Abstract
The C-terminal domain of SARS-CoV main protease (Mpro-C) can form 3D domain-swapped dimer by exchanging the α1-helices fully buried inside the protein hydrophobic core, under non-denaturing conditions. Here, we report that Mpro-C can also form amyloid fibrils under the 3D domain-swappable conditions in vitro, and the fibrils are not formed through runaway/propagated domain swapping. It is found that there are positive correlations between the rates of domain swapping dimerization and amyloid fibrillation at different temperatures, and for different mutants. However, some Mpro-C mutants incapable of 3D domain swapping can still form amyloid fibrils, indicating that 3D domain swapping is not essential for amyloid fibrillation. Furthermore, NMR H/D exchange data and molecular dynamics simulation results suggest that the protofibril core region tends to unpack at the early stage of 3D domain swapping, so that the amyloid fibrillation can proceed during the 3D domain swapping process. We propose that 3D domain swapping makes it possible for the unpacking of the amyloidogenic fragment of the protein and thus accelerates the amyloid fibrillation process kinetically, which explains the well-documented correlations between amyloid fibrillation and 3D domain swapping observed in many proteins.
Abbreviations: Mpro-C, C-terminal domain of SARS-CoV main protease; NMR, Nuclear magnetic resonance; SEC, Size-exclusion chromatography; CD, Circular dichroism; MD, Molecular dynamics; RMSF, Root mean square fluctuation; SASA, Solvent accessible surface area
Keywords: 3D domain swapping, Amyloid fibrillation, Protein folding
1. Introduction
Domain swapping, better defined as three-dimensional (3D) domain swapping, was first introduced by Eisenberg and his colleagues to describe the crystal structure of dimeric diphtheria toxin [1]. It is a unique mechanism for protein dimerization or oligomerization through exchanging identical structure elements between two or more molecules of the same protein [2]. As a result, the folding of each subunit in the domain-swapped dimer/oligomer is almost the same as that of the monomer, except for the hinge loop that links the swapped structural elements with the rest of the structure [2], [3].
3D domain swapping is also proposed to be a mechanism for proteins to form aggregates and amyloid fibrils, which are often associated with neurodegenerative diseases [3], [4], [5]. Notably, amyloidogenic proteins, including human prion [6], cystatin C [7], and β2-microglobulin [8], were reported to be able to form 3D domain-swapped dimer/oligomer. Meanwhile, some proteins capable of 3D domain swapping were found to be able to form amyloid fibrils in vitro, such as RNase A with poly-Q expansion at its hinge loop [9], GB1 [10], T7EI [11], and cystatin E [12], etc. Indeed, obvious correlations between 3D domain swapping and amyloid fibrillation have been found for some of these proteins. And continuous domain swapping with open ends (named as “runaway domain swapping” or “propagated domain swapping”) was suggested to be the possible molecular mechanism for their amyloid fibrillation [8], [11], [12], [13], [14], [15]. However, amyloid structures of 3D domain swapping proteins revealed by experimental techniques, such as solid-state NMR (ssNMR), cryogenic electron microscopy (cryo-EM), and electron paramagnetic resonance (EPR), commonly adopt non-native cross-β conformations that are inconsistent with the structural features of 3D domain swapping, including the fibrils of human prion [16], [17], β2-microglobulin [18], [19], dsGB1 [20], [21] and cystatin B [22]. In other words, there is still a lack of direct structural evidence to support the 3D domain swapping model as a mechanism for amyloid fibrillation, and the relationship between 3D domain swapping and amyloid fibrillation remains to be further investigated.
We have previously reported that the C-terminal domain of SARS-CoV main protease (Mpro-C, residues 187–306) can form 3D domain-swapped dimer by two monomers exchanging the first α-helices, with the following L1-loop as the hinge loop [23], [24]. And the full-length Mpro can be locked at a constantly active octameric conformation consisting of four domain-swapped dimers [25], which may play an important role in cleaving polyproteins during the initial period of SARS-CoV infection [26]. Although the swappable α1-helix is enwrapped inside the hydrophobic core of Mpro-C by the rest 4 α-helices, the interconversion between monomer and domain-swapped dimer can occur under non-denaturing conditions, which is mediated by the order-to-disorder transition of the α5-helix [24].
Here, we report that the all α-helical protein Mpro-C can form amyloid fibrils with cross-β features at the same non-denaturing conditions for the 3D domain swapping in vitro. We found that the hinge loop and its following helix α2 and loop L2, within the protofibril core region, display a strong tendency of transient structural unpacking during the 3D domain swapping process, which increase the chance for fibrillation. Our results suggest that 3D domain swapping makes it possible for the unpacking of the amyloidogenic fragment of the protein and thus accelerates the amyloid fibrillation process kinetically, and provide an explanation to the well-documented correlations between amyloid fibrillation and 3D domain swapping.
2. Materials and methods
2.1. Protein purification and mutagenesis
WT Mpro-C (resides 187–306) was expressed and purified as described previously [23]. All the mutants were constructed using Muta-Direct™ Site-Directed Mutagenesis Kit (SBS Genetech). The expression and purification procedures of all the mutants were the same as those of WT Mpro-C. The proteins were stored in 50 mM PBS (pH 7.0) with 1 mM DTT (Dithiothreitol), a buffer condition used for all following assays, unless otherwise stated.
2.2. Size-exclusion chromatography (SEC)
The interconversion between Mpro-C monomer and domain-swapped dimer, and the polymerization processes were monitored by a pre-packed Superdex 75 5/150 SEC column or a pre-packed Superose 6 10/300 SEC column (GE Healthcare). The initial protein concentration was 0.5 mM for the monomer or 0.25 mM for the domain-swapped dimer. Protein samples were incubated in a water bath at 37 °C (or 42 °C for the oxidized disulfide mutants). The fractions of these samples were taken out at different time points and put on ice to stop the interconversion and polymerization processes. Mass percentages of the Mpro-C monomer, domain-swapped dimer, and polymer were determined by elution peak area integration. The kinetic rate constants for 3D domain swapping dimerization (ka) were obtained by the nonlinear fitting of the concentrations of domain-swapped dimer at different time points to a second-order kinetic equation, as previously described [27].
2.3. Native PAGE analysis
The native PAGE gel was prepared with a 12% separating gel (12% acrylamide, 375 mM Tris-HCl, pH 8.8) and a 4% stacking gel (4% acrylamide, 125 mM Tris-HCl, pH 6.8). The 2× loading buffer contained 100 mM Tris-HCl (pH 6.8), 20% glycerol, and 0.2% bromophenol blue. The running buffer contained 125 mM Tris base and 1.25 M Glycine, pH 8.3. The protein samples were first mixed with the loading buffer and then loaded onto the gel, which was run using a Bio-Rad Mini-PROTEAN assembly with normal polarity at a constant current of 20 mA. Afterwards, the gel was stained with colloidal Coomassie (0.05% Coomassie Brilliant Blue R-250, 45% methanol, and 10% acetic acid) and destained with water.
2.4. Circular dichroism (CD) spectroscopy
A monomeric Mpro-C protein sample was incubated at 37 °C for 3 days (d). Then the fractions of Mpro-C monomer, domain-swapped dimer, and polymer were isolated and diluted to ~1.0 mg/mL in 50 mM PBS (pH 7.0) with 1 mM DTT. The CD spectra were recorded at 25 °C using a 0.1 mm path-length quartz cell in the range of 190–250 nm on a MOS-500 CD spectrometer (BioLogic Science).
2.5. Identification of amyloid fibrillation
Identification of amyloid fibrillation of Mpro-C was carried out as previously described [11]. For thioflavine T (ThT) binding assays, protein samples were diluted to a monomeric concentration of 5 μM in 50 mM PBS containing 50 μM ThT. The fluorescence scans were performed on an F-7000 fluorescence spectrophotometer (Hitachi). Excitation was at 440 nm, and emission was scanned from 460 to 650 nm. For Congo Red (CR) binding assays, protein samples were diluted to a monomeric concentration of 50 μM in 50 mM PBS containing 10 μM CR. The absorbance was recorded from 400 nm to 650 nm on a UNICO 3802 UV–Vis spectrophotometer. For 2D X-ray diffraction, a 20-μL suspension of protein precipitates (~20 mg/mL) was loaded between the wax-sealed ends of two glass capillaries, and allowed to dry overnight at room temperature. X-ray diffraction data were collected using a Rigaku R-AXIS IV++ diffractometer. For transmission electron microscopy (TEM), 10-μL protein samples were loaded onto glow discharged copper grids with 200 mesh carbon/formvar support film and stained with 2% uranyl acetate. The samples were examined under an FEI Tecnai G2 transmission electron microscope with an accelerating voltage of 120 kV.
