Abstract
Colorectal cancer (CRC) is one of the most heritable cancers, and genetic factors play an important role in the increased CRC risk. However, the well-established CRC-risk genes were limited for explaining the increased risk of CRC individuals. Germline mutations in DNA damage repair (DDR) genes have also been reported to be implicated in CRC heritability. Here, we aimed to determine the prevalence and significance of germline DDR and well-established CRC-risk gene variants in CRCs with paired somatic analyses. Next-generation sequencing (NGS) was performed on tumor tissues and paired white blood cells collected from 2160 Chinese patients with CRC using well-designed 381- or 733-cancer gene panel. Germline/somatic variations were identified and assessed for pathogenicity and likely pathogenicity. Of 2160 CRCs, 136 pathogenic germline mutations in 133 patients (133/2160, 6.1%) were identified in 21 genes, including 19 out of 32 examined DDR genes. Compared with non-carriers, individuals with germline variants were prone to a higher level of microsatellite instability (MSI) and tumor mutational burden (TMB), and an earlier age of onset. Somatic sequencing identified second hits in 24/133 (18%) patients with germline variants. Among the mismatch repair (MMR) genes with germline mutations, the second hit significantly increased MSI and TMB, particularly apparent in MSH6. All MMR germline variation carriers further with a second hit were all MSI-H and had an extraordinarily high level of TMB. Collectively, approximately 6.1% of CRC patients carried pathogenic germline variants, and additional somatic second hit increases the genomic instability in CRC, whereas the more clinical significance warrants further study.
Keywords: Colorectal cancer, DNA damage repair, germline mutation, somatic second hit
Introduction
Owning to the implementation of routine screening and the advancement of precise treatments, the incidence and mortality of colorectal cancer (CRC) have declined globally. However, the incidence and mortality in the subset of CRC individuals with age < 50 are increasing by approximately 2% and 1.3% annually, respectively [1]. Early onset is a sign of inherited cancer predisposition. CRC is one of the most heritable cancers, of which approximately 30% of CRC cases are attributed to inherited factors [2]. Nevertheless, among the inherited factors, the well-established CRC-risk genes, predominantly the mismatch repair (MMR) genes, APC and MUTYH, are limited for explaining the estimated heritability degree [3-5]. Interrogation of the genes with inherited risks is necessary to explain CRC heredity, and particularly essential for guide prevention and management for the patient and his family.
The DNA damage repair (DDR) is essential for the preservation of genomic integrity and stability, whose defection is considered to be the underlying mechanism behind the occurrence and development of various human malignant tumors, including CRC. Interrogation of the germline DDR gene mutation landscape is emerging in various types of human malignancies. Virtually, germline mutations in other DDR genes besides MMR genes have also been reported to be implicated in CRC heritability, such as BRCA, ATM, and PALB2 [6-8]. Whereas these findings are mainly based on data from Caucasian populations, the prevalence and spectrum of comprehensive DDR germline mutations in unselected Asian CRC patients remain largely unknown.
A ‘two-hit’ (i.e. the germline variation as the first hit, and somatic alteration in the wild-type allele of affected gene as the second hit) hypothesis for tumorigenesis is proposed by Knudson, in which germline variation and somatic variation are independent mutation events, and finally result in the function loss of a tumor suppressor gene [9]. However, for CRC patients with germline cancer susceptibility gene variants, second hits have not been rigorously assessed.
In this study, we determined the prevalence of germline variants and somatic second hits among 10 well-established CRC-risk genes and 32 DDR genes in 2160 CRC patients through next-generation sequencing (NGS), and assessed their correlations with tumor mutational burden (TMB), microsatellite instability (MSI), and onset age of CRC.
Materials and methods
Clinical tumor specimens
Patients were selected from the 3DMed Biobank according to the inclusion criteria: (i) histologically proven CRC by two independent pathologists via hematoxylin and eosin (H&E) staining; and (ii) available NGS data of formalin-fixed paraffin embedded (FFPE) tissues and paired white blood cells. Patients were excluded if they had a concurrent invasive malignant disease primary to other sites.
