Abstract
MEX3A is an RNA-binding protein that mediates mRNA decay through binding to 3′ untranslated regions. However, its role and mechanism in clear cell renal cell carcinoma remain unknown. In this study, we found that MEX3A expression was transcriptionally activated by ETS1 and upregulated in clear cell renal cell carcinoma. Silencing MEX3A markedly reduced clear cell renal cell carcinoma cell proliferation in vitro and in vivo. Inhibiting MEX3A induced G1/S cell-cycle arrest. Gene set enrichment analysis revealed that E2F targets are the central downstream pathways of MEX3A. To identify MEX3A targets, systematic screening using enhanced cross-linking and immunoprecipitation sequencing, and RNA-immunoprecipitation sequencing assays were performed. A network of 4,000 genes was identified as potential targets of MEX3A. Gene ontology analysis of upregulated genes bound by MEX3A indicated that negative regulation of the cell proliferation pathway was highly enriched. Further assays indicated that MEX3A bound to the CDKN2B 3′ untranslated region, promoting its mRNA degradation. This leads to decreased levels of CDKN2B and an uncontrolled cell cycle in clear cell renal cell carcinoma, which was confirmed by rescue experiments. Our findings revealed that MEX3A acts as a post-transcriptional regulator of abnormal cell-cycle progression in clear cell renal cell carcinoma.
Keywords: MEX3A, CDKN2B, eCLIP-seq, RIP-seq, G1/S arrest, RB/E2F, clear cell renal cell carcinoma
Graphical abstract

We described the comprehensive downstream targets of MEX3A using RIP-seq and eCLIP-seq. The data revealed that oncogene MEX3A was transcriptionally activated and mainly regulated G1/S transition in ccRCC, thus providing a novel molecular basis for clinical diagnosis and treatment.
Introduction
Renal cell carcinoma (RCC), originating from nephric epithelial cells, ranks as the sixth most common cancer in males and ninth in females.1 Clear cell RCC (ccRCC) is the most common type of RCC and accounts for approximately 75% of all cases.2 Recent studies have substantially increased our understanding of the genomic complexity and molecular features of ccRCC, such as chromosomal alterations, phosphoinositide 3-kinase (PI3K)-protein kinase B (AKT)-mammalian target of rapamycin (mTOR) pathway mutations, loss of oxygen sensing, and loss of cell-cycle regulation.3, 4, 5 Despite advances in both diagnostic and therapeutic strategies in ccRCC, very few prognostic markers for ccRCC are available in current clinical practice.6,7 Hence, further study is required to understand the molecular pathogenesis of ccRCC.
Mex-3 RNA-binding family member A (MEX3A), an RNA-binding protein containing two KH domains and one RING finger, is a post-translational repressor that is dysregulated in several tumors.8,9 Previous studies have reported that MEX3A was upregulated and associated with poor patient outcomes in multiple cancers, such as esophageal squamous cell carcinoma (ESCC), lung adenocarcinoma (LUAD), and glioma.10, 11, 12 Functionally, MEX3A acts as a facilitator of tumorigenesis by promoting cell-cycle progression and sustaining cell viability, but preventing cell apoptosis. For example, in pancreatic ductal adenocarcinoma (PDAC), MEX3A expression was higher in the G1 phase than in the S phase, and silencing MEX3A induced G1/S cell-cycle arrest.13 In triple-negative breast cancer, inhibition of MEX3A led to cell apoptosis and an increase in the proportion of cells in the G2/M phase.14 By contrast, in colorectal carcinoma, overexpression of MEX3A reduced the Caco-2 cell population at G0/G1 phase and increased the S phase population.15 Recently, although high MEX3A expression was reported to correlate positively with poor prognosis for patients with ccRCC,16 the exact molecular mechanisms by which MEX3A promotes ccRCC tumorigenesis remain largely unexplored.
In the present study, we aimed to explore the function and mechanism of MEX3A in ccRCC. We observed that MEX3A expression was upregulated in ccRCC, and a higher level of MEX3A correlated with poorer overall survival (OS). Functional assessment revealed that disruption of MEX3A expression impaired ccRCC cell proliferation, colony formation, cell-cycle progression, and migration. Using enhanced cross-linking and immunoprecipitation sequencing (eCLIP-seq) and RNA immunoprecipitation sequencing (RIP-seq) assays, we systematically analyzed the downstream targets of MEX3A in ccRCC and proved that MEX3A affected CDKN2B (cyclin dependent kinase inhibitor 2B) expression by modulating its mRNA degradation directly, thereby disturbing the cell cycle. In summary, we identified MEX3A as an important cell-cycle regulator, and provided new insights into the ccRCC tumor biology.
Results
MEX3A is upregulated in ccRCC
We first interrogated The Cancer Genome Atlas (TCGA) RNA sequencing (RNA-seq) data of kidney renal clear cell carcinoma (KIRC) using the University of California, Santa Cruz (UCSC) Xena browser, and found that MEX3A was significantly overexpressed in ccRCC samples compared with adjacent normal samples (Figure 1A). Further analysis of Gene Expression Omnibus (GEO) datasets (GSE16449 and GSE86095) showed a consistent expression pattern of MEX3A in ccRCC (Figures 1B and 1C). Clinically, although MEX3A expression did not correlate with ccRCC pathological stage (Figures S1A and S1B), higher MEX3A levels were associated significantly with shorter OS (Figure 1D). In addition, we randomly chose 10 pairs of ccRCC tissues to perform a western blotting assay. The results showed that MEX3A levels were upregulated significantly in six ccRCC samples, showed no remarkable alterations in three ccRCC samples, and were downregulated in one ccRCC sample (Figure 1E). This finding was consistent with the above results, which implied that MEX3A might serve as a prognostic factor for patients with ccRCC.
Figure 1.
MEX3A is significantly upregulated and activated by ETS1 in ccRCC
(A and B) Increased MEX3A expression in tumors compared with normal tissues. Data were obtained from the TCGA and GEO databases. (C) Before-after graph showing MEX3A expressions in paired samples (GSE86095). Tumors showed a higher value. (D) OS time of patients with ccRCC categorized according to MEX3A mRNA levels (p = 0.0251). (E) The protein expression of MEX3A in 10 pairs of ccRCC samples. N, normal; T, tumor. (F) IGV line plots of the H3K27ac, H3K4me3, and ETS1 ChIP-seq results in the indicated samples. Signal values of normalized peak intensity are shown in the upper left corner. Gene count values of tissue samples are shown in the upper right corner. All ChIP-seq data were from GSE86095 or GSE78113 (786-O H3K27ac). (G) Scatterplots showing the correlation between ATAC-seq peaks and MEX3A mRNA expression. Each dot is a TCGA ccRCC sample. The RNA expression and ATAC-seq data of ccRCC from the TCGA were analyzed. (H) mRNA levels of ETS1 and MEX3A after ETS1 knockdown. The means ± SD are shown, n = 3. (I) Western blotting validating the change in MEX3A levels after silencing ETS1. (J) Luciferase reporter assays in 786-O cells. The means ± SD are shown, n = 3. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.01, ∗∗∗∗p < 0.0001; n.s., not significant.
To determine the molecular mechanisms underlying MEX3A upregulation in ccRCC, we first categorized TCGA ccRCC samples into groups of “wild-type” (WT) and “mutation” based on the absence or presence of the von Hippel-Lindau (VHL) mutation, the most common and characteristic genetic change of ccRCC. However, no significant difference in MEX3A expression was observed between the groups, indicating that upregulation of MEX3A in ccRCC was independent of the VHL mutation status (Figures S1C and S1D). We also investigated possible genetic alterations of MEX3A itself by studying exome sequencing data of primary ccRCC samples using the cBioportal pan-Cancer database.17 Notably, a very low frequency (less than 1%) of either MEX3A mutation or amplification was observed in ccRCCs from various cohorts. These results suggested that dysregulation of MEX3A was not attributed to genetic disorders in ccRCC (Figure S1E).
Apart from genetic aberrations, epigenetic modifications, such as DNA methylation, also play a major role in reprogramming gene expression. Hence, we analyzed the DNA methylation status at the MEX3A gene locus using Infinium Human Methylation 450K data from the TCGA to determine whether DNA methylation affected MEX3A expression in ccRCC. Instead of the promoter regions, lower methylation levels were found in the second exon region of MEX3A among the majority of cancer samples compared with that in normal samples (Figure S1F). However, the DNA methylation levels in these regions were not inversely correlated with MEX3A expression levels (Figure S1G).
