Abstract
Candida glabrata is the second leading cause of candidemia in many countries and is one of the most concerning yeast species of nosocomial importance due to its increasing rate of antifungal drug resistance and emerging multidrug-resistant isolates. Application of multilocus sequence typing (MLST) to clinical C. glabrata isolates revealed an association of certain sequence types (STs) with drug resistance and mortality. The current C. glabrata MLST scheme is based on single nucleotide polymorphisms (SNPs) at six loci and is therefore relatively laborious and costly. Furthermore, only a few high-quality C. glabrata reference genomes are available, limiting rapid analysis of clinical isolates by whole genome sequencing. In this study we provide long-read based assemblies for seven additional clinical strains belonging to three different STs and use this information to simplify the C. glabrata MLST scheme. Specifically, a comparison of these genomes identified highly polymorphic loci (HPL) defined by frequent insertions and deletions (indels), two of which proved to be highly resolutive for ST. When challenged with 53 additional isolates, a combination of TRP1 (a component of the current MLST scheme) with either of the two HPL fully recapitulated ST identification. Therefore, our comparative genomic analysis identified a new typing approach combining SNPs and indels and based on only two loci, thus significantly simplifying ST identification in C. glabrata. Because typing tools are instrumental in addressing numerous clinical and biological questions, our new MLST scheme can be used for high throughput typing of C. glabrata in clinical and research settings.
Key words: Candida glabrata, MLST, Sequence type, Whole-genome sequencing
Introduction
Candida species are among the most prevalent mycobiome constituents (Romo & Kumamoto 2020) and also a major cause of invasive fungal infections in humans worldwide (Brown et al. 2012). It has been estimated that > 400 000 life-threatening infections are caused by Candida albicans alone (Brown et al. 2012). Although C. albicans once was the most prevalent cause of candidemia, recent studies have revealed a stark increase in candidemias caused by non-albicans Candida (NAC) species, especially C. glabrata, C. parapsilosis, and C. tropicalis (Astvad et al. 2018, Lamoth et al., 2018, Fuller et al. 2019; Pfaller et al., 2019, Song et al. 2020, Stavrou et al., 2019; Won et al. 2021). Candida glabrata has been identified as the second leading cause of candidemia in the US (Pfaller et al. 2009, Pfaller et al., 2019, Tsay et al. 2020), Canada (Fuller et al. 2019), Australia (Chapman et al. 2017), and some European (Astvad et al. 2018) and Asian countries (Taj-Aldeen et al. 2014, Arastehfar et al. 2020a, Kord et al. 2020). Candida glabrata has reduced susceptibility to fluconazole (Healey & Perlin 2018, Arastehfar et al. 2020b) and can rapidly develop drug resistance during infection (Healey & Perlin 2018, Ksiezopolska & Gabaldón 2018, Arastehfar et al. 2020b). Indeed, the increasing incidence of candidemia due to C. glabrata in recent years has coincided with a notable increase in the number of fluconazole-resistant (FLZR) isolates in many countries (Hou et al. 2017, Astvad et al. 2018, Pfaller et al., 2019, Arastehfar et al. 2020c, Won et al. 2021). Patients infected with FLZR C. glabrata isolates had the highest mortality rate and experienced a shorter median survival days after diagnosis compared to those infected with fluconazole-susceptible-dose-dependent (FLZ-SDD) isolates (Won et al. 2021). These observations are especially concerning for developing countries, where fluconazole is the frontline antifungal drug used to treat candidemia (Chakrabarti et al. 2015, Arastehfar et al. 2020d, Kord et al. 2020, Megri et al. 2020). However, the emerging echinocandin-resistant (ECR) and multidrug-resistant (MDR) C. glabrata isolates, which constitute > 30 % of the ECR isolates (Astvad et al. 2018, Won et al. 2021), may also threaten the clinical efficacy of echinocandins, the frontline antifungal drugs recommended by international guidelines (Pappas et al., 2016). Together, these features make C. glabrata one of the most challenging fungal pathogens of the present time.
Although C. glabrata is considered to be predominantly asexual, various types of genome analyses of C. glabrata clinical isolates revealed a high degree of genetic diversity both in terms of chromosome structure and sequence (Carreté et al. 2018, Xu et al. 2021), and this diversity was estimated to be greater than that of C. albicans (Carreté et al. 2018, Gabaldón & Fairhead 2019). Understanding the genetic diversity of C. glabrata isolates from both clinical and biological standpoints is of paramount importance, as it will help to answer key questions regarding its epidemiology, aid in implementing appropriate infection control strategies by identifying the route of infection (Megri et al. 2020), and also enables researchers to uncover the evolution of drug resistance of genetically-related isolates during the course of infection (Carreté et al. 2019). Additionally, several studies have uncovered an association between genotype and mortality (Byun et al. 2018, Arastehfar et al. 2019) and drug resistance (Won et al. 2021). Studies dissecting the genetic structure of C. glabrata isolates have employed various tools, such as whole-genome sequencing (WGS) (Biswas et al. 2018, Carreté et al. 2018, 2019), polymorphic locus sequence typing (Katiyar et al. 2016, Katiyar & Edlind 2021), pulsed field gel electrophoresis (PFGE) (Lin et al. 2007), amplified fragment length polymorphism analysis (Arastehfar et al. 2019), multilocus microsatellite typing (MLMT) (Hou et al., 2018, Bordallo-Cardona et al., 2019, Arastehfar et al. 2020c), and multilocus sequence typing (MLST) (Lott et al. 2010, 2012, Hou et al. 2017, Hou et al., 2018, Byun et al. 2018, Bordallo-Cardona et al., 2019, Khalifa et al. 2020, Won et al. 2021). Among these, WGS provides the most information and resolution, and short read based and nanopore long read based sequencing platforms are gradually becoming more accessible and affordable (Gabaldón 2019). However, the extensive chromosome structure diversity between clinical isolates makes it difficult to use standard tools for short read sequence analysis, which rely on mapping the reads to a reference genome with a common structure. Therefore, there is a need for additional high quality assembled C. glabrata genomes that can serve as references, facilitating WGS analysis of clinical isolates.
Although WGS provides the most information and resolution, the still high costs and sophisticated analysis hinder its wide use in clinical settings, especially among medical mycologists (Gabaldón et al. 2020). Therefore, most of the studies conducted so far have used MLMT and MLST as the most popular techniques to type clinical C. glabrata isolates (Gabaldón et al. 2020). The MLST method is highly reproducible and provides data that are consistent with WGS (Carreté et al. 2018). These data can be archived at “https://pubmlst.org/organisms/candida-glabrata” (Gabaldón et al. 2020), making the worldwide, temporal, and nationwide analysis of clinical C. glabrata isolates possible. The currently used MLST scheme (Dodgson et al. 2003) has identified 1 414 isolates of C. glabrata belonging to > 100 STs, which underscores its genetic diversity. This scheme is based on SNPs at six loci (FKS1, LEU2, NMT1, TRP1, UGP1, and URA3), together comprising > 3 000 bps. Of course, it is desirable to reduce the required number of loci to reduce the time and cost associated with MLST analysis, without losing resolutive power.
In this study, we obtained high quality genome data based on PacBio and Illumina sequencing for seven additional clinical isolates of C. glabrata belonging to ST10, 15, and 16. Comparative analysis of the seven genomes identified a number of highly polymorphic loci (HPL) characterized by a high number of insertions/deletions (indels). These loci were enriched for the functional categories governing stress response pathways, particularly those involved in membrane and cell wall stresses. Interestingly, we found that two of these HPLs were highly resolutive for ST identity. Using an additional set of 53 clinical isolates showed that each of these loci in combination with TRP1 provided the same resolution as that generated by the traditional six-locus MLST protocol (Dodgson et al. 2003). Therefore, we propose an improved MLST protocol for C. glabrata, which necessitates sequencing only two loci (totalling ≤1 Kbp) and offers a quicker and cheaper approach without the loss of resolution, and which can be used for high throughput typing studies of C. glabrata.
Materials and methods
Isolates and growth conditions
The 53 C. glabrata strains used in this study are listed in Table 1. The isolates originated from various geographical regions and had various antifungal susceptibility profiles. All isolates were grown on yeast peptone dextrose (YPD) agar and incubated at 37 °C for 24–48 h.
Table 1.
