Abstract
Introduction
Dried blood spot (DBS) sampling is a minimally invasive method for specimen collection with potential multifaceted uses, particularly for serosurveillance of previous SARS-CoV-2 infection. In this study, we assessed DBS as a potential specimen type for assessing IgG and total (including IgG and IgM) antibodies to SARS-CoV-2 in vaccinated and naturally infected patients.
Methods
Six candidate buffers were assessed for eluting blood from DBS cards. The study utilized one hundred and five paired plasma specimens and DBS specimens from prospectively collected SARS-CoV-2 vaccinated individuals, remnants from those with PCR confirmed SARS-CoV-2 infections, or remnants from those without history of infection or vaccination. All specimens were tested with the Siemens SARS-CoV-2 total assay (COV2T) or IgG assay (sCOVG).
Results
The lowest backgrounds were observed with water and PBS, and water was used for elution. Relative to plasma samples, DBS samples had a positive percent agreement (PPA) of 94.4% (95% CI: 94.9–100%) for COV2T and 79.2 (68.4–87.0) for sCOVG using the manufacturer’s cutoff. The NPA was 100 % (87.1–100.0 and 85.13–100) for both assays. Dilution studies revealed 100% (95% CI: 90.8–100%) qualitative agreement between specimen types on the COV2T assay and 98.0% (88.0–99.9%) with the sCOVG using study defined cutoffs.
Conclusion
DBS specimens demonstrated high PPA and NPA relative to plasma for SARS-CoV-2 serological testing. Our data support feasibility of DBS sampling for SARS-CoV-2 serological testing.
Keywords: DBS, SARS-CoV-2, IgG, Serology, Total IgG & IgM, COVID-19
1. Introduction
Correctly identifying, triaging, and isolating patients with active SARS-CoV-2 infection is paramount to minimize the risk of transmission to close contacts in the community and at health care centers. While diagnostic nucleic acid-based methods are the gold standard for detecting active COVID-19 infection, SARS-CoV-2 serological testing is an important tool for confirming the presence of current or previous infection in certain situations. According to the Infectious Diseases Society of America, detection of SARS-CoV-2 antibodies is useful for evaluating patients with high clinical suspicion of COVID-19 but persistently negative PCR testing and > 2 weeks of symptoms, for assessing multisystem inflammatory syndrome in children, and for conducting serosurveillance studies [1]. Seroprevalence studies are crucial for understanding the total disease burden of SARS-CoV-2 [2]; and have demonstrated 2–3 fold higher infections per reported COVID-19 case among U.S. blood donors [3]. Of particular importance for serosurveillance studies are the use of assays with high sensitivity and specificity [4] . While many high throughput, automated methods have sufficient analytical performance for this purpose [5], [6], [7], several limitations exist if serological assays are to be used for serosurveillance. Limitations include the requirement for traditional phlebotomy, inability to reach patients who are currently under quarantine, and difficulty in testing disadvantaged patients that lack access to COVID-19 testing centers. This has led some to use point of care based, lateral flow assays in population studies to easily assess SARS-CoV-2 prevalence [8], [9]. However several lateral flow based methods have demonstrated poor clinical performance [10], calling into question the results from surveillance studies performed using assays that have not been thoroughly validated.
Dried blood spot (DBS) sampling has emerged as an invaluable minimally invasive tool for clinical specimen collection. DBS sampling is currently performed clinically for newborn screening [11], [12]. DBS sampling has also been proposed for monitoring changes in erythropoiesis and other biologically relevant molecules in the fight against doping in professional sports [13], [14], [15], drug screening [16], and for the diagnoses of infectious diseases including hepatitis C virus [17], [18], cytomegalovirus [19] and HIV [20]. The varied utility of DBS sampling is further reflected by the stability and retrieval of analytes like microRNA [21], protein [22] and sphingolipids [23] for sensitive molecular assays and mass spectrometry. The clinical utility of DBS sampling lies in its accessibility to at risk populations like the homeless, those with substance use disorders, and patients with limited healthcare access [16]. This also includes resource limited countries where access to COVID-19 testing is often scarce [24], [25]. To obtain a specimen for conventional serology, health care workers perform venipuncture, collecting blood in tubes that require relatively significant storage and/or processing to maintain sample integrity. In contrast, DBS sampling eliminates the need for a phlebotomist since capillary specimens can be easily acquired through a fingerstick and has less stringent storage requirements. Use of dried sample as opposed to a liquid sample also avoids the risk of spills, facilitating handling outside of the typical clinical and laboratory environment. Lastly, amid the recent global shortages in blood tube and collection devices due to manufacturing and shipping delays induced by the coronavirus pandemic, DBS sampling may serve as a proxy for continued SARS-Cov-2 serological testing [26], [27]. Studies utilizing DBS sampling for SARS-CoV-2 serological testing have been reported, but they suffered from small sample size, did not assess seroprevalence, were limited to ELISA based, semi-automated assays for specimen analysis, or required specimen dilution prior to analyses [28], [29], [30].
In this study, we assess DBS as a potential specimen type for assessing IgG and total (including IgG and IgM) antibodies to SARS-CoV-2 in vaccinated and naturally infected patients.
