Abstract
Wastewater-based epidemiology (WBE) has proven to be an effective tool for epidemiological surveillance of SARS-CoV-2 during the current COVID-19 pandemic. Furthermore, combining WBE together with high-throughput sequencing techniques can be useful for the analysis of SARS-CoV-2 viral diversity present in a given sample. The present study focuses on the genomic analysis of SARS-CoV-2 in 76 sewage samples collected during the three epidemiological waves that occurred in Spain from 14 wastewater treatment plants distributed throughout the country. The results obtained demonstrate that the metagenomic analysis of SARS-CoV-2 in wastewater allows the detection of mutations that define the B.1.1.7 lineage and the ability of the technique to anticipate the detection of certain mutations before they are detected in clinical samples. The study proves the usefulness of sewage sequencing to track Variants of Concern that can complement clinical testing to help in decision-making and in the analysis of the evolution of the pandemic.
Keywords: SARS-CoV-2, Wastewater, Genome sequencing, Spike mutations, Variants of concern, Variants of interest
1. Introduction
The family Coronaviridae is a family of enveloped RNA viruses generally associated with mild respiratory and gastrointestinal infections (Shang et al., 2020). Nevertheless, in recent decades new and highly pathogenic zoonotic coronavirus (CoVs) have emerged such as the Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) (Drosten et al., 2003; Ksiazek et al., 2003), the Middle East Respiratory Syndrome Coronavirus (MERS-CoV) (Zaki et al., 2012) and, most recently, SARS-CoV-2 (Zhu et al., 2020) which has resulted in the Coronavirus Disease 2019 (COVID-19) pandemic. Transmission of SARS-CoV-2 occurs mainly through aerosols or respiratory secretions (Chan et al., 2020) but it has also been found that, due to its replication capacity in the gastrointestinal tract (Xiao et al., 2020), it is excreted in feces and urine, as was previously reported for its counterparts SARS-CoV and MERS-CoV. For this reason, it has been possible to detect the genetic material of the virus in the feces of not only symptomatic, but also asymptomatic people (Polo et al., 2020). These findings have led to the use of wastewater monitoring for SARS-CoV-2. As for other pathogens, the use of Wastewater-Based Epidemiology (WBE) has proven to be a very useful tool as an early detection warning system, allowing trend-estimations as well as establishing correlations between different epidemiological indicators (Bivins et al., 2020; Medema et al., 2020; Randazzo et al., 2020b, 2020a). One of the reasons for the success of WBE is that wastewater samples are a non-invasive and inexpensive source of information to investigate the spread of SARS-CoV-2 within a community. Moreover, it provides real-time information on the circulating lineages of SARS-CoV-2, which is essential for the development of vaccines and drugs. This is particularly relevant in view of the current situation where the world's population is being vaccinated against SARS-CoV-2 and where, due to the appearance of emerging lineages, vaccine effectiveness might be compromised (Zhou et al., 2021).
Massive parallel sequencing techniques applied to sewage samples allow us to analyze a large number of SARS-CoV-2 genomes, including those present in symptomatic and asymptomatic persons. Through the analysis of sequences, it is possible to detect low-frequency variants (LFV) and to infer which lineages are circulating at a certain time and place (Bar-Or et al., 2021; Crits-Christoph et al., 2021; Dharmadhikari et al., 2021; Herold et al., 2021; Izquierdo-Lara et al., 2021; La Rosa et al., 2021; Nemudryi et al., 2020; Rios et al., 2021). Additionally, genomic analyses may allow to detect the entry of described lineages or Variants of Concern (VOCs) into populations, as well as the appearance of emerging lineages, to characterize new outbreaks, and to aid in viral strains tracking (Bar-Or et al., 2021; Crits-Christoph et al., 2021; Izquierdo-Lara et al., 2021; La Rosa et al., 2021; Nemudryi et al., 2020; Rios et al., 2021). These studies also evidenced that improvement on sequencing techniques must be performed in order to reduce error rates, as the case of Nanopore sequencing (Nemudryi et al., 2020). Despite these limitations, the published works showed that genomic analysis of SARS-CoV-2 in wastewater should be used as a complementary tool in epidemiological surveillance. This aspect has grown in significance because during the spread of SARS-CoV-2, different mutations (i.e. D614G, Δ69/70, N501Y, E484K, K417N) present in VOCs (i.e. B.1.1.7, B.1.351, B.1.617.2, and B.1.1.28.1) have emerged (https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/), which have a considerable impact on transmissibility, infection severity (Singh et al., 2021), or immunity. These characteristics, if they occur, can aggravate the epidemiological situation in certain areas, so the detection of new lineages and the appearance of VOCs in any specific population is crucial to overcome the current pandemic situation and control the spread of the virus. Variants of interest (VOI) (for example, B.1.427) have also been defined which are currently under investigation and must be monitored to ensure a prompt response should they pose a greater threat to the population. The usefulness of these techniques is evident from the fact that the European Commission published, on March 17, 2021, recommendations for the establishment of SARS-CoV-2 surveillance in wastewater, highlighting the importance of SARS-CoV-2 sequencing in wastewater as a tool for the detection of VOC and VOI (https://eur-lex.europa.eu/legal-content/EN/ALL/?uri=uriserv:OJ.L_.2021.098.01.0003.01.ENG). The aim of this study was to analyze SARS-CoV-2 genomes in wastewater through high-throughput sequencing in order to monitor the emergence of mutations, lineages or the detection of signature mutations of VOCs and VOIs.