2.6. Measurement of fibrillation kinetics
Monomeric Mpro-C samples were diluted to a final concentration of 50 μM in 50 mM PBS (pH 7.0) containing 20 μM ThT, 0.03% NaN3, and cocktail proteinase inhibitors (Roche). Then the samples were loaded to a 96-well plate with an optical flat bottom in triplicate. The top of each well was sealed to prevent evaporation. Amyloid fibrillation was initiated by placing the 96-well plate into a FLUOstar OPTIMA microplate reader (BMG Labtech), set at a constant temperature of 37 °C (or 42 °C for the oxidized disulfide mutants) without shaking. ThT fluorescence was measured from the bottom of the 96-well plate per 30 min using an excitation wavelength of 430 nm and an emission wavelength of 485 nm. ThT fluorescence enhancements (∆F) were obtained by subtracting the fluorescence values of the blank sample without adding any proteins. A stretched exponential equation, expressed as ∆F = (∆Ff − m · t) − (∆Ff − m · t) exp (−[ksp · t]n), was used in the curve-fitting analysis of the fibrillation kinetics [28]. ksp is the spontaneous fibrillation rate, and n is the heterogeneity parameter. The final fluorescence change (∆Ff) was adjusted by a sedimentation coefficient (m), assuming that the precipitation of the fibrils is linear with time.
2.7. Hydrogen deuterium (H/D) exchange data processing
H/D exchange rates (kex) of the amide hydrogens in Mpro-C monomer were previously measured by solution NMR spectroscopy at 25 °C and 37 °C [24].
Based on the H/D exchange theory, the protected amide protons in a protein can only exchange with the solvent deuterons when they are transiently exposed to the solvent during some closed-to-open conformational changes [29], [30], as depicted in Eq. (1):
| (1) |
Here, k op is the rate constant of the opening motion, k cl is the rate constant of the closing motion, and k ch is the intrinsic chemical exchange rate constant at the open state, which can be determined from model compound studies. Here, the k ch value for each amino acid residue of Mpro-C was calculated using SPHERE (http://landing.foxchase.org/research/labs/roder/sphere/). For a folded protein, it is assumed that k op ≪ k cl, and thus the observed H/D exchange rate constant (kex) can be expressed as:
| (2) |
Most commonly, H/D exchange for well-folded proteins under native conditions occurs at an EX2 limit (k ch ≪ k cl), while the exchange under denatured conditions generally takes place at an EX1 limit (k ch ≫ k cl) with a rate-determining step [29], [30]. At the EX2 limit, Eq. (2) can be simplified to:
| (3) |
where Kop is the equilibrium constant for the opening [Kop ≪ 1, and fraction open = Kop / (Kop + 1)] [29]. The apparent free energy for the opening of the structure [29], [30] can be determined according to Eq. (4), where R is the gas constant, and T is the absolute temperature.
| (4) |
2.8. Molecular dynamics (MD) simulations
All the MD simulations were performed using an Amber 20 program [31] with AMBER ff14SB force field [32]. The initial conformation, taken from the solution structure of Mpro-C monomer (PDB ID: 2K7X) [23], was placed in the center of a cubic periodic water box using the TIP3P water model [33]. The box edges were set at least 1.0 nm around the protein molecule. Six Na+ ions were added to neutralize the net charges. 1500 steps of the steepest descent minimization and 500 steps of the conjugate gradient minimization were performed to remove the steric clashes between atoms. After that, the system was warmed up from 0 K to the target temperature in 0.5 ns under the NVT ensemble and further equilibrated for 0.5 ns at a constant pressure of 1 atm. Then 5 ns equilibration simulation was carried out under the NPT ensemble, followed by production simulations. For all the equilibrations and simulations in this work, a time step of 2 fs was used, the long-range electrostatic interactions were treated with the particle mesh Ewald method [34], the nonbonded cutoff distance was set to 10 Å, and the hydrogen-involved bonds were constrained using the SHAKE algorithm [35]. The pressure was controlled by coupling the system to a Berendsen barostat [36] at a reference pressure of 1 atm with a relaxation time of 2 ps. The temperature was controlled using the Langevin thermostat [37], and the collision frequency γ was set to be 2 ps−1.
For the classic MD (cMD) simulations, we performed a series of 250 ns simulations on the Mpro-C monomer at temperatures of 298 K, 313 K, 343 K, 373 K, and 403 K, without changing the potential energy surface. For the Gaussian accelerated MD (GaMD) simulations, the system potential energy surface was smoothened by adding a harmonic boost potential [38]. Before the GaMD simulations, cMD simulations were carried out to equilibrate the system and collect statistical boost parameters, each with 5.0 × 106 steps. Then the boost potential was added with initial boost parameters, and the system was equilibrated for 5.0 × 106 steps. Next, the boost parameters were updated every 1.0 × 105 steps, and the final parameters were obtained after 5.0 × 106 steps. Finally, the production simulation was run with the final boost parameters for 500 ns under the NPT ensemble. The user-specified upper limits of the standard deviations of the total boost potential and dihedral boost potential were set to 13 kcal/mol−1 and 10 kcal/mol−1, respectively.
3. Results
3.1. Mpro-C can form amyloid fibril at 37 °C
During the interconversion experiments of Mpro-C monomer and domain-swapped dimer in vitro (Fig. 1A), we noticed that high molecular weight (MW) fractions could be detected after 12 h at 37 °C. These fractions were eluted at the void volume (~1.13 mL) of the Superdex-75 column (Fig. S1 A–B). After 24 h, the high MW fractions account for ~1.5% of the total protein, while the monomer and the domain-swapped dimer account for ~92% and ~6.5%, respectively. Analysis performed using a Superose-6 column showed that the high MW fractions initially appeared with an elution peak centered at ~14.3 mL (Fig. 1B, magenta), corresponding to an MW of ~344 kDa based on the MW calibration curve (Fig. S2). With prolonged incubation time, the elution peaks of the high MW fractions became broader, ranging from 8 to 16 mL, with the center gradually shifting towards the void volume (Fig. 1B, blue & red). After 8 d, nearly all the monomer and the domain-swapped dimer were converted to high MW fractions (Fig. 1B, green), and ~90% of the protein precipitated out. The above results indicate that Mpro-C can form polymers under non-denaturing conditions, and their amounts and MWs increase with time.
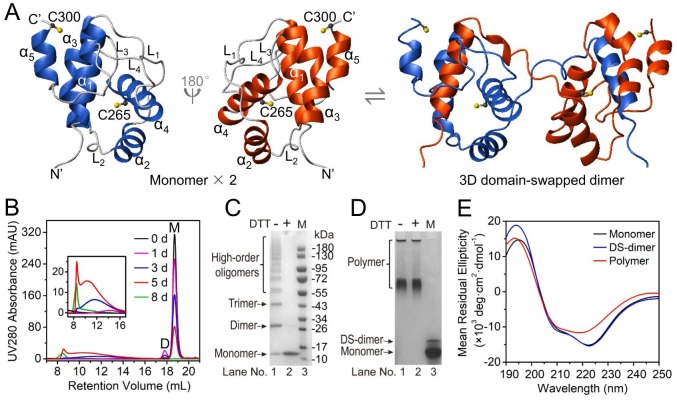
Fig. 1.
Mpro-C forms polymers under a non-denaturing condition in vitro. (A) Ribbon diagram of the structures of Mpro-C monomer and domain-swapped dimer (generated based on PDB ID: 2K7X and 3IWM). (B) Elution profiles of Mpro-C monomer incubated at 37 °C for 0 d (black), 1 d (magenta), 3 d (blue), 5 d (red), and 8 d (green). The sample contained 6.7 mg/mL protein in a PBS buffer (50 mM phosphate, 1 mM DTT, pH 7.0). Protein samples were analyzed using a Superoase-6 10/300 SEC column. (C) SDS-PAGE analysis of the soluble polymer without (−) and with (+) the addition of reducing agent DTT. (D) Native PAGE analysis of the soluble polymer without (−) and with (+) DTT. A protein sample consisting of Mpro-C monomer and domain-swapped dimer was loaded to lane 3 as a marker. (E) CD spectra of Mpro-C monomer (black), domain-swapped dimer (blue), and polymer (red). The Mpro-C polymer was isolated from a protein sample after 3 d at 37 °C.