DNA sequencing and interpretation
The NGS sequencing data from 2160 patients were analyzed. A well-designed 381- or 733-cancer gene panel was utilized for NGS as previously described in 3DMed Clinical Laboratory Inc., a College of American Pathologists (CAP) and Clinical Laboratory Improvement Amendments (CLIA) certified laboratory [10]. Briefly, DNA were extracted and quantified using the ReliaPrep™ FFPE gDNA Miniprep System (Promega) and the Qubit™ dsDNA HS Assay Kit (Thermo Fisher Scientific), respectively. After shearing to 250 bp fragments, KAPA Hyper Prep Kit (KAPA Biosystems) was used to construct the libraries. Indexed libraries were subjected to probe-based hybridization with a customized NGS panel targeting 381 or 733 cancer-related genes (3D Medicines). The captured libraries were sequenced on a NovaSeq 6000 platform (Illumina) in 100 bp paired-end mode with a minimum coverage of 500 X. Raw data were mapped to the reference human genome hg19 using the Burrows-Wheeler Aligner (v0.7.12). PCR duplicate reads were removed using Picard (v1.130) and sequence metrics were collected using SAMtools (v1.1.19). Only the targeted regions were searched for variant calling. Single nucleotide variants were recognized using an in-house developed R package to execute a variant detection model based on binomial test, while indels were detected by local realignment. Then variants were filtered by their unique supporting read depth, strand bias, and base quality, followed by continually narrowed down using an automated false positive filtering pipeline to ensure sensitivity and specificity at an allele frequency of ≥ 5%. Single-nucleotide polymorphism and indels were annotated by ANNOVAR against the dbSNP (v138), 1000Genome and ESP6500 databases (population frequency >0.015). Only stop-gain, missense, frameshift and non-frameshift indel mutations were kept. Copy number variations and gene rearrangements were detected as described previously [10]. Identification and analysis of germline alternations were restricted to 10 well-established CRC-risk genes, as well as 32 DDR genes (Table 1). Germline variants were identified using white blood cells; deleterious or suspected deleterious alterations were ascertained by the bioinformatics specialist as per a joint consensus of the latest literature or reports from clinical trials and the recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMP-AMP), which collectively defined as pathogenic mutations [11]. Analyses were also performed to identify the somatic alteration. Patients with deleterious germline variants were regarded to have a second hit if a pathogenic somatic deleterious alteration in the same gene was detected (e.g., missense, stop-gain, frameshift and non-frameshift indel mutations, copy number loss, or gene rearrangements).
Table 1.
The gene list of identified germline alternations
| 10 CRC-risk genes | APC MLH1 MSH2 MSH6 PMS2 MUTYH PTEN TP53 STK11 SMAD |
| 32 DDR genes | MRE11A RAD50 MLH1 MSH2 MSH6 PMS2 POLD1 ATM ATR BARD1 BLM BRCA1 BRCA2 BRIP1 CHEK1 CHEK2 FANCA FANCC FANCD2 FANCG MUTYH PALB2 POLE TP53 PRKDC CUL3 ERCC1 FANCE FANCF FANCL RAD51 WEE1 |
TMB
TMB was defined as the number of somatic single nucleotide variants (SNVs) and indels in examined coding regions, with driver mutations excluded. All SNVs and indels in the coding region of targeted genes, including missense, silent, stop gain, stop loss, in-frame and frameshift mutations, were considered.
MSI
After NGS testing of tumor DNA in FFPE samples, the identification of MSI-high (MSI-H) or microsatellite-stable (MSS) was performed via detecting the status of 100 microsatellite loci as previously published [12]. For each sample, the percentage of MSI loci was calculated, and a percentage of above 0.4 was considered as MSI-H and otherwise MSS.
Statistical analysis
Data were presented as median values (range minimum-maximum) or as numbers (%). All analyses were performed on R version 3.4.4 (The R Foundation of Statistical Computating) by using the t test or the non-parametric Mann-Whitney U test for continuous variables, and chi-square test or fisher’s exact test for categorical variables. All comparisons were two-sided, and a P value < 0.05 was considered statistically significant.
Results
Patient characteristics
Demographics and baseline characteristics of CRC patients were summarized in Table 2. This cohort was constituted by 60.2% male and 39.8% female. For patients with age at diagnosis documented, 95 individuals (7.4%) were at or under age of 35; 308 (24.0%) cases were aged between 35 and 50; and 883 (68.7%) patients were aged over 50 according to the classification criteria of previous articles [13-15]. Among all tumor tissues, 1536 (71.1%) were from surgery, and 624 (28.9%) were obtained by biopsy. The majority (80.1%) were primary tumors. NGS data showed that in 2160 CRC patients, 141 (6.5%) cases were MSI-H, and the remaining 2019 (93.5%) were MSS while the median TMB in the overall population was 7 mutations per MB ranging 0 to 600 mutations per MB.
Table 2.