2. Materials and methods
2.1. Sample processing
In the framework of SARS-CoV-2 wastewater monitoring in Spain, grab samples were collected from 14 treatment plants located in different parts of the Spanish territory, with equivalent inhabitant values ranging from 60,600 to 1900,800. The samples taken between April 2020 and January 2021 encompass the three waves that have affected the country. The first wave occurred between March and April of 2020, the second wave in November 2020, and the third wave between January and February 2021. For each sample, 200 mL of wastewater samples were artificially inoculated with porcine epidemic diarrhea virus (PEDV) as process control with a final concentration of 4.5 log (PCRU/L), and concentrated following an aluminum-based adsorption precipitation method (AAVV, 2018; Pérez-Cataluña et al., 2021; Randazzo et al., 2020b). Then, 200 mL of wastewater was adjusted to pH 6.0. Precipitation by Al(OH)3 was carried out by mixing 1 part of 0.9 N AlCl3 per 100 parts of sample. Next, the solution was mixed at 150 rpm for 15 min, centrifuged at 1700 × g for 20 min, and the resulting pellet was resuspended in 10 mL of 3% beef extract (pH 7.4) then stirred at 150 rpm for 10 min at room temperature (RT). Finally, the suspension was centrifuged at 1900 × g for 30 min and the pellet resuspended in 1 mL of phosphate buffered saline solution (PBS, pH 7.4). After this, concentrated samples were stored at −80 °C until analysis.
2.2. Nucleic acid extraction and SARS-CoV-2 RT-qPCR quantification
Nucleic acid extraction from wastewater concentrates was performed using an automated method with the Maxwell RSC Pure Food GMO and authentication kit (Promega) with slight modifications (Pérez-Cataluña et al., 2021). Firstly, 300 μL of concentrated samples were mixed with 400 μL of cetyltrimethyl ammonium bromide (CTAB) and 40 μL of proteinase K solution. The mixed sample was incubated at 60 °C for 10 min and centrifuged for 10 min at 16,000 × g. Next, the resulting supernatant was transferred to the loading cartridge and 300 μL of lysis buffer added. The cartridge was then loaded in the Maxwell® RSC Instrument (Promega) using the “Maxwell RSC Viral total Nucleic Acid” running program for the nucleic acid extraction. The obtained RNA was eluted in 100 μL nuclease-free water. Negative controls were included by using nuclease-free water instead of concentrated sample.
SARS-CoV-2 nucleic acid detection was performed by RT-qPCR using One Step PrimeScriptTM RT-PCR Kit (Perfect Real Time) (Takara Bio, USA) targeting a genomic region of the nucleocapsid gene (N1 region) using primers, probes and conditions previously described (CDC, 2020). The complete genomic RNA of SARS-CoV-2 (ATCC VR-1986D) and nuclease free water were used as positive and negative controls, respectively.
2.3. SARS-CoV-2 genome sequencing and analysis
Samples with RT-qPCR cycle threshold (Ct) values below 36 were selected for sequencing analysis. Genomic sequencing of SARS-CoV-2 present in selected wastewater samples was carried out following ARTIC protocol version 3 for retrotranscription and amplification by multiplex PCR (Quick, 2020; https://www.protocols.io/view/ncov-2019-sequencing-protocol-v3-locost-bh42j8ye). Sequencing libraries were built using the Nextera Flex kit (Illumina) and sequenced on Illumina MiSeq platform by paired-end reads (2 × 200).
Raw reads were cleaned for adaptors and low quality nucleotides by using cutadapt software (Martin, 2011) and reformat.sh from bbmap (sourceforge.net/projects/bbmap/), respectively. Nucleotides with Phred score lower than 30 were discarded. Clean reads were aligned to the genome of SARS-CoV-2 isolate Wuhan-Hu-1 (MN908947.3) using the Burrows-Wheeler Aligner v0.7.17-r1188 with default parameters (Li and Durbin, 2009) and indexed by samtools (Li et al., 2009). For the analysis of genomic coverage for each sample, only nucleotides with at least 20X depth were taken into account. Nucleotide substitutions and deletions regarding SARS-CoV-2 isolate Wuhan-Hu-1 genome (MN908947.3) were detected with the aligned reads using mpileup from samtools (Li, 2011) and the command variants of ivar software (Grubaugh et al., 2019). For the assumption of one nucleotide polymorphism, at least a 50X depth of the alternative nucleotide and quality score higher than 30 were used as cutoff. Alignments were manually curated to avoid nucleotide substitutions that corresponded to incorrectly trimmed adaptors (Nemudryi et al., 2020). Information about SARS-CoV-2 mutation distribution worldwide was obtained from outbreak.info (Mullen et al., 2020).
