Abstract
We investigated six cases of enterovirus infection in a neonatal unit. The index patient, a 5-day-old boy, was admitted with aseptic meningitis due to echovirus 30 (E30). Secondary infections with E30 occurred in five babies. Comparison of the complete VP1 sequences showed that the isolates recovered from the index patient and his mother were closely related to those recovered from the five babies with secondary infections, demonstrating a nosocomial transmission of the virus. In the phylogenetic tree reconstructed from the VP1 sequences, the isolates formed a monophyletic cluster related to an E30 strain collected in June 1997 during an outbreak of aseptic meningitis.
Enterovirus infections through nosocomial transmission are well documented and involve different serotypes of group A and group B coxsackieviruses and echoviruses (10). Increased vigilance is particularly necessary in neonatal nurseries and neonatal and intensive care units because neonates represent a population at risk for severe enterovirus diseases (1, 6–8, 16).
Echovirus 30 (E30) is one of the most prevalent enteroviruses (2, 11, 15). It caused several outbreaks of aseptic meningitis (3, 12, 21), but was not often involved in infections through nosocomial transmission (14). In a recent study, we reported clinical and epidemiological data strongly suggesting the nosocomial transmission, in 1997, of an E30 strain in one area of a neonatal unit (9). In the present study, we report molecular evidence of the nosocomial transmission of the virus from the index patient to five other babies. The evidence is based on the phylogenetic analysis of the complete nucleotide sequence encoding the VP1 capsid polypeptide.
MATERIALS AND METHODS
Clinical specimen and virus identification.
Detection of the enterovirus genome in cerebrospinal fluid (CSF) with the AMPLICOR EV test kit (Roche Molecular Systems) was routinely performed according to the manufacturer's recommendations. During an outbreak of aseptic meningitis in a neonatal unit, E30 isolates were collected from stool specimens or CSF from six neonates (Table 1). A virus isolate was also obtained from a stool specimen from the index patient's mother. Virus isolation was performed as described elsewhere (9). Propagation of the isolates was limited to a maximum of two passages in MRC5 cell cultures (human lung embryonic fibroblasts; Bio-Mérieux, Marcy l'Etoile, France). Virus identification by neutralization tests with the Lim-Benyesh-Melnick antiserum pools was performed as described elsewhere (9, 13).
TABLE 1.
Patients, clinical observations, and virus isolates
| Patient (age/sex)a | Admission date (1997)b | Clinical feature(s) on admission | Date of:
|
Virus isolate (designation/LN/AN)d | |
|---|---|---|---|---|---|
| Diagnosis by PCRc | Virus isolation from stools | ||||
| Index patient (5 days/M) | 2 October | Aseptic meningitis | 2 October | 8 October | IP/97CF528/AJ241449 |
| Patient 1 (1 day/F) | 18 September | Prematurity, hypotrophy | 24 October | 24 October | P1/97CF555/AJ241452 |
| Patient 2 (1 day/F) | 14 September | Prematurity, hypotrophy | NDe | 30 October | P2/97CF566/AJ241454 |
| 30 Octoberf | |||||
| Patient 3 (1 day/F) | 14 September | Prematurity, hypotrophy | ND | 30 October | P3/97CF572/AJ241455 |
| 30 Octoberf | |||||
| Patient 4 (2 days/M) | 8 October | Proteus mirabilis infection | 21 October | 24 and 30 October | P4/97CF551/AJ241451 |
| Patient 5 (4 days/M) | 4 October | Escherichia coli infection | ND | 24 October | P5/97CF556/AJ241453 |
Patients 1, 2, and 3 were triplet babies, born by cesarean section at 34 weeks of gestation. Patient 1 was first admitted to an intensive care unit. M, male; F, female.
Admission to the neonatal unit.
Positive detection of enterovirus genome in CSF.
LN, laboratory number; AN, accession number.
ND, not determined.
Patients were discharged on 24 October and readmitted because of a 3-day recurrence of fever and seizure.
