Abstract
Neuropathic pain, which is accompanied by an unpleasant sensation, affects the patient’s quality of life severely. Considering the complexity of the neuropathic pain, there are huge unmet medical needs for it while current effective therapeutics remain far from satisfactory. Accordingly, exploration of mechanisms of neuropathic pain could provide new therapeutic insights. While numerous researches have pointed out the contribution of sensory neuron-immune cell interactions, other mechanisms of action, such as long non-coding RNAs (lncRNAs), also could contribute to the neuropathic pain observed in vivo. LncRNAs have more than 200 nucleotides and were originally considered as transcriptional byproducts. However, recent studies have suggested that lncRNAs played a significant role in gene regulation and disease pathogenesis. A substantial number of long non-coding RNAs were expressed differentially in neuropathic pain models. Besides, therapies targeting specific lncRNAs can significantly ameliorate the development of neuropathic pain, which reveals the contribution of lncRNAs in the generation and maintenance of neuropathic pain and provides a new therapeutic strategy. The primary purpose of this review is to introduce recent studies of lncRNAs on different neuropathic pain models.
Keywords: neuropathic pain, long non-coding RNAs, pain model, treatment, microarray analysis
Introduction
Overview of Neuropathic Pain
Pain is a common disease in clinical practice, which has important implications for the human body. Pain perception has evolved as a warning role to respond to organismic danger and tissue damage in a timely manner, then it presents a protective effect by generating related danger signals.1 Patients are deeply troubled by the continuous generation of pain signals and the dysfunction of the nervous system. As generally known, neuropathic Pain is a common chronic pain in clinical practice. It is generally caused by the dysfunction of somatosensory neurons in the central or peripheral nervous systems.2 Recently, neuropathic pain was redefined as “pain initiated or caused by a primary lesion or dysfunction in the nervous system” by the International Association for the Study of Pain (IASP).3 The primary diseases include neurological damage caused by cancer, diabetes, infection, autoimmune diseases, chemotherapy, or trauma.4 These conditions usually lead to chronic pain syndrome, manifesting as spontaneous pain, hyperalgesia and allodynia. In addition, chronic pain has been shown to cause sleep disturbances, mental illnesses and difficulty in concentrating.5 Nearly 8% of the world’s people suffer from chronic pain, which seriously affects the quality of life.6
The primary causes for those symptoms of chronic pain include alteration of ion channels (such as calcium and sodium channels) in neurons,7,8 activation of immune cells (like microglia, macrophage and CD4+T cell et al) and systemic inflammatory imbalance.1,9,10 These results ultimately lead to imbalance between the ascending pain pathways and the descending inhibitory pathway. In sensory transduction, dorsal root ganglion (DRG) is the gathering place of the somatosensory primary afferent neuron cell body and it is of vital importance in the signal transmission between peripheral stimuli and central sensory system.11 Admittedly, the pathogenesis of neuropathic pain is complicated. In the beginning, it seems that few neurons were damaged, which led to abnormal electrical activity, synaptic changes and continuous inflammation.12,13 Further studies have found that the production of chronic pain includes the abnormal production of pro-inflammatory mediators and the interaction between neurons and immune cells as well.14,15 It contains a complex network system composed of microglia and macrophages.15,16 For example, colony-stimulating factor 1 (CSF1) induces microglial proliferation and aggravates the pain dramatically.9 Reduction of macrophages in DRG can relieve mechanical and thermal hyperalgesia.17 Besides, Some new studies show that receptors such as melanocortin 4 receptor (MC4R) and Toll-like receptor 4 (TLR4) are also helpful.18,19 Here, we briefly reviewed recent studies of lncRNAs in neuropathic pain, focusing on different rodent neuropathic pain models.
Biological Functions of Long Non-Coding RNAs
The human transcriptome has pointed that only 1–2% of the human genome is used to encode proteins, and the remaining 98% is non-coding RNA. LncRNAs is one of the non-coding RNA without protein-coding function. However, an important progression has been the emergence of lncRNAs as key post-transcriptional regulators.20 Crucial roles for lncRNAs have been identified in conditions such as cardiovascular diseases, various cancers, autoimmune disorders, kidney and liver system, etc.21–23 Subsequent studies have shown that lncRNAs play an essential role in gene expression by co-transcriptional regulation, post-transcriptional bridging proteins and chromatin, pairing with other RNA and chromosomal modification,24,25 especially in the development, proliferation, differentiation, migration and invasion of tumors cells.26 What’s more, some lncRNAs could play a role in an indirect way, which interacts with microRNAs (miRNAs) and proteins to regulate signal transduction pathways downstream, affecting the development of various diseases.27,28
Bioinformatic analysis and silico inference methods reveal potential interactions between lncRNAs-miRNAs and lncRNAs-RNA binding protein, which have attracted intense interest. MiRNAs, small non-coding RNAs, play an important role in the post-transcriptional regulation.27 The possible mechanisms for lncRNAs to regulate miRNA are as follows: (1) lncRNAs indirectly inhibits negative regulation of miRNA by competitively binding to the 3ʹ-UTR of target mRNA.29 (2) LncRNAs could act as a competitive endogenous RNA to play the “molecular sponge” and inhibit the expression of miRNA.30,31 (3) LncRNAs is a potential pri-miRNA, which indirectly regulates the expression of target genes.32 RNA-Protein interactions are key aspects in many biological processes.28 The primary concern of the present studies is shifting the attention from RNAs (like tRNAs, rRNAs) involved in translation and regulation towards lncRNAs.20 The Encyclopedia of DNA Elements shows that the correspondence between lncRNAs and RBP is not one-to-one. LncRNAs can interact with multiple RBP and generate biological efficacy via regulating expression of genes or preventing association with DNA.33 The association between them could be with potential therapeutic implications. While interactions of lncRNAs are more complex than previously thought, it still caused considerable concern and interest. It has been proven that lncRNAs plays an imperative role in tumors and participates in the generation and development of many diseases. In the meantime, a large amount of recent evidence shows that lncRNAs is also essential in neuropathic pain.34
The Profiles of Differentially Expressed LncRNAs in Neuropathic Pain
In recent years, with the rapid improvement of sequencing technology, many studies have conducted more in-depth research on neuropathic pain from the perspective of genes. Stacks of differentially expressed genes were found in the dorsal horn of the spinal cord, DRG and other regions in neuropathic pain models.35 Among them, it would have indicated that long-chain non-coding RNAs play a fundamental role in disease regulation. Therefore, the lncRNAs will become crucial targets for the treatment of neuropathic pain.36,37
Gene sequencing technology was being used in the rat neuropathic pain model to find many differentially expressed genes in the spinal cord and DRG. Therefore, bioinformatics analysis technology can be an ideal method to perform enrichment analysis and construct interactive network, revealing the molecular mechanism of lncRNAs in nerve damage and repair process.36 LncRNAs still, in turn, regulate neurite outgrowth of DRG neuron.36 Hu et al identified the differentially expressed lncRNAs in the spared nerve injury (SNL) model and found 1259 down-regulated and 2473 up-regulated lncRNAs in SNL rats. Among them, lncRNAs PKIA-AS1, small nucleolar RNA host gene 4(SNHG4), SH3 and cysteine-rich containing protein 3(STAC3) and CIRBP-AS1 were significantly up-regulated.38 Besides, more than 800 differentially expressed genes have been discovered in the chronic constriction injury model.39 A variety of differentially expressed genes were also detected in the neuropathic pain model of mice42–44 (Table 1). These studies provide the premise for studying the genetic level of neuropathic pain.
Table 1.
The Number of Differentially Expressed lncRNAs by Using Sequencing Technology in Multiple Models
| Model | Method | Total DE lncRNAs | Up-Regulation | Down-Regulation | Tissue | Species | Ref. |
|---|---|---|---|---|---|---|---|
| CCI | Microarray analysis | 876 | 339 | 537 | L3-L5 spinal cord | Rat | [39] |
| SNI | Microarray analysis | 105 | 81 | 24 | L4-L6 DRGs | Rat | [36] |
| SNI | Microarray analysis | lncRNAs change over time. | / | / | L4-L5 spinal cord | Rat | [40] |
| SNI | Microarray analysis | 112 | / | / | L4-L6 DRGs | Rat | [41] |
| SNL | Microarray analysis | 511 | 366 | 145 | L5 spinal cord | Mice | [42] |
| SNI | Microarray analysis | 1200 | / | / | L4-L5 spinal cord | Mice | [43] |
| SNI | Microarray analysis | 144 | 15 | 129 | L4−L5spinal cord | Rat | [44] |
| SNL | Microarray analysis | 1597 | / | / | Bilateral L4 DRGs | Mice | [45] |
| SNL | Microarray analysis | 3732 | 2473 | 1259 | L5 spinal cord | Rat | [38] |
| DNP | High throughput sequencing | 9 | 1 | 8 | L4‑L5 spinal cord | Mice | [46] |
| PIPN | High throughput sequencing | 412 | 145 | 267 | L4-L6 spinal cord dorsal horn | Rat | [47] |
Note: “/” Indicates no mention in the article.