2. Materials and methods
2.1. Specimen collection and handling
This study was approved by the Institutional Review Board of Washington University in St. Louis. We utilized 105 paired plasma (in lithium heparin tubes) and DBS patient specimens (on Whatman 903 Cards, 5 spots per sample) divided into 3 groups. The first group comprised of 48 prospectively recruited post-SARS-CoV-2-vaccine recipients within 126 ± 18.75 days after their second dose of the Pfizer SARS-CoV-2 vaccine. Prospective patients had a paired lithium heparin plasma specimen collected and DBS card spotted from capillary blood. Plasma specimens were processed within 4 h of draw and DBS cards were allowed to dry overnight, sealed in a plastic bag with desiccant, and frozen at −80 °C until analyses. The second group consisted of 34 remnant clinical specimen in lithium heparin syringes from patients diagnosed with COVID-19 via positive SARS-CoV-2 RT-PCR or total SARS-CoV-2 antibody 9 (COV2T) results. Two of the remnant samples came from patients who were vaccinated prior to testing positive for SARS-CoV-2 via RT-PCR. Both specimens were classed in the vaccinated cohort for all subsequent analyses. The third group consisted of 23 remnant clinical patient specimens in lithium heparin syringes from patients with no history of SARS-CoV-2 vaccination or positive SARS-CoV-2 RT-PCR or positive COV2T serology results. To obtain DBS samples from remnant specimens, 60ul was aliquoted onto each DBS Whatman card spot, dried overnight, sealed in plastic bags, and stored at −80 °C until analyses. The residual specimens were subsequently centrifuged to obtain plasma which was stored at −80 °C until analyses.
2.2. Eluent selection
To determine the appropriate eluent for DBS samples, six candidate buffers were assessed: phosphate buffered saline (PBS), PBS + tween 20 at 5% weight / volume (W/V), deionized water, Brij L23 (Sigma Aldrich), Tergitol 15-S-9 (Sigma Aldrich), and ECOSURF EH (Sigma Aldrich). Undiluted (Neat) buffer, 300ul buffer eluates of blank Whatman card punches, and 300ul buffer eluates from 5 punches of Whatman cards spotted with 60 ul of SARS-CoV-2 negative whole blood were analyzed in triplicate to assess overall background by comparing the relative light units (RLU) from each eluent.
2.3. Sample analysis
To elute specimens from DBS cards, five round 6 mm punches per single patient card were acquired using the Arrayit Disc Punch (Arrayit Corporation, Sunnyvale, CA, USA), and placed into a cryovial. Based on deionized water having the lowest RLUs from our eluent selection protocol, 300ul of deionized water was utilized as eluent. Cryovials were then placed on a shaker for 4 h at room temperature and centrifuged at 14,000 rpm for 10 min at room temperature. Supernatants were immediately analyzed.
2.4. Materials
All analyses were conducted on the ADVIA Centaur® XP Immunoassay System (Siemens, Healthineers) using reagents [SARS-CoV-2 IgG (sCOVG) assay, and SARS-CoV-2 Total (COV2T) assay], calibrators, controls and consumables as recommended by the manufacturer. The ADVIA Centaur COV2T assay is a qualitative assay which detects total antibodies (including IgG and IgM) to SARS-CoV-2 in human serum and plasma, while the sCOVG assay qualitatively and semi-quantitatively identifies SARS-CoV-2 IgG antibodies. Both assays generate numerical output which is reported in this manuscript as number indices.
2.5. Serial dilution assay
The linearity of COV2T and sCOVG recovery following serial dilutions was assayed using both plasma and DBS eluates. The following dilution series was performed on 10 SARS-CoV-2 positive remnant whole blood specimen in lithium heparin syringes: no dilution (Neat), 1:2, 1:4, 1:8, and 1:16. Each positive specimen was diluted into a pool of remnant whole blood that was negative for antibodies to SARS-CoV-2 (as determined by both the COV2T and sCOVG assays). After dilution, aliquots (60ul) of whole blood dilution sequence were spotted (5 spots per specimen) onto Whatman 903 DBS cards and frozen at −80 °C. The remaining whole blood for each dilution was then centrifuged, plasma separated, and stored at −80 °C. Eluates were acquired as described above.
2.6. Statistics
Positive and negative percent agreement (PPA and NPA respectively) of the DBS result was assessed relative to the plasma specimen. ROC curves were generated relative to the paired plasma specimen and ideal cutoffs for each assay to maximize PPA and NPA were identified using Youdens Index. For linearity studies, if the neat specimen had a signal greater than the limit of detection, the relative recovery for sCOVG was determined by dividing subsequent dilutions with the value from the 1:2 specimen. Concordance was determined using Cohens Kappa. Significance was determined using unpaired student T test or a Mann Whitney-U test for non-normal distributed data as indicated. All statistical analyses were performed with GraphPad Prism 8.
3. Results
The lowest background RLUs were observed for PBS (15,066 AU) and deionized water (15,905 AU) for the SCOVG assay (Supplemental Table 1 ). The highest background was observed with ECOSURF (102,252 AU). We then compared background RLUs for both the COV2T and sCOVG assays with capillary blood specimen from 3 patients with negative molecular testing for SARS-CoV-2. Deionized water eluates exhibited lower overall background RLUs compared to PBS and was used as the eluent for the remainder of the study. The imprecision of extraction with water over three replicates was 19.2%. Similar background RLUs were observed for PBS (32,879 AU) and water (16,396 AU) as eluents using the COV2T assay (Supplemental Table 2).
Table 1.
Assay agreement between plasma and DBS samples.
| Dilution Factor | DBS COV2T | DBS sCOVG |
|---|---|---|
| Agreement1 | Agreement2 | |
| Neat | 9/9 3 | 10/10 |
| 1:2 | 10/10 | 10/10 |
| 1:4 | 10/10 | 10/10 |
| 1:8 | 9/9 3 | 10/10 |
| 1:16 | 10/10 | 9/10 |
| TOTAL | 48/48 (100%, 90.8–100) | 49/50 (98.0%, 88.0–99.9) |
Positive DBS COV2T signal is ≥ 0.56.
Positive when DBS sCOVG is ≥ 0.1.
Specimen was QNS to repeat after instrument error.