The E30 prototype strain, E30/1958/USA/Bastianni (5), and 10 E30 isolates (78CF1260, 78CF1074, 91CF670, 92CF495, 94CF1845, 97CF1261, 97CF1308, 97CF1377, 97CF1619, and 98CF746) collected between 1978 and 1998 from patients with aseptic meningitis were included in the study as controls for the phylogenetic analysis.
Amplification of the complete VP1 sequence.
Synthesis of cDNAs was performed according to previously described methods (4, 5). Two synthetic oligonucleotide primers, ECOX01 (5′-GC GGA TCC GCG GCC GCG AGC TCI GCR TGC AAY GAY TTY TCW G-3′) and ECOX02 (5′-GCT GCA GGG CGC GCC TCT AGA RTC YCT RTT RTA RTC YTC CCA-3′), were constructed to amplify the entire sequence of VP1 from the genome of echoviruses and coxsackie B viruses (J.-L. Bailly, A. Béguet, M. Chambon, C. Henquell, and H. Peigue-Lafeuille, submitted for publication). Each oligonucleotide has a composite sequence: a group-specific sequence (underlined) at the 3′ end and a heterologous sequence with a high guanosine and cytosine content at the 5′ end. The group-specific sequences were constructed from nucleotide stretches conserved in the genome of echoviruses and coxsackie B viruses for which genome sequences were available from international databases. ECOX01 was designed from a stretch conserved in the VP3 encoding sequence and ECOX02 was designed from the 2A encoding sequence (Bailly et al., submitted). Group-specific sequences contained mixed bases or inosine residues at degenerate sites. The heterologous sequence in the oligonucleotides allowed hybridization during the amplifications to be performed at a stringent temperature. PCR products were synthesized by using the Expand Long Template PCR System (Roche Molecular Biochemicals) with 10× buffer no. 3. The amplification reactions were performed with 2 μl of the cDNA in a mixture containing 200 μM each of the four deoxynucleotides and 1.75 U of the enzyme mix (thermostable Taq and Pwo DNA polymerases). Thermal cycling comprised 40 cycles as follows. The first five cycles consisted of denaturation for 2 min at 94°C, hybridization for 20 s at 51°C, and elongation for 50 s at 68°C (denaturation was lowered to 15 s in cycles 2 to 5). Amplification of the cDNA was then performed in 35 cycles of 15 s at 94°C, 20 s at 64°C, and 35 s at 72°C. Amplifications were carried out with an Omnigene thermocycler (Hybaid).
Nucleotide sequencing of PCR products.
Amplification products were electrophoresed on a 1% NuSieve GTG agarose (FMC Bioproducts, Rockland, Maine) preparative gel and purified from low-melting-point agarose by conventional phenol-chloroform extractions. Nucleotide sequences were determined on both strands of the purified PCR products. Sequencing reactions were carried out at Nucleica SA (Clermont-Ferrand, France) with an ABI PRISM Dye Terminator Cycle Sequencing Ready Reaction kit (PE Applied Biosystems).
Phylogenetic analysis of the VP1 sequences.
The phylogenetic analysis of VP1 sequences was performed with the Tree-Puzzle computer program (22). The phylogenetic tree was reconstructed from the VP1 sequences with Tamura-Nei's model of sequence evolution (24) by the quartet puzzling method (22), which estimates pairwise distances by maximum likelihood. To ensure phylogenetic accuracy, a data set was constructed with the nucleotide sequences of the Bastianni reference strain (5), the 7 isolates recovered during the neonatal outbreak, and the 10 control isolates collected in our laboratory between 1978 and 1998.
Nucleotide sequence accession number.
The nucleotide sequences determined for isolates IP, MO, and P1 to P5 were deposited with the EMBL data library under accession no. AJ241449 to AJ241455. The sequences determined for the E30 control isolates were deposited under accession no. AJ241439, AJ241441, AJ241444, AJ241448, AJ241456, AJ276626, and AJ276812 to AJ276815.