Abbreviations: CCI, chronic constriction injury; SNI, spared nerve injury; SNL, spinal nerve ligation; DNP, diabetic neuropathic pain; PIPN, paclitaxel-induced peripheral neuropathy.
Major Neuropathic Pain Models
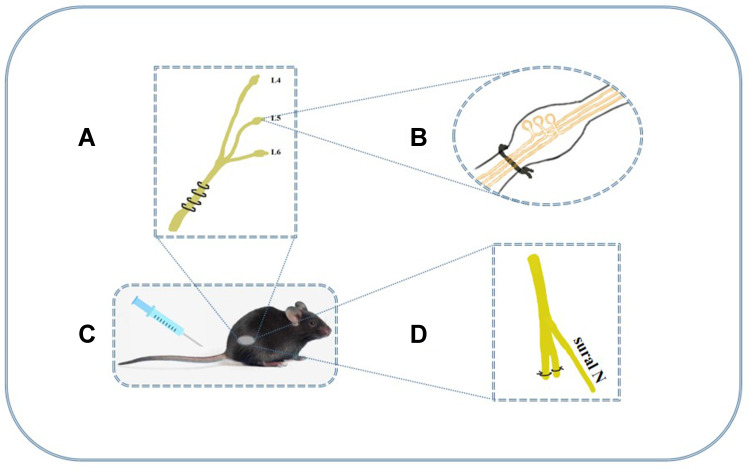
Promising new models for chronic pain provided a good base for further in-depth study of mechanisms. Photochemical and excitotoxic spinal cord injury are main central pain models. But the peripheral models seem to be more popular, which include chronic constriction injury (CCI), SNL, diabetic neuropathic pain (DNP), sciatic nerve injury (SCI) models and so on. In those studies, mechanical and thermal thresholds are important indicators to evaluate the hyperalgesia; and the content of inflammatory mediators usually reflects the degree of neuroinflammation. Nowadays, mature model of peripheral neuropathic pain has been established and major animal models are shown in Figure 1.48
Figure 1.
The major animal models of neuropathic pain. (A) The CCI model loosely ties the unilateral sciatic nerve below the mid-femur region with four chromic gut ligatures. It shows behavioral signs of spontaneous pain and hyperalgesia (thermal and tactile allodynia) and duration of pain signs persist over 2 months. (B) L5 and L6 spinal nerves of SNL model are tied unilaterally and closely at the distal side of dorsal root ganglia. It could indicate allodynia and hyperalgesia and last for at least 4 months. (C) DNP model was established by a single i.p. injection of streptozocin (STZ) and it induces thermal and mechanical hyperalgesia for 2–3 weeks.49 (D) In SNI model, the common peroneal and tibial nerves was ligated and axotomized, but sural nerve remained intact. Paw withdrawal threshold after SNI is significantly lower than sham group and hyperpathia lasts for at least 1 month.50
Latest Research Advances of LncRNAs in Neuropathic Pain Models
Based on the numerous researches on neuropathic pain-related lncRNAs, a large number of studies have found that lncRNAs are highly correlated with neuropathic pain, which provides a new target for its treatment.
SNL Model
There seems many lncRNAs expressed differentially in the SNL model (Table 2). Interaction between the lncRNAs and proteins is important. Firstly, Zhao et al discovered that the voltage-dependent potassium channel-related lncRNAs, KCNA2 antisense RNA, could regulate the excitability of DRG neurons by acting on zinc finger protein in rat, affecting the development and maintenance of neuropathic pain.51 The expression of lncRNA H19 in the DRG increased after SNL, and it is mainly up-regulated in Schwann cells—a non-primary sensory neuron, which likely be closely related to neuropathic pain.52 Recent studies have found that lncRNA embryonic stem cell expression 1 (lncenc1) is significantly upregulated in the DRG. Lncenc1 could bind to the RNA binding protein (RBP) enhancer of Zeste homologous protein 2 (EZH2), affecting the activation of microglia and the occurrence of neuropathic pain.53 Hu et al performed array analysis on the spinal cord and found that lncRNA PKIA-AS1 is highly expressed in SNL rat model. It has proved that PKIA-AS1 can mediate neuropathic pain by regulating the expression and function of CDK6 in experiments. It seems that most of lncRNAs may interact with receptor indirectly in SNL model.
Table 2.
Differential Expression of lncRNAs in SNL Model
| Model | Name of LncRNA | Location of the Gene | Expression and Significance | Pathway/Receptor | Biological Function | Tissue | Species | Gender | Ref. |
|---|---|---|---|---|---|---|---|---|---|
| SNL | H19 | Chromosome 1 1q41 |
↑ * | STAT3/TRPA1? | Activating inflammatory processes | Schwann cells/L5 DRG | Rat | Male | [52] |
| SNL | KCNA2 | Chromosome 2 2q34 |
↓ ** | MZF1 | Regulation of Kcna2 channel expression and neuronal excitability | L4-L5 DRGs | Rat | - | [51] |
| SNL | Linc00052 | / | ↑ * | miR-448/JAK1 axis | Regulating the behaviors of neuropathic pain and processes of neuroinflammation | L4-L6 dorsal spinal cord | Rat | Male | [54] |
| SNL | Lncenc1 | Chromosome13 13 D1; 13 |
↑ ** | RBP/EZH2 axis | The activation of microglia and production of inflammatory cytokines | L4-L6 DRGs and microglia | Mice | Male | [53] |
| SNL | P21 | / | ↑ ** | MiR181b/AKT/CREB axis | Promoting apoptosis and inflammatory factors secretion | L4-L6 dorsal spinal cords and microglia | Rat | Male | [55] |
| SNL | PKIA-AS1 | / | ↑ * | CKD6 | Directly regulating the expression and function of CDK6 to maintain neuroinflammation and neuropathic pain | L5 spinal cord | Rat | Male | [38] |
| SNL | SNHG1 | / | ↑ * | CKD4 | Influencing the development of NP by regulating neuroinflammation | L5 spinal cord | Rat | Male | [56] |
| SNL | SNHG4 | Chromosome 18 18p11 |
↑ * | miR-423-5p | Regulating behaviors of neuropathic pain and neuroinflammation | L4-L6 dorsal spinal cord | Rat | Male | [57] |
| SNL | SNHG5 | Chromosome 9 9; 9 E3.1 |
↑ ** | miR-154-5p/CXCL13 Axis | SNHG5 knockdown alleviated neuropathic pain and inhibited the activation of astrocytes and microglia | L5 DRG | Mice | Male | [58] |
Notes: “?” Means possible pathway or receptor but has no studies in this article. “/” means no related location of genes were found in specific genera but it does not means absent. “-” indicates no mention in the article; * means P < 0.05, ** means P < 0.01.
Abbreviations: KCNA2-AS, KCNA2 antisense RNA; Lncenc1, LncRNA embryonic stem cells expressed 1; MZF1, myeloid zinc finger protein 1; SNHG, small nucleolar RNA host gene; EZH2, enhancer of zeste 2 polycomb repressive complex 2 subunits; CDK, cyclin dependent kinase; CXCL13, C-X-C motif chemokine ligand 13.
