Graphical abstract
Abbreviations: ABC, ATP-binding cassette; AD, Alzheimer’s disease; APP, amyloid precursor protein; ATP, Adenosine-triphosphate; BBB, blood-brain barrier; BODIPY-cholesterol, 4,4-difluoro-4-bora-3a,4a-diaza-s-indacene-cholesterol; cryo-EM, cryogenic-electron microscopy; EH, extracellular helix; ECD, extracellular domain; GSH, reduced glutathione; HTS, high-throughput screening; IC, intracellular helix; MOE, Molecular Operating Environment; MSD, membrane spanning domain; NBD, nucleotide binding domain; NBD-cholesterol, 7-nitro-2-1,3-benzoxadiazol-4-yl-cholesterol; PDB, protein data bank; PET, positron emission tomography; PLIF, protein ligand interaction; PSO, particle swarm optimization; R-domain/region, regulatory domain/region; RMSD, root mean square distance; SNP, single-nucleotide polymorphism; TM, transmembrane helix
Keywords: ABC transporter (ABCA1, ABCA4, ABCA7); Alzheimer’s disease (AD); Multitarget modulation (PANABC); PET tracer (PETABC); Polypharmacology; Rational drug design and development
Abstract
The adenosine-triphosphate-(ATP)-binding cassette (ABC) transporter ABCA7 is a genetic risk factor for Alzheimer’s disease (AD). Defective ABCA7 promotes AD development and/or progression. Unfortunately, ABCA7 belongs to the group of ‘under-studied’ ABC transporters that cannot be addressed by small-molecules. However, such small-molecules would allow for the exploration of ABCA7 as pharmacological target for the development of new AD diagnostics and therapeutics. Pan-ABC transporter modulators inherit the potential to explore under-studied ABC transporters as novel pharmacological targets by potentially binding to the proposed ‘multitarget binding site’. Using the recently reported cryogenic-electron microscopy (cryo-EM) structures of ABCA1 and ABCA4, a homology model of ABCA7 has been generated. A set of novel, diverse, and potent pan-ABC transporter inhibitors has been docked to this ABCA7 homology model for the discovery of the multitarget binding site. Subsequently, application of pharmacophore modelling identified the essential pharmacophore features of these compounds that may support the rational drug design of innovative diagnostics and therapeutics against AD.
1. Introduction
ABC transporters form the backbone of systemic barrier formation [1], [2] and are key factors in inter- and intracellular compartmentalization, allowing for the existence of opposing cellular processes. ABC transporter function has not only been demonstrated in the cellular membrane but also intracellularly in vesicular bodies [3], [4], [5]. Several representatives of the ABC transporter superfamily stand in association with the constitution and composition of cellular as well as vesicular membranes and microparticles [4], [6], such as certain A and G subfamily members. Furthermore, a contribution of these transporters to the formation and organization of lipid rafts has been discussed [4], [6], [7]. These lipid rafts are the venue of very important membrane-associated cellular processes, for example, secretase-mediated amyloid precursor protein (APP) processing, a process critical for the initiation and progress of AD [8], [9], [10].
Most ABCA and ABCG transporters are cholesterol and/or phospholipid transporters that regulate cellular lipid homeostasis [11], [12], which is not only linked to membrane permeability and barrier function but also to human diseases, such as AD [6], [13], [14], [15], [16], [17], [18], [19], [20], [21]. The lipid transporters ABCA1–2, ABCA5, ABCA7, ABCG1, and ABCG4 have particularly been associated with AD development [6], [13], [19], [22], [23], [24], [25], [26], [27], [28]. Genetic variant association studies regarding ABCA1 have revealed controversial information related to certain single-nucleotide polymorphisms (SNPs) and AD [6], [25], [29], [30], [31], [32], [33]. However, genome-wide association studies provided statistical proof that ABCA7 is strongly associated with AD development and/or progression [13], [24], [25], [34], [35], [36], [37]. Certain polymorphisms were associated with plaque formation in patients, which was correlated to an increased expression of ABCA7, supposedly as compensation for the increased Aβ load, suggesting an inhibition of Aβ deposition [22]. In addition, Abca7 knock-out led to Aβ load increase, while the overexpression of ABCA7 led to reduction of Aβ in mice [22]. ABCA7 has functionally be linked to cholesterol metabolism and phagocytosis, which may influence Aβ distribution and degradation [22].
Unfortunately, ABCA7 belongs to the so-called ‘under-studied’ ABC transporters that cannot be addressed by small-molecules [38]. Besides the substrates cholesterol and phospholipids [12], [13], [24], [34], the only small-molecules associated with ABCA7 are regulators of ABCA7 expression. These regulators include the inducers ponasterone A [39], pravastatin [40], [41], and rosuvastatin [40], as well as the downregulators digoxin [42] and 25-hydroxycholesterol [43] (all Fig. 1). These regulators are only of very limited use to monitor and/or influence the transport process, as they do not directly interact with ABCA7. However, deciphering this particular functional background would allow for the targeted development of novel AD diagnostics and therapeutics.
Fig. 1.
The only known small-molecule modulators of ABCA7.
A recently proposed approach to promote the development of directly-interacting under-studied ABC transporter-targeting agents is the generation and application of multitarget ABC transporter (‘pan-ABC transporter’) inhibitors to obtain novel lead structures for future diagnostic and therapeutic development [4], [38], [44], [45], [46]. A common ‘multitarget binding site’ of these modulators amongst the different ABC transporter subfamilies has been proposed [4], [38], [44], [45], which may serve as a starting point for rational drug design, exploration, and exploitation of under-studied ABC transporters as (polypharmacological) drug targets.
Since ABCA7 activity is believed to be reduced in AD [13], [25], activators (increase transport function) or correctors (restore functional conformation) are needed, and the use of ABC transporter inhibitors seems counterintuitive. However, although activators (no separation of, for example, ABCB1 and [47], [48], ABCC1 [47], [49], [50], ABCC2 [51], ABCC3 [51], ABCC8 [52], [53], [54], [55], or ABCC9 [52], [53], [55]) and correctors (e.g., for ABCA3 [56], ABCA4 [57], [58], ABCB4 [59], ABCB11 [60], or ABCC7 [61]) have been described in the literature, the knowledge of such agents is very scarce, undermining the concept of exploration of under-studied ABC transporters by a multitargeting approach. Luckily, the knowledge of pan-ABC transporter inhibitors is broader [45].
Additionally, it must be noted that several activators of ABC transporters share molecular-structural features of ABC transporter inhibitors. For example, tetrahydroisoquinolines (ABCB1) [48], [62], [63] and pyrrolopyrimidines (ABCC1) [49], [64] have been described for both activators [48], [49] and inhibitors [62], [63], [64] of ABCB1 and ABCC1, respectively. Other examples are the compound classes of phenothiazines [65], [66] and purines [49], [67], [68], which have been described to comprise of ABCC1 activators [49], [65], [66] and ABCB1 inhibitors [65], [66], [67], [68]. More examples can be found in the literature [47], [69]. Considering these findings, gained knowledge about the multitarget binding site by the use of multitargeting inhibitors may indeed provide necessary molecular-structural information of active scaffolds/substructures for the targeted design of ABCA7 activators (and/or correctors). In addition, imaging techniques using ABC transporter inhibitors have already been demonstrated to trace the expression of the respective ABC transporter in vivo [63], making ABC transporter inhibitors that are not substrates of the respective ABC transporter (e.g., ABCA7) eligible for, e.g., positron emission tomography (PET) as diagnostics in patients.
Several moderately potent pan-ABC transporter inhibitors have been discovered in recent years. Table 1 summarizes selected candidates (6–28) and their associated physicochemical properties. These pan-ABC transporter inhibitors can principally be divided into two groups: (i) truly multitarget pan-ABC transporter inhibitors and (ii) focused pan-ABC transporter inhibitors. Truly multitarget pan-ABC transporter inhibitors are mostly drugs and drug-like candidates that have been known for a long period of time to interfere with specific ABC transporters [45], [69], [70], [71], [72], [73] and were broadly evaluated toward several members. These compounds include benzbromarone (6; ABCB1 [74], ABCB11 [75], ABCC1–6 [69], [73], [76], [77], [78], ABCG2 [74]), imatinib (7; ABCA3 [79], ABCB1 [80], ABCB11 [75], ABCC1 [80], ABCC10 [80], ABCG2 [80]), quercetin (8; ABCB1 [76], ABCC1–2 [69], [76], ABCC4–5 [81], [82], ABCC11 [83], ABCG2 [69], ABCG6 [84]), verapamil (9; ABCA8 [85], ABCB1 [70], ABCB4–5 [86], [87], ABCB11 [88], ABCC1 [69], ABCC4 [89], ABCC10 [90], ABCG2 [76]), and verlukast (MK571, 10; ABCA8 [85], ABCB4 [86], ABCB11 [75], ABCC1–5 [69], [76], [91], [92], [93], ABCC10–11 [73], [94], ABCG2 [76]), all Fig. 2.
Table 1.
Selected pan-ABC transporter inhibitors that have been reported in the literature as well as associated physicochemical and molecular-structural properties determined by using InstandJChem version 20.15.0.
|
Compd. No. |
Original Name |
Molecular Weight |
Calc Log P |
Rotatable Bonds |
H-bond Acceptors |
H-bond Donors |
Targeted ABC Transporters |
|---|---|---|---|---|---|---|---|
| Truly Pan-ABC Transporter Inhibitors | |||||||
| 6 | benzbromarone | 424.09 | 5.55 | 3 | 2 | 1 | B1, B11, C1–6, G2 |
| 7 | imatinib | 493.62 | 4.38 | 7 | 7 | 2 | A3, B1, B11, C1, C10, G2 |
| 8 | quercetin | 302.24 | 2.16 | 1 | 7 | 5 | B1, C1–2, C4–5, C11, G2, G6 |
| 9 | verapamil | 454.61 | 5.04 | 13 | 6 | 0 | A8, B1, B4–5, B11, C1, C4, C11, G2 |
| 10 | verlukast | 515.08 | 5.67 | 11 | 4 | 1 | A8, B4, B11, C1–C5, C10–C11, G2 |
| Focused Pan-ABC Transporter Inhibitors | |||||||
| 11 | imidazole/pyrimidine 23 | 438.54 | 4.35 | 5 | 5 | 0 | B1, C1, G2 |
| 12 | quinoline/1,2,4-oxadiazole 15 | 421.45 | 4.11 | 7 | 7 | 0 | B1, C1, G2 |
| 13 | quinoline/1,3,4-thiadiazole 18 | 446.50 | 4.85 | 4 | 6 | 0 | B1, C1, G2 |
| 14 | quinazoline/1,2,4-oxadiazole 21 | 444.42 | 5.24 | 7 | 7 | 0 | B1, C1, G2 |
| 15 | quinoline/thieno[3,2-c]pyridine 22 | 424.54 | 3.31 | 2 | 5 | 0 | B1, C1, G2 |
| 16 | quinoline/1,2,4-oxadiazole 26 | 391.48 | 4.12 | 5 | 6 | 0 | B1, C1, G2 |
| 17 | pyrimidine 26 | 379.42 | 5.82 | 5 | 6 | 3 | B1, C1, G2 |
| 18 | tariquidar-related derivative 40 | 501.54 | 3.52 | 8 | 8 | 1 | B1, C1, G2 |
| 19 | pyrimdine 37 | 312.38 | 4.68 | 3 | 4 | 1 | B1, C1, G2 |
| 20 | amino aryl ester (S)-9 | 631.76 | 5.84 | 23 | 9 | 0 | B1, C1, G2 |
| 21 | thienopyrimidine 14 | 421.56 | 4.32 | 4 | 5 | 1 | B1, C1, G2 |
| 22 | pyrrolopyrimidine 55 | 489.63 | 5.73 | 8 | 5 | 1 | B1, C1, G2 |
| 23 | indolopyrimidine 69 | 357.46 | 4.12 | 4 | 4 | 1 | B1, C1, G2 |
| 24 | quinazoline 52 | 387.44 | 5.12 | 6 | 6 | 1 | B1, C1, G2 |
| 25 | quinoline 29 | 342.46 | 6.23 | 4 | 2 | 1 | B1, C1, G2 |
| 26 | thienopyridine 6r | 566.63 | 4.52 | 12 | 9 | 1 | B1, C1, G2 |
| 27 | benzoflavone 16 | 332.36 | 3.64 | 3 | 4 | 0 | B1, C1, G2 |
| 28 | MC18 | 393.53 | 4.90 | 6 | 4 | 0 | B1, C1, G2 |
Fig. 2.
Drugs and drug-like compounds that were discovered as truly multitarget pan-ABC transporter inhibitors.
Focused pan-ABC transporter inhibitors are high-throughput screening-(HTS)- and/or organic synthesis-derived small-molecules specifically designed to target the well-studied ABC transporters ABCB1, ABCC1, and ABCG2 [38], [45]. These are very modern molecules that were not evaluated at other transporters yet. The most potent representatives were designated as so-called ‘class 7′ molecules [38], [44], [45] – which exert their inhibitory power against ABCB1, ABCC1, and ABCG2 below 10 µM. A selection of candidates (11–28), which were also used in the present study, are shown in Fig. 3.
Fig. 3.
Selection of focused pan-ABC transporter inhibitors used in the present study.
