatory diseases (Cochran et al., 2019; Wang & Filippakopoulos, 2015).
While several BET inhibitors are currently evaluated clinically (Smith & Zhou, 2016), some adverse effects, including thrombocytopenia, have been reported in early clinical studies (Doroshow et al., 2017). Therefore, further development of alternative BET inhibitors with distinct structures and better safety and efficacy profiles remains of high interest. We recently developed a novel quinazoline-based BET inhibitor, CN210 (Figure 1b) with a therapeutically suitable target affinity (Kd = 70 nM for BRD4 containing tandem bromodomains), cellular potency (IC50 of 0.94 μM for the viability assay in a leukemia cell line, MV4–11) and a pharmacokinetic profile (t1/2 of 1.7 h; Cmax of 2163 ng/ml; AUC0–∞ of 8927 ng h/ml; and oral bioavailability of 75% following a 10 mg/kg oral administration in mice) (Yang et al., 2019). CN210 is highly effective in inhibiting tumor growth in a Kasumi-1 ×enograft mouse model and in improving arthritis in a collagen-induced arthritis mouse model (Yang et al., 2019).
FIG U R E 1.
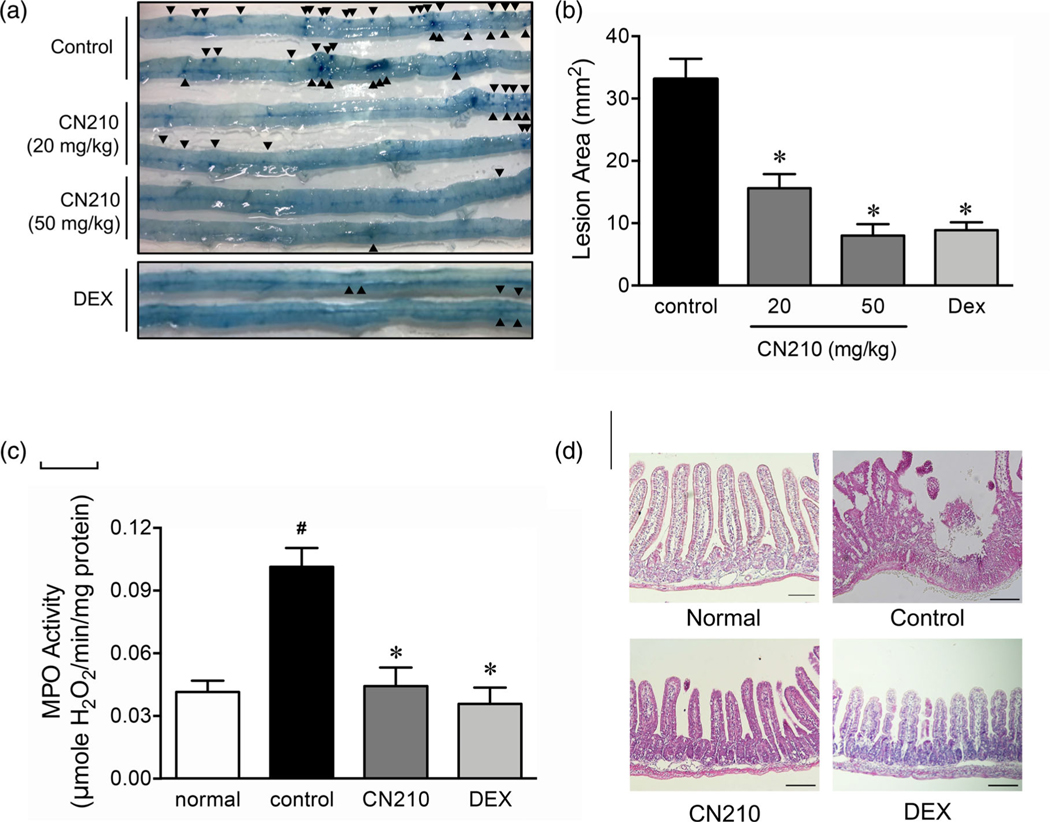
Effect of CN210 on indomethacin-induced ileitis. Animals received indomethacin (10 mg/kg) subcutaneously and were sacrificed 48 h later. CN210 (20 and 50 mg/kg) and dexamethasone (3 mg/kg) were administered 30 min before and 24 h after indomethacin treatment. (a) Representative macroscopic observation of indomethacin-induced ileitis. Arrowheads indicate Evans blue-stained ileal lesions. (b) Lesion area and (c) MPO activity of indomethacin-induced ileitis. (d) Representative histological observation of indomethacin-induced ileitis at ×100 magnification. Scale bar: 50 μm. Data are shown as means ± standard errors of the mean (ns = 5 to 8). One-way ANOVA with Holm-Sidak’s test revealed a statistically significant difference between the experimental mice and the controls (vehicle alone; *P < 0.05)
Nonsteroidal anti-inflammatory drugs (NSAIDs), including indomethacin, produce tissue damages in the stomach and small intestine in human and animals (Bjarnason et al., 1993; Konaka et al., 1999; Yamada et al., 1993). Because of the pathomorphological similarity to human Crohn’s disease in the small intestine, indomethacin-induced ileitis in rodents is proposed as a model for studying human Crohn’s disease in addition to NSAID-induced enteropathy (Anthony et al., 2000).
Inflammatory bowel disease (IBD), including ulcerative colitis and Crohn’s disease, is characterized by chronic inflammation of gastrointestinal tract as a result of abnormal immune responses including aberrantly increased amounts of inflammatory cytokines. Recent introduction of biologic agents, such as anti–tumor necrosis factor α (TNF-α) and anti–IL-12/IL-23p40 monoclonal antibodies, has significantly improved the outcome of IBD (Feagan et al., 2016; Hanauer et al., 2002; Rutgeerts et al., 2005); however, complete remission still is a challenge because patients with IBD often develop anti-drug antibodies, which reduce the long-term effectiveness of these agents. Thus, development of small-molecule drugs that complement current armamentarium to treat IBD is of high medical interest.
Recent studies have shown that BET inhibitors ameliorate colitis by inhibiting the differentiation of Th17 cells, the maturation of dendritic cells and the suppression of inflammatory cytokine expressions, at least in part, by blocking the activation of nuclear factor κB (NF-κB) and mitogen-activated protein kinase (MAPK) (Cheung et al., 2017; Huang, Zeng, et al., 2017; Schilderink et al., 2016).
In the current study, we demonstrated that CN210 suppressed indomethacin-induced ileitis in mice with concomitant suppression of gene expression of proinflammatory cytokines. Furthermore, CN210 inhibited not only the bromodomains of BET proteins but also the bromodomains of CREB-binding protein (CBP) and p300, two paralogous histone acetyltransferases.
2 |. METHODS
2.1 |. Materials
Indomethacin and dexamethasone were obtained from Sigma-Aldrich (St. Louis, MO, United States). CN210 was synthesized according to the route illustrated in a previous report (Yang et al., 2019). The structure of CN210 is shown in Figure 1b. Indomethacin was dissolved in physiological saline with a drop of Tween 80 (Wako, Osaka, Japan) and administered subcutaneously in a volume of 0.1 ml/10 g body weight. CN210 and dexamethasone were suspended in carboxymethylcellulose (CMC) (Nacalai Tesque, Kyoto, Japan) and administered orally in a volume of 0.1 ml/10 g body weight.
2.2 |. Animals
Male C57BL/6 mice weighing 22–26 g (SLC Co., Shizuoka, Japan) were acclimated to standard laboratory conditions with 12-h light–dark cycles and a temperature of 22 ± 1°C. Experiments were performed with five to eight unanesthetized mice per group. This study was performed in accordance with the Animal Research: Reporting In Vivo Experiments (ARRIVE) guidelines (Kilkenny et al., 2010). The protocols were approved by the Committee on the Ethics of Animal Research of Kyoto Pharmaceutical University (Permit Number: PETH-19–004).