Transcriptional control represents another molecular mechanism pertaining to gene expression regulation. In search of potential transcriptional changes underlying MEX3A hyper-activation, we retrieved chromatin immunoprecipitation sequencing (ChIP-seq) data from GES86095 and found that strong coincident H3K27ac and H3K4me3 signals were deposited at both the promoter (region 2) and second exon (region 1) regions of MEX3A in all ccRCC tumor samples. In contrast, such H3K27ac or H3K4me3 signals were largely diminished in paired normal samples (Figure 1F).18 The H3K27ac mark is widely accepted as an indicator of active cis-regulatory DNA and the H3K4me3 signal is considered as a marker of promoters. Therefore, we suspected that hyper-activation of MEX3A was triggered at the transcriptional level. Unsurprisingly, we demonstrated lower MEX3A expression in all paired normal samples through further interrogating the RNA-seq data (Figure 1F). Assay for transposase-accessible chromatin using sequencing (ATAC-seq) data from the TCGA showed a strong signal in both regions of MEX3A in ccRCC (Figure 1F). Further analysis of the ATAC-seq and RNA-seq data showed a highly positive correlation between ATAC-seq peak intensity and the expression level of MEX3A in genomic region 2 rather than region 1 (Figure 1G). Hence, we were particularly interested in unveiling transcription factors that directly regulate the promoter region (region 2) of MEX3A. Fortunately, we found that the ChIP-seq data of ETS1 (encoding ETS proto-oncogene 1, transcription factor) in 786-O cells showed a strong binding peak in the promoter region of MEX3A (Figure 1F). Analysis of the TCGA database showed that ETS1 is a highly expressed transcription factor in ccRCC (Figure S2A). Mikami et al. also confirmed that ETS1 mRNA expression was upregulated in ccRCC samples.19 Functional assay revealed that overexpressing ETS1 enhanced cell proliferation and migration in ccRCC cell lines, which indicated that ETS1 is an oncogenic transcription factor in ccRCC.20,21 To validate whether ETS1 regulates MEX3A expression depending on binding to the MEX3A promoter region in ccRCC, we constructed a short hairpin RNA (shRNA) to decrease ETS1 expression in ccRCC cell lines. The results showed that silencing ETS1 in 786-O and A498 cells significantly inhibited MEX3A expression (Figures 1H and 1I). However, no significant alterations in MEX3A expression were found after knocking down ETS1 in ACHN cells (Figures S2B and S2C). Referring to a previous study, we constructed two pGL3-enhancer vectors, one containing the WT MEX3A promoter region and another with mutations (MUT) in the ETS1 binding sequence (from GGAA to TCGA), to detect the binding of ETS1 in ccRCC cell lines (Figure 1J).22 The results showed that the firefly luciferase signal was remarkably reduced after silencing ETS1 expression in 786-O cells transfected with the WT plasmid (Figure 1J). However, no significant difference in the firefly luciferase signal was found in the MUT groups (Figure 1J). In summary, instead of gene mutation, amplification, or DNA methylation, the main reason for the high expression of MEX3A was transcriptional activation. Our results also proved that ETS1 activated MEX3A transcription in ccRCC.
Silencing MEX3A suppresses ccRCC cell proliferation, migration, stemness, and tumor growth
To test the functional roles of aberrant MEX3A expression, MEX3A was silenced using small interfering RNA (siRNA) transfection from a lentivirus vector in ccRCC cells. The efficiency of siRNA interference was confirmed using western blotting (Figure 2A). According to previous studies, MEX3A is considered to be a stem cell marker. Therefore, we tested the impact of silencing MEX3A on cell stemness. Sphere-formation assays showed that silencing of MEX3A depressed cell stemness significantly (Figures 2B and 2C). In addition, the quantitative real-time reverse transcription PCR (qRT-PCR) results revealed that knocking down MEX3A remarkably downregulated the expression of stemness-related genes, including OCT4 and NANOG (Figure 2D). We then explored the effect of MEX3A on cell growth. Inhibition of MEX3A expression impeded ccRCC cell viability in(3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) and colony formation assays (Figures 2E, 2F, and S3A). The effect of MEX3A on cell motility was also examined. Compared with that in the control groups, silencing MEX3A significantly decreased the cell migration ability of ccRCC cells (Figures 2G and S3B). Then, we overexpressed MEX3A in ccRCC cells, which markedly promoted cell viability in the colony formation assay (Figures 2H, 2I, and S3C). Similarity, upregulation of MEX3A promoted cell migration in A498 and 786-O cells (Figure S3D). To explore the effect of knockdown of MEX3A on tumorigenesis in vivo, constitutive pLKO.1-MEX3A-sh1-expressing or pLKO.1-Control-expressing ACHN cells were injected into mice. Tumors formed from MEX3A-silenced cells were markedly smaller than those formed from the control cells (Figure 2J–2L). Overall, these results demonstrated that MEX3A promotes cell viability and tumor growth in ccRCC.
Figure 2.
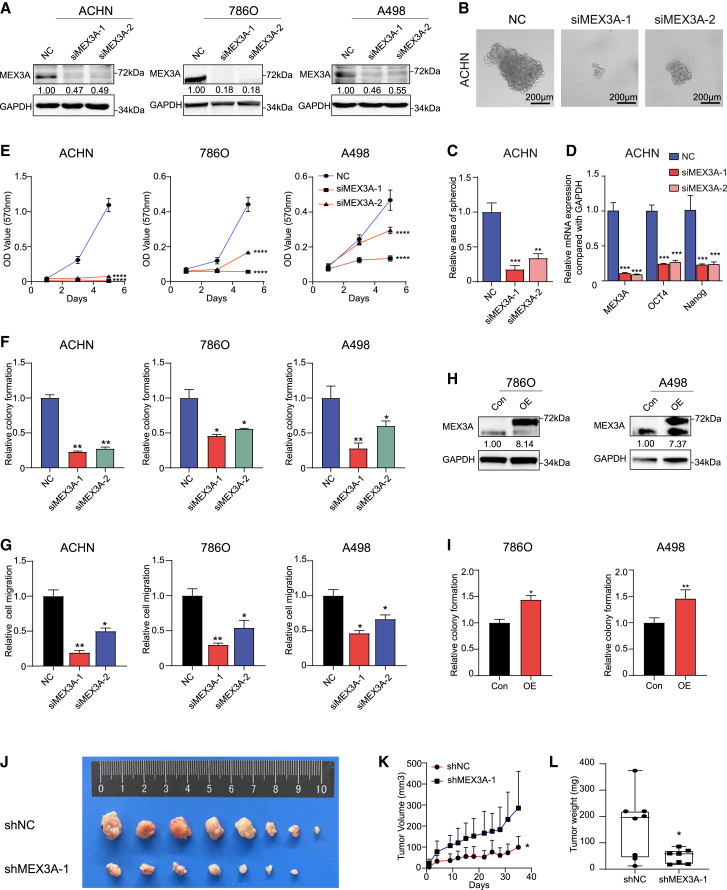
Inhibition of MEX3A suppresses ccRCC cell viability
(A) MEX3A silencing by siRNAs was verified by western blotting in different ccRCC cell lines. (B and C) Cell stemness was detected by silencing MEX3A in ACHN cells. Mean ± SD, n = 3. Scale bar, 200 μm. (D) The mRNA expression of stemness markers after silencing MEX3A in ACHN cells. Knockdown of MEX3A inhibited (E) MTT, (F) colony formation, and (G) cell migration in ccRCC cell lines. Mean ± SD, n = 3. (H) Western blotting validation of MEX3A overexpression. (I) Overexpressed (OE) MEX3A promoted cell colony formation compared with control group in ccRCC cell lines. Mean ± SD, n = 3. (J) Xenograft images, (K) tumor growth, and (L) tumor weight of xenografts expressing either scramble shRNA or MEX3A-shRNA in mice. Means ± SD are shown, n = 8, ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001.