Besides, lncRNAs directly affects neuropathic pain as well as interacts with miRNAs’ downstream. Linc00052 is significantly expressing in the spinal cord of SNL rats. The knockout of Linc00052 can inhibit the neuroinflammatory process through the miR-448-JAK1 axis, alleviating the development of neuropathic pain.54 Studies have found that lncRNA p21 has been confirmed in vivo and in vitro experiments that it may act on miR-181b/Tnfaip1 to activate the downstream AKT/CREB axis, regulating cell apoptosis and inflammatory factor secretion, then affecting neuropathic pain.55 The small nucleolar RNA host gene (SNHG) family not only involves in the proliferation, metastasis and prognosis of tumors cells, but also affects the progression of neuropathic pain. LncRNA SNHG1 mainly induces pain by directly regulating the level of CDK4 in the spinal cord of SNL rats.56 SNHG4 regulates miR-423-5p negatively, which promotes neuro-inflammation and affects the progression of neuropathic pain.57 In addition, a significant up-regulation of SNHG5 has been revealed in the mouse SNL model. After knocking out the SNHG5 gene, the mechanical threshold and thermal threshold were significantly increasing. At the same time, the studies suggested that the activation of microglia could be inhibited by regulating the miR-154-5p/CXCL13 axis, then neuropathic pain is reduced as well.58 Such upregulation or downregulation of miRNA was highly correlated with the expression level of lncRNAs in the SNL model. As aforementioned, lncRNAs usually act as a “molecular sponge” to regulate neuroinflammation by lncRNAs–miRNAs axis. The result proven herein, affirms the indispensable role played by miRNA in development of neuropathic pain. As a result, lncRNAs is a new target for the treatment of neuroinflammatory neuropathic pain.38
CCI Model
A large number of differentially expressed lncRNAs genes related to neuropathic pain were found in the spinal cord and DRG in CCI models (Table 3). Many studies have suggested that lncRNA X–inactive specific transcript (XIST) is significantly up-regulated in the spinal cord tissue and microglia of the CCI model, and it plays a significant role by regulating downstream molecules. For one thing, the development of neuropathic pain can be affected by acting on downstream miR-154-5p, which directs the expression of TLR5.59 For another, lncRNA XIST was also reported that it could increase the expression of STAT3 (Signal transducer and activator of transcription 3) by regulating downstream miR-544, which then causes pain aggravation.60 Besides, lncRNA XIST can not only negatively regulate miR-150 and zinc-finger E-box-binding (ZEB),61 but also act on miR-137 to enhance the expression of TNFAIP1, which can promote inflammation and aggravate the development of neuropathic pain.62 It demonstrates that lncRNA XIST plays an essential role in neuropathic pain. The targeted miRNAs of lncRNAs could be predicted by bioinformatics analysis technology, and the interrelations were followed up with animal experiments, which is helpful for the prediction of therapeutic targets. lncRNA MALAT1 (metastasis-associated lung adenocarcinoma transcript 1) is one of the diagnostic markers in many diseases.63,64 Research indicated that the expression of lncRNA MALAT1 was significantly up-regulated in the spinal cord and microglia of the CCI model, and it is considered to be a prominent target for pain regulation. Neuropathic pain could be regulated by changing the expression of miR-154-5p and AQP9 by lncRNA MALAT1.65 LncRNA MALAT1 also interacts with downstream miR-129-5p to regulate the expression of HMGB1 (High-mobility group box 1), which can promote hyperalgesia.66 In addition, deletion of MALAT1 gene can alleviate the development of neuroinflammation by inhibiting expression of COX-2, IL-1β and IL-6, while promoting up-regulation of miR-206 and inhibiting overexpression of ZEB2, which can improve the progression of neuropathic pain.67 LncRNA XIST exists in a variety of biological processes as a ceRNA regulator network of miRNA-mRNA,68 and it could potentially be involved in the therapy of neuropathic pain. As can be observed, the pain networks are highly complex and interconnected. For example, the expression of HMGB1 gene can be affected by lncRNA FIRRE and NEAT1 at the same time. Besides, other significantly differentially expressed lncRNAs and the downstream pathway in the CCI model provide new ideas and targets for its treatment.
Table 3.
Differential Expression of lncRNAs in CCI Model
| Model | Name of lncRNAs | Location of the Gene | Expression and Significance | Pathway/Receptor | Biological Function | Tissue | Species | Gender | Ref |
|---|---|---|---|---|---|---|---|---|---|
| CCI | XIST | Chromosome X | ↑ ** | miR-154-5p/TLR5 axis | XIST can inhibit development of neuropathic pain | L4-L6 dorsal spinal cord | Rat | Female | [59] |
| CCI | XIST | Chromosome X | ↑ ** | miR-544/STAT3 axis | Knockdown of XIST can relieve pain hyperalgesia and suppress expression of inflammatory cytokines | L4-L6 dorsal spinal cord | Rat | Female | [60] |
| CCI | XIST | Chromosome X | ↑ * | miR-150/ZEB1 axis | Inhibition of XIST can inhibit neuropathic pain behaviors and alleviate neuroinflammation | L4-L6 dorsal spinal cord | Rat | Female | [61] |
| CCI | XIST | Chromosome X | ↑ ** | miR-137/TNFAIP1 axis | XIST silencing relieve hyperpathia by regulating the expression of miR-137 and TNFAIP1 | L4-L6 dorsal spinal cord | Rat | Female | [62] |
| CCI | MALAT1 | / | ↑ *** | miR-154-5p/AQP9 axis | MALAT1 will promote neuropathic pain progression | Spinal cord and microglia | Rat | Female | [65] |
| CCI | MALAT1 | / | ↑ ** | miR-129-5p/HMGB1 axis | MALAT1 increase the development of neuropathic pain and neuroinflammation | L4-L6 dorsal spinal cord | Rat | Female | [66] |
| CCI | MALAT1 | ↑ * | miR-206/ZEB2 axis | Loss of MALAT1 can relieve pain behaviors and depress the neuroinflammation | L4-L6 dorsal spinal cord | Rat | Female | [67] | |
| CCI | CRNDE | Chromosome19 19p11 |
↑ *** | miR-136/IL6R axis | Overexpression of CRNDE enhanced neuropathic pain behaviors and neuroinflammation | L4-L6 dorsal spinal cord | Rat | - | [69] |
| CCI | DGCR5 | / | ↓ * | miR-330-3p/PDCD4 axis | DGCR5 overexpression can alleviate neuropathic pain development | L4-L6 dorsal spinal cord | Rat | - | [70] |
| CCI | FIRRE | Chromosome X X; X A5 |
↑ * | HMGB1 | LncRNA FIRRE downregulation inhibits the secretion of microglial cells-rived inflammatory cytokines | L4-L6 spinal cord | Mice | Female | [71] |
| CCI | Gas5 | Chromosome13 13q22 |
↓ * | miR-452-5p/CELF2 axis | GAS5 overexpression will mitigate mechanical allodynia and thermal hyperalgesia | Spinal cord and microglia | Rat | Female | [72] |
| CCI | LINC00657 | / | ↑ * | miR-136/ZEB1 axis | Knockdown of LINC00657 will inhibit the neuroinflammation and alleviate hyperalgesia | L4-L6 dorsal spinal cord | Rat | Female | [73] |
| CCI | NEAT1 | / | ↑ * | miR-381/HMGB1 axis | Knockdown of NEAT1 can repress neuropathic pain behaviors and inhibiting expression of IL-6, IL-1β, and TNF-α | L4-L6 dorsal spinal cord | Rat | Female | [74] |
| CCI | DLEU1 | / | ↑ * | miR-133a-3p/SRPK1 axis | DLEU1 knockdown reduces the neuroinflammation and inhibited neuropathic pain behaviors | Dorsal spinal cord | Rat | Female | [75] |
| CCI | H19 | Chromosome 1 1q41 |
↑ ** | miR-196a-5p/CDK5 axis | Overexpression of H19 can alleviate hyperalgesia and upregulate inflammatory factors and GFAP levels, and reduced expression of 5-HT2A and GABAB2 | L4-L5 dorsal spinal cord | Rat | - | [76] |
| CCI | MRAK009713 | / | ↑ ** | P2X3 receptor | Highly expressed MRAK009713 markedly increased pain behaviors and expression of P2X3 | DRG | Rat | Male | [77] |
| CCI | SNHG16 | / | ↑ *** | miR-124-3p and miR-141-3p/JAG1 axis | SNHG16 could increase the mechanical allodynia and thermal hyperalgesia and mRNA expression of inflammatory factors | Dorsal spinal cord | Rat | Female | [78] |
| CCI | uc.153 | / | ↑ *** | miR-182-5p/EphB1-NMDA axis | uc.153 will promote pain behaviors and neuronal sensitization | L4-L5 dorsal spinal cord | Mice | Male | [79] |
| CCI | CRNDE | Chromosome19 19p11 |
↑ *** | miR-146a-5p/WNT5A | Silencing of CRNDE relieved mechanical allodynia, thermal hyperalgesia, and neuroinflammation | Dorsal spinal cord | Rat | Male | [80] |
| CCI | MEG3 | / | ↑ *** | miR-130a-5p/CXCL12/CXCR4 | Up-regulating MEG3 aggravated NP, enhanced inflammatory cytokines expression and release. | L4-L6 dorsal spinal cord | Rat | Female | [80] |
| CCI | PCAT19 | / | ↑ *** | miR-182-5p/JMJD1A | Mechanical pain and thermal hyperalgesia as well as neuroinflammation can be reduced by knocking down PCAT19 | L4-L6 dorsal spinal cord | Rat | Male | [80] |
Notes: “/” Means no related location of genes were found in specific genera but it does not means absent. “-” indicates no mention in the article; * means P < 0.05, ** means P < 0.01, and *** means P < 0.001.