Prospective computational approaches, such as molecular docking, support the search for potential binding sites of ligands, and thus, rational drug design in general [38]. However, molecular docking and binding pose analyses require structural information of the respective transporters or closely related siblings. Unfortunately, no X-ray structure of ABCA7 is available. Indeed, a cryo-EM structure of ABCA7 has already in 2020 been announced [95] on the protein data bank (https://www.rcsb.org; PDB ID: 7KQC; PDB status: unreleased deposition withdrawn) and bioRxiv (https://www.biorxiv.org), however, the respective structural information is at the time of writing of this manuscript not publicly available and cannot be used for subsequent computational operations. Luckily, cryo-EM structures of ABCA1 (4.1 Å) [96] and ABCA4 (3.3–3.6 Å) [97], [98], [99] have recently been reported, allowing for homology modelling and generation of new structural information. Homology models of other less- and under-studied ABC transporters in combination with molecular docking experiments led already to the discovery of novel bioactive molecules, as for example, for ABCC5 [100] or ABCG2 [101]. This poses a good perspective for the design and discovery of new interactors and putative binding sites of ABCA7.
The present study aimed for the development of an ABCA7 homology model and subsequent blind docking analysis of the membrane-spanning domains (MDDs) with the pan-ABC transporter inhibitors 6–28, as the multitarget biding site that is supposed to be shared amongst several subfamily members of ABC transporters was postulated to be located in the proximity of the MSDs [4]. This postulation is supported by cryo-EM (ABCB1 [102], ABCB2 [103], ABCB4 [104], ABCC8 [105], and ABCG2 [106]) and homology model data (ABCB1 [107], ABCB2 [108], ABCB5 [109], ABCC1 [110], ABCC4 [111], ABCC5 [100], ABCC6 [112], ABCC7 [113], ABCC11 [114], and ABCG2 [101]) of other ABC transporters which had their primary substrate/inhibitor binding sites within the transmembrane regions.
The herein presented blind docking experiments identified a putative binding pocket of a majority of the 23 selected molecules that reflected a common pharmacophore. These discoveries have broad implications regarding the development of novel AD diagnostics and therapeutics.
2. Results and discussion
2.1. Sequence alignment
Supplementary Fig. S1 compares the canonical amino acid sequences of ABCA1 (UniProt ID: O95477), ABCA4 (UniProt ID: P78363), as well as ABCA7 (UniProt ID: Q8IZY2), and important structural elements have been marked. ABCA7 is from the structural perspective a ‘common’ ABCA transporter. It is large in size, consisting of 2,146 amino acids, comparable to the 2,261 and 2,273 amino acids of ABCA1 [96] and ABCA4 [97], [98], [99], respectively. ABCA7 shares 52.8% and 48.2% sequence homology with ABCA1 and ABCA4, respectively, which is in agreement with the literature data (54% and 49%, respectively) [24], [34], [115]. Functionally, ABCA1 and ABCA7 are closely related, recognizing and translocating cholesterol and phospholipids [12], [13], [24], [34], while ABCA4 is mainly a retinoid transporter [12]. Considering its greater similarity in terms of the amino acid sequence and its closer functional similarity, the generation of the ABCA7 homology model has been accomplished taking only the cryo-EM structure of ABCA1 into account [96]. However, for the sequential and structural interpretation, in particular in terms of phospholipid binding, specific structural aspects of the ABCA4 cryo-EM structures have been compared to the obtained homology model as model validation [97], [98], [99].
2.2. Homology model
Fig. 4 (A)–(D) provides all structural information available on ABCA transporters, namely ABCA1 [(A), PDB ID: 5XJY [96]] and ABCA4 [(B), PDB ID: 7E7I [98]; (C), PDB ID: 7LKP [99]; (D), PDB ID: 7E7O [97]]. The underlying cryo-EM structures of ABCA1 and ABCA4 had resolutions of 4.1 Å [96] and 3.3–3.6 Å [97], [98], [99], respectively. Furthermore, Fig. 4 (E) shows the modelled ABCA7 structure according to the structural composition and orientation of ABCA1 [96], and Supplementary Fig. S2 provides the Ramachandran plot of the modelled ABCA7. The overall quality factor [116] of the model is 87%. This is a rather low value that stems from the fact that ABCA7 is a very large protein of 2146 amino acids. Structural templates with high resolutions of large proteins like ABCA7 have rarely been assessed in terms of model quality, and hence, these results cannot accurately be put into perspective. Nevertheless, the low value is exclusively based on the highly-flexible loop and ECD regions, which were particularly not focused in the present study.
Fig. 4.
Structural information and conformational representation of ABCA transporters that can be found in the literature [96], [98], [99] using Molecular Operating Environment (MOE) version 2019.01 [118]. (A) Cryo-EM structure of human ABCA1 as reported by Qian et al. in 2017 [96]. (B) Cryo-EM structure of human ABCA4 as reported by Xie et al. in 2021 [98]. (C) Cryo-EM structure of human ABCA4 as reported by Liu et al. in 2021 [99]. (D) Cryo-EM structure of human ABCA4 as reported by Scortecci et al. in 2021 [97] (E) Homology model of human ABCA7 generated from the cryo-EM structure of human ABCA1 [96]. The membrane bilayers in (A)–(E) are indicated as brown balls and light brown areas. (F) simplified scheme of the organization of ABCA7 and its structural components. The inter-membrane space is indicated as light brown area, and the border to the cytosol and lumen is indicated by brown lines.
In terms of its structural organization, ABCA7 is – like ABCA1 [96] and ABCA4 [97], [98], [99] as well – a so-called ‘full transporter’, consisting of two membrane spanning domains (MSD1 and MSD2), each composed of six transmembrane helices (TM1–6 and TM7–12). As already described for ABCA1 [96] and ABCA4 before [98], TM1–6 (MSD1) as well as TM7–12 (MSD2) represent separate entities without large intermolecular interactions between the two MSDs, which is in contrast to a swapped/twisted transmembrane structure, as this is the case, for example, for ABCB1 [102], [106]. This separation of the MSDs is a typical construct of so-called ‘type II’ transporters and has first been described for the model type II transporter, ABCG5/ABCG8 (PDB ID: 5DO7 [117]), which is also a lipid transporter [11]. Only TM5 and TM11 form a contact zone, as demonstrated for ABCA1 [96] and ABCA4 [97], [98], [99]. The other helices of ABCA7 are completely exposed to the hydrophobic inter-membrane space, potentially for attraction of and interaction with membrane-bound/solved cholesterol and phospholipids before these molecules are guided into the substrate/drug translocation cavity. This conformational orientation is called the ‘lateral-opening’ conformation [98]. The substrate/drug translocation cavity is formed by the two MSDs and stretches from the cytoplasmic to the luminal side through the whole membrane bilayer. Indeed, residues of TM1–2 and TM6 of ABCA1 have been proposed to form a ‘phospholipid binding pocket’ [96], and the structural data of ABCA4 revealed several binding sites for phospholipids and N-retinylidene-phosphatidylethanolamine [97], [98] within the cavity built by the two MSDs.
Structural similarity between ABCA1 [96], ABCA4 [97], [98], [99], ABCG5/ABCG8 [117], and ABCA7 can also be seen in terms of 4 intracellular helices (IHs) between TM1–2, TM3–4, TM7–8, and TM9–10. These IHs are typical in type II transporters and are believed to provide the necessary flexibility for the allosteric communication between the MSDs and NBDs in the substrate/drug translocation process [117]. The same accounts for 4 extracellular helices (EHs) between both TM5–6 and TM11–12.
MSD1 and MSD2 of ABCA7 are flanked by one nucleotide-binding domain (NBD1 and NBD2) each at the carboxy terminus of the respective MSD within the cytoplasm. These NBDs bind and cleave ATP to ADP and Pi and generate the necessary energy for active transport. Both NBDs of ABCA7 bear the Walker A (GXXGXGKS/T; X = any amino acid) [119], [120] and Walker B (XXXXD; X = hydrophobic amino acid) [119], [120] motifs as well as the ABC signature motif (NBD1: LSGGM; NBD2: YSGGN) [99], [119], which together form the most highly conserved features amongst all ABC transporters [119], [120]. Interestingly, the carboxy termini of both NBDs are occupied by one regulatory domain each (‘R-domain’/‘R-region’; R1 and R2), which are suggested to stabilize NBD1/NBD2 interaction [96], [98] and strongly interact with one another in the absence of ATP [99].
ABCA7 bears also two extracellular domains (ECD1 and ECD2), which represent special structural features of ABCA transporters [96], [97], [98], [99]. These domains are not present in other ABC transporter subfamilies. ECD1 and ECD2 elongate ABCA7 to a total height of ∼216 Å, comparable to ∼200 Å of ABCA1 [96] and ∼230–240 Å of ABCA4 [97], [98], [99]. ECD1 is located between TM1–2, while ECD2 is located between TM7–8. Both ECDs form a hydrophobic channel embedded in the intraluminal body of ∼105 Å height as already found particularly for ABCA1 (∼100 Å)[96] and ABCA4 (∼130 Å) [99] before. The ECDs are composed of 506 and 273 amino acids, respectively, compared to 583 and 270 amino acids of ABCA1 [96] as well as 600 and 290 amino acids of ABCA4 [99]. This channel is similar to the discovered ‘base-tunnel-lid’ flame-shaped body as described for ABCA1 [96] and ABCA4 [98] before, with strong interactions of the ‘base’ and the formed ‘tunnel’. The tunnel of ABCA7 is composed of – similar to ABCA1 [96] – mostly hydrophobic amino acids. The ECDs are suggested to bind the main acceptor of cholesterol from the reverse cholesterol transport mediated by ABCA1, APOA1 [121]. Specifically the tunnel was proposed as (temporary) storage of cholesterol and/or phospholipids [96], which is also likely in terms of the cholesterol and phospholipid transporter ABCA7 [12], [13], [24], [34]. In comparison to the ABCA1 template, ECD1 of ABCA7 is shorter. This reflects in only two disulfide bonds compared to ABCA1, which contains three. The third disulfide bond formed between Cys75 and Cys309 (ABCA1) is not present in the homology model of ABCA7. On the other hand, the forth disulfide bond in ECD2 is present in both ABCA1 and ABCA7. Interestingly, the interruption in sequence of ABCA7 between the putative transmembrane tunnel/binding cavity and the ECD-tunnel hints to necessary conformational changes for the substrate/drug translocation process, which was also suggested for its functional counterpart, ABCA1 [96]. Fig. 4 (F) gives the schematic representation of the structural components of ABCA7.
The transporter ABCA1 selected as template for the homology modelling approach has been resolved in an ATP-unbound state [96]. This is of major importance, as this state allows for substrate/drug/modulator recognition/binding before ATP-dependent translocation of the substrate. This was demonstrated for other ABC transporters as well, e.g., ABCB1 [102], ABCB2 [103], ABCB6 [122], ABCB11 [123], ABCC7 [124], ABCC8 [105], ABCG2 [106], and ABCG5/G8 [125], forming either an inward-facing [105], [106], [122], [123], [124], [125] or (drug-induced) occluded [102], [103] conformation. In contrast, most ABC transporters in the ATP-bound state are not able to recognize substrates, as shown for ABCB4 [104], as these form an outward-facing conformation. Only in exceptional cases an ATP-bound state was observed with an inward-facing conformation, for example, the mitochondrial ABC transporters ABCB8 [126] and ABCB10 [127]. In the case of the lysosomal ABC transporter ABCD4, the outward-facing conformation (bound to two molecules of ATP) was suggested as the actual state of acceptance of the substrate cobalamin [128]. These findings could be peculiarities of vesicular transporters [3], [4] and are likely to have no implications for other ABC transporters, such as ABCA7.
Interestingly, ABCA1 was found in a ‘(pseudo-)outward-facing’ conformation, which means that – unlike other ABC transporters in the ATP-unbound state [102], [103], [106], [122] – the MSDs seem not to be exposed to the cytosol for substrate/drug/modulator attraction and translocation. This structural feature was distinct from the other known type II ABC transporter, ABCG5/ABCG8 [117]. However, the found state for ABCA1 [96] was supported by the cryo-EM data of ABCA4 [97], [98], [99]. In addition, as most TMs are oriented toward the inter-membrane space – potentially to attract and bind cholesterol and phospholipids present within the membrane – a classical ‘inward-facing’ conformation seems unnecessary. In conclusion, this (pseudo-)outward-facing conformation appears not to be a peculiarity of type II transporters in general but of ABCA transporters in particular, and we were confident with the generated homology model to be in the most relevant conformational state for ligand binding to continue with molecular modelling studies.
2.3. Molecular docking
Directly interacting small-molecules of ABCA7 are unknown except for the substrates cholesterol and phospholipids [12], [13], [24], [34]. No genuine structural information of ABCA7 is available from which a potential binding site of its substrates could have been deduced. A lipid-filled gap could be identified in the recently announced cryo-EM structure of ABCA7 (PDB ID: 7KQC; PDB status: unreleased deposition withdrawn), however, neither the identity nor the binding region/site of these lipids could be determined [95]. A binding site for the most studied ABCA transporter ABCA1 has only been suggested [96], and only the very recently published ABCA4 cryo-EM structures outlined potential phospholipid and retinoid binding sites [97], [98]. However, a common or overlapping (multitarget) binding site amongst all ABC transporter superfamily members has recently been suggested acknowledging the multitarget affinity and potency of truly multitarget as well as focused pan-ABC transporter inhibitors (Table 1) [4], [38], [45]. These pan-ABC transporter inhibitors inhere the molecular-structural information for multitarget inhibition – and therefore the potential of multitarget exploration of under-studied ABC transporters [38], [45], such as ABCA7. Thus, we conducted blind docking studies considering the entire MSDs of the ABCA7 homology model using AutoDock [129]. The MSDs as docking space were particularly chosen because they cover the suggested/outlined binding sites in other ABCA transporters [96], [97], [98], which is generally supported by the structural information of other ABC transporters subfamilies [100], [101], [102], [103], [104], [105], [106], [107], [108], [109], [110], [111], [112], [113], [114]. Fig. 5 (A) outlines the focused space for the blind docking experiments.