Inhibition of MEX3A induces G1/S arrest in ccRCC
To gain a further insight into MEX3A-regulated gene and signaling pathways, RNA-seq was performed to explore the target genes affected by MEX3A. A total of 488 differentially expressed genes were identified, including 310 downregulated and 178 upregulated genes, in MEX3A-silenced ACHN cells (p < 0.05, count ≥ 100, and log2 fold-change [FC] ≥ 1 or ≤ −1; Figure 3A). Certain genes from the RNA-seq data were chosen for qRT-PCR verification and the results showed that their expression levels were significantly altered after silencing MEX3A in ACHN cells (Figure S4A). Gene set enrichment analysis (GSEA) of TCGA RNA-seq data showed that MEX3A expression correlated highly with several pathways, including epithelial-mesenchyme transition (EMT), Notch signaling, and DNA repair. To gain a further understanding of MEX3A highly correlated pathways, we combined the GSEA results of RNA-seq in ACHN cells, TCGA RNA-seq data of ccRCC, and transcriptome sequencing data in GSE86095, and found that higher expression of MEX3A was associated with E2F-targets, MYC-targets, and G2/M checkpoint pathways in ccRCC (Figures 3B, 3C, and S4B). Accordingly, we detected the cell-cycle alterations after silencing MEX3A in ccRCC cell lines. Compared with the control groups, a larger proportion of cells were in the G1 phase in ACHN, A498, and 786-O cells (Figures 3D, 3E, and S4C). In addition, monitoring of cell-cycle progression at the single-cell level in MEX3A-silenced ACHN cells demonstrated that MEX3A knockdown prolonged the G1 phase significantly (Figures 3F and 3G). The E2F transcription factor plays an important role in controlling the transition from the G1 to S phase.23 However, E2F1 protein levels were not significantly changed in ccRCC cells (Figure S4D). Considering that the phosphorylation of retinoblastoma 1 (RB) is crucial to E2F transcription factor activity,24 we detected the phosphorylation of RB at S795 and S807/811 after silencing MEX3A in ccRCC cells. The results showed that silencing MEX3A decreased the levels of phosphorylated RB, cyclin A2, and cell-dependent kinase (CDK) 1 in ccRCC, which indicated that silencing MEX3A induced G1/S arrest by regulating the RB/E2F pathway (Figure S4D). A study reported that CDK6 regulates G1/S transition and is a downstream target of MEX3A.13 Hence, we detected the levels of CDK1, CDK2, CDK4, and CDK6 in ccRCC cell lines. The results showed that CDK1 levels decreased significantly after knocking down MEX3A in ccRCC cell lines (Figure S4E). However, the levels of CDK2, CDK4, or CDK6 were not consistent after silencing MEX3A using two different siRNAs or different cell lines, which might be because of the different siRNA sequence or cell specificity (Figure S4E). Despite the different alterations of CDK4 or CDK6 induced by two different siRNAs of MEX3A, we found that both siRNAs induced G1/S arrest and inhibited the RB/E2F pathway in ccRCC cell lines. Hence, these results indicated that CDK6 or CDK4 are not the key proteins in MEX3A-mediated G1/S transition in ccRCC. In summary, these data indicated that RNA-binding protein MEX3A is a cell-cycle regulator and silencing MEX3A induced cell arrest at G1 phase by regulating the RB/E2F pathway in ccRCC.
Figure 3.
MEX3A is a cell regulator in ccRCC
(A) Heatmap of the RNA-seq results in ACHN cells. (B) Heatmap of the GSEA results of top 20 enriched hallmark pathways in the MEX3A high-expression groups (top 25%) versus MEX3A low-expression (bottom 25%) ccRCC samples from the TCGA samples. MEX3A-Tumor groups (17621953T, 70528835T, 74575859T) versus MEX3A-Normal groups (17621953N, 70528835N, 74575859N) from GSE86095. (C) Individual GSEA plots of E2F-target pathways as in (B). (D) Fluorescence-activated cell sorting (FACS) analysis showing the DNA content (PI) of ACHN cells after silencing MEX3A. (E) Bar graphs show the percentage of cells in G1, S, and G2 phase of the cell cycle by FACS analysis, as in (D). Mean ± SD, n = 3. (F) Representative images of cells at the indicated times post transfection with the NC (top) or MEX3A-siRNAs (bottom). Green, S and G2/M phase; yellow, G1 phase. Scale bar, 50 μm. (G) Bar graphs showing the time that the cells stalled in G1 phase after mitosis. n = 10. ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001.
Screening for the downstream targets of MEX3A
To further understand which RNA sequences are directly recognized by MEX3A, we first established an ACHN cell line stably expressing N-terminal FLAG-tagged MEX3A and performed single-nucleotide resolution eCLIP (Figure S5A). The eCLIP sequencing results showed that a total of 7,350 peaks were identified by the FLAG antibody, and the annotation of these eCLIP peaks revealed that about 68% of MEX3A binding sites were located in the 3′ UTR of targets (Figure 4A). Analyses of these peaks showed that most of the MEX3A-captured RNAs were protein-coding RNAs (Figure 4B). We used eCLIP-seq to find the downstream targets of MEX3A in ccRCC; however, the repeatability of this technique is not as good as RIP-seq. To eliminate this limitation of the eCLIP-seq assay, we also performed RIP-seq to detect the downstream RNAs that interacted with MEX3A. Analyses of the RIP-seq data showed that 4,713 RNAs were enriched by the MEX3A protein compared with the immunoglobulin (Ig) G group, and the majority of the MEX3A-captured RNAs encoded functional proteins; less than 3% of encoded pseudogenes, antisense, long non-coding RNAs (lncRNAs), and others (Figure 4C, immunoprecipitation (IP) gene count ≥100, log FC ≤ 1). Studies had found that CDX2 (encoding caudal type homeobox 2), CDK6, and CCL2 (encoding C-C motif chemokine ligand 2) were the downstream targets of MEX3A.10,11,15 Except for CDX2, which is not expressed in ACHN cells, both the eCLIP-seq and RIP-seq data showed that MEX3A could bind to the 3′ UTR of CDK6 and CCL2 mRNA (Figures S5B and S5C). Then, we performed motif analyses of the eCLIP peaks located in the 3’ UTRs, which showed that 5′-UUUAUAAA-3′ was the MEX3A high-affinity recognition motif (Figure 4D). Interestingly, MEX3A binding sites were mainly enriched in 3′ UTR region near the end of the CDS region and stop codon (Figure 4E). By comparing the distribution of peaks of upregulated or downregulated genes after silencing MEX3A, we found that the distribution of peaks was altered (Figure 4F). However, we could not think of a reasonable explanation for this phenomenon, thus this aspect requires further exploration.
Figure 4.
Identification of direct downstream targets of MEX3A in ccRCC
(A) Distribution of MEX3A eCLIP-binding sites across different regions of gene bodies. (B and C) Distribution of MEX3A targets in eCLIP-Seq or RIP-seq experiments. (D) Weblogos depicting the significant MEX3A eCLIP-binding motifs in ACHN cells identified through de novo motif analysis. (E) Metagene profiles of enrichment of all peaks of MEX3A. (F) Metagene profiles of enrichment of alterations genes after silencing MEX3A. (G) Overlap of RIP-seq, RNA-seq, and eCLIP-seq in ACHN cells. (H and I) Bubble chart showing the top 10 enriched pathways in upregulated genes after silencing MEX3A in ACHN cells (GO term).
MEX3A was identified as an RNA-binding protein that degrades its target mRNAs via binding their 3′ UTRs.15 Therefore, we chose genes that could be both enriched by eCLIP-seq and RIP-seq and whose expression levels were changed after silencing MEX3A for study. The results showed that 79 genes were directly regulated by MEX3A (Figures 4G, 47 out of 178 upregulated; 32 out of 310 downregulated). Then, Gene Ontology (GO) analyses of these upregulated and downregulated genes were performed. The GO results for the upregulated genes suggested an overrepresentation of genes involved in tissue development and the negative regulation of cell proliferation (Figure 4H). GO analysis of the downregulated genes' results showed that bone development and cartilage development were highly enriched (Figure 4I). Considering that MEX3A correlates highly with the cell cycle, the genes associated with negative regulation of cell proliferation pathways were regarded as major research objects.