Abbreviations: DGCR5, DiGeorge syndrome critical region gene 5; NEAT1, Nuclear paraspeckle assembly transcript1; XIST, X inactivation-specific transcript; DLEU1, deleted in lymphocytic leukemia 1; MALAT1, metastasis-associated lung adenocarcinoma transcript 1; SNHG, small nucleolar RNA host gene; CRNDE, Colorectal neoplasia differentially expressed; FIRRE, functional intergenic repeating RNA element; Gas5, growth arrest-specific transcript 5. MEG3, maternally expressed 3; PCAT19, prostate cancer associated transcript 19; TLR5, toll like receptor 5; STAT3, signal transducer and activator of transcription 3; TNFAIP1, TNF alpha induced protein 1; AQP9, aquaporin 9; HMGB1, high mobility group box 1; ZEB2L, zinc finger E-box binding homeobox 2; PDCD4, programmed cell death 4; CELF2, CUGBP Elav-like family member 2; SRPK1, SRSF protein kinase 1; CDK5, cyclin dependent kinase 5; WNT5A, Wnt family member 5A; CXCL12, C-X-C motif chemokine ligand 12; CXCR4, C-X-C motif chemokine receptor 4; JMJD1A, lysine demethylase 3A.
DNP Model
The dominating manifestation of DNP model, common complications of type 2 diabetes, is accompanied by a burning sensation, acupuncture sensation and other abnormal sensations.81,82 The pathogenesis of diabetic neuropathic pain is complicated.82 Most recently, lncRNAs have attracted attention due to the continuous research on the mechanism of diabetic peripheral nerve pain. Related differentially expressed lncRNAs genes were parading in the DNP rat model (Table 4). Liu et al showed that inhibition of lncRNA BC168687 could reverse the threshold of mechanical pain and thermal pain in DNP rats. It mainly interferes with the expression of TRPV1 in DRG and inhibits phosphorylation (P)-ERK and p-p38 signaling pathways at the same time;83 They also found that lncRNA BC168687 acts on the P2X7 receptor in DRG, which relieves neuropathic pain by inhibiting its expression.84 In addition, lncRNA NONRATT021972 is a gene that plays a pivotal role in DNP. After its expression was inhibited, in the DNP model, results showed that it could interact with the P2X3 and P2X7 receptors in DRG to alleviate the development of peripheral pain complications in diabetes.85,86 Increased expression of lncRNA uc.48+ was also demonstrated in DNP rat DRG and serum of diabetic patients, which can up-regulate the P2X3 receptor of DRG to regulate the excitability of neurons to affect the development of pain.87 Besides, Xiong et al found that lncRNA uc.48+ could promote the release of calcitonin gene-related peptide (CGRP) in the spinal cord. When the expression of lncRNA uc.48+ is down-regulated, it can inhibit the phosphorylation of downstream p38 and ERK1/2, which will reduce the release of IL-1β and TNF-α as well as alleviate hyperalgesia in DNP rats.88 It is known that diabetic neuropathy is one of the diabetic complications and the patients are concomitant with maintaining a low quality of life. Now this illustrates neuroinflammation and microglia activation are critical in origination and development of DNP. Therefore, these related lncRNAs may become noteworthy targets for the treatment of DNP in the future.
Table 4.
Differential Expression of lncRNAs in CCI Model. GENE
| Model | Name of lncRNAs | Expression and Significance | Pathway/Receptor | Biological Function | Tissue | Species | Gender | Ref. |
|---|---|---|---|---|---|---|---|---|
| DNP | BC168687 | ↑ - | TRPV1 | It could increase diabetic neuropathic pain by acting TRPV1 receptor | DRG | Rat | Male | [83] |
| DNP | BC168687 | ↑ ** | P2X7 | BC168687 could increase the release of NO and the activation of SGCs, thus regulate the excitability of DRG neurons and the pain behaviors | DRG | Rat | Male | [84] |
| DNP | NONRATT021972 | ↑ ** | P2X3 | It could aggravate thermal and mechanical hyperalgesia vis activating P2X3 receptor of ERK1/2 | DRG | Rat | Male | [85] |
| DNP | NONRATT021972 | ↑ *** | P2X7 | It can increase the expression levels of P2X7 and activation of satellite glial cells in DRG | DRG | Rat | - | [86] |
| DNP | uc.48+ | ↑ ** | P2X3 | uc.48+ may regulate the excitatory of primary sensory neurons by the P2X3 receptor | DRG | Rat | Male | [87] |
| DNP | uc.48+ | ↑ ** | p38 and ERK1/2 | lncRNA uc.48+ may regulate DNP by promoting the release of CGRP | L4-L5 spinal cords | Rat | Male | [88] |
Notes: ** Means P < 0.01, and *** means P < 0.001. “-” indicates no mention in the article.
Abbreviation: TRPV1, transient receptor potential cation channel subfamily V member 1.
Other Neuropathic Pain Models
Moreover, lncRNAs are found to be actively involved in other models (Table 5). Trigeminal neuralgia (TN) is also a prevalent neuropathic pain in clinical practice. Cai et al found that lncRNA Gm14461 is related to the activation and autophagy of astrocytes in a mouse model of trigeminal neuralgia. When the lncRNA was silencing, loss of Gm14461 increased autophagy in astrocytes and played an analgesic effect by regulating AMPK and Akt/mTOR signaling pathways.89 LncRNA Gm14461 is also a factor to regulate CGRP (calcitonin gene-related peptide) and P2X3/7 receptors, which affect the release of inflammatory mediators and pain.90 Similar to the role of lncRNA uc.48+ in the DNP model, in the TN model, lncRNA uc.48+ will up-regulate the expression of P2X7 receptors and enhance the phosphorylation of ERK1/2 in the trigeminal nerve meridian, which will heighten pain sensitization.91 In the rat brachial plexus injury (BPI) model, the expression of lncRNA Malat1 decreases in the spinal cord, which may affect the excitability of neurons by regulating the level of calcium ions.92 In addition, up-regulation of JHDM1D-AS1 can reduce the inflammation and neuronal damage of the spinal cord after brachial plexus injury by targeting miR-101-3p-DUSP.93 In the rat model of cancer pain, the expression of lncRNA-NONRATT021203.2 was increased, which can participate in the production of pain by targeting CXCL9 (C-X-C motif chemokine ligand 9).94 Postherpetic neuralgia (PHN) is also a considerable neuropathic pain. It confirmed that KCNA2-AS is highly expressing in the spinal cord of the PHN (postherpetic neuralgia) rat model. The pain is relieved when KCNA2-AS was knocking out. The possible reason is that it reduces the translocation of pSTAT3 to the nucleus and inhibits the activation of astrocytes.95 However, lncRNA NEAT1 can target the expression of AQP4 by inhibiting miR-128-3p in the rat SCI model, which affects the development of neuropathic pain.96 SNI (spare nerve injury) model as a common model in neuropathic pain research. But there are few reliable studies of lncRNAs in this model. Silencing of LINC01119 can alleviate the hypersensitivity and reduced the increase in IL−6, IL−1β, and TNF−α via binding with ELAVL1 and increasing BDNF mRNA stability and expression level. Here, we briefly reviewed recent studies of lncRNAs in neuropathic pain, focusing on different rodent neuropathic pain models. Many expressions of lncRNAs are altered in rodent rats after the establishment of chronic pain. Therefore, lncRNAs might be a good potential therapeutic target for neuropathic pain.
Table 5.