RESULTS
Outbreak description.
In October 1997, the index patient, a 5-day-old boy (Table 1), was admitted to a neonatal unit with aseptic meningitis due to an enterovirus, thought to have been acquired from his infected mother. Detection of the enterovirus genome in CSF was positive on admission (9). A virus isolate was recovered 6 days later from MRC5 cell cultures inoculated with a stool specimen. Virus identification showed the virus isolate to be E30. E30 was also identified from a stool specimen of the index patient's mother obtained after admission of her child.
Secondary infections occurred subsequently in five babies (Table 1). Triplet babies (patients 1, 2, and 3) had been hospitalized for about 15 days in the neonatal unit when the index patient was admitted on 2 October. Patients 4 and 5 were admitted 6 and 2 days later, respectively. In all five patients, infection with an enterovirus was evidenced either by detection of the genome in the CSF, isolation of a virus from stool specimens, or both. Virus identification showed that the virus isolates from the five patients were E30.
Molecular evidence of nosocomial transmission of the secondary infections.
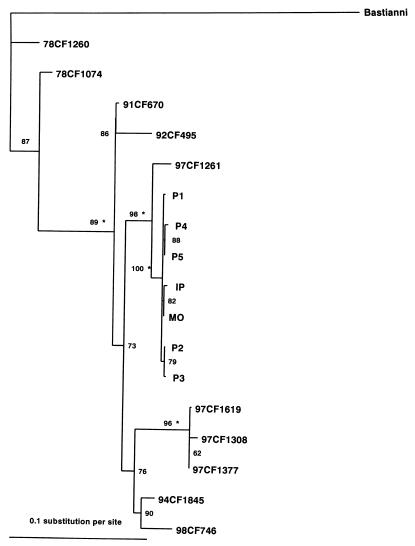
The chronology of the infections, the clinical symptoms, and isolation of the same enterovirus serotype in all patients strongly suggested a nosocomial transmission of an E30 strain in the neonatal unit. To determine whether the patients had been infected independently or whether the infections had a nosocomial origin, virus isolates were characterized by sequencing the complete VP1 encoding sequence after reverse transcription-PCR. Overall, virus isolates IP, MO, and P1 to P5 differed at nine nucleotide sites in the VP1 encoding sequence (Table 2). Virus isolates IP and MO differed at only 2 nucleotide sites (positions 441 and 667) in the 876 nucleotides of the VP1 sequence (99.8% nucleotide similarity). In addition, both isolates had a cytosine at position 714, whereas the five isolates P1 to P5 recovered from patients with suspected nosocomial infection had a uracil at this site. These observations demonstrate the close relatedness of the two isolates and the transmission of the virus from the mother to her child, most probably at the end of gestation or during birth. Isolates P1 to P5 differed at two, three, or four sites from isolate MO and at four, five, and six sites from isolate IP (Table 2). This strongly suggests that all isolates were derived from a common ancestor (isolate IP or MO). To assess the genetic relationships between the different viruses and to determine their origin, the VP1 sequences were compared with homologous sequences determined in four E30 control isolates (97CF1261, 97CF1308, 97CF1377, and 97CF1619) recovered during the outbreak of aseptic meningitis, which began in May and ended at the beginning of August 1997. The seven E30 isolates involved in the neonatal outbreak shared about 98% nucleotide identity with isolate 97CF1261 and 93.1 to 94.1% identity with the other three control isolates. A phylogenetic tree (Fig. 1) reconstructed from maximum likelihood distances showed a close relatedness between the five isolates P1 to P5 and isolates IP and MO, which all clustered in a monophyletic group, thereby demonstrating that the infections in the neonates had a single origin. Moreover, phylogenetic analysis showed conclusively that the readmission of patients 2 and 3 was a direct consequence of an infection acquired in the neonatal unit just before they were discharged. The seven isolates grouped with control isolate 97CF1261 recovered in June 1997 from a CSF specimen from a patient during an outbreak of aseptic meningitis (the internal branch had a very high reliability). Overall, the analysis showed that the index patient's mother was infected with an E30 strain that had been circulating in the general population since the summer outbreak and that she had transmitted the virus to her baby. Finally, the phylogenetic relationships observed in Fig. 1 show the existence of another lineage (isolates 97CF1308, 97CF1377, and 97CF1619) in the E30 strain recovered during the outbreak of aseptic meningitis.