2.3 |. Induction of ileitis
After fasting for 18 h and refeeding for 1 h, the animals were administered 10 mg/kg of indomethacin subcutaneously. CN210 (20 and 50 mg/kg) and dexamethasone (3 mg/kg) were administered orally 30 min before and 24 h after indomethacin injection. The animals were sacrificed 48 h after indomethacin administration. The small intestine was removed and fixed with 2% formalin. The fixed small intestine was opened along the antimesenteric attachment and examined for lesions under a dissecting microscope (S6D; Leica Microsystems, Wetzlar, Germany) at a magnification of ×20. The area of macroscopically visible lesions (mm2) was measured and summed per ileum. To visualize lesions, 0.5% of Evans blue solution was injected intravenously in a volume of 0.2 ml in each animal 30 min before sacrifice. The measurement of area was performed by two investigators unaware of experimental groups. Subsequently, the ileum was immersed in 10% formalin overnight, and tissue samples were excised, embedded in paraffin, sectioned into 4-μm slices and stained with hematoxylin and eosin. Histological ileitis was observed under a light microscope (BX51; Olympus, Tokyo, Japan) fitted with a digital camera system (DS-Ri1; Nikon, Tokyo, Japan).
2.4 |. MPO activity
Ileal tissues were rinsed with cold phosphate-buffered saline, weighed and homogenized in 50 mM of phosphate buffer, pH 6.0, containing 0.5% hexadecyltrimethylammonium bromide. A spectrophotometric assay kit (Pierce, Rockford, IL, United States) was used to estimate total protein in lysates, whereas MPO activity was determined with the use of o-dianisidine hydrochloride (Sigma-Aldrich), as previously described (Yasuda et al., 2011).
2.5 |. Cytokine expression
Ileal tissues were washed with cold phosphate-buffered saline and stored in RNAlater Stabilization Solution (Thermo Fisher Scientific) at 4°C until use. Total RNA was extracted with the use of Sepasol-RNA I Super G (Nacalai Tesque) according to the manufacturer’s instructions and reverse-transcribed with the use of PrimeScript Reverse Transcriptase (Takara Bio, Shiga, Japan). Thermal Cycler Dice Real-Time System (Takara Bio) with SYBR Premix ExTaq II (Takara Bio) was used to quantify the expression of cytokines. Predesigned primer sets for mouse TNF-α (primer set MA097070), IL-6 (primer set MA039013), interferon-γ (IFN-γ) (primer set MA025911), IL-17 (primer set MA157056) and TATA-binding protein (TBP) (primer set MA050367) were obtained from the Perfect Real-Time Supporting System (Takara Bio). The mRNA expression level was normalized to the expression of TBP.
2.6 |. Cell culture
Mouse monocyte/macrophage cell line RAW264 was obtained from RIKEN BRC (Ibaraki, Japan). The cells were grown in Dulbecco’s modified Eagle medium (DMEM: Wako) containing 10% fetal bovine serum with 100 U/ml of penicillin G and 100 μg/ml of streptomycin at 37°C in 5% CO2 with a humidified atmosphere. Subconfluent cells in 12-well plates were exposed to LPS (Sigma-Aldrich) at a concentration of 1 μg/ml, and total RNA was extracted 4 h later. Subsequently, reverse transcription and PCR were performed as described in the previous section. Predesigned primer sets for mouse TNF-α (primer set MA097070), IL-6 (primer set MA039013), IL-12A (primer set MA0287533), IL-23A (primer set MA095159), and TBP (primer set MA050367) were obtained from the Perfect Real-Time Supporting System (Takara Bio). The messenger RNA expression level was normalized to the expression of TBP. CN210 (0.1, 1 and 10 μM) and dexamethasone (1 μM) were cotreated with LPS in RAW264 cells.
2.7 |. Western blotting
Nuclear proteins were isolated from RAW264 cells with the Nuclear Extract Kit (Active Motif, Carlsbad, CA, United States) according to the manufacturer’s protocols. To extract total proteins from RAW264 cells, we used radioimmunoprecipitation assay (RIPA) buffer containing protease inhibitors (Complete Mini; Roche Diagnostics GmbH, Mannheim, Germany) and phosphatase inhibitors (PhosSTOP EASYPack; Roche Diagnostics GmbH) as described previously (Tsukahara et al., 2017). Nuclear and total protein samples were separated on sodium dodecyl sulfate–polyacrylamide gel (SDS-PAGE) electrophoresis and subsequently transferred to polyvinylidene difluoride (PVDF) membranes (Bio-Rad, Hercules, CA, United States). After blocking with 5% of bovine serum albumin or skim milk for 1 h at room temperature, the specific primary antibodies for NF-κB, p65, p38, phospho-p38, extracellular signal-regulated kinase (ERK), phospho-ERK, c-JUN-terminal kinase (JNK), phosphor-JNK (Cell Signaling, Danvers, MA, United States), lamin B1 (ProteinTech, Chicago, IL, United States) or β-actin (Gene Tex, Irvine, CA, United States) were used to probe the PVDF membranes at 4°C overnight. Subsequently, the membranes were incubated with horseradish peroxidase–conjugated secondary antibodies (Cell Signaling) for 1 h at room temperature; a chemiluminescence detection kit (Perkin Elmer, Waltham, MA, United States) was used to detect protein expression signals, and a chemiluminescence imaging system (Fusion Solo S, Viber, Collegien, France) was used to visualization and quantification. The electrophoresis and western blotting were performed separately for each protein (i.e., nine gels and membranes), while total protein concentration was used to ensure equal loading (20 μg/lane). The levels of protein expression were normalized to the expression of lamin B1 for p65 or β-actin for the other proteins.
2.8 |. Competitive ligand binding assay
Competitive ligand binding assays for CN210 against CBP and p300 proteins were performed with BROMOscan® service at Eurofins DiscoverX Corporation (San Diego, CA, United States) described previously (Yang et al., 2018). Kd values were obtained by using a threefold serial dilution across 11 compound concentrations ranging from 0 to 10 μM and by calculating a standard dose–response curve according to the Hill equation, with a slope of −1. Curves were fitted according to a nonlinear least-squares fit with the Levenberg–Marquardt algorithm.
2.9 |. Docking study
The co-crystallized structure of N-terminal bromodomain of human BRD4 (BRD4 BD1) in complex with CN210 was previously reported (PDB code: 6MAU) (Yang et al., 2019). Docking studies of CN210 to the bromodomains of CBP and p300 were performed with the Molecular Operating Environment (MOE) program (Chemical Computing Group, Montreal, QC, Canada) using the co-crystallized structures of SGC-CBP30 (Figure 5b) with CBP (PDB code: 4NR7) and p300 (PDB code: 5BT3) (Hammitzsch et al., 2015). Prior to molecular modeling and docking, the structures were prepared and energy-minimized using the Protein Preparation module in the MOE program. The ligand-induced fit protocol was used and the binding affinity was evaluated using the GBVI/WSA score.
2.10 |. Statistical analysis
All the data were calculated as means ± SEs of the mean. Statistical analyses were performed with one-way analysis of variance (ANOVA), followed by a Holm–Sidak multiple comparison test. A p level of less than .05 was considered statistically significant. The GraphPad Prism 6.0h program (GraphPad Software, La Jolla, CA, United States) was used for statistical analyses.
3 |. RESULTS
3.1 |. Effect of CN210 on indomethacin-induced ileitis and increase in MPO activity
Single subcutaneous administration of indomethacin in mice produced ileitis characterized by severe lesions in the small intestine, mostly between the distal jejunum and the ileum (Figure 1a). CN210 (20 and 50 mg/kg), given orally 30 min before and 24 h after indomethacin administration, reduced the severity of ileal lesions in a dose-dependent manner while the effects were statistically significant at both dose levels (Figure 1a,b). Likewise, dexamethasone (3 mg/kg), a reference corticosteroid, given orally 30 min before and 24 h after indomethacin administration, significantly attenuated the ileitis. Single subcutaneous administration of indomethacin markedly increased MPO activity in the ileal mucosa (Figure 1c). This increase of MPO activity, a hallmark of neutrophil activation, was suppressed significantly by CN210 (50 mg/kg) and dexamethasone (3 mg/kg). Histological observation indicated that severe mucosal lesions as deep as the muscle layer were found 48 h after indomethacin administration (Figure 1c). In contrast, such formation of mucosal lesions was attenuated by the treatment with either CN210 (50 mg/kg) or dexamethasone (3 mg/kg); as a result, the mucosa appeared mostly normal.