3. Results and discussion
3.1. SARS-CoV-2 quantification and genome coverage
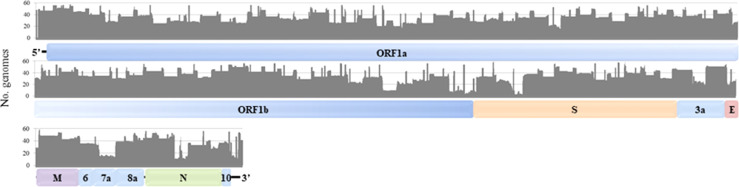
A total of 76 sewage samples positive for SARS-CoV-2 by RT-qPCR (Ct < 36) collected throughout the three epidemiological waves were sequenced during this study (Supplementary Fig. S1). Samples were grouped in three regions: north (2 WWTPs, n = 8), center (7 WWTPs, n = 39), and south (5 WWTPs, n = 29). Results showed Ct values of SARS-CoV-2 target N1 ranged from 26.59 to 34.75 (Table S1). From the 76 sequenced samples, 11 (14.5%) showed percentages of 20X coverage values higher than 90% (Figs. 1 and S2), and a mean genomic percentage of coverage of 50.1 ± 30.6%. In order to study the potential correlation between viral loads and genome coverage in wastewater samples, correlation analyses between RT-qPCR outputs (genome copies (gc) per liter) versus genome coverage were carried out for each individual sample. No correlations were found for the analyzed target, as occurred in the study of Izquierdo-Lara et al. (2021) for Illumina reads. Fig. 1 shows the number of samples that covered a certain nucleotide position with depths higher than 20X among the samples with genomic coverage greater than 20% (n = 59). However, some areas were only covered by less than 20 samples (< 33.90%), as is the case of two regions at the end of ORF 1b (nucleotides 21,456–21,467 and several regions between nucleotides 21,162 and 21,600), the regions of the S gene from nucleotides 22,303 to 22,342 and from 22,364 to 22,523, most of the ORF7a (nucleotides 27,529–27,790), and the central region of the N gene (nucleotides 28,773–28,853 and 28,901–28,993) (Fig. 1).
Fig. 1.
Representation of the number of samples that covered each nucleotide of the SARS-CoV-2 isolate Wuhan-Hu-1 genome (MN908947.3) with coverage values higher than 20X.
3.2. Overview of detected nucleotide substitutions and deletions
Sequence analysis showed a total of 627 nucleotide substitutions and 20 deletions (Table 1 ) in comparison with the reference genome of SARS-CoV-2 isolate Wuhan-Hu-1 (MN908947.3). Among detected nucleotide substitutions: 248 were found in ORF1a polyprotein; 171 in ORF1b; 71 in the spike glycoprotein; 32 in ORF3a; 29 in the membrane glycoprotein; 20 in ORF8; 39 in the nucleocapsid gene; 31 in intergenic regions; 3 in ORF10; and one in the envelope protein, ORF6, and ORF7a each (Table 1, Fig. 2 ). Regarding deletions, a total of 8 deletions were found in samples of the first and second waves: 5 of them in the ORF1a region (Δ21–23, Δ82–84, Δ84–86, Δ141–143, and Δ682); one in the spike glycoprotein (Δ385); and two in the ORF3a (Δ80 and Δ11–20). Two of these deletions were found in two samples: ORF1a:Δ141–143 in samples N1–18–2020 and C3–40–2020; and ORF3a:Δ11–20 in samples N1–18–2020 and N2–37–2020. None of these deletions were previously reported according to GISAID database after searching each deletion in the outbreak.info database. In samples from the third wave, a total of 12 deletions were detected: three in ORF1a (Δ141–143 found in two samples of first and second waves and in three samples of different locations from the third wave (C4–2–2021, C4–3–2021, and C5–4–2021), Δ2037, and Δ3675–3677 found in three samples (C2–1–2021, C4–1–2021, and C5–2–2021))), and nucleocapsid gene (Δ266–273, Δ352–356, and Δ392); and two in ORF1b (Δ176 and Δ1111), spike glycoprotein (Δ69/70 and Δ144, both found in samples S1–2–2021 and C4–2–2021), and ORF3a (Δ234 and Δ259/262). The percentage of non-synonymous substitutions ranged from 45.5% (in membrane glycoprotein) to 100% (in ORF7a and ORF10) in samples from the first and second waves, while in samples from the third wave this percentages ranged from 28.0% (in membrane glycoprotein) to 82.4% (in ORF8) (Table 1).
Table 1.
Overview of the nucleotide substitutions and deletions detected in SARS-CoV-2 genomes from wastewater samples (n = 76) as compared to the SARS-CoV-2 isolate Wuhan-Hu-1 reference genome (MN908947.3). NA, not applicable.