TABLE 2.
Nucleotide differences observed in the VP1 encoding sequences
| Virus isolatea | Nucleotide site at position:
|
No. of nucleotide differences from IP | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 72 | 245 | 398 | 441 | 486 | 667 | 714 | 772 | 859 | ||
| IP | C | A | C | C | U | C | C | C | A | 0 |
| MO | C | A | C | U | U | A | C | C | A | 2 |
| P1 | C | A | C | U | U | A | U | C | U | 4 |
| P2 | U | A | C | U | C | A | U | C | A | 5 |
| P3 | U | A | C | U | C | A | U | U | A | 6 |
| P4 | C | G | A | U | U | A | U | C | A | 5 |
| P5 | C | A | G | U | U | A | U | C | A | 4 |
Virus isolates recovered from the index patient (IP), the index patient's mother (MO), and five patients with secondary infections (P1 to P5).
FIG. 1.
Genetic relationships in E30 VP1 sequences for virus isolates recovered during a nosocomial outbreak. Pairwise maximum likelihood distances were estimated from the complete VP1 sequences (876 nucleotides). To account for nucleotide substitutions, Tamura-Nei's model (24) was used with a gamma distribution of rate heterogeneity across sites (transition/transversion ratio, κ = 8.47; shape parameter for gamma distribution, α = 0.3). To ensure phylogenetic accuracy, parameters α and κ were estimated with the Tree-Puzzle computer program (22) from a data set containing 27 E30 VP1 sequences. Nucleotide frequencies estimated from the data set were as follows: 29.1% for A, 24.3% for C, 23.5% for G, and 23.1% for U. The tree was constructed by the quartet puzzling method. The reliability value (as a percentage) for internal branches indicates how often the corresponding cluster was found among the 10,000 intermediate trees. In 3,060 quartets analyzed, 367 (12.0%) were unresolved. Branch length was drawn to the indicated scale. The sequence of the Bastianni reference strain was used as an outgroup to root the tree. An asterisk indicates a reliability of 100%, estimated by the bootstrap method in the neighbor-joining tree (data not shown). The phylogenetic tree was edited with the Treeview program (19).
DISCUSSION
In a recent study, the nosocomial transmission of an echovirus 7 strain in a neonatal nursery was shown by sequencing a fragment of the 5′ noncoding region (23). This part of the genome is extremely useful for detecting enterovirus genomes in clinical specimens by reverse transcription-PCR (see reference 17 and references therein), but is not suitable for accurate molecular epidemiological studies (5, 17, 18). We based our study on the phylogenetic analysis of the VP1 encoding sequence, one of the most variable sequences of the enterovirus genome (20). The analysis was performed for the complete VP1 sequence, because the mean number of nucleotide differences between virus isolates related to the nosocomial outbreak was very low (minimum of 2, maximum of 6). Hence, we confirmed the usefulness of the method designed for the molecular epidemiology of echoviruses (Bailly et al., submitted). The phylogenetic relationships between E30 isolates related to the nosocomial outbreak were consistent with clinical observations (9). Although it was not possible to identify the exact origin (either isolate IP or isolate MO) of the secondary infections, our results strongly support the hypothesis of the nosocomial transmission of an E30 strain in the neonatal unit. A transmission of the virus from the index patient's mother cannot be excluded, because at the time of the infections, she was still excreting the virus, and all of the nosocomial isolates are more related to isolate MO than to isolate IP. As a consequence of the nosocomial outbreak, the hospital stay of the patients was prolonged by about 6 weeks. All of the patients made a complete recovery; however, the outbreak underlines the necessity of a strict hygiene policy in hospital units for medical staff and visitors.