Fig. 5.
Blind docking using the herein described homology model of ABCA7 applying AutoDock [129]. (A) Space chosen for the blind docking experiments within the membrane bilayer indicated with brown balls and a light brown area. (B) Superimposed top ranking docking poses of the ten chosen pan-ABC transporter inhibitors 9–11, 14, 17, and 22–26[38], [45], [101], [134], [135], [136], [137] amongst the 50 docking poses generated by AutoDock [129] (colored cyan, stick representation) within the MSDs of the ABCA7 homology model. (C) Close-up of the superimposed top ranking poses of the docked molecules 9–11, 14, 17, and 22–26[38], [45], [101], [134], [135], [136], [137]. Nonpolar hydrogen atoms were omitted, and polar hydrogen, carbon, nitrogen, oxygen, as well as sulfur atoms were colored in silver white, cyan, blue, red, and dark yellow, respectively. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
In a first step, we assembled a list of 23 diverse, modern, and potent pan-ABC transporter inhibitors (6–28; Fig. 2; Table 1) covering the maximal structural diversity possible amongst known pan-ABC transporter inhibitors [38], [45], [101], [130], [131], [132], [133], [134], [135], [136], [137], [138], [139], [140]. From these 23 molecules, a first set of 10 diverse molecular representatives was chosen for the blind docking experiments (9–11, 14, 17, and 22–26) [38], [45], [101], [134], [135], [136], [137] using AutoDock [129]. Compounds 9–10 were chosen because these two molecules are the most promiscuous known in terms of ABC transporter inhibition (9 [69], [70], [76], [85], [86], [87], [88], [89], [90] and 11 [69], [73], [75], [76], [85], [86], [91], [92], [93], [94] targets, respectively). Amongst the targets of compounds 9–10 is also another ABCA transporter, namely ABCA8 [85]. Compound 11 was taken due to its novelty and structural uniqueness [38]. Finally, compounds 14, 17, and 22–26 were taken due to their high multitarget potency, as these compounds are all class 7 molecules (IC50 (ABCB1, ABCC1, and ABCG2)<10 µM) [45], [101], [134], [135], [136], [137].
Fig. 5 (B) provides the top ranking docking poses of these 10 docked molecules 9–11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] amongst the 50 docking poses generated for each molecule by AutoDock [129] in the ABCA7 homology model. The top ranking docking pose of each individual molecule can be found in Supplementary Fig. S3 (A)–(J), and Supplementary Table 1 provides the docking scores of the top ranking docking pose of each molecule obtained from AutoDock [129]. In addition, Supplementary Table 1 contains also the docking scores of the two phospholipids PL1 and PL2 that were found complexed in one of the ABCA4 cryo-EM structures [98]. These scores were better in comparison to the docked pan-ABC transporter inhibitors; however, this may be due to the very strong electrostatic interactions of the phosphate group and the amino acids, which did not occur in the case of pan-ABC transporter inhibitors. As can be seen from the docking poses, the molecular-structural orientation of the majority of the pan-ABC transporter modulators resembled or partially overlapped each other [Fig. 5 (C)]. The docking poses of the molecules largely bind between the two MSDs interacting with residues of TM1–2, TM5, TM7–TM8, and TM11, majorly through hydrophobic interactions, which is in agreement with the binding pocket identified for phospholipids in ABCA4 [97], [98]. The identified molecular-structural orientations of compounds 9–10, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] of the top ranking docking poses suggest common or partially overlapping pharmacophore features amongst these compounds, which was subsequently explored by applying pharmacophore modelling.
2.4. Pharmacophore modelling
In order to explore common or overlapping pharmacophore features between pan-ABC transporter modulators, pharmacophore modelling was applied using MOE 2019.01 [118]. The model was generated by utilizing the top ranking docking poses of compounds 9–11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] obtained from the molecular docking studies with AutoDock [129] [Fig. 5 (B)–(C)]. Fig. 6 (A)–(B) outlines the four identified pharmacophore features F1–F2 (aromatic/hydrophobic), F3 (aromatic), and F4 (acceptor/donor), as well as the distances between the individual features.
Fig. 6.
Pharmacophore model using the top ranking docking poses of compounds 9–11, 14, 17, and 22–26[38], [45], [101], [134], [135], [136], [137] obtained from AutoDock [129]. (A) Superimposed top ranking poses of the docked compounds 9–11, 14, 17, and 22–26[38], [45], [101], [134], [135], [136], [137] from which the four pharmacophore features F1–F4 could be deduced (colored cyan, stick representation). Nonpolar hydrogen atoms were omitted, and polar hydrogen, carbon, nitrogen, oxygen, as well as sulfur atoms were colored in silver white, cyan, blue, red, and dark yellow, respectively. (B) The four pharmacophore features F1–F2 (aromatic/hydrophobic), F3 (aromatic), and F4 (acceptor/donor) are depicted in orange (F1–F3) as well as silver (F4). The distances between the individual features are indicated as light green lines and are outlined in the table. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
As the top ranking binding pose of a molecule is not necessarily the most relevant one, the other 49 binding poses generated from AutoDock [129] needed to be explored. This was accomplished by a pharmacophore search amongst the 50 docked poses of compounds 9–11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] obtained from AutoDock [129] by screening against the four identified pharmacophore features F1–F4. Very interestingly, at least one of the 50 AutoDock-generated [129] conformations of each of the 10 molecules matched in full with the four identified pharmacophore features. The best fitting conformations of compounds 9–11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] in terms of the pharmacophore features F1–F4 are shown in Supplementary Fig. S4 (A)–(J), and the root mean square distance (RMSD) of the best fitting docking poses of the 10 molecules in terms of the pharmacophore features F1–F4 with the respective docking scores are given in Supplementary Table 1.
2.5. Model validation
To further validate the generated pharmacophore model, we applied another docking tool, Glide [141], [142], implemented in Schrödinger 2020–3 [143], and the very same compounds 9–11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] were docked. Fig. 7 demonstrates the top ranking poses of the docked molecules amongst the 10 docking poses generated by Glide [141], [142]. Supplementary Fig. S5 (A)–(J) provides each individual top ranking docking pose of the compounds, and Supplementary Table 2 provides the respective docking scores as well as the docking scores of the two phospholipids that were found complexed in one of the ABCA4 cryo-EM structures [98]. With respect to Glide [141], [142], the docking scores of the phospholipids were comparable to the docking scores of the pan-ABC transporter inhibitors.
Fig. 7.
Blind docking using the herein described homology model of ABCA7 applying Glide [141], [142]. (A) Superimposed top ranking docking poses of the pan-ABC transporter inhibitors 9–11, 14, 17, and 22–26[38], [45], [101], [134], [135], [136], [137] amongst the 10 docking poses generated by Glide [141], [142] (colored yellow, stick representation) within the MSDs of the ABCA7 homology model. (B) Close-up of the superimposed top ranking poses of the docked molecules 9–11, 14, 17, and 22–26[38], [45], [101], [134], [135], [136], [137]. Nonpolar hydrogen atoms were omitted, and polar hydrogen, carbon, nitrogen, oxygen, as well as sulfur atoms were colored in silver white, yellow, red, blue, and dark yellow, respectively. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
Similar to AutoDock [129], the top ranking docking poses generated from Glide [141], [142] do not need to be the most relevant ones in terms of the binding of the molecules. Thus, in our next step, the 10 resultant docking poses of each of the compounds 9–11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] obtained from Glide [141], [142] were screened against the established pharmacophore model [Fig. 6 (A)–(B)]. The best fitting conformations of compounds 9–11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] in terms of the pharmacophore features F1–F4 are shown in Supplementary Fig. S6 (A)–(J), and the RMSD of the best fitting docking poses of the 10 molecules in terms of the pharmacophore features F1–F4 with the respective docking scores are given in Supplementary Table 2.
Strikingly, only compound 10 did not fully match the searched pharmacophore features F1–F4, while at least one of the 10 generated docking poses from Glide [141], [142] of each of the compounds 9, 11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] were identified as hit in terms of the four pharmacophore features F1–F4 [Fig. 6 (A)–(B)]. The reason for this may lie in the presence of an acid function, which is unique amongst the docked compounds 9, 11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137] and forms different possible interactions.
Strikingly, pharmacophore modelling taking the top ranking docking poses generated from Glide [141], [142] into account resulted in similar four pharmacophore features [F1–F3 (aromatic/hydrophobic) and F4 (acceptor/donor); Supplementary Fig. S7], which were reflected in full in both the generated poses from Autodock [129] (Supplementary Fig. S8) and Glide [141], [142] (Supplementary Fig. S9) of compounds 9, 11, 14, 17, and 22–26 [38], [45], [101], [134], [135], [136], [137]. The elucidated pharmacophore features identified from Glide [141], [142] are closer to each other than the features from AutoDock [129]. Although no direct overlap could be observed between the two pharmacophore models, the identified features (particularly aromatic/hydrophobic) seem to complement each other. These results suggest that the specified pharmacophore model and the respective binding site could be explored as common or overlapping binding site of pan-ABC transporter inhibitors.
In the next step of model validation, a second set of 13 pan-ABC transporter inhibitors, compounds 6–8, 12–13, 15–16, 18–21, and 27–28 [45], [130], [131], [132], [133], [138], [139], [140] (Fig. 2; Table 1), was docked with both AutoDock [129] and Glide [141], [142]. The resultant top ranking docking poses are shown in Fig. 8 (A)–(D). Supplementary Figs. S10 (A)–(M) and S11 (A)–(M) provide the individual top ranking docking poses of the compounds applying either AutoDock [129] or Glide [141], [142], and the docking scores are provided in Supplementary Tables 3 (AutoDock [129]) and 4 (Glide [141], [142]). Furthermore, conformers of these 13 molecules were generated using the conformer generation tool implemented in MOE 2019.01 [118].
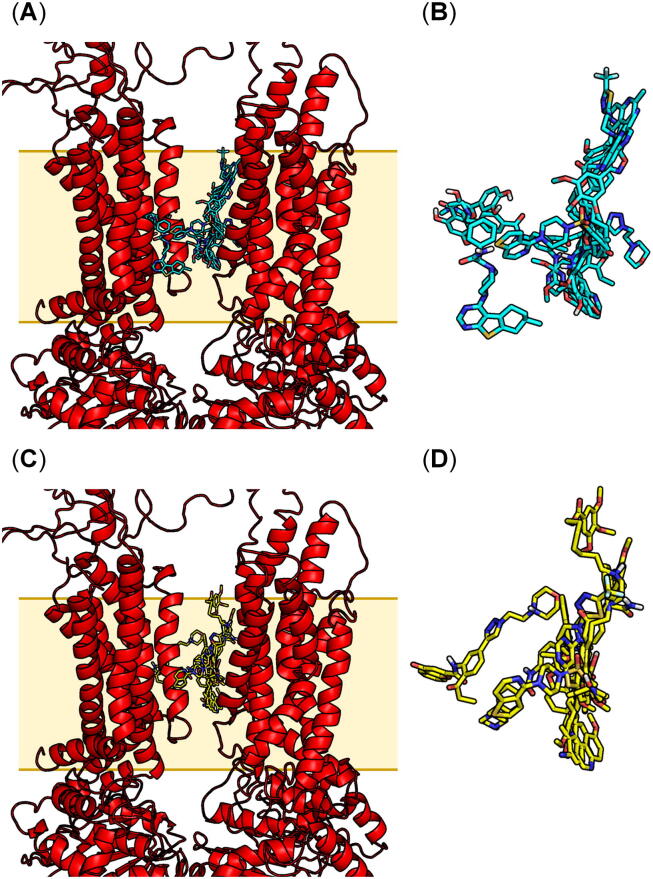
Fig. 8.
Extended docking experiments with compounds 6–8, 12–13, 15–16, 18–21, and 27–28[45], [130], [131], [132], [133], [138], [139], [140] for model validation purposes. (A) Superimposed top ranking docking poses of the compounds obtained from AutoDock [129]. (B) Close-up of the superimposed top ranking poses of the docked molecules. (C) Superimposed top ranking docking poses of the compounds obtained from Glide [141], [142]. (D) Close-up of the superimposed top ranking poses of the docked molecules. The compounds are colored in cyan (A)–(B) as well as yellow (C)–(D) and are shown in stick representation. Nonpolar hydrogen atoms were omitted, and polar hydrogen, carbon, nitrogen, oxygen, as well as sulfur atoms were colored in silver white, cyan/yellow, blue, red, and dark yellow, respectively. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
The 50 and 10 docked poses generated from AutoDock [129] and Glide [141], [142], respectively, as well as the 2,445 conformers generated with the conformer generation tool (MOE 2019.01) [118] for each molecule were screened against the developed pharmacophore model generated from the top ranking docking poses as obtained from Autodock [149] as described above [Fig. 6 (A)–(B)]. From these 13 candidates, 9 (AutoDock [129]), 7 (Glide [141], [142]), and 10 [conformer generation tool (MOE 2019.01) [118]] molecules were identified as hit [118], [129], [141], [142]. Supplementary Figs. S12 (A)–(I), S13 (A)–(G), and S14 (A)–(J) provide the best fitting conformations of compounds 6–8, 12–13, 15–16, 18–21, and 27–28 [45], [130], [131], [132], [133], [138], [139], [140] in terms of the pharmacophore features F1–F4 [Fig. 6 (A)–(B)], and the RMSD of the best fitting docking poses of the molecules in terms of the pharmacophore features F1–F4 with the respective docking scores are given in Supplementary Tables 3 (AutoDock [129]) and 4 (Glide [141], [142]). In essence, 7 compounds (12–13, 15–16, 18, 21, 27 [45], [130], [133], [138]) identified from these three approaches reflected the four pharmacophore features F1–F4 in full. On the other hand, compounds 6–8, 19–20, and 27 matched only in part with the four pharmacophore features F1–F4. As different binding sites and (sub)pockets of ABC transporter ligands generally exist [47], [98], [101], the proposed binding site of multitarget inhibitors is potentially favorable for some of the pan-ABC transporter inhibitors and results in strong binding affinities. Alternatively, weaker affinities may result for other molecules that mutually and/or additionally bind to (a) distinct binding site(s) as they bear only a part of the ‘necessary’ features.