| Wave | 5′ UTR | ORF1a polyprotein | ORF1b polyprotein | Surface glycoprotein | ORF3a protein | Intergenic region | Envelope protein | Membrane glycoprotein | ORF6 protein | ORF7a protein | Intergenic region | ORF8 protein | Nucleocapsid phosphoprotein | ORF10 protein | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1st and 2nd (a) | Synonymous | NA | 32 | 15 | 6 | 1 | NA | 1 | 6 | 0 | 0 | NA | 1 | 6 | 0 |
| Non synonymous | NA | 52 | 43 | 14 | 4 | NA | 0 | 5 | 0 | 1 | NA | 2 | 2 | 1 | |
| Deletions | NA | 5 | 0 | 1 | 2 | NA | 0 | 0 | 0 | 0 | NA | 0 | 0 | 0 | |
| Total | 10 | 89 | 58 | 21 | 7 | 1 | 1 | 11 | 0 | 1 | 1 | 3 | 8 | 1 | |
| 3rd (b) | Synonymous | NA | 53 | 33 | 12 | 7 | NA | 0 | 13 | 1 | 0 | NA | 3 | 11 | 1 |
| Non synonymous | NA | 103 | 78 | 36 | 16 | NA | 0 | 5 | 0 | 0 | NA | 14 | 17 | 1 | |
| Deletions | NA | 3 | 2 | 2 | 2 | NA | 0 | 0 | 0 | 0 | NA | 0 | 3 | 0 | |
| Total | 16 | 159 | 113 | 50 | 25 | 2 | 0 | 18 | 1 | 0 | 1 | 17 | 31 | 2 | |
| Total | 26 | 248 | 171 | 71 | 32 | 3 | 1 | 29 | 1 | 1 | 2 | 20 | 39 | 3 |
1st wave: Samples from week 15 to 19 of 2020, n = 5; 2nd wave: Samples from week 34 to 42 of 2020, n = 35.
3rd wave: Samples from week 50 of 2020 to week 8 of 2021, n = 36.
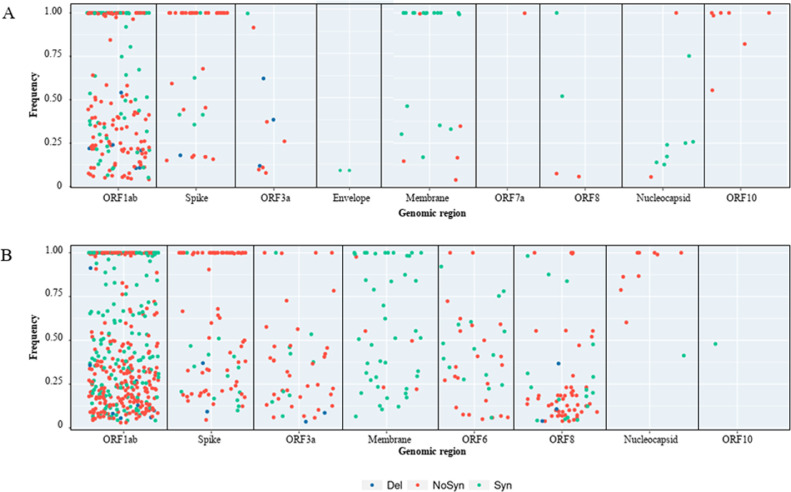
Fig. 2.
Frequency of the different nucleotide substitutions and deletions obtained for samples of each epidemiological wave grouped by genomic region. A, first and second waves; B, third wave.
Some of these nucleotide substitutions and deletions were present along with the homologous nucleotide of SARS-CoV-2 isolate Wuhan-Hu-1 genome (Fig. 2) evidencing the presence of multiple genomes in wastewaters. Mean values of the frequency of these nucleotide polymorphisms were 70±35% for synonymous substitutions, 56±38% for non-synonymous substitutions and 27±18% for deletions in samples from the first and second waves (Fig. 2A). These values in the third wave samples were 53±34%, 43±34%, and 19±22% for synonymous substitutions, non-synonymous substitutions, and deletions, respectively (Fig. 2B).
3.3. Nucleotide substitutions and deletions in the spike glycoprotein gene
Table 2 shows the non-synonymous nucleotide substitutions (n = 49) and deletions (n = 3) found in the spike glycoprotein gene. Among these polymorphisms, 18 of them were not previously described in genomes obtained from Spanish sequences, according to the database available at https://outbreak.info/ (Mullen et al., 2020). However, two of these nucleotide substitutions (amino acid substitutions G404V and G648V) have been found at low frequencies among the reads obtained in the sequencing of Spanish genomes from clinical samples. These results evidence the ability of this technique to detect mutations that are in low percentage in the viral population and from different lineages.
Table 2.
Non-synonymous nucleotide substitutions and deletions detected in the spike glycoprotein region as compared to the SARS-CoV-2 isolate Wuhan-Hu-1 reference genome (MN908947.3). Reference (ref) and alternative (alt) depth relate to the percentage of the total depth that corresponded to the nucleotide present in the reference genome MN908947.3 and the alternative nucleotide, respectively. Mixed samples related to the number of samples showing nucleotides according to reference and alternative sequence. Light gray, region of the receptor binding domain; dark gray, region of the S1/S2 cleavage region. NA, not applicable.
 |
 |
Interestingly, some of these amino acid substitutions in the spike protein were found in sewage at the same time or even weeks or months before their appearance in genomes from clinical samples. For example, among nucleotide substitutions that have been detected in Spain in clinical samples, two spike mutations (G639S and V642G) were found in waters around the same time that they appeared in clinical genomes, while spike mutations A648V was found in waters 6 weeks before, and mutations S884F, G404V, and A372T were found in waters between 4 and 5 months before their detection in clinical genomes. It should be noted that, in the case of G404V, its first detection occurred at very low percentages of sequencing reads in some clinically obtained genomes (n = 2), and its appearance at higher frequencies in one clinical genome was 5 months later. Additionally, for these genomic mutations, the number of clinical cases was very low, ranging from 1 to 6 cases. Moreover, mutations A893T, L1152S, and N1173K had not been detected in Spanish clinical genomes but their detection in other countries occurred after detection in Spanish wastewater, more specifically 3, 4, and 8 months before, respectively. These results, along with those obtained by other authors who found genomes and single nucleotide polymorphisms (SNPs) widely described in the clinical samples (Crits-Christoph et al., 2021; Izquierdo-Lara et al., 2021), show that high-throughput sequencing of SARS-CoV-2 in wastewater is a very useful complementary tool for studies and decision-making related to the epidemiology of the virus.