Patient demographics are listed in Supplemental Table 3. The median age of naturally infected males and females were 58 (Range ± 10) and 59 (Range ± 20) years respectively; and 42 (Range ± 9.5) and 42 (Range ± 23.5) years for vaccinated males and females, respectively. 83.3% and 70.5% of female and male patients who were naturally infected with SARS-CoV-2 were symptomatic at the time of sample collection. The median number of days from positive PCR test to sample collection in naturally infected subjects were 15.5 ± 23 and 12 ± 16.5 days, for males and females respectively. In vaccinated patients, median sample collection from the date of receipt of the 2nd dose was 122 ± 18.25 and 129 ± 22.25 days for males and females respectively.
At the manufacturer defined cutoff of 1.0 for the COV2T assay, the PPA was 94.4% (95% CI: 94.9–100%) and the NPA was 100% (95% CI: 85.13–100) for DBS specimens relative to plasma (Fig. 1 A, 1B). At the manufacturer defined cutoff Index of 1.0 for the sCOVG assay, the PPA was 79.2 (95% CI; 68.4–87.0) and the NPA was 100 (95% CI; 87.1–100.0) for DBS cards relative to plasma specimens (Fig. 1C, 1D).
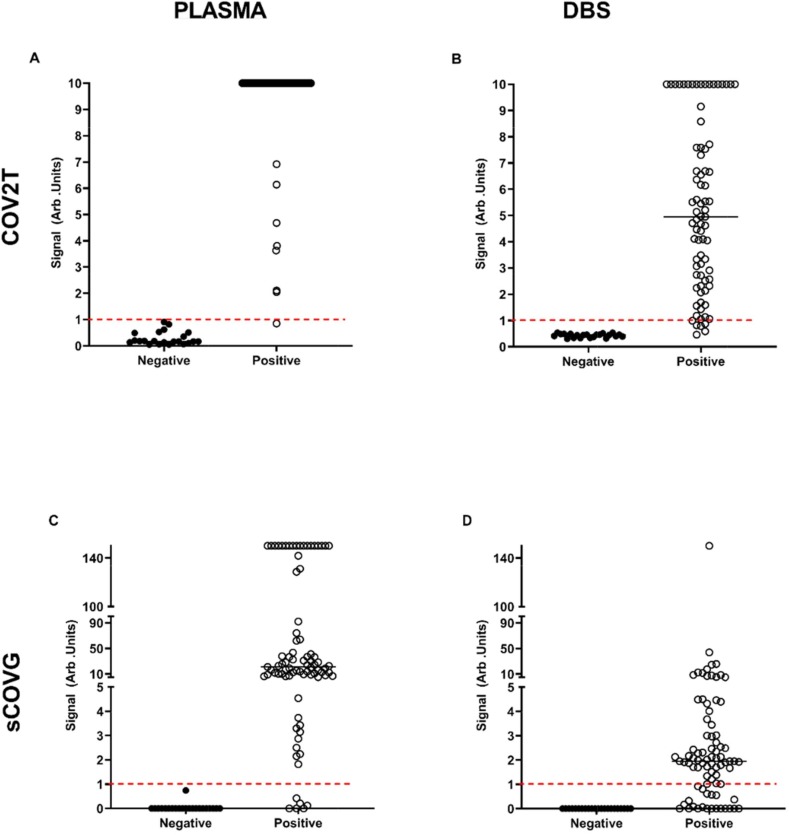
Fig. 1.
Signal indices for COV2T in A) Plasma and B) DBS specimens, and sCOVG in C) Plasma and D) DBS specimens. Dotted red line indicates the manufacturer’s recommended cutoff for positivity. n = 23–78. (For interpretation of the references to colour in this figure legend, the reader is referred to the web version of this article.)
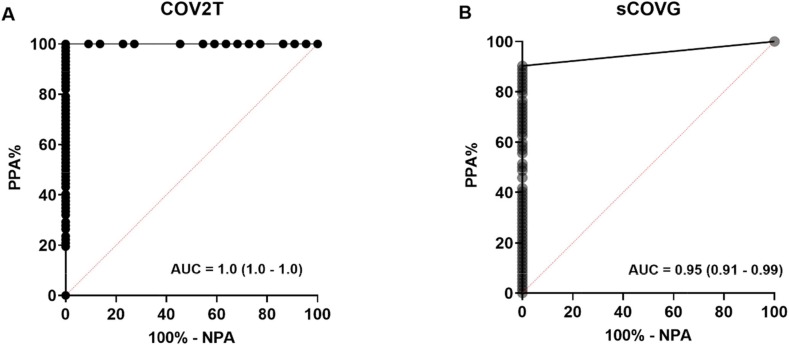
ROC curves for DBS specimens analyzed by the COV2T assay revealed an AUC of 1.0 (95% CI: 1.0–1.0) (Fig. 2 A) and by the sCOVG assay revealed an AUC of 0.95 (95% CI: 0.91–0.99) (Fig. 2B). Using Youdens Index, an ideal cutoff of 0.56 Index on the COV2T assay demonstrated a PPA of 100.0% (95% CI: 94.9–100.0) and an NPA of 100% (95% CI: 85.1–100.0) for DBS specimens. An Index of 0.1 on the sCOVG assay revealed a PPA of 90.3% (95% CI; 81.3–95.2%) and an NPA of 100% (95% CI; 87.1–100.0%). Using these cutoffs, an observed Kappa of 0.87 (95% CI: 0.76–0.98) was calculated between sCOVG and COV2T assays in plasma samples, and 0.66 (95% CI: 0.50–0.82) with DBS specimens (Supplemental Tables 4 and 5). At the manufacturer Limit of Detection of 0.5 Index for DBS specimens, the PPA with plasma was 85.7% (95%; 76.2–91.8) with no change in NPA.
Fig. 2.