2.6. Putative binding site
In the last step, we analyzed in detail the putative interactions of the 23 compounds with the amino acids in the binding pocket of the multitarget binding site. For this, we used the protein–ligand interaction fingerprint (PLIF) analysis tool implemented in MOE 2019.01 [118]. Fig. 9 provides the 2D interaction diagram of a representative compound, 15, in its top ranking docking pose as obtained from docking with AutoDock [129] (A) and Glide [141], [142] (B). Supplementary Figs. S15 (A)–(J), S16 (A)–(J), S17 (A)–(M), and S18 (A)–(M) show the putative interactions of all evaluated 23 molecules in their top ranking docking poses as obtained from AutoDock [129] and Glide [141], [142].
Fig. 9.
2D interaction diagram of the top ranking docking poses of compound 15 and possible strong interactions with amino acids of the multitarget binding site generated by PLIF implemented in MOE 2019.01 [118]. (A) Possible interactions obtained from AutoDock [129] formed with phenylalanine 1544 and cysteine 1653. (B) Possible interactions obtained from Glide [141], [142] formed with leucine 662 and cysteine 1653. Hydrophobic and polar amino acids are shown in green circles and purple circles, respectively. Interactions with the amino acids are depicted as green dotted arrows (side chain acceptor) or green dotted lines (aromatic). (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
Compound 15 possibly formed strong hydrophobic interactions with leucine 662 and phenylalanine 1544 with its quinoline basic scaffold as demonstrated in AutoDock [129] and Glide [141], [142], respectively. In addition, both docking tools revealed cysteine 1653 as an important amino acid that possibly interacts with the sulfone of compound 15. Both the hydrophobic and acceptor interaction of the quinoline and sulfone substructures, respectively, are two major parts of the developed pharmacophore models [Fig. 6 (B) and Supplementary Fig. S7]. These three amino acids are of importance as they are shared as interacting amino acids amongst several selected compounds and are highlighted in yellow in Supplementary Fig. S1. The bar code diagram in Fig. 10 provides insights into the most frequently occurring interactions with amino acids and the 23 docked compounds. Besides the mentioned amino acids, AudoDock [129] revealed also cysteine 659, valine 1649, and threonine 1652 as putatively important. These amino acids are located in TM5, TM8, and TM11 and are also highlighted in yellow in Supplementary Fig. S1. None of these six amino acids appeared in a critical region as outlined in Supplementary Fig. S2. Interestingly, cysteine 659, phenylalanine 1544, and valine 1649 are highly conserved amongst ABCA1 [96], ABCA4 [98], [99], and ABCA7 and may form the backbone of the multitarget binding site. A further detailed analysis of these amino acids with different methodologies is in need to validate the found results.
Fig. 10.
Distribution of the 23 docked compounds that may form possible interactions with amino acids in the MSDs of the generated homology model of ABCA7. (A) Bar code diagram and (B) population of the 23 docked compounds as obtained from AutoDock [129]). (C) Bar code diagram and (D) population of the 23 docked compounds as obtained from Glide [141], [142]. Single or multiple interactions are indicated with single and multiple columns, respectively. The respective engaged compound is indicated in the bar code diagram.
3. Conclusions
Multitarget agents have generally received increased interest over the last years, not only in terms of polypharmacology [46], [144], [145], [146] but also in terms of exploration of under-studied pharmacological targets [38], [44], [45]. Most members of the ABC transporter superfamily are considered as under-studied and cannot be addressed by small-molecules [38], [44], [45]. Except for ABCA1, which is a so-called ‘less-studied’ ABC transporter [38] with only 14 known inhibitors of this transport protein [147], [148], [149], [150], all members of the ABCA subfamily are under-studied. Half of the ABCA transporter sub-family is associated with AD [13], [19], [24], [25], [26], [27], [28], [34], and defective ABCA7 in particular has statistically been proven to contribute to AD development and/or progression [13], [24], [25], [34], [35], [37]. The lack of directly interfering small-molecules hinders research in developing molecular tools to functionally study this particularly important ABC transporter, hampering the development of novel and innovative AD diagnostics and therapeutics.
This lack of small-molecule interactors – of under-studied ABC transporters in general, and ABCA7 in particular – can be explained by three aspects: (i) the lack of multitargeting agents that share affinities to several ABC transporters of different sub-families, in particular ABCA transporters. The development of multitarget inhibitors has only very recently been focused [38], [44], [45], which limits the number of molecules. To this date, only 140 focused pan-ABC transporter inhibitors exist [38], [44], [45], [130], [151], [152], and non-focused pan-ABC transporter inhibitors (independent of targeted transporters) with at least 3 targeted transporters have only been described for roughly 120 compounds. Generally, modulators of ABCA transporters are less known, only ABCA1 (14 inhibitors [147], [148], [149], [150]) and ABCA8 (7 inhibitors [85], [153]) can be addressed by inhibitors, and small-molecule activators are unknown; (ii) the lack of structural information of ABCA7 in particular and ABCA transporters in general from which information can be deduced to design novel directly-interacting agents. The four cryo-EM structures of ABCA1 [96] and ABCA4 [97], [98], [99] have been reported rather recently in 2017 and 2021, and hence, have not been used for the design of novel agents yet. The already in 2020 announced [95] cryo-EM structure of ABCA7 (PDB ID: 7KQC; PDB status: unreleased deposition withdrawn) is to this date not publicly available, and thus, cannot be used for structural analyses; and (iii) the lack of standardized HTS assays to monitor the function of ABCA7 and to determine the inhibiting (or activating) property of compounds, as this is the case, for example, for the well-studied ABC transporters [38] ABCB1, ABCC1, and ABCG2 [154]. Assays using labelled tracers of ABCA transporter (particularly ABCA1) function, mostly NBD- [155] or BODIPY-labeled [156] cholesterol, indeed exist, however, have to this day not led to small-molecule inhibitors or activators. One reason is that these assays have primarily been developed to evaluate inducers of ABCA1 [155]. Another reason may be that the used tracers (NBD- [155] or BODIPY-cholesterol [156]) are derivatives of the rather unspecific and ubiquitously present intrinsic substrate cholesterol. Taken together, these three obstacles hinder the targeted development of selective and potent agents for under-studied ABC transporters involved in human diseases, such as ABCA7. However, the present study in combination with recent advances provides an initial hypothesis and further evaluation of these three stated problems.
First of all, computer-aided pattern analysis (‘C@PA’) has very recently been invented to rationally discover and design novel multitargeting pan-ABC transporter inhibitors to obtain potential ligands for under-studied ABC transporters [38], [44], [45]. Hereby, compounds 11–16 [38], [45] have been discovered as moderately potent class 7 [38], [44], [45] and semi class 7 (IC50 ≤ 15 µM) [56] molecules. These molecules represent a novel generation of broad-spectrum ligands of the well-studied ABC transporters [38] ABCB1, ABCC1, and ABCG2. These pan-ABC transporter inhibitors have not been evaluated toward other transporters yet, however, might be the key as transition molecules for the generation of selective and potent agents for under-studied ABC transporters. A common binding site of pan-ABC transporter inhibitors has been proposed [4], [38], [44], [45], which was focused in the present study and is subject to current research.
Second, to explore this potential multitarget binding site, the complete available structural information of human ABCA1 [96] and ABCA4 [98], [99] has been used and extended for the first time to the target of interest of the present study, ABCA7. The generated homology model of ABCA7 could be applied for blind docking experiments using diverse truly multitarget [69], [70], [73], [74], [75], [76], [77], [78], [79], [80], [81], [82], [83], [84], [85], [86], [87], [88], [89], [90], [91], [92], [93], [94] as well as focused [38], [45], [101], [130], [131], [132], [133], [134], [135], [136], [137], [138], [139], [140] pan-ABC transporter inhibitors with two different docking tools, Autodock [129] and Glide [141], [142]. Supported by pharmacophore modelling, the present report suggests that 16 (9, 11–18, 21–27) [38], [45], [101], [130], [133], [134], [135], [136], [137], [138]) of the 23 compounds had consensus features that could be attributed to a common or partially overlapping multitarget binding site. These features F1–F4 (Fig. 6) may be essential to interact with the amino acids in the binding site, specifically cysteine 659, leucin 662, phenylalanine 1544, valine 1649, threonine 1652, and cysteine 1653 (Figs. 10 and 11 as well as Supplementary Fig. S1). The discovery of the consensus features amongst these structurally diverse pan-ABC transporter inhibitors 9, 11–18, 21–27 [38], [45], [101], [130], [133], [134], [135], [136], [137], [138]) in two different in silico tools and systematic cross-validation strongly supports the postulated multitarget binding site amongst ABC transporters of different sub-families, which may generally overlap with the binding site of ‘regular’ substrates of the respective transporter (Supplementary Fig. S19).
Amongst these 16 molecules is the truly-multitarget inhibitor 9, which targets 9 different ABC transporters of 4 different sub-families. [69], [70], [76], [85], [86], [87], [88], [89], [90]. Compound 9 is of particular interest, not only due to its truly multitargeting nature in terms of ABC transporters in general, but also because it was demonstrated to activate ABCC1-mediated transport of the ABCC1 substrate glutathione (GSH) [69]. Defective ABCA7 calls for modulators that increase its transport capability. Generally, activators of ABC transporters have been found before [47], [48], [49], [51], [52], [53], [54], [55]. However, the particular association of an ABC transporter activator with pan-ABC transporter inhibition opens a new perspective on multitargeting in general. Besides inhibitory activities and physicochemical properties, the mode of modulation, in particular the activating nature of compounds, can supplement the newly invented C@PA [38], [44], [45] methodology, expanding bioactivity space of small-molecules and associating their molecular-structural patterns with increase of functional ABC transporter activity. Amongst the known focused pan-ABC transporter inhibitors [38], [45], [130], [151], [152], molecular-structural patterns can be found that have already been associated with activation of ABC transporters, such as tetrahydroisoquinoline [48], pyrrolopyrimidine [49], and purine [49].
While these aspects above were mostly relevant for therapeutic drug development, the discovery of inhibitors of ABCA7 by addressing the multitarget binding site and their simultaneous derivatization to selective agents would allow for in vivo imaging of ABCA7. Imaging of ABC transporters of the BBB has already been accomplished before [63], and the results of the present study found the basis for PET tracer development of other, yet under-studied ABC transporters involved in neurological disorders like AD, as for example, ABCA7.
The results presented in this report have broad implications for addressing other under-studied ABC transporters [38] as well as their exploration and exploitation as potential pharmacological drug targets to tackle major human diseases. Specifically the combination of both ligand-based pattern analysis and structure-based docking studies may provide novel, innovative, and effective drugs for the curation of human diseases in general.
4. Experimental section
4.1. Sequence alignment and homology model
The sequences of the human ABC transporters ABCA1 (UniProt ID: O95477), ABCA4 (UniProt ID: P78363), and ABCA7 (Uniprot ID: Q8IZY2) were downloaded from Uniprot Knowledgebase [157]. The sequence identity and similarity between the three ABCA transporter subtypes were analyzed using Align/Superpose module implemented in Molecular Operating Environment (MOE) 2019.01 [118]. The sequence similarity of ABCA7 was 52.8% and 48.2% compared to ABCA1 and ABCA4, respectively. The cryo-EM structures of ABCA1 (PDB ID: 5XJY [96]) and ABCA4 (PDB ID: 7E7I [98]; PDB ID: 7LKP [99]; PDB ID: 7M1Q [97]) were downloaded from the RCSB protein data bank (PDB) [158]. For homology modelling purposes, the ABCA1 cryo-EM structure was selected due to its functionally and structurally closer similarity to ABCA7. As an initial step, the co-crystallized sugar molecules of ABCA1 have been removed. Ionization and hydrogen atoms were added using the Protonate-3D tool implemented in MOE 2019.01 [118], and the structures were subsequently energy-minimized by keeping the heavy atoms fixed at their crystallographic positions applying the Amber99 force field [159] until the root-mean-square of the conjugate gradient was less than 0.05 kcal·mol−1 · Å−1. A total of 250 models were generated using the GB/VI scoring function with a gradient limit of 0.5.
4.2. Molecule preparation and conformer generation
All 2D representation of the molecules were drawn using ChemDraw version 20.1.1. and optimized using the builder module implemented in MOE 2019.01. The hydrogen atoms were added, energy minimized by applying the AMBER10:EHT forcefield. The conformers of these molecules were generated using the conformer generation tool by selecting the stochastic search method implemented in MOE 2019.01 [118]. The default parameters were applied for the conformational search with a maximum limit of 10,000. The lowest energy conformer obtained were selected for further preparation in the respective docking tools. Finally for the 13 pan-ABC transporter inhibitors 6–8, 12–13, 15–16, 18–21, and 27–28 [45], [130], [131], [132], [133], [138], [139], [140] selected for validation the conformers generated were utilized for the analysis. In total, 2,445 conformers of the 13 molecules were generated.