3.2 |. Effect of CN210 on indomethacin-induced increase in cytokine expressions
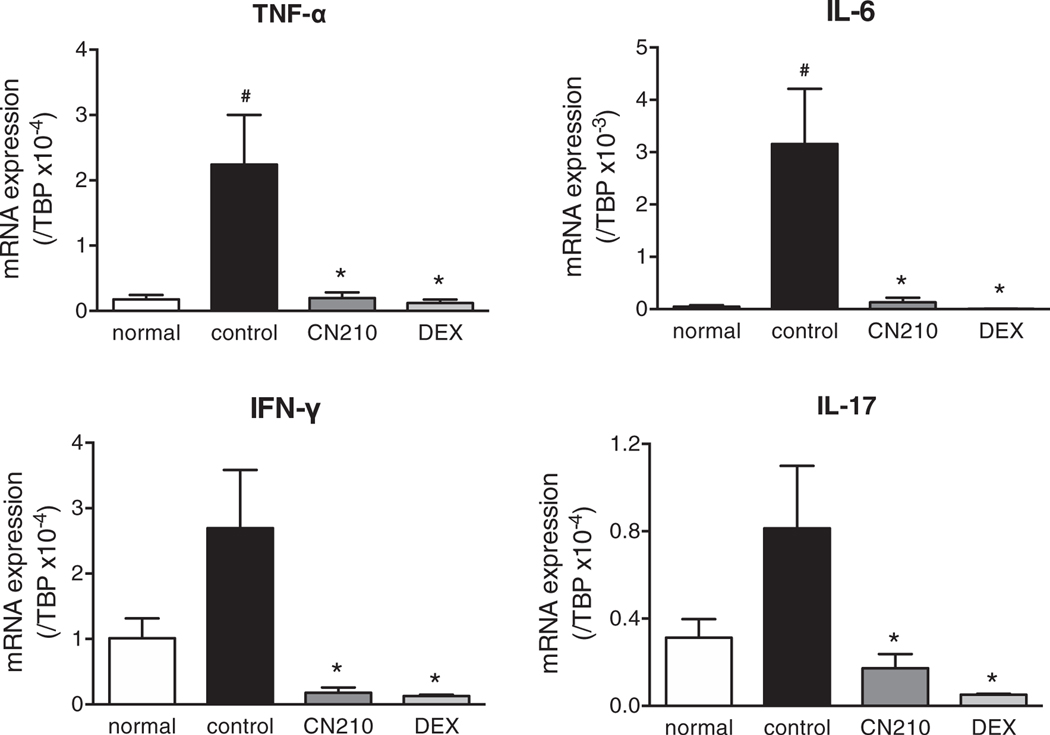
To confirm the suppressive effect of CN210 on indomethacin-induced ileitis, we examined the effects of CN210 on cytokine expressions in the ileal mucosa. Concomitantly with the increase in MPO activity, gene expression of cytokines such as TNF-α, IL-6, IFN-γ and IL-17 in the ileal mucosa was markedly enhanced 48 h after indomethacin administration (Figure 2). These responses were abolished by the treatment with CN210 (50 mg/kg) or dexamethasone (3 mg/kg).
FIG U R E 2.
Effect of CN210 on indomethacin-induced increase in cytokine expression in the ileal mucosa. Animals received indomethacin (10 mg/kg) subcutaneously, and messenger RNA expression for TNF-α, IL-6, IFN-γ, and IL-17 was determined by quantitative real-time RT-PCR 48 h later. CN210 (50 mg/kg) and dexamethasone (3 mg/kg) were administered 30 min before and 24 h after indomethacin treatment. Data are shown as means ± standard errors of the mean (n = 8). One-way ANOVA with Holm-Sidak’s test revealed a statistically significant difference between experimental mice and controls (vehicle alone; *P < 0.05) and between experimental mice and those not given indomethacin (#P < 0.05)
3.3 |. Effect of CN210 on LPS-stimulated cytokine expressions in RAW264 cells
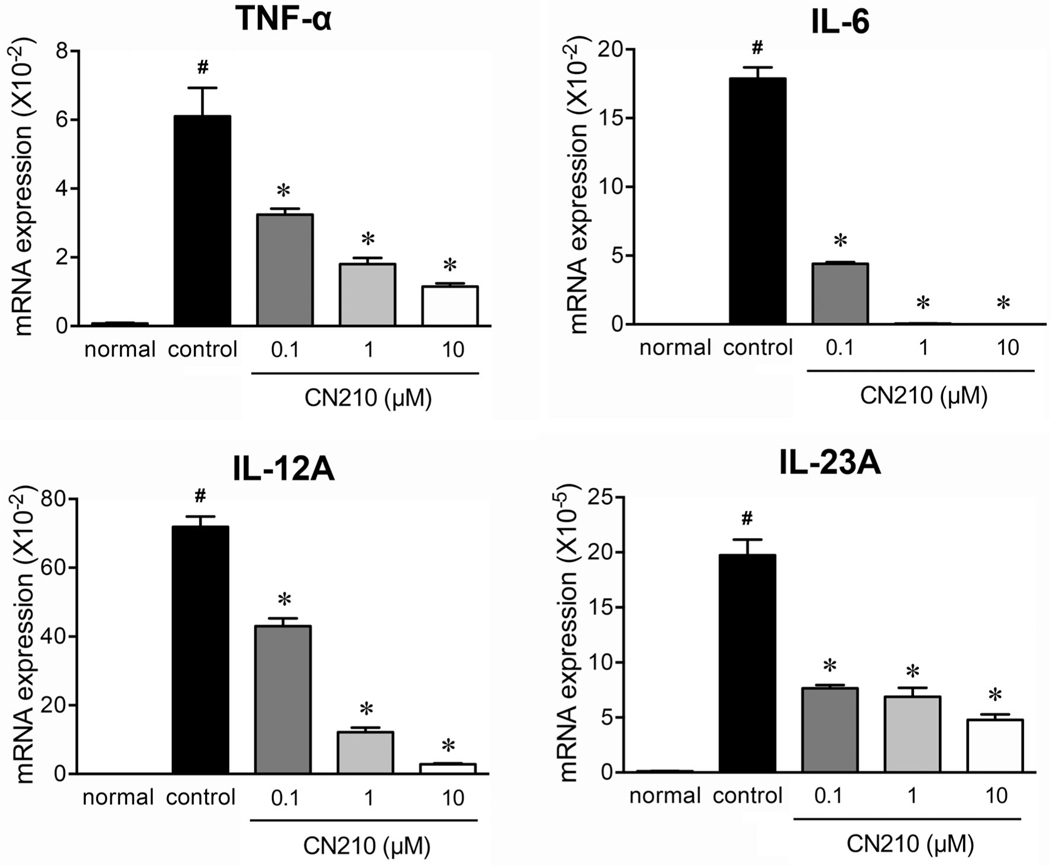
To further characterize the effects of CN210 on inflammatory cellular responses, we next examined the effects of CN210 in a monocyte/macrophage cell line, RAW264. The exposure of RAW264 cells to LPS (1 μg/ml) markedly upregulated the gene expression of cytokines such as TNF-α, IL-6, IL-12α and IL-23α (Figure 3). These responses were suppressed by cotreatment with CN210 (0.1–10 μM), in a concentration-dependent manner, while statistically significant effects already taking place at 0.1 μM.
FIG U R E 3.