Patient demographics and baseline characteristics
| Characteristic | All patients (N=2160) | Carrier (N=133) | Non-carrier (N=2027) | P value | |
|---|---|---|---|---|---|
| Age at diagnosis | ≤ 35 | 95 (7.4%) | 13 (14.8%) | 82 (6.8%) | < 0.001 |
| 35-50 | 308 (24.0%) | 22 (25.0%) | 286 (23.9%) | ||
| >50 | 883 (68.7%) | 53 (60.2%) | 830 (69.3%) | ||
| NA | 874 | 45 | 829 | ||
| Sex | Male | 1301 (60.2%) | 80 (60.1%) | 1221 (60.2%) | 0.984 |
| Female | 859 (39.8%) | 53 (39.9%) | 806 (39.8%) | ||
| Sampling type | Resection | 1536 (71.1%) | 1432 (70.6%) | 104 (78.2%) | 0.628 |
| Biopsy | 624 (28.9%) | 595 (29.4%) | 119 (21.8%) | ||
| Sample lesion | Primary | 1731(80.1%) | 106 (79.7%) | 1625 (80.2%) | 0.896 |
| Metastatic | 430 (19.9%) | 27 (20.30%) | 402 (19.8%) | ||
| MSI status | MSI-H | 141 (6.5%) | 44 (33.1%) | 97 (4.8%) | < 0.001 |
| MSS | 2019 (93.5%) | 89 (66.9%) | 1930 (95.2%) | ||
| TMB, median (min, max) | 7 (0,600) | 10 (0,400) | 7 (2,600) | < 0.001 |
Abbreviations: MSI-H, high-level microsatellite instability; MSS, stable-level microsatellite instability; TMB, tumor mutation burden; Carrier, patients with pathogenic germline variants; Non-carrier, patients without pathogenic germline variants; NA, not available.
Germline and somatic genetic feature of CRC patients
NGS was performed to interrogate the germline mutation landscape of CRC patients, with 10 well-established CRC-risk genes and 32 DDR genes being inspected. Among 2160 CRC patients, 136 pathogenic germline mutations in 133 patients (6.1%) were identified in 21 genes. These mutations fell predominantly in MLH1 (n=22, 16.2%), MSH2 (n=14, 10.3%), MSH6 (n=11, 8.1%), MUTYH (n=10, 7.4%), APC (n=10, 7.4%), BRCA (9 in BRCA2, and 1 in BRCA1, n=10, 7.4%), ATM (n=8, 5.9%), BLM (n=7, 5.1%), RAD50 (n=7, 5.1%), and CHEK2 (n=7, 5.1%) (Figure 1A and 1B). Among the 133 carriers with a median TMB of 10 mutations per Mb, 44 (33.1%) were MSI-H. Meanwhile, in the 2027 non-carriers with a median TMB of 7 mutations per Mb, only 97 (4.8%) were MSI-H (Table 2). The results of evaluation of a second hit showed that in 133 cases with pathogenic germline variants, 24 had evidence of a second hit, including 13 in MMR genes (Table 3).
Figure 1.
The spectrum of pathogenic germline mutations and somatic second hits in our cohort. A. The identified pathogenic germline mutations and somatic second hits among 133 patients with colorectal cancer; B. Distribution of 136 pathogenic germline mutations that occurred in 21 genes; C. Location of pathogenic germline mutations (black) and somatic second hits (red) are shown by lollipop plots.
Table 3.
Clinical information for patients with somatic second hit mutation
| Patient No. | Sex | Age | Gene | Pathogenic germline mutation | Somatic second hits | MSI status | TMB |
|---|---|---|---|---|---|---|---|
| 1 | Male | 48 | MLH1 | c.2059C>T | c.298C>T | MSI-H | 55 |
| 2 | Male | 58 | MLH1 | c.1558+1G>A | c.829G>T | MSI-H | 110 |
| 3 | Female | 24 | MSH2 | c.387_388del | c.G1661A | MSI-H | 156 |
| 4 | Male | 31 | MSH6 | c.3056_3057del | c.2892T>A | MSI-H | 108 |
| 5 | Male | 48 | MSH6 | c.3755dup | c.3312dup | MSI-H | 600 |
| c.1188_1189del | |||||||
| 6 | Male | NA | MSH6 | c.3226C>T | c.3261dup | MSI-H | 94 |
| 7 | Male | NA | MSH6 | c.425G>A | c.1912delinsTT | MSI-H | 364 |
| 8 | Female | 54 | MSH6 | c.505_506dup | c.3226C>T | MSI-H | 83 |
| c.3991C>T | |||||||
| 9 | Male | 33 | MSH6 | c.1483C>T | c.3261dup | MSI-H | 164 |
| 10 | Male | 65 | MSH6 | c.1033_1034del | c.3261dup | MSI-H | 116 |
| 11 | Female | 34 | MSH6 | c.1347_1372dup | c.399_402del | MSI-H | 77 |
| 12 | Male | 26 | PMS2 | c.451dup | c.861_864del | MSI-H | 73 |
| 13 | Female | 33 | PMS2 | c.1738A>T | c.3G>A | MSI-H | 76 |
| 14 | Male | NA | TP53 | c.524G>A | c.371dup | MSS | 14 |
| 15 | Male | 82 | TP53 | c.743G>A | c.524G>A | MSS | 12 |
| 16 | Female | NA | APC | c.4391_4394del | c.1504G>T | MSS | 10 |
| 17 | Male | NA | APC | c.3927_3931del | c.4393_4394del | MSS | 7 |
| 18 | Male | 33 | APC | c.4133_4134del | c.2626C>T | MSS | 7 |
| 19 | Male | 74 | APC | c.6245del | c.1411G>T | MSS | 8 |
| c.3902_3912del | |||||||
| 20 | Female | 30 | APC | c.2677G>T | c.4285C>T | MSS | 6 |
| 21 | Male | 66 | ATM | c.6910del | c.742C>T | MSS | 3 |
| 22 | Male | NA | ATM | c.8641C>T | c.5692C>T | MSS | 5 |
| 23 | Male | 46 | ATM | c.7141_7151del | Loss of copy number | MSS | 4 |
| 24 | Female | 48 | BRCA2 | c.8400_8402delinsAAAA | c.956dup | MSS | 14 |
Abbreviations: TMB, tumor mutation burden; MSI status, microsatellite instability status; NA, not available.