4.3. Molecular docking
4.3.1. PSO@AutoDock
The generated homology model of the human ABCA7 was used for the docking procedure applying the particle swarm optimization (PSO) tool, PSO@AutoDock [129], implemented in AutoDock 4.2 [160]. The AutoDockTools package was employed to generate the docking input files and to analyze the docking results. The search algorithm, varCPSO-ls [129], from PSO@AutoDock [129] implemented in AutoDock4.2 [160] was employed for docking calculations. Three-dimensional energy scoring grids of 0.375 Å resolution and a dimension of 120 Å × 120 Å × 120 Å were computed. The grids were centered based on the two MSDs of the ABCA7 homology model. The parameters of the varCPSO-ls algorithm [129], cognitive coefficient (c1) and social coefficient (c2), were set to 6.05 and the swarm size as 60 individual particles. The default values were applied for the remaining parameters of the algorithm. A total of 50 independent docking calculations were performed by setting the termination criteria as 50,000 evaluation steps. The docked poses obtained from docking studies for the selected compounds were explored by visual inspection and the putative binding poses were selected.
4.3.2. Glide
As a in initial step, the homology model of human ABCA7 was prepared using the Protein Preparation Wizard module implemented in Schrödinger 2020–3 [143]. The preparation of the protein included three steps: (i) assigning bond orders; (ii) adding the missing hydrogen atoms; (iii) and generating het states with the support of Epik module, a tool which can predict the protonation state of the protein structure at the physiological pH of 7.4. Then in the next step, assignment of H-bond and optimization by using the PROPKA module implement in Schrödinger 2020–3. Finally, the protein is minimized by setrestraining the heavy atoms with a maximum RMSD value of 0.30 Å using the Liquid Simulations Version 3 (OPLS3) force field [161].
The selected ligands were prepared using LigPrep module implemented in Schrödinger 2020–3 [143] and were optimized with the OPLS force field [161]. The grid files required for the docking procedure were prepared using the Receptor Grid Generation panel of Glide [141], [142] with grid points calculated enclosing a box covering the entire transmembrane region similar to PSO@AutoDock [129]. After grid preparation, the selected molecules were docked into the generated transporter grids using Glide XP (extra precision) docking. For the docking procedure, the default parameter settings were applied. A total of 10 independent docking calculations were performed.
4.4. Pharmacophore modelling
The top ranking docking poses of the ten selected molecules from PSO@AutoDock [129] were considered for generating the pharmacophore model. The pharmacophore model was established using the pharmacophore elucidator module implemented in MOE 2019.01 [118]. In this module, the pharmacophore features were automatically calculated using the consensus pharmacophore function. This function clusters the features into potential pharmacophore features which are more conserved than a tolerance and threshold distance of 1.5 Å and the threshold of 30%, respectively.
CRediT authorship contribution statement
Vigneshwaran Namasivayam: Conceptualization, Methodology, Validation, Formal analysis, Investigation, Data curation, Writing – original draft, Writing – review & editing, Visualization, Project administration. Katja Stefan: Writing – review & editing, Funding acquisition. Jens Pahnke: Resources, Writing – review & editing, Project administration, Funding acquisition. Sven Marcel Stefan: Conceptualization, Data curation, Writing – original draft, Writing – review & editing, Visualization, Project administration, Funding acquisition.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
Acknowledgements
KS receives a Walter Benjamin fellowship from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation, Germany; 466106904).
JP received funding from the DFG (263024513); EFRE und Ministerium für Wirtschaft, Wissenschaft und Digitalisierung Sachsen-Anhalt (Germany; ZS/2016/05/78617; Latvian Council of Science (Latvia; lzp-2018/1-0275); Nasjonalforeningen (Norway; 16154), HelseSØ (Norway; 2016062, 2019054, 2019055); Barnekreftforeningen (Norway; 19008); EEA grant/Norway grants Kappa programme (Iceland, Liechtenstein, Norway, Czech Republic; TAČR TARIMAD TO100078); Norges forskningsrådet [Norway; 260786 (PROP-AD), 295910 (NAPI), and 327571 (PETABC)].
PROP-AD (2016–2021) and PETABC (2021–2024) are EU Joint Programme - Neurodegenerative Disease Research (JPND) projects. PROP-AD was supported through the following funding organizations under the aegis of JPND – www.jpnd.eu: AKA #301228 – Finland, BMBF #01ED1605 – Germany; CSO-MOH #30000-12631 – Israel; NFR #260786 – Norway; SRC #2015-06795 – Sweden). PETABC is supported through the following funding organizations under the aegis of JPND – www.jpnd.eu: NFR #327571 – Norway; FFG #882717 – Austria; BMBF #01ED2106 – Germany; MSMT #8F21002 – Czech Republic; VIAA #ES RTD/2020/26 – Latvia; ANR #20-JPW2-0002-04 – France, SRC #2020-02905 – Sweden. The projects receive funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement #643417 (JPco-fuND).
SMS receives a Walter Benjamin fellowship from the DFG (446812474).
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.csbj.2021.11.035.
Contributor Information
Jens Pahnke, Email: jens.pahnke@medisin.uio.no, http://www.pahnkelab.eu.
Sven Marcel Stefan, Email: s.m.stefan@medisin.uio.no.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- 1.Abdallah I.M., Al-Shami K.M., Yang E., Kaddoumi A. Blood-brain barrier disruption increases amyloid-related pathology in TgSwDI mice. Int J Mol Sci. 2021;22:1231. doi: 10.3390/ijms22031231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Gomez-Zepeda D., Taghi M., Scherrmann J.M., Decleves X., Menet M.C. ABC Transporters at the Blood-Brain Interfaces, Their Study Models, and Drug Delivery Implications in Gliomas. Pharmaceutics. 2019;12:20. doi: 10.3390/pharmaceutics12010020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Szakacs G., Abele R. An inventory of lysosomal ABC transporters. FEBS Lett. 2020;594:3965–3985. doi: 10.1002/1873-3468.13967. [DOI] [PubMed] [Google Scholar]
- 4.Stefan K., Leck L.Y.W., Namasivayam V., Bascuñana P., Huang M.L.-H., Riss P.J., Pahnke J., Jansson P.J., Stefan S.M. Vesicular ATP-binding cassette transporters in human disease: relevant aspects of their organization for future drug development. Future Drug Discovery. 2020;2:FDD51. [Google Scholar]
- 5.Stefan S.M., Jansson P.J., Kalinowski D.S., Anjum R., Dharmasivam M., Richardson D.R. The growing evidence for targeting P-glycoprotein in lysosomes to overcome resistance. Future Med Chem. 2020;12:473–477. doi: 10.4155/fmc-2019-0350. [DOI] [PubMed] [Google Scholar]
- 6.Jacobo-Albavera L., Domínguez-Pérez M., Medina-Leyte D.J., González-Garrido A., Villarreal-Molina T. The Role of the ATP-Binding Cassette A1 (ABCA1) in Human Disease. Int J Mol Sci. 2021;22:1593. doi: 10.3390/ijms22041593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wu A., Wojtowicz K., Savary S., Hamon Y., Trombik T. Do ABC transporters regulate plasma membrane organization? Cell Mol Biol Lett. 2020;25:37. doi: 10.1186/s11658-020-00224-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Poejo J., Salazar J., Mata A.M., Gutierrez-Merino C. Binding of Amyloid β(1–42)-Calmodulin Complexes to Plasma Membrane Lipid Rafts in Cerebellar Granule Neurons Alters Resting Cytosolic Calcium Homeostasis. Int J Mol Sci. 2021;22 doi: 10.3390/ijms22041984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cho Y.Y., Kwon O.H., Chung S. Preferred endocytosis of amyloid precursor protein from cholesterol-enriched lipid raft microdomains. Molecules. 2020;25 doi: 10.3390/molecules25235490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Fantini J., Chahinian H., Yahi N. Progress toward Alzheimer's disease treatment: Leveraging the Achilles' heel of Aβ oligomers? Protein Sci. 2020;29:1748–1759. doi: 10.1002/pro.3906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kerr I.D., Hutchison E., Gerard L., Aleidi S.M., Gelissen I.C. Mammalian ABCG-transporters, sterols and lipids: To bind perchance to transport? Biochim Biophys Acta Mol Cell Biol Lipids. 2021;1866 doi: 10.1016/j.bbalip.2020.158860. [DOI] [PubMed] [Google Scholar]
- 12.Pasello M., Giudice A.M., Scotlandi K. The ABC subfamily A transporters: Multifaceted players with incipient potentialities in cancer. Semin Cancer Biol. 2020;60:57–71. doi: 10.1016/j.semcancer.2019.10.004. [DOI] [PubMed] [Google Scholar]
- 13.Dib S., Pahnke J., Gosselet F. Role of ABCA7 in human health and in Alzheimer's disease. Int J Mol Sci. 2021;22 doi: 10.3390/ijms22094603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Agarwal M., Khan S. Plasma lipids as biomarkers for Alzheimer's disease: a systematic review. Cureus. 2020;12 doi: 10.7759/cureus.12008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lyssenko N.N., Praticò D. ABCA7 and the altered lipidostasis hypothesis of Alzheimer's disease. Alzheimers Dement. 2021;17:164–174. doi: 10.1002/alz.12220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Behl T., Kaur I., Sehgal A., Kumar A., Uddin M.S., Bungau S. The interplay of ABC transporters in Aβ translocation and cholesterol metabolism: implicating their roles in Alzheimer's disease. Mol Neurobiol. 2021;58:1564–1582. doi: 10.1007/s12035-020-02211-x. [DOI] [PubMed] [Google Scholar]
- 17.Abe-Dohmae S., Yokoyama S. ABCA7 links sterol metabolism to the host defense system: Molecular background for potential management measure of Alzheimer's disease. Gene. 2021;768 doi: 10.1016/j.gene.2020.145316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Picard C., Julien C., Frappier J., Miron J., Théroux L., Dea D., et al. Alterations in cholesterol metabolism-related genes in sporadic Alzheimer's disease. Neurobiol Aging. 2018;66(180):e181–180.e189. doi: 10.1016/j.neurobiolaging.2018.01.018. [DOI] [PubMed] [Google Scholar]
- 19.Piehler A.P., Ozcürümez M., Kaminski W.E. A-Subclass ATP-Binding Cassette Proteins in Brain Lipid Homeostasis and Neurodegeneration. Front Psychiatry. 2012;3:17. doi: 10.3389/fpsyt.2012.00017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Beel A.J., Sakakura M., Barrett P.J., Sanders C.R. Direct binding of cholesterol to the amyloid precursor protein: An important interaction in lipid-Alzheimer's disease relationships? Biochim Biophys Acta. 1801;2010:975–982. doi: 10.1016/j.bbalip.2010.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shobab L.A., Hsiung G.Y., Feldman H.H. Cholesterol in Alzheimer's disease. Lancet Neurol. 2005;4:841–852. doi: 10.1016/S1474-4422(05)70248-9. [DOI] [PubMed] [Google Scholar]
- 22.Pahnke J, Bascuñana P, Brackhan M, Stefan K, Namasivayam V, Koldamova R, Wu J, Möhle L, Stefan SM, Strategies to Gain Novel Alzheimer’s Disease Diagnostics and Therapeutics Using Modulators of ABCA Transporters, Free Neuropathology, accepted November 12, 2021. 10.17879/freeneuropathology-2021-3528. [DOI] [PMC free article] [PubMed]
- 23.Katzeff J.S., Kim W.S. ATP-binding cassette transporters and neurodegenerative diseases. Essays Biochem. 2021 doi: 10.1042/EBC20210012. [DOI] [PubMed] [Google Scholar]
- 24.Surguchev A.A., Surguchov A. ABCA7-A Member of the ABC Transporter Family in Healthy and Ailing Brain. Brain Sci. 2020;10:121. doi: 10.3390/brainsci10020121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pereira C.D., Martins F., Wiltfang J., da Cruz E.S.O.A.B., Rebelo S. ABC transporters are key Players in Alzheimer's disease. J Alzheimers Dis. 2018;61:463–485. doi: 10.3233/JAD-170639. [DOI] [PubMed] [Google Scholar]
- 26.Pahnke J, Langer O, Krohn M, Alzheimer's and ABC transporters--new opportunities for diagnostics and treatment, Neurobiol Dis, 72 Pt A (2014) 54-60. [DOI] [PMC free article] [PubMed]
- 27.Abuznait A.H., Kaddoumi A. Role of ABC transporters in the pathogenesis of Alzheimer's disease. ACS Chem Neurosci. 2012;3:820–831. doi: 10.1021/cn300077c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wolf A., Bauer B., Hartz A.M. ABC transporters and the Alzheimer's disease enigma. Front Psychiatry. 2012;3:54. doi: 10.3389/fpsyt.2012.00054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.H. Holstege, M. Hulsman, C. Charbonnier, B. Grenier-Boley, O. Quenez, D. Grozeva, J.G.J. van Rooij, R. Sims, S. Ahmad, N. Amin, P.J. Norsworthy, O. Dols-Icardo, H. Hummerich, A. Kawalia, P. Amouyel, G.W. Beecham, C. Berr, J.C. Bis, A. Boland, P. Bossù, F. Bouwman, D. Campion, A. Daniele, J.-F. Dartigues, S. Debette, J.-F. Deleuze, N. Denning, A.L. DeStefano, L.A. Farrer, N.C. Fox, D. Galimberti, E. Genin, J.L. Haines, C. Holmes, M.A. Ikram, M.K. Ikram, I. Jansen, R. Kraaij, M. Lathrop, E. Lemstra, A. Lleó, L. Luckcuck, R. Marshall, E.R. Martin, C. Masullo, R. Mayeux, P. Mecocci, A. Meggy, M.O. Mol, K. Morgan, B. Nacmia, A.C. Naj, P. Pastor, M.A. Pericak-Vance, R. Raybould, R. Redon, A.-C. Richard, S.G. Riedel-Heller, F. Rivadeneira, S. Rousseau, N.S. Ryan, S. Saad, P. Sanchez-Juan, G.D. Schellenberg, P. Scheltens, J.M. Schott, D. Seripa, G. Spalleta, B. Tijms, A.G. Uitterlinden, S.J. van der Lee, M. Wagner, D. Wallon, L.-S. Wang, A. Zarea, M.J.T. Reinders, J. Clarimon, J.C. van Swieten, J.J. Hardy, A. Ramirez, S. Mead, W.M. van der Flier, C.M. van Duijn, J. Williams, G. Nicolas, C. Bellenguez, J.-C. Lambert, Exome sequencing identifies novel AD-associated genes, medRxiv, (2020) 2020.2007.2022.20159251.