ROC Curves of percent agreement between plasma and DBS-extracted specimens. A) The COV2T assay displayed high sensitivity and specificity (AUC = 1) with DBS sampling. B) The sCOVG assay displayed high sensitivity and specificity (AUC = 0.95) with DBS sampling.
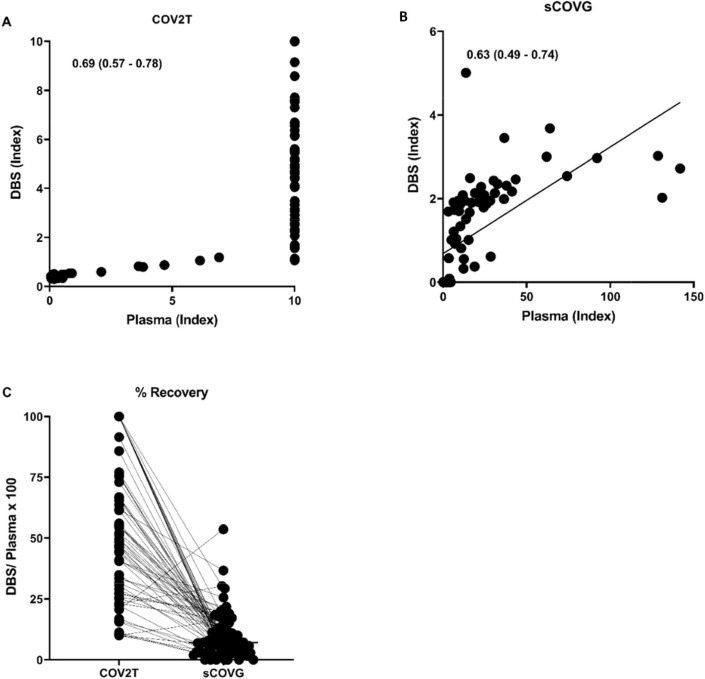
A positive correlation was observed between the signal Index from DBS eluates and plasma samples on the COV2T assay (Pearson r = 0.69, 95% CI; 0.57–0.78) and sCOVG assay (Pearson r = 0.63, (95% CI; 0.49–0.74) (Fig. 3 A-B). Overall, higher recovery of the chemiluminescent signal was observed with the COV2T assay [Median 51.4% (95% CI: 41.1–63.7)] compared to the sCOVG assay [Median 7.085% (95% CI: 5.473–9.130)] (Fig. 3C).
Fig. 3.
Correlation and recovery in DBS relative to plasma specimen. (A) COV2T recovery from DBS relative to plasma on the COV2T assay, (B) sCOVG recovery in DBS is relative to sCOVG recovery from plasma specimens. Values represent Pearson r and 95% CI. C) Percent recovery of COV2T versus sCOVG in SARS-CoV-2 positive patients.
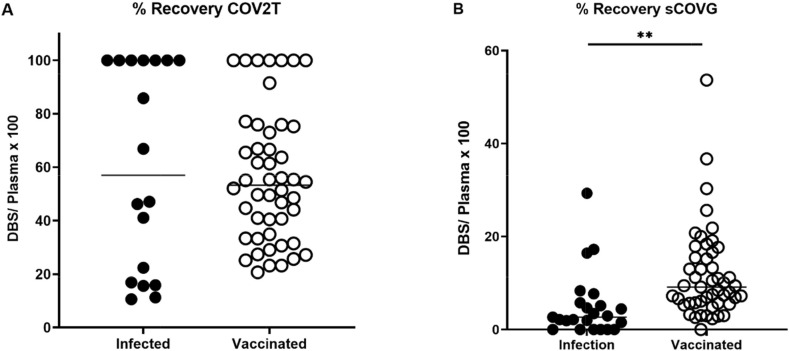
Relative to patients with natural infection, vaccinated patients had on average a higher Index by the COV2T and sCOVG assays in both plasma and DBS eluates (Supplemental Fig. 1). Nonetheless, similar recovery from DBS eluates relative to plasma was observed for the COV2T assay in infected patient samples (Median = 46.2%, 95% CI: 20.7 – 100) relative to vaccinated patient samples (Median = 51.75%, 95% CI: 44.1 – 6i1.7%, Fig. 4 A). However, sCOVG recovery from DBS eluates was lower relative to plasma (P-value < 0.0001) when patients were naturally infected (Median = 2.67%, 95% CI: 1.541 – 5.133) compared to vaccinated (Median = 9.13%, 6.94 – 11.22, Fig. 4B).
Fig. 4.
Recovery of COV2T and sCOVG in infected or vaccinated patients. (A) COV2T recovery and B) sCOVG recovery in DBS eluates in infected or vaccinated patients. n = 33 (infected), n = 50 (vaccinated).
Serial dilutions revealed a qualitative agreement of 100% (95% CI: 90.8–100%) between the plasma specimen with a cutoff Index of 1.0 and the DBS specimen with a cutoff Index of 0.56 on the COV2T assay (Table 1). Similarly, qualitative agreement using the sCOVG assay was 98.0% (88.0–99.9%) across all dilutions with a cutoff Index of 1.0 for plasma and 0.1 for DBS eluates. Interestingly, the single discordant sample was negative in plasma at a dilution of 1:16 but positive by DBS.