The pathogenic germline mutation frequencies of patients at or under age of 35, between 35 and 50, and older than 50 were 13.7%, 7.1%, and 6.0%, respectively (Table 2). Early-onset CRCs with age ≤ 50 had a trend of higher germline mutation frequency than average-onset CRC patients (9.6% vs 6.0%, P=0.094), especially in MMR, APC and CHEK2 (Table 4). As compared with average-onset CRC patients (>50 years old), early-onset CRCs had more cases with MSI-H (13.6% vs 6.7%, P < 0.001).
Table 4.
Prevalence of germline pathogenic mutations in CRC patients with age ≤ 50 or age >50
| ≤ 50 (N=403) | >50 (N=883) | |
|---|---|---|
| APC | 2 (0.5%) | 4 (0.5%) |
| ATM | 1 (0.2%) | 4 (0.5%) |
| ATR | 1 (0.2%) | 3 (0.3%) |
| BRCA1 | 1 (0.2%) | 0 (0%) |
| BRCA2 | 1 (0.2%) | 6 (0.7%) |
| CHEK2 | 4 (1%) | 3 (0.3%) |
| FANCD2 | 1 (0.2%) | 1 (0.1%) |
| MLH1 | 15 (3.7%) | 3 (0.3%) |
| MSH2 | 4 (1.0%) | 6 (0.7%) |
| MSH6 | 4 (3.7%) | 3 (0.3%) |
| PMS2 | 3 (0.7%) | 0 (0%) |
| BARD1 | 0 (0%) | 1 (0.1%) |
| BLM | 0 (0%) | 1 (0.1%) |
| BRIP1 | 0 (0%) | 1 (0.1%) |
| FANCA | 0 (0%) | 4 (0.5%) |
| MUTYH | 0 (0%) | 6 (0.7%) |
| PALB2 | 0 (0%) | 2 (0.2%) |
| RAD50 | 0 (0%) | 3 (0.3%) |
| TP53 | 0 (0%) | 3 (0.3%) |
Notes: only patients with surgical specimens of primary tumors were included.
Of 2160 CRC patients, 123 (5.7%) participants carried at least one pathogenic germline variants in the DDR genes, 19 of whom had a second hit (Figure 1A). Patients with pathogenic germline DDR gene mutations were more likely to present higher TMB level (median TMB 11.3 vs 7.26, P < 0.001), MSI-H phenotype (35.8% vs 4.8%, P < 0.001) and an earlier onset age of CRC (median age 54.0 vs 57.0, P=0.016) compared to the non-carriers (Figure 2A and 2B). When a second hit occurred in DDR genes, TMB (median TMB 77.4 vs 10.5, P=0.002) and MSI-H incidence (68.4% vs 29.8%, P=0.003) tended to be higher but there was no significant difference in the age of onset of CRC, indicating that somatic mutation increased the instability of genome, but was not associated with the onset age of CRC (Figure 2C and 2D).
Figure 2.
Effects of germline DNA damage repair (DDR) gene mutations and somatic second hits on TMB and age at diagnosis. (A, B) The effects of germline DDR gene mutations on TMB (A) and age at diagnosis (B); (C, D) The effects of DDR somatic second hits on TMB (A) and age at diagnosis (B). *P < 0.05, **P < 0.01, ****P < 0.001.