- 30.Andrews S.J., Fulton-Howard B., Goate A. Protective Variants in Alzheimer's Disease. Curr Genet Med Rep. 2019;7:1–12. doi: 10.1007/s40142-019-0156-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Teresa J.C., Fernado C., Nancy M.R., Gilberto V.A., Alberto C.R., Roberto R.R. Association of genetic variants of ABCA1 with susceptibility to dementia: (SADEM study) Metab Brain Dis. 2020;35:915–922. doi: 10.1007/s11011-020-00577-4. [DOI] [PubMed] [Google Scholar]
- 32.Jiang S., Zhang C.Y., Tang L., Zhao L.X., Chen H.Z., Qiu Y. Integrated Genomic Analysis Revealed Associated Genes for Alzheimer's Disease in APOE4 Non-Carriers. Curr Alzheimer Res. 2019;16:753–763. doi: 10.2174/1567205016666190823124724. [DOI] [PubMed] [Google Scholar]
- 33.Chen Q., Liang B., Wang Z., Cheng X., Huang Y., Liu Y., et al. Influence of four polymorphisms in ABCA1 and PTGS2 genes on risk of Alzheimer's disease: a meta-analysis. Neurol Sci. 2016;37:1209–1220. doi: 10.1007/s10072-016-2579-9. [DOI] [PubMed] [Google Scholar]
- 34.Aikawa T., Holm M.L., Kanekiyo T. ABCA7 and Pathogenic Pathways of Alzheimer's Disease. Brain Sci. 2018;8 doi: 10.3390/brainsci8020027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Li H., Karl T., Garner B. Understanding the function of ABCA7 in Alzheimer's disease. Biochem Soc Trans. 2015;43:920–923. doi: 10.1042/BST20150105. [DOI] [PubMed] [Google Scholar]
- 36.Steinberg S., Stefansson H., Jonsson T., Johannsdottir H., Ingason A., Helgason H., et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer's disease. Nat Genet. 2015;47:445–447. doi: 10.1038/ng.3246. [DOI] [PubMed] [Google Scholar]
- 37.Piaceri I., Nacmias B., Sorbi S. Genetics of familial and sporadic Alzheimer's disease. Front Biosci (Elite Ed) 2013;5:167–177. doi: 10.2741/e605. [DOI] [PubMed] [Google Scholar]
- 38.Namasivayam V., Silbermann K., Pahnke J., Wiese M., Stefan S.M. Scaffold fragmentation and substructure hopping reveal potential, robustness, and limits of computer-aided pattern analysis (C@PA) Comput Struct Biotechnol J. 2021;19:3269–3283. doi: 10.1016/j.csbj.2021.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hayashi M., Abe-Dohmae S., Okazaki M., Ueda K., Yokoyama S. Heterogeneity of high density lipoprotein generated by ABCA1 and ABCA7. J Lipid Res. 2005;46:1703–1711. doi: 10.1194/jlr.M500092-JLR200. [DOI] [PubMed] [Google Scholar]
- 40.Tanaka N., Abe-Dohmae S., Iwamoto N., Fitzgerald M.L., Yokoyama S. HMG-CoA reductase inhibitors enhance phagocytosis by upregulating ATP-binding cassette transporter A7. Atherosclerosis. 2011;217:407–414. doi: 10.1016/j.atherosclerosis.2011.06.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Iwamoto N., Abe-Dohmae S., Sato R., Yokoyama S. ABCA7 expression is regulated by cellular cholesterol through the SREBP2 pathway and associated with phagocytosis. J Lipid Res. 2006;47:1915–1927. doi: 10.1194/jlr.M600127-JLR200. [DOI] [PubMed] [Google Scholar]
- 42.Wakaumi M., Ishibashi K., Ando H., Kasanuki H., Tsuruoka S. Acute digoxin loading reduces ABCA8A mRNA expression in the mouse liver. Clin Exp Pharmacol Physiol. 2005;32:1034–1041. doi: 10.1111/j.1440-1681.2005.04301.x. [DOI] [PubMed] [Google Scholar]
- 43.Wu C.A., Wang N., Zhao D.H. An evaluation of the mechanism of ABCA7 on cellular lipid release in ABCA7-HEC293 cell. Chin Med J (Engl) 2013;126:306–310. [PubMed] [Google Scholar]
- 44.Namasivayam V, Silbermann K, Pahnke J, Wiese M, Stefan S.M., Feature-driven Pattern Analysis for Multitarget Modulator Landscapes. , Bioinformatics, accepted December 1, 2021. 10.1093/bioinformatics/btab832. [DOI] [PMC free article] [PubMed]
- 45.Namasivayam V., Silbermann K., Wiese M., Pahnke J., Stefan S.M. C@PA: Computer-Aided Pattern Analysis to Predict Multitarget ABC Transporter Inhibitors. J Med Chem. 2021;64:3350–3366. doi: 10.1021/acs.jmedchem.0c02199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Stefan S.M. Multi-target ABC transporter modulators: what next and where to go?, Future. Med Chem. 2019;11:2353–2358. doi: 10.4155/fmc-2019-0185. [DOI] [PubMed] [Google Scholar]
- 47.Wiese M., Stefan S.M. The A-B-C of small-molecule ABC transport protein modulators: From inhibition to activation-a case study of multidrug resistance-associated protein 1. ABCC1Med Res Rev. 2019;39:2031–2081. doi: 10.1002/med.21573. [DOI] [PubMed] [Google Scholar]
- 48.Contino M., Cantore M., Capparelli E., Perrone M.G., Niso M., Inglese C., et al. A benzopyrane derivative as a P-glycoprotein stimulator: a potential agent to decrease β-amyloid accumulation in Alzheimer's disease. ChemMedChem. 2012;7:391–395. doi: 10.1002/cmdc.201100469. [DOI] [PubMed] [Google Scholar]
- 49.Schmitt S.M., Stefan K., Wiese M. Pyrrolopyrimidine derivatives and purine analogs as novel activators of Multidrug Resistance-associated Protein 1 (MRP1, ABCC1) Biochim Biophys Acta Biomembr. 1859;2017:69–79. doi: 10.1016/j.bbamem.2016.10.017. [DOI] [PubMed] [Google Scholar]
- 50.Krohn M., Lange C., Hofrichter J., Scheffler K., Stenzel J., Steffen J., et al. Cerebral amyloid-β proteostasis is regulated by the membrane transport protein ABCC1 in mice. J Clin Invest. 2011;121:3924–3931. doi: 10.1172/JCI57867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Csandl M.A., Conseil G., Cole S.P. Cysteinyl Leukotriene Receptor 1/2 Antagonists Nonselectively Modulate Organic Anion Transport by Multidrug Resistance Proteins (MRP1-4) Drug Metab Dispos. 2016;44:857–866. doi: 10.1124/dmd.116.069468. [DOI] [PubMed] [Google Scholar]
- 52.Trechot P., Conart J.B., Trechot F. ATP sensitive potassium channel openers: A new class of ocular hypotensive agents. Exp Eye Res. 2019;178:223–224. doi: 10.1016/j.exer.2016.06.014. [DOI] [PubMed] [Google Scholar]
- 53.Wang H., Tang Y., Wang L., Long C.L., Zhang Y.L. ATP-sensitive potassium channel openers and 2,3-dimethyl-2-butylamine derivatives. Curr Med Chem. 2007;14:133–155. doi: 10.2174/092986707779313390. [DOI] [PubMed] [Google Scholar]
- 54.Hansen J.B. Towards selective Kir6.2/SUR1 potassium channel openers, medicinal chemistry and therapeutic perspectives. Curr Med Chem. 2006;13:361–376. doi: 10.2174/092986706775527947. [DOI] [PubMed] [Google Scholar]
- 55.Moreau C., Prost A.L., Dérand R., Vivaudou M. SUR, ABC proteins targeted by KATP channel openers. J Mol Cell Cardiol. 2005;38:951–963. doi: 10.1016/j.yjmcc.2004.11.030. [DOI] [PubMed] [Google Scholar]
- 56.Kinting S., Höppner S., Schindlbeck U., Forstner M.E., Harfst J., Wittmann T., et al. Functional rescue of misfolding ABCA3 mutations by small molecular correctors. Hum Mol Genet. 2018;27:943–953. doi: 10.1093/hmg/ddy011. [DOI] [PubMed] [Google Scholar]
- 57.Liu Q., Sabirzhanova I., Bergbower E.A.S., Yanda M., Guggino W.G., Cebotaru L. The CFTR Corrector, VX-809 (Lumacaftor), Rescues ABCA4 Trafficking Mutants: a Potential Treatment for Stargardt Disease. Cell Physiol Biochem. 2019;53:400–412. doi: 10.33594/000000146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Sabirzhanova I., Lopes Pacheco M., Rapino D., Grover R., Handa J.T., Guggino W.B., et al. Rescuing Trafficking Mutants of the ATP-binding Cassette Protein, ABCA4, with Small Molecule Correctors as a Treatment for Stargardt Eye Disease. J Biol Chem. 2015;290:19743–19755. doi: 10.1074/jbc.M115.647685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Delaunay J.L., Bruneau A., Hoffmann B., Durand-Schneider A.M., Barbu V., Jacquemin E., et al. Functional defect of variants in the adenosine triphosphate-binding sites of ABCB4 and their rescue by the cystic fibrosis transmembrane conductance regulator potentiator, ivacaftor. VX-770Hepatology. 2017;65:560–570. doi: 10.1002/hep.28929. [DOI] [PubMed] [Google Scholar]
- 60.Mareux E., Lapalus M., Amzal R., Almes M., Aït-Slimane T., Delaunay J.L., et al. Functional rescue of an ABCB11 mutant by ivacaftor: A new targeted pharmacotherapy approach in bile salt export pump deficiency. Liver Int. 2020;40:1917–1925. doi: 10.1111/liv.14518. [DOI] [PubMed] [Google Scholar]
- 61.Bardin E., Pastor A., Semeraro M., Golec A., Hayes K., Chevalier B., et al. Modulators of CFTR. Updates on clinical development and future directions. Eur J Med Chem. 2021;213 doi: 10.1016/j.ejmech.2021.113195. [DOI] [PubMed] [Google Scholar]
- 62.Teodori E., Dei S., Bartolucci G., Perrone M.G., Manetti D., Romanelli M.N., et al. Structure-Activity Relationship Studies on 6,7-Dimethoxy-2-phenethyl-1,2,3,4-tetrahydroisoquinoline Derivatives as Multidrug Resistance Reversers. ChemMedChem. 2017;12:1369–1379. doi: 10.1002/cmdc.201700239. [DOI] [PubMed] [Google Scholar]
- 63.Leopoldo M., Contino M., Berardi F., Perrone R., Colabufo N.A. PET radiotracers for imaging P-glycoprotein: the challenge for early diagnosis in AD. ChemMedChem. 2014;9:38–42. doi: 10.1002/cmdc.201300362. [DOI] [PubMed] [Google Scholar]
- 64.Schmitt S.M., Stefan K., Wiese M. Pyrrolopyrimidine Derivatives as Novel Inhibitors of Multidrug Resistance-Associated Protein 1 (MRP1, ABCC1) J Med Chem. 2016;59:3018–3033. doi: 10.1021/acs.jmedchem.5b01644. [DOI] [PubMed] [Google Scholar]
- 65.Wesołowska O. Interaction of phenothiazines, stilbenes and flavonoids with multidrug resistance-associated transporters, P-glycoprotein and MRP1. Acta Biochim Pol. 2011;58:433–448. [PubMed] [Google Scholar]
- 66.Wesołowska O., Molnar J., Ocsovszki I., Michalak K. Differential effect of phenothiazines on MRP1 and P-glycoprotein activity. In Vivo. 2009;23:943–947. [PubMed] [Google Scholar]
- 67.Namanja H.A., Emmert D., Hrycyna C.A., Chmielewski J. Homodimers of the Antiviral Abacavir as Modulators of P-glycoprotein Transport in Cell Culture: Probing Tether Length. Medchemcomm. 2013;4:1344–1349. doi: 10.1039/C3MD00196B. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Dhainaut A., Regnier G., Tizot A., Pierre A., Leonce S., Guilbaud N., et al. New purines and purine analogs as modulators of multidrug resistance. J Med Chem. 1996;39:4099–4108. doi: 10.1021/jm960361i. [DOI] [PubMed] [Google Scholar]
- 69.Stefan S.M., Wiese M. Small-molecule inhibitors of multidrug resistance-associated protein 1 and related processes: A historic approach and recent advances. Med Res Rev. 2019;39:176–264. doi: 10.1002/med.21510. [DOI] [PubMed] [Google Scholar]
- 70.Zhang H., Xu H., Ashby C.R., Jr., Assaraf Y.G., Chen Z.S., Liu H.M. Chemical molecular-based approach to overcome multidrug resistance in cancer by targeting P-glycoprotein. P-gpMed Res Rev. 2021;41:525–555. doi: 10.1002/med.21739. [DOI] [PubMed] [Google Scholar]
- 71.Peña-Solórzano D., Stark S.A., König B., Sierra C.A., Ochoa-Puentes C. ABCG2/BCRP: Specific and Nonspecific Modulators. Med Res Rev. 2017;37:987–1050. doi: 10.1002/med.21428. [DOI] [PubMed] [Google Scholar]
- 72.Palmeira A., Sousa E., Vasconcelos M.H., Pinto M.M. Three decades of P-gp inhibitors: skimming through several generations and scaffolds. Curr Med Chem. 2012;19:1946–2025. doi: 10.2174/092986712800167392. [DOI] [PubMed] [Google Scholar]
- 73.Zhou S.F., Wang L.L., Di Y.M., Xue C.C., Duan W., Li C.G., et al. Substrates and inhibitors of human multidrug resistance associated proteins and the implications in drug development. Curr Med Chem. 2008;15:1981–2039. doi: 10.2174/092986708785132870. [DOI] [PubMed] [Google Scholar]