List of clinical Candida glabrata isolates used in this study.
| Isolate # | Original # | Sequence type | Minimum inhibitory concentration (μg/ml) |
Susceptibility profiles | Comments | |||
|---|---|---|---|---|---|---|---|---|
| Fluconazole | Micafungin | Anidulafungin | AMB | |||||
| 1 | CAS08-0293 | ST3 | 64 | 4 | 4 | 1 | MDR | |
| 2 | CAS08-0725 (CMD00311) | ST3 | 64 | 0.5 | 2 | 1 | MDR | |
| 3 | CAS09-0869 (CMD00373) | ST3 | 64 | 0.5 | 2 | 1 | MDR | |
| 4 | BG2 | ST3 | 4 | 0.015 | 0.125 | 1 | S | |
| 5 | Qatar 36 | ST3 | 4 | 0.015 | 0.125 | 0.5 | S | |
| 6 | DPK 159 | ST3 | 4 | 0.015 | 0.125 | 1 | S | |
| 7 | DPL27 (5962) | ST3 | 4 | 2 | 4 or 2 | 1 | ECR | |
| 8 | ATCC 66032 | ST6 | 4 | 0.015 | 0.06 | 0.5 | S | |
| 9 | CAS09-1225 (CGA00908) | ST6 | > 64 | 4 | 2 | 2 | MDR | |
| 10 | CAS09-1786 (CGA01258) | ST6 | 64 | 4 | 2 | 1 | MDR | |
| 11 | DPL155 (M234) | ST6 | 1 | 0.25 | 1 | 1 | ECR | |
| 12 | DPL157 (17351, CGC2) | ST6 | 64 | 4 | 4 | 1 | MDR | |
| 16 | CAS08-0089 | ST8 | 4 | 0.015 | 0.125 | 1 | S | |
| 17 | CAS08-494 | ST8 | 4 | 0.015 | 0.125 | 1 | S | |
| 18 | CAS08-528 | ST8 | 4 | 0.015 | 0.125 | 1 | S | |
| 19 | CAS11-3112 (CGA02083) | ST8 | 64 | 0.25 | 0.5 | 1 | MDR | |
| 20 | DPL209 (M2952) | STX | 64 | 4 | 4 | 1 | MDR | |
| 25 | CAS08-0205 | ST10 | 4 | 0.015 | 0.125 | 0.5 | S | |
| 26 | CAS09-1083 (CGA00822) | ST10 | 64 | 0.5 | 1 | 0.5 | MDR | |
| 27 | CAS09-1437 (CGA01045) | ST10 | 64 | 2 | 1 or 0.5 | 1 | MDR | |
| 28 | DPK 305 | ST10 | 4 | 0.015 | 0.125 | 1 | S | WGS |
| 29 | CAS08-0425 (CMD00008) | ST10 | 64 | 4 | 4 | 1 | MDR | WGS |
| 35 | DPL1021 (ATCC 90030) | ST10 | 4 | 0.015 | 0.125 | 1 | S | WGS |
| 21 | Qatar 38 | ST15 | 4 | 0.015 | 0.25 | 1 | S | |
| 22 | Qatar 57 | ST15 | 4 | 0.015 | 0.125 | 1 | S | |
| 23 | Qatar 71 | ST15 | > 64 | 0.06 | 0.25 | 1 | FLZR | |
| 24 | ATCC 2001 (CBS 138) | ST15 | 4 | 0.015 | 0.125 | 0.5 | S | |
| 30 | CAS08-0027 | ST15 | 64 | 0.015 | 0.125 | 1 | FLZR | WGS |
| 31 | CAS08-485 | ST15 | 4 | 0.015 | 0.125 | 1 | S | |
| 32 | CAS09-755 | ST15 | 4 | 0.015 | 0.125 | 1 | S | |
| 33 | DPK 762 | ST15 | 4 | 0.015 | 0.125 | 1 | S | WGS |
| 34 | DPL274 (3-CPH-W20800) | ST15 | 1 | 1 | 4 | 4 | ECR | |
| 36 | CAS08-0092 | STY | 2 | 0.015 | 0.125 | 1 | S | |
| 37 | CAS11-3129 (CMD01408 | STY | 64 | 4 | 4 | 0.5 | MDR | |
| 38 | DPL245 (1611) | STY | 16 | 4 | 4 | 0.5 | ECR | WGS |
| 39 | DPL38 (42997) | STY | 32 | 1 | 2 | 0.25 | ECR | |
| 40 | CAS08-0016 (CGA00019) | STY | 64 | 2 | 1 | 1 | MDR | WGS |
| 41 | DPL217 | ST17 | 2 | 0.5 | 1 | 0.5 | ECR | |
| 42 | DPL219 | ST17 | 2 | 0.5 | 1 | 0.5 | ECR | |
| 44 | LB599-02 | ST46 | 4 | 0.015 | 0.125 | 1 | S | |
| 45 | LB906-05 | ST46 | 64 | 0.015 | 0.125 | 1 | FLZR | |
| 46 | Qatar 51 | ST46 | 2 | 0.015 | 0.125 | 1 | S | |
| 48 | Qatar 59 | ST46 | 2 | 0.015 | 0.06 | 1 | S | |
| 50 | Qatar 62 | ST46 | 2 | 0.015 | 0.06 | 1 | S | |
| 53 | Qatar 69 | ST46 | 4 | 0.03 | 0.25 | 1 | S | |
| 54 | Qatar 72 | ST46 | 64 | 0.015 | 0.125 | 1 | FLZR | |
| 56 | Qatar 76 | ST46 | 4 | 0.015 | 0.125 | 1 | S | |
| 47 | Qatar 53 | ST7 | 2 | 0.015 | 0.06 | 1 | S | |
| 49 | Qatar 61 | ST7 | 4 | 0.015 | 0.06 | 1 | S | |
| 51 | Qatar 63 | ST7 | 4 | 0.015 | 0.06 | 1 | S | |
| 52 | Qatar 64 | ST7 | 2 | 0.015 | 0.06 | 1 | S | |
| 55 | Qatar 73 | ST7 | 4 | 0.015 | 0.125 | 1 | S | |
| 57 | CAS08-0094 | ST76 | > 64 | 2 | 4 | 1 | MDR | |
AMB: Amphotericin B, MDR: Multidrug-resistant, S: Susceptible, ECR: Echinocandin-resistant, FLZR: Fluconazole-resistant, WGS: Whole-genome sequenced.
To enrich the genomes available for comparative purposes and due to the limited number of high-quality genome data, we performed whole-genome PacBio and Illumina sequencing of seven isolates, DPK305, CAS08-0425, DPK762, DPL1021, CAS080027, DPL245, CAS08-0016 (Table 1), which belonged to three STs (10, 15, and 16) and had different susceptibility profiles.
Antifungal susceptibility testing (AFST)
Antifungal susceptibility testing (AFST) followed the CLSI-M60 (Clinical and Laboratory Standards Institute. 2017) protocol and included fluconazole (Pfizer, New York, NY, USA), amphotericin B (AMB) (Sigma-Aldrich, Milwaukee, WI, USA), micafungin (Astellas Pharma, Tokyo, Japan), and anidulafungin (Pfizer). Plates containing antifungal drug-C. glabrata cell suspensions were incubated at 37 °C for 24 h and minimum inhibitory concentrations (MICs) were visually recorded. Type strains of C. krusei (ATCC 6258) and C. parapsilosis (ATCC 22019) were used in each individual AFST experiment for quality control purposes. Susceptibility profiles were denoted based on the availability of the clinical breakpoints and epidemiological cut-off values (ECVs) as suggested (Pfaller & Diekema 2012). Briefly, isolates showing MIC ≥ 64 μg/ml, ≥ 0.5 μg/ml, and ≥ 0.25 μg/ml were regarded as fluconazole-resistant, anidulafungin-resistant, and micafungin-resistant, respectively (Pfaller & Diekema 2012). The MIC data of AMB was interpreted based on ECVs and isolates showing a MIC > 2μg/ml were regarded as non-wild-type (Pfaller & Diekema 2012).
DNA extraction
The DNA extraction method varied depending on the sequencing method used. DNA samples subjected to Illumina and Sanger sequencing were extracted using the Quick-DNA Fungal/Bacterial Miniprep Kit (ZymoResearch, Irvine, CA, USA) following the protocol suggested by manufacturer. DNA isolation for PacBio sequencing followed an old-fashioned DNA isolation protocol. Briefly, overnight C. glabrata cultures were harvested by centrifugation, the cells were disrupted by vortexing in lysis buffer (100 mM Tris pH 8.0; 50 mM EDTA; 1 % SDS), and the liquid phase was collected. This was followed by incubation with ammonium sulphate, followed by chloroform extraction, and isopropanol precipitation. Finally, the pellets were washed with 70 % ethanol, air-dried, and resuspended in RNase/DNase free water.