Effect of CN210 on LPS-stimulated cytokine expression in RAW264 cells. The cells were treated with LPS (1 μg/mL), and total RNA was isolated 4 h later and messenger RNA expression for TNF-α, IL-6, IFN-γ, and IL-17 was determined by quantitative real-time RT-PCR. Data are shown as means ± standard errors of the mean (n = 6). One-way ANOVA with Holm-Sidak’s test revealed a statistically significant difference between experimental mice and controls (vehicle alone; *P < 0.05) and between experimental mice and those not given indomethacin (#P < 0.05)
3.4 |. Effect of CN210 on LPS-activated NF-κB, extracellular signal-regulated kinase and c-JUN N-terminal kinase in RAW264 cells
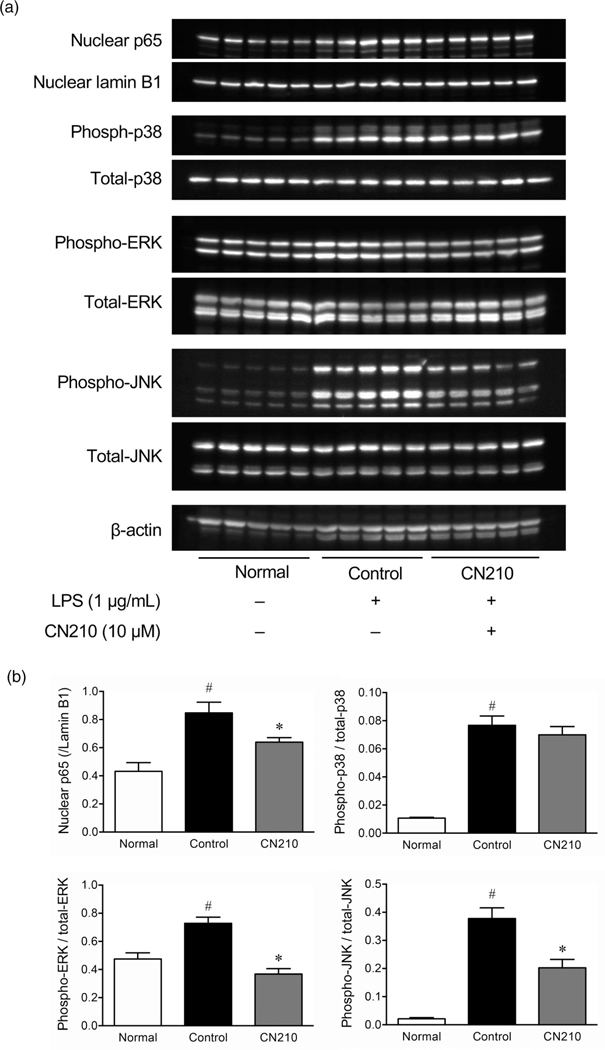
To elucidate the intracellular signaling pathways involved in CN210-induced anti-inflammatory effects, we examined the effects of CN210 on NF-κB, ERK, p38, and JNK activations in RAW264 cells. The exposure of RAW264 cells to LPS (1 μg/ml) induced translocation of the NF-κB p65 subunit to cell nuclei (Figure 4). This translocation was impaired significantly by cotreatment with CN210 (10 μM). Likewise, LPS treatment enhanced phosphorylation of p38, ERK and JNK in RAW264 cells. Although otreatment with CN210 (10 μM) significantly suppressed phosphorylation of ERK and JNK, the suppressive effect was more prominent with ERK phosphorylation than with JNK phosphorylation. Interestingly, CN210 did not suppress LPS-induced phosphorylation of p38 in RAW264 cells.
FIG U R E 4.
Effects of CN210 on LPS-activated NF-κB and MAPK in RAW264 cells. The cells were treated with LPS (1 μg/mL) and activation of NF-κB and MAPK was determined by Western blotting 4 h later. Mice given CN210 (10 μM) were cotreated with LPS. (a) Western blot analyses and (b) quantitative results for the effect of CN210 on LPS-activated NF-κB and MAPK in RAW264 cells (5 replicates at 4 h). Data are shown as mean ± standard errors of the mean (n = 5). One-way ANOVA with Holm-Sidak’s test revealed a statistically significant difference between experimental mice and controls (vehicle alone; *P < 0.05) and between experimental mice and those not given indomethacin (#P < 0.05)
3.5 |. Competitive binding effects of CN210 on the bromodomains of CBP and p300
We previously reported that CN210 potently inhibited the bromodomains of BET proteins, with Kd values ranging from 11 to 200 nM (Villar-Prados et al., 2019; Yang et al., 2019). In this study, we found that CN210 also inhibited the bromodomains of two paralogous histone acetyltransferases, CBP and p300, with Kd values of 260 and 190 nM, respectively (Table 1). In comparison, the previously reported Kd value was 70 nM for BRD4 containing tandem bromodomains (Yang et al., 2019). Of note, CBP and p330 are proximal to BET proteins in a phylogenetic tree generated with three-dimensional structure-based alignments (Filippakopoulos et al., 2012).
TA BL E 1.
The competitive binding affinity (Kd) for BRD4, CBP and p300
| BROMOscan® target | Kd (nM) |
|---|---|
| BRD4 (BD1, 2) | 70a |
| CBP | 260 |
| p300 | 190 |
Note: The Competitive ligand binding assay was conducted with the BROMOScan® service (Eurofins DiscoverX, San Diego, CA, United States). BROMOscan® was conducted against bromodomain in histone acetyltransferases, CBP and p300.
The Kd for BRD4 (BD1, 2), which contains tandem bromodomain BD1 and BD2, was from our previous study (Yang et al., 2019) and is shown for comparison.
3.6 |. Docking studies of CN210 in bromodomain of CBP and p300
To examine the binding of CN210 to the bromodomains of CBP and p300, we conducted a series of docking studies and compared the results to the co-crystal structure of BRD4 BD1 with CN210 and the co-crystal structures of CBP/p300 bromodomains with SGC-CBP30, a specific inhibitor of the bromodomains of CBP/p300 containing a 3,5-dimethylisoxazole group as a mimic of acetyllysine (Hammitzsch et al., 2015). Although BRD4 BD1 shares only ~31% of the sequence identity with the bromodomains of CBP and p300, structural analyses have shown that these bromodomains adopt the same three dimensional fold (Filippakopoulos et al., 2010) (Figure 5a–c).
FIG U R E 5.