MEX3A regulates cell cycle via degrading CDKN2B mRNA
Next, we decided to verify cell cycle-related targets of MEX3A in ccRCC cell lines using RIP qPCR. The genes CDKN1A, CDKN2B, E2F7, MAX, DCUN1D3, and BTG2, which were enriched by MEX3A, were further validated in ACHN, A498, and 786-O cells. The results showed that CDKN2B, E2F7, and BTG2 were specifically enriched by the anti- FLAG antibody in several cells (Figures 5A–5C). The qRT-PCR results showed that CDKN2B and BTG2 were both upregulated in ACHN and A498 cells after silencing MEX3A (Figure 5D). However, E2F7 was only slightly increased in ACHN cells (Figure 5D). CDKN2B is a cell-cycle regulator that controls G1/S transition and regulates the RB/E2F pathway.25 Therefore, we constructed a psi-Check2 vector containing the 3′ UTR sequence of CDKN2B, and the luciferase assay results showed that overexpressing MEX3A inhibited the firefly luciferase activity significantly, which suggested that MEX3A degraded CDKN2B mRNA by binding to its 3′ UTR (Figure 5E). After 6 h of actinomycin D (ActD) treatment, the rate of CDKN2B mRNA degradation in the MEX3A-silenced group was significantly slower than that in the control group (Figure 5F). As shown in Figure 5G, we synthesized three biotin-labeled RNA probes (negative control [NC], WT, and mutant [Mut]), to detect binding between MEX3A and 3′ UTR region of CDKN2B. The RNA pull-down assays showed that MEX3A was enriched by the WT probes instead of other probes or beads, which indicated that MEX3A interacted with CDKN2B mRNA via recognizing the 5′-UUUAUAAA-3′ sequence (Figure 5G). Then, we tested the mRNA and protein expression levels after silencing MEX3A in ccRCC cells. The results showed that silencing MEX3A markedly increased the mRNA expression levels of CDKN2B and decreased MCM2 (encoding minichromosome maintenance complex component 2), MCM5, and CDK1 mRNA levels. Knockdown of CDKN2B reversed these variations induced by silencing MEX3A in ACHN and A498 cells (Figure 5H). In addition, western blotting assays revealed that silencing CDKN2B could reverse the MEX3A-mediated inhibition of the RB/E2F pathway in ACHN and A498 cells (Figure 6A). Then, cell-cycle assays confirmed that silencing CDKN2B decreased the inhibition on G1/S transition caused by knocking down MEX3A in ccRCC cells (Figures 6B, S5D, and S5E). MTT and colony formation assays showed similar effects of silencing CDKN2B in ccRCC cells (Figures 6C, S5F, and S5G).
Figure 5.
MEX3A directly binds to the 3′UTR of CDKN2B mRNA
(A and B) RIP-seq assays showing several targets enriched by MEX3A in different ccRCC cell lines. Mean ± SD, n = 3. (C) IGV line plots of MEX3A RIP-seq and eCLIP-seq in ACHN cells. Signal values of normalized peak intensity are shown in the upper left corner. (D) The mRNA expressions of CDKN2B, E2F7, and BTG2 were detected after silencing MEX3A in ccRCC cells. Mean ± SD, n = 3. (E) Luciferase reporter assays in ACHN and 293T cells. The 3′ UTR of CDKN2B was cloned into a reporter vector. Mean ± SD, n = 3. (F) RNA degradation rate after silencing MEX3A in ACHN cells. Mean ± SD, n = 3. (G) Diagram of three RNA probes and the western blotting results of RNA pull-down. (H) mRNA expression of MEX3A, CDKN2B, MCM2, MCM5, and CDK1 after MEX3A and CDKN2B knockdown in different ccRCC cell lines. Mean ± SD, n = 3. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001; n.s., not significant.
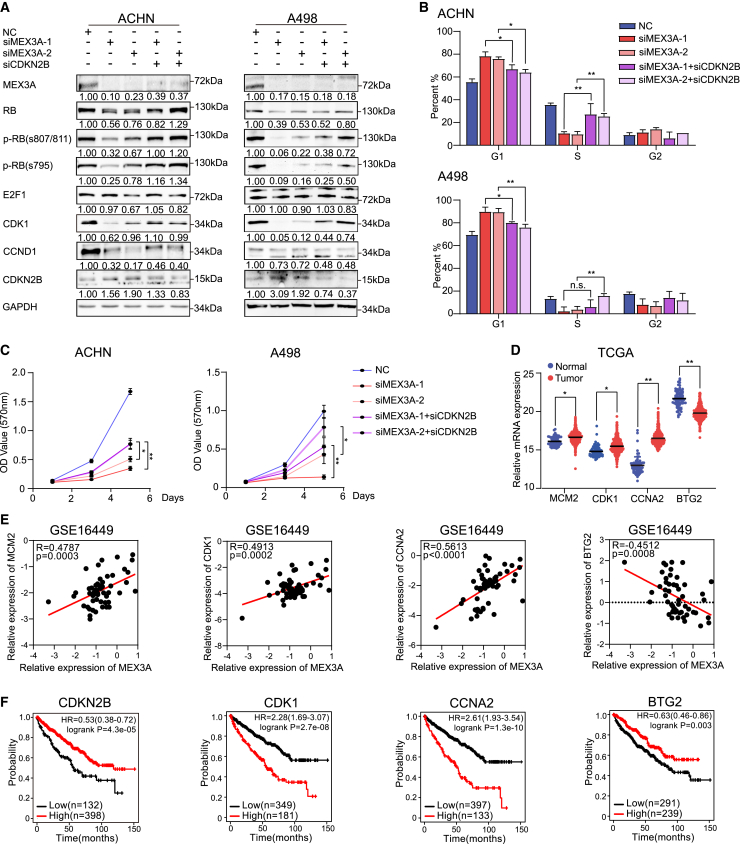
Figure 6.
MEX3A controls cell cycle via regulating CDKN2B mRNA decay in ccRCC
(A) Levels of MEX3A, CDKN2B, and E2F/RB-related proteins after MEX3A and CDKN2B knockdown. Inhibition of CDKN2B recovered the inhibition induced by silencing MEX3A on G1/S transition (B) and proliferation (C). (D) The expression of MCM2, CDK1, CCNA2, and BTG2 in TCGA. (E) The mRNA expression correlation between MEX3A and its downstream targets. (F) OS time of patients with ccRCC categorized according to mRNA levels.
Our results revealed that MEX3A promoted the expression of E2F targets via degrading CDKN2B mRNA. To better understand this relationship between MEX3A and E2F targets in vivo, we analyzed these gene expression correlations in the TCGA and GSE16449 datasets. Several representative results, including differential expression analysis, correlations analysis, and survival curve analysis, were presented. In detail, we found that the expression levels of CDK1, MCM2, and CCNA2 were increased significantly in the ccRCC tumor group compared with those in the control group (Figures 6D and S6A). Besides, there were positive correlations between MEX3A expression and the downstream effectors, including CDK1, MCM2, and CCNA2 (Figures 6E and S6B). In the above results, we mainly focused on the regulation between MEX3A and CDKN2B. Unfortunately, we did not find a negative correlation between MEX3A and CDKN2B in the TCGA or GSE16449 data. However, BTG2, another potential target of MEX3A, correlated negatively with MEX3A expression in GSE16449 (Figure 6E). Lastly, higher expression of CDK1 or CCNA2 was associated with a shorter survival time and higher expression of CDKN2B or BTG2 was associated with a longer survival time in the TCGA data (Figure 6F). Above all, these results revealed that MEX3A regulated G1/S transition and promoted the expression of E2F downstream targets by degrading CDKN2B mRNA expression and supported the view that MEX3A is crucial for cell-cycle regulation in ccRCC (Figure 7).
Figure 7.
Schematic summary of the role of MEX3A in ccRCC
Proposed working model showed MEX3A was activated by ETS1 and promoted G1/S transition by degrading target mRNAs, which regulates cell proliferation negatively.