Illumina library preparation and WGS
Genomic DNA was quantified using the Qubit v. 2.0 Fluorometer (Life Technologies, Carlsbad, CA, USA). The DNA integrity was checked with ∼1 % agarose gel with 50–100 ng sample loaded in each well. Samples were then chosen for library preparation based on the QC results. NEBNext® Ultra™ II DNA Library Prep Kit for Illumina, clustering, and sequencing reagents were used throughout the process following the manufacturer’s recommendations. Briefly, the genomic DNA was fragmented by acoustic shearing with a Covaris S220 instrument. Fragmented DNA was cleaned up and end repaired. Adapters were ligated after adenylation of the 3’ ends followed by enrichment by limited cycle PCR. DNA libraries were validated using a DNA 1000 Chip on the Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA, USA), and were quantified using Qubit v. 2.0 Fluorometer. The DNA libraries were also quantified by real-time PCR (Applied Biosystems, Carlsbad, CA, USA), clustered on one lane of a flowcell, and loaded on the Illumina HiSeq instrument according to manufacturer’s instructions. The samples were sequenced using a 2× 150 paired-end (PE) configuration. Image analysis and base calling were conducted by the HiSeq Control Software (HCS) on the HiSeq instrument. Raw sequence data (.bcl files) generated from Illumina HiSeq were converted into fastq files and de-multiplexed using Illumina's bcl2fastq v. 2.17 software. One mismatch was allowed for index sequence identification.
PacBio library preparation and genome assembly
PacBio DNA sequencing of C. glabrata strains was performed at the Waksman Institute, Rutgers University. Library preparation closely followed the multiplexed microbial library protocol provided by Pacific Biosciences. Briefly, the DNA quality was evaluated using a Qubit fluorometer (Thermo Fisher Scientific, Waltham, MA, USA) and Nanodrop spectrophotometer (Thermo Fisher Scientific). Two micrograms genomic DNA was fragmented using Magaruptor (Diagenode, Glen Ridge, NJ, USA) with the 10 kbp fragments settings according to the manufacturer's instructions. This is followed by 0.45× AMPure PB Bead Purification and removal of Single-Strand Overhangs. Samples were then subjected to DNA Damage Repair, End Repair, A-tailing and ligation to prepare SMRTbell DNA template libraries. After another round of 0.45× PB Bead Purification libraries were pooled in equimolar ratio and size selected with BluePippin (Sage Science, Beverly, MA, USA). Library quality was analysed by Qubit, and average fragment size was estimated using an Agilent Fragment Analyzer (Agilent, Santa Clara, CA, USA). SMRT sequencing was performed using a Pacific Biosciences Sequel I sequencer (PacBio, Menlo Park, CA, USA) and standard protocols (MagBead Standard Seq v. 2 loading, 600 min movie) using SMRT Cell 1M v. 2. Single molecule real-time sequencing reads were demultiplexed using SMRT Analysis software suite v. 5.1 (Pacific Biosciences Inc., Menlo Park, CA, USA) and de novo assembled using the CANU v. 1.7 workflow (Koren et al. 2017). Scaffolding was performed using the SSPACE long-read scaffolder (Boetzer & Pirovano 2014). Genome assembly was further improved for assembly contiguity using PBJelly (English et al. 2012). The PBJelly running parameters were as follows: -minMatch 8 -sdpTupleSize 8 -minPctIdentity 75 -bestn 1 -nCandidates 10 -maxScore -500 -nproc 13 -noSplitSubreads. WGS data obtained from Illumina was used to correct the those generated by PacBio. SNP calling was performed by aligning reads from each strain to each of the other strains using BWA mem v. 0.7.17-r1188 (Li & Durbin 2009) and GATK v. 4.0.4.0 (DePristo et al. 2011).
Annotation and gene prediction
Gene prediction was performed using a combination of methods. Reference sets were formed by the annotation of the reference genome of C. glabrata, and the collection of proteomes used in YGOB (Byrne & Wolfe 2005). Exonerate v. 2.4.0 (Slater & Birney 2005) was used to search for the presence of each protein in YGOB against each of the genomes. RATT (Otto et al. 2011) (downloaded 2018) from the PAGIT package was used to transfer annotations from the C. glabrata reference genome to the strain genomes; it was run in three modes: strain, species and multi, and the best transference was kept. YGAP web server (Proux-Wéra et al. 2012) (accessed October 2020) was then used to obtain a second gene prediction. MAKER2 v. 2.31.10 (Holt & Yandell 2011) was the third annotation program used. Results from RATT, YGAP, MAKER and exonerate were joined together into a single gene prediction using EVM (Haas et al. 2008) (downloaded 2018). This annotation was then improved by searching for the presence of genes that were predicted in the C. glabrata reference but not included in the strain annotation. As C. glabrata strains tend to be highly syntenic, the pipeline first associated each predicted protein in the strain genome to the reference genome using a best reciprocal hit approach. Then, based on location, it tried to associate unmatched genes if enough similarity was found even if they were not best reciprocal hits. In the same way it corrected spurious matches that were not congruent with the gene order conservation. Once the list of missing genes was found, the pipeline scanned the RATT annotation for those genes. If found they were incorporated to the gene prediction, if not the pipeline located surrounding genes and then used GenomeThreader v. 1.7.1 (Gremme et al. 2005) to search the intergenic space between those genes for the presence of the missing genes. In a last step, for genes still not found, the whole genome was scanned for their presence.
Identification of highly polymorphic loci (HPL) and primer design
Individual chromosomes of the seven clinical isolates (Table 1) were aligned with those of the reference strain CBS138/ATCC2001 (Xu et al. 2021) and strain DSY562 (Vale-Silva et al. 2017) using the Mauve alignment tool (DNASTAR, Madison, USA). Highly polymorphic loci (HPL) were defined based on the presence of indels that were different between strains of different STs. To avoid overlap with previously published sets of loci, HPL associated with satellite regions, megasatellites, minisatellites, and microsatellites, were excluded (Thierry et al. 2008). Loci obtained were subjected to gene ontology (GO) enrichment using FungiFun (Priebe et al. 2015), and those involved in cellular integrity pathways (osmotic stress tolerance, cell wall integrity and membrane integrity pathways, etc.) were selected for primer design. The primers were chosen to amplify the shortest possible regions without losing any indel information. All primer information is listed in Table 2.
Table 2.
Oligos used in this study, their functions, and their PCR conditions.
| Oligo name | Function | Oligo sequence (5'--3') |
|---|---|---|
| USA1-F | PCR/Sequencing | CCTGGAGAAGATGTATGTGTT |
|
USA1-R |
PCR/Sequencing |
TCTTCATGGTCGTGCTGAT |
| DUF1-F | PCR/Sequencing | TATCAAGTGTGTGGTGCC |
|
DUF1-R |
PCR/Sequencing |
CGTGTTCGACAGATTGTCC |
| SLG1-F | PCR/Sequencing | GATGCTACTTATACTGGCGG |
|
SLG1-R |
PCR/Sequencing |
TCAATCGGTTTCGTCTGG |
| HKR1-F | PCR/Sequencing | GAGAAAGCTATTGCTTTTGGT |
|
HKR1-R |
PCR/Sequencing |
TCAATAGATGATGGCTGCAC |
| H09053-F | PCR/Sequencing | AATACGAAAGCCACGACG |
|
H09053-R |
PCR/Sequencing |
ATATCTGGAATGCACTACCTG |
| A02255-F | PCR/Sequencing | TGGATTCGAATGAGGGACT |
|
A02255-R |
PCR/Sequencing |
CGAGTTCACACCGATTATCC |
| G00825-Fex | PCR/Sequencing | TCAAATGCTCCTCCTGGC |
| C00825-F1 | Sequencing | CTCTACAGGCGGCAAAA |
| G00825-R1 | Sequencing | GAATAATTAGAGTCGCCTCCG |
| 25F-N | Sequencing | TGG CAG AAA TAA ACG CCA G |
|
G00825-Rex |
PCR/Sequencing |
CCAACATCAATTTCAGGAGC |
| SSR1-F | PCR/Sequencing | GAGCTGGAACTCGATCCG |
|
SSR1-R |
PCR/Sequencing |
AGGAAGGGGAGTAATGATGG |
| PIR2-F | PCR/Sequencing | ATGCAATACAAAAAGACTCTAGC |
|
PIR2-R |
PCR/Sequencing |
TGGATCATTTGGTGCCTTAC |
| PIR1-F | PCR/Sequencing | CTTCTTCCTCTGTCGCTAAG |
|
PIR1-R |
PCR/Sequencing |
CAATTTGAGAAGCAGCGC |
| C00715-F | PCR/Sequencing | AGCCTCTGTCCCTACTTTATC |
|
C00715-R |
PCR/Sequencing |
GTCGCTGTTGGTGTGTAA |
| G03839-F | PCR/Sequencing | CCCAATCCCTTTCTCTGCT |
|
G03839-R |
PCR/Sequencing |
TATCCTCTTCTCATCGCTCG |
| G06281-F | PCR/Sequencing | GATGTTATGGTCTAGCTTTGC |
| G06281-R | PCR/Sequencing | CTGCCTATCTTAGATTGCTAGA |
Note that except for G00825 and PIRI, the rest of the loci had the same PCR program (94 °C 5 mins, 35 cycles of (94 °C 30 s, 60 °C 30 s, 72 °C 40 s), 72 °C 8 mins). PCR programs for G00825 was 94 °C 5 mins, 35 cycles of (94 °C 30 s, 58 °C 30 s, 72 °C 2 mins), 72 °C 8 mins and for PIRI was 94 °C 5 mins, 35 cycles of (94 °C 30 s, 58 °C 30 s, 72 °C 40 s), 72 °C 8 mins.
MLST schemes used and tree construction
The original MLST scheme developed by Dodgson et al. (2003) was used as the gold standard (Dodgson et al. 2003). Strain types (STs) were determined with the use of the https://pubmlst.org/about-us database. For each strain, a BLAST search was performed to determine the allele identity of each gene. Then the allelic combination was introduced in order to identify the STs. In one case, the allelic combination did not produce a known sequence type, and was named STX.
The trees obtained from the MLST scheme based on HPL were categorised as follows: trees that were obtained based on the concatenation of all 13 HPL, those constructed based on each individual HPL, those generated based on combination of two most resolutive HPL, and trees constructed based on the combination of the most resolutive HPL in combination with each loci of the original MLST scheme (Dodgson et al. 2003).
Robinson and Foulds comparison
ETE v. 3 was used to calculate the normalized Robinson and Foulds (RF) distances between trees (Huerta-Cepas et al. 2010). To calculate this value trees were first rooted to one of the leaves so that all trees had potentially the same structure. The RF calculates the number of common partitions between two trees. Normalization was carried out by dividing this value over the total number of partitions found in both trees. A high RF indicates little overlap between the two trees, although a single leaf moving to a very different position in the tree can cause a marked increase in the RF metric as it will affect many partitions.
Clade comparison
We used ETE v. 3 to assess the monophyly of sequences from the same ST. This would indicate that the sequences used to reconstruct the tree have the capacity to correctly classify strains into the same clades as the MLST genes. In order to quantify the consistency between clades we calculated the precision, recall and F1 values of each ST in the following way: for each node in the tree, we established to which ST most of the sequences belong. Considering all OTUs (strains) contained within this partition, we considered as true positives (TP) the strains that belonged to the ST, and false positives (FP) those belonging to alternative STs, how many strains from the same ST appeared outside that partition, false negatives (FN) and how many leaves from the rest of the tree were from a different ST, true negatives (TN). For each node we calculated the precision, recall and F1 for the chosen ST. Then, for each ST we selected the node that offered the best F1. So for a ST that was found completely monophyletic we would obtain a F1 of 1.0. For each tree, we calculated the average precision, recall and F1 of all the ST groups.
Results
Genome variation among the isolates
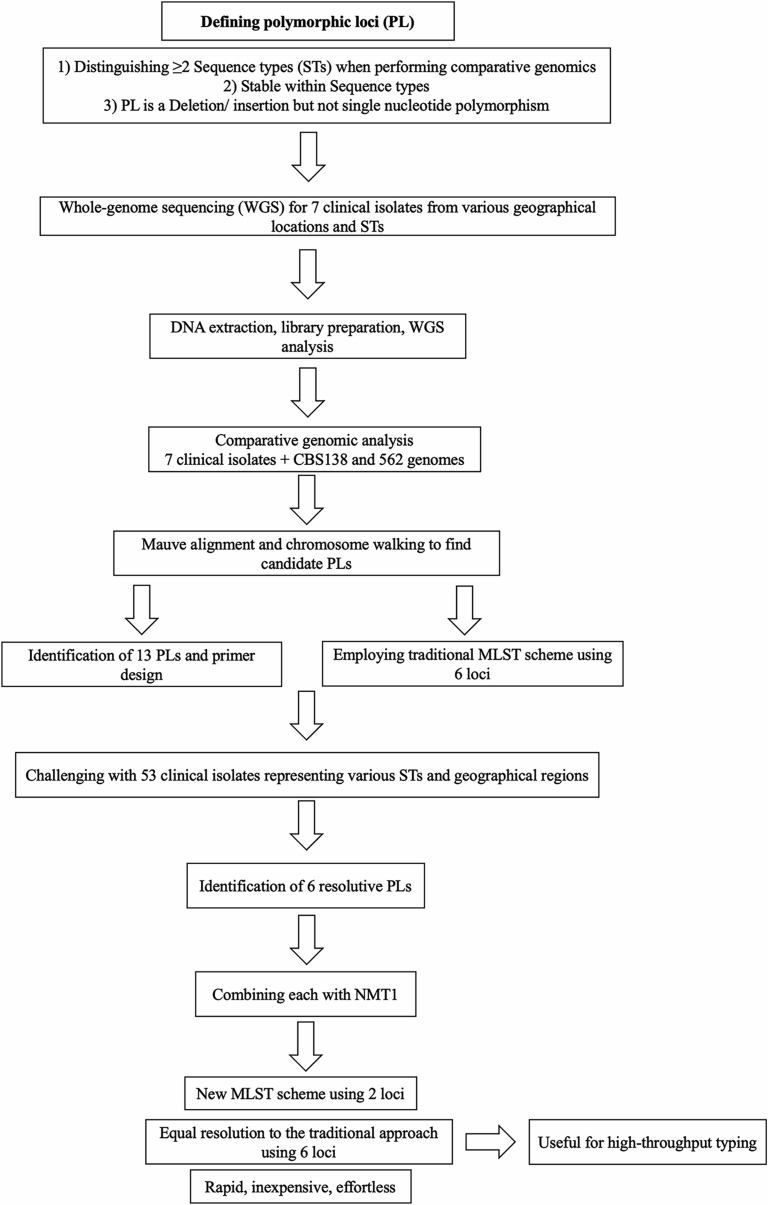
The workflow of this study is shown in Fig. 1. Seven clinical isolates belonging to ST10 (28, 29, 35), 15 (30 and 33), and 16 (38 and 40) (Table 1), were processed for WGS by long-read (PacBio) and short-read (Illumina) methods. Their genomes were then assembled and compared to each other and to the reference strain CBS138/ATCC2001 (Xu et al. 2021). Consistent with expectations, this analysis showed that 387 to 1 063 SNPs separated strains of the same ST, whereas strains of different STs were 57 476 to 75 811 SNPs apart.
Fig. 1.
The workflow of finding hyperpolymorphic loci (HPL) in this study.
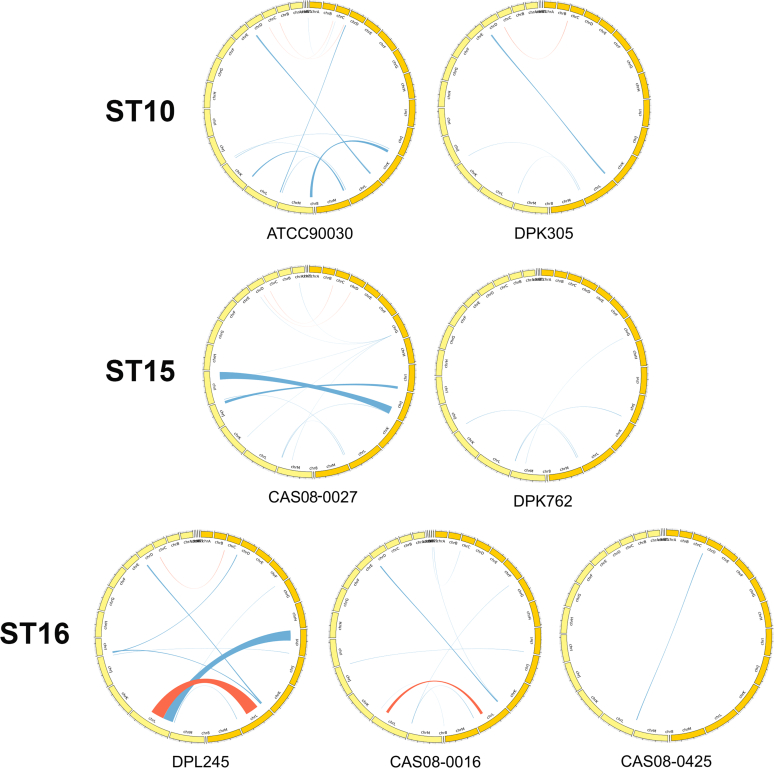
We also analysed the genomes for large-scale chromosomal variations known to frequently characterize C. glabrata strains (Carreté et al. 2018) (Fig. 2). Figure 2 shows the comparison between the seven newly sequenced genomes of C. glabrata and the CBS138 reference genome. Inversions, in red, were not very common. The three main inversions observed were a small inversion in chromosome C present in four of the seven strains (ATCC90030 (#35), DPK305 (#28), DPL245 (#38) and CAS08-0027 (#30)), a large (> 600 Kb) inversion in chromosome L in DPL245 (#38), and a smaller inversion in chromosome L in CAS08-0016 (#40) just adjacent to previous one. None of the inversions were specific to a given ST. For instance, DPL245 (#38) and CAS08-0016 (#40) belong to the same ST and they both have an inversion in chromosome L, but these inversions do not affect the same region. The inversion in chromosome C is shared by two isolates of clade ST10 (ATCC90030 (#35) and DPK305 (#28)), but not present in the third one (CAS08-0425 (#29)). However, this inversion is also present in DPK762 (#33) (ST15) but not in the other ST15 strains (CAS08-0027 (#30) and the reference strain CBS138). Translocations are more common, though they predominantly affect telomeric regions and therefore could be the result of either miss-assemblies or miss-alignments. Still, some of the translocations are consistent across isolates and had already previously been described (Carreté et al. 2018). For instance, in four of the seven strains we see a translocation from chromosome L to chromosome D. This translocation is found in two of the three isolates of ST10 (ATCC90030 (35) and DPK305 (28) and in the two strains of ST16 (DPL245 (38) and CAS08-0016 (40)). A second translocation moved a piece of chromosome D to chromosome L, affecting two of the three strains of ST10 (ATCC90030 (35) and CAS08-0425 (29)). Three additional translocations affected DPL245 (38): pieces of chromosome D and chromosome L moved to chromosome I and a large part of chromosome I was found attached to chromosome L. Finally, pieces of chromosomes I and J were interchanged in the genome of CAS08-0027 (30). This translocation was not observed previously.
Fig. 2.
Circos plots representing inversions (red) and translocations (blue) between the genomes of the C. glabrata reference strain (dark yellow on the right of each circle) and each individual strain (light yellow on the left of each circle). Chromosomes are ordered as mirror images of each other, starting with chromosome A at the top and ending with chromosome M at the bottom of the image.
Although changes in genome structure were not consistent with ST assignments, ST16 is the one most distantly related to the reference (CBS 138), both in terms of chromosome structure and DNA sequence (Fig. 2).
HPL selection and GEO
To identify discriminative HPL in C. glabrata that may be resolutive for ST, we aligned individual chromosomes from the seven newly assembled genomes with those from reference strain CBS138 and strain DSY562, for which high quality WGS is available (Vale-Silva et al. 2017, Xu et al. 2021). Highly polymorphic loci (HPL) were defined based on the presence of indels that were different between strains of different STs. To avoid overlap with previously published sets of loci, HPL associated with satellite regions, megasatellites, minisatellites, and microsatellites, were excluded (Thierry et al. 2008). In total 33 HPL were identified (Table 3), which were subjected to GO analysis using FungiFun (Priebe et al. 2015). Since the functions of most of the genes in C. glabrata are not characterized, their orthologs in Saccharomyces cerevisiae (https://www.yeastgenome.org) were used for GO analysis. These loci were found to be significantly enriched for stress response pathways, particularly those responding to cell surface (cell wall and plasma membrane) stress (Fig. 3), with 14 out of 33 loci containing genes involved in these processes. Our subsequent analyses focused on this subset of loci, with the exception of UBI4, which showed variation even among strains of the same ST and was therefore omitted, leaving 13 loci (Table 3).
Table 3.
Polymorphic loci of interest identified through comparative whole-genome sequencing.
| Polymorphic loci | Standard names | Systematic names |
|---|---|---|
| CHS5/ CAGL0G00814g | CHS5 | YLR330W |
| G00715/ CAGL0G00858g1 | IEA with MID2 | YLR332W |
| G00825/ CAGL0G00968g1 | VRP1 | YLR337C |
| SRP40/ CAGL0G02409g | SRP40 | YKR092C |
| G03839/ CAGL0G03993g1 | N/A | Not available |
| G06281/ CAGL0G06446g1 | N/A | Not available |
| G07117/ CAGL0G07293g1 | USA1 | YML029W |
| SSR1/ CAGL0H06413g1 | CCW14 | YLR390W-A |
| H09053/ CAGL0H09152g1 | PTK1 | YKL198C |
| PIR2/ CAGL0I06182g | SHP150 | YJL159W |
| I07645/ CAGL0I07821g1 | DUF1 | YOL087C |
| K01793/ CAGL0K01947g | YER137C | YER137C |
| K06501/ CAGL0K06655g | NGR1 | YBR212W |
| K09845/ CAGL0K10010g | PRP8 | YDR083W |
| PIR3/ CAGL0M08492g1 | PIR1 | YKL164C |
| M09053/ CAGL0M09086g | BUD4 | YJR092W |
| M09075/ CAGL0M09108g | JSN1 | YJR091C |
| GVI51_A02255/ CAGL0A02486g1 | SEB2 | YDR351W |
| A00825/ CAGL0A01001g1 | YLR326W | YLR326W |
| A02431/ CAGL0A02651g | EAF1 | YDR359C |
| C01617/ CAGL0C01859g | RER1 | YCL001W |
| ARB1/ CAGL0C02343g | ARB1 | YER036C |
| C02321/ CAGL0C02541g | BDF1 | YLR399C |
| POP2/ CAGL0C03399g | POP2 | YNR052C |
| ABP1/ CAGL0C03597g | ABP1 | YCR088W |
| D02871/ CAGL0D02926g | BRE2 | YLR015W |
| D05049/ CAGL0D05082g | UBI4 | YLL039C |
| E00341/ CAGL0E00561g | TUP1 | YCR084C |
| SWI5/ CAGL0E01331g | SWI5 | YDR146C |
| STP8/ CAGL0F01837g | SPT8 | YLR055C |
| SLG11 | SLG1 | YOR008C |
| HKR1/ CGAL0F03003g1 | HKR1 | YDR420W |
| F03333/ CAGL0F03641g | YML018C | YML018C |
These were the 13 polymorphic loci that were selected for MLST analysis in this study.
Fig. 3.
The gene ontology enrichment to select the polymorphic loci (PL) of interest among a pool of candidate loci. Eleven loci involved in cellular integrity and stress response pathways and two loci lacking orthologs in Saccharomyces cerevisiae were selected for further analysis.
Comparison of concatenated original MLST and HPL trees
To identify the most resolutive HPL, we sequenced the 13 HPL (Table 3) as well as the six loci used in the traditional MLST scheme in 53 additional C. glabrata clinical isolates belonging to 13 STs (Table 1). We built a tree based on the concatenation of the six traditional MLST genes and mapped ST information on it in different colors (Fig. 4A). The same was done with the 13 HPL (Fig. 4B). The trees were rooted at strain 57 as it is the only member of ST76. The two trees were very different in terms of the Robinson and Foulds calculation (RF), which is 0.72, meaning that 72 % of the nodes defined in the tree were different. These differences could be due to either the re-arrangements within STs, re-arrangements among STs, or strains moving between STs.
Fig. 4.
A. Tree resulting from the concatenation of six traditional MLST genes. Colours represent the ST group each strain belongs to. Bootstraps below 90 % are shown on the tree. B. Tree resulting from the concatenation of 13 hypervariable regions.
In order to compare the performance of the two means of generating phylogenetic trees (HPL vs traditional MLST loci), we computed the precision, recall and F1 for each ST and then calculated the average precision, recall and F1 for the whole tree. The MLST concatenated tree, which can serve as baseline, showed a precision of 1.0, a recall of 1.0 and a F1 of 1.0, whereas the concatenated of the HPL had a slightly lower precision 0.99, a recall of 0.1.0 and a F1 of 0.99. While the tree based on MLST data recovered all ST perfectly, the tree based on HPL only failed in retrieving ST03 as a monophyletic clade due to the presence of STX.
Comparison of single gene trees
We repeated the same comparison with trees built for each single gene in the analysis. As seen in Table 4, out of the six MLST genes, NMT1 was best able, on its own, to capture the variability of ST, though it did not perform as well as the concatenated assembly. Interestingly, two HPL were able to capture ST information better than NMT1: G00825 and SLG1. As seen in Fig. 5, NMT1 was unable to distinguish between ST10 and ST15, which is not surprising because the distinction between these two ST is based on the classification of LEU2 and URA3. It is also unable to separate STX from ST03, whose separation is again determined by a different gene (TRP1). In contrast, the two best HPL were able to separate ST10 from ST15 but could not separate STX from ST03 (Fig. 5).
Table 4.
Statistics for trees based on individual genes. MLST refers to the conventional MLST approach employing six loci to type C. glabrata and PLOI refers to the approach proposed in this study. The resolution of each approach and locus is evaluated by three factors, namely precision, recall, and F1. The higher the overall score, the more resolutive is the approach/locus.
| Loci | Approach | Precision | Recall | F1 |
|---|---|---|---|---|
| FKS1 | MLST | 0.96 | 0.87 | 0.88 |
| LEU2 | MLST | 0.87 | 0.75 | 0.74 |
| NMT1 | MLST | 0.93 | 0.93 | 0.93 |
| TRP1 | MLST | 0.96 | 0.81 | 0.83 |
| UGP1 | MLST | 0.79 | 0.85 | 0.77 |
| URA3 | MLST | 0.92 | 0.78 | 0.79 |
| A02255 | PLOI | 0.95 | 0.83 | 0.85 |
| DUF1 | PLOI | 0.88 | 0.87 | 0.84 |
| G00715 | PLOI | 0.83 | 0.91 | 0.86 |
| G00825 | PLOI | 0.99 | 1.0 | 0.99 |
| G03839 | PLOI | 0.92 | 0.82 | 0.82 |
| G06281 | PLOI | 0.92 | 0.81 | 0.82 |
| H09053 | PLOI | 0.92 | 0.81 | 0.83 |
| HKR1 | PLOI | 0.97 | 0.90 | 0.91 |
| PIR1 | PLOI | 0.90 | 0.86 | 0.85 |
| PIR2 | PLOI | 0.94 | 0.88 | 0.90 |
| SLG1 | PLOI | 0.99 | 1.0 | 0.99 |
| SSR1 | PLOI | 0.96 | 0.95 | 0.95 |
| USA1 | PLOI | 0.90 | 0.93 | 0.90 |
PLOI: Polymorphic loci of interest.
Fig. 5.
Single gene trees belonging to A. NMT1, B. G00825 and C. SLG1. Clades are colored according to the ST group. In NMT1 ST10 and ST15 are colored in the same color as they are identical in all strains.
Comparison of concatenated gene trees using combinations of loci
STs are currently assigned based on the allelic information provided by six genes. As single genes are not able to completely recover all ST groups as monophyletic clades, we tried to find a combination of six or fewer genes that could perform better than the concatenated MLST tree. To do that, we generated concatenated alignments of combinations of 2, 3, 4, 5 and 6 HPL and checked them for the distribution of ST. We then calculated the average precision, recall and F1 (Table 5). We found that none of the combinations performed as the concatenated MLST tree. However, 21 different combinations of two HPL genes were able to recover the same ST distribution as found by the concatenation of HPL, showing that few genes have as much resolutive power as the 13 selected HPL (Table 6 and Supplementary Table 1).
Table 5.
Summary statistics for each group of trees
| Statistics | Concatenated tree of MLST | Concatenated tree of PLOIs | 1 gene | 2 genes | 3 genes | 4 genes | 5 genes | 6 genes |
|---|---|---|---|---|---|---|---|---|
| Average Precision | - | - | 0.93 | 0.97 | 0.98 | 0.98 | 0.98 | 0.98 |
| Average Recall | - | - | 0.89 | 0.94 | 0.97 | 0.98 | 0.99 | 0.995 |
| Average F1 | - | - | 0.89 | 0.95 | 0.97 | 0.98 | 0.98 | 0.99 |
| Maximum F1 | 1.0 | 0.99 | 0.99 | 0.99 | 0.99 | 0.99 | 0.99 | 0.99 |
PLOI: Polymorphic loci of interest.
Table 6.
Best combinations of two polymorphic loci.
| Polymorphic loci combination | Alignment length | Precision |
|---|---|---|
| A02255;SLG1 | 1 549 | 0.99 |
| DUF1;G00825 | 2 120 | 0.99 |
| DUF1;SLG1 | 1 573 | 0.99 |
| G00825;G03839 | 2 280 | 0.99 |
| G00825;G06281 | 2 229 | 0.99 |
| G00825;HKR1 | 2 874 | 0.99 |
| G00825;SLG1 | 3 165 | 0.99 |
| G00825;SRR1 | 2 519 | 0.99 |
| G06281;SRR1 | 1 036 | 0.99 |
| H09053;HKR1 | 1 252 | 0.99 |
| H09053;SLG1 | 1 543 | 0.99 |
| HKR1;PIR2 | 2 035 | 0.99 |
| HKR1;SLG1 | 2 327 | 0.99 |
| HKR1;SRR1 | 1 681 | 0.99 |
| HKR1;USA1 | 1 558 | 0.99 |
| PIR2;SLG1 | 2 326 | 0.99 |
| PIR2;SRR1 | 1 680 | 0.99 |
| PIR2;USA1 | 1 557 | 0.99 |
| SLG1;SRR1 | 1 972 | 0.99 |
| SLG1;USA1 | 1 849 | 0.99 |
| SRR1;USA1 | 1 203 | 0.99 |
Finally, we tested whether combining MLST genes with HPL would produce better results. Two combinations of TRP1 with a single HPL (either SLG1 or G00825) resulted in a tree which was as good as the concatenated tree based on the six MLST loci (Fig. 5, Fig. 6).
Fig. 6.
Trees based on the concatenation of two genes. A.G00825 + TRP1. B.SLG1 + TRP1.
Conclusions
Understanding the genetic diversity of C. glabrata isolates has broad clinical and biological implications ranging from aiding effective implementation of infection control strategies in hospitals to understanding within-host microevolution, which can help answer key questions regarding host-pathogen interaction and the development of antifungal resistance. In this study we used WGS and comparative genomic analyses to identify a new MLST scheme based not only on SNPs but also indels, and, using only two loci, offers the same resolution as that provided by the widely used six-locus MLST scheme. Using 53 clinical C. glabrata isolates originating from various geographical regions and belonging to different STs, we showed that the two-locus scheme was in 100 % agreement with the traditional MLST approach. Also, we note that except for isolate CAS08-0293 for which the ST could not be distinguished with HPL, the ST of the rest of the isolates were successfully determined when testing only SLG1 or G00825. In summary, the newly described MLST scheme can be used as a reliable approach for high throughput typing purposes in clinical settings and large-scale studies aiming to understand the genetic diversity of C. glabrata globally. Additionally, the high-quality whole genomes of seven clinical strains reported in this study can serve as references for future short read based WGS analyses of C. glabrata, helping the field develop new diagnostic tools and address fundamental biological questions.
Data availability
The project has been deposited in GenBank under the Bioproject number PRJNA718446. The raw read data are deposited under accession number SRR8146337. Read data and genome assemblies can be found under SRA codes SRR14180810 to SRR14180816 for the Illumina reads and codes SRR14163270 to SRR14163276 for PacBio reads. MLST and PL sequences were deposited in GenBank under codes MW970414 to MW971421. Accession numbers associated with the isolates whole-genome sequenced were as follows, JAGTUA000000000 (CAS08-0425), JAGTTZ000000000 (CAS80027), JAGTTY000000000 (DPK762), JAGTTX000000000 (ATCC 90030), JAGTTW000000000 (DPL245), JAGTTV000000000 (CAS08-0016), JAGTTU000000000 (DPK 305).
Acknowledgements
We thank Dibyendu Kumar (Rutgers University) for help with C. glabrata PacBio sequencing. This work was supported by NIH 5R01AI109025 to D.S.P. TG group acknowledges support from the Spanish Ministry of Science and Innovation for grant PGC2018-099921-B-I00, cofounded by European Regional Development Fund (ERDF); from the Catalan Research Agency (AGAUR) SGR423; from the European Union's Horizon 2020 research and innovation programme (ERC-2016-724173); from the Gordon and Betty Moore Foundation (Grant GBMF9742) and from the Instituto de Salud Carlos III (INB Grant PT17/0009/0023 – ISCIII-SGEFI/ERDF).
Footnotes
Peer review under responsibility of Westerdijk Fungal Biodiversity Institute.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.simyco.2021.100133.
Contributor Information
E. Shor, Email: erika.shor@hmh-cdi.org.
T. Gabaldón, Email: toni.gabaldon.bcn@gmail.com.
Appendix A. Supplementary data
The following is the supplementary data to this article:
References
- Arastehfar A., Yazdanpanah S., Bakhtiari M., et al. Epidemiology of candidemia in Shiraz, southern Iran: A prospective multicenter study (2016-2018) Medical Mycology. 2020;59:422–430. doi: 10.1093/mmy/myaa059. [DOI] [PubMed] [Google Scholar]
- Arastehfar A., Lass-flörl C., Garcia-rubio R., et al. The quiet and underappreciated rise of drug-resistant invasive fungal pathogens. Journal of Fungi. 2020;6:138. doi: 10.3390/jof6030138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arastehfar A., Daneshnia F., Salehi M.R., et al. Low level of antifungal resistance of Candida glabrata blood isolates in Turkey: Fluconazole minimum inhibitory concentration and FKS mutations can predict therapeutic failure. Mycoses. 2020;63:911–920. doi: 10.1111/myc.13104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arastehfar A., Daneshnia F., Najafzadeh J., et al. Evaluation of molecular epidemiology, clinical characteristics, antifungal susceptibility profiles, and molecular mechanisms of antifungal resistance of Iranian Candida parapsilosis species complex blood isolates. Frontiers in Cellular and Infection Microbioliogy. 2020;10:206. doi: 10.3389/fcimb.2020.00206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arastehfar A., Daneshnia F., Zomorodian K., et al. Low level of antifungal resistance in Iranian isolates of Candida glabrata recovered from Blood samples in a multicenter study from 2015 to 2018 and potential prognostic values of genotyping and sequencing of PDR1. Antimicrobial Agents Chemotherapy. 2019;63:e02503–e02518. doi: 10.1128/AAC.02503-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Astvad K.M.T., Johansen H.K., Røder B.L., et al. Update from a 12-year nationwide fungemia surveillance: increasing intrinsic and acquired resistance causes concern. Journal of Clinical Microbiology. 2018;56:e01564–e01617. doi: 10.1128/JCM.01564-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biswas C., Marcelino V.R., Van Hal S., et al. Whole genome sequencing of australian Candida glabrata isolates reveals genetic diversity and novel sequence types. Frontiers in Microbiology. 2018;9:2946. doi: 10.3389/fmicb.2018.02946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boetzer M., Pirovano W. SSPACE-LongRead: scaffolding bacterial draft genomes using long read sequence information. BMC Bioinformatics. 2014;15:211. doi: 10.1186/1471-2105-15-211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bordallo-Cardona M.Á., Agnelli C., Gómez-Nuñez A., et al. MSH2 gene point mutations are not antifungal resistance markers in Candida glabrata. Antimicrobial Agents and Chemotherapy. 2019;63:e01876–e01918. doi: 10.1128/AAC.01876-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown G.D., Denning D.W., Gow N.A.R., et al. Hidden killers: Human fungal infections. Science Translational Medicine. 2012;4:165rv13. doi: 10.1126/scitranslmed.3004404. [DOI] [PubMed] [Google Scholar]
- Byrne K.P., Wolfe K.H. The yeast gene order browser: combining curated homology and syntenic context reveals gene fate in polyploid species. Genome Research. 2005;15:1456–1461. doi: 10.1101/gr.3672305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Byun S.A., Won E.J., Kim M.N., et al. Multilocus sequence typing (MLST) genotypes of Candida glabrata bloodstream isolates in Korea: association with antifungal resistance, mutations in mismatch repair gene (Msh2), and clinical outcomes. Frontiers in Microbiology. 2018;9:1523. doi: 10.3389/fmicb.2018.01523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carreté L., Ksiezopolska E., Gómez-Molero E., et al. Genome comparisons of Candida glabrata serial clinical isolates reveal patterns of genetic variation in infecting clonal populations. Frontiers in Microbiology. 2019;10:112. doi: 10.3389/fmicb.2019.00112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carreté L., Ksiezopolska E., Pegueroles C., et al. Patterns of genomic variation in the opportunistic pathogen Candida glabrata suggest the existence of mating and a secondary association with humans. Current Biology. 2018;28:15–27.e7. doi: 10.1016/j.cub.2017.11.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chakrabarti A., Sood P., Rudramurthy S.M., et al. Incidence, characteristics and outcome of ICU-acquired candidemia in India. Intensive Care Medicine. 2015;41:285–295. doi: 10.1007/s00134-014-3603-2. [DOI] [PubMed] [Google Scholar]
- Chapman B., Slavin M., Marriott D., et al. Changing epidemiology of candidaemia in Australia. Journal Antimicrobial Chemotherapy. 2017;72:1103–1108. doi: 10.1093/jac/dkw422. [DOI] [PubMed] [Google Scholar]
- Clinical and Laboratory Standards Institute . CLSI; Wayne, PA: 2017. Performance standards for antifungal susceptibility testing of yeasts. Approved standard M60. [Google Scholar]
- DePristo M.A., Banks E., Poplin R., et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genetics. 2011;43:491–498. doi: 10.1038/ng.806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dodgson A.R., Pujol C., Denning D.W., et al. Multilocus sequence typing of Candida glabrata reveals geographically enriched clades. Journal of Clinical Microbiology. 2003;41:5709–5717. doi: 10.1128/JCM.41.12.5709-5717.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- English A.C., Richards S., Han Y., et al. Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS One. 2012;7 doi: 10.1371/journal.pone.0047768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuller J., Dingle T.C., Bull A., et al. Species distribution and antifungal susceptibility of invasive Candida isolates from Canadian hospitals: results of the CANWARD 2011-16 study. Journal of Antimicrobial Chemotherapy. 2019;74:iv48–iv54. doi: 10.1093/jac/dkz287. [DOI] [PubMed] [Google Scholar]
- Gabaldón T. Recent trends in molecular diagnostics of yeast infections: from PCR to NGS. FEMS Microbiology Reviews. 2019;43:517–547. doi: 10.1093/femsre/fuz015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gabaldón T., Fairhead C. Genomes shed light on the secret life of Candida glabrata: not so asexual, not so commensal. Current Genetics. 2019;65:93–98. doi: 10.1007/s00294-018-0867-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gabaldón T., Gómez-Molero E., Bader O. Molecular typing of Candida glabrata. Mycopathologia. 2020;185:755–764. doi: 10.1007/s11046-019-00388-x. [DOI] [PubMed] [Google Scholar]
- Gremme G., Brendel V., Sparks M.E.S. Engineering a software tool for gene structure prediction in higher organisms. Information and Software Technology. 2005;47:965–978. [Google Scholar]