Differential Expression of lncRNAs in Other Models
| Model | Name of lncRNAs | Location of the Gene | Expression and Significance | Pathway/Receptor | Biological Function | Tissue | Species | Gender | Ref. |
|---|---|---|---|---|---|---|---|---|---|
| TN | Gm14461 | Chromosome2 2 C3; 2 |
↑ ** | AMPK and Akt/mTOR signaling | Gm14461 silencing relieves mechanical withdrawal threshold and inhibits astrocyte activation and enhances autophagy via regulating of the AMPK and Akt/mTOR signaling | Trigeminal ganglia | Mice | - | [89] |
| TN | Gm14461 | Chromosome2 2 C3; 2 |
↑ * | CGRP and P2X3/7 receptor | Gm14461 overexpression decreases MWT and upregulates mRNA levels of proinflammatory cytokine and protein levels of CGRP and P2X3/7 receptor | Trigeminal ganglia | Mice | - | [90] |
| TN | uc.48+ | / | ↑ ** | P2X7/ERK1/2 | uc.48+ can activate the P2X7 receptor to regulate the MWT | Trigeminal ganglia | Rat | Male | [91] |
| BPI | Malat1 | / | ↓ * | Unknown | Down-regulation of Malat1 increases the frequency of spontaneous electric activity via changing of intracellular calcium concentration in neurons but have no effect on neuronal apoptosis. | C5-T1 spinal cord | Rat | Male | [92] |
| CIP | NONRATT 021203.2 |
/ | ↑ * | CXCL9 | LncRNA-NONRATT021203.2 could target CXCL9 to regulate hyperalgesia | DRG | Rat | Female | [94] |
| PHN | KCNA2 | Chromosome 2 2q34 |
↓ ** | STAT3 | KCNA2-AS relieved mechanical allodynia partly by combining with pSTAT3 to regulate cytoplasmic/nuclear translocation. | Spinal cord | Rat | Female | [95] |
| SCI | NEAT1 | / | ↑ *** | miR-128-3p/AQP4 axis | Over-expression of NEAT1 enhance the expression of inflammatory factors (IL-6, IL-1β, and TNFα) | Spinal cord | Rat | Female | [96] |
| SCI | PVT1 | Chromosome 6 7q33 |
↑ *** | miR‑186‑5p/CXCL13/CXCR5 Axis | PVT1 depletion could alleviated pain behaviors, astrocytic activation and reduced the expression of neuroinflammatory cytokine | T10 spinal cord | Rat | Male | [97] |
| BPI | JHDM1D-AS1 | Chromosome 6 4q22 |
↓ *** | miR-101-3p/DUSP1 axis | JHDM1D-AS1 have a neuroprotective effect by suppressing neuronal apoptosis and microglial activation | Spinal cord | Rat | - | [93] |
| SNI | LINC01119 | Chromosome 6 2p21 |
↑ ** | ELAVL1/BDNF | Silencing of LINC01119 can alleviate the hypersensitivity and reduced the increase in IL−6, IL−1β, and TNF−a via binding with ELAVL1 and increasing BDNF mRNA stability and expression level. | L4–L5spinal cord; serum | Rat; Human | Male | [98] |
Notes: “/” Means no related location of genes were found in specific genera but it does not means absent. * means P < 0.05, ** means P < 0.01, and *** means P < 0.001; “-” indicates no mention in the article.
Abbreviations: TN, trigeminal neuralgia; BPI, brachial plexus injury; PHN, post-herpetic neuralgia; SCI, spinal cord injury; PHN, postherpetic neuralgia; CIP, Cancer-induced pain; MWT, mechanical withdrawal threshold; SNI, spare nerve injury; MALAT1, metastasis-associated lung adenocarcinoma transcript 1; PVT1, plasmacytoma variant translocation 1; KCNA2-AS, KCNA2 antisense RNA; NEAT1, Nuclear paraspeckle assembly transcript 1; JHDM1D-AS1, JHDM1D antisense 1; Mtor, mechanistic target of rapamycin kinase; STAT3, signal transducer and activator of transcription 3; DUSP1, dual specificity phosphatase 1; ELAVL1, ELAV like RNA binding protein 1; BDNF, brain derived neurotrophic factor.
Discussion and Future Perspectives
Chronic neuropathic pain is a severe situation, accompanying by injury and dysfunction of the somatosensory nervous system.6 However, at present, a lack of efficient curative treatments remains a severe challenge for neuropathic pain. It proved that the rodent models of chronic neuropathic pain play an essential role in parent studies. And detailed inter-molecular interactions such as lncRNAs and miRNAs were revealed, which could provide therapeutic promise for the treatment of chronic neuropathic pain.
With the advance in sequencing technology, the studies of lncRNAs in have become a hotspot of the filed. It has been demonstrated that played an essential role in neuropathic pain. Therefore, lncRNAs are likely expected to become a potential target for neuropathic pain. We wonder if these sequencing results of lncRNAs are reliable. For instance, even within the same model there are major differences among sequencing studies published in the literature. The sequencing technology should exhibit excellent sensitivity, repeatability, and stability. In other words, the current study confirms that lncRNAs lack specificity in different models. Perhaps due to complex influencing factors in the system, a few discrepancies are evident in the sequencing results.
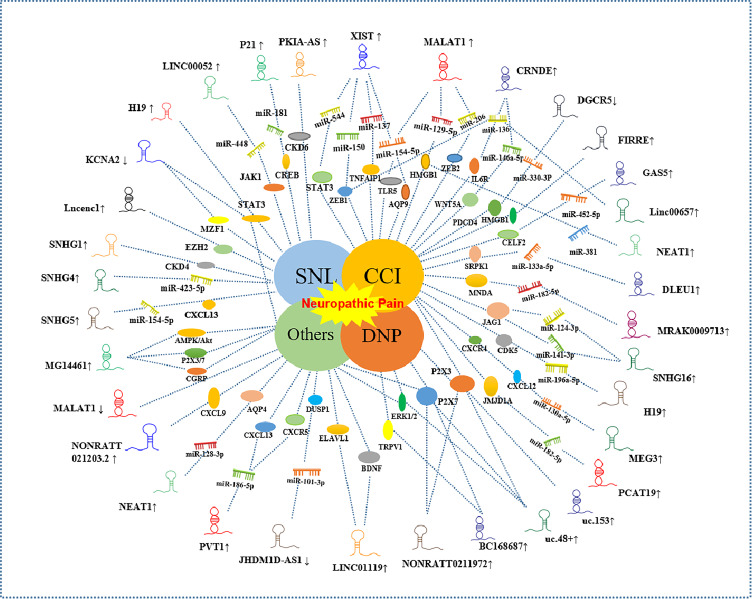
As is well known, lncRNAs do not participate in protein-coding directly. However, it plays an imperative part in co-transcriptional regulation, post-transcriptional and Post-translational protein modifications and so on.20 Related research found that the potential regulatory effect of lncRNAs is very distinguished in various diseases. In the above experiments of different neuropathic pain animal models, but also present a significant role in regulating the generation as well as the progression of neuropathic pain. The previous focus was mainly on the changes in the excitability of central and peripheral sensory neurons and the interaction of surrounding microglia, macrophages and other immune cells in the research of neuropathic pain, which led to the persistence of pain signals. What’s more, we have shown in this review that the interactions of the lncRNAs, miRNA and proteins are very complicated. As such, the studies of those non-coding RNAs can be used to understand better the mechanisms of molecular interaction. Current works of lncRNAs have been shown in Figure 2. However, there is limited research focused on clinical prospective studies of lncRNAs in neuropathic pain. Because of neurocytes are nonrenewable, human specimens are generally available. Currently, several researches are focusing attention on the animal and cell studies.
Figure 2.
Differentially expressed lncRNA in differential neuropathic pain models. ↑: up-regulation; ↓: down-regulation. Differential shapes of lncRNAs denote gene symbol without biological meaning. Center works represent different models of neuropathic pain.
Gender differences of immune cells were found among the neuropathic pain. The magnitude of the injury-induced macrophage expansion was different between the female and male mice.16 Microglia also showed sex differences. It is critical to the development of the neuropathic pain in male mice but not in female.99 No previous studies have, however, investigated the differentially expressed lncRNAs in neuropathic pain models of different sex. The exact role of lncRNAs in mediating hypersensitivity in sexually dimorphic is still to be defined in the in the subsequent studies.
In conclusion, current results of lncRNAs in neuropathic pain indicate the potential of using sequencing technology. Despite the current researches lacking clinical implications and specificity of lncRNAs in different models, sufficient evidence has been presented to prove that lncRNAs have critical roles in the development of neuropathic pain. Therefore, additional studies are required to look for potential specific biomarkers and treatment in neuropathic pain from basic research to clinical applications.
Acknowledgments
The authors are grateful for support from the Qingdao University Medical Research Center.
The authors thank Citexs (www.citexs.com) for English language editing.
This work was supported by Youth Research Fund of Affiliated Hospital of Qingdao University (for YZ).
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Disclosure
The authors have no conflicts of interest to declare.