- 74.Ivnitski-Steele I., Larson R.S., Lovato D.M., Khawaja H.M., Winter S.S., Oprea T.I., et al. High-throughput flow cytometry to detect selective inhibitors of ABCB1, ABCC1, and ABCG2 transporters. Assay Drug Dev Technol. 2008;6:263–276. doi: 10.1089/adt.2007.107. [DOI] [PubMed] [Google Scholar]
- 75.Pedersen J.M., Matsson P., Bergström C.A., Hoogstraate J., Norén A., LeCluyse E.L., et al. Early identification of clinically relevant drug interactions with the human bile salt export pump. BSEP/ABCB11Toxicol Sci. 2013;136:328–343. doi: 10.1093/toxsci/kft197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Matsson P., Pedersen J.M., Norinder U., Bergström C.A., Artursson P. Identification of novel specific and general inhibitors of the three major human ATP-binding cassette transporters P-gp, BCRP and MRP2 among registered drugs. Pharm Res. 2009;26:1816–1831. doi: 10.1007/s11095-009-9896-0. [DOI] [PubMed] [Google Scholar]
- 77.El-Sheikh A.A., van den Heuvel J.J., Koenderink J.B., Russel F.G. Effect of hypouricaemic and hyperuricaemic drugs on the renal urate efflux transporter, multidrug resistance protein 4. Br J Pharmacol. 2008;155:1066–1075. doi: 10.1038/bjp.2008.343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Zelcer N., Saeki T., Reid G., Beijnen J.H., Borst P. Characterization of drug transport by the human multidrug resistance protein 3. ABCC3J Biol Chem. 2001;276:46400–46407. doi: 10.1074/jbc.M107041200. [DOI] [PubMed] [Google Scholar]
- 79.Hupfeld T., Chapuy B., Schrader V., Beutler M., Veltkamp C., Koch R., et al. Tyrosinekinase inhibition facilitates cooperation of transcription factor SALL4 and ABC transporter A3 towards intrinsic CML cell drug resistance. Br J Haematol. 2013;161:204–213. doi: 10.1111/bjh.12246. [DOI] [PubMed] [Google Scholar]
- 80.Beretta G.L., Cassinelli G., Pennati M., Zuco V., Gatti L. Overcoming ABC transporter-mediated multidrug resistance: The dual role of tyrosine kinase inhibitors as multitargeting agents. Eur J Med Chem. 2017;142:271–289. doi: 10.1016/j.ejmech.2017.07.062. [DOI] [PubMed] [Google Scholar]
- 81.Tanaka N., Yamaguchi H., Mano N. Transport of eicosapentaenoic acid-derived PGE3, PGF(3α), and TXB3 by ABCC4. PLoS ONE. 2014;9 doi: 10.1371/journal.pone.0109270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Wu C.P., Calcagno A.M., Hladky S.B., Ambudkar S.V., Barrand M.A. Modulatory effects of plant phenols on human multidrug-resistance proteins 1, 4 and 5. ABCC1, 4 and 5Febs j. 2005;272:4725–4740. doi: 10.1111/j.1742-4658.2005.04888.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Saito H., Toyoda Y., Hirata H., Ota-Kontani A., Tsuchiya Y., Takada T., et al. Soy Isoflavone Genistein Inhibits an Axillary Osmidrosis Risk Factor ABCC11. In Vitro Screening and Fractional Approach for ABCC11-Inhibitory Activities in Plant Extracts and Dietary Flavonoids. Nutrients. 2020;12 doi: 10.3390/nu12082452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.BoseDasgupta S., Ganguly A., Roy A., Mukherjee T., Majumder H.K. A novel ATP-binding cassette transporter, ABCG6 is involved in chemoresistance of Leishmania. Mol Biochem Parasitol. 2008;158:176–188. doi: 10.1016/j.molbiopara.2007.12.007. [DOI] [PubMed] [Google Scholar]
- 85.Tsuruoka S., Ishibashi K., Yamamoto H., Wakaumi M., Suzuki M., Schwartz G.J., et al. Functional analysis of ABCA8, a new drug transporter. Biochem Biophys Res Commun. 2002;298:41–45. doi: 10.1016/s0006-291x(02)02389-6. [DOI] [PubMed] [Google Scholar]
- 86.Bieczynski F., Burkhardt-Medicke K., Luquet C.M., Scholz S., Luckenbach T. Chemical effects on dye efflux activity in live zebrafish embryos and on zebrafish Abcb4 ATPase activity. FEBS Lett. 2021;595:828–843. doi: 10.1002/1873-3468.14015. [DOI] [PubMed] [Google Scholar]
- 87.Saeed M.E.M., Boulos J.C., Elhaboub G., Rigano D., Saab A., Loizzo M.R., et al. Cytotoxicity of cucurbitacin E from Citrullus colocynthis against multidrug-resistant cancer cells. Phytomedicine. 2019;62 doi: 10.1016/j.phymed.2019.152945. [DOI] [PubMed] [Google Scholar]
- 88.Horikawa M., Kato Y., Tyson C.A., Sugiyama Y. Potential cholestatic activity of various therapeutic agents assessed by bile canalicular membrane vesicles isolated from rats and humans. Drug Metab Pharmacokinet. 2003;18:16–22. doi: 10.2133/dmpk.18.16. [DOI] [PubMed] [Google Scholar]
- 89.Bai J., Lai L., Yeo H.C., Goh B.C., Tan T.M. Multidrug resistance protein 4 (MRP4/ABCC4) mediates efflux of bimane-glutathione. Int J Biochem Cell Biol. 2004;36:247–257. doi: 10.1016/s1357-2725(03)00236-x. [DOI] [PubMed] [Google Scholar]
- 90.Zhou Y., Hopper-Borge E., Shen T., Huang X.C., Shi Z., Kuang Y.H., et al. Cepharanthine is a potent reversal agent for MRP7(ABCC10)-mediated multidrug resistance. Biochem Pharmacol. 2009;77:993–1001. doi: 10.1016/j.bcp.2008.12.005. [DOI] [PubMed] [Google Scholar]
- 91.Dalpiaz A., Pavan B. Nose-to-Brain Delivery of Antiviral Drugs: A Way to Overcome Their Active Efflux? Pharmaceutics. 2018;10:39. doi: 10.3390/pharmaceutics10020039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Videmann B., Mazallon M., Prouillac C., Delaforge M., Lecoeur S. ABCC1, ABCC2 and ABCC3 are implicated in the transepithelial transport of the myco-estrogen zearalenone and its major metabolites. Toxicol Lett. 2009;190:215–223. doi: 10.1016/j.toxlet.2009.07.021. [DOI] [PubMed] [Google Scholar]
- 93.Borst P., de Wolf C., van de Wetering K. Multidrug resistance-associated proteins 3, 4, and 5. Pflugers Arch. 2007;453:661–673. doi: 10.1007/s00424-006-0054-9. [DOI] [PubMed] [Google Scholar]
- 94.Tun-Yhong W., Chinpaisal C., Pamonsinlapatham P., Kaewkitichai S. Tenofovir Disoproxil Fumarate Is a New Substrate of ATP-Binding Cassette Subfamily C Member 11. Antimicrob Agents Chemother. 2017;61:e01725–01716. doi: 10.1128/AAC.01725-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.L.T. My Le, J.R. Thompson, T. Aikawa, T. Kanikeyo, A. Alam, Cryo-EM structure of lipid embedded human ABCA7 at 3.6Å resolution, bioRxiv, (2021) 2021.2003.2001.433448.
- 96.Qian H., Zhao X., Cao P., Lei J., Yan N., Gong X. Structure of the Human Lipid Exporter ABCA1. Cell. 2017;169:1228–1239.e1210. doi: 10.1016/j.cell.2017.05.020. [DOI] [PubMed] [Google Scholar]
- 97.Scortecci J.F., Molday L.L., Curtis S.B., Garces F.A., Panwar P., Van Petegem F., et al. Cryo-EM structures of the ABCA4 importer reveal mechanisms underlying substrate binding and Stargardt disease. Nat Commun. 2021;12:5902. doi: 10.1038/s41467-021-26161-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Xie T., Zhang Z., Fang Q., Du B., Gong X. Structural basis of substrate recognition and translocation by human ABCA4. Nat Commun. 2021;12:3853. doi: 10.1038/s41467-021-24194-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Liu F., Lee J., Chen J. Molecular structures of the eukaryotic retinal importer ABCA4. Elife. 2021;10 doi: 10.7554/eLife.63524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Kashgari F.K., Ravna A., Sager G., Lyså R., Enyedy I., Dietrichs E.S. Identification and experimental confirmation of novel cGMP efflux inhibitors by virtual ligand screening of vardenafil-analogues. Biomed Pharmacother. 2020;126 doi: 10.1016/j.biopha.2020.110109. [DOI] [PubMed] [Google Scholar]
- 101.Silbermann K., Li J., Namasivayam V., Stefan S.M., Wiese M. Rational drug design of 6-substituted 4-anilino-2-phenylpyrimidines for exploration of novel ABCG2 binding site. Eur J Med Chem. 2021;212 doi: 10.1016/j.ejmech.2020.113045. [DOI] [PubMed] [Google Scholar]
- 102.Alam A., Kowal J., Broude E., Roninson I., Locher K.P. Structural insight into substrate and inhibitor discrimination by human P-glycoprotein. Science. 2019;363:753–756. doi: 10.1126/science.aav7102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Oldham M.L., Grigorieff N., Chen J. Structure of the transporter associated with antigen processing trapped by herpes simplex virus. Elife. 2016;5 doi: 10.7554/eLife.21829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Olsen J.A., Alam A., Kowal J., Stieger B., Locher K.P. Structure of the human lipid exporter ABCB4 in a lipid environment. Nat Struct Mol Biol. 2020;27:62–70. doi: 10.1038/s41594-019-0354-3. [DOI] [PubMed] [Google Scholar]
- 105.Martin G.M., Sung M.W., Yang Z., Innes L.M., Kandasamy B., David L.L., et al. Mechanism of pharmacochaperoning in a mammalian K(ATP) channel revealed by cryo-EM. Elife. 2019;8 doi: 10.7554/eLife.46417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Kowal J., Ni D., Jackson S.M., Manolaridis I., Stahlberg H., Locher K.P. Structural Basis of Drug Recognition by the Multidrug Transporter ABCG2. J Mol Biol. 2021;433 doi: 10.1016/j.jmb.2021.166980. [DOI] [PubMed] [Google Scholar]
- 107.Jeevitha Priya M., Vidyalakshmi S., Rajeswari M. Study on reversal of ABCB1 mediated multidrug resistance in Colon cancer by acetogenins: An in-silico approach. J Biomol Struct Dyn. 2020:1–12. doi: 10.1080/07391102.2020.1855249. [DOI] [PubMed] [Google Scholar]
- 108.Corradi V., Singh G., Tieleman D.P. The human transporter associated with antigen processing: molecular models to describe peptide binding competent states. J Biol Chem. 2012;287:28099–28111. doi: 10.1074/jbc.M112.381251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Tangella L.P., Arooj M., Deplazes E., Gray E.S., Mancera R.L. Identification and characterisation of putative drug binding sites in human ATP-binding cassette B5 (ABCB5) transporter, Comput Struct. Biotechnol J. 2021;19:691–704. doi: 10.1016/j.csbj.2020.12.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Conseil G., Arama-Chayoth M., Tsfadia Y., Cole S.P.C. Structure-guided probing of the leukotriene C(4) binding site in human multidrug resistance protein 1 (MRP1; ABCC1) Faseb j. 2019;33:10692–10704. doi: 10.1096/fj.201900140R. [DOI] [PubMed] [Google Scholar]
- 111.Becerra E., Aguilera-Durán G., Berumen L., Romo-Mancillas A., García-Alcocer G. Study of Endogen Substrates, Drug Substrates and Inhibitors Binding Conformations on MRP4 and Its Variants by Molecular Docking and Molecular Dynamics. Molecules. 2021;26:1051. doi: 10.3390/molecules26041051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Szeri F., Corradi V., Niaziorimi F., Donnelly S., Conseil G., Cole S.P.C., et al. Mutagenic Analysis of the Putative ABCC6 Substrate-Binding Cavity Using a New Homology Model. Int J Mol Sci. 2021;22:6910. doi: 10.3390/ijms22136910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Yeh H.I., Qiu L., Sohma Y., Conrath K., Zou X., Hwang T.C. Identifying the molecular target sites for CFTR potentiators GLPG1837 and VX-770. J Gen Physiol. 2019;151:912–928. doi: 10.1085/jgp.201912360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Honorat M., Terreux R., Falson P., Di Pietro A., Dumontet C., Payen L. Localization of putative binding sites for cyclic guanosine monophosphate and the anti-cancer drug 5-fluoro-2'-deoxyuridine-5'-monophosphate on ABCC11 in silico models. BMC Struct Biol. 2013;13:7. doi: 10.1186/1472-6807-13-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Kaminski W.E., Orsó E., Diederich W., Klucken J., Drobnik W., Schmitz G. Identification of a novel human sterol-sensitive ATP-binding cassette transporter. ABCA7Biochem Biophys Res Commun. 2000;273:532–538. doi: 10.1006/bbrc.2000.2954. [DOI] [PubMed] [Google Scholar]
- 116.Vihinen M, Guidelines for reporting protein modelling studies, Authorea, (2021) Peer review.