This study is novel in that it shows an epidemiological connection between three infectious events that took place over a period of about 6 months. The molecular analysis of the VP1 sequences of the virus isolates enabled us to establish a link between a summer outbreak of aseptic meningitis, the vertical transmission of an epidemic virus 3 months later, and the horizontal transmission of the virus in a neonatal unit about 20 days after admission of the index patient.
ACKNOWLEDGMENTS
We are grateful to Danielle Thouvenot of the World Health Organization Collaborating Center, National Reference Center for Enteroviruses (Lyon, France), for providing us with the reference strain of E30. We thank Jeffrey Watts for his revision of the English in the manuscript.
This work was supported in part by a grant from Ministère de l'Education Nationale, de la Recherche et de la Technologie (EA2148).
REFERENCES
- 1.Abzug M J. Perinatal enterovirus infections. In: Rotbart H A, editor. Human enterovirus infections. Washington, D.C.: ASM Press; 1995. pp. 221–238. [Google Scholar]
- 2.Atkinson P J, Sharland M, Maguire H. Predominant enteroviral serotypes causing meningitis. Arch Dis Child. 1998;78:373–374. doi: 10.1136/adc.78.4.373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Aymard M, Chomel J-J, Lina B, Thouvenot D. Annual report 1997. Lyon, France: National Reference Center for Enteroviruses and Hepatitis A; 1997. [Google Scholar]
- 4.Bailly J-L, Borman A M, Peigue-Lafeuille H, Kean K M. Natural isolates of echovirus type 25 with extensive variations in IRES sequences and different translational efficiencies. Virology. 1996;215:83–96. doi: 10.1006/viro.1996.0009. [DOI] [PubMed] [Google Scholar]
- 5.Bailly J-L, Chambon M, Henquell C, Icart J, Peigue-Lafeuille H. Genomic variations in echovirus 30 persistent isolates recovered from a chronically infected immunodeficient child: comparison with the reference strain. J Clin Microbiol. 2000;38:552–557. doi: 10.1128/jcm.38.2.552-557.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bergman I, Painter M J, Wald E R, Chiponis D, Holland A L, Taylor H G. Outcome in children with enteroviral meningitis during the first year of life. J Pediatr. 1987;110:705–709. doi: 10.1016/s0022-3476(87)80006-9. [DOI] [PubMed] [Google Scholar]
- 7.Berlin L E, Rorabaugh M L, Heldrich F, Roberts K, Doran T, Modlin J F. Aseptic meningitis in infants <2 years of age: diagnosis and etiology. J Infect Dis. 1993;168:888–892. doi: 10.1093/infdis/168.4.888. [DOI] [PubMed] [Google Scholar]
- 8.Chambon M, Delage C, Bailly J-L, Gaulme J, Dechelotte P, Henquell C, Jallat C, Peigue-Lafeuille H. Fatal hepatic necrosis in a neonate with echovirus 20 infection: use of the polymerase chain reaction to detect enterovirus in liver tissue. Clin Infect Dis. 1997;24:523–524. doi: 10.1093/clinids/24.3.523. [DOI] [PubMed] [Google Scholar]
- 9.Chambon M, Bailly J-L, Béguet A, Henquell C, Archimbaud C, Gaulme J, Labbé A, Malpuech G, Peigue-Lafeuille H. An outbreak due to echovirus type 30 in a neonatal unit in France in 1997 and usefulness of PCR diagnosis. J Hosp Infect. 1999;43:63–68. doi: 10.1053/jhin.1999.0634. [DOI] [PubMed] [Google Scholar]
- 10.Cherry J D. Enteroviruses: polioviruses (poliomyelitis), coxsackieviruses, echoviruses, and enteroviruses. In: Feigin R D, Cherry J D, editors. Textbook of pediatric infectious diseases. 3rd ed. Philadelphia, Pa: WB Saunders; 1992. pp. 1705–1753. [Google Scholar]