Discussion
To date, research on MEX3A has concentrated mainly on its role in intestinal stem cells and tumor cells. In intestinal stem cells, MEX3A affects cell stemness in a post-transcriptional manner.26,27 Studies have found that MEX3A is upregulated in several tumors and regulates cell proliferation, migration, drug resistance, and tumor growth by regulating mRNA degradation or protein ubiquitylation.10,12,28,29 In this study, we found that MEX3A was overexpressed in ccRCC, which was caused by transcriptional activation. After a comprehensive analysis of public databases and ACHN RNA-seq data, we found that MEX3A mainly regulates the RB/E2F pathway in ccRCC. CDKN2B, located on chromosome 9p21, encodes a well-known cell regulator that controls G1/S transition in cell-cycle progression.25 Overexpression of CDKN2B significantly inhibited cell proliferation and decreased the phosphorylation level of RB s807/811 by directly binding CDK4 or CDK6.30 Studies have demonstrated that the RB/E2F pathway is critical for control of the cell cycle and tumor progression.31,32 pRB is phosphorylated on S807/811 during the S phase, and phosphorylation of RB's C-terminal serine (S795) destabilizes the interaction between RB and the E2F1-DP1 heterodimer.33,34 Upon phosphorylation of RB at S795 and S807/811, downstream targets of E2F, such as CCNA2, CDK1, and MCM2, will be transcriptionally activated.35 Our results revealed that silencing MEX3A inhibited the phosphorylation of RB at S795 and S807/811 through increasing CDKN2B expression and decreased the expression of CCNA2, CDK1, and MCM2, which indicated that inhibition of MEX3A induced G1/S arrest in ccRCC. Notably, loss of 9p21 is frequent and causes poorer survival in patients with ccRCC. However, 9p21 loss of heterozygosity does not lead to downregulation of CDKN2B.36 Higher CDKN2B expression correlated positively with a longer OS in patients with ccRCC, which indicated the anti-tumor role of CDKN2B in ccRCC. The results of the present study indicated that MEX3A regulates the cell cycle by degrading CDKN2B mRNA. Unfortunately, we did not find a negative correlation between MEX3A and CDKN2B expression. BTG2, encoding BTG anti-proliferation factor 2, was another potential target of MEX3A, which was upregulated after MEX3A silencing and downregulated in ccRCC samples. Similar to CDKN2B, BTG2 acts as a cell-cycle regulator. In ccRCC, overexpression of BTG2 suppressed cell proliferation and decreased CCND1 expression, which could inhibit activation of the RB/E2F pathway.37 In this study, we found that BTG2 expression was upregulated after knocking down MEX3A in ccRCC cells, and higher expression of BTG2 was related to a longer survival time in patients. However, we did not further verify the regulatory relationship between MEX3A and BTG2. Therefore, further research is needed to reveal the detailed regulatory network of MEX3A, which could provide a molecular theoretical basis for clinical diagnosis and treatment.
In a previous study, MEX3A was found to be mainly located in the cytoplasm and colocalized with Decapping MRNA 1A (DCP1A) in P bodies in MCF7 cells.9 MEX3A interacts with argonaute (AGO) 1 and AGO2, which could be interrupted by RNase treatment.9 These results indicated that MEX3A plays a role in mRNA degradation, which has been confirmed by several studies.12,15,38 However, a recent study reported that depletion of MEX3A decreased the expression of its direct target, CDK6, by promoting mRNA decay in pancreatic ductal adenocarcinoma cells.13 In addition, Santovito et al. found that MEX3A was enriched in the nucleus and coimmunoprecipitation (coIP) assessments did not detect the interaction between MEX3A and DCP1A in human umbilical vein endothelial cells.39
Despite identifying a list of MEX3A downstream targets in ACHN cells, it is still hard to confirm the RNA-binding function of MEX3A. On the one hand, the binding status of MEX3A to different genes is diverse. For example, the binding region of MEX3A in the 3′ UTR of CDK6 was not completely covered (Figure S4B). In addition, miRNAs could also bind to the 3′ UTR of mRNAs to regulate their expression. Therefore, if the binding region of MEX3A coincides with those of miRNAs, gene expression would be highly likely to be downregulated after silencing MEX3A. Similarly, if the binding region of MEX3A did not overlap with an miRNA-binding region, MEX3A might cooperate with miRNAs to regulate mRNA expression. Furthermore, MEX3A has been proved to form a complex with miR-126-5p and AGO2 without a target mRNA via its two KH domains, which might provide a novel insight into post-transcriptional regulation of MEX3A.39 In short, the regulatory functions of MEX3A and miRNAs on mRNAs might be mutually inhibitory when they form a complex. Upon loss of MEX3A or miRNAs, another one will regulate the expression of their targets by binding to the 3′ UTR of downstream targets. On the other hand, the RING domain of MEX3A might also play an important role in the regulation of downstream targets. In MEX3C, another member of the MEX3 family, loss of the RING domain did not interrupt the interaction between MEX3C and HLA-A2 mRNA.40,41 However, loss of the RING domain would not shorten the HLA-A2 mRNA poly(A) tail.41 These data also suggested that the function of MEX3A in degrading mRNA might be dependent on its RING domain. Here, we performed a genome-wide RNA screen to identify the downstream targets of MEX3A and described the molecular mechanism of RNA regulation of MEX3A to some extent. However, the function of post-transcriptional regulation of MEX3A has not been fully explained and needs further investigation.
Regarding the MEX3-recognition element (MRE), Pagano et al. identified a high-affinity MRE ((A/G/U) (G/U)AGN0-8U(U/A/C)UA)) by mixing MEX3 and several pools of ssRNA.42 Then, a study demonstrated that the KH1 domain of MEX3C could only bind to MRE10a (5′-CAGAGU-3′), whereas the KH2 domain could bind to both MRE10a and MRE10b (5′-GUUUAG-3′).43 Similarly, MEX3A was reported to bind to the 3′ UTR of CDX2 through recognizing the sequence 5′-AGAGUUUUUA-3′.15 In the present study, we identified a MEX3A recognition element (5′-UUUAUAAA-3′) using eCLIP-seq, which will be useful to screen targets of MEX3A in other research models.
Lastly, Barriga et al. found that MEX3A high-expression cells were resistant to chemotherapy (5-fluorouracil) and gamma radiation in intestinal stem cells.27 Knocking down MEX3A enhanced the sensitivity to gemcitabine treatment in PDAC.13 However, the mechanisms underlying these findings are largely unknown. In conclusion, further research should focus on the RNA-binding function and the clinical value of MEX3A in ccRCC.
Materials and methods
Cell culture and ccRCC tissue samples
ACHN, A498, and 786-O are the most commonly used ccRCC cell lines.44 ACHN, 786-O, and 293T were purchased from the Cell Bank of the Chinese Academy of Science, and A498 cells were obtained from the American Type Culture Collection (ATCC; Manassas, VA). All cell lines were authenticated by short tandem repeat45 profiles provided by IGE Biotechnology (Guangzhou, China). Cells were cultured in Dulbecco's modified Eagle's medium (DMEM) (Gibco, Grand Island, NY) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (Beyotime, Shanghai, China) at 37°C in 5% CO2. 10 pairs of ccRCC and adjacent tissues were obtained by surgical operation from Chinese Academy of Medical Sciences and Peking Union Medical College. This study was approved by the Review Board of Chinese Academy of Medical Sciences and Peking Union Medical College. All patients provided signed informed consent.
Cell transfections
Transfection of cells with plasmid DNA or siRNA was performed using ViaFect (Promega, Madison, WI) or Lipofectamine RNAiMAX (Thermo Fisher Scientific, Waltham, MA) according to the manufacturer's instructions. siRNAs targeting MEX3A and CDKN2B (siMEX3A-1, GCGAGGAACCAGTGTTCAT; siMEX3A-2, GGCAAGGCTGCAAGATTAA; siCDKN2B, CCAACGGAGTCAACCGTTT), and NC siRNA were purchased from GenePharma (Suzhou, China).
Antibodies and reagents
Western blotting was performed as previously described.24 All the blots were incubated with primary and secondary antibodies (Transgene biotech, 1:10,000) for 1 h. After incubating with the ECL reagent (Thermo Fisher), the blots were visualized using iBright CL1000 (Thermo Fisher). The primary antibodies recognized cyclin-dependent kinase (CDK) 1 (BD Bioscience, San Jose, CA; #610037), CDK4 (CST, Danvers, MA; #12790), CDK2 (CST, #18048), CDK6 (CST, #13331), cyclin-dependent kinase inhibitor 2B (CDKN2B) (Immunoway, Plano, TX; #YT3492), cyclin D1 (CCND1) (BD, #556470), retinoblastoma 1 (RB) (CST, #9309), phosphorylated (p)-RB S795 (CST, #9301), p-RB S807/811 (CST, #8516), E2F transcription factor 1 (E2F1) (Abcam, Cambridge, MA; #ab112580), Mex-3 RNA-binding family member A (MEX3A) (Sigma, St Louis, MO; #HPA062703), FLAG (Sigma, #F1804), and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (Transgene Biotech, Beijing, China; #HC301-01).