The soluble polymeric fraction, isolated from a Mpro-C protein sample incubated at 37 °C for 3 d, displayed multiple bands of monomer, dimer, trimer, and higher-order oligomers on the non-reducing SDS-PAGE gel (Fig. 1C, lane 1), while only a single band corresponding to the Mpro-C monomer was observed on the reducing gel (Fig. 1C, lane 2). However, native PAGE analysis revealed that the polymeric states were not affected by adding the reducing agent (Fig. 1D). As there are only two cysteine residues (C265 and C300) in Mpro-C (Fig. 1A), these results indicate that the polymers should have cross-linked intermolecular disulfide bonds involving both cysteines, but these disulfide bonds are not necessary for maintaining the polymeric states. Remarkably, the residue C265 is fully buried inside the hydrophobic core (Fig. 1A) in the Mpro-C monomer or domain-swapped dimer. The fact that C265 is involved in disulfide bond formation suggests that Mpro-C in the polymer should not adopt the native fold of the monomer or the domain-swapped dimer. Indeed, the CD spectrum of the soluble polymer has a negative band at ~218 nm (Fig. 1E), typical of β-sheet structures [39]. This is quite different from those of the monomer and the domain-swapped dimer, which both exhibit negative bands at ~210 nm and 222 nm (Fig. 1E), consistent with their all α-helical fold. According to the deconvolution analysis of the CD spectra using CDPro software [40], both the monomer and the domain-swapped dimer contain ~43% α-helix and ~9% β-sheet, while the polymer has ~30% α-helix and ~21% β-sheet. Thus, there is an increase in β-sheet content and a decrease in α-helix content during the polymerization process, also suggesting that the polymer is likely not formed due to runaway/propagated domain swapping.
As the α-helix to β-sheet conversion is often associated with protein amyloid fibrillation [41], [42], ThT and CR binding assays were used to monitor the polymerization process. The ThT assay starting from Mpro-C monomer at 37 °C showed that the fluorescence intensity was slightly elevated during the first day of incubation and then dramatically increased afterward, with maximum emission at ~482 nm (Fig. 2A). Similarly, the CR assay also showed a significant increase in optical absorbance after incubating the monomer sample at 37 °C for 1 d. The absorbance maximum displayed red-shifts from ~495 nm to ~515 nm, with a shoulder peak gradually appearing at ~540 nm (Fig. 2B). In addition, both the ThT fluorescence enhancements at 482 nm and the CR absorbance increments at 540 nm are positively correlated with the proportions of the polymeric fractions in the corresponding samples (Fig. S3A–B). All these observations are typical indications of the formation of amyloid fibrils [43], [44].
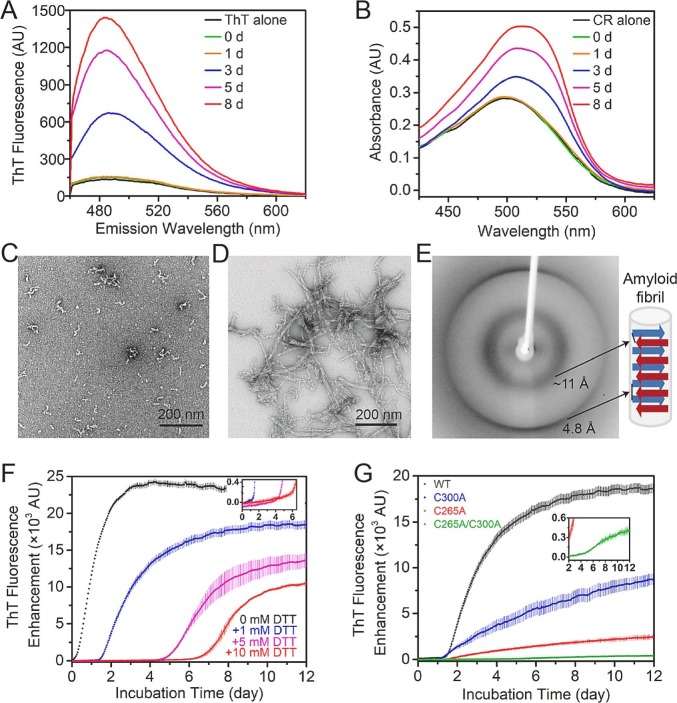
Fig. 2.
Amyloid fibril formation of Mpro-C. (A) Fluorescence spectra of ThT solution alone (black) and with Mpro-C monomer incubated at 37 °C for 0 d (green), 1 d (orange), 3 d (blue), 5 d (magenta), and 8 d (red). The Samples contain 67 μg/mL protein and 50 μM ThT. (B) Absorbance spectra of CR solution alone (black) and with Mpro-C monomer incubated at 37 °C for 0 d (green), 1 d (orange), 3 d (blue), 5 d (magenta), and 8 d (red). The samples contain 0.67 mg/mL protein and 10 μM CR. (C) TEM images of the Mpro-C polymer obtained from a protein sample after 3 d incubation at 37 °C. (D) TEM images of the Mpro-C precipitates obtained from a protein sample after 8 d incubation at 37 °C. (E) X-ray diffraction of the Mpro-C precipitates. A diagram of the cross-β structure is shown on the right. (F) Time courses of the amyloid fibrillation of WT Mpro-C monomer with 0, 1, 5, and 10 mM DTT at 37 °C, determined from ThT fluorescence measurements (excitation, 430 nm; emission, 485 nm). The lag phase region is enlarged in the insert. (G) Time courses of the amyloid fibrillation of WT Mpro-C (gray) and cysteine mutants C300A (blue), C265A (red), and C265A/C300A (green). The fibrillation curve of the C265A/C300A mutant is enlarged in the insert.
Next, we examined the morphology of the soluble polymers and the precipitates with TEM. The soluble polymers of Mpro-C displayed short curvilinear morphology with 5–10 nm in diameter and <100 nm in length (Fig. 2C), resembling the morphology of classic Aβ protofibrils [45], [46]. The precipitates formed in a protein sample incubated at 37 °C for 8 d were fibrils with diameters of 15–20 nm and lengths of several hundred nanometers (Fig. 2D). Then, we analyzed the precipitates with X-ray diffraction (Fig. 2E), which produced a typical cross-β diffraction pattern [47]. The outer 4.8 Å diffraction ring reflects the minimum distance between two β-sheets along the long fibril axis, and the inner ~11 Å diffraction ring reflects the minimum distance between two β-sheets perpendicular to the long fibril axis (Fig. 2E). Therefore, the precipitates formed by Mpro-C are mature amyloid fibrils with typical cross-β structures, and the soluble polymers should be protofibrils.
3.2. Disulfide formation accelerates amyloid fibrillation
As intermolecular disulfide bonds were found in the protofibrils (Fig. 1C), we have investigated the role of disulfide bond formation in the amyloid fibrillation of Mpro-C. The monomeric Mpro-C protein samples were incubated at 37 °C without or with the reducing agent DTT at different concentrations, in the presence of ThT. The fibrillation processes were studied by monitoring the ThT fluorescence change. The addition of DTT introduced a lag phase for the initial fibrillation process, during which the ThT fluorescence increases very slowly, followed by a dramatic increase of the fluorescence (Fig. 2F). As the duration of the fibrillation lag phase is positively correlated to the concentration of DTT, we wondered if the lag phase was related to the oxidation of DTT. We then measured the time for DTT oxidation at 1-, 5-, and 10-mM concentrations, in the same buffer condition at 37 °C (Fig. S4). The results showed that the duration of the lag phase is about the same as the time for DTT to be approximately fully oxidized, suggesting that the fibrillation rate couldn't be dramatically increased until DTT is fully oxidized and intermolecular disulfide bonds could be formed. These results indicate that the formation of intermolecular disulfide bonds can promote the amyloid fibrillation of Mpro-C on kinetics.