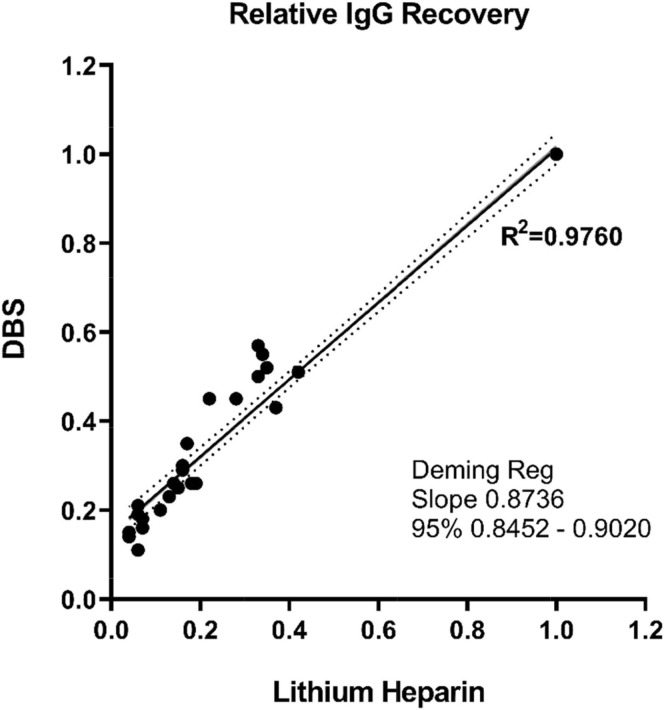
Since the sCOVG assay is designated as semi-quantitative, relative IgG recovery was assessed. Deming regression of the relative IgG recovery with plasma relative to DBS eluates demonstrated an R2 = 0.976 (Fig. 5 ). The Deming regression best fit line had a slope of 0.87 (95% CI 0.84–0.90) and an intercept of 0.14 (0.12–0.17).
Fig. 5.
Regression of sCOVG from serial dilutions. The solid line represents the Deming Regression slope, the dashed lines represent the 95% CI of the slope. The black solid line represents the line of best fit with R2 = 0.9760.
4. Discussion
DBS sampling has been proposed as a stable, cost-effective, and minimally invasive sampling technique with potential for broad usage in the clinical laboratory [13], [15], [17], [20]. In this study, we demonstrate that DBS specimens have excellent agreement with plasma specimens using the Siemens COV2T and sCOVG assays. Furthermore, we demonstrated strong correlation between the Index signal generated by both assays assessed in plasma and DBS. Together, these results imply a potential utility for DBS in serological testing.
It is crucial for any assay, particularly those to be used as a screening assay or for seroprevalence studies, to have high sensitivity and specificity for the disease of interest [4], [31]. To this end, we observed high PPA and NPA at the manufacturer defined cutoff index of 1.0 between plasma and DBS specimens. Furthermore, we demonstrate that reducing the cutoff Index to 0.56 and 0.1 for the COV2T and sCOVG respectively increased the PPA and NPA to almost 100%. The reduction in signal was necessary for both assays given the reduction in % recovery for both total and IgG antibodies. Reduced recovery was likely multifactorial and includes differences in antibody concentrations in whole blood specimen versus plasma specimen. Since antibodies are found in the plasma fraction, the presence of hematocrit in whole blood reduces antibody concentrations by ∼ 40–50% relative to plasma. Also, we demonstrate water as a simple elution buffer anticipated to maximize specificity for the sCOVG and COV2T Siemens assays since it resulted in less background signal relative to other buffers tested. Other studies have used 0.1% Tween-20/5% nonfat milk protein/PBS eluent [29] or 1% Tween-20/PBS [32]. Reducing background signal is likely to enhance specificity and positive predictive value; parameters crucial for serosurveillance studies. Interestingly, better recovery was observed with the COV2T assay which detects total antibodies than the sCOVG assay which detects IgG antibodies. This is unexpected, as one would anticipate that the pentameric IgM would be less likely to diffuse from the DBS card than IgG antibodies. This could have been impacted by the buffer chosen for elution. Nonetheless, this demonstrates the need to validate any assay and analyte from a DBS card prior to analysis with patient specimens. Importantly, by reducing the Index signal required for a qualitatively positive DBS specimen to 0.56 and 0.1 for the COV2T and sCOVG respectively, PPA was enhanced without sacrificing NPA. This approach has been used in other studies to maximize the sensitivity and specificity of DBS eluents for SARS-CoV-2 serological testing but with a lower number of patients to validate the established cutoffs [33]. Furthermore, one large study found that DBS eluents had a 30-fold higher limit of detection relative to plasma using the Roche Elecsys immunoassay [34]. Together, these studies reinforce the approach used here to maximize PPA and NPA by reducing the cutoff for a positive test result. However, follow-up studies are required to demonstrate the imprecision of both the COV2T and the SCOVG assays at cutoffs that are below the limit of quantitation.
An interesting finding from this study was the potential for semi-quantitative results from DBS eluates. We demonstrate a strong Pearson correlation with patient comparisons, excellent agreement with dilution studies, and linearity using the sCOVG assay. To our knowledge, little data exists in the available literature demonstrating linear results after elution from a DBS card. In the context of SARS-CoV-2 serology, we previously demonstrated that higher signals generated on automated serological assays are generally associated with higher neutralizing antibody titers [35], [36]. Furthermore, studies in the context of mRNA vaccine have demonstrated lower neutralizing antibody titers in those with breakthrough SARS-CoV-2 infections than those without [37]. In the absence of peer reviewed data on the use of SARS-Cov-2 serological testing as a marker of immunity from infection, the FDA does not currently recommend serological assays to assess for immunity after COVID-19 vaccination or primary infection [38]. However, recent recommendations for immunocompromised patients to receive an additional SARS-Cov-2 vaccine, coupled with the release of the SARS-CoV-2 antibody reference material by the WHO [39], may help standardize assays and cutoffs in the future leading to recommendations regarding serological testing cutoffs and protection. Nonetheless, the quantitative results from DBS testing demonstrated here imply that the same technique may be useful for other, quantitative analytes.