Cell-cycle analysis
After siRNA transfection for 48 h, cells were harvested using trypsin, and fixed overnight in 70% ethanol at 4°C. DNA was stained with propidium iodide (PI; 5 μg/μL; Beyotime, Shanghai, China) at 4°C for 15 min in 1× phosphate-buffered saline supplemented with RNase A. PI-labeled samples were measured using a CytoFLEX flow cytometer (Beckman Coulter, Indianapolis, IN). The ModFit LT 5.0 software (Verity Software House, Topsham, ME) was used to analyze the cell-cycle distribution.
Cell colony formation, migration, and MTT assays
After siRNA transfection for 48 h, cells were harvested using trypsin. For the colony-formation assay, 500 cells per well were seeded into six-well plates and incubated for 2 weeks, then fixed using 4% paraformaldehyde for 15 min and stained with 0.1% crystal violet for 15 min. For the MTT assay, 2 × 103 cells per well were seeded into 96-well plates, and cell proliferation was measured using a standard MTT assay, as described previously.24 For the migration assay, 550 μL of DMEM medium containing 20% FBS was added to the lower chamber of a Transwell insert. Cells suspended in 100 μL of DMEM medium without FBS (ACHN, 1 × 105 cells per well; A498, 3 × 104 cells per well; 786-O, 3 × 104 cells per well) were seeded into the upper chambers of 24-well Transwell inserts and incubated for 24 h. Cell in the upper chamber were fixed with 4% paraformaldehyde for 15 min and stained with 0.1% crystal violet for 15 min. After removing cells in the upper chamber, cells that passed through the membrane were counted under a light microscope (Nikon, Ni-U, Tokyo, Japan).
Luciferase reporter assay
To detect the binding of ETS1, the sequences of NC and WT were separately cloned into pGL3-enhancer vector. The mutant pGL3-enhancer was cloned using MutanBEST kit (Takara, Dalian, China, #R401). After co-transfection with pRK-TK for 48 h, luciferase activity was measured using a dual-luciferase reporter assay system (Promega). To detect the binding of MEX3A to the CDKN2B 3’ UTR, the sequences were cloned into the psiCHECK-2 vector purchased from IGE Biotechnology. After co-transfection with constructed psicheck-2-3′ UTR-CDKN2B and PCDH-CopGFP-puro-MEX3A in cells for 48 h, luciferase activity was measured using a dual-luciferase reporter assay system (Promega). Primers used for PCR amplification during luciferase reporter vector construction are listed in Table S1.
Lentivirus packaging and infection
Lentiviral plasmids PCDH-CopGFP-puro-MEX3A and pLKO.1-EGFP-puro-MEX3A, and control plasmids were purchased from IGE Biotechnology. These pLKO-based and PCDH-based lentiviral plasmids were co-transfected with packing plasmids (psPAX2 and pMD2.G) into 293T cells. Culture medium containing lentivirus was collected and filtered. Stable knockdown or overexpression cells were selected using 1–2 ng/μL puromycin for 3 days.
RNA-seq and RIP-seq assay
After transfection for 48 h, total RNA was extracted using RNAiso Plus (Takara, #9108Q) according to the manufacturer's instructions. mRNA library construction and single-end sequencing were executed by BGI (Wuhan, China). The primers used to validate altered genes using qRT-PCR are listed in Table S2.
The RIP-seq assay was performed as previously described.46 Briefly, 1 × 107 cells stably expressing FLAG-tagged MEX3A were treated with 0.3% formaldehyde and the cross-linking reaction was stopped by adding 0.125 M glycine. The complexes of RNA and protein were enriched using 5 μg of anti-FLAG antibody with Dynabeads protein G (Invitrogen, Waltham, MA; #10004D) in cell lysis buffer overnight at 4°C. Then, total RNA was isolated by the phenol-chloroform method after treatment with proteinase K (Thermo Fisher). Library construction and sequencing were completed by BGI.
RNA-seq data and RIP-seq data were aligned and quantified using RSEM47 and STAR48 against the GRCh37 human reference genome and annotation (GTF format) downloaded from GENCODE official website. Gene counts were then used for downstream analysis.
eCLIP assay and data analysis
FLAG-tagged MEX3A eCLIP was performed in ACHN cells, following a previously published protocol.49 Briefly, input and FLAG antibody IP fractions were run on an NuPAGE Bis-Tris protein gel (Thermo Fisher). Protein-RNA complexes between 72 kDa and 150 kDa were collected for RNA isolation, followed by library generation. eCLIP libraries were sent to IGE Biotechnology for paired-end sequencing (PE150). eCLIP data processing was conducted using eCLIP pipeline version 0.2.1a (https://github.com/YeoLab/eclip/releases/tag/0.2.1a). High-confidence eCLIP peaks were called by selecting MEX3A-binding peaks with a minimum IP count ≥10 and a minimum log2 fold-change (FC) IP/input value ≥1. De novo motif analysis was conducted by using HOMER v4.10 findMotifsGenome.pl script with the following parameters: -rna -S 10 -size 200 -len 8 -p 4. The ChIPSeeker R package was used to determine the distribution of peaks.50
RNA pull-down assay
The 3′ biotin-labeled RNA probes were synthesized by IGE Biotechnology. According to a previous study, ACHN cells in a 100-mm dish were lysed by adding 1 mL of lysis buffer (10 mM HEPES pH 7.0, 200 mM NaCl, 1% Triton X-100, 10 mM MgCl2, 1 mM DTT with protease inhibitors and RNase inhibitor).51 After incubation for 30 min on ice, supernatants were transferred to new tubes using high-speed centrifugation (12,000 × g, 15 min, 4°C). RNA probes were applied to 30 μL of washed streptavidin beads and incubated on a rotating shaker at 4°C for at least 20 min. Then, 200 μg of protein lysate was incubated with 30 μL of RNA-Beads (New England Biolabs, Ipswich, MA; S1420S) at 4°C for 2 h. Beads were washed three times with lysis buffer. Captured proteins were heated for denaturation and validated using western blotting. The sequence of RNA probes is listed in Table S3.
Sphere-formation assay
To evaluate sphere-formation potential, ACHN cells were treated with siRNAs for 24 h. Then, cells were plated at a density of 500 cells per well in the ultra-low attachment 12 plate (Corning Incorporated, Corning, NY) and incubated in DMEM/F12 (1:1) supplemented with 1× B-27 (Gibco, United States), 20 ng/mL epidermal growth factor (EGF; Gibco, United States), 10 ng/mL fibroblast growth factor-basic (bFGF; Gibco, United States), 1× L-glutamine, and 1× penicillin/streptomycin. Every 3 days, half of the medium was replaced. Cells were cultured for 10 days, and the spheroid of each well was photographed with an Olympus fluorescent microscope and counted.
Murine xenograft model
The studies were approved by the Animal Management Committee of Sun Yat-Sen University. Cells (3 × 106) were injected subcutaneously in nude mice (3–4 weeks old). The tumors were harvested and their sizes and weights were measured.
Statistics
All results were analyzed statistically using GraphPad Prism v8 (GraphPad, La Jolla, CA). A p value <0.05 was considered as statistically significant. OS was computed using the Kaplan-Meier method, and p values of OS were calculated using the log rank test.
Data availability
ChIP-seq data were downloaded from GEO (H3K4me3 ChIP-seq and H3K27ac ChIP-seq data of ccRCC tissues, H3K4me3 and H3K27ac ChIP-seq data of A498 and H3K4me3 and ETS1 ChIP-seq data of 786-O were download from GSE86095; H3K27ac ChIP-seq data of 786-O were download from GSE78113). ATAC-seq data were download from TCGA. RNA-seq of silencing MEX3A, and the RIP-seq and eCLIP-seq data, have been deposited at GEO with the accession number GSE175409. Any other data are available upon request.
Acknowledgments
This research was supported in part through computational resources provided by the Bioinformatics and Omics Center, Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University. This work was supported by grants from the National Natural Science Foundation of China (82073067 and 81872140 to D.Y., 81872306 to LH.L.); Guangdong Science and Technology Department (2019B020226003 and 2021A0505030084 to D.Y.; 2020B1212060018 and 2020B1212030004]; Chinese Academy of Medical Sciences (CAMS) Initiative for Innovative Medicine (2016-I2M-2–005).