To verify whether disulfide bond formation is necessary for the amyloid fibrillation of Mpro-C, mutagenesis studies were carried out for the two cysteine residues. The fibrillation kinetics curves of Cys-to-Ala mutants, C265A, C300A, and C265A/C300A, were measured at 37 °C in the presence of 1 mM DTT. As shown in Fig. 2G, all the mutants exhibited much slower fibrillation rates than WT Mpro-C, in the order of C300A > C265A > C265A/C300A. Protofibrils of these mutants were isolated after 12 d and confirmed with TEM (Fig. S5), even for the cysteine-free mutant C265A/C300A with the slowest fibrillation rate. These results indicate that although both C265 and C300 residues play significant roles in accelerating the amyloid fibrillation, disulfide bond formation is not necessary for the amyloid fibrillation.
The isolated protofibrils formed by either C265A or C300A mutant mainly displayed a dimer band on the non-reducing SDS-PAGE gel (Fig. S6). With the addition of DTT, only a monomer band can be observed. These indicate that the majority of both mutants, each with one cysteine residue left, form disulfide-linked dimers during their amyloid fibrillation processes. As the C265A mutant has a much slower fibrillation rate than that of the C300A mutant (Fig. 2G), the intermolecular disulfide bond formation involving the C265 residue should be more important than that of the C300 residue, in promoting amyloid fibrillation kinetically. In the structures of both Mpro-C monomer and domain-swapped dimer, the C265 residue is protected by loop L1, helix α2, and loop L2 (L1-α2-L2) from exposing to the solvent, while the C300 residue is partially exposed (Fig. 1A). Thus, the formation of disulfide bond by the C265 residue will irreversibly disrupt the native fold of Mpro-C, which may accelerate amyloid fibrillation more efficiently.
The limited proteolysis analysis using Proteinase K showed that both Mpro-C monomer and domain-swapped dimer have the same digestion pattern, which is quite different from those of the protofibrils and the mature fibrils that are more resistant to Proteinase K digestion (Fig. S7A). Mass spectroscopy (MS) analyses revealed that the major digestion products obtained after 24 h for both the monomer and the domain-swapped dimer consist of residues 195–299 (Figs. S7B & S8). In comparison, for the protofibril, after 24 h of Proteinase K digestion, there were a major product consisting of residues 206–253 and a minor product covering residues 206–237. And the digestion product of the mature fibril covered residues 206–306 (Fig. S7B & Tables S1–3). These are also evidences that the fibrils are not formed through the 3D domain swapping. Remarkably, the core region (residues 206–253) of protofibrils corresponds to the C-terminal part of α1-helix, L1-α2-L2, and the N-terminal part of α3-helix in the Mpro-C monomer, which covers on top of α4-helix where the C265 residue is located (Fig. 1A). Consistently, the α1-L1-α2 region is also predicted to be rich in amyloidogenic segments by different online amyloidogenicity prediction methods (Fig. S7C & Table S4) [48], [49], [50], [51], [52], [53]. Therefore, the formation of the disulfide bonds involving the C265 residue will result in irreversible unpacking of the structure elements consisting of the core region of protofibrils, which makes it easier to form amyloid fibrils. This should explain why the intermolecular disulfide formation involving the C265 residue plays a more significant role in accelerating the amyloid fibrillation of Mpro-C.
3.3. Amyloid fibrillation is related to 3D domain swapping
To explore the relationship between the amyloid fibrillation and the 3D domain swapping, we first examined the temperature dependence of the Mpro-C fibrillation process (Fig. 3A) as the 3D domain swapping rate of Mpro-C is highly dependent on temperature [24]. The spontaneous fibrillation rate (ksp) values, at different temperatures from 25 °C to 39 °C, were determined by fitting the ThT fluorescence enhancement curves to a modified stretched exponential function, as previously reported [28]. The results showed that ksp is also highly dependent on temperature (Table S5). The apparent activation energy (Ea) from the Arrhenius plot analysis is 232 kJ/mol (Fig. 3B), much lower than that of the 3D domain swapping dimerization (~375 kJ/mol) [24]. In addition, based on the Arrhenius plot analysis, the fibrillation rate increased ~37 times from 25 °C to 37 °C, while the rate of 3D domain swapping dimerization increased ~350 times. Thus, for Mpro-C, both the 3D domain swapping and the amyloid fibrillation processes are highly dependent on temperature (Fig. 3C), with the 3D domain swapping process more sensitive to temperature changes.
Fig. 3.
Correlations between 3D domain swapping and amyloid fibrillation. (A) Time courses of the amyloid fibrillation of WT Mpro-C at different temperatures ranging from 25 °C to 39 °C. (B) Arrhenius plot of ln(ksp) against 1000/T, where ksp is the spontaneous fibrillation rate and T is the absolute temperature. (C) Linear plot of ln(ksp) against ln(ka) at different temperatures, where ka is the rate constant for the 3D domain swapping dimerization. (D) Mass percentage of the domain-swapped dimer as a function of time during the monomer to domain-swapped dimer conversion at 37 °C for the C300 mutants. (E) Time courses of the amyloid fibrillation of the C300 mutants at 42 °C. (F) Linear fitting of ksp against ka for the C300 mutants.
We also carried out fibrillation experiments at three different protein concentrations 10 μM, 20 μM, and 50 μM, and determined the ksp values (Fig. S9A, D). The results show that ksp is positively correlated with the protein concentration (Fig. S9B). The final fluorescence change (∆Ff) is linearly dependent on the initial monomeric protein concentration (Fig. S9C), suggesting that the amount of fibrils formed should be proportional to the total protein. In comparison, as the monomer and the domain-swapped dimer are in equilibrium exchange, both the monomer association rate constant ka and dimer dissociation rate constant kd are independent of the protein concentration. Therefore, the monomer-to-dimer and dimer-to-monomer conversion rates should be ka*[M]2 and kd*[D], in which [M] and [D] are the concentrations of monomer and dimer, respectively. The protein concentration will affect the molar ratio between monomer and dimer at equilibrium for the 3D domain swapping process.
Next, we compared the effect of the C300 residue mutations on the amyloid fibrillation rate and the 3D domain swapping rate. The domain swapping dimerization kinetics curves of mutants C300D, C300K, C300A, C300S, C300V, and C300N, were measured at 37 °C (Fig. 3D), while the corresponding fibrillation kinetics curves were monitored at 42 °C to accelerate the fibrillation process (Fig. 3E). The results indicate a positive correlation between the 3D domain swapping and the amyloid fibrillation, as mutants with faster domain swapping dimerization rates also display higher fibrillation rates (Fig. 3F & Table S5).
Furthermore, we tested whether amyloid fibrillation could be blocked for Mpro-C mutants incapable of 3D domain swapping due to engineered intermolecular disulfide bonds. We generated three mutants (I213C, T198C/E240C/C300S, and T199C/D289C/C300S) of Mpro-C, in which each would form an intramolecular disulfide bond (C213-C300, C198-C240, or C199-C289) that cross-links the swappable α1-helix with the rest of the structural elements of the monomer (Figs. 4A & S10). We also designed a mutant Q256C that can form an intramolecular disulfide bond C256-C300, which would lock the swappable α1-helix inside the hydrophobic core of the Mpro-C monomer by cross-linking the α3-helix and α5-helix. Indeed, these intramolecular disulfide bonds can block the 3D domain swapping of Mpro-C. SEC analysis revealed no dimeric form for the monomeric samples of all the four disulfide mutants after 5 d of incubation at 42 °C, when the intramolecular disulfide bond is present at the oxidized state (ox) (Fig. 4B). For comparison, these mutants could still undergo 3D domain swapping at the reduced state (re), when the intermolecular disulfide bonds were reduced with DTT (Fig. S10). Interestingly, it was found that higher-order oligomers could be observed after incubation for the monomeric mutant with disulfide bond C199-C289 or C256-C300, but not for that with disulfide bond C198-C240 or C213-C300 (Fig. 4B). Consistently, the monomeric mutant with disulfide bond C199-C289 or C256-C300 exhibited continuous ThT fluorescence enhancement during the incubation at 42 °C, while there was no ThT fluorescence enhancement for the mutant with disulfide bond C198-C240 or C213-C300 (Fig. 4C). Fibrillar species could be observed using TEM for the monomeric mutant with disulfide C199-C289 or C256-C300 after 3 d at 42 °C, but not for that with C198-C240 or C213-C300 (Fig. 4D). These results indicate that the 3D domain swapping is not a prerequisite for the amyloid fibrillation of Mpro-C, although blocking 3D domain swapping may sometimes inhibit the amyloid fibrillation.