DBS collection has been used clinically for NBS screening for several decades now [11]. This is due, at least in part, to ease of collection in pediatric populations, simplicity of transport, and robust testing methods for multiple analytes. While other studies have used DBS cards for research, few have transitioned to translational or clinical applications [13], [14], [15], [16], [17], [18], [19], [20]. To this end, serological testing from DBS cards has the potential to be useful early in an epidemic or pandemic to better understand both symptomatic and asymptomatic spread of disease in difficult to reach populations particularly those far from testing centers, or in undeveloped countries. With regards to underdeveloped countries, there are limited available studies assessing seroprevalence in these populations, likely obscuring the global impact of the COVID-19 pandemic.
Despite implications of this data in DBS-based SARS-Cov-2 serology, our study has several limitations. The use of remnant specimens may have impacted findings, with the lithium heparin from the syringe altering the matrix relative to a whole blood capillary samples. However, we generally observed comparable or better recovery of signal from DBS relative to plasma in the remnant specimens relative to the prospectively collected, vaccinated individuals. Of note, % recovery of sCOVG was two-fold higher in DBS specimens from vaccinated relative to infected patients although no difference in % recovery was observed with the COV2T assay. These results may be due to enhanced recovery with higher starting antibody concentrations in vaccinated individuals, a matrix effect from using remnant lithium heparin whole blood, and differing SARS-Cov-2 seroprevalence in infected patients due to differences in disease severity with breakthrough infections. Larger, prospective studies to tease out the cause of this potential discrepancy are required. Another limitation of this study was the selection of diluent. Given the general concern for low specificity of SARS-CoV-2 immunoassays when used for seroprevalence, we selected elution buffers based on the lowest signal produced in a negative sample. However, we did not assess the buffer most suited to recover immunoglobulins from DBS cards. This may have resulted in lower recoveries than could have been achieved using other buffers. Furthermore, the use of water to elute DBS cards should be validated across other platforms before implementation for studies or testing.
In conclusion, we demonstrate a high percent agreement between plasma and DBS specimens for SARS-COV-2 total and IgG antibodies using the Siemens Centaur XP COV2T and sCOVG assays. The benefits of DBS specimen collection include the minimally invasive nature of sampling, rapid screening of patients, reduced exposure of healthcare workers to infected patients during venipuncture sampling, sample collection for patients under quarantine, and expanding access to vulnerable populations with limited access to COVID-19 testing centers and hospitals.
5. Disclaimer
The information in this paper is based on research results that are not commercially available
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.clinbiochem.2021.12.012.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- 1.Hanson K.E., Caliendo A.M., Arias C.A., Englund J.A., Hayden M.K., Lee M.J., Loeb M., Patel R., Altayar O., El Alayli A., Sultan S., Falck-Ytter Y., Lavergne V., Morgan R.L., Murad M.H., Bhimraj A., Mustafa R.A. Infectious Diseases Society of America Guidelines on the Diagnosis of Coronavirus Disease 2019 (COVID-19): Serologic Testing. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa1343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.CDC, Cases, Data, and Surveillance, Cent. Dis. Control Prev. (2020). https://www.cdc.gov/coronavirus/2019-ncov/covid-data/serology-surveillance/index.html (accessed September 7, 2021).
- 3.Jones J.M., Stone M., Sulaeman H., Fink R.V., Dave H., Levy M.E., Di Germanio C., Green V., Notari E., Saa P., Biggerstaff B.J., Strauss D., Kessler D., Vassallo R., Reik R., Rossmann S., Destree M., Nguyen K.-A., Sayers M., Lough C., Bougie D.W., Ritter M., Latoni G., Weales B., Sime S., Gorlin J., Brown N.E., Gould C.V., Berney K., Benoit T.J., Miller M.J., Freeman D., Kartik D., Fry A.M., Azziz-Baumgartner E., Hall A.J., MacNeil A., Gundlapalli A.V., Basavaraju S.V., Gerber S.I., Patton M.E., Custer B., Williamson P., Simmons G., Thornburg N.J., Kleinman S., Stramer S.L., Opsomer J., Busch M.P. Estimated US Infection- and Vaccine-Induced SARS-CoV-2 Seroprevalence Based on Blood Donations, July 2020-May 2021. JAMA. 2021;326(14):1400. doi: 10.1001/jama.2021.15161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Farnsworth C.W., Anderson N.W. SARS-CoV-2 Serology: Much Hype, Little Data. Clin. Chem. 2020;66:875–877. doi: 10.1093/clinchem/hvaa107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Florin L., Maelegheer K., Vandewal W., Bernard D., Robbrecht J. Performance Evaluation of the Siemens SARS-CoV-2 Total Antibody and IgG Antibody Test. Lab. Med. 2021;52(6):e147–e153. doi: 10.1093/labmed/lmab027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.GeurtsvanKessel C.H., Okba N.M.A., Igloi Z., Bogers S., Embregts C.W.E., Laksono B.M., Leijten L., Rokx C., Rijnders B., Rahamat-Langendoen J., van den Akker J.P.C., van Kampen J.J.A., van der Eijk A.A., van Binnendijk R.S., Haagmans B., Koopmans M. An evaluation of COVID-19 serological assays informs future diagnostics and exposure assessment. Nat. Commun. 2020;11:3436. doi: 10.1038/s41467-020-17317-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tang M.S., Hock K.G., Logsdon N.M., Hayes J.E., Gronowski A.M., Anderson N.W., Farnsworth C.W. Clinical Performance of Two SARS-CoV-2 Serologic Assays. Clin. Chem. 2020;66:1055–1062. doi: 10.1093/clinchem/hvaa120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jia X., Zhang P., Tian Y., Wang J., Zeng H., Wang J., Liu J., Chen Z., Zhang L., He H., He K., Liu Y. Clinical Significance of an IgM and IgG Test for Diagnosis of Highly Suspected COVID-19. Front. Med. 2021;8 doi: 10.3389/fmed.2021.569266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sidiq Z., Hanif M., Dwivedi K.K., Chopra K.K. Benefits and limitations of serological assays in COVID-19 infection. Indian J. Tuberc. 