3.4. Identification of B.1.1.7 (VOC 202,012/01) mutation signatures
The highly transmissible B.1.1.7 lineage of SARS-CoV-2 contains 16 characteristic non-synonym nucleotide substitutions and deletions (Rambaut et al., 2020) and was first detected in Spain in week 52 of 2020 in different Spanish regions (Madrid, Basque Country, and the Balearic islands).
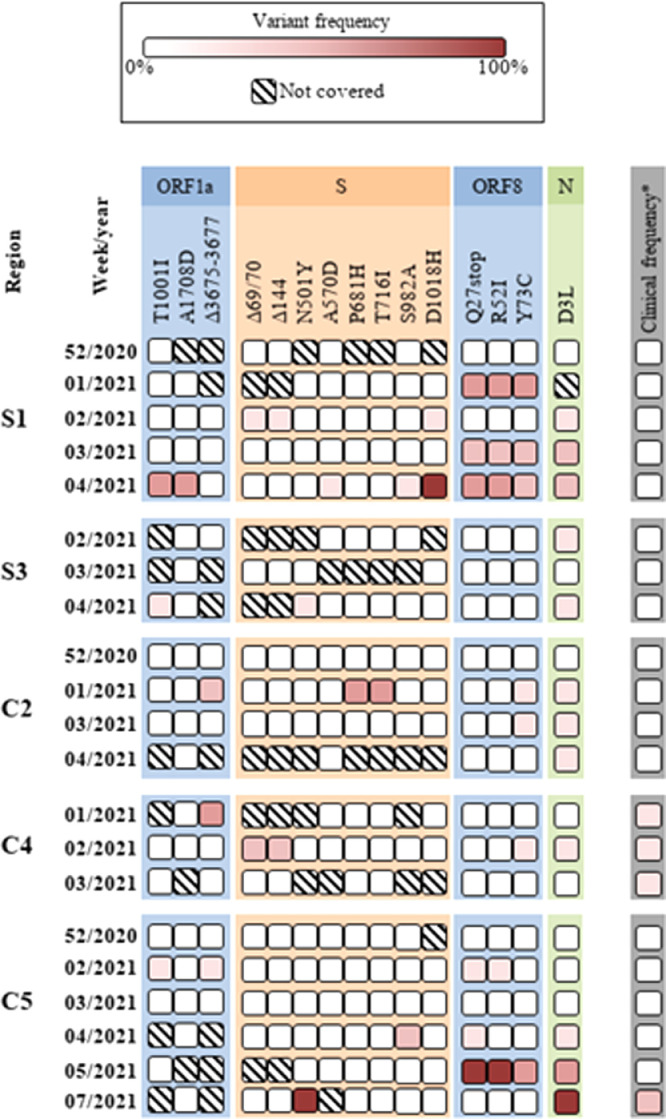
The characteristic mutations described in the genome of the B.1.1.7 lineage were searched for in our sequencing data. These mutations corresponded to 2 nucleotide substitutions and one deletion in ORF1a, 6 nucleotide substitutions and 2 deletions in spike gene, 3 nucleotide substitutions in ORF8, and one nucleotide substitution in N gene (Fig. 3 ). Amino acid substitutions S235F of nucleocapsid protein was not shown because it was absent or not covered. Samples with Ct values below 36 for N1 were analyzed, starting from the week 52 of 2020 up to week 7 in the case of samples from region C5. Among the analyzed samples, only samples from regions S1, S3, C3, C4, and C5 showed characteristic mutations of the B.1.1.7 lineage. None of the samples showed all the 18 markers that were searched for, at the same time. The highest presence and frequency of mutations was found in nucleocapsid (detected in 60% of the samples) and in ORF8 region (detected in 45% of the samples), and the lowest in ORF1a. Interestingly, the three characteristic mutations in ORF8 (Q27stop, R52I, and Y73C) were detected together in 4 of the 9 samples. Only one sample, S1–4–2021, showed 9 out of 15 characteristic mutations. The deletion of spike amino acids 69 and 70 (S:Δ69/70) was found in 2 samples from different geographical regions, that were present along with deletion ΔY144. Although some of these mutations can belong to different lineages (Table 2), their absence in samples from the first and second waves denotes the introduction of new mutations that could be related with the lineage B.1.1.7. This idea is reinforced by the fact that some mutations, such as the three present in ORF8, appeared together in some samples. Additionally, as occurred with the detection of new amino acid substitutions in the spike gene (Section 3.3), some of these mutations associated to the lineage B.1.1.7. appeared even 4 weeks before their detection in clinical samples, according to the results obtained by S-gene target failure (SGTF) marker or detection of N501Y amino acid substitution (Fig. 3). This was the case for samples from regions S1, S3, and C2, where detection of the lineage B.1.1.7 in clinical samples occurred as of the fifth week.