Author contributions
Research designation, D.Y., X.B., and L.L.; experimental performance and data analyses, Y.Q., M.M., C.C., J.Z., X.C., Y.H., H.C., Y.L., D.T., Y.H., L.P., K.H., Y.Z., J.L., J.H., and X.W.; figure preparation, Y.Q.; manuscript writing, Y.Q., L.L., and D.L.
Declaration of interests
The authors declare no competing interests.
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.omtn.2021.11.026.
Contributor Information
Lehang Lin, Email: linlh7@mail2.sysu.edu.cn.
Xingang Bi, Email: bixingang@csco.org.cn.
Dong Yin, Email: yind3@mail.sysu.edu.cn.
Supplemental information
References
- 1.Siegel R.L., Miller K.D., Fuchs H.E., Jemal A. Cancer statistics, 2021. CA Cancer J. Clin. 2021;71:7–33. doi: 10.3322/caac.21654. [DOI] [PubMed] [Google Scholar]
- 2.Ricketts C.J., De Cubas A.A., Fan H., Smith C.C., Lang M., Reznik E., Bowlby R., Gibb E.A., Akbani R., Beroukhim R., et al. The Cancer Genome Atlas comprehensive molecular characterization of renal cell carcinoma. Cell Rep. 2018;23:313–326 e315. doi: 10.1016/j.celrep.2018.03.075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Linehan W.M., Ricketts C.J. The Cancer Genome Atlas of renal cell carcinoma: findings and clinical implications. Nat. Rev. Urol. 2019;16:539–552. doi: 10.1038/s41585-019-0211-5. [DOI] [PubMed] [Google Scholar]
- 4.Sato Y., Yoshizato T., Shiraishi Y., Maekawa S., Okuno Y., Kamura T., Shimamura T., Sato-Otsubo A., Nagae G., Suzuki H., et al. Integrated molecular analysis of clear-cell renal cell carcinoma. Nat. Genet. 2013;45:860–867. doi: 10.1038/ng.2699. [DOI] [PubMed] [Google Scholar]
- 5.Cancer Genome Atlas Research Network Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature. 2013;499:43–49. doi: 10.1038/nature12222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Jonasch E., Gao J., Rathmell W.K. Renal cell carcinoma. BMJ. 2014;349:g4797. doi: 10.1136/bmj.g4797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Jonasch E., Walker C.L., Rathmell W.K. Clear cell renal cell carcinoma ontogeny and mechanisms of lethality. Nat. Rev. Nephrol. 2021;17:245–261. doi: 10.1038/s41581-020-00359-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lederer M., Muller S., Glass M., Bley N., Ihling C., Sinz A., Huttelmaier S. Oncogenic potential of the dual-function protein MEX3A. Biology (Basel) 2021;10:415. doi: 10.3390/biology10050415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Buchet-Poyau K., Courchet J., Le Hir H., Seraphin B., Scoazec J.Y., Duret L., Domon-Dell C., Freund J.N., Billaud M. Identification and characterization of human Mex-3 proteins, a novel family of evolutionarily conserved RNA-binding proteins differentially localized to processing bodies. Nucleic Acids Res. 2007;35:1289–1300. doi: 10.1093/nar/gkm016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Yang C., Zhan H., Zhao Y., Wu Y., Li L., Wang H. MEX3A contributes to development and progression of glioma through regulating cell proliferation and cell migration and targeting CCL2. Cell Death Dis. 2021;12:14. doi: 10.1038/s41419-020-03307-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wei L., Wang B., Hu L., Xu Y., Li Z., Shen Y., Huang H. MEX3A is upregulated in esophageal squamous cell carcinoma (ESCC) and promotes development and progression of ESCC through targeting CDK6. Aging (Albany NY) 2020;12:21091–21113. doi: 10.18632/aging.103196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Liang J., Li H., Han J., Jiang J., Wang J., Li Y., Feng Z., Zhao R., Sun Z., Lv B., et al. Mex3a interacts with LAMA2 to promote lung adenocarcinoma metastasis via PI3K/AKT pathway. Cell Death Dis. 2020;11:614. doi: 10.1038/s41419-020-02858-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Panzeri V., Manni I., Capone A., Naro C., Sacconi A., Di Agostino S., de Latouliere L., Montori A., Pilozzi E., Piaggio G., et al. The RNA-binding protein MEX3A is a prognostic factor and regulator of resistance to gemcitabine in pancreatic ductal adenocarcinoma. Mol. Oncol. 2021;15:579–595. doi: 10.1002/1878-0261.12847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jiang S., Meng L., Chen X., Liu H., Zhang J., Chen F., Zheng J., Liu H., Wang F., Hu J., et al. MEX3A promotes triple negative breast cancer proliferation and migration via the PI3K/AKT signaling pathway. Exp. Cell Res. 2020;395:112191. doi: 10.1016/j.yexcr.2020.112191. [DOI] [PubMed] [Google Scholar]
- 15.Pereira B., Sousa S., Barros R., Carreto L., Oliveira P., Oliveira C., Chartier N.T., Plateroti M., Rouault J.P., Freund J.N., et al. CDX2 regulation by the RNA-binding protein MEX3A: impact on intestinal differentiation and stemness. Nucleic Acids Res. 2013;41:3986–3999. doi: 10.1093/nar/gkt087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jasinski-Bergner S., Steven A., Seliger B. The role of the RNA-binding protein family MEX-3 in tumorigenesis. Int. J. Mol. Sci. 2020;21:5209. doi: 10.3390/ijms21155209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cerami E., Gao J., Dogrusoz U., Gross B.E., Sumer S.O., Aksoy B.A., Jacobsen A., Byrne C.J., Heuer M.L., Larsson E., et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–404. doi: 10.1158/2159-8290.CD-12-0095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yao X., Tan J., Lim K.J., Koh J., Ooi W.F., Li Z., Huang D., Xing M., Chan Y.S., Qu J.Z., et al. VHL deficiency drives enhancer activation of oncogenes in clear cell renal cell carcinoma. Cancer Discov. 2017;7:1284–1305. doi: 10.1158/2159-8290.CD-17-0375. [DOI] [PubMed] [Google Scholar]
- 19.Mikami S., Oya M., Mizuno R., Murai M., Mukai M., Okada Y. Expression of Ets-1 in human clear cell renal cell carcinomas: implications for angiogenesis. Cancer Sci. 2006;97:875–882. doi: 10.1111/j.1349-7006.2006.00268.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang R., Ma Y., Yu D., Zhao J., Ma P. miR-377 functions as a tumor suppressor in human clear cell renal cell carcinoma by targeting ETS1. Biomed. Pharmacother. 2015;70:64–71. doi: 10.1016/j.biopha.2015.01.012. [DOI] [PubMed] [Google Scholar]
- 21.Luo Y., Liu F., Yan C., Qu W., Zhu L., Guo Z., Zhou F., Zhang W. Long non-coding RNA CASC19 sponges microRNA-532 and promotes oncogenicity of clear cell renal cell carcinoma by increasing ETS1 expression. Cancer Manag. Res. 2020;12:2195–2207. doi: 10.2147/CMAR.S242472. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 22.Scoles D.R., Pflieger L.T., Thai K.K., Hansen S.T., Dansithong W., Pulst S.M. ETS1 regulates the expression of ATXN2. Hum. Mol. Genet. 2012;21:5048–5065. doi: 10.1093/hmg/dds349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nevins J.R. The Rb/E2F pathway and cancer. Hum. Mol. Genet. 2001;10:699–703. doi: 10.1093/hmg/10.7.699. [DOI] [PubMed] [Google Scholar]
- 24.Li Y., Xiao X., Chen H., Chen Z., Hu K., Yin D. Transcription factor NFYA promotes G1/S cell cycle transition and cell proliferation by transactivating cyclin D1 and CDK4 in clear cell renal cell carcinoma. Am. J. Cancer Res. 2020;10:2446–2463. [PMC free article] [PubMed] [Google Scholar]
- 25.Kim W.Y., Sharpless N.E. The regulation of INK4/ARF in cancer and aging. Cell. 2006;127:265–275. doi: 10.1016/j.cell.2006.10.003. [DOI] [PubMed] [Google Scholar]