Fig. 4.
Amyloid fibrillation of disulfide mutants that abolishes 3D domain swapping. (A) Illustration of the cysteine residues introduced to form intramolecular disulfide bonds in the structure of Mpro-C monomer. The swappable element is colored in pale green on the ribbon diagram of the structure, while the rest is colored in gray. The protofibril core region (residues 206–253) is indicated in blue on the diagram showing the secondary structural elements, and the engineered disulfide bonds are indicated by black dash lines. (B) SEC elution profiles of four oxidized monomeric disulfide mutants incubated at 42 °C for 0 d (black), 3 d (blue), and 5 d (red). (C) Time courses of the amyloid fibrillation of the monomeric disulfide mutants at oxidized state (left, at 42 °C) and reduced state (right, at 37 °C). (D) TEM images of the polymeric fractions obtained from the oxidized disulfide mutants after 3 d incubation at 42 °C. The black bars represent 200 nm.
It appears that the two disulfide bonds (C198-C240 and C213-C300) that block both 3D domain swapping and amyloid fibrillation involve residues from the protofibril core region (residues 206–253) (Fig. 4A, blue). It is possible that the two disulfide bonds could significantly restrict the conformational dynamics and prevent the amyloidogenic core region from unpacking and forming the intermolecular cross-β structure. The other two disulfide bonds (C199-C289 and C256-C300) do not interfere with the protofibril core region, which may account for why they only block the 3D domain swapping, but not the amyloid fibrillation.
3.4. Both amyloid fibrillation and 3D domain swapping depend on dynamic native fold unpacking
We have previously shown that the 3D domain swapping of Mpro-C is activated by an order-to-disorder transition of its C-terminal α5-helix foldon, which displays the most dramatic increase in H/D exchange rate (kex), i.e., a dramatic decrease in the conformational stability, from 25 °C to 37 °C [24]. To further characterize the local conformational stability of the Mpro-C monomer, we reanalyzed the previous NMR H/D exchange experimental data, and calculated the free energy for the structural opening reaction (∆Gop) and the corresponding equilibrium opening constant (Kop) at the two temperatures [30].
Consistent with our previous studies [24], the α5-helix exhibited a significant decline in its conformational stability with the temperature rising from 25 °C to 37 °C, as the ∆Gop of the V296 residue is greatly reduced by over 20 kJ/mol (Fig. 5A). The averaged ∆Gop value of residues L205-V212 from the α1-helix is significantly decreased by ~11 kJ/mol with the temperature increased from 25 °C to 37 °C (Fig. 5A, orange box), while that of residues M264-K269 and L271 from the α4-helix is decreased by ~6 kJ/mol (Fig. 5A, green box). As a result, at 25 °C, the ∆Gop values of the α1-helix residues are much higher than those of the α4-helix residues, but at 37 °C, the ∆Gop values of the two regions are comparable. These findings indicate that the stability of the swappable α1-helix becomes comparable to that of the α4-helix at 37 °C, even though α1-helix is the most stable structure element at 25 °C. It is worth noting that the Kop values of L205-V212 for the α1-helix were increased by about two orders of magnitude (50–900 times) from 25 °C to 37 °C (Fig. 5B, orange box), correspondingly, the 3D domain swapping rate involving the exchange of α1-helices between two molecules was estimated to increase by ~350 times. Meanwhile, the Kop values of the α4-helix residues M264-L268 (including the critical C265 residue for fibrillation) were increased by one order of magnitude (10–35 times) from 25 °C to 37 °C (Fig. 5B, green box), which is comparable to the estimated ~37 times increase of the fibrillation rate.
Fig. 5.
The structural dynamics of Mpro-C monomer. (A) The free energy ∆Gop for H/D exchange of the amide hydrogens in Mpro-C monomer at 25 °C (blue) and 37 °C (red). For amides that exchange too slowly, their kex values were taken as 1/5 of the slowest kex value determined, and their estimated ∆Gop values are indicated by circles or triangles with a dot in the center. For amides that exchange too fast at 37 °C, their kex values were taken as 5-fold of the fastest kex value determined, and their estimated ∆Gop values are indicated by red open triangles. ∆GU values of the whole molecule unfolding at 25 °C and 37 °C [24] are indicated by blue and red dash lines, respectively. (B) Comparison of the equilibrium constant Kop for H/D exchange of the amide hydrogens in Mpro-C monomer at 25 °C and 37 °C. (C) Time evolutions of backbone heavy atom RMSDs of the α1-α4 region (residues 201–274) in 250-ns cMD simulations of the Mpro-C monomer at 298 K (black), 313 K (blue), 343 K (green), 373 K (orange), and 403 K (red). (D) RMSFs of Cα atoms for each residue during the cMD simulations at 298 K (black), 313 K (blue), 343 K (green), 373 K (orange), and 403 K (red). (E) Time proportions of RMSD >2 Å for the secondary structural elements of the Mpro-C monomer during the cMD simulations. (F) Time evolutions of backbone RMSDs of α1-α4 region (residues 201–274) in two parallel runs of 500 ns GaMD simulations for the Mpro-C monomer at 310 K. (G) RMSFs of Cα atoms for each residue during the GaMD simulations. (H) Time evolutions of SASAs of the sidechain of the C265 residue in the GaMD simulations.
To evaluate the dynamics of Mpro-C structure, we first performed cMD simulations with the structure of the Mpro-C monomer at different temperatures ranging from 298 K to 403 K. During the 250 ns simulations, α1-α4 helices are intact, while the α5-helix appears less stable at all temperatures (Fig. S11), consistent with our previous experimental findings [24]. As expected, the backbone heavy atom RMSDs for the α1-α4 region (residues 201 to 274) are generally larger at higher simulation temperatures (Fig. 5C), especially at 373 K and 403 K. The root mean square fluctuations (RMSFs) of Cα atoms indicate that residues of the L4-α5 region have the largest structural fluctuations under the higher temperatures, followed by residues at L1-α2 junction region (Fig. 5D), which are located right on top of the C265 residue and within the fibrillation core region (Fig. 1A). At 403 K, the whole loop L1 displays relatively larger RMSFs (>1 Å), and L1 is just the hinge loop of 3D domain swapping. We also calculated the time proportions of RMSD >2 Å for the 5 α-helices and the 4 loops linking them during the above cMD simulations, and it is apparent that structure elements α5, L4, L1, and α2 are more dynamic (Figs. 5E & S12). These data are consistent with that the α5-helix unfolds first during the 3D domain swapping process [24], and then the residues of the L1-α2 region within the protofibril core region tend to unpack.
We also performed two parallel runs of Gaussian accelerated MD (GaMD) simulations on the Mpro-C monomer at 310 K [38], in order to sample a larger conformational space. According to the backbone RMSD evolutions of the α1-α4 region, significant conformational changes occurred after 300 ns simulations (Fig. 5F). It is found that the unpacking of the L4-α5 region occurred within 300 ns, followed by the unpacking of the L1-α2-L2 region after 300 ns, while the conformations of α1-helix and α4-helix are relatively stable (Fig. S13). RMSFs of the Cα atoms also reveal that the L4-α5 region and the L1-α2-L2 region are more flexible during the GaMD simulations (Fig. 5G). It should be noted that the L1-α2-L2 region (residues 215–243) corresponds to the protofibril core region (residues 206–253), and the unpacking of this region leads to the significant increase in the solvent accessible surface area (SASA) of the C265 residue (Fig. 5H, >300 ns).
Taken together, with the temperature rising, the L4-α5 region of Mpro-C displays a strong tendency to unfold, followed by the L1-α2-L2 region that also shows a tendency to unpack. As the L1-α2-L2 region initially packs on top of the α4-helix where C265 residue resides (Fig. 1A), its unpacking results in the exposure of C265 residue to solvent, which makes it possible for the formation of the intermolecular disulfide bonds involving this cysteine residue. As a result, the protein molecule cannot fold back to the native monomeric or domain-swapped dimeric structure any more, and should be kept at a misfolded state in favor of the amyloid fibrillation.