Fig. 3.
Frequency of nucleotide substitutions and deletions related with B.1.1.7 linage according to the sampling week. *Clinical frequency based on data obtained from the Spanish Ministry of Health according to the results obtained by S-gene target failure (SGTF) marker or detection of N501Y mutation.
4. Conclusions
SARS-CoV-2 has created a pandemic scenario unprecedented in modern times. The rapid spread of this virus together with the appearance of emerging linages has also mobilized the scientific community like never before. Its detection in wastewater has been very helpful for the epidemiological study in large populations and is currently being implemented worldwide. However, few studies using mass sequencing have been published.
-
•
The present study describes the mutations found in SARS-CoV-2 genomes isolated from wastewater in 14 different regions of Spain. This is the first study carried out in Spain that analyzes the diversity of SARS-CoV-2 present in wastewater in the three epidemiological waves which occurred between 2020 and 2021.
-
•
These results confirm the potential of sewage sequencing to detect new mutations and lineages of SARS-CoV-2, which is of utmost relevance for the monitoring efforts of emerging vaccine-escape SARS-CoV-2 mutants in the forthcoming post-vaccination era.
-
•
Genomic sequencing of viruses found in wastewater provides complementary results to those of clinical laboratories, as has been demonstrated in various ways such as the confirmation of the initial detection of low number of reads on genomes from clinical specimens that was later confirmed in wastewater samples; the detection of amino acid substitutions in the spike protein weeks or months before their discovery in clinical samples; or the known amino acid substitutions in the spike protein detected for the first time in Spain.
-
•
This technique provides complementary information for SARS-CoV-2 surveillance, allowing both the control of lineages including VOC and VOI already described and the detection and control of new emerging lineages.
-
•
This data supports the hypothesis that the study of wastewater using high-throughput sequencing techniques is a useful and effective tool that can be implemented worldwide in support of public health for the epidemiological control of SARS-CoV-2.
CRediT authorship contribution statement
Alba Pérez-Cataluña: Investigation, Formal analysis, Writing – review & editing, Conceptualization. Álvaro Chiner-Oms: Investigation, Formal analysis, Writing – review & editing. Enric Cuevas-Ferrando: Investigation, Formal analysis, Writing – review & editing. Azahara Díaz-Reolid: Investigation, Formal analysis, Writing – review & editing. Irene Falcó: Resources, Writing – review & editing. Walter Randazzo: Writing – review & editing. Inés Girón-Guzmán: Resources, Writing – review & editing. Ana Allende: Writing – review & editing. María A. Bracho: Writing – review & editing. Iñaki Comas: Funding acquisition, Writing – review & editing. Gloria Sánchez: Conceptualization, Funding acquisition, Writing – review & editing.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
This study was supported by projects “VIRIDIANA” (AGL2017–82909/ AEI/FEDER, UE) and MCEC-WATER (PID2020–116789RB-C42/ AEI/FEDER, UE) funded by Spanish Ministry of Science, Innovation and Universities; CSIC (202070E101), Generalitat Valenciana (Covid_19-SCI), the European Union – NextGenerationEU (SGL2103034), the COVID-19 wastewater surveillance project (VATar COVID19), funded by the Spanish Ministry for the Ecological Transition and the Demographic Challenge and the Spanish Ministry of Health, the Instituto de Salud Carlos III (Ref COV20/00140), ERC TB-RECONNECT 101001038 funded by The European Union – NextGenerationEU and the Lagoon project (PROMETEO/2021/044). EC-F is recipient of a predoctoral contract from the MICINN, Call 2018 and AP-C was supported by a postdoctoral fellowship (APOSTD/21/292). We acknowledge NILSA, FACSA, and MITECO for authorizing the sampling. The authors thank Agustin Garrido Fernández, and Andrea Lopez de Mota for their technical support.
Footnotes
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.watres.2021.118007.