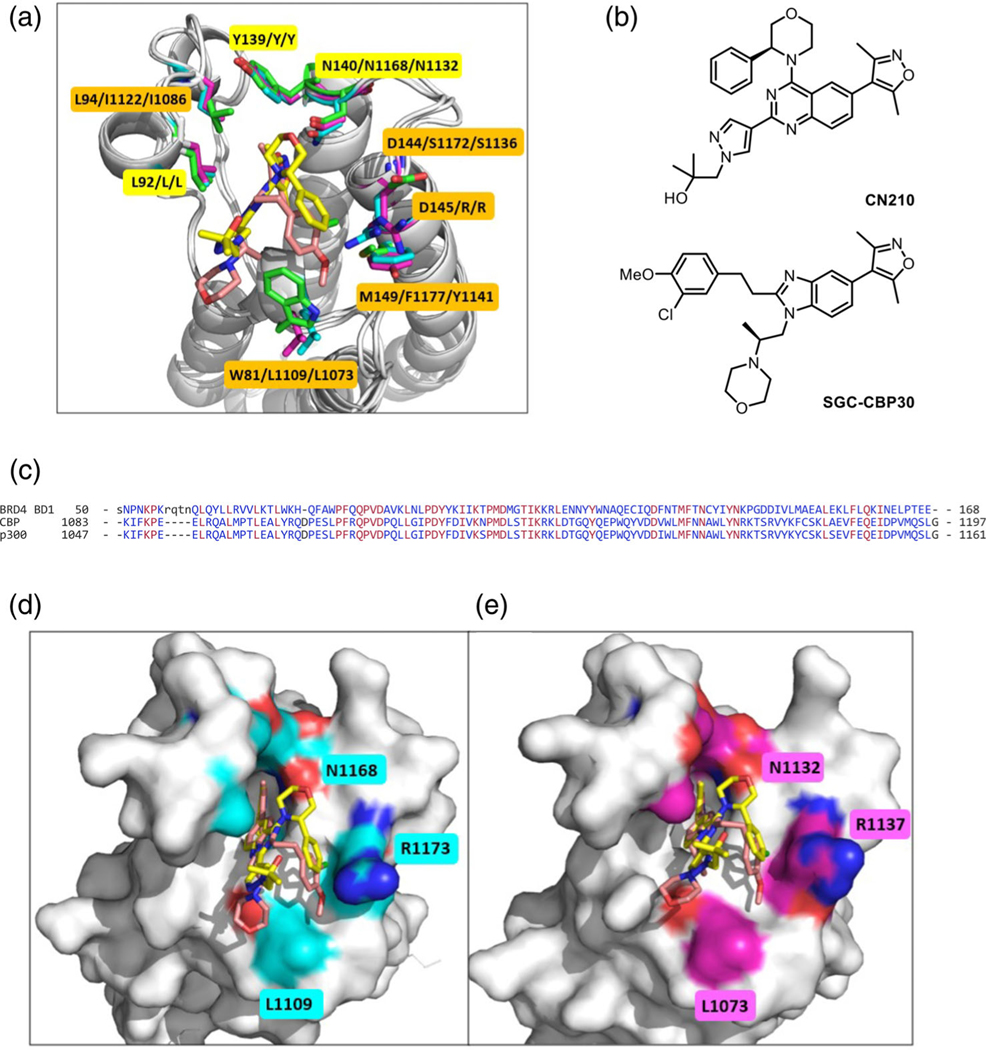
Docking study of CN210 to the bromodomain of CBP and p300. (a) Structural comparison of BRD4 BD1 (6MAU), CBP (4NR7) and p300 (5BT3). Proteins are shown in cartoons in gray. Key residues surrounding the binding pocket are shown in sticks and the carbon atoms are colored in green BRD4 BD1/cyan (CBP)/magenta (p300), respectively. Conserved residues are highlighted in bright yellow and the varied residues are highlighted in yellow-orange in the order of BRD4 BD1/CBP/p300. CN210 in the co-crystal with BRD4 BD1 is shown in yellow, and SGC-CBP30 in the co-crystals with CBP/p300 is shown in salmon. (b) Structures of CN210 and SGC-CBP30. (c) Amino acid sequence alignment of bromodomains of three proteins; N-terminal bromodomain of BRD4 (BRD4 BD1), CBP and p300. (d) Predicted binding model of CN210 (yellow) in the binding pocket of the bromodomain of CBP (4NR7). SGC-CBP30 is shown in salmon. (e) Predicted binding model of CN210 (yellow) in the binding pocket of the bromodomain of p300 (5BT3). SGC-CBP30 is shown in salmon
Since CN210 and SGC-CBP30 share the same 3,5-dimethylisoxazole and a bicyclic core structure (Figure 5b), a similar binding mode between CN210 and SGC-CBP30 to the bromodomains of CBP/p300 was anticipated. Based on the docking studies showed in Figure 5d,e, the predicted binding model of CN210 bound to the bromodomains of CBP and p300 adopted in a similar orientation as observed with SGC-CBP30 in the co-crystallized structures of CBP (4NR7, Figure 5d) and p300 (5BT3, Figure 5e). The conserved Asn residue—that is, N140 (BRD4), N1168 (CBP) and N1132 (p300), the key residue for binding acetyllysine on target proteins—formed potential H-bond interactions with the isoxazole moiety of CN210 and SGC-CBP30 in a same manner. In the co-crystal with BRD4 BD1, the pyrazole ring of CN210 formed an important π-edge interaction with Trp residue (W81) at the WPF shelf. However, such interaction was absent in the docked binding to the bromodomains of CBP and p300 since Trp residue was substituted with a Leu residue— that is, L1109 (CBP) and L1073 (p300)—at this position (Figure 5d,e). Another significant binding difference was seen at the region corresponding to D144/D145/M149 in BRD4 BD1, where varied residues (e.g., Ser, Arg, Phe, or Tyr) were found in CBP and p300 (Figure 1d,e).
4 |. DISCUSSION
In recent years, BET inhibitors have attracted attention as a novel class of drug candidates not only for cancer but also for various inflammatory diseases (Cochran et al., 2019; Wang & Filippakopoulos, 2015). Several studies demonstrated that BET inhibitors were effective in alleviating arthritis, vascular inflammation, acute pancreatitis, hepatic fibrosis and renal damage (Ding et al., 2015; Huang, Haynes, et al., 2017; Huang, Zeng, et al., 2017; Klein et al., 2016; Suarez-Alvarez et al., 2017), as well as in T cell transfer models of colitis (Cheung et al., 2017; Schilderink et al., 2016). In this study, we demonstrated that a novel quinazoline-based bromodomain inhibitor, CN210, potently attenuated indomethacin-induced like ileitis in mice.
BET proteins, such as BRD2 and BRD4, epigenetically regulate transcription of progrowth genes and proinflammatory cytokine genes (Muller et al., 2011). Several studies demonstrated that small-molecule BET inhibitors and small interfering RNAs (siRNAs) targeting BET protein transcripts suppressed cytokine production in inflammatory cells such as macrophages and synovial fibroblasts (Belkina et al., 2013; Klein et al., 2016). Furthermore, it is known that BET proteins regulate gene transcriptions during the differentiation of naïve CD4+ cells to mature helper T cells, and that BET inhibitors, such as JQ1, attenuated the differentiation of Th1, Th2 and Th17 cells (Bandukwala et al., 2012; Mele et al., 2013). BET inhibitors have also reported to suppress inflammatory cytokine production and T-cell activation by inhibiting dendritic cell maturation and promotion of regulatory T cell differentiation (Schilderink et al., 2016).
In this study, we induced ileitis in mice to assess CN210 as a potential drug candidate in the treatment of NSAID-induced enteropathy and IBD including Crohn’s disease. We demonstrated that CN210 interfered with the upregulation of the genes that encode TNF-α, IL-6, IFN-γ, and IL-17, which are known to play key roles in the pathogenesis of indomethacin-induced ileitis in experimental animals (Fukumoto et al., 2011; Kato et al., 2012; Yamada et al., 2011). Importantly, the suppression of IFN-γ and IL-17 genes by CN210 may provide long-term benefits by interfering with T cell differentiation and dendritic cell maturation.
To study the anti-inflammatory effects of CN210 in more detail, we examined its effects on LPS-stimulated cytokine expression in monocyte/macrophage cell line RAW264 in vitro. Previous studies showed that a BET inhibitor, JQ1, and knockdown of BET family proteins with siRNAs blocked cytokine productions in LPS-treated RAW264 cells (Belkina et al., 2013). CN210 potently suppressed the upregulation of the genes encoding TNF-α, IL-6, IL-12α and IL-23α in RAW264 cells. Because IL-12 and IL-23 are important regulators of the differentiation and maintenance of Th1 and Th17 cells, respectively (Szabo et al., 2003; Zhu et al., 2010), CN210 may also provide prolonged therapeutic benefits by interfering with the immune responses by antigen-presented cells that induce differentiation and maintenance of Th1 and Th17 cells.
CN210 also reduced LPS-induced translocation of the NF-κB p65 subunit to cell nuclei. It has been shown that acute stimulation of endothelial cells with TNF-α results in a NF-κB-guided redistribution of BRD4 in the genome, resulting in the formation of a de novo super-enhancer complex, which, in turn, can be abrogated by JQ1 (Brown et al., 2014). In addition, inhibition by JQ1 or knockdown of BRD2 and BRD4 genes with short hairpin RNA reduced NF-κB activation in response to TNF-α in human vascular endothelial cells (Huang, Zeng, et al., 2017). Lastly, in addition to binding acetylated histones, BRD4 was shown to directly bind the NF-κB p65 subunit acetylated by p300, thereby coactivating the genes involved in the inflammatory response (Huang et al., 2009). Taken together, the interference of NF-κB nuclear translocation by CN210 may play a role in its anti-inflammatory effects.