References
- 1.Inoue K, Tsuda M. Microglia in neuropathic pain: cellular and molecular mechanisms and therapeutic potential. Nat Rev Neurosci. 2018;19(3):138–152. doi: 10.1038/nrn.2018.2 [DOI] [PubMed] [Google Scholar]
- 2.Ochoa JL. Neuropathic pain: redefinition and a grading system for clinical and research purposes. Neurology. 2009;72(14):1282–1283. doi: 10.1212/01.wnl.0000346325.50431.5f [DOI] [PubMed] [Google Scholar]
- 3.Scholz J, Finnerup NB, Attal N, et al. The IASP classification of chronic pain for ICD-11: chronic neuropathic pain. Pain. 2019;160(1):53–59. doi: 10.1097/j.pain.0000000000001365 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Borzan J, Meyer RA. Neuropathic Pain. Encycl Neurosci. 2009;Imi:749–757. [Google Scholar]
- 5.Andersen ML, Araujo P, Frange C, Tufik S. Sleep disturbance and pain: a tale of two common problems. Chest. 2018;154(5):1249–1259. doi: 10.1016/j.chest.2018.07.019 [DOI] [PubMed] [Google Scholar]
- 6.Gilron I, Baron R, Jensen T. Neuropathic pain: principles of diagnosis and treatment. Mayo Clin Proc. 2015;90(4):532–545. doi: 10.1016/j.mayocp.2015.01.018 [DOI] [PubMed] [Google Scholar]
- 7.Li Y, North RY, Rhines LD, et al. DRG voltage-gated sodium channel 1.7 is upregulated in paclitaxel-induced neuropathy in rats and in humans with neuropathic pain. J Neurosci. 2018;38(5):1124–1136. doi: 10.1523/JNEUROSCI.0899-17.2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Li N, Liu B, Wu W, et al. Upregulation of transcription factor 4 downregulates NaV1.8 expression in DRG neurons and prevents the development of rat inflammatory and neuropathic hypersensitivity. Exp Neurol. 2020;327:113240. doi: 10.1016/j.expneurol.2020.113240 [DOI] [PubMed] [Google Scholar]
- 9.Guan Z, Kuhn JA, Wang X, et al. Injured sensory neuron-derived CSF1 induces microglial proliferation and DAP12-dependent pain. Nat Neurosci. 2015;19(1):94–101. doi: 10.1038/nn.4189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Davoli-Ferreira M, De Lima KA, Fonseca MM, et al. Regulatory T cells counteract neuropathic pain through inhibition of the Th1 response at the site of peripheral nerve injury. Pain. 2020;161(8):1730–1743. doi: 10.1097/j.pain.0000000000001879 [DOI] [PubMed] [Google Scholar]
- 11.Hogan QH. Labat lecture: the primary sensory neuron: where it is, what it does, and why it matters. Reg Anesth Pain Med. 2010;35(3):306–311. doi: 10.1097/AAP.0b013e3181d2375e [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Taylor BK. Pathophysiologic mechanisms of neuropathic pain. Curr Pain Headache Rep. 2001;5(2):151–161. doi: 10.1007/s11916-001-0083-1 [DOI] [PubMed] [Google Scholar]
- 13.Sommer C, Leinders M, Üçeyler N. Inflammation in the pathophysiology of neuropathic pain. Pain. 2018;159(3):595–602. doi: 10.1097/j.pain.0000000000001122 [DOI] [PubMed] [Google Scholar]
- 14.Scholz J, Woolf CJ. The neuropathic pain triad: neurons, immune cells and glia. Nat Neurosci. 2007;10(11):1361–1368. doi: 10.1038/nn1992 [DOI] [PubMed] [Google Scholar]
- 15.Tozaki-Saitoh H, Tsuda M. Microglia-neuron interactions in the models of neuropathic pain. Biochem Pharmacol. 2019;169:113614. doi: 10.1016/j.bcp.2019.08.016 [DOI] [PubMed] [Google Scholar]
- 16.Yu X, Liu H, Hamel KA, et al. Dorsal root ganglion macrophages contribute to both the initiation and persistence of neuropathic pain. Nat Commun. 2020;11(1):1–12. doi: 10.1038/s41467-019-13993-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yu X, Leff J, Guan Z. Rapid isolation of dorsal root ganglion macrophages. J Vis Exp. 2019;2019(151):2–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhao Y, Xin Y, Chu H. MC4R is involved in neuropathic pain by regulating JNK signaling pathway after chronic constriction injury. Front Neurosci. 2019;13:1–11. doi: 10.3389/fnins.2019.00919 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bruno K, Woller SA, Miller YI, et al. Targeting toll-like receptor-4 (TLR4)-an emerging therapeutic target for persistent pain states. Pain. 2018;159(10):1908–1915. doi: 10.1097/j.pain.0000000000001306 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bali KK, Kuner R. Noncoding RNAs: key molecules in understanding and treating pain. Trends Mol Med. 2014;20(8):437–448. doi: 10.1016/j.molmed.2014.05.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Nandwani A, Rathore S, Datta M. LncRNAs in cancer: regulatory and therapeutic implications. Cancer Lett. 2021;501:162–171. doi: 10.1016/j.canlet.2020.11.048 [DOI] [PubMed] [Google Scholar]
- 22.Wu GC, Pan HF, Leng RX, et al. Emerging role of long noncoding RNAs in autoimmune diseases. Autoimmun Rev. 2015;14(9):798–805. doi: 10.1016/j.autrev.2015.05.004 [DOI] [PubMed] [Google Scholar]
- 23.Lorenzen JM, Thum T. Long noncoding RNAs in kidney and cardiovascular diseases. Nat Rev Nephrol. 2016;12(6):360–373. doi: 10.1038/nrneph.2016.51 [DOI] [PubMed] [Google Scholar]
- 24.Ulitsky I, Bartel DP. XLincRNAs: genomics, evolution, and mechanisms. Cell. 2013;154(1):26. doi: 10.1016/j.cell.2013.06.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li Z, Li X, Chen X, et al. Emerging roles of long non-coding RNAs in neuropathic pain. Cell Prolif. 2019;52(1):1–7. doi: 10.1111/cpr.12528 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Peng WX, Koirala P, Mo YY. LncRNA-mediated regulation of cell signaling in cancer. Oncogene. 2017;36(41):5661–5667. doi: 10.1038/onc.2017.184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Paraskevopoulou MD, Hatzigeorgiou AG. Analyzing MiRNA – lncRNA interactions. Methods Mol Biol. 2016;1402:271–286. [DOI] [PubMed] [Google Scholar]
- 28.Ferrè F, Colantoni A, Helmer-Citterich M. Revealing protein-lncRNA interaction. Brief Bioinform. 2016;17(1):106–116. doi: 10.1093/bib/bbv031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mayr C. Regulation by 3'-Untranslated Regions. Annu Rev Genet. 2017;51:171-194. doi: 10.1146/annurev-genet-120116-024704 [DOI] [PubMed] [Google Scholar]
- 30.Qi X, Zhang DH, Wu N, Xiao JH, Wang X, Ma W. ceRNA in cancer: possible functions and clinical implications. J Med Genet. 2015;52(10):710–718. doi: 10.1136/jmedgenet-2015-103334 [DOI] [PubMed] [Google Scholar]
- 31.Thomson DW, Dinger ME. Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet. 2016;17(5):272–283. doi: 10.1038/nrg.2016.20 [DOI] [PubMed] [Google Scholar]
- 32.Prasad A, Sharma N, Prasad M. Noncoding but coding: pri-miRNA into the action. Trends Plant Sci. 2021;26(3):204–206. doi: 10.1016/j.tplants.2020.12.004 [DOI] [PubMed] [Google Scholar]
- 33.Geisler S, Coller J. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol. 2013;14(11):699–712. doi: 10.1038/nrm3679 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wu W, Ji X, Zhao Y. Emerging roles of long non-coding RNAs in chronic neuropathic pain. Front Neurosci. 2019;13. doi: 10.3389/fnins.2019.01097 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wu S, Bono J, Tao YX. Long noncoding RNA (lncRNA): a target in neuropathic pain. Expert Opin Ther Targets. 2019;23(1):15–20. doi: 10.1080/14728222.2019.1550075 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yu B, Zhou S, Hu W, et al. Altered long noncoding RNA expressions in dorsal root ganglion after rat sciatic nerve injury. Neurosci Lett. 2013;534(1):117–122. doi: 10.1016/j.neulet.2012.12.014 [DOI] [PubMed] [Google Scholar]
- 37.Tang S, Jing H, Huang Z, et al. Identification of key candidate genes in neuropathic pain by integrated bioinformatic analysis. J Cell Biochem. 2020;121(2):1635–1648. doi: 10.1002/jcb.29398 [DOI] [PubMed] [Google Scholar]