MMR germline mutation and second hit in CRC patients
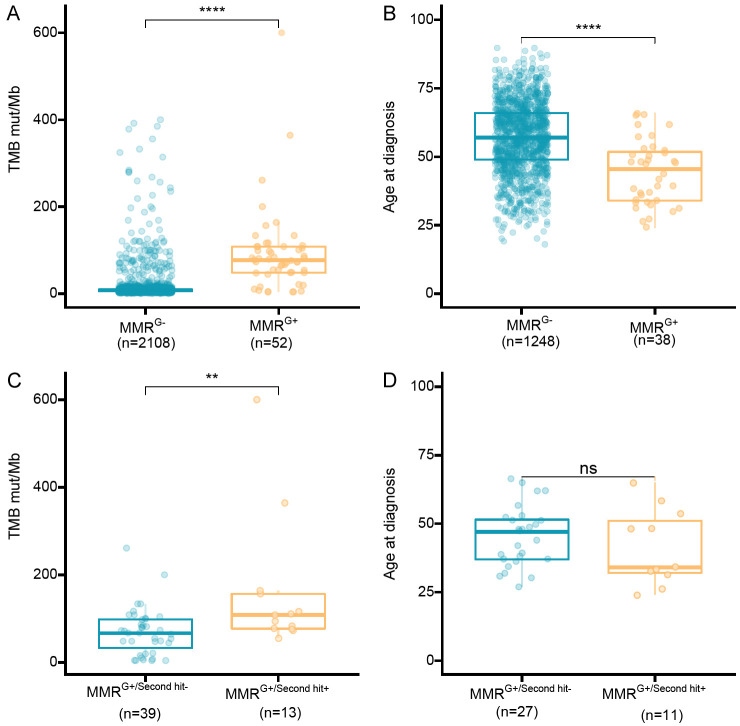
Among 2160 CRC individuals, 52 participants carried at least one pathogenic germline variants in MMR genes, of whom 41 (78.8%) were MSI-H. Compared with the non-carriers, the individuals with germline MMR variants had a higher TMB level (median TMB 76.7 VS 7.26, P < 0.001) and an earlier onset age (median age 45.5 vs 57, P < 0.001) (Figure 3A and 3B). Among the subset of 52 individuals with germline MMR gene variants, 13 carried with a second hit were all MSI-H, including 8 (72.7%) of 11 MSH6 carriers, 2 (40%) of 5 PMS2 carriers, 2 (9.1%) of 22 MLH1 carriers, and 1 (7.1%) of 14 MLH2 carriers, and had significantly higher TMB (median TMB 108 VS 66.9, P=0.004) but comparable onset age (data not shown) versus 39 second-hit non-carriers (Table 3; Figures 1C, 3C and 3D).
Figure 3.
Effects of germline mismatch repair (MMR) gene mutations and somatic second hits on tumor mutational burden (TMB) and age at diagnosis. (A, B) The effects of germline MMR gene mutations on TMB (A) and age at diagnosis (B); (C, D) The effects of MMR somatic second hits on TMB (A) and age at diagnosis (B). **P < 0.01, ****P < 0.001.
Discussion
In this study, 136 pathogenic germline variants were identified in 21 genes including well-known CRC-risk genes and DDR genes from 133 patients among 2160 Chinese CRC patients. Among the 133 CRC probands with germline variants, 24 had a somatic second hit in the wild-type allele of the affected gene in the tumor. To our knowledge, this is the first systematic study for pathogenic DDR germline mutations in the Chinese population with CRC, and somatic second hits in the tumors of CRC patients with the pathogenic germline mutations.
It has been well-acknowledged that the prevalence of hereditary syndrome in early-onset CRC is higher than that in the average-onset CRC population [13,15-17]. In line with this, the patients aged ≤ 50 had a higher frequency of pathogenic germline mutations compared with the average-onset CRC population, especially in inherited predisposition genes; and the individuals carried with pathogenic DDR germline variants in the overall population had a younger onset age of CRC than the non-carriers.