2020;67(4):S163–S166. doi: 10.1016/j.ijtb.2020.07.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Whitman J.D., Hiatt J., Mowery C.T., Shy B.R., Yu R., Yamamoto T.N., Rathore U., Goldgof G.M., Whitty C., Woo J.M., Gallman A.E., Miller T.E., Levine A.G., Nguyen D.N., Bapat S.P., Balcerek J., Bylsma S.A., Lyons A.M., Li S., Wong A.-y., Gillis-Buck E.M., Steinhart Z.B., Lee Y., Apathy R., Lipke M.J., Smith J.A., Zheng T., Boothby I.C., Isaza E., Chan J., Acenas D.D., Lee J., Macrae T.A., Kyaw T.S., Wu D., Ng D.L., Gu W., York V.A., Eskandarian H.A., Callaway P.C., Warrier L., Moreno M.E., Levan J., Torres L., Farrington L.A., Loudermilk R.P., Koshal K., Zorn K.C., Garcia-Beltran W.F., Yang D., Astudillo M.G., Bernstein B.E., Gelfand J.A., Ryan E.T., Charles R.C., Iafrate A.J., Lennerz J.K., Miller S., Chiu C.Y., Stramer S.L., Wilson M.R., Manglik A., Ye C.J., Krogan N.J., Anderson M.S., Cyster J.G., Ernst J.D., Wu A.H.B., Lynch K.L., Bern C., Hsu P.D., Marson A. Evaluation of SARS-CoV-2 serology assays reveals a range of test performance. Nat. Biotechnol. 2020;38(10):1174–1183. doi: 10.1038/s41587-020-0659-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.De Jesús V.R., Mei J.V., Bell C.J., Hannon W.H. Improving and Assuring Newborn Screening Laboratory Quality Worldwide: 30-Year Experience at the Centers for Disease Control and Prevention. Semin. Perinatol. 2010;34(2):125–133. doi: 10.1053/j.semperi.2009.12.003. [DOI] [PubMed] [Google Scholar]
- 12.Malsagova K., Kopylov A., Stepanov A., Butkova T., Izotov A., Kaysheva A. Dried Blood Spot in Laboratory: Directions and Prospects. Diagnostics. 2020;10:248. doi: 10.3390/diagnostics10040248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Cox H.D. Dried Blood Spots May Improve Detection of Blood Doping. Clin. Chem. 2019;65:1481–1483. doi: 10.1373/clinchem.2019.311902. [DOI] [PubMed] [Google Scholar]
- 14.Cox H.D., Miller G.D., Manandhar A., Husk J.D., Jia X., Marvin J., Ward D.M., Phillips J., Eichner D. Measurement of Immature Reticulocytes in Dried Blood Spots by Mass Spectrometry. Clin. Chem. 2021;67:1071–1079. doi: 10.1093/clinchem/hvab058. [DOI] [PubMed] [Google Scholar]
- 15.Thevis M. Broadening the Horizon of Antidoping Analytical Approaches Using Dried Blood Spots. Clin. Chem. 2021;67:1041–1043. doi: 10.1093/clinchem/hvab074. [DOI] [PubMed] [Google Scholar]
- 16.Gaugler S., Al-Mazroua M.K., Issa S.Y., Rykl J., Grill M., Qanair A., Cebolla V.L. Fully Automated Forensic Routine Dried Blood Spot Screening for Workplace Testing. J. Anal. Toxicol. 2019;43:212–220. doi: 10.1093/jat/bky074. [DOI] [PubMed] [Google Scholar]
- 17.Coats J.T., Dillon J.F. The effect of introducing point-of-care or dried blood spot analysis on the uptake of hepatitis C virus testing in high-risk populations: A systematic review of the literature. Int. J. Drug Policy. 2015;26(11):1050–1055. doi: 10.1016/j.drugpo.2015.05.001. [DOI] [PubMed] [Google Scholar]
- 18.Jack K., Irving W.L. Using dried blood spot testing for diagnosing viral hepatitis. Br. J. Nurs. Mark Allen Publ. 2020;29:1155–1158. doi: 10.12968/bjon.2020.29.20.1155. [DOI] [PubMed] [Google Scholar]
- 19.Bottino P., Balloco C., Rittà M., Bianco G., Sidoti F., Cavallo R., Costa C. Evaluation of CMV DNA in dried blood spot. New Microbiol. 2020;43:93–95. [PubMed] [Google Scholar]
- 20.Tuaillon E., Kania D., Pisoni A., Bollore K., Taieb F., Ontsira Ngoyi E.N., Schaub R., Plantier J.-C., Makinson A., Van de Perre P. Dried Blood Spot Tests for the Diagnosis and Therapeutic Monitoring of HIV and Viral Hepatitis B and C. Front. Microbiol. 2020;11:373. doi: 10.3389/fmicb.2020.00373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Diener C., Galata V., Keller A., Meese E. MicroRNA profiling from dried blood samples. Crit. Rev. Clin. Lab. Sci. 2019;56(2):111–117. doi: 10.1080/10408363.2018.1561641. [DOI] [PubMed] [Google Scholar]
- 22.Vreeker G.C.M., Bladergroen M.R., Nicolardi S., Mesker W.E., Tollenaar R.A.E.M., van der Burgt Y.E.M., Wuhrer M. Dried blood spot N-glycome analysis by MALDI mass spectrometry. Talanta. 2019;205:120104. doi: 10.1016/j.talanta.2019.06.104. [DOI] [PubMed] [Google Scholar]
- 23.Malvagia S., Ferri L., Della Bona M., Borsini W., Cirami C.L., Dervishi E., Feriozzi S., Gasperini S., Motta S., Mignani R., Trezzi B., Pieruzzi F., Morrone A., Daniotti M., Donati M.A., la Marca G. Multicenter evaluation of use of dried blood spot compared to conventional plasma in measurements of globotriaosylsphingosine (LysoGb3) concentration in 104 Fabry patients. Clin. Chem. Lab. Med. CCLM. 2021;59:1516–1526. doi: 10.1515/cclm-2021-0316. [DOI] [PubMed] [Google Scholar]
- 24.Ondoa P., Kebede Y., Loembe M.M., Bhiman J.N., Tessema S.K., Sow A., Sall A.A., Nkengasong J. COVID-19 testing in Africa: lessons learnt. Lancet Microbe. 2020;1(3):e103–e104. doi: 10.1016/S2666-5247(20)30068-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Songok E. A locally sustainable approach to COVID-19 testing in Africa. Lancet Microbe. 2020;1(5):e197. doi: 10.1016/S2666-5247(20)30118-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gosselin R.C., Bowyer A., Favaloro E.J., Johnsen J.M., Lippi G., Marlar R.A., Neeves K., Rollins‐Raval M.A. Guidance on the critical shortage of sodium citrate coagulation tubes for hemostasis testing. J. Thromb. Haemost. JTH. 2021;19(11):2857–2861. doi: 10.1111/jth.15514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.C. for D. and R. Health, Medical Device Shortages During the COVID-19 Public Health Emergency, FDA. (2021). https://www.fda.gov/medical-devices/coronavirus-covid-19-and-medical-devices/medical-device-shortages-during-covid-19-public-health-emergency (accessed December 6, 2021).