- 38.Hu JZ, Rong ZJ, Li M, et al. Silencing of incRNA PKIA-AS1 attenuates spinal nerve ligation-induced neuropathic pain through epigenetic downregulation of CDK6 expression. Front Cell Neurosci. 2019;13:1–12. doi: 10.3389/fncel.2019.00050 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Cao S, Yuan J, Zhang D, et al. Transcriptome changes in dorsal spinal cord of rats with neuropathic pain. J Pain Res. 2019;12:3013–3023. doi: 10.2147/JPR.S219084 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhou J, Fan Y, Chen H. Analyses of long non-coding RNA and mRNA profiles in the spinal cord of rats using RNA sequencing during the progression of neuropathic pain in an SNI model. RNA Biol. 2017;14(12):1810–1826. doi: 10.1080/15476286.2017.1371400 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Mao P, Li CR, Zhang SZ, Zhang Y, Liu BT, Fan BF. Transcriptomic differential lncRNA expression is involved in neuropathic pain in rat dorsal root ganglion after spared sciatic nerve injury. Braz J Med Biol Res. 2018;51(10):16–21. doi: 10.1590/1414-431x20187113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Jiang BC, Sun WX, He LN, Cao DL, Zhang ZJ, Gao YJ. Identification of lncRNA expression profile in the spinal cord of mice following spinal nerve ligation-induced neuropathic pain. Mol Pain. 2015;11(1):1–18. doi: 10.1186/s12990-015-0047-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Liu Z, Liang Y, Wang H, et al. LncRNA expression in the spinal cord modulated by minocycline in a mouse model of spared nerve injury. J Pain Res. 2017;10:2503–2514. doi: 10.2147/JPR.S147055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhou J, Xiong Q, Chen H, Yang C, Fan Y. Identification of the spinal expression profile of non-coding RNAs involved in neuropathic pain following spared nerve injury by sequence analysis. Front Mol Neurosci. 2017;10:1–22. doi: 10.3389/fnmol.2017.00091 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wu S, Marie Lutz B, Miao X, et al. Dorsal root ganglion transcriptome analysis following peripheral nerve injury in mice. Mol Pain. 2016;12:1–14. doi: 10.1177/1744806916629048 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.He J, Wang H, Huang J, et al. Diabetic neuropathic pain induced by streptozotocin alters the expression profile of non‑coding RNAs in the spinal cord of mice as determined by sequencing analysis. Exp Ther Med. 2021;22(1):1–11. doi: 10.3892/etm.2021.10207 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Li Y, Yin C, Liu B, et al. Transcriptome profiling of long noncoding RNAs and mRNAs in spinal cord of a rat model of paclitaxel-induced peripheral neuropathy identifies potential mechanisms mediating neuroinflammation and pain. J Neuroinflammation. 2021;18(1):1–22. doi: 10.1186/s12974-020-02040-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wang LX, Wang ZJ. Animal and cellular models of chronic pain. Adv Drug Deliv Rev. 2003;55(8):949–965. doi: 10.1016/S0169-409X(03)00098-X [DOI] [PubMed] [Google Scholar]
- 49.Jaggi AS, Jain V, Singh N. Animal models of neuropathic pain. Fundam Clin Pharmacol. 2011;25(1):1–28. doi: 10.1111/j.1472-8206.2009.00801.x [DOI] [PubMed] [Google Scholar]
- 50.Cichon J, Sun L, Yang G. Spared nerve injury model of neuropathic pain in mice. Bio-Protocol. 2018;8(6). doi: 10.21769/BioProtoc.2777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Zhao X, Tang Z, Zhang H, et al. A long noncoding RNA contributes to neuropathic pain by silencing Kcna2 in primary afferent neurons. Nat Neurosci. 2013;16(8):1024–1031. doi: 10.1038/nn.3438 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Iwasaki H, Sakai A, Maruyama M, Ito T, Sakamoto A, Suzuki H. Increased H19 long non-coding RNA expression in Schwann cells in peripheral neuropathic pain. J Nippon Med Sch. 2019;86(4):215–221. doi: 10.1272/jnms.JNMS.2018_86-402 [DOI] [PubMed] [Google Scholar]
- 53.Zhang Z, Sun X, Zhao G, Ma Y, Zeng G. LncRNA embryonic stem cells expressed 1 (Lncenc1) is identified as a novel regulator in neuropathic pain by interacting with EZH2 and downregulating the expression of Bai1 in mouse microglia. Exp Cell Res. 2021;399(1):112435. doi: 10.1016/j.yexcr.2020.112435 [DOI] [PubMed] [Google Scholar]
- 54.Wang L, Zhu K, Yang B, Cai Y. Knockdown of Linc00052 alleviated spinal nerve ligation-triggered neuropathic pain through regulating miR-448 and JAK1. J Cell Physiol. 2020;235(10):6528–6535. doi: 10.1002/jcp.29465 [DOI] [PubMed] [Google Scholar]
- 55.Liu Y, Sun H, Sun Y. LncRNA p21, downregulating miR-181b, aggravates neuropathic pain by upregulating Tnfaip1 and inhibit the AKT/CREB axis. Brain Res Bull. 2021;171:150–161. doi: 10.1016/j.brainresbull.2021.03.005 [DOI] [PubMed] [Google Scholar]
- 56.Zhang JY, Lv DB, Su YN, et al. LncRNA SNHG1 attenuates neuropathic pain following spinal cord injury by regulating CDK4 level. Eur Rev Med Pharmacol Sci. 2020;24(23):12034–12040. doi: 10.26355/eurrev_202012_23992 [DOI] [PubMed] [Google Scholar]
- 57.Pan X, Shen C, Huang Y, Wang L, Xia Z. Erratum: Loss of SNHG4 attenuated spinal nerve ligation-triggered neuropathic pain through sponging miR-423-5p. Mediators Inflamm. 2020;2020:2094948. Mediators Inflamm. 2020;2020. doi: 10.1155/2020/2094948 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chen M, Yang Y, Zhang W, et al. Long noncoding RNA SNHG5 knockdown alleviates neuropathic pain by targeting the miR-154-5p/CXCL13 axis. Neurochem Res. 2020;45(7):1566–1575. doi: 10.1007/s11064-020-03021-2 [DOI] [PubMed] [Google Scholar]
- 59.Wei M, Li L, Zhang Y, Zhang ZJ, Liu HL, Bao HG. LncRNA X inactive specific transcript contributes to neuropathic pain development by sponging miR-154-5p via inducing toll-like receptor 5 in CCI rat models. J Cell Biochem. 2019;120(2):1271–1281. doi: 10.1002/jcb.27088 [DOI] [PubMed] [Google Scholar]
- 60.Jin H, Du XJ, Zhao Y, Xia DL. XIST/miR-544 axis induces neuropathic pain by activating STAT3 in a rat model. J Cell Physiol. 2018;233(8):5847–5855. doi: 10.1002/jcp.26376 [DOI] [PubMed] [Google Scholar]
- 61.Yan XT, Lu JM, Wang Y, et al. XIST accelerates neuropathic pain progression through regulation of miR-150 and ZEB1 in CCI rat models. J Cell Physiol. 2018;233(8):6098–6106. doi: 10.1002/jcp.26453 [DOI] [PubMed] [Google Scholar]
- 62.Zhao Y, Li S, Xia N, Shi Y, Zhao CM. Effects of XIST/miR-137 axis on neuropathic pain by targeting TNFAIP1 in a rat model. J Cell Physiol. 2018;233(5):4307–4316. doi: 10.1002/jcp.26254 [DOI] [PubMed] [Google Scholar]
- 63.Tian X, Xu G. Clinical value of lncRNA MALAT1 as a prognostic marker in human cancer: systematic review and meta-analysis. BMJ Open. 2015;5(9):e008653. doi: 10.1136/bmjopen-2015-008653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Zhao Y, Yu YQ, You S, et al. Long non-coding RNA MALAT1 as a detection and diagnostic molecular marker in various human cancers: a pooled analysis based on 3255 subjects. Onco Targets Ther. 2020;13:5807–5817. doi: 10.2147/OTT.S250796 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Wu J, Wang C, Ding H. LncRNA MALAT1 promotes neuropathic pain progression through the miR‑154‑5p/AQP9 axis in CCI rat models. Mol Med Rep. 2020;21(1):291–303. doi: 10.3892/mmr.2019.10829 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Ma X, Wang H, Song T, Wang W, Zhang Z. lncRNA MALAT1 contributes to neuropathic pain development through regulating miR-129-5p/HMGB1 axis in a rat model of chronic constriction injury. Int J Neurosci. 2020;130(12):1215–1224. doi: 10.1080/00207454.2020.1731508 [DOI] [PubMed] [Google Scholar]
- 67.Chen ZL, Liu JY, Wang F, Jing X. Suppression of MALAT1 ameliorates chronic constriction injury-induced neuropathic pain in rats via modulating miR-206 and ZEB2. J Cell Physiol. 2019;234(9):15647–15653. doi: 10.1002/jcp.28213 [DOI] [PubMed] [Google Scholar]
- 68.Wang W, Min L, Qiu X, et al. Biological function of long non-coding RNA (LncRNA) Xist. Front Cell Dev Biol. 2021;9:1–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Zhang D, Mou JY, Wang F, Liu J, Hu X. CRNDE enhances neuropathic pain via modulating miR-136/IL6R axis in CCI rat models. J Cell Physiol. 2019;234(12):22234–22241. doi: 10.1002/jcp.28790 [DOI] [PubMed] [Google Scholar]