4. Discussion
Mpro-C is a unique system for studying the mechanism of 3D domain swapping, since its monomeric form and domain-swapped dimeric form can exchange reversibly under non-denaturing conditions, with both forms adopting all α-helical structures. In this study, we have demonstrated that Mpro-C can form amyloid fibrils under the non-denaturing conditions that favor the 3D domain swapping event. The amyloid fibrils formed by Mpro-C have the typical cross-β structure, and thus are not formed due to runaway/propagated domain swapping. We also demonstrated that the 3D domain swapping is not essential for the amyloid fibrillation, since two intramolecular disulfide mutants (C199-C289 or C256-C300) that are incapable of domain swapping can still form amyloid fibrils. However, the amyloid fibrillation process is not totally independent of the 3D domain swapping process, as the other two intramolecular disulfide mutations (C198-C240 or C213-C300) designed to abolish 3D domain swapping also eliminate amyloid fibrillation. The spontaneous amyloid fibrillation rate and the 3D domain swapping dimerization rate are positively correlated at different temperatures for WT Mpro-C. In addition, the two rates are also positively correlated for mutant proteins with residue 300 mutated from cysteine to other amino acids.
We have previously shown that foldon unfolding mediates the interconversion between Mpro-C monomer and 3D domain-swapped dimer, in which the unfolding of α5-helix promotes self-association of Mpro-C monomers and the disordered “α5-helix” functions to mediate the 3D domain swapping [24]. Here, our MD simulation results suggested that α5-helix and its preceding loop L4 have the highest tendency to unpack at higher temperatures, followed by the L1-α2-L2 region, which is included in the protofibril core region consisting of the most part of α1-L1-α2-L2-α3. The NMR H/D exchange experiment results also showed that the folds of increase for Kop values of L205-V212 residues of the α1-helix are consistent with the fold of increase for the 3D domain swapping rate, from 25 °C to 37 °C, while the folds of increase for the Kop values of the α4-helix residues are comparable to the fold of increase for the amyloid fibrillation rate. As the C265 residue residing α4-helix is buried underneath the L1-α2-L2 region in the structure of Mpro-C, the change of Kop values should be associated with increased unpacking frequency of the L1-α2-L2 region. As the L1-α2-L2 region is right within the protofibril core region, its unpacking will make the amyloid fibrillation possible, without completing the 3D domain swapping process. It also explains why the formation of intermolecular disulfide bonds involving the C265 residue can dramatically accelerate amyloid fibrillation.
Taken all together, we update our previously proposed model for the 3D domain swapping mechanism of Mpro-C, to illustrate its relationship with the amyloid fibrillation (Fig. 6 ). Since the swappable element α1-helix is buried inside the hydrophobic core for both the monomer and domain-swapped dimer of Mpro-C, it is obvious that the structures have to unpack for two Mpro-C molecules to exchange their α1-helices, during the 3D domain swapping process. Therefore, the structure of Mpro-C must be intrinsically very dynamic, and there should be frequent unpacking of structure elements associated with the 3D domain swapping. As the temperature increases, α5-helix first unfolds, together with the unpacking of loop L4, so as to mediate the 3D domain swapping. Meanwhile, the hinge loop (L1 loop) and the whole L1-α2-L2 region display increased conformation fluctuation and can unpack reversibly, which is also required for 3D domain swapping. As the primary sequence of L1-α2-L2 region has a high propensity for fibrillation, the opening of this region enables the formation of the cross-β structure and thus the protofibril. The exposure of the C265 residue and its participation in the formation of an intermolecular disulfide bond will make the unpacking of the L1-α2-L2 region irreversible, and thus speed up the fibrillation process. The formation of an intermolecular disulfide bond involving C300 should interrupt the ability of unfolded “α5-helix” in mediating the 3D domain swapping, in favor of the amyloid fibrillation process. This model can explain the positive correlation between the domain swapping rate and the corresponding amyloid fibrillation rate at different temperatures and for the C300 mutants. The model can also explain why an intramolecular disulfide bond preventing the 3D domain swapping can abolish the amyloid fibrillation only when it involves residues from the protofibril core region. The amyloid fibrillation process of Mpro-C can be viewed as a bypass misfolding process during the 3D domain swapping process.
Fig. 6.
Proposed model for amyloid fibrillation mechanism of Mpro-C and its relationship with 3D domain swapping. When the temperature increases, α5-helix is firstly unfolded, followed by the transient unpacking of the hinge loop and succeeding structure elements, consisting of the protofibril core region. The transient opening of the protofibril core region makes it possible to form cross-β structures and thus protofibrils. The protofibril core region is indicated in pink colour. The cysteine residues C265 and C300 are indicated by red and gold circles, respectively.
The positive correlations between the 3D domain swapping and the amyloid fibrillation can be found on different proteins, such as cystatin C [14], [54], GB1 [10], prion [13], [55], and CarD [56]. And runaway/propagated domain swapping has been proposed to be a potential amyloid fibrillation mechanism for a number of proteins, including cystatin C [15], prion [13], T7EI [11], and β2-microglobulin [8]. However, increasing evidence suggested that the amyloid fibrils of domain swapping proteins may not be formed through a runaway/propagated domain swapping mechanism. For example, it was recently reported that the V57N mutant of cystatin C, stabilized against 3D domain swapping, can still form amyloid fibrils [57]. Moreover, all the experimental amyloid structures of domain swapping proteins, including human prion [16], [17], β2-microglobulin [18], [19], and cystatin B [22], are inconsistent with the structural features of 3D domain swapping. For instance, the fibrils formed by GB1 were found to lack a native-like structure [20], although a model of extensively intermolecular β-sheets that contain the native-like β-sheets observed in the domain-swapped dimeric GB1 was initially proposed [10]. Consistent with our studies, it was also suggested that 3D domain swapping may play a role in destabilizing the native conformation at the early step of the GB1 fibrillation process [21].
Our studies on the amyloid fibrillation mechanism of Mpro-C provide insights into why 3D domain swapping is frequently associated with amyloid fibrillation. The 3D domain swapping phenomenon reflects the ability of a protein to undergo structure unpacking, which also facilitates the amyloid fibrillation for a folded protein when the unpacked structure elements have an intrinsic propensity for fibrillation. As a result, the ability of 3D domain swapping enables frequent unpacking of amyloidogenic fragments in a folded protein, and thus accelerate the amyloid fibrillation process kinetically, but not thermodynamically.
CRediT authorship contribution statement
Zhiliang Yuan: Conceptualization, Validation, Formal analysis, Investigation, Writing - Original Draft, Visualization; Zhi Qu: Validation, Investigation; Bo Duan: Writing - Original Draft; Tianyi Wang: Investigation; Jiajun Xu: Investigation; Bin Xia: Conceptualization, Validation, Formal analysis, Writing - Original Draft, Supervision, Funding acquisition.
Declaration of competing interest
The authors declare no competing interests.
Acknowledgments
We thank Dr. Weibin Gong for his help with X-ray diffraction experiments, and thank Dr. Xi Wu and Prof. Na Li for their assistant with ThT fluorescence assays. We also thank the Analytical Instrumentation Center in Peking University for the assistance with MS experiments and Dr. Wen Zhou for her help with LC/MS/MS data analysis. This work was supported by Grant 2016YFA0501202 from the Ministry of Science and Technology of China, and Grant 31770797 from the National Natural Science Foundation of China to Bin Xia.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.ijbiomac.2021.12.072.