Appendix. Supplementary materials
References
- AAVV. A.D.E. In: Standard Methods for the Examination of Water and Wastewater. Rice E.W., Baird R.B., editors. American Water Works Association; Denver, CO: 2018. Section 9510 D. Virus concentration by aluminum hydroxide adsorption-precipitation, chapter detection of enteric viruses. [Google Scholar]
- Bar-Or I., Weil M., Indenbaum V., Bucris E., Bar-Ilan D., Elul M., Levi N., Aguvaev I., Cohen Z., Shirazi R., Erster O., Sela-Brown A., Sofer D., Mor O., Mendelson E., Zuckerman N.S. Detection of SARS-CoV-2 variants by genomic analysis of wastewater samples in Israel. Sci. Total Environ. 2021;789 doi: 10.1016/J.SCITOTENV.2021.148002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bivins A., North D., Ahmad A., Ahmed W., Alm E., Been F., Bhattacharya P., Bijlsma L., Boehm A.B., Brown J., Buttiglieri G., Calabro V., Carducci A., Castiglioni S., Cetecioglu Gurol Z., Chakraborty S., Costa F., Curcio S., De Los Reyes F.L., Delgado Vela J., Farkas K., Fernandez-Casi X., Gerba C., Gerrity D., Girones R., Gonzalez R., Haramoto E., Harris A., Holden P.A., Islam M.T., Jones D.L., Kasprzyk-Hordern B., Kitajima M., Kotlarz N., Kumar M., Kuroda K., La Rosa G., Malpei F., Mautus M., McLellan S.L., Medema G., Meschke J.S., Mueller J., Newton R.J., Nilsson D., Noble R.T., Van Nuijs A., Peccia J., Perkins T.A., Pickering A.J., Rose J., Sanchez G., Smith A., Stadler L., Stauber C., Thomas K., Van Der Voorn T., Wigginton K., Zhu K., Bibby K. Wastewater-based epidemiology: global collaborative to maximize contributions in the fight against COVID-19. Environ. Sci. Technol. 2020 doi: 10.1021/acs.est.0c02388. [DOI] [PubMed] [Google Scholar]
- Chan J.F.W., Kok K.H., Zhu Z., Chu H., To K.K.W., Yuan S., Yuen K.Y. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 2020;9:221–236. doi: 10.1080/22221751.2020.1719902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crits-Christoph A., Kantor R.S., Olm M.R., Whitney O.N., Al-Shayeb B., Lou Y.C., Flamholz A., Kennedy L.C., Greenwald H., Hinkle A., Hetzel J., Spitzer S., Koble J., Tan A., Hyde F., Schroth G., Kuersten S., Banfield J.F., Nelson K.L. Genome sequencing of sewage detects regionally prevalent SARS-CoV-2 variants. MBio. 2021;12 doi: 10.1128/mbio.02703-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dharmadhikari T., Rajput V., Yadav R., Boargaonkar R., Patil D., Kale S., Kamble S.P., Dastager S.G., Dharne M.S. High throughput sequencing based direct detection of SARS-CoV-2 fragments in wastewater of Pune, West India. Sci. Total Environ. 2021 doi: 10.1016/j.scitotenv.2021.151038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drosten C., Günther S., Preiser W., van der Werf S., Brodt H.R., Becker S., Rabenau H., Panning M., Kolesnikova L., Fouchier R.A.M., Berger A., Burguière A.M., Cinatl J., Eickmann M., Escriou N., Grywna K., Kramme S., Manuguerra J.C., Müller S., Rickerts V., Stürmer M., Vieth S., Klenk H.D., Osterhaus A.D.M.E., Schmitz H., Doerr H.W. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003;348:1967–1976. doi: 10.1056/nejmoa030747. [DOI] [PubMed] [Google Scholar]
- Grubaugh N.D., Gangavarapu K., Quick J., Matteson N.L., De Jesus J.G., Main B.J., Tan A.L., Paul L.M., Brackney D.E., Grewal S., Gurfield N., Van Rompay K.K.A., Isern S., Michael S.F., Coffey L.L., Loman N.J., Andersen K.G. An amplicon-based sequencing framework for accurately measuring intrahost virus diversity using PrimalSeq and iVar. Genome Biol. 2019;20:8. doi: 10.1186/s13059-018-1618-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herold M., d’Hérouël A.F., May P., Delogu F., Wienecke-Baldacchino A., Tapp J., Walczak C., Wilmes P., Cauchie H.M., Fournier G., Ogorzaly L. Genome Sequencing of SARS-CoV-2 allows monitoring of variants of concern through wastewater. Water (Basel) 2021;13:3018. doi: 10.3390/w13213018. [DOI] [Google Scholar]
- Izquierdo-Lara R., Elsinga G., Heijnen L., Munnink B.B.O., Schapendonk C.M.E., Nieuwenhuijse D., Kon M., Lu L., Aarestrup F.M., Lycett S., Medema G., Koopmans M.P.G., de Graaf M. Monitoring SARS-CoV-2 circulation and diversity through community wastewater sequencing, the Netherlands and Belgium. Emerg. Infect. Dis. 2021;27:1405–1415. doi: 10.3201/eid2705.204410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ksiazek T.G., Erdman D., Goldsmith C.S., Zaki S.R., Peret T., Emery S., Tong S., Urbani C., Comer J.A., Lim W., Rollin P.E., Dowell S.F., Ling A.E., Humphrey C.D., Shieh W.J., Guarner J., Paddock C.D., Rota P., Fields B., DeRisi J., Yang J.Y., Cox N., Hughes J.M., LeDuc J.W., Bellini W.J., Anderson L.J. A novel coronavirus associated with severe acute respiratory syndrome. N. Engl. J. Med. 2003;348:1953–1966. doi: 10.1056/nejmoa030781. [DOI] [PubMed] [Google Scholar]
- La Rosa G., Brandtner D., Mancini P., Veneri C., Ferraro G.B., Bonadonna L., Lucentini L., Suffredini E. Key SARS-CoV-2 mutations of alpha, gamma, and eta variants detected in Urban Wastewaters in Italy by long-read amplicon sequencing based on nanopore technology. Water. 2021;13:2503 13. doi: 10.3390/W13182503. [DOI] [Google Scholar]
- Li H., Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H., Handsaker B., Wysoker A., Fennell T., Ruan J., Homer N., Marth G., Abecasis G., Durbin R. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–2079. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.Journal. 2011;17:10. doi: 10.14806/ej.17.1.200. [DOI] [Google Scholar]
- Medema G., Heijnen L., Elsinga G., Italiaander R., Brouwer A. Presence of SARS-Coronavirus-2 RNA in sewage and correlation with reported COVID-19 prevalence in the early stage of the epidemic in the Netherlands. Environ. Sci. Technol. Lett. 2020;7:511–516. doi: 10.1021/acs.estlett.0c00357. [DOI] [PubMed] [Google Scholar]
- Mullen, J., Tsueng, G., Latif, A.A., Alkuzweny, M., Cano, M., Haag, E., Zhou, J., Zeller, M., Matteson, N., Andersen, K.G., Wu, C., Su, A.I., Gangavarapu, K., Hughes, L.D., 2020. https://outbreak.info/.