Mounting evidence points that germline DDR gene deficiency is related with increased cancer risk across tumor types. However, previous research for identification of germline DDR variants in CRC focused primarily on limited genes, a small subset or Caucasian populations. For example, it had been documented that the alterations of BRCA1/2, two homologous recombination repair (HRR) genes, were associated with early-onset CRC, and germline pathogenic variants of BRCA1 as a risk factor for CRC is on the rise [7,18]. The potentially pathogenic variants in Fanconi anemia (FA) pathway genes were identified in the familial CRC cohort, primarily located in BRCA2/FANCD1, BRIP1/FANCJ, FANCC, FANCE, and REV3L/POLZ, highlighting FA DDR pathway may play an important role in the inherited predisposition to CRC [19]. Another two studies documented that carriers of deleterious mutations of BLM, one HRR gene, or NTHL1, one base-excision repair (BER) gene, were at increased risk to develop CRC [20,21]. A recent study systematically examined germline DDR gene mutations in CRCs with Caucasian populations, and identified ATM and PALB2, two HRR genes, as CRC-risk genes [8]. Here, we investigated the germline variants of 32 DDR genes in 2160 CRCs from China via NGS. Among the examined DDRs, germline variants primarily fell in MLH1, MSH2, MSH6, and PMS2 in MMR pathway, BRCA2, BRCA1, PALB2, BLM, RAD50, ATM, BARD1, and BRIP1 in HRR pathway, and FANCD2, FANCA, and FANCG in the FA pathway, CHEK2, TP53, and ATR in DDR checkpoint signaling pathway, and MUTYH in BER pathway. Pathogenic DDR germline mutations increased TMB level and MSI-H incidence, and associated with early-onset of CRC. Most of the germline mutations identified in this study have been reported to be linked to inherited CRC susceptibility (e.g., MMR, BRCA1/2, ATM, PALB2, TP53, FANCD2, BLM, CHEK2) [7,8,18,20,21]. However, surprisingly, only 18% of the CRC probands with germline variants had an identified somatic second hit in the wild-type allele of the tumor, which might be mainly due to detection omission of the second hits caused by other forms of allelic inactivation (e.g., hypermethylation).
Pathogenic germline variants in the MMR genes are the major etiology of Lynch syndrome, the most common inherited cancer predisposition syndrome in CRC. MSI-H is the hallmark of tumors in patients with Lynch syndrome. More than 90% of Lynch-associated CRC tumors are MSI-H, which is mainly caused by germline mutations of MLH1 and MSH2 genes. Meanwhile, some of Lynch-associated CRC tumors caused by germline MSH6 mutations exhibit the phenotypes of MSS or MSI-L [22]. This phenomenon is anastomotic with the mechanism of action of MMR. MMR proteins usually work two by two, MLH1 with PMS2 and MSH2 with MSH6. However, the lack of PMS2 and MSH6 can be compensated by other proteins, and MLH1 and MSH2 proteins could still remain stable in the absence of their partners. As a consequence, mutations in MLH1 or MSH2 usually cause the loss of MLH1/PMS2 or MSH2/MSH6 protein, respectively, whereas mutations in PMS2 or MSH6 only affected its own expression [23]. Here, we found that MSI-H account for 78.8% in the patients with germline MMR variants. Interesting, 13 carried with a second hit in the affected MMR genes were all MSI-H, implying that the second hits substantially increased MSI-H occurrence. Notably, among the 13 carriers with a second hit, there were 8 (72.7%) of 11 MSH6 carriers, and 2 (40%) of 5 PMS2 carriers. Combined with the above findings, it is not irrational to speculate that the second hit is crucial for tumors with MSH6 or PMS2 germline mutation to induce MSI-H phenotype, while the MSI-H phenotype in tumors with MLH1 and MSH2 germline mutations might be driven by various mechanisms. Additionally, these patients with a second hit in MMR genes are more likely to have an extraordinarily high TMB phenotype which have a better chance to benefit from immunotherapy.
There were several limitations in our study. As a retrospective study, selection bias may have existed in the sample population, although all patients receiving genetic testing were enrolled in this study. Thirty-two frequently mutated DDRs were chosen instead of all DDRs, which may result in the omission of important findings. Moreover, due to a low mutation frequency of each specific gene, we did not perform a comprehensive analysis of the specific gene mutations.
Overall, we presented the spectrum of pathogenic germline mutations in DDR genes and well-known CRC-risk genes in Chinese CRCs, suggesting their important role in CRC susceptibility and management. In addition, the second hits in MMR genes significantly increased MSI and TMB, hinting that immunotherapy has more chances to be effective in the patients with MMR second hits.
Acknowledgements
The authors wish to thank all the patients that provided samples for the study. This study was supported by the Sanming Project of Medicine in Shenzhen, China (SZSM201812062).
Informed consent was exempted for this retrospective study.
Disclosure of conflict of interest
Bei Zhang, Shuang Tong, Jinping Cai, Feilong Zhao, Xiaochen Zhao, and Shiqing Chen are employed by 3D Medicines Inc.