The MAPKs are a family of serine/threonine kinases that are activated by diverse stimulations and induce various cellular functions, including inflammatory responses. In mammals, the three major classes of MAPKs are ERK, p38 and JNK (Johnson & Lapadat, 2002). Previous studies demonstrated that BET inhibition reduced the activation of all three classes of MAPKs (Huang, Zeng, et al., 2017; Sun et al., 2018). In contrast, CN210 suppressed LPS-induced phosphorylation of JNK and ERK but not that of p38. The reason for this difference is unknown and warrants further investigation.
In this study, we found that CN210 showed competitive binding effects for the bromodomains of CBP and p300, in addition to its activity against BET proteins. From the docking studies, the isoxazole moiety of CN210 formed the same key H-bonding interaction with Asn residues, that is, N1168 in CBP and N1132 in p300. While it was predicted from the structure-based docking, the binding model suggests a potential potency improvement by the replacement of dimethylisoxazole, which serves as the acetyllysine mimic. Another putative π-edge binding interaction was predicted between W81 at the WPF shelf in BRD4 BD1 and the pyrazole ring of CN210, a Leu residue is found at this position in CBP and p300 proteins. The lack of this aromatic stacking interaction may contribute to the reduced binding affinity of CN210 against bromodomain of CBP/p300. Of note, the region at the opening of the binding pocket corresponding to D144/D145/M149 residues in BRD4 BD1 are varied to S1172/R1173/F1177 in CBP and S1136/R1137/Y1141 in p300, respectively. A bulkier 3-Cl-4-OMe substituted phenyl group of SGC-CBP30 was preferably accommodated at this region in CBP and p300, suggesting additional potentials for further optimization at this site to improve potency and selectivity against these bromodomain-containing proteins.
CBP and p300 are paralogous histone acetyltransferases and transcription coactivators (Giles et al., 1998; Shiama, 1997). CBP and p300 play important regulatory roles in gene expression through NF-κB- and MAPK-dependent pathways, especially JNK-dependent pathways (Hassa et al., 2005; Wang et al., 2015). A study with spleen cells derived from mice that lacked one allele of CBP demonstrated impaired induction of TNF-α gene expression in T cells in response to anti-CD3 antibody stimulation, implying that CBP has a key role in T cell immune response (Falvo et al., 2000). Another study showed that SGC-CBP30, an inhibitor specific for CBP/p300, reduced the production of IL-17α and other proinflammatory cytokines in immune cells (Hammitzsch et al., 2015). SGC-CBP30 also inhibited IL-17α secretion by Th17 cells from healthy donors, as well as from patients with ankylosing spondylitis and psoriatic arthritis (Hammitzsch et al., 2015). Therefore, the anti-inflammatory effects of CN210 may be in part mediated by inhibition of CBP and p300 in addition to BET proteins. Additional studies, including the examination of the effects of CN210 on histone acetyltransferase activity of CBP and p300, as well as on the transcriptional processes, are ongoing.
In conclusion, CN210 potently attenuates indomethacin-induced ileitis in mice by inhibiting cytokine expression, at least in part, by suppressing the activation of NF-κB and MAPK. CN210 is a BET bromodomain inhibitor that also inhibits the bromodomains of two paralogous histone acetyltransferases, CBP and p300. Thus, CN210 is a potential drug candidate with a unique mechanism of action to treat NSAID-induced ileitis and IBD.
Supplementary Material
ACKNOWLEDGMENTS
This research was supported in part by the JSPS KAKENHI Grant Number 20K07078 (Satsuki Kobayashi) and the Intramural research program of the National Center for Advancing Translational Sciences (NCATS), National Institutes of Health (Xin Hu, David J. Maloney, Shyh-Ming Yang).
Funding information
Intramural research program of the NCATS, NIH; Japan Society for the Promotion of Science, Grant/Award Number: 20K07078
Abbreviations:
- ANOVA
analysis of variance
- ARRIVE
Animal Research: Reporting In Vivo Experiments
- BET proteins
bromodomain and extra-terminal motif proteins
- BRD
bromodomaincontaining protein
- BRDT
bromodomain-testis-containing protein
- cAMP
cyclic adenosine monophosphate
- CBP
CREB binding protein
- CMC
carboxymethylcellulose
- CREB
cAMP-responsive element binding protein
- DMEM
Dulbecco’s modified Eagle medium
- DMSO
dimethyl sulfoxide
- ERK
extracellular signal-regulated kinase
- GBVI/WSA
generalized-born volume integral/weighted surface area
- IBD
inflammatory bowel disease
- IFN
interferon
- IL
interleukin
- JNK
c-JUN N-terminal kinase
- LPS
lipopolysaccharide
- MAPK
mitogen-activated protein kinases
- MOE
molecular operating environment
- MPO
myeloperoxidase
- NF-κB
nuclear factor κB
- NSAID
nonsteroidal anti-inflammatory drug
- PCR
quantitative polymerase chain reaction
- PVDF
polyvinylidene difluoride
- RIPA
radioimmunoprecipitation assay
- RT-PCR
reverse transcription-PCR
- SDS-PAGE
sodium dodecyl sulfate–polyacrylamide gel
- TBP
TATA-binding protein
- Th17
T helper 17
- TNF-α
tumor necrosis factor-α
Footnotes
CONFLICT OF INTEREST
The authors declare the following competing financial interests: Makoto Yoshioka is employed by ConverGene LLC and holds equity in ConverGene LLC.
DATA AVAILABILITY STATEMENT
The data that support the findings of this study are available from the corresponding author on reasonable request.
SUPPORTING INFORMATION
Additional supporting information may be found online in the Supporting Information section at the end of this article.
REFERENCES
- Anthony A, Pounder RE, Dhillon AP, & Wakefield AJ (2000). Similarities between ileal Crohn’s disease and indomethacin experimental jejunal ulcers in the rat. Alimentary Pharmacology & Therapeutics, 14, 241–245. [DOI] [PubMed] [Google Scholar]
- Bandukwala HS, Gagnon J, Togher S, Greenbaum JA, Lamperti ED, Parr NJ, Molesworth AM, Smithers N, Lee K, Witherington J, Tough DF, Prinjha RK, Peters B, & Rao A. (2012). Selective inhibition of CD4+ T-cell cytokine production and autoimmunity by BET protein and c-Myc inhibitors. Proceedings of the National Academy of Sciences of the United States of America, 109, 14532–14537. 10.1073/pnas.1212264109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belkina AC, Nikolajczyk BS, & Denis GV (2013). BET protein function is required for inflammation: Brd2 genetic disruption and BET inhibitor JQ1 impair mouse macrophage inflammatory responses. Journal of Immunology, 190, 3670–3678. 10.4049/jimmunol.1202838 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bjarnason I, Hayllar J, MacPherson AJ, & Russell AS (1993). Side effects of nonsteroidal anti-inflammatory drugs on the small and large intestine in humans. Gastroenterology, 104, 1832–1847. 10.1016/0016-5085(93)90667-2 [DOI] [PubMed] [Google Scholar]