- 70.Peng C, Zhang C, Su Z, Lin D. DGCR5 attenuates neuropathic pain through sponging miR-330-3p and regulating PDCD4 in CCI rat models. J Cell Physiol. 2019;234(5):7292–7300. doi: 10.1002/jcp.27487 [DOI] [PubMed] [Google Scholar]
- 71.Wen Y, Fan X, Bu H, et al. Downregulation of lncRNA FIRRE relieved the neuropathic pain of female mice by suppressing HMGB1 expression. Mol Cell Biochem. 2021;476(2):841–852. doi: 10.1007/s11010-020-03949-7 [DOI] [PubMed] [Google Scholar]
- 72.Tian Y, Sun L, Qi T. Long noncoding RNA gas5 ameliorates chronic constriction injury induced neuropathic pain in rats by modulation of the mir-452-5p/celf2 axis. Can J Physiol Pharmacol. 2020;98(12):870–877. doi: 10.1139/cjpp-2020-0036 [DOI] [PubMed] [Google Scholar]
- 73.Shen F, Zheng H, Zhou L, Li W, Zhang Y, Xu X. LINC00657 expedites neuropathic pain development by modulating miR-136/ZEB1 axis in a rat model. J Cell Biochem. 2019;120(1):1000–1010. doi: 10.1002/jcb.27466 [DOI] [PubMed] [Google Scholar]
- 74.Xia LX, Ke C, Lu JM. NEAT1 contributes to neuropathic pain development through targeting miR-381/HMGB1 axis in CCI rat models. J Cell Physiol. 2018;233(9):7103–7111. doi: 10.1002/jcp.26526 [DOI] [PubMed] [Google Scholar]
- 75.Li Z, Li A, Yan L, Yang T, Xu W, Fan P. Downregulation of long noncoding RNA DLEU1 attenuates hypersensitivity in chronic constriction injury-induced neuropathic pain in rats by targeting miR-133a-3p/SRPK1 axis. Mol Med. 2020;26(1). doi: 10.1186/s10020-020-00235-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Li K, Jiao Y, Ren X, You D, Cao R. Long noncoding RNA h19 induces neuropathic pain by upregulating cyclin-dependent kinase 5-mediated phosphorylation of camp response element binding protein. J Pain Res. 2020;13:2113–2124. doi: 10.2147/JPR.S240273 [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 77.Li G, Jiang H, Zheng C, et al. Long noncoding RNA MRAK009713 is a novel regulator of neuropathic pain in rats. Pain. 2017;158(10):2042–2052. doi: 10.1097/j.pain.0000000000001013 [DOI] [PubMed] [Google Scholar]
- 78.Li H, Fan L, Zhang Y, Cao Y, Liu X. SNHG16 aggravates chronic constriction injury-induced neuropathic pain in rats via binding with miR-124-3p and miR-141-3p to upregulate JAG1. Brain Res Bull. 2020;165:228–237. doi: 10.1016/j.brainresbull.2020.09.025 [DOI] [PubMed] [Google Scholar]
- 79.Zhang C, Peng Y, Wang Y, Xu H, Zhou X. Transcribed ultraconserved noncoding RNA uc.153 is a new player in neuropathic pain. Pain. 2020;161(8):1744–1754. doi: 10.1097/j.pain.0000000000001868 [DOI] [PubMed] [Google Scholar]
- 80.Zhang Q, Zhu D, Li Q. LncRNA CRNDE exacerbates neuropathic pain in chronic constriction injury-induced(CCI) rats through regulating miR-146a-5p/WNT5A pathway. Bioengineered. 2021;12(1):7348–7359. doi: 10.1080/21655979.2021.1972901 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Callaghan BC, Cheng HT, Stables CL, Smith AL, Feldman EL. Diabetic neuropathy: clinical manifestations and current treatments. Lancet Neurol. 2012;11(6):521–534. doi: 10.1016/S1474-4422(12)70065-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Feldman EL, Nave KA, Jensen TS, Bennett DLH. New horizons in diabetic neuropathy: mechanisms, bioenergetics, and pain. Neuron. 2017;93(6):1296–1313. doi: 10.1016/j.neuron.2017.02.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Liu C, Li C, Deng Z, Du E, Xu C. Long non-coding RNA BC168687 is involved in TRPV1-mediated diabetic neuropathic pain in rats. Neuroscience. 2018;374:214–222. doi: 10.1016/j.neuroscience.2018.01.049 [DOI] [PubMed] [Google Scholar]
- 84.Liu C, Tao J, Wu H, et al. Effects of LncRNA BC168687 siRNA on diabetic neuropathic pain mediated by P2X 7 receptor on SGCs in DRG of rats. Biomed Res Int. 2017;2017:1–10. doi: 10.1155/2017/7831251 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Peng H, Zou L, Xie J, et al. lncRNA NONRATT021972 siRNA decreases diabetic neuropathic pain mediated by the P2X3 receptor in Dorsal root Ganglia. Mol Neurobiol. 2017;54(1):511–523. doi: 10.1007/s12035-015-9632-1 [DOI] [PubMed] [Google Scholar]
- 86.Liu S, Zou L, Xie J, et al. LncRNA NONRATT021972 siRNA regulates neuropathic pain behaviors in type 2 diabetic rats through the P2X7 receptor in dorsal root ganglia. Mol Brain. 2016;9(1):1–13. doi: 10.1186/s13041-016-0226-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Wang S, Xu H, Zou L, et al. LncRNA uc.48+ is involved in diabetic neuropathic pain mediated by the P2X3 receptor in the dorsal root ganglia. Purinergic Signal. 2016;12(1):139–148. doi: 10.1007/s11302-015-9488-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Xiong W, Huang L, Shen Y, et al. Effects of lncRNAuc.48+ siRNA on the release of CGRP in the spinal cords of rats with diabetic neuropathic pain. Int J Clin Exp Pathol. 2017;10(9):9960–9969. [PMC free article] [PubMed] [Google Scholar]
- 89.Cai J, Yan Y, Zhang D, et al. Silencing of lncRNA Gm14461 alleviates pain in trigeminal neuralgia through inhibiting astrocyte activation. IUBMB Life. 2020;72(12):2663–2671. doi: 10.1002/iub.2395 [DOI] [PubMed] [Google Scholar]
- 90.Xu M, Yan Y, Zhu M, Wang Z, Zhang X, Zhang D. Effects of long non-coding RNA Gm14461 on pain transmission in trigeminal neuralgia. J Inflamm. 2020;17(1):1–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Xiong W, Tan M, Tong Z, et al. Effects of long non-coding RNA uc.48+ on pain transmission in trigeminal neuralgia. Brain Res Bull. 2019;147:92–100. doi: 10.1016/j.brainresbull.2019.02.009 [DOI] [PubMed] [Google Scholar]
- 92.Meng C, Yang X, Liu Y, et al. Decreased expression of lncRNA Malat1 in rat spinal cord contributes to neuropathic pain by increasing neuron excitability after brachial plexus avulsion. J Pain Res. 2019;12:1297–1310. doi: 10.2147/JPR.S195117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Liu LP, Zhang J, Pu B, Li WQ, Wang YS. Upregulation of JHDM1D-AS1 alleviates neuroinflammation and neuronal injury via targeting miR-101-3p-DUSP1 in spinal cord after brachial plexus injury. Int Immunopharmacol. 2020;89(374):106962. doi: 10.1016/j.intimp.2020.106962 [DOI] [PubMed] [Google Scholar]
- 94.Sun RM, Wei J, Wang SS, Xu GY, Jiang GQ. Upregulation of lncRNA-NONRATT021203.2 in the dorsal root ganglion contributes to cancer-induced pain via CXCL9 in rats. Biochem Biophys Res Commun. 2020;524(4):983–989. doi: 10.1016/j.bbrc.2020.01.163 [DOI] [PubMed] [Google Scholar]
- 95.Kong C, Du J, Bu H, Huang C, Xu F, Ren H. LncRNA KCNA2-AS regulates spinal astrocyte activation through STAT3 to affect postherpetic neuralgia. Mol Med. 2020;26(1). doi: 10.1186/s10020-020-00232-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Xian S, Ding R, Li M, Chen F. LncRNA NEAT1/miR-128-3p/AQP4 axis regulating spinal cord injury-induced neuropathic pain progression. J Neuroimmunol. 2021;351(169):577457. doi: 10.1016/j.jneuroim.2020.577457 [DOI] [PubMed] [Google Scholar]
- 97.Zhang P, Sun H, Ji Z. Downregulating lncRNA PVT1 relieves astrocyte overactivation induced neuropathic pain through targeting miR-186-5p/CXCL13/CXCR5 axis. Neurochem Res. 2021;46(6):1457–1469. [DOI] [PubMed] [Google Scholar]
- 98.Zhang L, Feng H, Jin Y, et al. Long non-coding RNA LINC01119 promotes neuropathic pain by stabilizing BDNF transcript. Front Mol Neurosci. 2021;14:1–13. doi: 10.3389/fnmol.2021.673669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Mapplebeck JCS, Beggs S, Salter MW. Review articles from the 5th International Meeting of the IASP Special Interest Group on Neuropathic Pain (NeuPSIG) Sex differences in pain: a tale of two immune cells. Pain. 2015;157:2–6. [DOI] [PubMed] [Google Scholar]