- Nemudryi A., Nemudraia A., Wiegand T., Surya K., Buyukyoruk M., Cicha C., Vanderwood K.K., Wilkinson R., Wiedenheft B. Temporal detection and phylogenetic assessment of SARS-CoV-2 in municipal wastewater. Cell Rep. Med. 2020;1 doi: 10.1016/j.xcrm.2020.100098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pérez-Cataluña A., Cuevas-Ferrando E., Randazzo W., Falcó I., Allende A., Sánchez G. Comparing analytical methods to detect SARS-CoV-2 in wastewater. Sci. Total Environ. 2021;758 doi: 10.1016/j.scitotenv.2020.143870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polo D.D., Quintela-Baluja M., Corbishley A., Jones D.L., Singer A.C., Graham D.W., Romalde P.J.L. Making waves: wastewater-based epidemiology for SARS-CoV-2 – Developing robust approaches for surveillance and prediction is harder than it looks. Water Res. 2020;186 doi: 10.1016/j.watres.2020.116404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quick, J., 2020. nCoV-2019 sequencing protocol v3 (LoCost). protocols.io https://protocols.io/view/ncov-2019-sequencing-protocol-v3-locost-bh42j8ye.
- Rambaut, A., Loman, N., Pybus, O., Barclay, W., Barrett, J., Carabelli, A., Connor, T., Peacock, T., Robertson, D.L., Volz, E., (CoG-UK)9., on behalf of C.-19 G.C.U., 2020. Preliminary genomic characterisation of an emergent SARS-CoV-2 lineage in the UK defined by a novel set of spike mutations - SARS-CoV-2 coronavirus /nCoV-2019 Genomic Epidemiology - Virological [WWW Document]. URL https://virological.org/t/preliminary-genomic-characterisation-of-an-emergent-sars-cov-2-lineage-in-the-uk-defined-by-a-novel-set-of-spike-mutations/563 (accessed 6.14.21).
- Randazzo W., Cuevas-Ferrando E., Sanjuán R., Domingo-Calap P., Sánchez G. Metropolitan wastewater analysis for COVID-19 epidemiological surveillance. Int. J. Hyg. Environ. Health. 2020;230:1438–4639. doi: 10.1016/j.ijheh.2020.113621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Randazzo W., Truchado P., Cuevas-Ferrando E., Simón P., Allende A., Sánchez G. SARS-CoV-2 RNA in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 2020;181 doi: 10.1016/j.watres.2020.115942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rios G., Lacoux C., Leclercq V., Diamant A., Lebrigand K., Lazuka A., Soyeux E., Lacroix S., Fassy J., Couesnon A., Thiery R., Mari B., Pradier C., Waldmann R., Barbry P. Monitoring SARS-CoV-2 variants alterations in Nice neighborhoods by wastewater nanopore sequencing. Lancet Reg. Heal. Eur. 2021;0 doi: 10.1016/J.LANEPE.2021.100202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shang Z., Chan S.Y., Liu W.J., Li P., Huang W. Recent insights into emerging coronavirus: SARS-CoV-2. ACS Infect. Dis. 2020 doi: 10.1021/acsinfecdis.0c00646. [DOI] [PubMed] [Google Scholar]
- Singh, J., Samal, J., Kumar, V., Sharma, J., Agrawal, U., Ehtesham, N.Z., Sundar, D., Rahman, S.A., Hira, S., Hasnain, S.E., 2021. Structure-function analyses of new SARS-CoV-2 Variants B.1.1.7, B.1.351 and B.1.1.28.1: clinical, diagnostic, therapeutic and public health implications. 10.3390/v13030439. [DOI] [PMC free article] [PubMed]
- Xiao F., Tang M., Zheng X., Liu Y., Li X., Shan H. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology. 2020;158:1831–1833.e3. doi: 10.1053/j.gastro.2020.02.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaki A.M., van Boheemen S., Bestebroer T.M., Osterhaus A.D.M.E., Fouchier R.A.M. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012;367:1814–1820. doi: 10.1056/nejmoa1211721. [DOI] [PubMed] [Google Scholar]
- Zhou W., Xu C., Wang P., Anashkina A.A., Jiang Q. Impact of mutations in SARS-COV-2 spike on viral infectivity and antigenicity. Brief Bioinform. 2021;2021:1–8. doi: 10.1093/bib/bbab375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu N., Zhang D., Wang W., Li X., Yang B., Song J., Zhao X., Huang B., Shi W., Lu R., Niu P., Zhan F., Ma X., Wang D., Xu W., Wu G., Gao G.F., Tan W. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020;382:727–733. doi: 10.1056/nejmoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.