- Brown JD, Lin CY, Duan Q, Griffin G, Federation A, Paranal RM, Bair S, Newton G, Lichtman A, Kung A, Yang T, Wang H, Luscinskas FW, Croce K, Bradner JE, & Plutzky J. (2014). NFkappaB directs dynamic super enhancer formation in inflammation and atherogenesis. Molecular Cell, 56, 219–231. 10.1016/j.molcel.2014.08.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung K, Lu G, Sharma R, Vincek A, Zhang R, Plotnikov AN, Zhang F, Zhang Q, Ju Y, Hu Y, Zhao L, Han ×, Meslamani J, Xu F, Jaganathan A, Shen T, Zhu H, Rusinova E, Zeng L, … Zhou MM (2017). BET N-terminal bromodomain inhibition selectively blocks Th17 cell differentiation and ameliorates colitis in mice. Proceedings of the National Academy of Sciences of the United States of America, 114, 2952–2957. 10.1073/pnas.1615601114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cochran AG, Conery AR, & Sims RJ 3rd. (2019). Bromodomains: A new target class for drug development. Nature Reviews. Drug Discovery, 18, 609–628. 10.1038/s41573-019-0030-7 [DOI] [PubMed] [Google Scholar]
- Delmore JE, Issa GC, Lemieux ME, Rahl PB, Shi J, Jacobs HM, Kastritis E, Gilpatrick T, Paranal RM, Qi J, Chesi M, Schinzel AC, McKeown MR, Heffernan TP, Vakoc CR, Bergsagel PL, Ghobrial IM, Richardson PG, Young RA, … Mitsiades CS (2011). BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell, 146, 904–917. 10.1016/j.cell.2011.08.017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding N, Hah N, Yu RT, Sherman MH, Benner C, Leblanc M, He M, Liddle C, Downes M, & Evans RM (2015). BRD4 is a novel therapeutic target for liver fibrosis. Proceedings of the National Academy of Sciences of the United States of America, 112, 15713–15718. 10.1073/pnas.1522163112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doroshow DB, Eder JP, & LoRusso PM (2017). BET inhibitors: a novel epigenetic approach. Annals of Oncology, 28, 1776–1787. 10.1093/annonc/mdx157 [DOI] [PubMed] [Google Scholar]
- Falvo JV, Brinkman BM, Tsytsykova AV, Tsai EY, Yao TP, Kung AL, & Goldfeld AE (2000). A stimulus-specific role for CREBbinding protein (CBP) in T cell receptor-activated tumor necrosis factor alpha gene expression. Proceedings of the National Academy of Sciences of the United States of America, 97, 3925–3929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feagan BG, Sandborn WJ, Gasink C, Jacobstein D, Lang Y, Friedman JR, Blank MA, Johanns J, Gao LL, Miao Y, Adedokun OJ, Sands BE, Hanauer SB, Vermeire S, Targan S, Ghosh S, de Villiers WJ, Colombel JF, Tulassay Z, … Group U-IUS. (2016). Ustekinumab as induction and maintenance therapy for Crohn’s disease. The New England Journal of Medicine, 375, 1946–1960. 10.1056/NEJMoa1602773 [DOI] [PubMed] [Google Scholar]
- Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert JP, Barsyte-Lovejoy D, Felletar I, Volkmer R, Muller S, Pawson T, Gingras AC, Arrowsmith CH, & Knapp S. (2012). Histone recognition and large-scale structural analysis of the human bromodomain family. Cell, 149, 214–231. 10.1016/j.cell.2012.02.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O, Morse EM, Keates T, Hickman TT, Felletar I, Philpott M, Munro S, McKeown MR, Wang Y, Christie AL, West N, Cameron MJ, Schwartz B, Heightman TD, … Bradner JE (2010). Selective inhibition of BET bromodomains. Nature, 468, 1067–1073. 10.1038/nature09504 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukumoto K, Naito Y, Takagi T, Yamada S, Horie R, Inoue K, Harusato A, Hirata I, Omatsu T, Mizushima K, Hirai Y, Yoshida N, Uchiyama K, Ishikawa T, Handa O, Konishi H, Wakabayashi N, Yagi N, Kokura S, … Yoshikawa T. (2011). Role of tumor necrosis factor-alpha in the pathogenesis of indomethacininduced small intestinal injury in mice. International Journal of Molecular Medicine, 27, 353–359. 10.3892/ijmm.2011.602 [DOI] [PubMed] [Google Scholar]
- Giles RH, Peters DJ, & Breuning MH (1998). Conjunction dysfunction: CBP/p300 in human disease. Trends in Genetics, 14, 178–183. [DOI] [PubMed] [Google Scholar]
- Hammitzsch A, Tallant C, Fedorov O, O’Mahony A, Brennan PE, Hay DA, Martinez FO, Al-Mossawi MH, de Wit J, Vecellio M, Wells C, Wordsworth P, Muller S, Knapp S, & Bowness P. (2015). CBP30, a selective CBP/p300 bromodomain inhibitor, suppresses human Th17 responses. Proceedings of the National Academy of Sciences of the United States of America, 112, 10768–10773. 10.1073/pnas.1501956112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanauer SB, Feagan BG, Lichtenstein GR, Mayer LF, Schreiber S, Colombel JF, Rachmilewitz D, Wolf DC, Olson A, Bao W, Rutgeerts P, & Group AIS. (2002). Maintenance infliximab for Crohn’s disease: The ACCENT I randomised trial. Lancet, 359, 1541–1549. 10.1016/S0140-6736(02)08512-4 [DOI] [PubMed] [Google Scholar]
- Hassa PO, Haenni SS, Buerki C, Meier NI, Lane WS, Owen H, Gersbach M, Imhof R, & Hottiger MO (2005). Acetylation of poly(ADP-ribose) polymerase-1 by p300/CREB-binding protein regulates coactivation of NF-kappaB-dependent transcription. The Journal of Biological Chemistry, 280, 40450–40464. [DOI] [PubMed] [Google Scholar]
- Huang B, Yang ×D, Zhou MM, Ozato K, & Chen LF (2009). Brd4 coactivates transcriptional activation of NF-kappaB via specific binding to acetylated RelA. Molecular and Cellular Biology, 29, 1375–1387. 10.1128/MCB.01365-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang M, Zeng S, Zou Y, Shi M, Qiu Q, ×iao Y, Chen G, Yang ×, Liang L, & ×u H. (2017). The suppression of bromodomain and extraterminal domain inhibits vascular inflammation by blocking NF-kappaB and MAPK activation. British Journal of Pharmacology, 174, 101–115. 10.1111/bph.13657 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang W, Haynes AC, Mukherjee R, Wen L, Latawiec D, Tepikin AV, Criddle DN, Prinjha RK, Smithers N, & Sutton R. (2017). Selective inhibition of BET proteins reduces pancreatic damage and systemic inflammation in bile acidand fatty acid ethyl esterbut not caerulein-induced acute pancreatitis. Pancreatology, 17, 689–697. 10.1016/j.pan.2017.06.005 [DOI] [PubMed] [Google Scholar]
- Johnson GL, & Lapadat R. (2002). Mitogen-activated protein kinase pathways mediated by ERK, JNK, and p38 protein kinases. Science, 298, 1911–1912. 10.1126/science.1072682 [DOI] [PubMed] [Google Scholar]
- Kato S, Matsuda N, Matsumoto K, Wada M, Onimaru N, Yasuda M, Amagase K, Horie S, & Takeuchi K. (2012). Dual role of serotonin in the pathogenesis of indomethacin-induced small intestinal ulceration: Pro-ulcerogenic action via 5-HT3 receptors and anti-ulcerogenic action via 5-HT4 receptors. Pharmacological Research, 66, 226–234. 10.1016/j.phrs.2012.06.002 [DOI] [PubMed] [Google Scholar]
- Kilkenny C, Browne WJ, Cuthill IC, Emerson M, & Altman DG (2010). Improving bioscience research reporting: The ARRIVE guidelines for reporting animal research. PLoS Biology, 8, e1000412. 10.1371/journal.pbio.1000412 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein K, Kabala PA, Grabiec AM, Gay RE, Kolling C, Lin LL, Gay S, Tak PP, Prinjha RK, Ospelt C, & Reedquist KA (2016). The bromodomain protein inhibitor I-BET151 suppresses expression of inflammatory genes and matrix degrading enzymes in rheumatoid arthritis synovial fibroblasts. Annals of the Rheumatic Diseases, 75, 422–429. 10.1136/annrheumdis-2014-205809 [DOI] [PubMed] [Google Scholar]
- Konaka A, Kato S, Tanaka A, Kunikata T, Korolkiewicz R, & Takeuchi K. (1999). Roles of enterobacteria, nitric oxide and neutrophil in pathogenesis of indomethacin-induced small intestinal lesions in rats. Pharmacological Research, 40, 517–524. 10.1006/phrs.1999.0550 [DOI] [PubMed] [Google Scholar]