- 28.Gaugler S., Sottas P.-E., Blum K., Luginbühl M. Fully automated dried blood spot sample handling and extraction for serological testing of SARS-CoV-2 antibodies. Drug Test. Anal. 2021;13(1):223–226. doi: 10.1002/dta.2946. [DOI] [PubMed] [Google Scholar]
- 29.Moat S.J., Zelek W.M., Carne E., Ponsford M.J., Bramhall K., Jones S., El-Shanawany T., Wise M.P., Thomas A., George C., Fegan C., Steven R., Webb R., Weeks I., Morgan B.P., Jolles S. Development of a high-throughput SARS-CoV-2 antibody testing pathway using dried blood spot specimens. Ann. Clin. Biochem. 2021;58(2):123–131. doi: 10.1177/0004563220981106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Turgeon C.T., Sanders K.A., Granger D., Nett S.L., Hilgart H., Matern D., Theel E.S. Detection of SARS-CoV-2 IgG antibodies in dried blood spots. Diagn. Microbiol. Infect. Dis. 2021;101(1):115425. doi: 10.1016/j.diagmicrobio.2021.115425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Theel E.S., Slev P., Wheeler S., Couturier M.R., Wong S.J., Kadkhoda K., McAdam A.J. The Role of Antibody Testing for SARS-CoV-2: Is There One? J. Clin. Microbiol. 2020;58(8) doi: 10.1128/JCM.00797-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Amini F., Auma E., Hsia Y., Bilton S., Hall T., Ramkhelawon L., Heath P.T., Le Doare K., Paxton W.A. Reliability of dried blood spot (DBS) cards in antibody measurement: A systematic review. PloS One. 2021;16(3):e0248218. doi: 10.1371/journal.pone.0248218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fontaine E., Saez C. Analysis of SARS-CoV-2 antibodies from dried blood spot samples with the Roche Elecsys Immunochemistry method. Pract. Lab. Med. 2021;25:e00234. doi: 10.1016/j.plabm.2021.e00234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mulchandani R., Brown B., Brooks T., Semper A., Machin N., Linley E., Borrow R., Wyllie D., Taylor-Philips S., Jones H., Oliver I., Charlett A., Hickman M., Brooks T., Mulchandani R., Wyllie D. EDSAB-HOME Study Investigators, Use of dried blood spot samples for SARS-CoV-2 antibody detection using the Roche Elecsys ® high throughput immunoassay. J. Clin. Virol. Off. Publ. Pan Am. Soc. Clin. Virol. 2021;136:104739. doi: 10.1016/j.jcv.2021.104739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Farnsworth C.W., Case J.B., Hock K., Chen R.E., O'Halloran J.A., Presti R., Goss C.W., Rauseo A.M., Ellebedy A., Theel E.S., Diamond M.S., Henderson J.P. Assessment of serological assays for identifying high titer convalescent plasma. Transfusion (Paris). 2021;61(9):2658–2667. doi: 10.1111/trf.16580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tang M.S., Case J.B., Franks C.E., Chen R.E., Anderson N.W., Henderson J.P., Diamond M.S., Gronowski A.M., Farnsworth C.W. Association between SARS-CoV-2 neutralizing antibodies and commercial serological assays. Clin. Chem. 2020;66(12):1538–1547. doi: 10.1093/clinchem/hvaa211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lange B., Gerigk M., Tenenbaum T. Breakthrough Infections in BNT162b2-Vaccinated Health Care Workers. N. Engl. J. Med. 2021;385(12):1145–1146. doi: 10.1056/NEJMc2108076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.C. for D. and R. Health, Antibody Testing Is Not Currently Recommended to Assess Immunity After COVID-19 Vaccination: FDA Safety Communication, FDA. (2021). https://www.fda.gov/medical-devices/safety-communications/antibody-testing-not-currently-recommended-assess-immunity-after-covid-19-vaccination-fda-safety (accessed September 3, 2021).
- 39.First WHO International Standard Anti-SARS-CoV-2 Immunoglobulin (Human), (n.d.). https://www.nibsc.org/products/brm_product_catalogue/detail_page.aspx?catid=20/136 (accessed March 22, 2021).
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.