- 26.Pereira B., Amaral A.L., Dias A., Mendes N., Muncan V., Silva A.R., Thibert C., Radu A.G., David L., Maximo V., et al. MEX3A regulates Lgr5(+) stem cell maintenance in the developing intestinal epithelium. EMBO Rep. 2020;21:e48938. doi: 10.15252/embr.201948938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Barriga F.M., Montagni E., Mana M., Mendez-Lago M., Hernando-Momblona X., Sevillano M., Guillaumet-Adkins A., Rodriguez-Esteban G., Buczacki S.J.A., Gut M., et al. Mex3a marks a slowly dividing subpopulation of Lgr5+ intestinal stem cells. Cell Stem Cell. 2017;20:801–816 e807. doi: 10.1016/j.stem.2017.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bufalieri F., Caimano M., Lospinoso Severini L., Basili I., Paglia F., Sampirisi L., Loricchio E., Petroni M., Canettieri G., Santoro A., et al. The RNA-binding ubiquitin ligase MEX3A affects glioblastoma tumorigenesis by inducing ubiquitylation and degradation of RIG-I. Cancers (Basel) 2020;12:321. doi: 10.3390/cancers12020321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Jiang H., Zhang X., Luo J., Dong C., Xue J., Wei W., Chen J., Zhou J., Gao Y., Yang C. Knockdown of hMex-3A by small RNA interference suppresses cell proliferation and migration in human gastric cancer cells. Mol. Med. Rep. 2012;6:575–580. doi: 10.3892/mmr.2012.943. [DOI] [PubMed] [Google Scholar]
- 30.Xia Y., Liu Y., Yang C., Simeone D.M., Sun T.T., DeGraff D.J., Tang M.S., Zhang Y., Wu X.R. Dominant role of CDKN2B/p15INK4B of 9p21.3 tumor suppressor hub in inhibition of cell-cycle and glycolysis. Nat. Commun. 2021;12:2047. doi: 10.1038/s41467-021-22327-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wang W., Zhang R., Wang X., Wang N., Zhao J., Wei Z., Xiang F., Wang C. Suppression of KIF3A inhibits triple negative breast cancer growth and metastasis by repressing Rb-E2F signaling and epithelial-mesenchymal transition. Cancer Sci. 2020;111:1422–1434. doi: 10.1111/cas.14324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kent L.N., Leone G. The broken cycle: E2F dysfunction in cancer. Nat. Rev. Cancer. 2019;19:326–338. doi: 10.1038/s41568-019-0143-7. [DOI] [PubMed] [Google Scholar]
- 33.Polit J.T., Kazmierczak A., Walczak-Drzewiecka A. Cell cycle-dependent phosphorylation of pRb-like protein in root meristem cells of Vicia faba. Protoplasma. 2012;249:131–137. doi: 10.1007/s00709-011-0272-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Heilmann A.M., Dyson N.J. Phosphorylation puts the pRb tumor suppressor into shape. Genes Dev. 2012;26:1128–1130. doi: 10.1101/gad.195552.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mok M.T., Zhou J., Tang W., Zeng X., Oliver A.W., Ward S.E., Cheng A.S. CCRK is a novel signalling hub exploitable in cancer immunotherapy. Pharmacol. Ther. 2018;186:138–151. doi: 10.1016/j.pharmthera.2018.01.008. [DOI] [PubMed] [Google Scholar]
- 36.Baietti M.F., Zhao P., Crowther J., Sewduth R.N., De Troyer L., Debiec-Rychter M., Sablina A.A. Loss of 9p21 regulatory hub promotes kidney cancer progression by upregulating HOXB13. Mol. Cancer Res. 2021;19:979–990. doi: 10.1158/1541-7786.MCR-20-0705. [DOI] [PubMed] [Google Scholar]
- 37.Sima J., Zhang B., Sima X.Y., Mao Y.X. Overexpression of BTG2 suppresses growth, migration, and invasion of human renal carcinoma cells in vitro. Neoplasma. 2016;63:385–393. doi: 10.4149/307_150822N455. [DOI] [PubMed] [Google Scholar]
- 38.Draper B.W., Mello C.C., Bowerman B., Hardin J., Priess J.R. MEX-3 is a KH domain protein that regulates blastomere identity in early C. elegans embryos. Cell. 1996;87:205–216. doi: 10.1016/s0092-8674(00)81339-2. [DOI] [PubMed] [Google Scholar]
- 39.Santovito D., Egea V., Bidzhekov K., Natarelli L., Mourao A., Blanchet X., Wichapong K., Aslani M., Brunssen C., Horckmans M., et al. Noncanonical inhibition of caspase-3 by a nuclear microRNA confers endothelial protection by autophagy in atherosclerosis. Sci. Transl. Med. 2020;12:eaaz2294. doi: 10.1126/scitranslmed.aaz2294. [DOI] [PubMed] [Google Scholar]
- 40.Cano F., Bye H., Duncan L.M., Buchet-Poyau K., Billaud M., Wills M.R., Lehner P.J. The RNA-binding E3 ubiquitin ligase MEX-3C links ubiquitination with MHC-I mRNA degradation. EMBO J. 2012;31:3596–3606. doi: 10.1038/emboj.2012.218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cano F., Rapiteanu R., Sebastiaan Winkler G., Lehner P.J. A non-proteolytic role for ubiquitin in deadenylation of MHC-I mRNA by the RNA-binding E3-ligase MEX-3C. Nat. Commun. 2015;6:8670. doi: 10.1038/ncomms9670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Pagano J.M., Farley B.M., Essien K.I., Ryder S.P. RNA recognition by the embryonic cell fate determinant and germline totipotency factor MEX-3. Proc. Natl. Acad. Sci. U S A. 2009;106:20252–20257. doi: 10.1073/pnas.0907916106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Yang L., Wang C., Li F., Zhang J., Nayab A., Wu J., Shi Y., Gong Q. The human RNA-binding protein and E3 ligase MEX-3C binds the MEX-3-recognition element (MRE) motif with high affinity. J. Biol. Chem. 2017;292:16221–16234. doi: 10.1074/jbc.M117.797746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wolf M.M., Kimryn Rathmell W., Beckermann K.E. Modeling clear cell renal cell carcinoma and therapeutic implications. Oncogene. 2020;39:3413–3426. doi: 10.1038/s41388-020-1234-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Klughammer J., Kiesel B., Roetzer T., Fortelny N., Nemc A., Nenning K.H., Furtner J., Sheffield N.C., Datlinger P., Peter N., et al. The DNA methylation landscape of glioblastoma disease progression shows extensive heterogeneity in time and space. Nat. Med. 2018;24:1611–1624. doi: 10.1038/s41591-018-0156-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Tsai M.C., Manor O., Wan Y., Mosammaparast N., Wang J.K., Lan F., Shi Y., Segal E., Chang H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329:689–693. doi: 10.1126/science.1192002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Li B., Dewey C.N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics. 2011;12:323. doi: 10.1186/1471-2105-12-323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Dobin A., Davis C.A., Schlesinger F., Drenkow J., Zaleski C., Jha S., Batut P., Chaisson M., Gingeras T.R. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Van Nostrand E.L., Pratt G.A., Shishkin A.A., Gelboin-Burkhart C., Fang M.Y., Sundararaman B., Blue S.M., Nguyen T.B., Surka C., Elkins K., et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP) Nat. Methods. 2016;13:508–514. doi: 10.1038/nmeth.3810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Yu G., Wang L.G., He Q.Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics. 2015;31:2382–2383. doi: 10.1093/bioinformatics/btv145. [DOI] [PubMed] [Google Scholar]
- 51.Zhang Y., Huang Y.X., Wang D.L., Yang B., Yan H.Y., Lin L.H., Li Y., Chen J., Xie L.M., Huang Y.S., et al. LncRNA DSCAM-AS1 interacts with YBX1 to promote cancer progression by forming a positive feedback loop that activates FOXA1 transcription network. Theranostics. 2020;10:10823–10837. doi: 10.7150/thno.47830. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
ChIP-seq data were downloaded from GEO (H3K4me3 ChIP-seq and H3K27ac ChIP-seq data of ccRCC tissues, H3K4me3 and H3K27ac ChIP-seq data of A498 and H3K4me3 and ETS1 ChIP-seq data of 786-O were download from GSE86095; H3K27ac ChIP-seq data of 786-O were download from GSE78113). ATAC-seq data were download from TCGA. RNA-seq of silencing MEX3A, and the RIP-seq and eCLIP-seq data, have been deposited at GEO with the accession number GSE175409. Any other data are available upon request.