- Loven J, Hoke HA, Lin CY, Lau A, Orlando DA, Vakoc CR, Bradner JE, Lee TI, & Young RA (2013). Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell, 153, 320–334. 10.1016/j.cell.2013.03.036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mele DA, Salmeron A, Ghosh S, Huang HR, Bryant BM, & Lora JM (2013). BET bromodomain inhibition suppresses TH17-mediated pathology. The Journal of Experimental Medicine, 210, 2181–2190. 10.1084/jem.20130376 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mertz JA, Conery AR, Bryant BM, Sandy P, Balasubramanian S, Mele DA, Bergeron L, & Sims RJ 3rd. (2011). Targeting MYC dependence in cancer by inhibiting BET bromodomains. Proceedings of the National Academy of Sciences of the United States of America, 108, 16669–16674. 10.1073/pnas.1108190108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muller S, Filippakopoulos P, & Knapp S. (2011). Bromodomains as therapeutic targets. Expert Reviews in Molecular Medicine, 13, e29. 10.1017/S1462399411001992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicodeme E, Jeffrey KL, Schaefer U, Beinke S, Dewell S, Chung CW, Chandwani R, Marazzi I, Wilson P, Coste H, White J, Kirilovsky J, Rice CM, Lora JM, Prinjha RK, Lee K, & Tarakhovsky A. (2010). Suppression of inflammation by a synthetic histone mimic. Nature, 468, 1119–1123. 10.1038/nature09589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rutgeerts P, Sandborn WJ, Feagan BG, Reinisch W, Olson A, Johanns J, Travers S, Rachmilewitz D, Hanauer SB, Lichtenstein GR, de Villiers WJ, Present D, Sands BE, & Colombel JF (2005). Infliximab for induction and maintenance therapy for ulcerative colitis. The New England Journal of Medicine, 353, 2462–2476. 10.1056/NEJMoa050516 [DOI] [PubMed] [Google Scholar]
- Schilderink R, Bell M, Reginato E, Patten C, Rioja I, Hilbers FW, Kabala PA, Reedquist KA, Tough DF, Tak PP, Prinjha RK, & de Jonge WJ (2016). BET bromodomain inhibition reduces maturation and enhances tolerogenic properties of human and mouse dendritic cells. Molecular Immunology, 79, 66–76. 10.1016/j.molimm.2016.09.010 [DOI] [PubMed] [Google Scholar]
- Shiama N. (1997). The p300/CBP family: Integrating signals with transcription factors and chromatin. Trends in Cell Biology, 7, 230–236. 10.1016/S0962-8924(97)01048-9 [DOI] [PubMed] [Google Scholar]
- Smith SG, & Zhou MM (2016). The bromodomain: A new target in emerging epigenetic medicine. ACS Chemical Biology, 11, 598–608. 10.1021/acschembio.5b00831 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suarez-Alvarez B, Morgado-Pascual JL, Rayego-Mateos S, Rodriguez RM, Rodrigues-Diez R, Cannata-Ortiz P, Sanz AB, Egido J, Tharaux PL, Ortiz A, Lopez-Larrea C, & Ruiz-Ortega M. (2017). Inhibition of bromodomain and extraterminal domain family proteins ameliorates experimental renal damage. Journal of the American Society of Nephrology, 28, 504–519. 10.1681/ASN.2015080910 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun L, Liu J, Yuan Y, Zhang ×, & Dong Z. (2018). Protective effect of the BET protein inhibitor JQ1 in cisplatin-induced nephrotoxicity. American Journal of Physiology. Renal Physiology, 315, F469–F478. 10.1152/ajprenal.00527.2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szabo SJ, Sullivan BM, Peng SL, & Glimcher LH (2003). Molecular mechanisms regulating Th1 immune responses. Annual Review of Immunology, 21, 713–758. 10.1146/annurev.immunol.21.120601.140942 [DOI] [PubMed] [Google Scholar]
- Tsukahara T, Hamouda N, Utsumi D, Matsumoto K, Amagase K, & Kato S. (2017). G protein-coupled receptor 35 contributes to mucosal repair in mice via migration of colonic epithelial cells. Pharmacological Research, 123, 27–39. 10.1016/j.phrs.2017.06.009 [DOI] [PubMed] [Google Scholar]
- Villar-Prados A, Wu SY, Court KA, Ma S, LaFargue C, Chowdhury MA, Engelhardt MI, Ivan C, Ram PT, Wang Y, Baggerly K, Rodriguez-Aguayo C, Lopez-Berestein G, Ming-Yang S, Maloney DJ, Yoshioka M, Strovel JW, Roszik J, & Sood AK (2019). Predicting novel therapies and targets: Regulation of Notch3 by the bromodomain protein BRD4. Molecular Cancer Therapeutics, 18, 421–436. 10.1158/1535-7163.MCT-18-0365 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang CY, & Filippakopoulos P. (2015). Beating the odds: BETs in disease. Trends in Biochemical Sciences, 40, 468–479. 10.1016/j.tibs.2015.06.002 [DOI] [PubMed] [Google Scholar]
- Wang Y, Wang Y, Luo M, Wu H, Kong L, ×in Y, Cui W, Zhao Y, Wang J, Liang G, Miao L, & Cai L. (2015). Novel curcumin analog C66 prevents diabetic nephropathy via JNK pathway with the involvement of p300/CBP-mediated histone acetylation. Biochimica et Biophysica Acta, 1852, 34–46. 10.1016/j.bbadis.2014.11.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, Rahl PB, Lee TI, & Young RA (2013). Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell, 153, 307–319. 10.1016/j.cell.2013.03.035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu SY, & Chiang CM (2007). The double bromodomain-containing chromatin adaptor Brd4 and transcriptional regulation. The Journal of Biological Chemistry, 282, 13141–13145. 10.1074/jbc.R700001200 [DOI] [PubMed] [Google Scholar]
- Yamada S, Naito Y, Takagi T, Mizushima K, Hirai Y, Horie R, Fukumoto K, Inoue K, Harusato A, Yoshida N, Uchiyama K, Handa O, Ishikawa T, Konishi H, Wakabayashi N, Yagi N, Kokura S, Kita M, & Yoshikawa T. (2011). Reduced small-intestinal injury induced by indomethacin in interleukin-17A-deficient mice. Journal of Gastroenterology and Hepatology, 26, 398–404. 10.1111/j.1440-1746.2010.06496.x [DOI] [PubMed] [Google Scholar]
- Yamada T, Deitch E, Specian RD, Perry MA, Sartor RB, & Grisham MB (1993). Mechanisms of acute and chronic intestinal inflammation induced by indomethacin. Inflammation, 17, 641–662. [DOI] [PubMed] [Google Scholar]
- Yang SM, Urban DJ, Yoshioka M, Strovel JW, Fletcher S, Wang AQ, Xu X, Shah P, Hu X, Hall MD, Jadhav A, & Maloney DJ (2018). Discovery and lead identification of quinazoline-based BRD4 inhibitors. Bioorganic & Medicinal Chemistry Letters, 28, 3483–3488. 10.1016/j.bmcl.2018.08.039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang SM, Yoshioka M, Strovel JW, Urban DJ, Hu X, Hall MD, Jadhav A, & Maloney DJ (2019). Lead optimization and efficacy evaluation of quinazoline-based BET family inhibitors for potential treatment of cancer and inflammatory diseases. Bioorganic & Medicinal Chemistry Letters, 29, 1220–1226. 10.1016/j.bmcl.2019.03.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yasuda M, Kawahara R, Hashimura H, Yamanaka N, Iimori M, Amagase K, Kato S, & Takeuchi K. (2011). Dopamine D2–receptor antagonists ameliorate indomethacin-induced small intestinal ulceration in mice by activating α7 nicotinic acetylcholine receptors. Journal of Pharmacological Sciences, 116, 274–282. 10.1254/jphs.11037FP [DOI] [PubMed] [Google Scholar]
- Zhu J, Yamane H, & Paul WE (2010). Differentiation of effector CD4 T cell populations (*). Annual Review of Immunology, 28, 445–489. 10.1146/annurev-immunol-030409-101212 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.