- 11.Druyts-Voets E. Epidemiological features of entero non-poliovirus isolations in Belgium 1980–94. Epidemiol Infect. 1997;119:71–77. doi: 10.1017/s0950268897007656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Leonardi G P, Greenberg A J, Costello P, Szabo K. Echovirus type 30 infection associated with aseptic meningitis in Nassau County, New York, USA. Intervirology. 1993;36:53–56. doi: 10.1159/000150321. [DOI] [PubMed] [Google Scholar]
- 13.Lim K A, Benyesh-Melnick M. Typing of viruses by combination of antiserum pools. Application to typing of enteroviruses (coxsackie and ECHO) J Immunol. 1960;84:309–317. [PubMed] [Google Scholar]
- 14.Matsumoto K, Yokochi T, Matsuda S, Itoi Y, Kimura Y. Characterization of an echovirus type 30 variant isolated from patients with aseptic meningitis. Microbiol Immunol. 1986;30:333–342. doi: 10.1111/j.1348-0421.1986.tb00950.x. [DOI] [PubMed] [Google Scholar]
- 15.Melnick J L. Enteroviruses: polioviruses, coxsackieviruses, echoviruses and newer enteroviruses. In: Fields B N, Knipe D M, editors. Virology. 2nd ed. New York, N.Y: Raven Press; 1990. pp. 549–605. [Google Scholar]
- 16.Modlin J F. Update on enterovirus infections in infants and children. Adv Pediatr Infect Dis. 1997;12:155–180. [PubMed] [Google Scholar]
- 17.Muir P, Kämmerer U, Korn K, Mulders M N, Pöyry T, Weissbrich B, Kandolf R, Cleator G M, van Loon A M for the European Union Concerted Action on Virus Meningitis and Encephalitis. Molecular typing of enteroviruses: current status and future requirements. Clin Microbiol Rev. 1998;11:202–227. doi: 10.1128/cmr.11.1.202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Oberste M S, Maher K, Kilpatrick D R, Flemister M R, Brown B A, Pallansch M A. Typing of human enteroviruses by partial sequencing of VP1. J Clin Microbiol. 1999;37:1288–1293. doi: 10.1128/jcm.37.5.1288-1293.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Page R D M. TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci. 1996;12:357–358. doi: 10.1093/bioinformatics/12.4.357. [DOI] [PubMed] [Google Scholar]
- 20.Palmenberg A C. Sequence alignments of picornaviral capsid proteins. In: Semler B L, Ehrenfeld E, editors. Molecular aspects of picornavirus infection and detection. Washington, D.C.: ASM Press; 1989. pp. 211–241. [Google Scholar]
- 21.Reintjes R, Pohle M, Vieth U, Lyytikainen O, Timm H, Schreier E, Petersen L. Community-wide outbreak of enteroviral illness caused by echovirus 30: a cross-sectional survey and a case-control study. Pediatr Infect Dis J. 1999;18:104–108. doi: 10.1097/00006454-199902000-00005. [DOI] [PubMed] [Google Scholar]
- 22.Strimmer K, von Haeseler A. Quartet puzzling: a quartet maximum likelihood method for reconstructing tree topologies. Mol Biol Evol. 1996;13:964–969. [Google Scholar]
- 23.Takami T, Kawashima H, Takei Y, Miyajima T, Mori T, Nakayama T, Takekuma K, Hoshika A. Usefulness of nested PCR and sequence analysis in a nosocomial outbreak of neonatal enterovirus infection. J Clin Virol. 1998;11:67–75. doi: 10.1016/s0928-0197(98)00022-1. [DOI] [PubMed] [Google Scholar]
- 24.Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 1993;10:512–526. doi: 10.1093/oxfordjournals.molbev.a040023. [DOI] [PubMed] [Google Scholar]