Abbreviations
- BER
base-excision repair
- CAP
College of American Pathologists
- CLIA
Clinical Laboratory Improvement Amendments
- CRC
colorectal cancer
- DDR
DNA damage repair
- FA
Fanconi anemia
- FFPE
Formalin-Fixed Paraffin-Embedded
- H&E
hematoxylin and eosin
- HRR
homologous recombination repair
- MMR
mismatch repair
- MSI
microsatellite instability
- NGS
Next-generation sequencing
- TMB
tumor mutational burden
References
- 1.Siegel RL, Miller KD, Goding Sauer A, Fedewa SA, Butterly LF, Anderson JC, Cercek A, Smith RA, Jemal A. Colorectal cancer statistics, 2020. CA Cancer J Clin. 2020;70:145–164. doi: 10.3322/caac.21601. [DOI] [PubMed] [Google Scholar]
- 2.Lichtenstein P, Holm NV, Verkasalo PK, Iliadou A, Kaprio J, Koskenvuo M, Pukkala E, Skytthe A, Hemminki K. Environmental and heritable factors in the causation of cancer--analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med. 2000;343:78–85. doi: 10.1056/NEJM200007133430201. [DOI] [PubMed] [Google Scholar]
- 3.Lorans M, Dow E, Macrae FA, Winship IM, Buchanan DD. Update on hereditary colorectal cancer: improving the clinical utility of multigene panel testing. Clin Colorectal Cancer. 2018;17:e293–e305. doi: 10.1016/j.clcc.2018.01.001. [DOI] [PubMed] [Google Scholar]
- 4.Boland PM, Yurgelun MB, Boland CR. Recent progress in Lynch syndrome and other familial colorectal cancer syndromes. CA Cancer J Clin. 2018;68:217–231. doi: 10.3322/caac.21448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Stoffel EM, Mangu PB, Gruber SB, Hamilton SR, Kalady MF, Lau MW, Lu KH, Roach N, Limburg PJ American Society of Clinical Oncology; European Society of Clinical Oncology. Hereditary colorectal cancer syndromes: American society of clinical oncology clinical practice guideline endorsement of the familial risk-colorectal cancer: European society for medical oncology clinical practice guidelines. J. Clin. Oncol. 2015;33:209–217. doi: 10.1200/JCO.2014.58.1322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Soyano AE, Baldeo C, Kasi PM. BRCA mutation and its association with colorectal cancer. Clin Colorectal Cancer. 2018;17:e647–e650. doi: 10.1016/j.clcc.2018.06.006. [DOI] [PubMed] [Google Scholar]
- 7.Mauri G, Sartore-Bianchi A, Russo AG, Marsoni S, Bardelli A, Siena S. Early-onset colorectal cancer in young individuals. Mol Oncol. 2019;13:109–131. doi: 10.1002/1878-0261.12417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.AlDubayan SH, Giannakis M, Moore ND, Han GC, Reardon B, Hamada T, Mu XJ, Nishihara R, Qian Z, Liu L, Yurgelun MB, Syngal S, Garraway LA, Ogino S, Fuchs CS, Van Allen EM. Inherited DNA-repair defects in colorectal cancer. Am J Hum Genet. 2018;102:401–414. doi: 10.1016/j.ajhg.2018.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Knudson AG. Two genetic hits (more or less) to cancer. Nat Rev Cancer. 2001;1:157–162. doi: 10.1038/35101031. [DOI] [PubMed] [Google Scholar]
- 10.Su D, Zhang D, Chen K, Lu J, Wu J, Cao X, Ying L, Jin Q, Ye Y, Xie Z, Xiong L, Mao W, Li F. High performance of targeted next generation sequencing on variance detection in clinical tumor specimens in comparison with current conventional methods. J Exp Clin Cancer Res. 2017;36:121. doi: 10.1186/s13046-017-0591-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E, Voelkerding K, Rehm HL ACMG Laboratory Quality Assurance Committee. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American college of medical genetics and genomics and the Association for molecular pathology. Genet Med. 2015;17:405–424. doi: 10.1038/gim.2015.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Xiao J, Li W, Huang Y, Huang M, Li S, Zhai X, Zhao J, Gao C, Xie W, Qin H, Cai S, Bai Y, Lan P, Zou Y. A next-generation sequencing-based strategy combining microsatellite instability and tumor mutation burden for comprehensive molecular diagnosis of advanced colorectal cancer. BMC Cancer. 2021;21:282. doi: 10.1186/s12885-021-07942-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Stoffel EM, Koeppe E, Everett J, Ulintz P, Kiel M, Osborne J, Williams L, Hanson K, Gruber SB, Rozek LS. Germline genetic features of young individuals with colorectal cancer. Gastroenterology. 