- 117.Lee J.Y., Kinch L.N., Borek D.M., Wang J., Wang J., Urbatsch I.L., et al. Crystal structure of the human sterol transporter ABCG5/ABCG8. Nature. 2016;533:561–564. doi: 10.1038/nature17666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Molecular Operating Environment (MOE) West; Suite #910, Montreal, QC, Canada: 2019.01;. Chemical Computing Group ULC, 1010 Sherbooke St. [Google Scholar]
- 119.Oswald C., Holland I.B., Schmitt L. The motor domains of ABC-transporters. What can structures tell us? Naunyn Schmiedebergs Arch Pharmacol. 2006;372:385–399. doi: 10.1007/s00210-005-0031-4. [DOI] [PubMed] [Google Scholar]
- 120.Schmitt L., Tampé R. Affinity, specificity, diversity: a challenge for the ABC transporter TAP in cellular immunity. ChemBioChem. 2000;1:16–35. doi: 10.1002/1439-7633(20000703)1:1<16::AID-CBIC16>3.0.CO;2-Y. [DOI] [PubMed] [Google Scholar]
- 121.Fitzgerald M.L., Morris A.L., Rhee J.S., Andersson L.P., Mendez A.J., Freeman M.W. Naturally occurring mutations in the largest extracellular loops of ABCA1 can disrupt its direct interaction with apolipoprotein A-I. J Biol Chem. 2002;277:33178–33187. doi: 10.1074/jbc.M204996200. [DOI] [PubMed] [Google Scholar]
- 122.Wang C., Cao C., Wang N., Wang X., Wang X., Zhang X.C. Cryo-electron microscopy structure of human ABCB6 transporter. Protein Sci. 2020;29:2363–2374. doi: 10.1002/pro.3960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Wang L., Hou W.T., Chen L., Jiang Y.L., Xu D., Sun L., et al. Cryo-EM structure of human bile salts exporter ABCB11. Cell Res. 2020;30:623–625. doi: 10.1038/s41422-020-0302-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Liu F., Zhang Z., Csanády L., Gadsby D.C., Chen J. Molecular Structure of the Human CFTR Ion Channel. Cell. 2017;169:85–95.e88. doi: 10.1016/j.cell.2017.02.024. [DOI] [PubMed] [Google Scholar]
- 125.Zhang H., Huang C.S., Yu X., Lee J., Vaish A., Chen Q., et al. Cryo-EM structure of ABCG5/G8 in complex with modulating antibodies. Commun Biol. 2021;4:526. doi: 10.1038/s42003-021-02039-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Li S., Ren Y., Lu X., Shen Y., Yang X. Cryo-EM structure of human ABCB8 transporter in nucleotide binding state. Biochem Biophys Res Commun. 2021;557:187–191. doi: 10.1016/j.bbrc.2021.04.007. [DOI] [PubMed] [Google Scholar]
- 127.Shintre C.A., Pike A.C., Li Q., Kim J.I., Barr A.J., Goubin S., et al. Structures of ABCB10, a human ATP-binding cassette transporter in apo- and nucleotide-bound states. Proc Natl Acad Sci U S A. 2013;110:9710–9715. doi: 10.1073/pnas.1217042110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Xu D., Feng Z., Hou W.T., Jiang Y.L., Wang L., Sun L., et al. Cryo-EM structure of human lysosomal cobalamin exporter ABCD4. Cell Res. 2019;29:1039–1041. doi: 10.1038/s41422-019-0222-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Namasivayam V., Günther R. pso@autodock: a fast flexible molecular docking program based on Swarm intelligence. Chem Biol Drug Des. 2007;70:475–484. doi: 10.1111/j.1747-0285.2007.00588.x. [DOI] [PubMed] [Google Scholar]
- 130.Antoni F., Wifling D., Bernhardt G. Water-soluble inhibitors of ABCG2 (BCRP) - A fragment-based and computational approach. Eur J Med Chem. 2021;210 doi: 10.1016/j.ejmech.2020.112958. [DOI] [PubMed] [Google Scholar]
- 131.Silbermann K., Li J., Namasivayam V., Baltes F., Bendas G., Stefan S.M., et al. Superior Pyrimidine Derivatives as Selective ABCG2 Inhibitors and Broad-Spectrum ABCB1, ABCC1, and ABCG2 Antagonists. J Med Chem. 2020;63:10412–10432. doi: 10.1021/acs.jmedchem.0c00961. [DOI] [PubMed] [Google Scholar]
- 132.Teodori E., Contino M., Riganti C., Bartolucci G., Braconi L., Manetti D., et al. Design, synthesis and biological evaluation of stereo- and regioisomers of amino aryl esters as multidrug resistance (MDR) reversers. Eur J Med Chem. 2019;182 doi: 10.1016/j.ejmech.2019.111655. [DOI] [PubMed] [Google Scholar]
- 133.Silbermann K., Stefan S.M., Elshawadfy R., Namasivayam V., Wiese M. Identification of Thienopyrimidine Scaffold as an Inhibitor of the ABC Transport Protein ABCC1 (MRP1) and Related Transporters Using a Combined Virtual Screening Approach. J Med Chem. 2019;62:4383–4400. doi: 10.1021/acs.jmedchem.8b01821. [DOI] [PubMed] [Google Scholar]
- 134.Stefan K., Schmitt S.M., Wiese M. 9-Deazapurines as Broad-Spectrum Inhibitors of the ABC Transport Proteins P-Glycoprotein, Multidrug Resistance-Associated Protein 1, and Breast Cancer Resistance Protein. J Med Chem. 2017;60:8758–8780. doi: 10.1021/acs.jmedchem.7b00788. [DOI] [PubMed] [Google Scholar]
- 135.Krapf M.K., Gallus J., Wiese M. Synthesis and biological investigation of 2,4-substituted quinazolines as highly potent inhibitors of breast cancer resistance protein. ABCG2Eur J Med Chem. 2017;139:587–611. doi: 10.1016/j.ejmech.2017.08.020. [DOI] [PubMed] [Google Scholar]
- 136.Krapf M.K., Wiese M. Synthesis and Biological Evaluation of 4-Anilino-quinazolines and -quinolines as Inhibitors of Breast Cancer Resistance Protein. ABCG2J Med Chem. 2016;59:5449–5461. doi: 10.1021/acs.jmedchem.6b00330. [DOI] [PubMed] [Google Scholar]
- 137.Krauze A., Grinberga S., Krasnova L., Adlere I., Sokolova E., Domracheva I., et al. Thieno[2,3-b]pyridines–a new class of multidrug resistance (MDR) modulators. Bioorg Med Chem. 2014;22:5860–5870. doi: 10.1016/j.bmc.2014.09.023. [DOI] [PubMed] [Google Scholar]
- 138.Juvale K., Stefan K., Wiese M. Synthesis and biological evaluation of flavones and benzoflavones as inhibitors of BCRP/ABCG2. Eur J Med Chem. 2013;67:115–126. doi: 10.1016/j.ejmech.2013.06.035. [DOI] [PubMed] [Google Scholar]
- 139.Colabufo N.A., Berardi F., Perrone M.G., Cantore M., Contino M., Inglese C., et al. Multi-drug-resistance-reverting agents: 2-aryloxazole and 2-arylthiazole derivatives as potent BCRP or MRP1 inhibitors. ChemMedChem. 2009;4:188–195. doi: 10.1002/cmdc.200800329. [DOI] [PubMed] [Google Scholar]
- 140.Colabufo N.A., Pagliarulo V., Berardi F., Contino M., Inglese C., Niso M., et al. Bicalutamide failure in prostate cancer treatment: involvement of Multi Drug Resistance proteins. Eur J Pharmacol. 2008;601:38–42. doi: 10.1016/j.ejphar.2008.10.038. [DOI] [PubMed] [Google Scholar]
- 141.Halgren T.A., Murphy R.B., Friesner R.A., Beard H.S., Frye L.L., Pollard W.T., et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem. 2004;47:1750–1759. doi: 10.1021/jm030644s. [DOI] [PubMed] [Google Scholar]
- 142.Friesner R.A., Banks J.L., Murphy R.B., Halgren T.A., Klicic J.J., Mainz D.T., et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem. 2004;47:1739–1749. doi: 10.1021/jm0306430. [DOI] [PubMed] [Google Scholar]
- 143.Schrödinger Release 2020-3: Schrödinger, LLC, New York, NY, 2021., in.
- 144.Blaschke T., Bajorath J. Compound dataset and custom code for deep generative multi-target compound design. Future Sci OA. 2021;7:Fso715. doi: 10.2144/fsoa-2021-0033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Bajorath J, Structural characteristics of compounds with multitarget activity, Future Drug Discovery, 0 FDD60.
- 146.C. Feldmann, J. Bajorath, Compounds with multitarget activity: structure-based analysis and machine learning, Future Drug Discovery, 2 (2020) FDD44.
- 147.Nagao K., Maeda M., Mañucat N.B., Ueda K. Cyclosporine A and PSC833 inhibit ABCA1 function via direct binding. Biochim Biophys Acta. 1831;2013:398–406. doi: 10.1016/j.bbalip.2012.11.002. [DOI] [PubMed] [Google Scholar]
- 148.Wu C.A., Tsujita M., Hayashi M., Yokoyama S. Probucol inactivates ABCA1 in the plasma membrane with respect to its mediation of apolipoprotein binding and high density lipoprotein assembly and to its proteolytic degradation. J Biol Chem. 2004;279:30168–30174. doi: 10.1074/jbc.M403765200. [DOI] [PubMed] [Google Scholar]
- 149.Hamon Y., Luciani M.F., Becq F., Verrier B., Rubartelli A., Chimini G. Interleukin-1beta secretion is impaired by inhibitors of the Atp binding cassette transporter, ABC1. Blood. 1997;90:2911–2915. [PubMed] [Google Scholar]
- 150.Becq F., Hamon Y., Bajetto A., Gola M., Verrier B., Chimini G. ABC1, an ATP binding cassette transporter required for phagocytosis of apoptotic cells, generates a regulated anion flux after expression in Xenopus laevis oocytes. J Biol Chem. 1997;272:2695–2699. doi: 10.1074/jbc.272.5.2695. [DOI] [PubMed] [Google Scholar]
- 151.Telbisz Á., Ambrus C., Mózner O., Szabó E., Várady G., Bakos É., et al. Interactions of Potential Anti-COVID-19 Compounds with Multispecific ABC and OATP Drug Transporters. Pharmaceutics. 2021;13:81. doi: 10.3390/pharmaceutics13010081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Vagiannis D., Novotna E., Skarka A., Kammerer S., Küpper J.H., Chen S., et al. Ensartinib (X-396) Effectively Modulates Pharmacokinetic Resistance Mediated by ABCB1 and ABCG2 Drug Efflux Transporters and CYP3A4 Biotransformation Enzyme. Cancers (Basel) 2020;12:813. doi: 10.3390/cancers12040813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Von Eckardstein A., Langer C., Engel T., Schaukal I., Cignarella A., Reinhardt J., et al. ATP binding cassette transporter ABCA1 modulates the secretion of apolipoprotein E from human monocyte-derived macrophages. Faseb j. 2001;15:1555–1561. doi: 10.1096/fj.00-0798com. [DOI] [PubMed] [Google Scholar]
- 154.Gameiro M., Silva R., Rocha-Pereira C., Carmo H., Carvalho F., Bastos M.L., et al. Cellular Models and In Vitro Assays for the Screening of modulators of P-gp, MRP1 and BCRP. Molecules. 2017;22:600. doi: 10.3390/molecules22040600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Liu H., Jiang X., Gao X., Tian W., Xu C., Wang R., et al. Identification of N-benzothiazolyl-2-benzenesulfonamides as novel ABCA1 expression upregulators. RSC Med Chem. 2020;11:411–418. doi: 10.1039/c9md00556k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Sankaranarayanan S., Kellner-Weibel G., de la Llera-Moya M., Phillips M.C., Asztalos B.F., Bittman R., et al. A sensitive assay for ABCA1-mediated cholesterol efflux using BODIPY-cholesterol. J Lipid Res. 2011;52:2332–2340. doi: 10.1194/jlr.D018051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Apweiler R., Bairoch A., Wu C.H., Barker W.C., Boeckmann B., Ferro S., et al. UniProt: the Universal Protein knowledgebase. Nucleic Acids Res. 2004;32:D115–119. doi: 10.1093/nar/gkh131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Berman H.M., Westbrook J., Feng Z., Gilliland G., Bhat T.N., Weissig H., et al. The Protein Data Bank. Nucleic Acids Res. 2000;28:235–242. doi: 10.1093/nar/28.1.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Wang J., Cieplak P., Kollman P.A. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J Comp Chem. 2000;21:1049–1074. [Google Scholar]
- 160.Morris G.M., Huey R., Lindstrom W., Sanner M.F., Belew R.K., Goodsell D.S., et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem. 2009;30:2785–2791. doi: 10.1002/jcc.21256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Harder E., Damm W., Maple J., Wu C., Reboul M., Xiang J.Y., et al. OPLS3: A Force Field Providing Broad Coverage of Drug-like Small Molecules and Proteins. J Chem Theory Comput. 2016;12:281–296. doi: 10.1021/acs.jctc.5b00864. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.