Appendix A. Supplementary data
Supplementary material
References
- 1.Bennett M.J., Choe S., Eisenberg D. Domain swapping: entangling alliances between proteins. Proc. Natl. Acad. Sci. U. S. A. 1994;91(8):3127–3131. doi: 10.1073/pnas.91.8.3127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bennett M.J., Schlunegger M.P., Eisenberg D. 3D domain swapping: a mechanism for oligomer assembly. Protein Sci. 1995;4(12):2455–2468. doi: 10.1002/pro.5560041202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Liu Y., Eisenberg D. 3D domain swapping: as domains continue to swap. Protein Sci. 2002;11(6):1285–1299. doi: 10.1110/ps.0201402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bennett M.J., Sawaya M.R., Eisenberg D. Deposition diseases and 3D domain swapping. Structure. 2006;14(5):811–824. doi: 10.1016/j.str.2006.03.011. [DOI] [PubMed] [Google Scholar]
- 5.Zerovnik E., Stoka V., Mirtic A., Guncar G., Grdadolnik J., Staniforth R.A., Turk D. Mechanisms of amyloid fibril formation–focus on domain-swapping. FEBS J. 2011;278(13):2263–2282. doi: 10.1111/j.1742-4658.2011.08149.x. [DOI] [PubMed] [Google Scholar]
- 6.Knaus K.J., Morillas M., Swietnicki W., Malone M., Surewicz W.K., Yee V.C. Crystal structure of the human prion protein reveals a mechanism for oligomerization. Nat. Struct. Biol. 2001;8(9):770–774. doi: 10.1038/nsb0901-770. [DOI] [PubMed] [Google Scholar]
- 7.Janowski R., Kozak M., Jankowska E., Grzonka Z., Grubb A., Abrahamson M., Jaskolski M. Human cystatin C, an amyloidogenic protein, dimerizes through three-dimensional domain swapping. Nat. Struct. Biol. 2001;8(4):316–320. doi: 10.1038/86188. [DOI] [PubMed] [Google Scholar]
- 8.Liu C., Sawaya M.R., Eisenberg D. β2-microglobulin forms three-dimensional domain-swapped amyloid fibrils with disulfide linkages. Nat. Struct. Mol. Biol. 2011;18(1):49–55. doi: 10.1038/nsmb.1948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sambashivan S., Liu Y., Sawaya M.R., Gingery M., Eisenberg D. Amyloid-like fibrils of ribonuclease a with three-dimensional domain-swapped and native-like structure. Nature. 2005;437(7056):266–269. doi: 10.1038/nature03916. [DOI] [PubMed] [Google Scholar]
- 10.Louis J.M., Byeon I.J., Baxa U., Gronenborn A.M. The GB1 amyloid fibril: recruitment of the peripheral beta-strands of the domain swapped dimer into the polymeric interface. J. Mol. Biol. 2005;348(3):687–698. doi: 10.1016/j.jmb.2005.02.071. [DOI] [PubMed] [Google Scholar]
- 11.Guo Z., Eisenberg D. Runaway domain swapping in amyloid-like fibrils of T7 endonuclease I. Proc. Natl. Acad. Sci. U. S. A. 2006;103(21):8042–8047. doi: 10.1073/pnas.0602607103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Elfriede D., Hollerweger J.C., Dahms S.O., Haissi C., Katharina H.U.E., Hans B. Structural and functional analysis of cystatin E reveals enzymologically relevant dimer and amyloid fibril states. J. Biol. Chem. 2018;293(34):13151–13165. doi: 10.1074/jbc.RA118.002154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lee S., Eisenberg D. Seeded conversion of recombinant prion protein to a disulfide-bonded oligomer by a reduction-oxidation process. Nat. Struct. Biol. 2003;10(9):725–730. doi: 10.1038/nsb961. [DOI] [PubMed] [Google Scholar]
- 14.Nilsson M., Wang X., Rodziewicz-Motowidlo S., Janowski R., Lindstrom V., Onnerfjord P., Westermark G., Grzonka Z., Jaskolski M., Grubb A. Prevention of domain swapping inhibits dimerization and amyloid fibril formation of cystatin C - use of engineered disulfide bridges, antibodies, and carboxymethylpapain to stabilize the monomeric form of cystatin C. J. Biol. Chem. 2004;279(23):24236–24245. doi: 10.1074/jbc.M402621200. [DOI] [PubMed] [Google Scholar]
- 15.Wahlbom M., Wang X., Lindström V., Carlemalm E., Jaskolski M., Grubb A. Fibrillogenic oligomers of human cystatin C are formed by propagated domain swapping. J. Biol. Chem. 2007;282(25):18318–18326. doi: 10.1074/jbc.M611368200. [DOI] [PubMed] [Google Scholar]
- 16.Cobb N.J., Soennichsen F.D., McHaourab H., Surewicz W.K. Molecular architecture of human prion protein amyloid: a parallel, in-register beta-structure. Proc. Natl. Acad. Sci. U. S. A. 2007;104(48):18946–18951. doi: 10.1073/pnas.0706522104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wang L.-Q., Zhao K., Yuan H.-Y., Wang Q., Guan Z., Tao J., Li X.-N., Sun Y., Yi C.-W., Chen J., Li D., Zhang D., Yin P., Liu C., Liang Y. Cryo-EM structure of an amyloid fibril formed by full-length human prion protein. Nat. Struct. Mol. Biol. 2020;27(6):598–602. doi: 10.1038/s41594-020-0441-5. [DOI] [PubMed] [Google Scholar]
- 18.Ladner C.L., Min C., Smith D.P., Platt G.W., Radford S.E., Langen R. Stacked sets of parallel, in-register β-strands of β2-microglobulin in amyloid fibrils revealed by site-directed spin labeling and chemical labeling. J. Biol. Chem. 2010;285(22):17137–17147. doi: 10.1074/jbc.M110.117234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Iadanza M.G., Silvers R., Boardman J., Smith H.I., Karamanos T.K., Debelouchina G.T., Su Y., Griffin R.G., Ranson N.A., Radford S.E. The structure of a beta(2)-microglobulin fibril suggests a molecular basis for its amyloid polymorphism. Nat. Commun. 2018;9 doi: 10.1038/s41467-018-06761-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Li J., Hoop C.L., Kodali R., Sivanandam V.N., van der Wel P.C.A. Amyloid-like fibrils from a domain-swapping protein feature a parallel, in-register conformation without native-like interactions. J. Biol. Chem. 2011;286(33):28988–28995. doi: 10.1074/jbc.M111.261750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.van der Wel P.C.A. Domain swapping and amyloid fibril conformation. Prion. 2012;6(3):211–216. doi: 10.4161/pri.18987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Davis P.J., Holmes D., Waltho J.P., Staniforth R.A. Limited proteolysis reveals that amyloids from the 3D domain-swapping cystatin B have a non-native β-sheet topology. J. Mol. Biol. 2015;427(15):2418–2434. doi: 10.1016/j.jmb.2015.05.014. [DOI] [PubMed] [Google Scholar]
- 23.Zhong N., Zhang S., Xue F., Kang X., Zou P., Chen J., Liang C., Rao Z., Jin C., Lou Z., Xia B. C-terminal domain of SARS-CoV main protease can form a 3D domain-swapped dimer. Protein Sci. 2009;18(4):839–844. doi: 10.1002/pro.76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kang X., Zhong N., Zou P., Zhang S., Jin C., Xia B. Foldon unfolding mediates the interconversion between mpro-C monomer and 3D domain-swapped dimer. Proc. Natl. Acad. Sci. U. S. A. 2012;109(37):14900–14905. doi: 10.1073/pnas.1205241109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhang S., Zhong N., Xue F., Kang X., Ren X., Chen J., Jin C., Lou Z., Xia B. Three-dimensional domain swapping as a mechanism to lock the active conformation in a super-active octamer of SARS-CoV main protease. Protein & Cell. 2010;1(4):371–383. doi: 10.1007/s13238-010-0044-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Xia B., Kang X. Activation and maturation of SARS-CoV main protease. Protein Cell. 2011;2(4):282–290. doi: 10.1007/s13238-011-1034-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jerala R., Zerovnik E. Accessing the global minimum conformation of stefin a dimer by annealing under partially denaturing conditions. J. Mol. Biol. 1999;291(5):1079–1089. doi: 10.1006/jmbi.1999.3045. [DOI] [PubMed] [Google Scholar]
- 28.Hamada D., Dobson C.M. A kinetic study of beta-lactoglobulin amyloid fibril formation promoted by urea. Protein Sci. 2002;11(10):2417–2426. doi: 10.1110/ps.0217702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bai Y., Milne J.S., Mayne L., Englander S.W. Protein stability parameters measured by hydrogen exchange. Proteins: Struct., Funct., Bioinf. 1994;20(1):4–14. doi: 10.1002/prot.340200103. [DOI] [PubMed] [Google Scholar]