- Haas B.J., Salzberg S.L., Zhu W., et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biology. 2008;9:R7. doi: 10.1186/gb-2008-9-1-r7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Healey K.R., Perlin D.S. Fungal resistance to cchinocandins and the MDR phenomenon in Candida glabrata. Journal of Fungi. 2018;4:105. doi: 10.3390/jof4030105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holt C., Yandell M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics. 2011;12:491. doi: 10.1186/1471-2105-12-491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hou X., Xiao M., Chen S.C.A., et al. Molecular epidemiology and antifungal susceptibility of Candida glabrata in China (August 2009 to July 2014): a multi-center study. Frontiers in Microbiology. 2017;8:880. doi: 10.3389/fmicb.2017.00880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hou X., Xiao M., Wang H., et al. Profiling of PDR1 and MSH2 in Candida glabrata bloodstream isolates from a multicenter study in China. Antimicrobial Agents Chemotherapy. 2018;62:e00153–e00218. doi: 10.1128/AAC.00153-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huerta-Cepas J., Dopazo J., Gabaldón T. ETE: a python environment for tree exploration. BMC Bioinformatics. 2010;11:24. doi: 10.1186/1471-2105-11-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katiyar S., Edlind T. New locus for Candida glabrata sequence-based strain typing provides evidence for nosocomial transmission. Journal of Clinical Microbiology. 2021;59:e02933–e03020. doi: 10.1128/JCM.02933-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katiyar S., Shiffrin E., Shelton C., et al. Evaluation of polymorphic locus sequence typing for Candida glabrata epidemiology. Journal of Clinical Microbiology. 2016;54:1042–1050. doi: 10.1128/JCM.03106-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khalifa H.O., Arai T., Majima H., et al. Genetic basis of azole and echinocandin resistance in clinical Candida glabrata in Japan. Antimicrobial Agents and Chemotherapy. 2020;64:e00783–e00820. doi: 10.1128/AAC.00783-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kord M., Salehi M., Khodavaisy S., et al. Epidemiology of yeast species causing bloodstream infection in Tehran, Iran (2015 – 2017); superiority of 21-plex PCR over the Vitek 2 system for yeast identification. Journal of Medical Microbiology. 2020;69:712–720. doi: 10.1099/jmm.0.001189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koren S., Walenz B.P., Berlin K., et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Research. 2017;27:722–736. doi: 10.1101/gr.215087.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ksiezopolska E., Gabaldón T. Evolutionary emergence of drug resistance in Candida opportunistic pathogens. Genes. 2018;9:461. doi: 10.3390/genes9090461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamoth F., Lockhart S.R., Berkow E.L., et al. Changes in the epidemiological landscape of invasive candidiasis. The Journal of Antimicrobial Chemotherapy. 2018;73:i4–i13. doi: 10.1093/jac/dkx444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H., Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin C.Y., Chen Y.C., Lo H.J., et al. Assessment of Candida glabrata strain relatedness by pulsed-field gel electrophoresis and multilocus sequence typing. Journal of Clinical Microbiology. 2007;45:2452–2459. doi: 10.1128/JCM.00699-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lott T.J., Frade J.P., Lockhart S.R. Multilocus sequence type analysis reveals both clonality and recombination in populations of Candida glabrata bloodstream isolates from U.S. surveillance studies. Eukaryotic Cell. 2010;9:619–625. doi: 10.1128/EC.00002-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lott T.J., Frade J.P., Lyon G.M., et al. Bloodstream and non-invasive isolates of Candida glabrata have similar population structures and fluconazole susceptibilities. Medical Mycology. 2012;50:136–142. doi: 10.3109/13693786.2011.592153. [DOI] [PubMed] [Google Scholar]
- Megri Y., Arastehfar A., Boekhout T., et al. Candida tropicalis is the most prevalent yeast species causing candidemia in Algeria: the urgent need for antifungal stewardship and infection control measures. Antimicrobial Resistance and Infection Control. 2020;9:50. doi: 10.1186/s13756-020-00710-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otto T.D., Dillon G.P., Degrave W.S., et al. RATT: Rapid annotation transfer tool. Nucleic Acids Research. 2011;39:e57. doi: 10.1093/nar/gkq1268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pappas P.G., Kauffman C.A., Andes D.R., et al. Clinical practice guideline for the management of candidiasis : 2016 update by the Infectious Diseases Society of America. Clinical Infectious Diseases. 2016;62:1–50. doi: 10.1093/cid/civ933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfaller M.A., Diekema D.J. Progress in antifungal susceptibility testing of Candida spp. by use of Clinical and Laboratory Standards Institute broth microdilution methods, 2010 to 2012. Journal of Clinical Microbiology. 2012;50:2846–2856. doi: 10.1128/JCM.00937-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfaller M.A., Diekema D.J., Turnidge J.D., et al. Twenty years of the SENTRY antifungal surveillance program: results for Candida species from 1997-2016. Open Forum Infectious Diseases. 2019;6:S79–S94. doi: 10.1093/ofid/ofy358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfaller M.A., Messer S.A., Hollis R.J., et al. Variation in susceptibility of bloodstream isolates of Candida glabrata to fluconazole according to patient age and geographic location in the United States in 2001 to 2007. Journal of Clinical Microbiology. 2009;47:3185–3190. doi: 10.1128/JCM.00946-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Priebe S., Kreisel C., Horn F., et al. FungiFun2: a comprehensive online resource for systematic analysis of gene lists from fungal species. Bioinformatics. 2015;31:445–446. doi: 10.1093/bioinformatics/btu627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Proux-Wéra E., Armisén D., Byrne K.P., et al. A pipeline for automated annotation of yeast genome sequences by a conserved-synteny approach. BMC Bioinformatics. 2012;13:237. doi: 10.1186/1471-2105-13-237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Romo J.A., Kumamoto C.A. On commensalism of Candida. Journal of Fungi. 2020;6:16. doi: 10.3390/jof6010016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slater G.S.C., Birney E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics. 2005;6:31. doi: 10.1186/1471-2105-6-31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song Y., Chen X., Yan Y., et al. Prevalence and antifungal susceptibility of pathogenic yeasts in China: a 10-year retrospective study in a teaching hospital. Frontiers in Microbiology. 2020;11:1401. doi: 10.3389/fmicb.2020.01401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stavrou A.A., Lackner M., Lass-Flörl C., Boekhout T. The changing spectrum of Saccharomycotina yeasts causing candidemia: phylogeny mirrors antifungal susceptibility patterns for azole drugs and amphothericin B. FEMS Yeast Research. 2019;19:4. doi: 10.1093/femsyr/foz037. [DOI] [PubMed] [Google Scholar]
- Taj-Aldeen S.J., Kolecka A., Boesten R., et al. Epidemiology of candidemia in Qatar, the Middle East: performance of MALDI-TOF MS for the identification of Candida species, species distribution, outcome, and susceptibility pattern. Infection. 2014;42:393–404. doi: 10.1007/s15010-013-0570-4. [DOI] [PubMed] [Google Scholar]
- Thierry A., Bouchier C., Dujon B., et al. Megasatellites: a peculiar class of giant minisatellites in genes involved in cell adhesion and pathogenicity in Candida glabrata. Nucleic Acids Research. 2008;36:5970–5982. doi: 10.1093/nar/gkn594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsay S.V., Mu Y., Williams S., et al. Burden of candidemia in the United States, 2017. Clinical Infectious Diseases. 2020;71:e449–e453. doi: 10.1093/cid/ciaa193. [DOI] [PubMed] [Google Scholar]
- Vale-Silva L., Beaudoing E., Tran V.D.T., et al. Comparative genomics of two sequential Candida glabrata clinical isolates. G3. 2017;7:2413–2426. doi: 10.1534/g3.117.042887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Won E.J., Choi M.J., Kim M.N., et al. Fluconazole-resistant Candida glabrata bloodstream isolates, South Korea, 2008-2018. Emerging Infectious Diseases. 2021;27:779–788. doi: 10.3201/eid2703.203482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Z., Green B., Benoit N., et al. Cell wall protein variation, break induced replication, and subtelomere dynamics in Candida glabrata. Molecular Microbiology. 2021 doi: 10.1111/mmi.14707. Online ahead of print. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The project has been deposited in GenBank under the Bioproject number PRJNA718446. The raw read data are deposited under accession number SRR8146337. Read data and genome assemblies can be found under SRA codes SRR14180810 to SRR14180816 for the Illumina reads and codes SRR14163270 to SRR14163276 for PacBio reads. MLST and PL sequences were deposited in GenBank under codes MW970414 to MW971421. Accession numbers associated with the isolates whole-genome sequenced were as follows, JAGTUA000000000 (CAS08-0425), JAGTTZ000000000 (CAS80027), JAGTTY000000000 (DPK762), JAGTTX000000000 (ATCC 90030), JAGTTW000000000 (DPL245), JAGTTV000000000 (CAS08-0016), JAGTTU000000000 (DPK 305).