2018;154:897–905. e1. doi: 10.1053/j.gastro.2017.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Siegel RL, Fedewa SA, Anderson WF, Miller KD, Ma J, Rosenberg PS, Jemal A. Colorectal cancer incidence patterns in the United States, 1974-2013. J Natl Cancer Inst. 2017;109:djw322. doi: 10.1093/jnci/djw322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Xu T, Zhang Y, Zhang J, Qi C, Liu D, Wang Z, Li Y, Ji C, Li J, Lin X, Hou T, Liu H, Zhang L, Han-Zhang H, Shen L, Wang X. Germline profiling and molecular characterization of early onset metastatic colorectal cancer. Front Oncol. 2020;10:568911. doi: 10.3389/fonc.2020.568911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mork ME, You YN, Ying J, Bannon SA, Lynch PM, Rodriguez-Bigas MA, Vilar E. High prevalence of hereditary cancer syndromes in adolescents and young adults with colorectal cancer. J. Clin. Oncol. 2015;33:3544–3549. doi: 10.1200/JCO.2015.61.4503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pearlman R, Frankel WL, Swanson B, Zhao W, Yilmaz A, Miller K, Bacher J, Bigley C, Nelsen L, Goodfellow PJ, Goldberg RM, Paskett E, Shields PG, Freudenheim JL, Stanich PP, Lattimer I, Arnold M, Liyanarachchi S, Kalady M, Heald B, Greenwood C, Paquette I, Prues M, Draper DJ, Lindeman C, Kuebler JP, Reynolds K, Brell JM, Shaper AA, Mahesh S, Buie N, Weeman K, Shine K, Haut M, Edwards J, Bastola S, Wickham K, Khanduja KS, Zacks R, Pritchard CC, Shirts BH, Jacobson A, Allen B, de la Chapelle A, Hampel H Ohio Colorectal Cancer Prevention Initiative Study Group. Prevalence and spectrum of germline cancer susceptibility gene mutations among patients with early-onset colorectal cancer. JAMA Oncol. 2017;3:464–471. doi: 10.1001/jamaoncol.2016.5194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Cullinane CM, Creavin B, O’Connell EP, Kelly L, O’Sullivan MJ, Corrigan MA, Redmond HP. Risk of colorectal cancer associated with BRCA1 and/or BRCA2 mutation carriers: systematic review and meta-analysis. Br J Surg. 2020;107:951–959. doi: 10.1002/bjs.11603. [DOI] [PubMed] [Google Scholar]
- 19.Esteban-Jurado C, Franch-Exposito S, Munoz J, Ocana T, Carballal S, Lopez-Ceron M, Cuatrecasas M, Vila-Casadesus M, Lozano JJ, Serra E, Beltran S, Brea-Fernandez A, Ruiz-Ponte C, Castells A, Bujanda L, Garre P, Caldes T, Cubiella J, Balaguer F, Castellvi-Bel S. The Fanconi anemia DNA damage repair pathway in the spotlight for germline predisposition to colorectal cancer. Eur J Hum Genet. 2016;24:1501–1505. doi: 10.1038/ejhg.2016.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.de Voer RM, Hahn MM, Mensenkamp AR, Hoischen A, Gilissen C, Henkes A, Spruijt L, van Zelst-Stams WA, Kets CM, Verwiel ET, Nagtegaal ID, Schackert HK, van Kessel AG, Hoogerbrugge N, Ligtenberg MJ, Kuiper RP. Deleterious germline BLM mutations and the risk for early-onset colorectal cancer. Sci Rep. 2015;5:14060. doi: 10.1038/srep14060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Weren RD, Ligtenberg MJ, Kets CM, de Voer RM, Verwiel ET, Spruijt L, van Zelst-Stams WA, Jongmans MC, Gilissen C, Hehir-Kwa JY, Hoischen A, Shendure J, Boyle EA, Kamping EJ, Nagtegaal ID, Tops BB, Nagengast FM, Geurts van Kessel A, van Krieken JH, Kuiper RP, Hoogerbrugge N. A germline homozygous mutation in the base-excision repair gene NTHL1 causes adenomatous polyposis and colorectal cancer. Nat Genet. 2015;47:668–671. doi: 10.1038/ng.3287. [DOI] [PubMed] [Google Scholar]
- 22.Stoffel EM, Kastrinos F. Familial colorectal cancer, beyond Lynch syndrome. Clin Gastroenterol Hepatol. 2014;12:1059–1068. doi: 10.1016/j.cgh.2013.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.De’ Angelis GL, Bottarelli L, Azzoni C, De’ Angelis N, Leandro G, Di Mario F, Gaiani F, Negri F. Microsatellite instability in colorectal cancer. Acta Biomed. 2018;89:97–101. doi: 10.23750/abm.v89i9-S.7960. [DOI] [PMC free article] [PubMed] [Google Scholar]