- 30.Bai Y., Sosnick T.R. Protein folding intermediates: native-state hydrogen exchange. Science. 1995;269(5221):192. doi: 10.1126/science.7618079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Case D.A., Belfon K., Ben-Shalom I.Y., Brozell S.R., Cerutti D.S., Cheatham I.T.E., Cruzeiro V.W.D., Darden T.A., Duke R.E., Giambasu G., Gilson M.K., Gohlke H., Goetz A.W., Harris R., Izadi S., Izmailov S.A., Kasavajhala K., Kovalenko A., Krasny R., Kurtzman T., Lee T.S., LeGrand S., Li P., Lin C., Liu J., Luchko T., Luo R., Man V., Merz K.M., Miao Y., Mikhailovskii O., Monard G., Nguyen H., Onufriev A., Pan F., Pantano S., Qi R., Roe D.R., Roitberg A., Sagui C., Schott-Verdugo S., Shen J., Simmerling C., Skrynnikov N.R., Smith J., Swails J., Walker R.C., Wang J., Wilson L., Wolf R.M., Wu X., Xiong Y., Xue Y., York D.M., Kollman P.A. University of California; San Francisco: 2020. Amber 2020. [Google Scholar]
- 32.Simmerling, Carlos, Hauser, Kevin E., Maier, James A., Kasavajhala, Koushik ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. 2015;11(8):3696–3713. doi: 10.1021/acs.jctc.5b00255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Jorgensen W.L., Chandrasekhar J., Madura J.D., Impey R.W., Klein M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983;79(2):926–935. [Google Scholar]
- 34.Darden T., York D., Pedersen L. Particle mesh ewald - an n.LOG(n) method for ewald SUMS in large systems. J. Chem. Phys. 1993;98(12):10089–10092. [Google Scholar]
- 35.Ryckaert J.P., Ciccotti G., Berendsen H.J.C. Numerical-integration of cartesian equations of motion of a system with constraints - molecular-dynamics of n-alkanes. J. Comput. Phys. 1977;23(3):327–341. [Google Scholar]
- 36.Berendsen H.J.C., Postma J.P.M., Vangunsteren W.F., Dinola A., Haak J.R. Molecular-dynamics with coupling to an external bath. J. Chem. Phys. 1984;81(8):3684–3690. [Google Scholar]
- 37.Turq P., Lantelme F., Friedman H.L. Brownian dynamics - its application to ionic-solutions. J. Chem. Phys. 1977;66(7):3039–3044. [Google Scholar]
- 38.Miao Y., Feher V.A., Mccammon J.A. Gaussian accelerated molecular dynamics: unconstrained enhanced sampling and free energy calculation. J. Chem. Theory Comput. 2015;11(8):3584–3595. doi: 10.1021/acs.jctc.5b00436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Greenfie N., Fasman G.D. Computed circular dichroism spectra for the evaluation of protein conformation. Biochemistry. 1969;8(10):4016–4108. doi: 10.1021/bi00838a031. [DOI] [PubMed] [Google Scholar]
- 40.Sreerama N., Woody R.W. Estimation of protein secondary structure from circular dichroism spectra: comparison of CONTIN, SELCON, and CDSSTR methods with an expanded reference set. Anal. Biochem. 2000;287(2):252–260. doi: 10.1006/abio.2000.4880. [DOI] [PubMed] [Google Scholar]
- 41.Pan K.M., Baldwin M., Nguyen J., Gasset M., Serban A., Groth D., Mehlhorn I., Huang Z.W., Fletterick R.J., Cohen F.E., Prusiner S.B. Conversion of ALPHA-helices into BETA-sheets features in the formation of the scrapie prion proteins. Proc. Natl. Acad. Sci. U. S. A. 1993;90(23):10962–10966. doi: 10.1073/pnas.90.23.10962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Fandrich M., Fletcher M.A., Dobson C.M. Amyloid fibrils from muscle myoglobin. Nature. 2001;410(6825):165–166. doi: 10.1038/35065514. [DOI] [PubMed] [Google Scholar]
- 43.Hudson S.A., Ecroyd H., Kee T.W., Carver J.A. The thioflavin T fluorescence assay for amyloid fibril detection can be biased by the presence of exogenous compounds. FEBS J. 2009;276(20):5960–5972. doi: 10.1111/j.1742-4658.2009.07307.x. [DOI] [PubMed] [Google Scholar]
- 44.Tufail S., Owais M., Kazmi S., Balyan R., Khalsa J.K., Faisal S.M., Sherwani M.A., Gatoo M.A., Umar M.S., Zubair S. Amyloidform of ovalbumin evokes native antigen-specific immune response in the host prospective immuno-prophylactic potential. J. Biol. Chem. 2015;290(7):4131–4148. doi: 10.1074/jbc.M113.540989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Paranjape G., Gouwens L.K., Osborn D.C., Nichols M.R. Isolated amyloid-β(1–42) protofibrils, but not isolated fibrils, are robust stimulators of microglia. ACS Chem. Neurosci. 2012;3(4):302–311. doi: 10.1021/cn2001238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kheterpal I., Lashuel H.A., Hartley D.M., Wlaz T., Lansbury P.T., Wetzel R. A beta protofibrils possess a stable core structure resistant to hydrogen exchange. Biochemistry. 2003;42(48):14092–14098. doi: 10.1021/bi0357816. [DOI] [PubMed] [Google Scholar]
- 47.Riek R., Eisenberg D.S. The activities of amyloids from a structural perspective. Nature. 2016;539(7628):227–235. doi: 10.1038/nature20416. [DOI] [PubMed] [Google Scholar]
- 48.Fernandez-Escamilla A.M., Rousseau F., Schymkowitz J., Serrano L. Prediction of sequence-dependent and mutational effects on the aggregation of peptides and proteins. Nat. Biotechnol. 2004;22(10):1302–1306. doi: 10.1038/nbt1012. [DOI] [PubMed] [Google Scholar]
- 49.Walsh I., Seno F., Tosatto S.C., Trovato A. PASTA 2.0: an improved server for protein aggregation prediction. Nucleic Acids Res. 2014;42(Web Server issue):W301–W307. doi: 10.1093/nar/gku399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Conchillo-Sole O., de Groot N.S., Aviles F.X., Vendrell J., Daura X., Ventura S. AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides. BMC Bioinformatics. 2007;8:65. doi: 10.1186/1471-2105-8-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.O'Donnell C.W., Waldispuehl J., Lis M., Halfmann R., Devadas S., Lindquist S., Berger B. A method for probing the mutational landscape of amyloid structure. Bioinformatics. 2011;27(13):I34–I42. doi: 10.1093/bioinformatics/btr238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Maurer-Stroh S., Debulpaep M., Kuemmerer N., de la Paz M.L., Martins I.C., Reumers J., Morris K.L., Copland A., Serpell L., Serrano L., Schymkowitz J.W.H., Rousseau F. Exploring the sequence determinants of amyloid structure using position-specific scoring matrices. Nat. Methods. 2010;7(3):237–242. doi: 10.1038/nmeth.1432. [DOI] [PubMed] [Google Scholar]
- 53.Garbuzynskiy S.O., Lobanov M.Y., Galzitskaya O.V. FoldAmyloid: a method of prediction of amyloidogenic regions from protein sequence. Bioinformatics. 2010;26(3):326–332. doi: 10.1093/bioinformatics/btp691. [DOI] [PubMed] [Google Scholar]
- 54.Ostner G., Lindstrom V., Christensen P.H., Kozak M., Abrahamson M., Grubb A. Stabilization, characterization, and selective removal of cystatin C amyloid oligomers. J. Biol. Chem. 2013;288(23):16438–16450. doi: 10.1074/jbc.M113.469593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Hafner-Bratkovic I., Bester R., Pristovsek P., Gaedtke L., Veranic P., Gaspersic J., Mancek-Keber M., Avbelj M., Polymenidou M., Julius C. Globular domain of the prion protein needs to be unlocked by domain swapping to support prion protein conversion. J. Biol. Chem. 2011;286(14):12149–12156. doi: 10.1074/jbc.M110.213926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Gundeep K., Soni K., Srajan K., GJM, HJT, Gopal T.K. Mycobacterium tuberculosis CarD, an essential global transcriptional regulator forms amyloid-like fibrils. Sci. Rep. 2018;8 doi: 10.1038/s41598-018-28290-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Perlenfein T.J., Mehlhoff J.D., Murphy R.M. Insights into the mechanism of cystatin C oligomer and amyloid formation and its interaction with beta-amyloid. J. Biol. Chem. 2017;292(27):11485–11498. doi: 10.1074/jbc.M117.786558. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary material