Abstract
The coronavirus disease 2019 (COVID-19) pandemic is an exceptional public health crisis that demands the timely creation of new therapeutics and viral detection. Owing to their high specificity and reliability, monoclonal antibodies (mAbs) have emerged as powerful tools to treat and detect numerous diseases. Hence, many researchers have begun to urgently develop Ab-based kits for the detection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and Ab drugs for use as COVID-19 therapeutic agents. The detailed structure of the SARS-CoV-2 spike protein is known, and since this protein is key for viral infection, its receptor-binding domain (RBD) has become a major target for therapeutic Ab development. Because SARS-CoV-2 is an RNA virus with a high mutation rate, especially under the selective pressure of aggressively deployed prophylactic vaccines and neutralizing Abs, the use of Ab cocktails is expected to be an important strategy for effective COVID-19 treatment. Moreover, SARS-CoV-2 infection may stimulate an overactive immune response, resulting in a cytokine storm that drives severe disease progression. Abs to combat cytokine storms have also been under intense development as treatments for COVID-19. In addition to their use as drugs, Abs are currently being utilized in SARS-CoV-2 detection tests, including antigen and immunoglobulin tests. Such Ab-based detection tests are crucial surveillance tools that can be used to prevent the spread of COVID-19. Herein, we highlight some key points regarding mAb-based detection tests and treatments for the COVID-19 pandemic.
Keywords: Angiotensin converting enzyme II (ACE2), Coronavirus disease 2019 (COVID-19), Cytokine storm, Viral detection, Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), Spike, Receptor-binding domain (RBD), Receptor binding motif (RBM), Therapeutic antibody
Introduction
The coronavirus disease 2019 (COVID-19) pandemic is the result of widespread infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Compared to other highly transmissible viruses, SARS-CoV-2 is associated with high rates of morbidity and mortality, and it represents an unprecedented challenge to global public health [1]. Most people infected with SARS-CoV-2 experience mild to moderate respiratory illness similar to influenza or other virus infections, with symptoms such as fever, dry cough, and dyspnea. However, a considerable number of infected people develop pneumonia and acute lung injury or acute respiratory distress syndrome (ARDS); these conditions are closely associated with the relatively high mortality rate of COVID-19 [2]. Some patients also exhibit pulmonary alveolitis, bronchiolitis, accumulation of mucus and edema fluid, and different degrees of inflammation marked by infiltration of various immune cells into the pulmonary interstitium [3, 4].
The tissue distribution of the virus-targeted receptor protein, angiotensin converting enzyme II (ACE2), determines which organs will be attacked by SARS‐CoV‐2; lung, the immune system, heart, kidney, esophagus and small intestine all have high expression of ACE2 [5–8]. Based on this set of target tissues, SARS-CoV-2 can cause non‐respiratory clinical symptoms, such as diarrhea, sore throat, muscle aches, headache and vomiting, in a minority of patients [8, 9]. Moreover, patients with severe disease suffer from respiratory and lung function failure, and some even require extracorporeal membrane oxygenation (ECMO) and intensive care due to multiple organ failure and septic shock [6, 10]. Therefore, a pressing global need exists to develop vaccines and therapeutics that can mitigate the COVID-19 pandemic and cure infected patients.
Over the past year, extraordinary biomedical and financial resources have been devoted to the rapid development of diagnostic, prophylactic and therapeutic measures for this single disease. Due to their high specificity and versatility, monoclonal antibodies (mAbs) are at the fore of all three of these battlefronts in the fight against COVID-19. Recently, therapeutic mAbs have become essential tools to defeat various diseases, including virus infections, based on their abilities to prevent disease progression immediately after administration and to speed up recovery, regardless of whether the patient has fully developed immunity [11].
SARS-CoV-2 is a single-stranded RNA virus belonging to the betacoronavirus genus. As with other viruses in this genus, several critical points in the life cycle of SARS-CoV-2 can potentially be targeted and blocked by mAbs, making mAbs promising prophylactic and therapeutic agents for COVID-19. The first critical point is when the virus S protein binds to a host cell receptor, such as ACE2 [12] or cluster of differentiation 147 (CD147) [13]. After the initial binding event, host proteases, such as furin, transmembrane serine protease 2 (TMPRSS2) and cathepsin L, cleave the head of S protein, transforming it into a spring-like structure; this action allows the viral membrane to fuse with the host membrane and enables direct cell surface entry or via endosome by endocytosis [14, 15]. Once the virus enters the host cell, its RNA is translated and the innate immune response is immediately induced via host expression of type I/III interferon, chemokines and cytokines, such as tumor necrosis factor (TNF), interleukin 1 beta (IL-1β), interleukin 6 (IL-6), and granulocyte-macrophage colony-stimulating factor (GM-CSF) [6, 16, 17]. Upon continued viral replication, the cytokine levels may keep rising, leading to severe tissue damage and cytokine release syndrome (CRS) in some patients [18]. Thus, therapeutic Abs that inhibit the biological activities of cytokines may alleviate the harmful effects of over-stimulated host immune response and serve as treatments for COVID-19 [19–23].
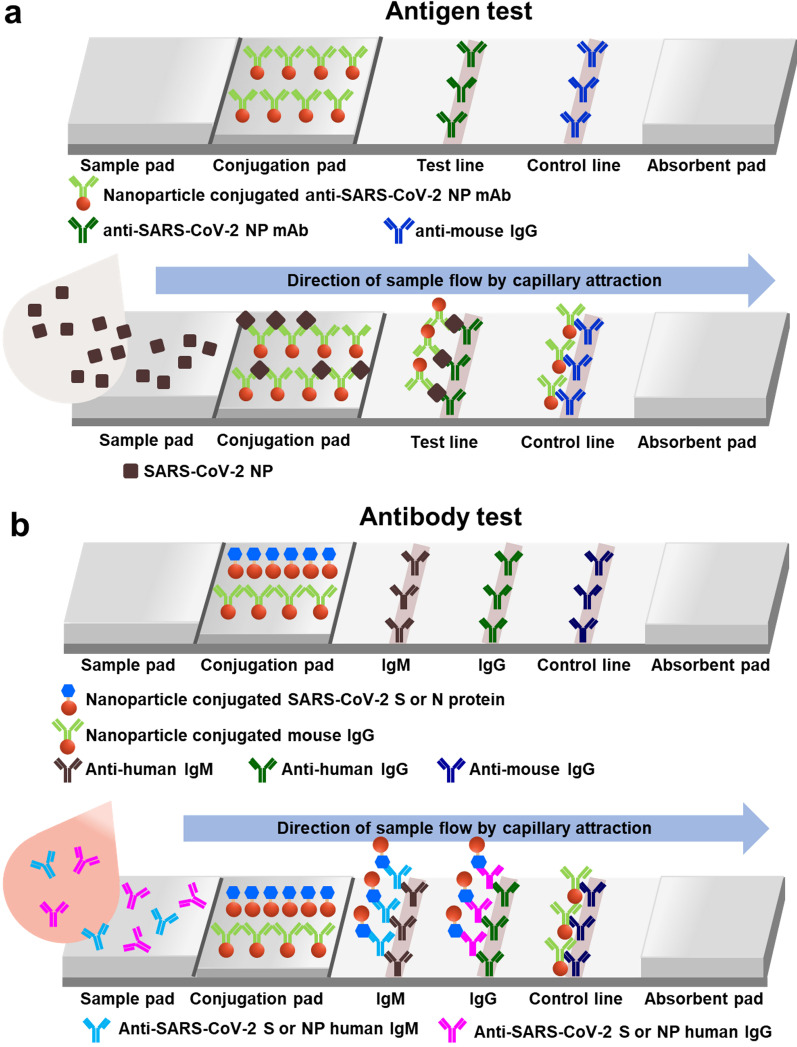
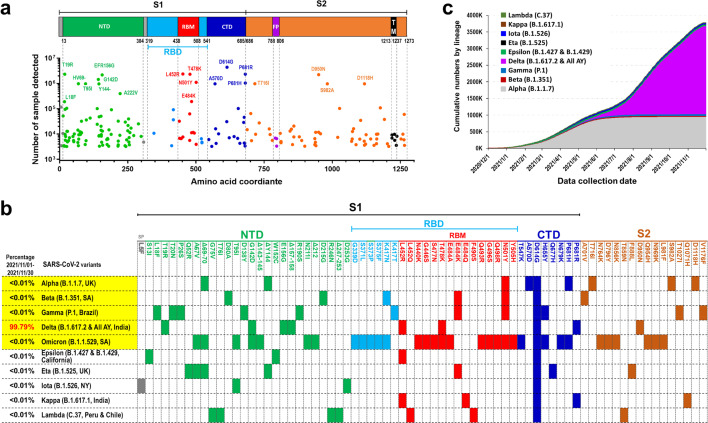
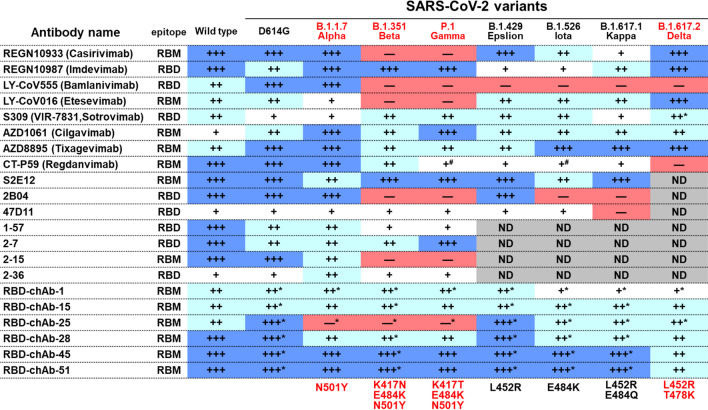
More than half of all people with SARS-CoV-2 infection have no symptoms; however, they may still be contagious in the asymptomatic state [24–26]. Four SARS-CoV-2 variants of concern that emerged in the United Kingdom (Alpha, B.1.1.7), South Africa (Beta, B.1.351), Brazil (Gamma, P.1) and India (Delta, B.1.617.2), have rapidly become dominant around the world and appear to display enhanced transmissibility and higher in-hospital mortality rates [27]. Moreover, B.1.1.529 was recently named Omicron and designated as a fifth variant of concern and by WHO after its emergence in South Africa [28]. Even more distressing, some other new SARS-CoV-2 variants that originally appeared in California (Epsilon, B.1.427 and B.1.429), Nigeria (Eta, B.1.525), New York (Iota, B.1.526), and India (Kappa, B.1.617.1 and Delta, B.1.617.2) are not only more transmissible but also exhibit reduced neutralization by convalescent and post-vaccination sera [29]. Thus, in addition to vaccines and therapeutic Abs, effective and rapid diagnostic tests for SARS-CoV-2 variants are necessary for timely medical and public health decisions, such as who should be placed in quarantine or hospitalized to reduce uncontrolled transmission. Molecular tests based on viral antigens can be used to identify individuals with acute phase SARS-CoV-2 infection, as well as control transmission when used in contact tracing, and allow for repeat testing in disease screening. Tests using Ab-antigen-formatted immunocomplexes are perhaps the most promising tools to accomplish this type of wide surveillance and control outbreaks of COVID-19. In this review, we summarize current knowledge about the use of neutralizing mAbs for prophylaxis, treatment and viral detection for COVID-19, especially focusing on those mAbs that are prime clinical candidates and have received emergency use authorization (EUA). We also describe how antibodies (Abs) can neutralize the virus in terms of S protein binding and structure. Finally, we propose strategies to combat the SARS-CoV-2 pandemic using therapeutic antibodies to overcome possible resistance of currently identified and potential mutants. The summarized information also provides insights into how therapeutic antibodies may be used against variants of SARS-CoV-2 in potential future pandemics.
Therapeutic Abs
Currently, the global effects of COVID-19 continue to grow, and the disastrous pandemic requires fast development and implementation of countermeasures. To address these needs, researchers around the world are racing to develop therapies and vaccines. Among the technologies under intensive development, neutralizing mAbs are expected to be especially useful in prophylactic and therapeutic applications, based on the success of previously developed mAb drugs [30–33].
Neutralizing Abs targeting spike (S) protein
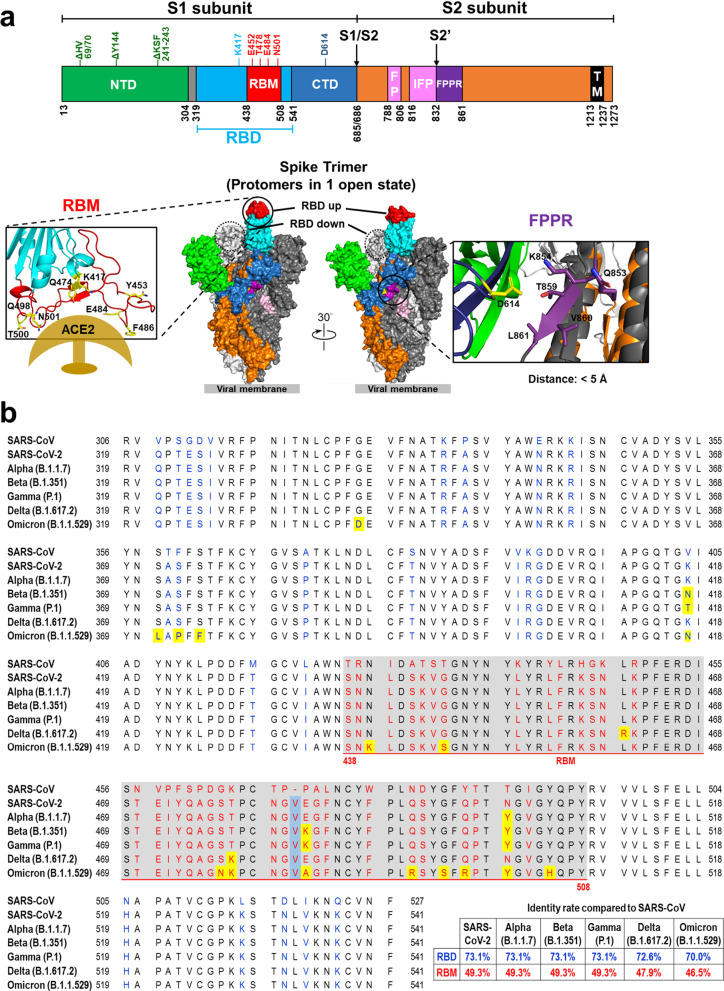
The SARS-CoV-2 S protein is a trimeric complex that is cleaved into S1 and S2 subunits (Fig. 1a). S1 is responsible for receptor binding, while S2 is responsible for membrane fusion. On human cells, the S protein targets ACE2, a key regulator of the renin-angiotensin system, which acts as the cell entry receptor for the virus. S1 protein contains an N-terminal domain (NTD) and a receptor binding domain (RBD), which interacts with the peptidase domain of ACE2 through a receptor-binding motif (RBM). Although the role of the NTD is not entirely clear, it may be responsible for the recognition of specific sugar moieties upon initial attachment; such recognition could facilitate the transition of S protein from a prefusion state to a postfusion conformation. Abs binding to certain epitopes on the NTD have been shown to inhibit SARS-CoV-2 infection [34, 35]. Moreover, SARS-CoV-2 infectivity may also be enhanced by specific antibodies against the NTD, and infectivity-enhancing antibodies have been detected in severe COVID-19 patients [36]. Neutralization of S protein function has drawn considerable attention as a means to disrupt viral entry, making the S protein the most common target for new vaccines and drugs against SARS-CoV-2.
Fig. 1.
SARS-CoV-2 Spike protein. a Structure SARS-CoV-2 spike protein. Different domains of the SARS-CoV-2 spike protein: N-terminal domain (NTD), receptor-binding domain (RBD), receptor-binding motif (RBM), subdomain 1 and 2, protease cleavage sites (S1/S2/S2′), fusion peptide (FP), internal fusion peptide (IFP), fusion peptide proximal region (FPPR), and transmembrane region (TM). HV69/70, Y144, and KSF241-243 are frequently deleted residues in the NTD of SARS-CoV-2 variants of concern. K417, E452, E484, T478, N501 and D614 are the most frequently mutated residues in the RBD of SARS-CoV-2 variants of concern. Key residues of the receptor-binding motif in the S protein of SARS-CoV-2 that interact with ACE2 are shown (lower left). The SARS-CoV-2 S protein trimeric complex is shown in a “one-up” RBD conformation. The two RBD-down protomers are depicted in light and dark gray. The RBD-up protomer is colored according to its domains; RBM in red, non-RBM RBD in light blue, NTD in green, S2 in orange, FP and IFP in pink, and FPPR in purple. The dashed circle indicates the RBD site of an RBD-down conformation protomer. Inter-atomic contacts between aspartate 614 (yellow) in a reference S monomer (dark blue) and five residues (purple) in its adjacent S protein monomer chain (dark gray) within 5 Å. These five contacts might be destabilizing and create a hydrophilic-hydrophobic repulsion that is lost upon replacement of aspartate by glycine in the D614G mutation (lower right). b RBD sequences of SARS-CoV (GeneBank: AAP30030.1), SARS-CoV-2 (GeneBank: QVW76257.1), and SARS-CoV-2 variants of concern, including B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), B.1.617.2 (Delta), and B.1.1.529 (Omicron). The amino acids encoded by SARS-CoV-2 that are altered in comparison to SARS-CoV are colored blue (RBD) or red (RBM). The amino acid inserted in SARS-CoV-2 is denoted by a light blue background. The amino acids substituted in variants of concern are denoted by a yellow background. The residues 438–508 comprise the RBM of SARS-CoV-2 and are shown with grey background
An abundance of new SARS-CoV-2 S protein-specific mAbs have been reported by different researchers [32, 33, 37] and many bind to the RBD (Fig. 2, Table 1). The major strategy used for rapid isolation of high-efficacy nAbs is reverse transcriptase-polymerase chain reaction (RT-PCR) from single human B cells [38, 39]. In this approach, the SARS-CoV-2 S or RBD protein-specific memory B cells from convalescent or acute-phase COVID-19 patients are sorted by flow cytometry, and single-cell RT-PCR for immunoglobulin genes is performed. Alternatively, nAbs have also been generated using human Ab transgenic mice [40–43], phage display library screening [44–48], yeast surface display library screening [49] or hybridoma and Ab engineering technology [50].
Fig. 2 .
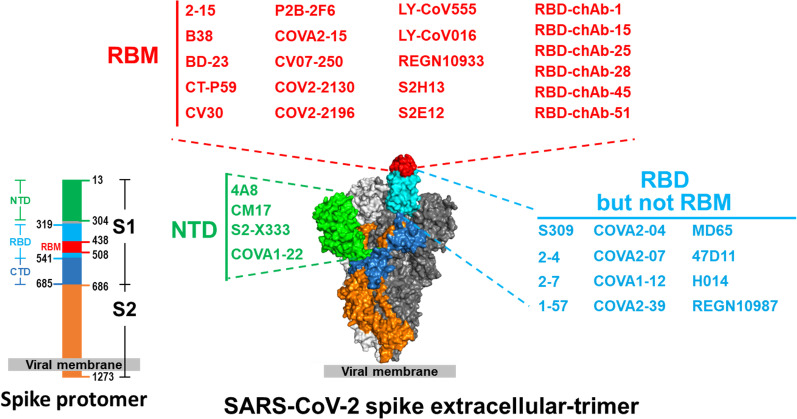
Epitopes of anti-spike and anti-RBD nAbs mapped to a surface model of SARS-CoV-2 spike trimer. The identified epitope regions are depicted as surface regions (PDB: 6VSB). Some of the shown anti-spike nAbs have known exact epitopes; for others the exact epitopes are unknown. Ab names are color-coded by the domains they recognized: N-terminal domain (NTD), light green; receptor binding motif (RBM), red; and receptor binding domain (RBD) but not RBM, cyan
Table 1.
Summary of published SARS-CoV-2-neutralizing Abs until October, 2021
| Ab name*/epitope | Source | Method for structure | In vitro neutralization | In vivo experiment | References | |
|---|---|---|---|---|---|---|
| 1 | B38/RBM | B cell, COVID-19 patient | X-ray crystallography | AV, CPE50 = 177.0 ng/ml |
hACE2 mice, Treatment, 25 mg/kg, ↓3.3 log |
[72] |
| 2 | 47D11/RBD | Hybridoma mice, SARS-CoV | Cryo-EM | AV, PRNT50 = 570.0 ng/ml |
Hamsters, Prophylactic, 3 mg/1 mL or 500 μL human convalescent plasma, TCID50 (Lung) ↓ 1–2 log |
[40, 67, 246] |
| 3 | S309 (VIR-7831, Sotrovimab)/RBD | Human patient, SARS-CoV, and Single B cell | Cryo-EM | AV, FRNT50 = 79.0 ng/ml |
Hamsters, Prophylactic, 5 mg/kg, viral load↓3 log |
[96, 97] |
| 4 |
311mab-31B5/RBD 311mab-32D4/RBD |
B cell, COVID-19 patient | N.D |
PSV, 311mab-31B5, IC50 = 33.8 ng/ml. 311mab-32D4, IC50 = 69.8 ng/ml |
N.D | [100] |
| 5 | BD-23/RBM and N165 glycan of the neighboring “down” RBD | B cell, COVID-19 patient | Cryo-EM | AV, PRNT50 = 15.0 ng/ml |
hACE2 mice, Prophylactic, 20 mg/kg, viral load, ↓ 7 log Treatment, 20 mg/kg, viral load, ↓ 4 log |
[194] |
| 6 |
2B04/RBD 1B07/RBD |
Immunized Mouse single B cell | N.D |
AV, 2B04, FRNT50 = 1.46 ng/ml 1B07, FRNT50 = 37.0 ng/ml |
hACE2 mice, Prophylactic, 10 mg/kg, RNA 107 → 106 |
[247] |
| 7 |
REGN10933(Casirivimab)/RBM REGN10987 (Imdevimab)/RBD |
REGN10933/Humanized mice REGN10987/Patient single B cell |
HDX-MS Cryo-EM |
AV, REGN10933, PRNT50 = 5.6 ng/ml REGN10987, PRNT50 = 6.3 ng/ml |
REGN10933 + REGN10987, Rhesus macaques, Prophylactic, 25 mg/kg, subgenomic RNA, ↓ 2 log Hamsters, Prophylactic, 50, 5, or 0.5 mg/kg, subgenomic RNA, ↓ 3, 2, 1 log Treatment, 50, 5, or 0.5 mg/kg, subgenomic RNA, ↓ 4, 4, 2 log |
[41, 82] |
| 8 | 4A8/NTD | B cell, COVID-19 patient | Cryo-EM |
AV, virus RNA by qPCR IC50 = 390 ng/ml |
N.D | [74] |
| 9 |
COVA1-22/NTD COVA1-18/RBD COVA2-15/RBM |
B cell, COVID-19 patient | Negative stain EM |
PSV, COVA1-18, IC50 = 8.0 ng/ml COVA2-15, IC50 = 8.0 ng/ml AV, VeroE6 cells staining, COVA1-18, IC50 = 7.0 ng/ml COVA2-15, IC50 = 9.0 ng/ml |
hACE2 mice, Prophylactic, 10 mg/kg, viral load, ↓ 4 log Treatment, 10 mg/kg, viral load, ↓ 4 log Hamsters, Treatment, 10 mg/kg, Viral titer, ↓ 3 log Cynomolgus macaques, Prophylactic, 10 mg/kg, absence of detectable sgRNA subgenomic RNA |
[71, 248] |
| 10 | CV30/RBM | B cell, COVID-19 patient | N.D | PSV, IC50 = 30 ng/ml | N.D | [70, 249] |
| 11 | P2B-2F6/RBD | B cell, COVID-19 patient | X-ray crystallography |
AV, P2B-2F6, PRNT50 = 50 ng/ml P2B-1F11, PRNT50 = 30 ng/ml |
N.D | [250] |
| 12 | C121/RBD C135/RBD C144/RBD | B cell, COVID-19 patient | Negative stain EM, X-ray crystallography, and Cryo-EM |
AV, VeroE6 cells infection (IFA) C121, IC50 = 1.64 ng/ml C135, IC50 = 2.98 ng/ml C144, IC50 = 2.55 ng/ml |
N.D | [197, 251] |
| 13 |
COV2-2130 (Cilgavimab)/RBM COV2-2196 (Tixagevimab)/RBM |
B cell, COVID-19 patient | Negative stain EM |
AV, FRNT COV2-2130, IC50 = 107 ng/ml COV2-2196, IC50 = 15 ng/ml PSV, COV2-2130, IC50 = 1.6 ng/ml COV2-2196, IC50 = 0.7 ng/ml |
hACE2 mice, 10 mg/kg Prophylactic, lung plaque assay (PFU) COV2-2130, ↓ 3 log COV2-2196, ↓ 3 log COV2-2130 + COV2-2196, ↓ 3 log Rhesus macaques, 50 mg/kg Prophylactic, subgenomic viral RNA COV2-2196, ↓ 3 log BALB/c mice, 20 mg/kg Treatment, lung plaque assay (PFU) COV2-2130, ↓ 1 log COV2-2196, ↓ 4 log COV2-2130 + COV2-2196, ↓ 4 log |
[100, 101] |
| 14 | IgG1 ab1/RBD | Fab, scFv, VH phage display libraries | N.D |
PSV, Luciferase reporter virus IC50 = 200 ng/ml |
hACE2 mice, 3 mg/kg Prophylactic, lung plaque assay (PFU) 104.5 → 101 |
[39] |
| 15 | rRBD-15/RBD | Phage display | N.D | PSV, IC50 = 1830 ng/ml | N.D | [252] |
| 16 | HbnC3t1p1_C6/RBD | B cell, COVID-19 patient | N.D | AV, CPE, IC100 = 40 ng/ml | N.D | [253] |
| 17 |
2–15/RBM 2–7/RBD |
B cell, COVID-19 patient | Cryo-EM |
AV, CPE 2–15, IC50 = 0.7 ng/ml 2–7, IC50 = 3.0 ng/ml PSV, CPE 2–15, IC50 = 5.0 ng/ml 2–7, IC50 = 10.0 ng/ml |
Hamsters, 0.3 ~ 1.5 mg/kg 2–15, Prophylactic, RNA copy 106 → 102, ↓ 4 log |
[66, 254] |
| 18 | S2H13/RBM | B cell, COVID-19 patient | Cryo-EM | PSV, IC50 = 500 ng/ml | N.D | [255] |
| 19 | S2M11/RBD S2E12/RBM | B cell, COVID-19 patient | Cryo-EM |
PSV, S2M11, IC50 = 2.1 ng/ml S2E12, IC50 = 2.3 ng/ml AV, Focus-forming assay S2M11, IC50 = 1.2 ng/ml S2E12, IC50 = 4.2 ng/ml |
Hamsters, 1 mg/kg, Prophylactic, S2M11, TCID50 105 → 101, ↓ 4 log S2E12, TCID50 105 → 101, ↓ 4 log S2M11 + S2E12, TCID50 105 → 101, ↓ 4 log 0.5 mg/kg, Prophylactic, S2M11 + S2E12, TCID50 105 → 103, ↓ 2 log |
[256] |
| 20 | CV07-209/N.D. CV07-250/RBM | B cell, COVID-19 patient | X-ray crystallography |
AV, CV07-209, PRNT50 = 3.1 ng/ml CV07-250, PRNT50 = 3.5 ng/ml |
Hamsters, CV07-209, 18 mg/kg Prophylactic, ↓ 4 ~ 5 log, Treatment, ↓ 3 ~ 4 log |
[257] |
| 21 | P008_056/NTD | B cell, COVID-19 patient | Cryo-EM and X-ray crystallography | AV, CPE50 = 30 ng/ml | N.D | [258] |
| 22 |
58G6/RBM 13G9/RBM |
B cell, COVID-19 patient | Cryo-EM |
AV, 58G6, PRNT50 = 6.0 ng/ml 13G9, PRNT50 = 9.2 ng/ml PSV, 58G6, IC50 = 4.0 ng/ml 13G9, IC50 = 5.9 ng/ml |
hACE2 mice, 10 mg/kg, Prophylactic, PRNT50 ↓ 3 log |
[207] |
| 23 |
S1D7/RBD S3D8/RBD |
Immunised Mouse | N.D |
AV, VeroE6 cells infection (IFA) S1D7, IC50 = 405 ng/ml S3D8, IC50 = 139 ng/ml S1D7 + S3D8, IC = 200 ng/ml |
N.D | [259] |
| 24 |
Wang-C387/RBD Wang-C437/RBD |
B cell, COVID-19 patient | N.D |
AV, VeroE6 cells infection (IFA) Wang-C387, IC50 = 8.4 ng/ml Wang-C437, IC50 = 2.0 ng/ml PSV, Wang-C387, IC50 = 10.6 ng/ml Wang-C437, IC50 = 4.9 ng/ml |
N.D | [260] |
| 25 | S2-X333/NTD | B cell, COVID-19 patient | Cryo-EM |
AV, S2-X333, IC50 = 2.0 ng/ml |
Hamsters, viral challenge Viral RNA copies/mg lung: 4 mg/kg, 106 → 103, ↓ 3 log TCID50/mg lung: 1 mg/kg, 104 → 101, ↓ 3 log 4 mg/kg, 104 → 101, ↓ 3 log |
[34] |
| 26 | C601/RBD | B cell, COVID-19 patient | Cryo-EM |
PSV, Luciferase assay IC50 = 2.0 ng/ml |
N.D | [191] |
| 27 | LY-CoV555 (Bamlanivimab)/RBD | B cell, COVID-19 patient |
Cryo-EM and X-ray crystallography |
AV, PRNT50 = 20 ng/ml (WA isolate) PRNT50 = 49 ng/ml (Italy isolate) PSV, stably transfected ACE2 IC50 = 12 ng/ml |
Rhesus macaques, 2.5 mg/kg Prophylactic, BAL viral replication (Day3): ↓ > 1 log RNA copies/ml BAL viral replication (Day6): ↓ > 2 log RNA copies/ml Lung viral replication (Day6): ↓ > 3 log RNA copies/ml |
[89] |
| 28 | XG003/RBD | B cell, COVID-19 patient | N.D |
AV, XG005, IC50 = ~ 100 ng/ml XG014, IC50 = 5.1 ng/ml PSV, Luciferase assay XG005, IC50 = 6.1 ng/ml XG014, IC50 = 14.4 ng/ml |
N.D | [261] |
| 29 | CM17/NTD | B cell, COVID-19 patient | Cryo-EM | AV, IC50 = 30 ng/ml |
MA10 mice, virus titer (PFU), 105 to 103, ↓ 2 log (MA10 mice: BALB/c mouse model, a pathogenic mouse ACE2-adapted SARS-CoV-2 variant) |
[262] |
| 30 | ABP18/RBD | Phage Display (Ab, human, non-immune) | N.D |
PSV, Luciferase assay IC50 = 60 ng/ml |
N.D | [263] |
| 31 | ION-360/RBD | B cell, COVID-19 patient | X-ray crystallography |
PSV, Luciferase assay IC50 = 25.5 ng/ml |
N.D | [264] |
| 32 | STE90-C11/RBD | Phage Display Library (Antibody, human, immune—CoV2) | X-ray crystallography | AV, PRNT50 = 84 ng/ml | N.D | [48] |
| 33 | FC05/NTD | Phage Display Library (Antibody, human, immune—CoV2) | Cryo-EM | N.D | N.D | [265] |
| 34 | P17/RBD | Phage Display (Ab, human, non-immune) | Cryo-EM |
PSV, IC50 = 24.8 ng/ml, AV, PRNT50 = 29.2 ng/ml |
hACE mice, 20 mg/kg Prophylactic + Treatment, ↓ 1.93 log RNA copies/g, > 2 log PFU/ml (lung) Treatment, ↓ 1 log RNA copies/g, > 2 log PFU/ml (lung) |
[266] |
| 35 | HB27/RBD | Humanized from Immunised Mouse | Cryo-EM |
PSV, IC50 = 6 ng/ml AV, PRNT50 = 33 ng/ml |
hACE mice, 20 mg/kg Prophylactic, Day3 (lung): ↓ 5 log RNA copies/g, > 3 log PFU/ml Day5 (lung): ↓ 3 log RNA copies/g, > 1 log PFU/ml Therapeutic treatment: Day3 (lung): ↓ 4 log RNA copies/g, > 3 log PFU/ml Day5 (lung): ↓ 3 log RNA copies/g, > 1 log PFU/ml |
[267] |
| 36 | 6D3/S1-S2 cleavage Site | Mouse Hybridoma | X-ray crystallography | N.D | N.D | [268] |
| 37 | P4A1/RBD | B cell, COVID-19 patient | X-ray crystallography | PSV, IC50 = 975 ng/ml |
Cynomolgus monkeys, 10 mg/kg, Day7 (lung): ↓ 3–4 log viral load (copies/g) |
[43] |
| 38 | P5A-1B9/RBM | B cell, COVID-19 patient | Cryo-EM |
AV, IC50 = 16.5 ng/ml PSV, IC50 = 12.0 ng/ml |
N.D | [269] |
| 39 | TAU-1109/RBD | B cell, COVID-19 patient | N.D |
PSV, pseudo-typed GFP SARS-CoV-2 IC50 = 45 ng/ml |
N.D | [270] |
| 40 | 58G6/RBD | B cell, COVID-19 patient | N.D |
AV, RT-qPCR IC50 = 9.98 ng/ml |
N.D | [271] |
| 41 | H014/RBD | Immunized Humanized (hACE2) Mouse | Cryo-EM | AV, PRNT50 = 5725.5 ng/ml |
hACE2 mice, 50 mg/kg, Viral load, Treatment ↓ 1 log, Prophylactic + therapeutic treatment ↓2 log |
[46] |
| 42 | BD-368–2/RBM | B cell, COVID-19 patient | Cryo-EM |
AV, IC50 = 15 ng/ml: PSV, IC50 = 1.2 ng/ml |
hACE2 mice, 20 mg/kg, Prophylactic, Viral load ↓6 log Treatment, Viral load ↓3 log |
[194, 272] |
| 43 | CnC2t1p1_B4/RBD | B cell, COVID-19 patient | N.D | AV, IC100 = ~ 10,000 ng/ml | N.D | [253] |
| 44 | 413–2/non-RBD | B cell, COVID-19 patient | N.D |
AV, IC50 = ~ 7500 ng/ml PSV, IC50 = 8198 ng/ml |
N.D | [273] |
| 45 | EY6A/RBD | B cell, COVID-19 patient | X-ray crystallography | AV, PRNT50 ~ 10,800 ng/ml | N.D | [60] |
| 46 | Fab1-20/RBD | B cell, COVID-19 patient | N.D | PSV, IC50 = 8 ng/ml | N.D | [66] |
| 47 | MD65/RBD | Phage Display Library (Antibody, human, immune—CoV2) | N.D | AV, PRNT50 = 220 ng/ml | N.D | [73] |
| 48 | CC12.1/RBD | B cell, COVID-19 patient | X-ray crystallography |
PSV, HeLa-ACE2, IC50 = 46 ng/ml VER0-6, IC50 = 120 ng/ml |
Hamsters, Prophylactic, 16.5 ~ 4.2 mg/kg Viral RNA, ↓2.5 log |
[39, 274] |
| 49 | CA521/RBD | Transgenic Mouse | Cryo-EM |
AV, PRNT50 = 0.73 ng/ml PSV, IC50 = 0.1 ng/ml |
C57BL/6 mice, 20 mg/kg, Prophylactic, Viral RNA ↓2–4 log |
[275] |
| 50 | BG10-19/RBD | B cell, COVID-19 patient | Cryo-EM and X-ray crystallography |
PSV, D614G, IC50 = 2.0 ng/ml B.1.1.7, IC50 = 1.0 ng/ml B.1.351, IC50 = 4.0 ng/ml |
N.D | [276] |
| 51 | COV2-2531/S2 | B cell, COVID-19 patient | Negative stain EM | PSV, IC50 = 1.6 ng/ml |
hACE2 mice, 10 mg/kg, Viral RNA ↓2 log |
[207] |
| 52 |
RBD-chAb-15/RBM RBD-chAb-25/RBM RBD-chAb-45/RBM |
Hybridoma screening and humanized | Cryo-EM |
AV, WT, RBD-chAb-15, PRNT50 = 30.3 ng/ml RBD-chAb-25, PRNT50 = 15.8 ng/ml RBD-chAb-45, PRNT50 = 9.9 ng/ml B.1.617.2, RBD-chAb-15, PRNT50 = 37.8 ng/ml RBD-chAb-45, PRNT50 = 18.0 ng/ml RBD-chAb-15 + 45, PRNT50 = 37.5 ng/ml PSV, WT, RBD-chAb-15, IC50 = 52.3 ng/ml RBD-chAb-25, IC50 = 25.44 ng/ml RBD-chAb-45, IC50 = 2.3 ng/ml B.1.617.2, RBD-chAb-15, PRNT50 = 103.6 ng/ml RBD-chAb-45, PRNT50 = 15.5 ng/ml RBD-chAb-15 + 45, PRNT50 = 25.7 ng/ml |
Hamsters, Prophylactic, WT, 3 mg/kg RBD-chAb-15: TCID50 ↓ 1 log RBD-chAb-45: TCID50 ↓ 3.5 log RBD-chAb-15 + 45: TCID50 ↓ 4 log WT, 4.5 mg/kg RBD-chAb25: TCID50 ↓ 2 log RBD-chAb45: TCID50 ↓ 2 log RBD-chAb25 + 45: TCID50 ↓ 4 log AAV-hACE2 mice, Treatment, WT, 3 mg/kg RBD-chAb25 + 45: TCID50 ↓ 1.5 log Hamsters, Treatment, WT, 3 mg/kg RBD-chAb-15 + 45: TCID50 ↓ 4 log RBD-chAb25 + 45: TCID50 ↓ 4 log B.1.617.2, 6 mg/kg RBD-chAb-45: TCID50 ↓ 3 log RBD-chAb-15 + 45: TCID50 ↓ 3.5 log |
[50, 75, 76] |
| 53 | CT-P59 | B cell, COVID-19 patient | X-ray crystallography | AV, PRNT50 = 8.4 ng/ml |
Ferrets, Treatment, 30 mg/kg TCID50 (Lung) ↓ 1 log Hamsters, Treatment, 30 mg/kg TCID50 (Lung) ↓ 7 log Rhesus monkeys, Treatment, 45 mg/kg TCID50 (Lung) unchanged |
[47] |
| 54 | LY-CoV016 (Etesevimab, CB6 JS016,)/RBM | B cell, COVID-19 patient | X-ray crystallography |
AV, CPE50 = 36 ng/ml PSV, CPE50 = 23 ng/ml |
Rhesus monkeys, Prophylactic, 50 mg/kg Day3 (lung): ↓ 4 log RNA Rhesus monkeys, Treatment, 50 mg/kg Day3 (lung): ↓ 2 log RNA |
[51] |
| 55 | 2C08/RBD | B-cell; SARS-CoV-2 Vaccinee | Cryo-EM | AV, FRNT50 = 5 ng/ml |
Hamsters, 2 mg/animal Prophylactic, viral RNA ↓ 3–4 log |
[277] |
| 56 | S2X259/RBD | B cell, COVID-19 patient | Cryo-EM |
AV, S2X259, PRNT50 = 144.2 ng/ml PSV, IC50 = 212.3 ng/ml |
Hamsters, B.1.351 viral challenge TCID50/mg lung: 1 mg/kg, 104 → 103, ↓ 1 log 4 mg/kg, 104 → 101, ↓ 3 log 1 + 1 mg/kg with S309, 104 → 101, ↓ 3 log |
[61] |
| 57 | A23-58–1/RBD | B cell, COVID-19 patient |
Cryo-EM and Negative stain EM |
AV, CPE, USA-WA1, IC50 = 2.0 ng/ml PSV, Luciferase assay D614G, IC50 = 1.8 ng/ml B.1.1.7, IC50 < 0.6 ng/ml B.1.351, IC50 = 1.6 ng/ml |
N.D | [278] |
| 58 | COV107-23/RBD | B-cells; SARS-CoV-2 Human patient | X-ray crystallography | N.D | N.D | [279] |
| 59 | 910–30/RBD | B-cells; SARS-CoV-2 Human patient | Cryo-EM |
AV, CPE, IC50 = 183 ng/ml PSV, Luciferase assay IC50 = 66 ng/ml |
N.D | [280] |
| 60 |
DH1043/RBD DH1052/NTD |
B-cells; SARS-CoV-2 Human patient |
Cryo-EM and Negative stain EM |
PSV, Luciferase assay DH1043, IC50 = 34 ng/ml DH1050, IC50 > 100,000 ng/ml |
BALB/c mice, 30 mg/kg, Prophylactic, DH1052, viral RNA, ↓ 1 log Macaque, Prophylactic, 10 mg/kg lung subgenomic RNA DH1043, ↓ 5 log DH1052 ↓ < 1 log |
[281] |
| 61 | C1A-B3/RBD | B-cells; SARS-CoV-2 Human patient | X-ray crystallography |
AV, PRNT50 = 62 ng/ml PSV, Lentivirus pseudotype D614G, IC50 = 81 ng/ml |
N.D | [282] |
| 62 | S2H97/RBD | B-cells; SARS-CoV-2 Human patient | X-ray crystallography and Cryo-EM |
AV, PRNT50 = 794 g/ml PSV, PRNT50 = 338 ng/ml |
Hamsters, 25 mg/animal Prophylactic, viral RNA ↓ 4 log |
[62] |
| 63 | 47D1/RBD | B-cells; SARS-CoV-2 Human patient | X-ray crystallography |
AV, PRNT50 = 12.7 ng/ml PSV, Luciferase assay IC50 = 6.0 ng/ml |
Hamsters, Prophylactic, 100, 25, or 6.25 mg/kg, lung viral RNA ↓ 1 log 1.6, or 0.4 mg/kg, lung viral RNA without difference |
[283] |
| 64 | S2P6/S2 | B-cells; SARS-CoV-2 Human patient | X-ray crystallography and Cryo-EM |
PSV, Luciferase assay D614G, IC50 ~ 10,000 ng/ml P.1, IC50 ~ 10,000 ng/ml B.1.1.7, IC50 ~ 100,000 ng/ml B.1.351, IC50 ~ 100,000 ng/ml B.1.617.1, IC50 ~ 20,000 ng/ml |
Hamsters, Prophylactic, Prototypic SARS-CoV-2 20 mg/kg, TCID50 (Lung) ↓ 2 log 2 mg/kg, TCID50 (Lung) < 1 log B.1.351 SARS-CoV-2 20 mg/kg, TCID50 (Lung) ↓ 1.5 log |
[284] |
| 65 | P5A-3C8/RBD | B-cells; SARS-CoV-2 Human patient | X-ray crystallography |
AV, FRNT50 = 11.2 ng/ml PSV, Luciferase assay IC50 = 20.6 ng/ml |
Hamsters, Prophylactic, 5 mg/kg, lung viral RNA ↓ 1 log |
[285] |
| 66 | 5A6/RBD | Phage Display (Ab, human, non-immune) | Cryo-EM |
AV, CPE, IC50 = 140.7 ng/ml PSV, Luciferase assay IC50 = 75.5 ng/ml |
N.D | [286] |
| 67 | BLN12/NTD | Phage Display (Ab, human, immune [SARS-CoV-2]) | N.D | AV, PRNT50 = 8.0 ng/ml |
hACE2 mice, Prophylactic 5 mg/kg, 100% protection of death 0.5 mg/kg, 80% protection of death |
[287] |
| 68 | N12-11/NTD | B-cells; SARS-CoV-2 Human patient | Cryo-EM |
PSV, Luciferase assay IC50 ~ 490 ng/ml |
N.D | [288] |
| 69 | 2B11/RBD | Phage Display (Ab, human, immune [SARS-CoV-2]) | X-ray crystallography |
AV, PRNT50 = 1.0 ng/ml PSV, Luciferase assay IC50 = 6.0 ng/ml B.1.1.7, IC50 = 12.2 ng/ml B.1.351, IC50 = 5091 ng/ml P.1, IC50 = 2527 ng/ml |
hACE2 mice, 25 or 75 mg/kg, Prophylactic, lung viral RNA ↓ 2 log Treatment, lung viral RNA ↓ 1 log |
[289] |
| 70 | mAb40/RBD | B-cells; SARS-CoV-2 Human patient | N.D |
AV, B.1.167.2, FRNT50 = 29 ng/ml PSV, Luciferase assay B.1.167.1, IC50 = 24 ng/ml B.1.167.2, IC50 = 24 ng/m B.1.1.519, IC50 = 17 ng/ml l B.1.429, IC50 = 11 ng/ml |
N.D | [216] |
| 71 | C549/RBD | B-cells; SARS-CoV-2 Human patient | N.D |
PSV, Luciferase assay WT, IC50 = 10.95 ng/ml Q493R, IC50 = 2.35 ng/m E484G, IC50 = 2.29 ng/ml R346S, IC50 = 8.33 ng/ml |
N.D | [220] |
| 72 | SARS2-38/RBD | Immunised Mouse | Cryo-EM |
AV, FRNT50 = 5.0 ng/ml PSV, FRNT50 = 6.0 ng/ml |
hACE2 mice, 5 mg/kg Ab, Treatment, viral RNA, ↓ 3 log Hamsters, 10 mg/kg, Treatment, viral RNA, ↓ 2 log |
[290] |
| 73 | 54,042–4/RBD | B-cells; SARS-CoV-2 Human patient | Cryo-EM |
PSV, Real-time cell analysis assay IC50 = 9.0 ng/ml AV, ELISA B.1.1.7, IC50 = 5.5 ng/ml B.1.351, IC50 = 9.7 ng/ml B.1.617.2, IC50 = 1.5 ng/ml P.1, IC50 = 3.7 ng/ml |
N.D | [291] |
| 74 | MA1/RBD | Immunised Mouse | Cryo-EM |
AV, Luciferase assay IC50 ~ 10 ng/ml PSV, Luciferase assay IC50 ~ 10 ng/ml |
N.D | [292] |
| 75 | C12A2/RBD C12C9/NTD | B-cells; SARS-CoV-2 Human patient | Cryo-EM |
AV, CPE USA-WA1 C12A2, IC50 = 2 ng/ml C12C9, IC50 = 43 ng/ml B.1.1.7, C12A2, IC50 = 8 ng/ml C12C9, IC50 = 6 ng/ml B.1.351, C12A2, IC50 > 50 ng/ml C12C9, IC50 > 500 ng/ml |
N.D | [293] |
| 76 | TRES6/RBD | Transgenic Mouse | N.D |
AV, CoV-2-ER1 (D614G) TRES6, IC50 = 102 ng/ml TRES6hu, IC50 = 33 ng/ml |
hACE2 mice, viral challenge 5.25 mg/kg Ab, Log10 viral load (RNA copies) reduction: 4 days post, lung 30x, BAL 40x 10 days post, lung 100x, BAL 400x Prevented body weight loss, Reduced clinical symptoms |
[294] |
| 77 | C1027/RBD | B-cells; SARS-CoV-2 Human patient | N.D |
PSV, after 12 month WT, IC50 = 20.8 ng/ml K417N, IC50 = 4.1 ng/m E484K, IC50 = 3.4 ng/ml N501Y, IC50 = 16.8 ng/ml AV, after 12 month WA1/2020, IC50 = 9.35 ng/ml B.1.351, IC50 = 6.08 ng/ml |
N.D | [295] |
| 78 | NT-193/RBD | Immunised mouse (TC-mAb) | X-ray crystallography |
PSV, WT IgG1, IC50 = ~ 5.0 ng/ml IgG3, IC50 = ~ 1.0 ng/ml AV, WT, IgG1, TCID50 = ~ 600 ng/ml IgG3, TCID50 = ~ 600 ng/ml D614G, IgG1, TCID50 = ~ 250 ng/ml IgG3, TCID50 = ~ 150 ng/ml |
Hamsters, IgG3, Viral RNA copies/mg lung: Prophylactic, 1.25 mg/kg, 106 → 105, ↓ 1 log 5 mg/kg, TCID50 106 → 105, ↓ 1 log Treatment, 1.25 mg/kg, 106 → 105, ↓ 1 log 5 mg/kg, TCID50 106 → 104, ↓ 2 log |
[296] |
| 79 | 7B8/RBD | Immunised mouse (RenMab) | Cryo-EM |
PSV, D614G, IC50 = ~ 100 ng/ml B.1.1.7, IC50 = ~ 100 ng/ml N501Y, IC50 = ~ 100 ng/ml |
N.D | [297] |
| 80 | CC40.1/RBD | B-cells; SARS-CoV-2 Human patient | X-ray crystallography |
PSV, IC50 < 100 ng/ml |
N.D | [298] |
| 81 | STE73-2E9/RBD | Phage Display Library (Antibody, human, immune-CoV2) | N.D | AV, TCID50 = 61.5 ng/ml | N.D | [48] |
| 82 | Fab-324/RBD | Phage Display Library (Antibody, human, non-immune) | Cryo-EM |
PSV, Multabody, IC50 = 24 ng/ml IgG, IC50 = 21,000 ng/ml |
N.D | [299] |
| 83 | P5C3/RBD | B-cells; SARS-CoV-2 Human patient | Cryo-EM |
PSV, WT, IC50 = 4.0 ng/ml D614G, IC50 = 14.0 ng/ml E484K/N501Y, IC50 = 4.0 ng/ml K417N/E484K/N501Y, IC50 = 13.0 ng/ml AV, CPE WT, IC50 = 5.0 ng/ml D614G, IC50 = 11.0 ng/ml B.1.1.7, IC50 = 8.0 ng/ml B.1.351, IC50 = 3.0 ng/ml |
Hamsters, Prophylactic, 5.0, 1.0, or 0.5 mg/kg Lung viral RNA, all 105 → 102, ↓ 3 log |
[300] |
| 84 |
PDI-222/RBD, WCSL-119/RBD |
PDI-222: B-cells; SARS-CoV-2 Human patient WCSL-119: Semi-synthetic Human Fab Library |
Cryo-EM |
AV, PDI-222, WT, PRNT50 = 5.0 ng/ml D614G, PRNT 50 = 11.0 ng/ml WCSL-119, WT, PRNT50 = 22.0 ng/ml D614G, PRNT 50 = 25.0 ng/ml |
B57BL mice, SARS-CoV-2 (D614G N501Y) Prophylactic, 5, 1, or 0.2 mg/kg PDI-222, TCID50 all ↓ 2 log WCSL-119, 5 or 1 mg/kg, TCID50 ↓ 2 log 0.2 mg/kg, TCID50 no change Hamsters, PDI-222, Prophylactic 5 mg/kg, TCID50 ↓ 5 log 0.25 mg/kg, TCID50 ↓ < 1 log |
[301] |
| 85 | C1207/RBD | B-cells (Human Naïve, mRNA vaccination) | N.D |
PSV, after 5 months vaccination WT, IC50 = 17.8 ng/ml K417N, IC50 = 7.2 ng/m N501Y, IC50 = 10.0 ng/ml E484K/R683G, IC50 = 3.3 ng/ml L525R/E484K/R683G, IC50 = 2.4 ng/ml |
N.D | [302] |
Note 1: In vitro neutralization experiment refers to authentic (AV) or pseudotyped (PSV) SARS-CoV-2 neutralization assay as indicated
Note 2: In vivo experiment refers to the animal type, Ab injected amount, and observed prophylactic or treatment efficacy as indicated
↓, decrease after compared to the control group; ~ , roughly estimated; n-log, n × 10 times; AV authentic SARS-CoV-2 virus, CPE cytopathic effect, IC50 half-maximal inhibitory concentrations, IC100 100% inhibitory concentration, IFA immunofluorescence assays, N.D. not determined, NTD N-terminal domain, PFU plaque-forming unit, PSV SARS-CoV-2 pseudovirus, PRNT50 50% reduction of plaque neutralization test, qPCR real-quantitative polymerase change reaction, RBD receptor binding domain, RBM receptor binding motif, scFv single-chain fragment variable, TCID50 median tissue culture infectious dose; *Only listed representative Abs in indicated published papers
In order to screen Ab candidates for neutralizing capability in vitro, most groups test Abs against authentic living SARS-CoV-2, while some use pseudovirus with reporter readouts. A few methods have been used to quantify inhibitory concentrations, such as plaque reduction neutralization test (PRNT), focus reduction neutralization assay (FRNT), cytopathic effect (CPE), luciferase luminescence quantification, immunofluorescence assay (IFA), and virus mRNA quantification by quantitative polymerase change reaction (qPCR). The use of such a wide variety of in vitro assay methods makes it difficult to directly compare Abs from different publications (Table 1). To bring nAbs one step closer to clinical trials, a handful of publications also include data from in vivo animal models, which demonstrate the efficacy of the Ab as a treatment or prophylactic agent. Mice are not affected by SARS-CoV-2, presumably due to differences in ACE2 amino acid sequence compared to humans. Hence, mouse models for testing SARS-CoV-2 neutralizing capability must be generated by introducing human ACE2 into the lung cells of mice, either by the use of transgenic methods or by infecting normal mice with adenovirus encoding the human ACE2 gene for transient expression. As an alternative to mice, Shi et al. performed animal experiments in a rhesus macaque model; in this model, nAbs administered in both protection and treatment contexts caused clear reductions in viral load and lung damage [51]. Moreover, hamsters develop severe and easily observed signs of illness after infection with SARS-CoV-2, including rapid weight loss, a very high viral load in the lungs, and severe lung pathology [52]. Therefore, hamsters have become a commonly used model to evaluate the prophylactic and therapeutic efficacy of Abs (Table 1).
Next, we introduce prominent nAbs that bind to the RBD or non-RBD sites on the S protein, focusing on Abs that have received EUA from the U.S. FDA [53–58].
Neutralizing Abs targeting the S protein RBD
The RBD of the S protein is a target of multiple nAbs that inhibit SARS-CoV-2 infection by disrupting the interaction between the RBD and ACE2. Notably, the RBD sequence of SARS-CoV-2 S protein shares 73% amino acid identity with that of SARS-CoV (Fig. 1b), and the two viruses both possess a conserved epitope in the RBD that allows for possible Ab cross-reactivity. However, most SARS-CoV-nAbs do not bind the SARS-CoV-2 RBD, nor do they neutralize SARS-CoV-2 [59]. Only a few Abs have been shown to bind both SARS-CoV and SARS-CoV-2 [40, 60–62]. Researchers have used cryogenic electron microscopy (cryo-EM) to reveal that the structure of the SARS-CoV-2 S protein is an asymmetric trimer, with two conformations for the RBD (“open” and “closed”) [63, 64]. This dynamic conformation of the RBD may be a key factor affecting the neutralizing potency of anti-RBD Abs.
H014
Abs against the SARS-CoV-2 RBD were identified by screening a phage-display single-chain fragment variable (scFv) library generated from spleen mRNA of mice immunized with recombinant SARS-CoV RBD [46]. Among the hits from this screen, a potent nAb, H014, was found to bind the RBDs of SARS-CoV-2 and SARS-CoV with extremely high affinities (sub-nM concentrations). Cryo-EM reconstruction showed that H014 recognizes a conformational epitope on one side of the open (up) RBD, distinct from the RBM, whereas the closed RBD is inaccessible to H014. The authors had previously established human ACE2 knock-in mice using CRISPR/Cas9 technology as a model for SARS-CoV-2 infection [65]. The hACE2-humanized mice were infected with 5 × 105 PFU of SARS-CoV-2 intranasally and then treated by intraperitoneal injection of H014 at 50 mg/kg. In therapeutic and prophylactic plus therapeutic groups, H014 treatment reduced viral titers in the lungs at day 5 by approximately tenfold and 100-fold, respectively. These results indicated a potential therapeutic use for H014 in treating COVID-19.
2-15
Dr. David D. Ho’s group reported a collection of 61 SARS-CoV-2-nAbs from five infected patients with high plasma virus-neutralizing titers [66]. Their strategy for isolating Abs included sorting of SARS-CoV-2 S-specific memory B cells by flow cytometry and single-cell sequencing. Nineteen of the reported Abs could neutralize SARS-CoV-2 in vitro, with nine exhibiting high potency. Epitope mapping showed that about half of the 19 Abs are directed against the RBD, while the other half target the NTD, the top region of S protein. The RBD-directed Abs were shown to neutralize authentic SARS-CoV-2 virus with IC50 values of 0.7 to 209 ng/ml; the most potent Abs were 2-15, 2-7, 1-57 and 1-20. The NTD-directed Abs showed similar neutralizing activities, with the most potent being 2-17, 5-24 and 4-8 Abs. Cryo-EM structures were determined for several of the mAbs in complex with the S trimer to clarify Ab epitopes. The 2-4 Ab targeted the RBD and lock it into a “down” conformation, also obstructing the interaction with ACE2. The 4-8 Ab recognized the tip of the NTD, and 2-43 Ab recognized the top of the RBD, bridging two separate RBDs. In a study to evaluate prophylaxis in SARS-CoV-2-infected hamster models, a dosage of 1.5 mg/kg 2-15 showed protective efficacy, as it could reduce virus titer by more than four orders of magnitude. Thus, a relatively modest dose of this Ab almost completely prevented infection of SARS-CoV-2 in vivo.
Unfortunately, SARS-CoV-2 variants B.1.1.7 (Alpha) and B.1.351 (Beta) are resistant to neutralization by most NTD-targeting Abs, including 2-17, 5-24, and 4-8 [53]. However, both 5-24, and 4-8 retain the ability to inhibit the P.1 (Gamma) variant from Brazil [54]. Anti-RBD Abs (i.e., 2-15, 1-20 and 2-43) have impaired function against B.1.1.7 (Alpha), and the neutralizing potency against B.1.351 (Beta) is fully lost. The activity of anti-RBD Ab 1-57 is diminished by 1.5-fold against B.1.1.7 (Alpha) and 5.2-fold against B.1.351 (Beta). Meanwhile, the activity of 2-7 is unaffected by the variations in B.1.1.7 (Alpha), B.1.351 (Beta) and P.1 (Gamma), but its IC50 is reduced 3.4-fold when used against the E484K-single mutation pseudovirus [53, 54].
47D11
Wang et al. characterized a human mAb, 47D11, which is capable of neutralizing both SARS-CoV and SARS-CoV-2 in vitro [40]. This Ab was generated from H2L2 human Ab transgenic mice, which were immunized with the S ectodomain of HCoV-OC43, SARS-CoV, and MERS-CoV. Cryo-EM structures showed that 47D11 binds specifically to the closed conformation of the RBD, distal to the ACE2 binding site [67]. Interestingly, 47D11 preferentially recognizes the partially open conformation of the SARS-CoV-2 S protein, suggesting that it could be used effectively in combination with other Abs that target the exposed RBM. AbbVie has a license for this Ab from Harbour BioMed and completed a phase I clinical trial for the prevention and treatment of COVID-19 [68, 69].
CV30
Hurlburt et al. isolated a potent neutralizing mAb, CV30, from a patient infected with SARS-CoV-2 [70]. CV30 binds the RBD, neutralizes pseudovirus with an IC50 of 0.03 μg/ml, and competes for binding sites with ACE2. The X-ray crystal structure revealed that CV30 almost exclusively binds to the RBM in the RBD. Notably, CV30 has minimal somatic mutations compared to the germline sequence; it has only a two-residue change in heavy chain of variable domain and no change in the light chain of variable domain.
COVA2-15
Brouwer et al. isolated 19 nAbs from three convalescent COVID-19 patients using a stabilized prefusion SARS-CoV-2 S protein [71]. These Abs target a diverse range of epitopes on the S protein, and two showed picomolar neutralizing activities against authentic SARS-CoV-2 virus. EM was used to reveal the structures of six RBD antigen-binding fragments (Fabs). Four interacted with a stoichiometry of one Fab per trimer, with RBDs in the up state. COVA2-15 was able to bind RBD domains in both the up and down states.
B38 and H4
Wu et al. isolated four nAbs from a convalescent COVID-19 patient. Two of the Abs, B38 and H4, blocked RBD binding to ACE2 [72]. The Kd for B38 binding to the RBD was measured using surface plasmon resonance (SPR) at 70.1 nM, while that of H4 was 4.48 nM. The abilities of B38 and H4 Abs to protect against SARS-CoV-2 in vivo were also explored. hACE2 transgenic mice were treated with a single dose of 25 mg/kg B38 or H4 Abs 12 h after viral challenge. The RNA copies of virus in both the B38-treated and H4-treated groups were significantly reduced (by 3.3 and 2.7 orders of magnitude, respectively). A competition assay indicated the B38 and H4 epitopes on the RBD are different, and a cocktail of both Abs exhibited synergistic neutralizing ability in Vero-E6 cells. This pair of Abs could therefore potentially be used together to prevent immune escape in clinical applications.
MD65
Phage display is a powerful technique that enables rapid, efficient, and high-throughput selection of Abs (scFv or Fab) against antigens in vitro [48]. Several human Ab drugs derived from phage display libraries have been approved and are currently on the market. Noy-Porat et al. constructed a phage display scFv library using peripheral circulatory lymphocytes collected from patients in the acute phase of disease [73]. The phage scFv library complexity was 9.2 × 106, and the library was used for affinity selection of Abs against RBD-human fragment crystallizable (Fc). Eight fully human, SARS-CoV-2-nAbs were isolated and characterized. These Abs target four distinct epitopes on the S protein RBD. Evaluation of the Ab affinities toward S1 by biolayer interferometry (BLI) revealed Kd values of these human Abs ranging from 0.4 to 5.8 nM. The neutralization potencies of the Abs were then evaluated by PRNT using VeroE6 cells infected with the SARS-CoV-2. MD65 displayed the highest neutralization potency with a PRNT50 concentration of 0.22 μg/ml.
4A8
Chi et al. identified three neutralizing mAbs from 10 convalescent COVID-19 patients [74]. Among these mAbs, 4A8 exhibits high neutralization potency against authentic SARS-CoV-2. Interestingly, however, 4A8 does not bind the RBD. Cryo-EM was used to determine the structure of 4A8 in complex with the S protein, revealing that its epitope is located in the NTD of S protein, and that the Ab binds to S1 with Kd of 92.7 nM. 4A8 exhibits moderate neutralizing capacity, with an EC50 of 0.61 μg/ml, but it does not block the binding of S protein to the ACE2 receptor. Thus, 4A8 functions via a mechanism that is independent of receptor binding inhibition. According to the structure of the complex, the mechanisms of neutralization may involve restraining conformational changes in S protein.
RBD-chAb-1, 15, 25, 28, 45 and 51
In a recent study, a panel of Abs against the SARS-CoV-2 RBD were generated from mouse hybridoma Ab screening and were engineered into human immunoglobulin G (IgG)1 chimeric Abs [50]. Among these Abs, six potent nAbs, RBD-chAb-1, 15, 25, 28, 45, and 51, were found to bind the RBD of SARS-CoV-2 with high affinities (KD values lower than 6.5 × 10–9 M) and high neutralizing activities (PRNT50 values lower than 10 ng/ml). Experiments using site-directed mutagenesis and competition-binding assays further indicated that these six chAbs bind to three distinct epitopes within the RBM. Cryo-EM reconstruction was then used to show that the epitopes of two highly potent Abs, RBD-chAb-25 and 45, are on one side of the open (up) RBD. This structural analysis suggested that RBD-chAb-25 and 45 can simultaneously bind to the same RBD, and the simultaneous binding was confirmed by size-exclusion chromatography. Importantly, the prophylactic effects of these Abs were demonstrated in an AAV-hACE2 mouse model and a hamster model, and the cocktail of RBD-chAb-25 and 45 showed highly promising therapeutic effects [50]. Notably, several antibody cocktails showed low IC50 values (3.35–27.06 ng/ml) against the SARS-CoV-2 variant pseudoviruses including Alpha, Beta, Gamma, Epsilon, Iota, Kappa and Delta variants [75]. Furthermore, the therapeutic treatment with an antibody cocktail of RBD-chAb-15 and 45 effectively protected hamsters from infection with the Delta SARS-CoV-2 variant [75].
Yang et al. further identified a unique salt bridge switch involving the B.1.1.7 (Alpha)-specific A570D mutation. The RBD-up state is stabilized by a double salt bridge involving A570D-K854 and D571-K964. Thus, introduction of the A570D mutation to S protein with D614G should lead to increased sensitivity of the virus to three RBD-up-specific Abs. Furthermore, the combined use of RBD-chAb-15 and 45, which simultaneously bind to distinct regions of the RBD, is also an attractive strategy for a prophylactic cocktail to prevent mutational viral escape [76].
EUA anti-SARS-CoV-2 therapeutic Abs
As of December 2021, the number of mAbs targeting S protein that were under evaluation in clinical trials was 25 (Table 2). At least 27 countries and 274 companies/institutions are developing Ab therapeutics [77], and these Abs have been comprehensively described in several review papers [33, 55, 56, 77–80]. Up to now, only seven Abs, including bamlanivimab, etesevimab, casirivimab, imdevimab, sotrovimab, cilgavimab and tixagevimab have been approved or received EUAs from the U.S. FDA (Table 2). In the following paragraphs, we introduce and update information regarding the development of these Ab treatments.
Table 2.
Clinical studies evaluating anti-SARS-CoV-2 mAbs
| No. | Name | Start date | Latest Status | Developer | Country | References |
|---|---|---|---|---|---|---|
| 1 | LY-CoV555 (Bamlanivimab) | 5/28/2020 |
EUA (11/09/2020) EUA revoked (4/9/2021) |
Eli Lilly/AbCellera | Canada/USA |
[89] |
| 2 | LY-CoV555 (Bamlanivimab) + LY-CoV016 (Etesevimab) | 6/17/2020 | EUA (2/09/2021) | Eli Lilly/AbCellera/Junshi | Canada/USA |
[94] |
| 3 |
REGN-COV2 (REGN10933/Casirivimab + REGN10987/Imdevimab) |
6/10/2020 |
EUA (11/21/2020) Approved (8/10/2021) |
Regeneron | USA | |
| 4 |
S309 (VIR-7831, Sotrovimab) |
8/27/2020 |
EUA (5/26/2021) |
Vir biotechnology/ GlaxoSmithKline | USA/UK |
[96] |
| 5 |
AZD7442 (COV2-2130/Cilgavimab + COV2-2196/Tixagevimab) |
8/17/2020 | EUA (12/08/2021) | AstraZeneca/Vanderbilt University Medical Center | UK/USA | NCT04501978, NCT04507256, NCT04625725, NCT04625972 |
| 6 | TY027 | 6/09/2020 | Phase III | Tychan Pte. LTD | Singapore |
[303] |
| 7 | BRII-196 + BRII-198 | 7/12/2020 | Phase III | Brii Bio/TSB Therapeutics/Tsinghua University | China/USA |
[250] |
| 8 | CT-P59 (Regdanvimab) | 7/18/2020 |
Phase II/III EUA (South Korea) |
Celltrion | South Korea |
NCT04525079, NCT04593641, NCT04602000 [47] |
| 9 | BI 767551 (DZIF-10c) | 11/23/2020 | Phase II/III | University of Cologne/he German Center for Infection Research/Boehringer Ingelheim | Germany |
NCT04631705, NCT04631666, NCT04822701 [253] |
| 10 | SCTA01 | 7/24/2020 | Phase II/III | Sinocelltech Ltd/Chinese Academy of Sciences | China |
[46] |
| 11 | ADG20 | 4/26/2021 | Phase II/III | Adagio Therapeutics | USA | NCT04805671, NCT04859517 |
| 12 | MAD0004J08 | 3/1/2021 | Phase II/III | Toscana Life Sciences Sviluppo s.r.l | Italia | NCT04932850, NCT04952805 |
| 13 | MW33 | 8/7/2020 | Phase II | Mabwell (Shanghai) Bioscience | China | NCT04533048 |
| 14 | DXP593 | 8/31/2020 | Phase II | Beigene | China |
[194] |
| 15 | COVI-AMG (STI-2020) | 2/2/2021 | Phase II | Sorrento Therapeutics | USA | NCT04734860 |
| 16 |
LY-CoV1404 + LY-CoV555 (Bamlanivimab) + LY-CoV016 (Etesevimab) |
11/18/2020 | Phase II | Eli Lilly/AbCellera/ Junshi | USA | NCT04634409 |
| 17 | XVR011 | 5/12/2021 | Phase I/II | Exevir Bio BV | Belgium | NCT05017168 |
| 18 |
LY-CoV016 (JS016, Etesevimab) |
6/5/2020 | Phase I | Junshi Biosciences/ Chinese Academy of Sciences/Eli Lilly | China/USA |
NCT04441918, NCT04441931, NCT04427501 [51] |
| 19 | 47D11 | 11/25/2020 | Phase I | Utrecht University/Abbvie/Erasmus MC/Harbor BioMed | Netherlands/China/USA |
NCT0464412 [40] |
| 20 | ADM03820 | 12/4/2020 | Phase II/III | Ology Bioservices | USA | NCT04592549, NCT05142527 |
| 21 | DXP604 | 12/15/2020 | Phase I | Beigene | China | NCT04669262 |
| 22 | C144-LS and C-135-LS | 1/11/2021 | Phase II/III | Bristol-Myers Squibb, Rockefeller University | USA |
[98] |
REGN-COV2 (casirivimab and imdevimab)
REGN-COV2 is a cocktail of the human Abs, casirivimab and imdevimab (formerly known as REGN10933 and REGN10987, respectively), which both target the S protein RBD but were identified by different methods [41]. Casirivimab was identified from VelocImmune hAb transgenic mice immunized with a DNA plasmid encoding SARS-CoV-2 S protein, followed by a booster of injected recombinant S protein. Meanwhile, imdevimab was identified from isolated PBMCs of three human donors previously infected with SARS-CoV-2. In both cases, the murine or human single B cells bound to S protein were sorted by FACS. The Kd values of casirivimab and imdevimab for S protein are both about 0.04 nM by measurement with Biacore T200. The PRNT50 of casirivimab and imdevimab are 0.0374 and 0.0421 nM, respectively. Novel S gene mutants rapidly appeared when virus was passaged in the presence of individual Abs, resulting in loss of neutralization. However, treatment of casirivimab and imdevimab together can prevent the selection of escape mutants in vitro since they comprise a non-competing Ab cocktail [81]. In vivo efficacy of this Ab cocktail has been evaluated in both rhesus macaques (used to model mild disease) and golden hamsters (model for more severe disease) [82]. In the rhesus macaques, REGN-COV2 greatly reduced virus load in the lower and upper airways and decreased virus-induced pathological sequelae when administered prophylactically (50 mg/kg dosage) or therapeutically (25 mg/kg dosage). Administration in hamsters (5 mg/kg dosage) inhibited weight loss and reduced viral titers in the lung.
Four separate large clinical trials are ongoing for REGN-COV2. One of the trials is a phase I–III adaptive, randomized, placebo-controlled, double-blind trial (NCT04425629) on non-hospitalized patients with COVID-19, aiming to reduce the risk of treatment-resistant mutant virus emergence [55]. Seven hundred ninety-nine patients were randomly assigned (1:1:1) to receive placebo, 2.4 g of REGN-COV2, or 8.0 g of REGN-COV2. The interim analysis showed that REGN-COV2 can indeed reduce viral load in patients. Safety outcomes were similar in the combined REGN-COV2 dose groups and the placebo group. The above results supported the EUA designation for the casirivimab and imdevimab cocktail, which was granted by the U.S. FDA on November 20, 2020 for COVID-19 therapy. Under the EUA, the recommended dose is 1.2 g of casirivimab and 1.2 g of imdevimab (2.4 g total), administered as a single intravenous infusion. The phase III data showed that the combined casirivimab and imdevimab treatment could reduce the risk of COVID-19-related hospitalization and death by 70% COVID-19 in non-hospitalized patients, and the median time of symptom duration was reduced from 14 to 10 days.
In April 2021, new data from a phase III treatment trial in recently infected asymptomatic COVID-19 patients demonstrated that subcutaneous injection of a 1.2 g total dose of REGN-COV2 (1:1, casirivimab:imdevimab) reduced the risk of progression to symptomatic COVID-19 by 31%, and the risk was reduced by 76% after the third day. Furthermore, another positive result from a phase III COVID-19 prevention trial in uninfected household contacts of SARS-CoV-2 infected individuals showed that the 1.2 g total dose of REGN-COV2 reduced the risk of symptomatic infections by 81% [83]. REGN-COV2 was granted an EUA by the U.S. FDA in December 2020 and gained full approval from Japan’s Ministry of Health, Labour and Welfare in July 2021 for the treatment of patients with mild to moderate COVID-19 [84].
As casirivimab and imdevimab were designed against the SARS-CoV-2 strains that were being transmitted at the beginning of the pandemic in 2020 [41, 81], there is some question as to the protective and therapeutic ability against newly emerged variant strains; however, the treatment remains effective or at least does not cause concern when used against new variants. Most recently, it has been reported that B.1.1.7 (Alpha) is not refractory to the neutralizing activity of casirivimab and imdevimab [53]. Notably, the B.1.351 (Beta) and P.1 (Gamma) variants are fully resistant to casirivimab and slightly resistant to the neutralization by imdevimab [53, 54]. However, the combination of casirivimab and imdevimab show prophylactic and therapeutic efficacy against SARS-CoV-2 variants including viruses with B.1.1.7 (Alpha), B.1.351 (Beta), or P.1 (Gamma) in animals [85]. With regard to the newly emerged B.1.617.2 (Delta) variant, casirivimab also exhibits reduced neutralizing ability; however, imdevimab and the cocktail of casirivimab and imdevimab can still efficiently block virus S protein entry into the host cell [86]. Moreover, according to the REGN-COV2 fact sheet authorized by the U.S. FDA, pseudovirus assays showed that the neutralizing activity of REGN-COV2 was not changed with regard to currently circulating variants, including B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), B.1.429 (Epsilon), and B.1.526 (Iota). On August 10, 2021, the U.S. FDA authorized REGN-COV2 for both treatment and post-exposure prophylaxis (prevention) of COVID-19; the approved dosage is 600 mg of casirivimab and 600 mg of imdevimab administered together [87].
In January 2021, the US government signed a contract to purchase 1.25 million doses of REGN-COV2, and is expected to pay US$2.625 billion to Regeneron ($2,100/dose). The company anticipates being able to provide at least 1 million doses by June 30, 2021 if the EUA is updated to the lower 1,200 mg dose. The European Medicines Agency (EMA) also approved the use of REGN-COV2 and stated that clinical results show that the use of REGN-COV2 treatment can reduce the amount of virus in the nose and throat of patients, thereby reducing the number of patient visits to health care providers. In January 2021, the German government purchased 200,000 doses at a price of US$488 million ($2,440/dose). In February 2021, the French government announced that it had distributed thousands of doses of REGN-COV2 to various hospitals for clinical treatment of patients. In May 2021, the governments of Belgium and Switzerland approved clinical use of REGN-COV2. Also in May, Japan completed an agreement with Roche to purchase REGN-COV2. Total sales for the first half of 2021 consisted of $4.156 billion for REGN-COV2 [88]
Bamlanivimab (LY-CoV555)
Bamlanivimab is a human IgG1 targeting the RBD of S protein. It was discovered by Eli-Lilly and AbCellera via a high-throughput microfluidic screen of antigen-specific B cells from the first U.S. patient to recover from COVID-19 [89]. In a rhesus macaque challenge model, prophylactic doses as low as 2.5 mg/kg reduced viral replication in the upper and lower respiratory tract. On May 28, 2020, a clinical trial for bamlanivimab was initiated on hospitalized patients with COVID-19, and the Ab became the world’s first SARS-CoV-2-specific Ab to be used for COVID-19 therapy.
In the phase II trial of Blocking Viral Attachment and Cell Entry with SARS-CoV-2 Neutralizing Antibodies (BLAZE-1; NCT04427501), 452 patients with mild to moderate COVID-19 were randomly assigned to receive a single intravenous infusion of bamlanivimab at one of three doses (700 mg, 2800 mg, or 7000 mg) or placebo; patients were evaluated for quantitative virologic endpoints and clinical outcomes [57]. Those patients treated with bamlanivimab showed reduced viral load and lower rates of symptoms and hospitalization. Based on data from the BLAZE-1 study, the U.S. FDA granted an EUA for a single infusion of 700 mg bamlanivimab for the treatment of mild to moderate COVID-19 in adults and pediatric patients on November 9, 2020 [56]. Thus, bamlanivimab was the first SARS-CoV-2-nAb authorized for clinical use. Eli Lilly has an agreement with the U.S. government to supply 300,000 vials of 700 mg doses of bamlanivimab for US$375 million ($1250/dose) [90]. According to Eli Lilly, the company plans to donate COVID-19 therapies to Direct Relief for use in low- and lower-middle-income countries, which have been heavily impacted by the pandemic.
There is some concern that while bamlanivimab activity is unaffected against the B.1.1.7 (Alpha) variant strain, its protective efficacy is lost against the B.1.351 (Beta), P.1 (Gamma), and B.1.617.2 (Delta) variants, due to the E484 mutation [53, 54, 85, 86]. The use of a yeast display library to comprehensively map mutations in the RBD that allow SARS-CoV-2 to escape Ab binding [91] revealed that the L452R mutation in the B.1.429 (Epsilon) lineage allows escape from bamlanivimab [92]. Because emerging data shows that common SARS-CoV-2 viral variants are resistant to bamlanivimab alone, the U.S. FDA revoked the EUA that allowed for bamlanivimab to be used as a monotherapy of COVID-19 patients on April 9, 2021.
Combination of bamlanivimab with etesevimab
Etesevimab (CB6, JS016, LY-CoV016) was identified by screening single B cells from a convalescent patient [51]. X-ray crystallography revealed that its epitope on SARS-CoV-2 RBD largely overlaps with ACE2 binding residues. To reduce the potential risk of an Ab-dependent enhancement (ADE) [93] and effector functions, the Fc of etesevimab was modified by two leucine-to-alanine substitutions at residues 234 and 235 (known as the LALA mutation), which abolished its affinity for the Fcγ receptor. In rhesus monkey models, treatment with etesevimab inhibited viral titers and reduced lung damage under both prophylactic and therapeutic usages. Etesevimab has been evaluated in a completed phase I clinical trial (NCT04441931) and a phase II/III study in combination with bamlanivimab (NCT04427501).
On January 26, 2021, Eli Lilly announced that the combination of bamlanivimab (2.8 g) and etesevimab (2.8 g) significantly reduced hospitalizations and deaths in high-risk patients recently diagnosed with COVID-19, reaching the primary endpoint of the Phase III BLAZE-1 trial (NCT04427501). In the 1035 patients enrolled in this trail, the treatment reduced hospitalizations and death by 70%. There were 10 deaths in total, all of which occurred in patients taking placebo, and no deaths were recorded in patients taking bamlanivimab and etesevimab together. The Phase III BLAZE-1 trial showed additional results to demonstrate combination of bamlanivimab 700 mg and etesevimab 1400 mg reduced the risk of COVID-19 related hospitalizations and deaths by 87% in high-risk patients aged 12 and older and recently diagnosed with the virus. The data were from 769 high-risk patients with mild to moderate COVID-19. Of those patients, 511 were randomly assigned to treatment with Ab cocktail, and the other 258 were assigned to placebo. The primary endpoint was percentage of participants who experience COVID-related hospitalizations or death from any cause by day 29.
Based on the BLAZE-1 trial, the U.S. FDA issued an EUA for combined bamlanivimab (700 mg) and etesevimab (1400 mg) for the treatment of mild to moderate COVID-19 in patients of at least 12 years old who weigh at least 40 kg and are at high risk of progressing to severe disease and/or hospitalization. This combination therapy is expected to reduce the risk of selecting for resistant viruses when compared to bamlanivimab administered alone [94]. While the combination of bamlanivimab and etesevimab can neutralize B.1.1.7 (Alpha), it is not protective against B.1.351 (Beta) and P.1 (Gamma) variants because of the K417N/T mutation [53, 54]. Regarding the newly emerged B.1.617.2 (Delta) variant, bamlanivimab loses neutralizing ability due to the E484Q mutation, whereas etesevimab is not influenced by this mutation and still retains neutralizing ability. Therefore, the cocktail of bamlanivimab and etesevimab has partially reduced ability to inhibit B.1.617.2 (Delta) variant [86].
The U.S. government agreed to purchase up to 1.2 million doses of bamlanivimab and etesevimab together by November 2021. One hundred thousand doses have been ordered for shipment by March 31 at a value of US$210 million ($2,100 USD/dose). According to Eli Lilly's financial report for the first quarter of 2021, bamlanivimab and etesevimab had global sales of US$810 million, ranking first among all product lines. In May 2021, Eli Lilly plans to provide bamlanivimab and etesevimab to low- and middle-income countries free of charge. Bamlanivimab and etesevimab has begun to be used in India, and the first Indian patient treated with this Ab cocktail was discharged from the hospital in Haryana on May 26, 2021. The Medanta hospital in India reported that the cocktail is also effective against B.1.617.2 (Delta) variant and that the price of each dose is US$815. Lilly reported that total sales for the first half of 2021 consisted of $959.1 million for bamlanivimab and etesevimab administered together [88]. However, results from in vitro assays show that bamlanivimab and etesevimab administered together are not active against either the P.1 (Gamma) or B.1.351 (Beta) variants. Therefore, the U.S. Department of Health and Human Services paused all distribution of etesevimab alone, and bamlanivimab and etesevimab together on June 25, 2021 [95].
Sotrovimab (VIR-7831, S309)
Sotrovimab is a derivative of the S309 mAb, which was engineered with an extended half-life and potentially improved biodistribution in the lungs by the introduction of a LS mutation in the Fc [96]. S309 was originally identified from memory B cells of an individual with SARS-CoV infection in 2003; this Ab was found to potently cross-neutralize authentic SARS-CoV-2 [97]. Cryo-EM analysis revealed that S309 can bind to the “up” and “down” states of the RBD in a single S trimer. However, the Fab engages an epitope distinct from the RBM and does not compete with ACE2 upon binding to S glycoprotein. It was proposed that the mechanism of S309-mediated neutralization may be the induction of S trimer cross-linking, steric hindrance, or aggregation of virions. S309 also showed strong Ab-dependent cell cytotoxicity and Ab-dependent cellular phagocytosis effector functions. The Fc-effector function was demonstrated to contribute to the neutralization of SARS-CoV-2 in mouse models [98].
A phase III COVID-19 mAb Efficacy Trial (COMET-ICE) evaluated sotrovimab (0.5 g, intravenous injection) as a monotherapy for the early treatment of COVID-19 in adults at high risk of hospitalization. The study was stopped early in March 2021 due to clear evidence of clinical efficacy. Interim study results demonstrated an 85% reduction in the primary endpoint of hospitalizations (more than 24 h) or death for those receiving sotrovimab (n = 291) compared to placebo (n = 292). On May 26, 2021, the U.S. FDA issued an EUA for the 0.5 g single-dose of sotrovimab for the treatment of mild-to-moderate COVID-19 in pediatric patients (12 years of age and older) who are at high risk for progression to severe COVID-19. In vitro testing showed that sotrovimab retains activity against currently circulating variants, including P.1 (Gamma), B.1.429 (Epsilon), B.1.526 (Iota) and B.1.617.2 (Delta) [53, 54, 99].
AZD7442 (tixagevimab and cilgavimab)
AZD7442 is the combination of two human mAbs initially isolated from convalescent patients after SARS-CoV-2 infection and later engineered to be long-acting IgG molecules. The mAbs, COV2-2130 (AZD1061/cilgavimab) and COV2-2196 (AZD8895/tixagevimab), recognize and simultaneously bind to two distinct non-overlapping epitopes on the virus RBD in the “up” configuration [100, 101]. COV2-2130 and COV2-2196 both have neutralizing abilities, with IC50 values of 1.6 ng/mL and 0.7 ng/mL in pseudovirus assays, and IC50 values of 107 ng/mL and 15 ng/mL in FRNT, respectively (Table 1). Furthermore, a dose of 50 mg/kg showed a major protective effect in Rhesus macaques, with no subgenomic viral RNA detected in the treated group. By contrast, the isotype control mAb group had high levels of subgenomic viral RNA after exposure to SARS-CoV-2. In a mouse experiment to evaluate the therapeutic effects of the combination, 80% of treated mice had undetectable levels of infectious virus in lung after receiving the most effective dose of approximately 20 mg/kg [100]. AstraZeneca licensed the combination in June 2020, and the mAbs were then further optimized by modifying amino acid residues in the Fc region [102]. First, L234F/L235E/P331S substitutions in the Fc region mitigate the potential risk of FcγR and complement binding [103]. Second, M252Y/S254T/T256E substitutions were made to increase the affinity for human FcRn at low endosomal pH, extending the half-lives of the mAbs [104]. After optimization, a single dose of AZD7442 was shown to provide protection against COVID-19 for 6 to 12 months [102].
On 15 June 2021, AstraZeneca announced results from a phase III trial (STORM CHASER) assessing the safety and efficacy of AZD7442 for the prevention of symptomatic COVID-19 in participants recently exposed to the SARS-CoV-2. AZD7442 reduced the risk of developing symptomatic COVID-19 by 33% compared to placebo, which did not meet the primary endpoint. However, other phase III trials PROVENT and TACKLE are still ongoing and will evaluate the respective efficacies of AZD7442 for pre-exposure prevention and preventing severe disease. Most recently, it has been reported that the combination of COV2-2130 and COV2-2196 can neutralize SARS-CoV-2 variants, including B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), B.1429 (Epsilon), B.1617.1, or B.1526 (Iota), in vitro. From the analysis of prophylactic and therapeutic efficacies against B.1.1.7 (Alpha), B.1.351 (Beta), or P.1 (Gamma) in animals, AZD7442 showed promising results [105]. In November 2021, new data from two phase III trials testing AZD7442 for prophylaxis and post-exposure prophylaxis were released (Table 2). The 6-month follow-up of the prevention trial showed that one 300 mg IM (intramuscular injection) dose of AZD7442 reduced risk of symptomatic COVID-19 by 83%, with no severe disease or deaths observed. The separate treatment trial showed 88% reduced risk of severe COVID-19 or death when treatments were given within three days of symptom onset [106]. Based on this progress, AstraZeneca has already signed an agreement with the U.S. government to supply up to 500,000 doses of AZD7442 for US$205 million ($410/dose), contingent on AZD7442 receiving EUA in post-exposure prophylaxis [107].
Antibodies to control the cytokine storm syndrome (CSS)
Cytokine storm syndrome (CSS) or CRS is an uncontrolled systemic inflammatory response associated with highly increased levels of inflammatory cytokines responding to different triggers, including therapies, pathogens or autoimmune disease. Critical COVID-19 patients often exhibit CSS-like syndromes, such as high fever, severe pneumonia leading to ARDS, multiple organ failure, or even death. Therefore, it is reasonable to suspect that the direct effects of CSS, triggered by exaggerated levels of inflammatory cytokines, are at least partially responsible for severe COVID-19 syndrome [108]. Although the role of these inflammatory factors in treatment of COVID-19 remains unclear, effectively neutralizing the overproduced inflammatory factors in CSS is essential to reduce mortality in patients with COVID-19 [109–111]. Here, we summarize the current clinical-stage therapeutic mAbs that can target cytokines to relieve CSS in COVID-19 patients (Table 3).
Table 3.
Clinical trials of therapeutic antibodies for COVID-19
| Target & mAb drug | ClinicalTrials.gov identification | Type | Phase |
|---|---|---|---|
| Anti-IL-6 | |||
| Clazakizumab | NCT04348500, 6 trials | Humanized rabbit IgG1 mAb | II |
| Siltuximab | NCT04329650, 3 trials | Chimeric IgGκ mAb | II/III |
| Olokizumab | NCT04452474, 2 trials | Humanized IgG4 mAb | II/III |
| Anti-IL-6R | |||
| Levilimab | NCT04397562 | Human mAb | III |
| Sarilumab | NCT04661527, 9 trials | Human IgG1 mAb | I/II/III |
| Sirukumab | NCT04380961 | Human IgG1κ mAb | II |
| Tocilizumab | NCT04372186, 56 trials | Humanized mouse IgG1 mAb | EUA |
| Anti-IL-1β | |||
| Canakinumab | NCT04362813, 5 trials | Human IgG1κ mAb | III |
| Anti-TNF | |||
| Infliximab | NCT04425538, 4 trials | Chimeric IgG1 mAb | II |
| Adalimumab | NCT04705844 | Human mAb | III |
| Anti-GM-CSF | |||
| Lenzilumab | NCT04351152 | Human IgG1 mAb | III |
| Otilimab | NCT04376684 | Human IgG1 mAb | II |
| TJ003234 | NCT04341116 | Human IgG1 mAb | II/III |
| Anti-GM-CSFR | |||
| Gimsilumab | NCT04351243 | Human IgG1 mAb | II |
| Anti-GM-CSFR-α | |||
| Mavrilimumab | NCT04447469, 5 trials | Human IgG4 mAb | II/III |
| Anti-C5 | |||
| Eculizumab | NCT04346797, 4 trials | Humanized mouse IgG2/4κ mAb | II |
| Anti-C5a | |||
| Vilobelimab | NCT04333420 | Chimeric IgG4 mAb | II/III |
| Anti-C5aR | |||
| Avdoralimab | NCT04371367, 2 trials | Human IgG1 mAb | II |
| Anti-PD-1 | |||
| Nivolumab | NCT04356508, 3 trials | Human IgG4 mAb | II |
Abs targeting interleukin-6 (IL-6)
The consistent observation of high IL-6 levels in CSS patients suggests that this cytokine is a key mediator of CSS, although the mechanisms of such action have not yet been fully elucidated [112]. IL-6 is known to be essential for the adaptive immune response in which T cells and B cells are recruited to the infected site. There are two main pathways of IL-6 signaling transduction, referred to as classic cis or trans signaling. In classic cis signaling, IL-6 and gp130 form a complex with membrane-bound IL-6 receptor (mIL-6R), while in the trans pathway, they bind to the soluble form of IL-6 receptor (sIL-6R). In either case, the IL-6 receptor (IL-6R) signaling complex activates intercellular signaling involved in a wide range of biological functions, such as immune regulation through downstream JAK-STAT3 signaling [113]. Importantly, elevated IL-6 level has been tightly associated with ARDS and high mortality of COVID-19 patients; therefore, IL-6 is thought to be a promising therapeutic target to reduce hyper inflammation and prevent the high mortalities of COVID-19 [112, 114–116]. According to the key role of IL-6 in CSS, several mAb drugs have been considered for the treatment of severe COVID-19, including sarilumab (Kevzara), tocilizumab (Actemra) and levilimab, which target IL-6R, as well as clazakizumab, siltuximab and olokizumab, which target IL-6 [19, 20, 117, 118].
These Abs specifically bind to both mIL-6R and sIL-6R and inhibit both cis and trans signal transduction. Several reports suggested that critically ill patients with COVID-19 who received tocilizumab or sarilumab had improved outcomes and lower rates of mortality [119, 120]. However, other studies on the efficacy of tocilizumab or sarilumab have shown conflicting results, as the drugs failed to reduce the risk of intubation or death in patients with COVID-19 in several clinical trials [118, 121–124]. Despite these inconclusive results, the U.S. FDA granted authorization for the emergency use of tocilizumab to treat patients hospitalized with COVID19 on June 24, 2021; the decision was based on the findings from a large clinical trial on tocilizumab [125, 126]. The EUA is specifically for treating certain hospitalized patients who are already receiving corticosteroids and need breathing support, but the drug is not approved as a general treatment for COVID-19. In the clinical trials on critically ill patients with COVID-19 in the intensive care unit, both tocilizumab and sarilumab improved survival [119, 127]. Furthermore, in clinical trials on hospitalized patients, tocilizumab used for the treatment of COVID-19 reduced the risk of death within 28 days by an absolute difference of 4% compared with usual care; this result was from patients with COVID-19 who required oxygen and had evidence of inflammation. Tocilizumab also reduced the time that patients remained in the hospital, and the probability of patient discharge within 28 days was raised from 50 to 57% (p < 0.0001) [125]. This trial provided the most definitive evidence that treatment with tocilizumab benefits hospitalized COVID-19 patients [120]. In addition, the WHO has recommended the use of tocilizumab and sarilumab plus corticosteroids to treat severe COVID-19 [127].
Targeting TNF
TNF is an important cytokine in many inflammatory diseases, and it is known to regulate IL-6 expression. In contrast to anti-IL-6 therapy, anti-TNF therapy has been shown to downregulate several inflammatory cytokines including IL-1, IL-6, and GM-CSF [128, 129]. Moreover, elevated levels of TNF in the blood and tissues of patients with COVID-19 have been indicated in previous reports [130]. Since blocking IL-6 met with limited success in COVID-19 patients, anti-TNF therapy has been recently considered as a means of reducing inflammation in COVID-19 [21, 131]. Early observations from clinical data support the idea that anti-TNF Abs, such as infliximab or adalimumab may reduce the mortality rate in patients with COVID-19 [132, 133]. Up to now, there have been four clinical trials on infliximab (NCT04344249, NCT04425538, NCT04593940, NCT04734678) and one on adalimumab (NCT04705844), all of which seek to evaluate their therapeutic potential in COVID-19.
Targeting IL-1β
There are three important cytokines in the IL-1 family that are especially relevant to cytokine storms: IL-1β, IL-18, and IL-33; among these cytokines, blocking IL-1β has great potential to counteract cytokine storms [22]. The IL-1 family members play different pro-inflammatory roles in patients with COVID-19, and these individual cytokines may be important mediators of many CSS symptoms, including fever, edema, and finally, organ dysfunction or death. Thus, blocking their function may possibly reverse the cytokine storm. Though the exact roles of IL-1 cytokines in the pathogenesis of CSS are unclear, it seems that IL-1 receptor blockade may help to maintain better control of inflammatory processes. Canakinumab is a human mAb that neutralizes IL-1β bioactivity by competing for IL-1RI binding; it is approved for the treatment of cryopyrin-associated periodic syndromes and several serious auto-inflammatory diseases [134, 135]. Clinical studies have been performed to examine the efficacy and safety of canakinumab in patients with COVID-19 [136, 137].
Others
Besides IL-1β, IL-6, and TNF, several cytokine storm-related factors are potential therapeutic targets for the treatment of severe COVID-19 patients. For example, GM-CSF is often found at a high level in COVID-19 patients. GM-CSF binding to GM-CSF receptor-α (GM-CSFR-α) stimulates IL-1, IL-6, and TNF production, promoting downstream Janus kinase 2 (JAK2) signal transduction [138]. Mavrilimumab is a human mAb targeted to GM‐CSFR-α that has been used as an investigational drug for the treatment of rheumatoid arthritis [139]. Recently, clinical data suggest that the condition of COVID-19 patients with pneumonia and systemic hyper inflammation can be improved by treatment of mavrilimumab and lenzilumab [23, 140, 141]. These results showed that therapeutic antibodies against GM-CSF can improve the clinical outcomes for COVID-19 patients with CSS.
In addition to GM-CSF, the complement system may be a valuable target for COVID-19 therapy, as it is an integral component of the innate immune response to virus infection. Complement signaling comprises three known axes, including the classical complement, alternative complement, and lectin pathways. All three pathways converge on the main component C3 of the complement pathway and result in the production of proinflammatory anaphylatoxins, C3a and C5a, and the formation of the terminal membrane attack complex (MAC) [142]. Patients with severe COVID-19 showed complement activation and high concentrations of C5a and MAC, suggesting that dysregulation of the complement pathway may participate in CSS and severe COVID-19 complications [143–147]. Notably, mechanistic studies showed the S or nucleocapsid protein of SARS-CoV-2 can activate the complement pathway [148, 149]. Based on the apparent involvement of complement in COVID-19, clinical studies have been initiated for several Abs, including avdoralimab, eculizumab, and vilobelimab (Table 3). Eculizumab is a humanized mAb with a high affinity to C5 that inhibits the generation of C5a and C5b proteins and prevents the formation of the inflammatory anaphylatoxin and the MAC [150]. In addition, avdoralimab and vilobelimab are mAbs targeting C5aR or C5a that prevent binding of C5a to C5aR and block the formation of the inflammatory anaphylatoxin associated with pulmonary pathology of ARDS in COVID-19 [145, 151]. Conceivably, these therapeutic antibodies could be effective treatments for severe COVID-19 with CSS.
Antibody-based SARS-CoV-2 detection
As the number of patients with COVID-19 continues to grow around the world, a major issue is monitoring and evaluating patients with diagnostic tests that can distinguish SARS-CoV-2 from other viruses causing common cold symptoms [152]. Tests for viral nucleic acids and antigens can specifically indicate the presence of the virus in patients during the acute phase of virus infection [153]. Moreover, the diagnostic sensitivity of each test varies depending on the duration of disease, viral load, and quality of specimen collection, in addition to the collection site [154]. Because SARS-CoV-2 mainly replicates in the respiratory tract, the U.S. Centers for Disease Control and Prevention (CDC) recommends collecting and analyzing patient specimens from the upper and lower respiratory tract [155]. The nucleic acid detection assays show better sensitivity when used on specimens from the lower respiratory tract, including bronchoalveolar lavage fluid and sputum, than for specimens from the upper respiratory tract [156]. Among upper respiratory specimens, swabs collected from the nasal cavity yield a higher detection rate than oropharyngeal swabs [157].
In one of the main nucleic acid amplification tests, RT-PCR is used to amplify a unique viral genome sequence with specific primers; this assay offers a high accuracy and was the first method developed for SARS-CoV-2 detection, making it the gold standard [158]. The CDCs from several different countries have provided RT-PCR protocols using oligonucleotide primers and probes that are complementary to several regions of the SARS-CoV-2 genome, including N, E, replicase ORF1a and ORF1b [159]. However, RT-PCR is time-consuming and requires trained personnel with specialized equipment in the laboratory. Therefore, rapid and sensitive point-of-care testing (POCT) assays have been developed, including many based on lateral flow immunoassay (LFIA).
In contrast to detection of the virus, COVID-19 serology tests detect Abs that are produced as part of the human immune response to antigen from the pathogen. Seroconversion for immunoglobulin M (IgM) and IgG may occur simultaneously or sequentially [160]. For COVID-19, seroconversion of IgM and IgG are observed an average of 13 days after onset of symptoms [160]. Serology tests using whole blood, plasma, or serum-containing abundant immunoglobulins can reveal a patient’s medical history after infection, which is useful for demonstrating Ab kinetics or assessing vaccine effectiveness [161]. One study analyzed millions of individuals diagnosed with COVID-19, showing that people aged 65 or older had higher rates of reinfection with SARS-CoV-2 [162]. Especially with regard to such vulnerable populations, serology tests can be applied to identify pre-asymptomatic individuals with SARS-CoV-2 reinfection, control transmission when used in contact tracing, and allow for repeat testing in disease screening.
Immunoassays based on antigen-Ab interactions include enzyme-linked immunosorbent assay (ELISA), chemiluminescence immunoassay (CLIA), and LFIA; these assays are widely applied for detection of specific antigens or Abs related to infectious agents [163]. The major antigens used for serology tests are purified N and S proteins, which can be applied alone or in combination to generate immunoassays that broadly detect different isotypes of Ab (Table 4). The nucleoprotein (NP) binds and packs the viral RNA genome into a helical nucleocapsid for viral replication [164]. Meanwhile, the S protein plays a significant role in viral fusion and entry into host cells and is composed of S1 RBD at N-terminus and S2 subunits at C-terminus [165]. Previous reports indicated that both NP and S protein are immunogenic, as Abs against NP and the RBD of S protein as well as their B cell epitopes were readily detected upon early seroconversion in COVID-19 patients [166–169]. ELISAs using NP and RBD of S protein show high specificity and no cross-reactivity with non-CoV, HCoV, MERS-CoV, or SARS-CoV [170, 171]. Furthermore, a meta-analysis identified 38 studies that showed the use of RBD as an antigen provides higher sensitivity than NP [172]. In addition, the presence of RBD-specific Abs is also highly associated with COVID-19 nAb response [173–176].
Table 4.
Serology tests with EUA from the U.S. FDA
| Name | Company | Source | Target | Accuracy | Method |
|---|---|---|---|---|---|
| Alinity i SARS-CoV-2 IgG | Abbott | IgG | NP |
IgG Sens 0–7 days: 49.3% 8–13 days: 80.4% ≥ 14 days: 98.1%, IgG Spec: 99.6% |
CLIA |
| Architect SARS-CoV-2 IgG | Abbott | IgG | NP |
IgG Sens 0–7 days: 49.3% 8–14 days: 82.6% ≥ 15 days: 98.1%, IgG Spec: 99.6% |
CLIA |
| AdviseDx SARS-CoV-2 IgM (Architect) | Abbott | IgM | S |
IgM Sens 0–7 days: 42.6% 8–14 days: 79% ≥ 15 days: 95%, IgM Spec: 99.6% |
CLIA |
| Babson Diagnostics aC19G1 | Babson Diagnostics, Inc | IgG | S |
IgG Sens 8–14 days: 66.7% ≥ 15 days: 100% IgG Spec: 100% |
CLIA |
| Access SARS-CoV-2 IgG | Beckman Coulter, Inc | IgG | S |
IgG Sens 0–7 days: 75.8% 8–14 days: 95.3% ≥ 15 days: 96.8%, IgG Spec: 99.6% |
CLIA |
| Access SARS-CoV-2 IgM | Beckman Coulter, Inc | IgM | S |
IgM Sens 0–7 days: 54.4% 8–14 days: 91.7% ≥ 15 days: 98.3%, IgM Spec: 99.9% |
CLIA |
| SARS-CoV-2 IgG and IgM Combo Test | BioCheck, Inc | IgM, IgG | S |
IgM Sens 0–7 days: 100% 8–14 days: 93.8% ≥ 15 days: 88.9%, IgM Spec: 97.2% IgG Sens 0–7 days: 100% 8–14 days: 100% ≥ 15 days: 100%, IgG Spec: 100% |
CLIA |
| SARS-CoV-2 IgG Antibody Test Kit | BioCheck, Inc | IgG | S |
IgG Sens 0–7 days: 100% 8–14 days: 100% ≥ 15 days: 100%, IgG Spec: 100% |
CLIA |
| SARS-CoV-2 IgM Antibody Test Kit | BioCheck, Inc | IgM | S |
IgM Sens 0–7 days: 100% 8–14 days: 93.8% ≥ 15 days: 88.9%, IgM Spec: 97.2% |
CLIA |
| LIAISON SARS-CoV-2 IgM Assay | DiaSorin, Inc | IgM | S |
IgM Sens 0–7 days: 64.4% 8–14 days: 90.2% ≥ 15 days: 92.6%, IgM Spec: 99.3% |
CLIA |
| LIAISON SARS-CoV-2 S1/S2 IgG | DiaSorin, Inc | IgG | S |
IgG Sens 0–5 days: 25% 6–14 days: 89.8% ≥ 15 days: 97.55%, IgG Spec: 99.3% |
CLIA |
| LIAISON SARS-CoV-2 TrimericS IgG | DiaSorin, Inc | IgG | S |
IgG Sens 0–7 days: 21.4% 8–14 days: 70.8% ≥ 15 days: 96.9%, IgG Spec: 99.5% |
CLIA |
| DZ-Lite SARS-CoV-2 IgG CLIA Kit | Diazyme Laboratories, Inc | IgG | S, NP |
IgG Sens 0–7 days: 43.5% 8–14 days: 91.7% ≥ 15 days: 100%, IgG Spec: 97.4% |
CLIA |
| DZ-Lite SARS-CoV-2 IgM CLIA Kit | Diazyme Laboratories, Inc | IgM | S, NP |
IgG Sens 0–7 days: 26.1% 8–14 days: 83.8% ≥ 15 days: 94.4%, IgG Spec: 98.3% |
CLIA |
| QUANTA Flash SARS-CoV-2 IgG | Inova Diagnostics, Inc | IgG | S, NP |
IgG Sens 0–7 days: 66.7% 8–14 days: 61.5% ≥ 15 days: 100%, IgG Spec: 99.9% |
CLIA |
| VITROS Anti-SARS-CoV-2 IgG test | Ortho-Clinical Diagnostics, Inc | IgG | S |
IgG Sens 12–15 days: 83.3% ≥ 16 days: 90%, IgG Spec: 100% |
CLIA |
| VITROS Immunodiagnostic Products Anti-SARS-CoV-2 Total Reagent | Ortho-Clinical Diagnostics, Inc | Pan-Ig | S |
Pan-Ig Sens 0–7 days: 80% ≥ 8 days: 100%, Pan-Ig Spec: 100% |
CLIA |
| Q-Plex SARS-CoV-2 Human IgG (4 Plex) | Quansys Biosciences, Inc | IgG | S |
IgG Sens 0–7 days: 100% 8–14 days: 100% ≥ 15 days: 95.2%, IgG Spec: 99.7% |
CLIA |
| MAGLUMI 2019-nCoV IgM/IgG | Shenzhen New Industries Biomedical Engineering Co., Ltd | IgM, IgG | S, NP |
IgM Sens 0–7 days: 43.8% 8–14 days: 78.3% ≥ 15 days: 77.5%, IgM Spec: 99.6% IgG Sens 0–7 days: 31.3% 8–14 days: 90.6% ≥ 15 days: 100%, IgG Spec: 99.1% |
CLIA |
| ADVIA Centaur SARS-CoV-2 IgG (COV2G) | Siemens Healthcare Diagnostics | IgG | S |
IgG Sens 0–6 days: 53.5% 7–13 days: 93.4% ≥ 14 days: 100%, IgG Spec: 99.9% |
CLIA |
| ADVIA Centaur SARS-CoV-2 Total (COV2T) | Siemens Healthcare Diagnostics | Pan-Ig | S |
IgG Sens 0–6 days: 61.1% 7–13 days: 97.5% ≥ 14 days: 100%, IgG Spec: 99.8% |
CLIA |
| Atellica IM SARS-CoV-2 IgG (COV2G) | Siemens Healthcare Diagnostics | IgG | S |
IgG Sens 0–6 days: 56% 7–13 days: 92.2% ≥ 14 days: 100%, IgG Spec: 99.9% |
CLIA |
| Atellica IM SARS-CoV-2 Total (COV2T) | Siemens Healthcare Diagnostics | Pan-Ig | S |
Pan-Ig Sens 0–6 days: 60.7% 7–13 days: 97.5% ≥ 14 days: 100%, Pan-Ig Spec: 99.8% |
CLIA |
| Vibrant COVID-19 Ab Assay | Vibrant America Clinical Labs | Pan-Ig | S, NP |
IgG/IgM Sens: 98.1%, IgG/IgM Spec: 98.6% |
CLIA |
| WANTAI SARS-CoV-2 Ab ELISA | Beijing Wantai Biological Pharmacy Enterprise Co., Ltd | Pan-Ig | S |
Pan-Ig Sens 0–7 days: 55.4% 8–14 days: 84.8% ≥ 15 days: 98.7%, Pan-Ig Spec: 97.5% |
ELISA |
| Platelia SARS-CoV-2 Total Ab | Bio-Rad Laboratories, Inc | Pan-Ig | NP |
Pan-Ig Sens 0–7 days: 100% 8–14 days: 96% ≥ 15 days: 100%, Pan-Ig Spec: 99.3% |
ELISA |
| SARS-CoV-2 RBD IgG test | Emory Medical Laboratories | IgG | S |
IgG Sens 0–7 days: 73% 8–14 days: 100% ≥ 15 days: 100%, IgG Spec: 97.7% |
ELISA |
| SARS-CoV-2 ELISA (IgG) | EUROIMMUN | IgG | S |
IgG Sens 0–4 days: 21.7% 5–10 days: 69.4% ≥ 11 days: 81.1%, IgG Spec: 100% |
ELISA |
| cPass SARS-CoV-2 Neutralization Antibody Detection Kit | GenScript USA Inc | Pan-Ig | S |
Pan-Ig Sens: 100%, Pan-Ig Spec: 100% |
ELISA |
| SCoV-2 Detect IgG ELISA | InBios International, Inc | IgG | S |
IgG Sens 8–14 days: 100% ≥ 15 days: 95.5%, IgG Spec: 100% |
ELISA |
| SCoV-2 Detect IgM ELISA | InBios International, Inc | IgM | S |
IgM Sens 0–7 days: 66.7% 8–14 days: 91.4% ≥ 15 days: 93.8%, IgM Spec: 98.8% |
ELISA |
| COVID-SeroKlir, Kantaro Semi-Quantitative SARS-CoV-2 IgG Antibody Kit | Kantaro Biosciences, LLC | IgG | S |
IgG Sens 0–7 days: 100% 8–14 days: 100% ≥ 15 days: 93%, IgG Spec: 99.6% |
ELISA |
| Mt. Sinai Laboratory COVID-19 ELISA Antibody Test | Mount Sinai Hospital Clinical Laboratory | IgM, IgG | S |
Combined Sens: 92.5%, Combined Spec: 100% |
ELISA |
| Simoa Semi-Quantitative SARS-CoV-2 IgG Antibody Test | Quanterix Corporation | IgG | S |
IgG Sens 0–7 days: 45.2% 8–14 days: 87.5% ≥ 15 days: 100%, IgG Spec: 99.2% |
ELISA |
| Dimension Vista SARS-CoV-2 Total Ab assay (COV2T) | Siemens Healthcare Diagnostics | Pan-Ig | S |
Pan-Ig Sens 0–6 days: 66.7% 7–13 days: 97.4% ≥ 14 days: 100%, Pan-Ig Spec: 99.8% |
ELISA |
| Dimension EXL SARS-CoV-2 Total Ab assay (CV2T) | Siemens Healthcare Diagnostics | Pan-Ig | S |
Pan-Ig Sens 0–6 days: 68.8% 7–13 days: 97.4% ≥ 14 days: 100%, Pan-Ig Spec: 99.9% |
ELISA |
| COVID-19 self-collected Ab test system | Symbiotica, Inc | IgG | S |
IgG Sens 8–14 days: 100% ≥ 15 days: 100%, Pan-Ig Spec: 98.04% |
ELISA |
| OmniPATH COVID-19 Total Antibody ELISA Test | Thermo Fisher Scientific | Pan-Ig | S |
Pan-Ig Sens 0–7 days: 19% 8–14 days: 76.7% ≥ 15 days: 100%, Pan-Ig Spec: 100% |
ELISA |
| COVID-19 ELISA pan-Ig Antibody Test | University of Arizona Genetics Core for Clinical Services | Pan-Ig | S |
Pan-Ig Sens ≥ 15 days: 97.5%, Pan-Ig Spec: 99.1% |
ELISA |
| ZEUS ELISA SARS-CoV-2 Total Test | ZEUS Scientific, Inc | Pan-Ig | S |
Pan-Ig Sens: 93.3%, Pan-Ig Spec: 100% |
ELISA |
| ZEUS ELISA SARS-CoV-2 IgG Test System | ZEUS Scientific, Inc | IgG | S |
IgG Sens 0–7 days: 100% 8–14 days: 100% ≥ 15 days: 100%, IgG Spec: 99.1% |
ELISA |
| CareStart COVID-19 IgM/IgG | Access Bio, Inc | IgM, IgG | S, NP |
IgM Sens 8–14 days: 100% ≥ 15 days: 88.7%, IgM Spec: 99.5% IgG Sens 8–14 days: 100% ≥ 15 days: 96.8%, IgG Spec: 99.5% |
LFIA |
| Assure COVID-19 IgG/IgM Rapid Test Device | Assure Tech | IgM, IgG | S, NP |
IgG/IgM Sens 0–7 days: 100% 8–14 days: 83.3% ≥ 15 days: 89.3%, IgG/IgM Spec: 100% |
LFIA |
| ACON SARS-CoV-2 IgG/IgM Rapid Test | ACON Laboratories, Inc | IgM, IgG | S, NP |
IgG/IgM Sens 0–7 days: 100% 8–14 days: 100% ≥ 15 days: 100%, IgG/IgM Spec: 95.9% |
LFIA |
| RapCov Rapid COVID-19 Test | ADVAITE, Inc | IgG | NP |
IgG Sens ≥ 15 days: 93.3%, IgG Spec: 99.5% |
LFIA |
| WANTAI SARS-CoV-2 Ab Rapid Test | Beijing Wantai Biological Pharmacy Enterprise Co., Ltd | Pan-Ig | S |
Pan-Ig Sens: 100%, Pan-Ig Spec: 98.8% |
LFIA |
| Tell Me Fast Novel Coronavirus (COVID-19) IgG/IgM Antibody Test | Biocan Diagnostics Inc | IgM, IgG | S, NP |
IgM Sens 8–14 days: 88.9% ≥ 15 days: 85.2%, IgM Spec: 98.7% IgG Sens 8–14 days: 100% ≥ 15 days: 100%, IgG Spec: 96.2% |
LFIA |
| Biohit SARS-CoV-2 IgM/IgG Antibody Test Kit | Biohit Healthcare (Hefei) | IgM, IgG | NP |
IgM Sens 0–7 days: 33.3% 8–14 days: 83% ≥ 15 days: 97.7%, IgM Spec: 99.5% IgG Sens 8–14 days: 56.6% ≥ 15 days: 96.2%, IgG Spec: 100% |
LFIA |
| qSARS-CoV-2 IgG/IgM Rapid Test | Cellex, Inc | IgM, IgG | S, NP |
Combined Sens: 93.8%, Combined Spec: 96% |
LFIA |
| COvAb SARS-CoV-2 Ab Test | Diabetomics, Inc | Pan-Ig | S |
Pan-Ig Sens 0–7 days: 41.6% 8–14 days: 84.2% ≥ 15 days: 97.6%, Pan-Ig Spec: 98.78% |
LFIA |
| RightSign COVID-19 IgG/IgM Rapid Test Cassette | Hangzhou Biotest Biotech | IgM, IgG | S |
IgG/IgM Sens 0–7 days: 66.7% 8–14 days: 100% ≥ 15 days: 88.9%, IgG/IgM Spec: 100% |
LFIA |
| LYHER Novel Coronavirus (2019-nCoV) IgM/IgG Antibody Combo | Hangzhou Laihe Biotech | IgM, IgG | S |
IgM Sens 0–6 days: 100% 7–14 days: 85.7% ≥ 15 days: 99.3%, IgM Spec: 99.4% IgG Sens 7–14 days: 76.2% ≥ 15 days: 98.5%, IgG Spec: 99.4% |
LFIA |
| COVID-19 IgG/IgM Rapid Test Cassette | Healgen Scientific, LLC | IgM, IgG | S |
IgM Sens: 100%, IgM Spec: 100% IgG Sens: 96.7%, IgG Spec: 97.5% Combined Sens: 100%, Combined Spec: 97.5% |
LFIA |
| Innovita 2019-nCoV Ab Test (Colloidal Gold) | Innovita (Tangshan) Biological Technology Co., Ltd | IgM, IgG | S, NP |
IgG/IgM Sens 0–7 days: 87.9% 8–14 days: 96.6% ≥ 15 days: 100%, IgG/IgM Spec: 98% |
LFIA |
| SCoV-2 Detect IgG Rapid Test | InBios International, Inc | IgG | S |
IgG Sens 0–7 days: 92.9% 8–14 days: 81.8% ≥ 15 days: 100%, IgG Spec: 97.7% |
LFIA |
| Orawell IgM/IgG Rapid Test | Jiangsu Well Biotech | IgM, IgG | S |
IgG/IgM Sens 8–14 days: 98.2% ≥ 15 days: 100%, IgG/IgM Spec: 98% |
LFIA |
| Rapid COVID-19 IgM/IgG Combo Test Kit | Megna Health, Inc | IgM, IgG | NP |
IgM Sens 0–7 days: 66.7% 8–14 days: 77.1% ≥ 15 days: 90.9%, IgM Spec: 99.6% IgG Sens 0–7 days: 62.3% 8–14 days: 85.7% ≥ 15 days: 90.9%, IgG Spec: 99.3% |
LFIA |
| Nirmidas COVID-19 (SARS-CoV-2) IgM/IgG Antibody Detection Kit | Nirmidas Biotech, Inc | IgM, IgG | S |
IgG Sens 0–7 days: 27.8% 8–14 days: 76.5% ≥ 15 days: 100%, IgM Sens 0–7 days: 27.8% 8–14 days: 82.4% ≥ 15 days: 97%, IgM/IgG Spec: 84.8% |
LFIA |
| ADEXUSDx COVID-19 Test | NOWDiagnostics, Inc | Pan-Ig | S |
Pan-Ig Sens: 93.3%, Pan-Ig Spec: 100% |
LFIA |
| QIAreach Anti-SARS-CoV-2 Total Test | QIAGEN, GmbH | Pan-Ig | S |
Pan-Ig Sens: 100% Pan-Ig Spec: 97.5% |
LFIA |
| Sienna-Clarity COVIBLOCK COVID-19 IgG/IgM Rapid Test Cassette | Salofa Oy | IgM, IgG | S |
IgM Sens: 90%, IgM Spec: 100% IgG Sens: 93.3%, IgG Spec: 98.8% Combined Sens: 93.3%, Combined Spec: 98.8% |
LFIA |
| SGTi-flex COVID-19 IgG | Sugentech, Inc | IgG | S, NP |
IgG Sens 0–7 days: 41.2% 8–14 days: 91.7% ≥ 15 days: 98.6%, IgG Spec: 100% |
LFIA |
| TBG SARS-CoV-2 IgG/IgM Rapid Test Kit | TBG Biotechnology Corp | IgM, IgG | S, NP |
IgM/IgG Sens ≥ 15 days: 96.4%, IgM/IgG Spec: 99.8% |
LFIA |
| BIOTIME SARS-CoV-2 IgG/IgM Rapid Qualitative Test | Xiamen Biotime Biotechnology Co., Ltd | IgM, IgG | S |
IgM Sens 0–7 days: 55.1% 8–14 days: 94.1% ≥ 15 days: 100%, IgG Sens 0–7 days: 46.4% 8–14 days: 67.7% ≥ 15 days: 100%, IgM/IgG Spec: 98.5% |
LFIA |
| VIDAS SARS-CoV-2 IgG | BioMérieux SA | IgG | S |
IgG Sens 0–7 days: 47.9% 8–14 days: 100% ≥ 15 days: 100%, IgG Spec: 99.9% |
ELFA |
| VIDAS SARS-CoV-2 IgM | BioMérieux SA | IgM | S |
IgM Sens 0–7 days: 53.8% 8–14 days: 100% ≥ 15 days: 100%, IgG Spec: 99.4% |
ELFA |
| Maverick SARS-CoV-2 Multi-Antigen Serology Panel v2 | Genalyte, Inc | Pan-Ig | S, NP |
Ig Sen 0–7 days: 66.7% 8–14 days: 90.9% ≥ 15 days: 96.1%, Ig Spec: 97.7% |
PRI |
| xMAP SARS-CoV-2 Multi-Antigen IgG Assay | Luminex Corporation | IgG | S, NP |
IgG Sens 0–7 days: 71.1% 8–14 days: 71.4% ≥ 15 days: 96.2%, IgG Spec: 100% |
FMIA |
| BioPlex 2200 SARS-CoV-2 IgG | Bio-Rad Laboratories | IgG | S |
IgG Sens 0–7 days: 81.3% 8–14 days: 96.3% ≥ 15 days: 93.9%, IgG Spec: 99.9% |
FIA |
| FREND COVID-19 total Ab | NanoEntek America, Inc | IgM, IgG | NP |
Combined Sens: 96.7%, Combined Spec: 98.8% |
FIA |
| MosaiQ COVID-19 Antibody Magazine | Quotient Suisse SA | Pan-Ig | S |
Ig Sens 0–7 days: 100% 8–14 days: 100% ≥ 15 days: 93%, Ig Spec: 99.8% |
PIA |
| Elecsys Anti-SARS-CoV-2 | Roche Diagnostics, Inc | Pan-Ig | NP |
Ig Sens 0–6 days: 60.2% 7–13 days: 85.3% ≥ 14 days: 99.5%, Ig Spec.: 99.7% |
ECLIA |
| Elecsys Anti-SARS-CoV-2 S | Roche Diagnostics, Inc | Pan-Ig | S |
Ig Sens 0–7 days: 90.6% 8–14 days: 87% ≥ 15 days: 96.6%, Ig Spec.: 100% |
ECLIA |
| New York SARS-CoV Microsphere Immunoassay for Antibody | Wadsworth Center, New York State Department of Health | Pan-Ig | NP |
Ig Sens 0–6 days: 17.9% 7–10 days: 31.3% 11–15 days: 48.9% 16–20 days: 49.2% > 20 days: 79.3%, Ig Spec: 99.6% |
MIA |
CLIA chemiluminescence immunoassay, ECLIA enzyme-enhanced chemiluminescence immunoassay, ELFA enzyme-linked fluorescence assay, FMIA fluorescent microsphere immunoassay, FIA fluorescence immunoassay, LFIA lateral flow immunoassay, MIA: magnetic immunoassay, NP nucleoprotein, PIA photometric immunoassay, PRI photonic ring immunoassay, S spike protein, Sens sensitivity (positive percent agreement), Spec specificity (negative percent agreement). The commercial kits granted EUA are updated based on the FDA. For each type of method, the products are listed in alphabetical order of the company names
In the next section, we will introduce prominent immunoassays, including ELISAs, CLIAs, and LFIAs, and comprehensively list the applications that have been granted EUA by the U.S. FDA for use as diagnostics for detection of SARS-CoV-2 and serology tests (Tables 4 and 5).
Table 5.
Antigen lateral flow assays with EUA from U.S. FDA
| Test name | Company | LoD (TCID50/ml) | Target | Device | SARS-CoV-2 accuracy information | Type of sample |
|---|---|---|---|---|---|---|
| BinaxNOW COVID-19 Ag Card Home Test | Abbott Diagnostics Scarborough, Inc | 140.6 | NP | NAVICA™ Mobile App |
Sens: 91.7% Spec: 100% |
ns |
| BinaxNOW COVID-19 Ag Card | Abbott Diagnostics Scarborough, Inc | 140.6 | NP | N/A |
Sens: 97.1% Spec: 98.5% |
ns |
| CareStart COVID-19 Antigen test | Access Bio, Inc | 800 | NP | N/A |
Sens: 88.4% Spec: 100% |
np |
| NIDS® COVID-19 Antigen Rapid Test Kit | ANP Technologies, Inc | 311 | NP | N/A |
Sens: 95.1% Spec: 97% |
ns |
| BD Veritor System for Rapid Detection of SARS-CoV-2 | Becton Dickinson & Company | 140.0 | NP | BD Veritor Plus Analyzer |
Sens: 84% Spec: 100% |
ns |
| BD Veritor™ At-Home COVID-19 Test | Becton Dickinson & Company | 187 | NP | Scanwell Health App |
Sens: 84.6% Spec: 99.8% |
ns |
| CelltrionDiaTrust™ COVID-19 Ag Rapid Test | Celltrion USA, Inc | 32 | NP, RBD | N/A |
Sens: 93.3% Spec: 99.0% |
np |
| Ellume COVID-19 Home Test | Ellume Limited | 6309 | NP | Ellume COVID-19 Home Test App |
Sens: 95% Spec: 97% |
ns |
| GenBody COVID-19 Ag | GenBody Inc | 111 | NP | N/A |
Sens: 91.1% Spec: 100% |
np |
| SCoV-2 Ag Detect Rapid Test | InBios International, Inc | 6300 | NP | N/A |
Sens: 86.7% Spec: 100% |
ns |
| iHealth COVID-19 Antigen Rapid Test | iHealth Labs, Inc | 20,000 | NP | N/A |
Sens: 94.3% Spec: 98.1% |
ns |
| Clip COVID Rapid Antigen Test | Luminostics, Inc | 88 | NP | Clip COVID Rapid Antigen Test |
Sens: 96.9% Spec: 100% |
ns |
| InteliSwab COVID-19 Rapid Test | OraSure Technologies, Inc | 250 | NP | N/A |
Sens: 84% Spec: 98% |
ns |
| InteliSwab COVID-19 Rapid Test Pro | OraSure Technologies, Inc | 250 | NP | N/A |
Sens: 84% Spec: 98% |
ns |
| Status COVID-19/Flu | Princeton BioMeditech Corp | 2700 | NP | N/A |
Sens: 93.9% Spec: 100% |
np |
| INDICAID COVID-19 Rapid Antigen Test | PHASE Scientific International, Ltd | 2800 | NP | N/A |
Sens: 84.4% Spec: 96.3% |
ns |
| QIAreach COVID-19 Rapid Antigen | QIAGEN GmbH | 50,000 | NP | QIAreach eHub |
NP: Sens: 80.7% Spec: 98.3% NS: Sens: 85% Spec: 99.1% |
np, ns |
| Sofia 2 SARS Antigen FIA | Quidel Corporation | 113 | NP | Sofia 2 |
Sens: 96.7% Spec: 100% |
np, ns |
| Sofia 2 Flu + SARS Antigen FIA | Quidel Corporation | 91.7 | NP | Sofia 2 |
Sens: 95.2% Spec: 100% |
np, ns |
| QuickVue SARS Antigen Test | Quidel Corporation | 7570 | NP | N/A |
Sens: 96.6% Spec: 99.3% |
ns |
| QuickVue At-Home COVID-19 Test | Quidel Corporation | 19,100 | NP | N/A |
Sens: 84.8% Spec: 99.1% |
ns |
| Sienna-Clarity COVID-19 Antigen Rapid Test Cassette | Salofa Oy | 1250 | NP | N/A |
Sens: 87.5% Spec: 98.9% |
np |
LoD limit of detection, NP nucleoprotein, np nasopharyngeal, ns nasal, Sens sensitivity (positive percent agreement), Spec specificity (negative percent agreement). The commercial kits granted with EUA are updated on the FDA and FIND websites. The assays are listed in alphabetical order of the company names
Enzyme-linked immunosorbent assay (ELISA)
The four main types of immunoassays include direct, indirect, sandwich, and competitive methods [177]. Most EUAs granted for Ab-based detection tests utilize the indirect ELISA strategy and probe for different human isotype immunoglobulins, such as IgG, IgM, and IgA. For example, some tests to detect virus in human serum or plasma consist of microplates coated with recombinant viral S1 protein. The interaction of antigen and Ab creates an immune-complex, which can be detected using horseradish peroxidase-conjugated Ab and tetramethylbenzidine substrate in a colorimetric reaction [178].
Chemiluminescence immunoassay (CLIA)
Indirect CLIAs use recombinant antigen-coated magnetic beads as a solid phase, which is incubated with liquid samples containing Ab to create immune-complexes. After the immune-complexes are formed, an enzyme-labelled anti-human Ab is added with the substrate to initiate a chemiluminescence reaction. The result is measured in relative light units and allows for quantification of Abs in the sample. CLIAs are conceptually similar to ELISAs but have a faster average time-to-result, i.e., 1–2 h for CLIA versus 3–5 h for ELISA [179]. In addition, Bastos et al. argued that CLIAs are generally more accurate than traditional ELISAs, according to a systematic review and meta-analysis [180].
Point-of-care rapid tests (POCTs) using lateral flow immunoassay (LFIA)
POCTs should be easy to operate, portable, and long-lasting, allowing patients to receive test reports in the care setting, rather than days later due to the need of transporting samples to a testing laboratory. POCTs LFIA can be conducted within 10 to 30 min, and it is measured by portable devices or visual observation, making it a potential application for large-scale surveys. However, LFIAs do not have signal amplification, resulting in low signal at low viral or Ab titers. Thus, in comparisons of serology tests, LFIAs often display lower sensitivity than ELISAs and CLIAs [180].
Typical LFIA formats involve capillary migration of sample through immunochromatography paper from a sample pad through a conjugation pad, where conjugated materials are released, and finally migrate to detection lines up to an absorbent pad [181]. An example LFIA with sandwich design for SARS-CoV-2 antigen detection is illustrated in Fig. 3a. NP serves as the target, and the conjugation pad contains capture Abs against NP, which are coupled to colored nanoparticles (e.g., colloidal gold or colored latex) [182]. Another detection Ab against a different epitope of NP is immobilized on the test line, and an anti-mouse IgG Ab is immobilized on the control line. When a sample containing viral NP is loaded, the anti-NP Ab on the conjugation pad captures the antigen, forming an immunocomplex. In this immunocomplex, the free epitope on NP is able to bind the second immobilized detection Ab at the test line. Unbound Abs (not in the immunocomplex) will bind to immobilized Abs on the control line, which captures the Fc of immunoglobulin. Several LFIA-based antigen detection tests for SARS-CoV-2 have been granted EUAs by the U.S. FDA (Table 5). In another example of a serology test (Fig. 3b), recombinant purified SARS-CoV-2 antigen protein (such as S protein) is labeled with nanoparticles and contained in the conjugation pad. Specific Abs from patient serum or plasma specimens are able to conjugate the labeled S protein and then bind to test lines recognizing specific isotype immunoglobulins, such as IgM, IgG, or IgA.
Fig. 3 .
Example of COVID-19 lateral flow immunoassays (LFIA). a For antigen detection, a sample containing viral antigens is dropped on the sample pad and flows by capillary action up to the absorbent pad. The sample with viral N protein (NP) directly binds to the anti-NP Ab conjugated with nanoparticles, such as colloidal gold particles or latex nanocomposites. Then, the nanoparticle-conjugated immunocomplexes are released from the conjugation pad. The free epitope of NP is captured to the second anti-NP Ab in the test line. Unbound conjugated Abs will be recognized by immobilized anti-mouse IgG in the control line. b For Ab detection, patient serum or plasma specimens are dropped on the sample pad. The sample fluid flows through the conjugation pad which contains nanoparticle-conjugated SARS-CoV-2 spike (S) or N proteins to form antigen-Ab immunocomplexes. The immunocomplexes flow to the test line and are then captured by specific isotype immunoglobulins such as IgM and IgG. Unbound control nanoparticle-conjugated mouse IgG is captured by anti-mouse IgG at the control line
Spike (S) protein structure-based Abs against SARS-CoV-2
The S protein of SARS-CoV-2 mediates host recognition and membrane fusion, so it has become a major target for the design of drugs and vaccines. S protein is a heavily glycosylated homo-trimeric membrane protein consisting of an extracellular S1 domain, an S2 domain, and a short cytosolic tail. The RBD is located at the top of the S1 domain and can fold as either an open or closed form [183, 184]; in the closed form, the RBM faces the N-terminal portion of its neighboring S1 protomer, while in the open form, it faces up. Consequently, the RBM of an open RBD can bind to the peptidase domain of ACE2, resulting in a trapping of the RBD in its open form and increased S1 subunit conformational dynamics [185]. Notably, the SARS-CoV-2 S trimer is much more sensitive than the SARS-CoV S trimer, with regard to ACE2 receptor-triggered transformation from the closed prefusion state to the fusion-prone open state; this difference might potentially account for the higher infectivity of SARS-CoV-2 compared to SARS-CoV [183, 186].
Structural biology of S protein
Since the beginning of the pandemic, development of nAbs to block virus entry has included utilization of structural biology tools like crystallography and cryo-EM single particle analysis to reveal viral protein structure and binding epitopes. To investigate the mechanisms of membrane fusion, the structures of various S protein isoforms and conformations have been solved by cryo-EM. Wrapp et al. solved the extracellular domain of S protein by cryo-EM, overcoming low S protein yield by removing the furin cleavage site and introducing two additional proline mutations for stability [63]. Another group reported the cryo-EM structure of full-length S protein, which includes the transmembrane domain and the cytosolic tail [187]. Ke and colleagues used cryo-EM tomography and single particle analysis to reconstruct the S structure directly from deactivated authentic SARS-CoV-2 virus containing the D614G mutation [184]. This structure was then superimposed with the extracellular domain structure. Despite lacking the S transmembrane domain and cytosolic tail, the cryo-EM structure for the extracellular domain showed good agreement with the structure of the S protein from the authentic virus. To gain a better understanding of the molecular conformation of the S protein during the infection process, Xu et al. solved the structure of the extracellular domain of S protein in complex with ACE2 [185]. Comparing the structure of the ACE2-S complex with the closed form of S and the one RBD up open form, the presence of ACE2 induced a swing motion in the ACE2-RBD interface and untwisting of the S trimer; as a result of this increased mobility, a missing density of the neighboring protomer fusion peptide was noted. Such a shift from a packed state to a dynamic state might make the TMPRSS2 cleavage site vulnerable to TMPRSS2 cleavage, thereby initiating the transition to postfusion and host membrane fusion [14].
Crystal structure of ACE2-RBD complex
The crystal structure of the RBD in complex with ACE2 suggests a molecular mechanism for the initial step of SARS-CoV-2 infection [188]. Between the antiparallel beta-sheet β4 and β7 of the RBD, there is an extended insertion that plays an important role influencing the residues of SARS-CoV-2 S protein that bind to ACE2; these residues are defined as the RBM and include: K417, Y453, Q474, F486, Q498, T500, and N501 (Fig. 1a). The RBDs of SARS-CoV-2 and SARS-CoV share 73% sequence similarity (Fig. 1b), and both viruses bind to ACE2 in an essentially identical manner [189, 190]. However, the RBM of SARS-CoV only shares 49% and 47% sequence similarities with wild-type and Delta SARS-CoV-2, respectively (Fig. 1b). These differences in the RBM sequence markedly increase the ACE2 binding affinity and infectivity of SARS-CoV-2 [53, 191–193]. Additionally, previous studies have shown that nAbs with overlapping epitopes can abolish binding between the RBD and ACE2 [41, 50, 72, 194]. Because the S protein on the SARS-CoV-2 membrane is the key viral component mediating receptor-binding and viral-host cell fusion, nAbs that specifically target the RBD or the RBM have garnered major attention as promising tools to block the fusion between SARS-CoV-2 and host cells.
Structure of nAb-S/RBD complexes
To summarize the recently published SARS-CoV-2-nAbs (Table 1), we defined three groups based on epitope mapping: Abs that (1) directly bind the RBM, (2) bind the RBD outside the RBM, or (3) bind S protein outside the RBD (Fig. 2). Among the reported structures for Abs in complex with SARS-CoV-2 S protein, most show S protein with 3-RBD down/closed, or 1-RBD up/open and 2-RBD down/closed. Only after stabilizing S protein with mutations do researchers see purified soluble S protein trimers with a 2-RBD up/open or all-RBD up/open conformation that could possibly lead to a more lethal SARS-CoV-2 infection [195]. Additionally, the S protein can display either RBD upward or downward conformations depending on pH. Under physiological conditions (pH 7.4), about 70% of the S protein RBDs have an upward conformation [196]. By lifting the RBD upward, a larger binding surface is made available to nAbs.
Using our classification system, the third group of nAbs may be effectively used in therapeutic combinations with Abs from the first or second groups. However, structural analyses predicting whether nAbs have overlapping epitopes showed that S protein has a dynamic nature, with movement of the NTD, RBD, S2 domain, and the stalk domain in different conformations. Thus, it may be insufficient to only examine Ab-RBD structures or even static images of the S trimer; one must also consider and test for possible simultaneous engagement of nAbs on S proteins with different combinations of up and down RBDs [184, 197]. With these data, researchers may sufficiently understand the SARS-CoV-2 S protein-Ab complexes and proceed to develop novel therapeutic measures against SARS-CoV-2.
S protein mutations
As an RNA virus, SARS-CoV-2 has a higher mutation rate than typical DNA viruses. Up to now, more than 4.45 million viral genomes have been sequenced from COVID-19-positive patients and uploaded to GISAID database (covidcg.org), and hundreds of mutations have been identified in S protein (Fig. 4a). Certain amino acid replacements might change the folding structure or conformation of a protein, potentially leading to increased virulence or evolutionary advantage. Among S protein mutants, those with the D614G point mutation are the most frequently identified in patient samples (Fig. 4a, Table 6), and this mutant has become the one of the dominant mutations of all new emerging SARS-CoV-2 variants worldwide (Fig. 4b). The mutated residue is located within the S1 domain, situated near the fusion peptide of the neighboring protomer (Fig. 1a). Cryo-EM analysis of D614G mutant S protein revealed a looser packing of the trimer structure and a more open form RBD [195]. Compared to the original SARS-CoV-2 S protein, the D614G mutation renders the S protein more stable and reduces the tendency for premature shape change [186]. Although this mutation causes weaker binding between S and the ACE2 receptor, the stability afforded by less frequent premature conformation changes makes the virus more infectious. Another mutation in S protein, A222V, occurred in the dominant D614G strain, and frequently appeared in patients from the recent second wave of the pandemic in Europe [198]. Currently, no structural analysis or patient data suggests that the protein structure of the D614G A222V S protein is different from D614G alone, nor is there evidence that the addition of the A222V mutation further increases infectivity.
Fig. 4.
SARS-CoV-2 spike mutations. a Top 200 identified SARS-CoV-2 spike mutations. Each dot indicates an amino acid mutation in the S protein. The colors indicate different domains of the SARS-CoV-2 spike protein; NTD N-terminal domain, RBD receptor-binding domain, RBM receptor-binding motif, CTD C-terminal domain, S2 subdomain 2, FP fusion peptide, TM transmembrane region. The altered amino acids of the top 20 SARS-CoV-2 spike mutations are shown as indicated. The data from 4,450,473 sequences were collected from GISAID and COVID CG (updated to 2021-11-22). b Nonsynonymous mutation positions in spike protein of newly emerged SARS-CoV-2 variants. The B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), B.1.617.2 (Delta) and B.1.1.529 (Omicron) variants are classified as variants of concern by WHO. The percentage of India B.1.617.2 (Delta) variant includes B.1.617.2 and its all AY sub-lineages. c Confirmed COVID-19 cases comprising SARS-CoV-2 variants. The data from 4,337,516 sequences were collected from GISAID and COVID CG (from 2020-12-01 to 2021-11-30) and grouped by lineage
Table 6.
Top 20 identified global SARS-CoV-2 spike protein mutations
| Mutant amino acid position | Domain of S protein | Number of sequences detected | Percentage in total cases (%) | Latest month increased (2021/10/22-11/22) |
|---|---|---|---|---|
| D614G | S1 CTD | 4,389,691 | 96.47 | 5.0 |
| L452R | RBM | 2,370,119 | 52.09 | 9.6 |
| P681R | S1 CTD | 2,330,791 | 51.22 | 9.9 |
| T478K | RBM | 2,315,734 | 50.89 | 9.9 |
| T19R | S1 NTD | 2,311,542 | 50.80 | 9.9 |
| D950N | S2 | 2,230,596 | 49.02 | 10.1 |
| EFR156G | S1 NTD | 2,178,419 | 47.87 | 10.3 |
| G142D | S1 NTD | 1,443,479 | 31.72 | 13.7 |
| N501Y | RBM | 1,103,940 | 24.26 | 0.0 |
| P681H | S1 CTD | 1,038,019 | 22.81 | 0.0 |
| HV69- | S1 NTD | 980,548 | 21.55 | 0.0 |
| T716I | S2 | 978,901 | 21.51 | 0.0 |
| A570D | S1 CTD | 972,330 | 21.37 | 0.0 |
| S982A | S2 | 971,209 | 21.34 | 0.0 |
| T95I | S1 NTD | 970,839 | 21.33 | 0.0 |
| D1118H | S2 | 970,596 | 21.33 | 1.0 |
| Y144- | S1 NTD | 960,965 | 21.12 | 13.3 |
| A222V | S1 NTD | 395,936 | 8.70 | 6.5 |
| E484K | RBM | 192,337 | 4.23 | 0.0 |
| L18F | S1 NTD | 190,391 | 4.18 | 0.5 |
S1 S1 subdomain, NTD N-terminal domain, RBD receptor-binding domain, RBM receptor-binding motif, CTD C-terminal domain, S2 subdomain 2. The data for 4,450,473 sequences from the COVID-19 started to November 22, 2021 were collected from GISAID and COVID CG
At the beginning of 2021, SARS-CoV-2 lineage B.1.1.7 (Alpha) received much attention because it is not only more transmissible than previous variants, but it also leads to increased mortality [199, 200]. Compared to patients with the original virus, B.1.1.7 (Alpha)-infected patients have higher viral loads and show less effective clearance by their immune responses [201]. It was found that this strain contains multiple mutations in the S1 NTD: deletion 69–70, deletion 144; RBM: N501Y; CTD: A570D, D614G, P681H; S2 domain: T716I, S982A, D1118H [202]. As one of the key residues of the RBD that interacts with ACE2 and nAbs (Fig. 4b) [203], mutation of N501 has been shown to increase ACE2 receptor affinity [91]. In particular, tyrosine substitution of asparagine (N501Y) was shown to not only enhance the binding affinity between S protein and ACE2, but it also increases virulence in mice [204–206]. In addition, amino acids 69 and 70 are commonly deleted from the NTD, often in combination with other mutations [207, 208], and the deletions may allosterically change the S protein conformation [63]. These deletions have been found to decrease the viral neutralization by serum from SARS-CoV-2 convalescent patients but not by serum from mRNA-1273 (Moderna)-vaccinated individuals [208, 209]. Another important site is the proline at position 681 (P681), within the furin cleavage site of S protein that exists between the receptor-binding and fusion domains [14]. Although it has not been shown to influence viral entry or transmission, the P681H mutation causes S protein cleavage to occur more efficiently [210].
Additionally, the South Africa lineage (B.1.351, Beta) includes three mutations in the RBD: K417N, E484K, and N501Y (two are in the RBM: E484K and N501Y); one in S1 NTD: D80A; one in S1 CTD: D614G and on in the S2 domain: A701V (Fig. 4b) [211]. This variant became dominant in the South African populations over the course of just a few weeks. Recently, a French research group found that the B.1.351 (Beta) variant has a significant transmission advantage over B.1.1.7 (Alpha) in some European regions [212]. Another variant of concern, the P.1 (Gamma) lineage, arose in Brazil and carries 17 unique amino acid changes, including five mutations in the S1 NTD: L18F, T20N, P26S, D138Y, R190S; three in the RBD: K417N, E484K, and N501Y (E484K and N501Y are in the RBM); one in S1 CTD: D614G; and two in the S2 domain: H655Y, T1027I (Fig. 4b) [211]. Both B.1.351 (Beta) and P.1 (Gamma) variants contain similar mutations in the RBM or RBD of S protein (K417N or K417T, E484K, N501Y), which may cause important conformational changes. The N501Y mutation is the same as the B.1.1.7 (Alpha) variant in terms of enhancing S protein binding to ACE2 and increasing virulence [204–206]. It has been found that the binding and neutralization effects of many SARS-CoV-2-nAbs can be abolished by the K417N and/or E484K mutations on S protein [206]. These effects may be due to structural changes in the receptor-binding site that prevent the interaction with nAbs. However, the binding sites of CR3022 and S309 are distant from K417 and E484, and their neutralizing abilities were unaffected by mutations at these sites [206].
The highly concerning B.1.617.2 (Delta) variant that first emerged in India was shown to be even more transmissible than the B.1.1.7 (Alpha) variant of SARS-CoV-2 (Fig. 4c) [213]. From May to July 2021, this variant spread to many countries at an alarming pace, not only affecting areas with lower vaccination rates, such as southern Africa and Asia, but also causing outbreaks in locations with high vaccination rates, such as the United Kindom and North America [213, 214]. B.1.617.2 (Delta) carries 11 unique amino acid changes in the S protein. The mutations include five in the S1 NTD (T19R, T95I, G142D, deletion 156/157, R158G), two in the RBM of RBD (L452R and T478K), two in the S1 CTD (D614G and P681R) close to the furin cleavage site, and one in the S2 domain (D950N) (Fig. 4b) [29]. It has been shown that B.1.617.2 (Delta) can totally or partially escape neutralization by many antibodies targeting the RBD or NTD of SARS-CoV-2 S protein [215, 216]. Fortunately, some antibodies with EUAs, such as REGN10933 (casirivimab), REGN10987 (imdevimab), and LY-CoV016 (Etesevimab), retain neutralization abilities against the B.1.617.2 (Delta) variant (Fig. 5). Moreover, the newly discovered potent Abs, RBD-chAb-1, 15, 28, 45, and 51, also retain neutralizing ability against the pseudovirus of B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), and B.1.617.2 (Delta) lineages of SARS-CoV-2 (Fig. 5) [50]. Therefore, the binding sites of these Abs are promising targets, as they are not subject to interference by SARS-CoV-2 mutations observed to date. Most recently, the B.1.1.529 (Omicron) variant has emerged in South Africa; it is reported to carry a large number of mutations, some of which are concerning [28]. In this variant, there are 32 mutations in the S protein, and 10 of these are within the RBM: N440K, G446S, S477N, T478K, E484A, Q493K, G496S, Q498R, N501Y and Y505H (Fig. 4b) [217]. Based on preliminary evidence that suggests an increased risk of reinfection with this variant, the WHO designated B.1.1.529 as a variant of concern on 26 November 2021 [28]. At the time of this writing, it is still unknown how effective current vaccines and nAbs are at protecting against the Omicron variant, though the topic is under intense investigation.
Fig. 5.
Neutralization of SARS-CoV-2 variants by therapeutic mAbs. The neutralization abilities of therapeutic mAbs against wild-type, D614G and newly emerged SARS-CoV-2 variants, including B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), B.1.427 and B.1.429 (Epsilon), B.1.526 (Iota), B.1.617.1 (Kappa), and B.1.617.2 (Delta). Symbols and colors indicate the range of half maximal inhibitory concentration (IC50) values toward authentic SARS-CoV-2 virus. +++ with blue, IC50 < 10 ng/ml; ++ with light blue, IC50 = 10–100 ng/ml; + with white, IC50 = 100–1000 ng/ml; —, IC50 with red > 1000 ng/ml; ND with grey, no determined; #, preliminary results reported on the website of Celltrion Healthcare Co., Ltd.; *, the range of IC50 values toward pseudotype SARS-CoV-2 virus. RBD, receptor-binding domain; RBM, receptor-binding motif. The mutant amino acids in RBD of each SARS-CoV-2 spike protein are shown as indicated
With regard to nAbs, point mutations or deletions in S protein might result in a local or global conformational change that could potentially disrupt the Ab epitope. The structures of mutated S protein-Ab complexes may reveal global conformational changes caused by the mutation and also whether the nAb can still bind to S protein in the same way as it binds to wild-type S protein. Indeed, the cryo-EM structure of D614G mutant S protein revealed that the RBD is shifted to a more open form as compared to the S protein from the original strain. Such a change in the exposed S protein surface might result in resistance to some Abs. To overcome or postpone the development of drug resistance, some strategies can be considered. The first is to map how all amino-acid mutations in the RBD affect the binding of nAbs [218]. The second approach is to further design escape-resistant Ab cocktails, which would consist of Abs that compete for binding to the same RBD surface but have different escape mutations. Indeed, recent publications have successfully demonstrated synergistic actions of nAbs in vitro. Baum et al. reported a synergistic effect with REGN10987 and REGN10933 in vitro, which was one factor allowing this combination to enter clinical trials even without animal experiments [81]. Despite their use of different neutralization assays, Wu et al. and Pinto et al. both were able to show synergistic effects in vitro for combinations of H4 + B38 and S304 + S309, respectively [72, 97]. Zost et al. further showed a synergistic effect of Abs in a mouse model with adenovirus-induced transient expression of human ACE2 in addition to their in vitro experiments [100]. In addition, Su et al., showed that the cocktail of RBD-chAb-25 and 45 not only exhibits synergistic neutralizing ability, but it is also likely to retain therapeutic potential for SARS-CoV-2 mutants [50, 75]. In addition, some recent studies showed that the accumulation of somatic mutations increased the diversity and potency of neutralizing Abs against SARS-CoV-2 [219–222]. There are also two broadly neutralizing anti-coronavirus antibodies, S2X259 and S2H97, which can neutralize SARS-CoV-2 variants of several pseudoviruses, including B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), and B.1.429 (Epsilon) [61, 62]. Therefore, the breadth and potency of nAbs may potentially be improved by using deep mutational scanning to comprehensively identify RBD mutations that lead to escape from binding by each Ab. Such improvements may help to prevent drug-resistant SARS-CoV-2 escape mutants.
Perspective
As of December 2021, the COVID-19 global pandemic has infected more than 280 million people and caused over 5.40 million deaths [223]. It is expected that the virus will continue to spread and circulate around the world for several years. Serious illnesses and deaths have been reported in all age groups and demographics. However, the percentage of COVID-19 patients that become hospitalized is less than 10% of active cases, except for populations over 65 years of age or with pre-existing diseases [224]. Thus, the majority of individuals with SARS-CoV-2 infection do not require hospitalization or need treatment, and the treatments and policy to control COVID-19 should focus on limiting transmission to reduce the medical burden.
Not only has COVID-19 caused a global health crisis, but it has also profoundly affected the global economy and financial markets [225]. Although more than 8.6 billion doses of SARS-CoV-2 vaccines have been administered around the world, no vaccine can be 100% effective at preventing symptomatic cases of COVID-19; after vaccination, some people may have no or low immune response [226], and others may experience vaccine breakthrough infection [227–229]. In addition, the uneven worldwide distribution of vaccinations, time-limited effects of immunization, and the emergence of new SARS-CoV-2 variants limit the effectiveness of vaccination as a stand-alone strategy to control the pandemic [230]. Even after global mass vaccination has been achieved, some public health measures, such as testing, tracing and isolation of patients still need to be maintained. Otherwise, new waves of infection may lead to even more morbidity and mortality.
Despite the success of COVID-19 vaccination efforts, there is still a need to provide prevention and treatment options for certain populations, including those who cannot be vaccinated or who may have an inadequate response to vaccination. Neutralizing Abs are likely to be critical tools for protecting against SARS-CoV-2 infection, and have been used successfully for this purpose in many animal studies (Table 1); the administration of passive nAbs also has preventive and therapeutic effects for SARS-CoV-2 infection [55, 57, 94]. To prevent work stoppages and alleviate work-related spread of disease, it would be beneficial to have reliable methods to prevent COVID-19 over the course of two to three weeks. Such methods could be used for short-term protection of people who need to work in COVID-19 high-risk areas. These populations can potentially be injected with preventive nAbs in advance of working as usual. There are now four Ab-based treatments authorized for emergency use in adults and teens with mild or moderate symptoms of COVID-19. Among these treatments, bamlanivimab from Eli Lily and REGN-COV2 (casirivimab and imdevimab) were the second (US$871.2 million) and fifth ($185.7 million) best-selling COVID-19 vaccines and drugs of 2020 [231]. In the first half of 2021, bamlanivimab and etesevimab (Eli Lily) and REGN-COV2 were still among the top 10 selling drugs of COVID-19, respectively earning US$959.1 million and $4.156 billion [88]. Analysts have forecasted full-year sales of about $7.0 billion for REGN-COV2 in 2021 [88, 232]. These sales figures suggest that more and more people and doctors believe nAbs have benefits in the prevention and treatment of COVID-19.
As the key determinant of host membrane fusion, S protein has become a major subject of research on SARS-CoV-2 infection mechanisms and a prime target for therapeutic Ab development. As one of only two exposed membrane proteins on SARS-CoV-2, it is also a predominant immunogen for the human immune system. A large scale survey of COVID-19 patient serum samples revealed that 96% to 98% of patient Abs recognize S protein [233], and 76% recognize the RBD in particular [234]. However, RNA viruses exhibit high mutation frequencies in the human body, and an increasing list of SARS-CoV-2 variants have been detected. To date, there have been hundreds of mutations identified in S protein (Fig. 4), and many are rapidly spreading in the population. Some of these point mutations might trigger local or global protein structure changes that enhance virulence or cause loss of efficacy for vaccines and nAbs [27, 53, 192, 235]. Therefore, Ab combination/cocktail therapies should be considered as a strategy to prevent the emergence of SARS-CoV-2 escape mutants. In addition to the REGN-COV2 cocktail Abs (casirivimab and imdevimab) [81], the combination treatment of bamlanivimab with etesevimab also reduced SARS-CoV-2 log viral load at day 11 in patients with mild to moderate COVID-19 [94]. Although it has been reported that casirivimab, bamlanivimab, and etesevimab lose neutralizing activity against B 1.351 (Alpha), and P.1 (Gamma) variants of SARS-CoV-2 [53, 54], the cocktails of Abs with non-overlapping epitopes on the RBD have been shown to exhibit great efficacy for neutralizing SARS-CoV-2 mutant escape variants [50, 76, 85, 236]. Continual development of potent nAbs against new variants of SARS-CoV-2 is therefore urgent and essential. Although no signs of ADE have been reported in human clinical trials, Liu et al. recently reported some specific Abs binding to the NTD of the open RBD might enhance virus infectivity independent of the Fc-receptor [36]. This observation is noteworthy because it means that infected or vaccinated people who generate such Abs may have a higher risk of future virus infection. Therefore, cocktails of Abs targeting multiple non-overlapping and avoiding infectivity-enhancing epitopes are a promising avenue for the development of COVID-19 therapies.
To date, enormous amounts of resources have been dedicated to studies focused on understanding the SARS-CoV-2 S protein and its RBD, to develop new tools to fight COVID-19. Many potent neutralizing mAbs have been shown to effectively inhibit virus binding to the host receptor, hACE2, both in vitro and in vivo. Cocktails of these neutralizing mAbs directed against non-competing epitopes are likely to improve the efficacy of Ab-based treatments while also preventing the emergence of SARS-CoV-2 escape mutants. Along with the ongoing vaccination efforts, easy and cheap detection systems are urgently needed to control disease spread, especially to control the unexpectedly rapid spread of escaped mutant viruses [237]. Antigen and Ab detection tests are promising candidates to fill this need and are widely used in many countries, giving the products an enormous market value [238]. The use of these products can represent a major clinical cost-saving practice, due to their low pricing, convenient and timely detection of infectious diseases with limited or no symptoms, acceleration of decisions regarding treatment or isolation, and reduction of other complications [239]. However, there has never been such a large-scale demand for these types of products, so the production and supply logistics will be key problems that need to be resolved. Assuredly, international partnerships for manufacturing and distribution, as well as new manufacturing platforms, will be required to address this pressing global need.
Conclusions
The COVID-19 pandemic is an ongoing global disaster and one of the leading causes of death in the past year, a distinction that is unprecedented in recent human history [240]. Fortunately, collaborative efforts within the worldwide scientific community have allowed the extraordinarily rapid development and authorization of vaccines and nAbs against COVID‑19. These successful efforts have greatly benefited from tremendous financial support by the governments of developed countries and the solid R&D and manufacturing capacities of pharmaceutical companies [241–244]. For example, the U.S. government has pre-ordered 100 million doses of Pfizer mRNA vaccine, BNT162b2, at a price of US$1.95 billion as early on July 22, 2020, and after one year, it has already ordered a total of 500 million doses of Pfizer mRNA vaccine. Furthermore, the U.S. government also purchased 100 million doses of mRNA-1273 of Moderna mRNA vaccine for US$2.48 billion as early on August 11, 2020, and up to July 2021, it reached to a total of 500 million doses of mRNA-1273 ordered by the U.S. government. As of February 2021, Sanofi with GlaxoSmithKline and Novavax had together received about US$2.1 billion from public and non-profit funding sources for vaccine development. Around the world, funding agencies have already paid over US$10 billion to vaccine developers [244]. In addition, the developers of therapeutic Abs, such as Regeneron, Eli Lilly and AstraZeneca, have received more than US$100 million for the production of therapeutic Abs against COVID-19 [90, 107, 245]. These extraordinary investments are one of the main reasons that hope for an end to the pandemic is beginning to shine in many countries around the world. However, the widespread emergence of highly communicable variants, such as B.1.617.2 (Delta) and B.1.1.529 (Omicron), and some uncontrolled outbreaks mean that much work remains to be done toward developing effective next-generation vaccines and medicines including therapeutic Abs for COVID-19. The advance purchase of therapeutic Abs by governments will speed the progress of pharmaceutical companies in obtaining authorizations or licensure from the FDA. Then, the clinical deployment of these therapeutic Abs can provide crucial tools for combatting SARS-CoV-2. As the emergence of variant lineages is now one of the most difficult obstacles to controlling the COVID-19 pandemic, the next generation of vaccines and therapeutic Abs must target epitopes on variants. It is especially important to predict and target epitopes with high potential to alter the transmission or infectivity of the virus, including those in recent or future emergent variants of SARS-CoV-2.
Acknowledgements
Not applicable.
Abbreviations
- Ab
Antibody
- ACE2
Angiotensin converting enzyme II
- ADE
Ab-dependent enhancement
- ARDS
Acute respiratory distress syndrome
- BLAZE-1
Blocking Viral Attachment and Cell Entry with SARS-CoV-2 Neutralizing Antibodies
- CDC
Centers for Disease Control and Prevention
- CLIA
Chemiluminescence immunoassay
- COVID-19
Coronavirus disease 2019
- CPE
Cytopathic effect
- CRS
Cytokine release syndrome
- CSS
Cytokine storm syndrome
- cryo-EM
Cryogenic electron microscopy
- ELISA
Enzyme-linked immunosorbent assay
- EUA
Emergency use authorization
- Fab
Antigen-binding fragment
- Fc
Fragment crystallizable
- FDA
Food and Drug Administration
- FP
Fusion peptide
- FPPR
Fusion peptide proximal region
- FRNT
Focus reduction neutralization assay
- GM-CSF
Granulocyte-macrophage colony-stimulating factor
- GM-CSFR-α
GM-CSF receptor-α
- IFA
Immunofluorescence assay
- IFP
Internal fusion peptide
- IgG
Immunoglobulin G
- IgM
Immunoglobulin M
- IL-1β
Interleukin 1 beta
- IL-6
Interleukin 6
- IL-6R
IL-6 receptor
- LFIA
Lateral flow immunoassay
- MAC
Membrane attack complex
- mIL-6R
Membrane-bound IL-6 receptor
- mAb
Monoclonal antibody
- NP
Nucleoprotein
- NTD
N-terminal domain
- POCT
Point-of-care testing
- PRNT
Plaque reduction neutralization test
- qPCR
Quantitative polymerase change reaction
- RBD
Receptor-binding domain
- RBM
Receptor-binding motif
- RT-PCR
Reverse transcriptase-polymerase chain reaction
- S
Spike
- SARS-CoV-2
Severe acute respiratory syndrome coronavirus 2
- scFv
Single-chain fragment variable
- sIL-6R
Soluble form of IL-6 receptor
- TMPRSS2
Transmembrane serine protease 2
- TNF
Tumor necrosis factor
Authors' contributions
Y-C H, R-M L, S-C S, P-Y C, S-H K, T-Y H and H-C W wrote the manuscript. Y-C H, R-M L, S-C S, P-Y C, S-H K, F-Y K, K-H L and T-Y H designed and illustrated figures and tables. H-C W obtained funding, provided overall direction and revised the manuscript. All authors read and approved the final manuscript.
Funding
This was supported by the funding supports from the Academia Sinica [AS-54H61] and the Ministry of Science and Technology [MOST-108-3114-Y-001-002] and [MOST 109-3114-Y-001-001], Taiwan (to H.-C. Wu).
Availability of data and materials
All the data and materials supporting the conclusions were included in the main paper.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
No potential conflicts of interest are disclosed.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yu-Chyi Hwang and Ruei-Min Lu authors contributed equally
References
- 1.WHO Coronavirus disease (COVID-2019) situation reports. World Health Organization. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports. Accessed 23 Mar 2021.
- 2.Fan E, Beitler JR, Brochard L, Calfee CS, Ferguson ND, Slutsky AS, Brodie D. COVID-19-associated acute respiratory distress syndrome: is a different approach to management warranted? Lancet Respir Med. 2020;8(8):816–821. doi: 10.1016/S2213-2600(20)30304-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bradley BT, Maioli H, Johnston R, Chaudhry I, Fink SL, Xu H, Najafian B, Deutsch G, Lacy JM, Williams T, Yarid N, Marshall DA. Histopathology and ultrastructural findings of fatal COVID-19 infections in Washington State: a case series. Lancet. 2020;396(10247):320–332. doi: 10.1016/S0140-6736(20)31305-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Struyf T, Deeks JJ, Dinnes J, Takwoingi Y, Davenport C, Leeflang MM, Spijker R, Hooft L, Emperador D, Dittrich S, Domen J, Horn SRA, Van den Bruel A, Cochrane C-DTAG. Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19 disease. Cochrane Database Syst Rev. 2020;7:13665. doi: 10.1002/14651858.CD013665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395(10223):507–513. doi: 10.1016/S0140-6736(20)30211-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA. 2020;323(11):1061–1069. doi: 10.1001/jama.2020.1585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gheblawi M, Wang K, Viveiros A, Nguyen Q, Zhong JC, Turner AJ, Raizada MK, Grant MB, Oudit GY. Angiotensin-converting enzyme 2: SARS-CoV-2 receptor and regulator of the renin-angiotensin system: celebrating the 20th anniversary of the discovery of ACE2. Circ Res. 2020;126(10):1456–1474. doi: 10.1161/CIRCRESAHA.120.317015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zhang H, Li HB, Lyu JR, Lei XM, Li W, Wu G, Lyu J, Dai ZM. Specific ACE2 expression in small intestinal enterocytes may cause gastrointestinal symptoms and injury after 2019-nCoV infection. Int J Infect Dis. 2020;96:19–24. doi: 10.1016/j.ijid.2020.04.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cheung KS, Hung IFN, Chan PPY, Lung KC, Tso E, Liu R, Ng YY, Chu MY, Chung TWH, Tam AR, Yip CCY, Leung KH, Fung AY, Zhang RR, Lin Y, Cheng HM, Zhang AJX, To KKW, Chan KH, Yuen KY, Leung WK. Gastrointestinal manifestations of SARS-CoV-2 infection and virus load in fecal samples from a Hong Kong cohort: systematic review and meta-analysis. Gastroenterology. 2020;159(1):81–95. doi: 10.1053/j.gastro.2020.03.065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Feng X, Li P, Ma L, Liang H, Lei J, Li W, Wang K, Song Y, Li S, Yang W, Yang C. Clinical characteristics and short-term outcomes of severe patients with COVID-19 in Wuhan, China. Front Med (Lausanne). 2020;7:491. doi: 10.3389/fmed.2020.00491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Lu RM, Chiu CY, Liu IJ, Chang YL, Liu YJ, Wu HC. Novel human antibody against vascular endothelial growth factor receptor 2 shows therapeutic potential for leukemia and prostate cancer. Cancer Sci. 2019;110(12):3773–3787. doi: 10.1111/cas.14208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270–273. doi: 10.1038/s41586-020-2012-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang K, Chen W, Zhang Z, Deng Y, Lian JQ, Du P, Wei D, Zhang Y, Sun XX, Gong L, Yang X, He L, Zhang L, Yang Z, Geng JJ, Chen R, Zhang H, Wang B, Zhu YM, Nan G, Jiang JL, Li L, Wu J, Lin P, Huang W, Xie L, Zheng ZH, Zhang K, Miao JL, Cui HY, Huang M, Zhang J, Fu L, Yang XM, Zhao Z, Sun S, Gu H, Wang Z, Wang CF, Lu Y, Liu YY, Wang QY, Bian H, Zhu P, Chen ZN. CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells. Signal Transduct Target Ther. 2020;5(1):283. doi: 10.1038/s41392-020-00426-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hoffmann M, Kleine-Weber H, Schroeder S, Kruger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Muller MA, Drosten C, Pohlmann S. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181(2):271–280 e278. doi: 10.1016/j.cell.2020.02.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Matsuyama S, Nao N, Shirato K, Kawase M, Saito S, Takayama I, Nagata N, Sekizuka T, Katoh H, Kato F, Sakata M, Tahara M, Kutsuna S, Ohmagari N, Kuroda M, Suzuki T, Kageyama T, Takeda M. Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc Natl Acad Sci U S A. 2020;117(13):7001–7003. doi: 10.1073/pnas.2002589117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kawai T, Akira S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat Immunol. 2010;11(5):373–384. doi: 10.1038/ni.1863. [DOI] [PubMed] [Google Scholar]
- 17.Stetson DB, Medzhitov R. Type I interferons in host defense. Immunity. 2006;25(3):373–381. doi: 10.1016/j.immuni.2006.08.007. [DOI] [PubMed] [Google Scholar]
- 18.Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395(10223):497–506. doi: 10.1016/S0140-6736(20)30183-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang C, Wu Z, Li JW, Zhao H, Wang GQ. Cytokine release syndrome in severe COVID-19: interleukin-6 receptor antagonist tocilizumab may be the key to reduce mortality. Int J Antimicrob Agents. 2020;55(5):105954. doi: 10.1016/j.ijantimicag.2020.105954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Khiali S, Rezagholizadeh A, Entezari-Maleki T. A comprehensive review on sarilumab in COVID-19. Expert Opin Biol Therapy. 2021;21(5):615–626. doi: 10.1080/14712598.2021.1847269. [DOI] [PubMed] [Google Scholar]
- 21.Robinson PC, Liew DFL, Liew JW, Monaco C, Richards D, Shivakumar S, Tanner HL, Feldmann M. The potential for repurposing anti-TNF as a therapy for the treatment of COVID-19. Med (N Y) 2020;1(1):90–102. doi: 10.1016/j.medj.2020.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Vardhana SA, Wolchok JD. The many faces of the anti-COVID immune response. J Exp Med. 2020;217(6):e20200678. doi: 10.1084/jem.20200678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.De Luca G, Cavalli G, Campochiaro C, Della-Torre E, Angelillo P, Tomelleri A, Boffini N, Tentori S, Mette F, Farina N, Rovere-Querini P, Ruggeri A, D'Aliberti T, Scarpellini P, Landoni G, De Cobelli F, Paolini JF, Zangrillo A, Tresoldi M, Trapnell BC, Ciceri F, Dagna L. GM-CSF blockade with mavrilimumab in severe COVID-19 pneumonia and systemic hyperinflammation: a single-centre, prospective cohort study. Lancet Rheumatol. 2020;2(8):e465–e473. doi: 10.1016/S2665-9913(20)30170-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Baggett TP, Keyes H, Sporn N, Gaeta JM. Prevalence of SARS-CoV-2 infection in residents of a large homeless shelter in Boston. JAMA. 2020;323(21):2191–2192. doi: 10.1001/jama.2020.6887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jung CY, Park H, Kim DW, Choi YJ, Kim SW, Chang TI. Clinical characteristics of asymptomatic patients with COVID-19: a nationwide cohort study in South Korea. Int J Infect Dis. 2020;99:266–268. doi: 10.1016/j.ijid.2020.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Oran DP, Topol EJ. Prevalence of asymptomatic SARS-CoV-2 infection : a narrative review. Ann Intern Med. 2020;173(5):362–367. doi: 10.7326/M20-3012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Abdool Karim SS, de Oliveira T. New SARS-CoV-2 variants—clinical, public health, and vaccine implications. N Engl J Med. 2021;384(19):1866–1868. doi: 10.1056/NEJMc2100362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Classification of Omicron (B.1.1.529): SARS-CoV-2 Variant of Concern. World Health Organization. https://www.who.int/news/item/26-11-2021-classification-of-omicron-(b.1.1.529)-sars-cov-2-variant-of-concern. Accessed 26 Nov 2021.
- 29.SARS-CoV-2 variant classifications and definitions. Centers for Disease Control and Prevention, USA. https://www.cdc.gov/coronavirus/2019-ncov/variants/variant-info.html. Accessed 1 Dec 2021.
- 30.Mejias A, Chavez-Bueno S, Sanchez PJ. Respiratory syncytial virus prophylaxis. NeoReviews. 2005;6(1):e26–e31. [Google Scholar]
- 31.Iversen PL, Kane CD, Zeng XK, Panchal RG, Warren TK, Radoshitzky SR, Kuhn JH, Mudhasani RR, Cooper CL, Shurtleff AC, Nasar F, Sunay MME, Duplantier AJ, Eaton BP, Zumbrun EE, Bixler SL, Martin S, Meinig JM, Chiang CY, Sanchez-Lockhart M, Palacios GF, Kugelman JR, Martins KA, Pitt ML, Crozier I, Saunders DL. Recent successes in therapeutics for Ebola virus disease: no time for complacency. Lancet Infect Dis. 2020;20(9):E231–E237. doi: 10.1016/S1473-3099(20)30282-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Taylor PC, Adams AC, Hufford MM, de la Torre I, Winthrop K, Gottlieb RL. Neutralizing monoclonal antibodies for treatment of COVID-19. Nat Rev Immunol. 2021;21(6):382–393. doi: 10.1038/s41577-021-00542-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Corti D, Purcell LA, Snell G, Veesler D. Tackling COVID-19 with neutralizing monoclonal antibodies. Cell. 2021;184(12):3086–3108. doi: 10.1016/j.cell.2021.05.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.McCallum M, De Marco A, Lempp FA, Tortorici MA, Pinto D, Walls AC, Beltramello M, Chen A, Liu Z, Zatta F, Zepeda S, di Iulio J, Bowen JE, Montiel-Ruiz M, Zhou J, Rosen LE, Bianchi S, Guarino B, Fregni CS, Abdelnabi R, Foo SC, Rothlauf PW, Bloyet LM, Benigni F, Cameroni E, Neyts J, Riva A, Snell G, Telenti A, Whelan SPJ, Virgin HW, Corti D, Pizzuto MS, Veesler D. N-terminal domain antigenic mapping reveals a site of vulnerability for SARS-CoV-2. Cell. 2021;184(9):2332–2347 e2316. doi: 10.1016/j.cell.2021.03.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Suryadevara N, Shrihari S, Gilchuk P, VanBlargan LA, Binshtein E, Zost SJ, Nargi RS, Sutton RE, Winkler ES, Chen EC, Fouch ME, Davidson E, Doranz BJ, Chen RE, Shi PY, Carnahan RH, Thackray LB, Diamond MS, Crowe JE., Jr Neutralizing and protective human monoclonal antibodies recognizing the N-terminal domain of the SARS-CoV-2 spike protein. Cell. 2021;184(9):2316–2331 e2315. doi: 10.1016/j.cell.2021.03.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liu Y, Soh WT, Kishikawa JI, Hirose M, Nakayama EE, Li S, Sasai M, Suzuki T, Tada A, Arakawa A, Matsuoka S, Akamatsu K, Matsuda M, Ono C, Torii S, Kishida K, Jin H, Nakai W, Arase N, Nakagawa A, Matsumoto M, Nakazaki Y, Shindo Y, Kohyama M, Tomii K, Ohmura K, Ohshima S, Okamoto T, Yamamoto M, Nakagami H, Matsuura Y, Nakagawa A, Kato T, Okada M, Standley DM, Shioda T, Arase H. An infectivity-enhancing site on the SARS-CoV-2 spike protein targeted by antibodies. Cell. 2021;184(13):3452–3466 e3418. doi: 10.1016/j.cell.2021.05.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kumar S, Chandele A, Sharma A. Current status of therapeutic monoclonal antibodies against SARS-CoV-2. PLoS Pathog. 2021;17(9):e1009885. doi: 10.1371/journal.ppat.1009885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hussain A, Hasan A, Nejadi Babadaei MM, Bloukh SH, Chowdhury MEH, Sharifi M, Haghighat S, Falahati M. Targeting SARS-CoV2 spike protein receptor binding domain by therapeutic antibodies. Biomed Pharmacother. 2020;130:110559. doi: 10.1016/j.biopha.2020.110559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rogers TF, Zhao F, Huang D, Beutler N, Burns A, He WT, Limbo O, Smith C, Song G, Woehl J, Yang L, Abbott RK, Callaghan S, Garcia E, Hurtado J, Parren M, Peng L, Ramirez S, Ricketts J, Ricciardi MJ, Rawlings SA, Wu NC, Yuan M, Smith DM, Nemazee D, Teijaro JR, Voss JE, Wilson IA, Andrabi R, Briney B, Landais E, Sok D, Jardine JG, Burton DR. Isolation of potent SARS-CoV-2 neutralizing antibodies and protection from disease in a small animal model. Science. 2020;369(6506):956–963. doi: 10.1126/science.abc7520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wang C, Li W, Drabek D, Okba NMA, van Haperen R, Osterhaus A, van Kuppeveld FJM, Haagmans BL, Grosveld F, Bosch BJ. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat Commun. 2020;11(1):2251. doi: 10.1038/s41467-020-16256-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hansen J, Baum A, Pascal KE, Russo V, Giordano S, Wloga E, Fulton BO, Yan Y, Koon K, Patel K, Chung KM, Hermann A, Ullman E, Cruz J, Rafique A, Huang T, Fairhurst J, Libertiny C, Malbec M, Lee WY, Welsh R, Farr G, Pennington S, Deshpande D, Cheng J, Watty A, Bouffard P, Babb R, Levenkova N, Chen C, Zhang B, Romero HA, Saotome K, Zhou Y, Franklin M, Sivapalasingam S, Lye DC, Weston S, Logue J, Haupt R, Frieman M, Chen G, Olson W, Murphy AJ, Stahl N, Yancopoulos GD, Kyratsous CA. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science. 2020;369(6506):1010–1014. doi: 10.1126/science.abd0827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dugan HL, Stamper CT, Li L, Changrob S, Asby NW, Halfmann PJ, Zheng NY, Huang M, Shaw DG, Cobb MS, Erickson SA, Guthmiller JJ, Stovicek O, Wang J, Winkler ES, Madariaga ML, Shanmugarajah K, Jansen MO, Amanat F, Stewart I, Utset HA, Huang J, Nelson CA, Dai YN, Hall PD, Jedrzejczak RP, Joachimiak A, Krammer F, Diamond MS, Fremont DH, Kawaoka Y, Wilson PC. Profiling B cell immunodominance after SARS-CoV-2 infection reveals antibody evolution to non-neutralizing viral targets. Immunity. 2021;54(6):1290–1303 e1297. doi: 10.1016/j.immuni.2021.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Guo Y, Huang L, Zhang G, Yao Y, Zhou H, Shen S, Shen B, Li B, Li X, Zhang Q, Chen M, Chen D, Wu J, Fu D, Zeng X, Feng M, Pi C, Wang Y, Zhou X, Lu M, Li Y, Fang Y, Lu YY, Hu X, Wang S, Zhang W, Gao G, Adrian F, Wang Q, Yu F, Peng Y, Gabibov AG, Min J, Wang Y, Huang H, Stepanov A, Zhang W, Cai Y, Liu J, Yuan Z, Zhang C, Lou Z, Deng F, Zhang H, Shan C, Schweizer L, Sun K, Rao Z. A SARS-CoV-2 neutralizing antibody with extensive Spike binding coverage and modified for optimal therapeutic outcomes. Nat Commun. 2021;12(1):2623. doi: 10.1038/s41467-021-22926-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Huo J, Le Bas A, Ruza RR, Duyvesteyn HME, Mikolajek H, Malinauskas T, Tan TK, Rijal P, Dumoux M, Ward PN, Ren J, Zhou D, Harrison PJ, Weckener M, Clare DK, Vogirala VK, Radecke J, Moynie L, Zhao Y, Gilbert-Jaramillo J, Knight ML, Tree JA, Buttigieg KR, Coombes N, Elmore MJ, Carroll MW, Carrique L, Shah PNM, James W, Townsend AR, Stuart DI, Owens RJ, Naismith JH. Neutralizing nanobodies bind SARS-CoV-2 spike RBD and block interaction with ACE2. Nat Struct Mol Biol. 2020;27(9):846–854. doi: 10.1038/s41594-020-0469-6. [DOI] [PubMed] [Google Scholar]
- 45.Chi X, Liu X, Wang C, Zhang X, Li X, Hou J, Ren L, Jin Q, Wang J, Yang W. Humanized single domain antibodies neutralize SARS-CoV-2 by targeting the spike receptor binding domain. Nat Commun. 2020;11(1):4528. doi: 10.1038/s41467-020-18387-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lv Z, Deng Y-Q, Ye Q, Cao L, Sun C-Y, Fan C, Huang W, Sun S, Sun Y, Zhu L, Chen Q, Wang N, Nie J, Cui Z, Zhu D, Shaw N, Li X-F, Li Q, Xie L, Wang Y, Rao Z, Qin C-F, Wang X. Structural basis for neutralization of SARS-CoV-2 and SARS-CoV by a potent therapeutic antibody. Science. 2020;369(6510):1505–1509. doi: 10.1126/science.abc5881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kim C, Ryu DK, Lee J, Kim YI, Seo JM, Kim YG, Jeong JH, Kim M, Kim JI, Kim P, Bae JS, Shim EY, Lee MS, Kim MS, Noh H, Park GS, Park JS, Son D, An Y, Lee JN, Kwon KS, Lee JY, Lee H, Yang JS, Kim KC, Kim SS, Woo HM, Kim JW, Park MS, Yu KM, Kim SM, Kim EH, Park SJ, Jeong ST, Yu CH, Song Y, Gu SH, Oh H, Koo BS, Hong JJ, Ryu CM, Park WB, Oh MD, Choi YK, Lee SY. A therapeutic neutralizing antibody targeting receptor binding domain of SARS-CoV-2 spike protein. Nat Commun. 2021;12(1):288. doi: 10.1038/s41467-020-20602-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bertoglio F, Meier D, Langreder N, Steinke S, Rand U, Simonelli L, Heine PA, Ballmann R, Schneider KT, Roth KDR, Ruschig M, Riese P, Eschke K, Kim Y, Schackermann D, Pedotti M, Kuhn P, Zock-Emmenthal S, Wohrle J, Kilb N, Herz T, Becker M, Grasshoff M, Wenzel EV, Russo G, Kroger A, Brunotte L, Ludwig S, Fuhner V, Kramer SD, Dubel S, Varani L, Roth G, Cicin-Sain L, Schubert M, Hust M. SARS-CoV-2 neutralizing human recombinant antibodies selected from pre-pandemic healthy donors binding at RBD-ACE2 interface. Nat Commun. 2021;12(1):1577. doi: 10.1038/s41467-021-21609-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Schoof M, Faust B, Saunders RA, Sangwan S, Rezelj V, Hoppe N, Boone M, Billesbolle CB, Puchades C, Azumaya CM, Kratochvil HT, Zimanyi M, Deshpande I, Liang J, Dickinson S, Nguyen HC, Chio CM, Merz GE, Thompson MC, Diwanji D, Schaefer K, Anand AA, Dobzinski N, Zha BS, Simoneau CR, Leon K, White KM, Chio US, Gupta M, Jin M, Li F, Liu Y, Zhang K, Bulkley D, Sun M, Smith AM, Rizo AN, Moss F, Brilot AF, Pourmal S, Trenker R, Pospiech T, Gupta S, Barsi-Rhyne B, Belyy V, Barile-Hill AW, Nock S, Liu Y, Krogan NJ, Ralston CY, Swaney DL, Garcia-Sastre A, Ott M, Vignuzzi M, Consortium QSB, Walter P, Manglik A. An ultrapotent synthetic nanobody neutralizes SARS-CoV-2 by stabilizing inactive Spike. Science. 2020;370(6523):1473–1479. doi: 10.1126/science.abe3255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Su SC, Yang TJ, Yu PY, Liang KH, Chen WY, Yang CW, Lin HT, Wang MJ, Lu RM, Tso HC, Chung MJ, Hsieh TY, Chang YL, Lin SC, Hsu FY, Ke FY, Wu YH, Hwang YC, Liu IJ, Liang JJ, Liao CC, Ko HY, Sun CP, Wu PY, Jan JT, Chang YC, Lin YL, Tao MH, Hsu SD, Wu HC. Structure-guided antibody cocktail for prevention and treatment of COVID-19. PLoS Pathog. 2021;17(10):e1009704. doi: 10.1371/journal.ppat.1009704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Shi R, Shan C, Duan X, Chen Z, Liu P, Song J, Song T, Bi X, Han C, Wu L, Gao G, Hu X, Zhang Y, Tong Z, Huang W, Liu WJ, Wu G, Zhang B, Wang L, Qi J, Feng H, Wang FS, Wang Q, Gao GF, Yuan Z, Yan J. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature. 2020;584(7819):120–124. doi: 10.1038/s41586-020-2381-y. [DOI] [PubMed] [Google Scholar]
- 52.Sia SF, Yan LM, Chin AWH, Fung K, Choy KT, Wong AYL, Kaewpreedee P, Perera R, Poon LLM, Nicholls JM, Peiris M, Yen HL. Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature. 2020;583(7818):834–838. doi: 10.1038/s41586-020-2342-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wang P, Nair MS, Liu L, Iketani S, Luo Y, Guo Y, Wang M, Yu J, Zhang B, Kwong PD, Graham BS, Mascola JR, Chang JY, Yin MT, Sobieszczyk M, Kyratsous CA, Shapiro L, Sheng Z, Huang Y, Ho DD. Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7. Nature. 2021;593(7857):130–135. doi: 10.1038/s41586-021-03398-2. [DOI] [PubMed] [Google Scholar]
- 54.Wang P, Casner RG, Nair MS, Wang M, Yu J, Cerutti G, Liu L, Kwong PD, Huang Y, Shapiro L, Ho DD. Increased resistance of SARS-CoV-2 variant P.1 to antibody neutralization. Cell Host Microbe. 2021;29(5):747–751 e744. doi: 10.1016/j.chom.2021.04.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Weinreich DM, Sivapalasingam S, Norton T, Ali S, Gao H, Bhore R, Musser BJ, Soo Y, Rofail D, Im J, Perry C, Pan C, Hosain R, Mahmood A, Davis JD, Turner KC, Hooper AT, Hamilton JD, Baum A, Kyratsous CA, Kim Y, Cook A, Kampman W, Kohli A, Sachdeva Y, Graber X, Kowal B, DiCioccio T, Stahl N, Lipsich L, Braunstein N, Herman G, Yancopoulos GD, Trial I. REGN-COV2, a neutralizing antibody cocktail, in outpatients with COVID-19. N Engl J Med. 2021;384(3):238–251. doi: 10.1056/NEJMoa2035002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Tuccori M, Ferraro S, Convertino I, Cappello E, Valdiserra G, Blandizzi C, Maggi F, Focosi D. Anti-SARS-CoV-2 neutralizing monoclonal antibodies: clinical pipeline. MAbs. 2020;12(1):1854149. doi: 10.1080/19420862.2020.1854149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Chen P, Nirula A, Heller B, Gottlieb RL, Boscia J, Morris J, Huhn G, Cardona J, Mocherla B, Stosor V, Shawa I, Adams AC, Van Naarden J, Custer KL, Shen L, Durante M, Oakley G, Schade AE, Sabo J, Patel DR, Klekotka P, Skovronsky DM, Investigators B SARS-CoV-2 neutralizing antibody LY-CoV555 in outpatients with COVID-19. N Engl J Med. 2021;384(3):229–237. doi: 10.1056/NEJMoa2029849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.ACTIV-3/TICO LY-CoV555 Study Group, Lundgren JD, Grund B, Barkauskas CE, Holland TL, Gottlieb RL, Sandkovsky U, Brown SM, Knowlton KU, Self WH, Files DC, Jain MK, Benfield T, Bowdish ME, Leshnower BG, Baker JV, Jensen JU, Gardner EM, Ginde AA, Harris ES, Johansen IS, Markowitz N, Matthay MA, Ostergaard L, Chang CC, Davey VJ, Goodman A, Higgs ES, Murray DD, Murray TA, Paredes R, Parmar MKB, Phillips AN, Reilly C, Sharma S, Dewar RL, Teitelbaum M, Wentworth D, Cao H, Klekotka P, Babiker AG, Gelijns AC, Kan VL, Polizzotto MN, Thompson BT, Lane HC, Neaton JD. A neutralizing monoclonal antibody for hospitalized patients with COVID-19. N Engl J Med. 2021; 384(10):905–914. [DOI] [PMC free article] [PubMed]
- 59.Tian X, Li C, Huang A, Xia S, Lu S, Shi Z, Lu L, Jiang S, Yang Z, Wu Y, Ying T. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg Microbes Infect. 2020;9(1):382–385. doi: 10.1080/22221751.2020.1729069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhou D, Duyvesteyn HME, Chen CP, Huang CG, Chen TH, Shih SR, Lin YC, Cheng CY, Cheng SH, Huang YC, Lin TY, Ma C, Huo J, Carrique L, Malinauskas T, Ruza RR, Shah PNM, Tan TK, Rijal P, Donat RF, Godwin K, Buttigieg KR, Tree JA, Radecke J, Paterson NG, Supasa P, Mongkolsapaya J, Screaton GR, Carroll MW, Gilbert-Jaramillo J, Knight ML, James W, Owens RJ, Naismith JH, Townsend AR, Fry EE, Zhao Y, Ren J, Stuart DI, Huang KA. Structural basis for the neutralization of SARS-CoV-2 by an antibody from a convalescent patient. Nat Struct Mol Biol. 2020;27(10):950–958. doi: 10.1038/s41594-020-0480-y. [DOI] [PubMed] [Google Scholar]
- 61.Tortorici MA, Czudnochowski N, Starr TN, Marzi R, Walls AC, Zatta F, Bowen JE, Jaconi S, Di Iulio J, Wang Z, De Marco A, Zepeda SK, Pinto D, Liu Z, Beltramello M, Bartha I, Housley MP, Lempp FA, Rosen LE, Dellota E, Jr, Kaiser H, Montiel-Ruiz M, Zhou J, Addetia A, Guarino B, Culap K, Sprugasci N, Saliba C, Vetti E, Giacchetto-Sasselli I, Fregni CS, Abdelnabi R, Foo SC, Havenar-Daughton C, Schmid MA, Benigni F, Cameroni E, Neyts J, Telenti A, Virgin HW, Whelan SPJ, Snell G, Bloom JD, Corti D, Veesler D, Pizzuto MS. Broad sarbecovirus neutralization by a human monoclonal antibody. Nature. 2021;597(7874):103–108. doi: 10.1038/s41586-021-03817-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Starr TN, Czudnochowski N, Liu Z, Zatta F, Park YJ, Addetia A, Pinto D, Beltramello M, Hernandez P, Greaney AJ, Marzi R, Glass WG, Zhang I, Dingens AS, Bowen JE, Tortorici MA, Walls AC, Wojcechowskyj JA, De Marco A, Rosen LE, Zhou J, Montiel-Ruiz M, Kaiser H, Dillen JR, Tucker H, Bassi J, Silacci-Fregni C, Housley MP, di Iulio J, Lombardo G, Agostini M, Sprugasci N, Culap K, Jaconi S, Meury M, Dellota E, Jr, Abdelnabi R, Foo SC, Cameroni E, Stumpf S, Croll TI, Nix JC, Havenar-Daughton C, Piccoli L, Benigni F, Neyts J, Telenti A, Lempp FA, Pizzuto MS, Chodera JD, Hebner CM, Virgin HW, Whelan SPJ, Veesler D, Corti D, Bloom JD, Snell G. SARS-CoV-2 RBD antibodies that maximize breadth and resistance to escape. Nature. 2021;597(7874):97–102. doi: 10.1038/s41586-021-03807-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh C-L, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367(6483):1260–1263. doi: 10.1126/science.abb2507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Benton DJ, Wrobel AG, Xu P, Roustan C, Martin SR, Rosenthal PB, Skehel JJ, Gamblin SJ. Receptor binding and priming of the spike protein of SARS-CoV-2 for membrane fusion. Nature. 2020;588(7837):327–330. doi: 10.1038/s41586-020-2772-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Sun SH, Chen Q, Gu HJ, Yang G, Wang YX, Huang XY, Liu SS, Zhang NN, Li XF, Xiong R, Guo Y, Deng YQ, Huang WJ, Liu Q, Liu QM, Shen YL, Zhou Y, Yang X, Zhao TY, Fan CF, Zhou YS, Qin CF, Wang YC. A mouse model of SARS-CoV-2 infection and pathogenesis. Cell Host Microbe. 2020;28(1):124–133124. doi: 10.1016/j.chom.2020.05.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Liu L, Wang P, Nair MS, Yu J, Rapp M, Wang Q, Luo Y, Chan JF, Sahi V, Figueroa A, Guo XV, Cerutti G, Bimela J, Gorman J, Zhou T, Chen Z, Yuen KY, Kwong PD, Sodroski JG, Yin MT, Sheng Z, Huang Y, Shapiro L, Ho DD. Potent neutralizing antibodies against multiple epitopes on SARS-CoV-2 spike. Nature. 2020;584(7821):450–456. doi: 10.1038/s41586-020-2571-7. [DOI] [PubMed] [Google Scholar]
- 67.Fedry J, Hurdiss DL, Wang C, Li W, Obal G, Drulyte I, Du W, Howes SC, van Kuppeveld FJM, Förster F, Bosch BJ. Structural insights into the cross-neutralization of SARS-CoV and SARS-CoV-2 by the human monoclonal antibody 47D11. Sci Adv. 2021 doi: 10.1126/sciadv.abf5632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Harbour BioMed and Utrecht University Announce License Agreement with AbbVie and Initiation of COVID-19 Antibody Clinical Trials. PR Newswire Association LLC. https://www.prnewswire.com/news-releases/harbour-biomed-and-utrecht-university-announce-license-agreement-with-abbvie-and-initiation-of-covid-19-antibody-clinical-trials-301192226.html. Accessed 14 Dec 2020.
- 69.Study to Assess Adverse Events and How Intravenous (IV) ABBV-47D11 and IV ABBV-2B04 Given Alone and in Combination Moves Through the Body of Adult Participants With Coronavirus Disease 2019 (COVID-19). https://ClinicalTrials.gov/show/NCT04644120.
- 70.Hurlburt NK, Seydoux E, Wan YH, Edara VV, Stuart AB, Feng J, Suthar MS, McGuire AT, Stamatatos L, Pancera M. Structural basis for potent neutralization of SARS-CoV-2 and role of antibody affinity maturation. Nat Commun. 2020;11(1):5413. doi: 10.1038/s41467-020-19231-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Brouwer PJM, Caniels TG, van der Straten K, Snitselaar JL, Aldon Y, Bangaru S, Torres JL, Okba NMA, Claireaux M, Kerster G, Bentlage AEH, van Haaren MM, Guerra D, Burger JA, Schermer EE, Verheul KD, van der Velde N, van der Kooi A, van Schooten J, van Breemen MJ, Bijl TPL, Sliepen K, Aartse A, Derking R, Bontjer I, Kootstra NA, Wiersinga WJ, Vidarsson G, Haagmans BL, Ward AB, de Bree GJ, Sanders RW, van Gils MJ. Potent neutralizing antibodies from COVID-19 patients define multiple targets of vulnerability. Science. 2020;369(6504):643–650. doi: 10.1126/science.abc5902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Wu Y, Wang F, Shen C, Peng W, Li D, Zhao C, Li Z, Li S, Bi Y, Yang Y, Gong Y, Xiao H, Fan Z, Tan S, Wu G, Tan W, Lu X, Fan C, Wang Q, Liu Y, Zhang C, Qi J, Gao GF, Gao F, Liu L. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science. 2020;368(6496):1274–1278. doi: 10.1126/science.abc2241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Noy-Porat T, Makdasi E, Alcalay R, Mechaly A, Levy Y, Bercovich-Kinori A, Zauberman A, Tamir H, Yahalom-Ronen Y, Israeli M, Epstein E, Achdout H, Melamed S, Chitlaru T, Weiss S, Peretz E, Rosen O, Paran N, Yitzhaki S, Shapira SC, Israely T, Mazor O, Rosenfeld R. A panel of human neutralizing mAbs targeting SARS-CoV-2 spike at multiple epitopes. Nat Commun. 2020;11(1):4303. doi: 10.1038/s41467-020-18159-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Chi X, Yan R, Zhang J, Zhang G, Zhang Y, Hao M, Zhang Z, Fan P, Dong Y, Yang Y, Chen Z, Guo Y, Zhang J, Li Y, Song X, Chen Y, Xia L, Fu L, Hou L, Xu J, Yu C, Li J, Zhou Q, Chen W. A neutralizing human antibody binds to the N-terminal domain of the Spike protein of SARS-CoV-2. Science. 2020;369(6504):650–655. doi: 10.1126/science.abc6952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Liang KH, Chiang PY, Ko SH, Chou YC, Lu RM, Lin HT, Chen WY, Lin YL, Tao MH, Jan JT, Wu HC. Antibody cocktail effective against variants of SARS-CoV-2. J Biomed Sci. 2021;28(1):80. doi: 10.1186/s12929-021-00777-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Yang TJ, Yu PY, Chang YC, Liang KH, Tso HC, Ho MR, Chen WY, Lin HT, Wu HC, Hsu SD. Effect of SARS-CoV-2 B.1.1.7 mutations on spike protein structure and function. Nat Struct Mol Biol. 2021;28(9):731–739. doi: 10.1038/s41594-021-00652-z. [DOI] [PubMed] [Google Scholar]
- 77.Yang L, Liu W, Yu X, Wu M, Reichert JM, Ho M. COVID-19 antibody therapeutics tracker: a global online database of antibody therapeutics for the prevention and treatment of COVID-19. Antib Ther. 2020;3(3):205–212. doi: 10.1093/abt/tbaa020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Gavor E, Choong YK, Er SY, Sivaraman H, Sivaraman J. Structural basis of SARS-CoV-2 and SARS-CoV antibody interactions. Trends Immunol. 2020;41(11):1006–1022. doi: 10.1016/j.it.2020.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Fouladirad S, Bach H. Development of coronavirus treatments using neutralizing antibodies. Microorganisms. 2021;9(1):165. doi: 10.3390/microorganisms9010165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Renn A, Fu Y, Hu X, Hall MD, Simeonov A. Fruitful neutralizing antibody pipeline brings hope to defeat SARS-Cov-2. Trends Pharmacol Sci. 2020;41(11):815–829. doi: 10.1016/j.tips.2020.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Baum A, Fulton BO, Wloga E, Copin R, Pascal KE, Russo V, Giordano S, Lanza K, Negron N, Ni M, Wei Y, Atwal GS, Murphy AJ, Stahl N, Yancopoulos GD, Kyratsous CA. Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies. Science. 2020;369(6506):1014–1018. doi: 10.1126/science.abd0831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Baum A, Ajithdoss D, Copin R, Zhou A, Lanza K, Negron N, Ni M, Wei Y, Mohammadi K, Musser B, Atwal GS, Oyejide A, Goez-Gazi Y, Dutton J, Clemmons E, Staples HM, Bartley C, Klaffke B, Alfson K, Gazi M, Gonzalez O, Dick E, Jr, Carrion R, Jr, Pessaint L, Porto M, Cook A, Brown R, Ali V, Greenhouse J, Taylor T, Andersen H, Lewis MG, Stahl N, Murphy AJ, Yancopoulos GD, Kyratsous CA. REGN-COV2 antibodies prevent and treat SARS-CoV-2 infection in rhesus macaques and hamsters. Science. 2020;370(6520):1110–1115. doi: 10.1126/science.abe2402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.O'Brien MP, Forleo-Neto E, Musser BJ, Isa F, Chan KC, Sarkar N, Bar KJ, Barnabas RV, Barouch DH, Cohen MS, Hurt CB, Burwen DR, Marovich MA, Hou P, Heirman I, Davis JD, Turner KC, Ramesh D, Mahmood A, Hooper AT, Hamilton JD, Kim Y, Purcell LA, Baum A, Kyratsous CA, Krainson J, Perez-Perez R, Mohseni R, Kowal B, DiCioccio AT, Stahl N, Lipsich L, Braunstein N, Herman G, Yancopoulos GD, Weinreich DM, Covid-19 Phase 3 Prevention Trial T Subcutaneous REGEN-COV antibody combination to prevent COVID-19. N Engl J Med. 2021;385(13):1184–1195. doi: 10.1056/NEJMoa2109682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Japan becomes first country to approve Ronapreve (casirivimab and imdevimab) for the treatment of mild to moderate COVID-19. F. Hoffmann-La Roche Ltd. https://www.roche.com/media/releases/med-cor-2021-07-20.htm?utm_source=T&utm_medium=E&utm_campaign=Ronapreve%20approval%20in%20Japan. Accessed 20 Jul 2021.
- 85.Chen RE, Winkler ES, Case JB, Aziati ID, Bricker TL, Joshi A, Darling TL, Ying B, Errico JM, Shrihari S, VanBlargan LA, Xie X, Gilchuk P, Zost SJ, Droit L, Liu Z, Stumpf S, Wang D, Handley SA, Stine WB, Jr, Shi PY, Davis-Gardner ME, Suthar MS, Knight MG, Andino R, Chiu CY, Ellebedy AH, Fremont DH, Whelan SPJ, Crowe JE, Jr, Purcell L, Corti D, Boon ACM, Diamond MS. In vivo monoclonal antibody efficacy against SARS-CoV-2 variant strains. Nature. 2021;596(7870):103–108. doi: 10.1038/s41586-021-03720-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Hoffmann M, Hofmann-Winkler H, Kruger N, Kempf A, Nehlmeier I, Graichen L, Arora P, Sidarovich A, Moldenhauer AS, Winkler MS, Schulz S, Jack HM, Stankov MV, Behrens GMN, Pohlmann S. SARS-CoV-2 variant B.1.617 is resistant to bamlanivimab and evades antibodies induced by infection and vaccination. Cell Rep. 2021;36(3):109415. doi: 10.1016/j.celrep.2021.109415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.FDA authorizes REGEN-COV monoclonal antibody therapy for post-exposure prophylaxis (prevention) for COVID-19. U.S. FDA. https://www.fda.gov/drugs/drug-safety-and-availability/fda-authorizes-regen-cov-monoclonal-antibody-therapy-post-exposure-prophylaxis-prevention-covid-19. Accessed 10 Aug 2021.
- 88.Philippidis A. Top 11 Best Selling COVID-19 Vaccines and Drugs of H1 2021. Genetic Engineering & Biotechnology News. https://www.genengnews.com/a-lists/top-11-best-selling-covid-19-vaccines-and-drugs-of-h1-2021/. Accessed 30 Sep 2021.
- 89.Jones BE, Brown-Augsburger PL, Corbett KS, Westendorf K, Davies J, Cujec TP, Wiethoff CM, Blackbourne JL, Heinz BA, Foster D, Higgs RE, Balasubramaniam D, Wang L, Zhang Y, Yang ES, Bidshahri R, Kraft L, Hwang Y, Zentelis S, Jepson KR, Goya R, Smith MA, Collins DW, Hinshaw SJ, Tycho SA, Pellacani D, Xiang P, Muthuraman K, Sobhanifar S, Piper MH, Triana FJ, Hendle J, Pustilnik A, Adams AC, Berens SJ, Baric RS, Martinez DR, Cross RW, Geisbert TW, Borisevich V, Abiona O, Belli HM, de Vries M, Mohamed A, Dittmann M, Samanovic MI, Mulligan MJ, Goldsmith JA, Hsieh CL, Johnson NV, Wrapp D, McLellan JS, Barnhart BC, Graham BS, Mascola JR, Hansen CL, Falconer E. The neutralizing antibody, LY-CoV555, protects against SARS-CoV-2 infection in nonhuman primates. Sci Transl Med. 2021 doi: 10.1126/scitranslmed.abf1906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Kaplon H, Reichert JM. Antibodies to watch in 2021. MAbs. 2021;13(1):1860476. doi: 10.1080/19420862.2020.1860476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Starr TN, Greaney AJ, Hilton SK, Ellis D, Crawford KHD, Dingens AS, Navarro MJ, Bowen JE, Tortorici MA, Walls AC, King NP, Veesler D, Bloom JD. Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell. 2020;182(5):1295. doi: 10.1016/j.cell.2020.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Starr TN, Greaney AJ, Dingens AS, Bloom JD. Complete map of SARS-CoV-2 RBD mutations that escape the monoclonal antibody LY-CoV555 and its cocktail with LY-CoV016. Cell Rep Med. 2021;2(4):100255. doi: 10.1016/j.xcrm.2021.100255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Liu L, Wei Q, Lin Q, Fang J, Wang H, Kwok H, Tang H, Nishiura K, Peng J, Tan Z, Wu T, Cheung KW, Chan KH, Alvarez X, Qin C, Lackner A, Perlman S, Yuen KY, Chen Z. Anti-spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight. 2019 doi: 10.1172/jci.insight.123158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Gottlieb RL, Nirula A, Chen P, Boscia J, Heller B, Morris J, Huhn G, Cardona J, Mocherla B, Stosor V, Shawa I, Kumar P, Adams AC, Van Naarden J, Custer KL, Durante M, Oakley G, Schade AE, Holzer TR, Ebert PJ, Higgs RE, Kallewaard NL, Sabo J, Patel DR, Klekotka P, Shen L, Skovronsky DM. Effect of Bamlanivimab as monotherapy or in combination with Etesevimab on viral load in patients with mild to moderate COVID-19: a randomized clinical trial. JAMA. 2021;325(7):632–644. doi: 10.1001/jama.2021.0202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Pause in the Distribution of bamlanivimab/etesevimab. U.S. Department of Health and Human Services. https://www.phe.gov/emergency/events/COVID19/investigation-MCM/Bamlanivimab-etesevimab/Pages/bamlanivimab-etesevimab-distribution-pause.aspx. Accessed 30 Sep 2021.
- 96.Cathcart AL, Havenar-Daughton C, Lempp FA, Ma D, Schmid MA, Agostini ML, Guarino B, Di iulio J, Rosen LE, Tucker H, Dillen J, Subramanian S, Sloan B, Bianchi S, Pinto D, Saliba C, Wojcechowskyj JA, Noack J, Zhou J, Kaiser H, Chase A, Montiel-Ruiz M, Dellota E, Park A, Spreafico R, Sahakyan A, Lauron EJ, Czudnochowski N, Cameroni E, Ledoux S, Werts A, Colas C, Soriaga L, Telenti A, Purcell LA, Hwang S, Snell G, Virgin HW, Corti D, Hebner CM. The dual function monoclonal antibodies VIR-7831 and VIR-7832 demonstrate potent in vitro and in vivo activity against SARS-CoV-2. bioRxiv. 2021. 2021.2003.2009.434607.
- 97.Pinto D, Park YJ, Beltramello M, Walls AC, Tortorici MA, Bianchi S, Jaconi S, Culap K, Zatta F, De Marco A, Peter A, Guarino B, Spreafico R, Cameroni E, Case JB, Chen RE, Havenar-Daughton C, Snell G, Telenti A, Virgin HW, Lanzavecchia A, Diamond MS, Fink K, Veesler D, Corti D. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature. 2020;583(7815):290–295. doi: 10.1038/s41586-020-2349-y. [DOI] [PubMed] [Google Scholar]
- 98.Schafer A, Muecksch F, Lorenzi JCC, Leist SR, Cipolla M, Bournazos S, Schmidt F, Maison RM, Gazumyan A, Martinez DR, Baric RS, Robbiani DF, Hatziioannou T, Ravetch JV, Bieniasz PD, Bowen RA, Nussenzweig MC, Sheahan TP. Antibody potency, effector function, and combinations in protection and therapy for SARS-CoV-2 infection in vivo. J Exp Med. 2021 doi: 10.1084/jem.20201993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.McCallum M, Bassi J, De Marco A, Chen A, Walls AC, Di Iulio J, Tortorici MA, Navarro MJ, Silacci-Fregni C, Saliba C, Sprouse KR, Agostini M, Pinto D, Culap K, Bianchi S, Jaconi S, Cameroni E, Bowen JE, Tilles SW, Pizzuto MS, Guastalla SB, Bona G, Pellanda AF, Garzoni C, Van Voorhis WC, Rosen LE, Snell G, Telenti A, Virgin HW, Piccoli L, Corti D, Veesler D. SARS-CoV-2 immune evasion by the B.1.427/B.1.429 variant of concern. Science. 2021;373(6555):648–654. doi: 10.1126/science.abi7994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Zost SJ, Gilchuk P, Case JB, Binshtein E, Chen RE, Nkolola JP, Schafer A, Reidy JX, Trivette A, Nargi RS, Sutton RE, Suryadevara N, Martinez DR, Williamson LE, Chen EC, Jones T, Day S, Myers L, Hassan AO, Kafai NM, Winkler ES, Fox JM, Shrihari S, Mueller BK, Meiler J, Chandrashekar A, Mercado NB, Steinhardt JJ, Ren K, Loo YM, Kallewaard NL, McCune BT, Keeler SP, Holtzman MJ, Barouch DH, Gralinski LE, Baric RS, Thackray LB, Diamond MS, Carnahan RH, Crowe JE., Jr Potently neutralizing and protective human antibodies against SARS-CoV-2. Nature. 2020;584(7821):443–449. doi: 10.1038/s41586-020-2548-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Dong J, Zost SJ, Greaney AJ, Starr TN, Dingens AS, Chen EC, Chen RE, Case JB, Sutton RE, Gilchuk P, Rodriguez J, Armstrong E, Gainza C, Nargi RS, Binshtein E, Xie X, Zhang X, Shi P-Y, Logue J, Weston S, McGrath ME, Frieman MB, Brady T, Tuffy KM, Bright H, Loo Y-M, McTamney PM, Esser MT, Carnahan RH, Diamond MS, Bloom JD, Crowe JE. Genetic and structural basis for SARS-CoV-2 variant neutralization by a two-antibody cocktail. Nat Microbiol. 2021;6(10):1233–1244. doi: 10.1038/s41564-021-00972-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Loo Y-M, McTamney PM, Arends RH, Gasser RA, Abram ME, Aksyuk A, Diallo S, Flores DJ, Kelly EJ, Ren K, Roque R, Rosenthal K, Streicher K, Tuffy KM, Bond NJ, Cornwell O, Bouquet J, Cheng LI, Dunyak J, Huang Y, Rosenbaum AI, Andersen H, Carnahan RH, Crowe JE, Kuehne AI, Herbert AS, Dye JM, Bright H, Kallewaard NL, Pangalos MN, Esser MT. AZD7442 demonstrates prophylactic and therapeutic efficacy in non-human primates and extended half-life in humans. medRxiv. 2021. 2021.2008.2030.21262666.
- 103.Oganesyan V, Gao C, Shirinian L, Wu H, Dall'Acqua WF. Structural characterization of a human Fc fragment engineered for lack of effector functions. Acta Crystallogr Sect D. 2008;64(6):700–704. doi: 10.1107/S0907444908007877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Robbie GJ, Criste R, Dall'acqua WF, Jensen K, Patel NK, Losonsky GA, Griffin MP. A novel investigational Fc-modified humanized monoclonal antibody, motavizumab-YTE, has an extended half-life in healthy adults. Antimicrob Agents Chemother. 2013;57(12):6147–6153. doi: 10.1128/AAC.01285-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Chen EC, Gilchuk P, Zost SJ, Suryadevara N, Winkler ES, Cabel CR, Binshtein E, Chen RE, Sutton RE, Rodriguez J, Day S, Myers L, Trivette A, Williams JK, Davidson E, Li S, Doranz BJ, Campos SK, Carnahan RH, Thorne CA, Diamond MS, Crowe JE., Jr Convergent antibody responses to the SARS-CoV-2 spike protein in convalescent and vaccinated individuals. Cell Rep. 2021;36(8):109604. doi: 10.1016/j.celrep.2021.109604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Mahase E. COVID-19: AstraZeneca says its antibody drug AZD7442 is effective for preventing and reducing severe illness. BMJ. 2021;375:n2860. doi: 10.1136/bmj.n2860. [DOI] [PubMed] [Google Scholar]
- 107.AstraZeneca to supply the US with up to half a million additional doses of the potential COVID-19 antibody treatment AZD7442. AstraZeneca 2021. https://www.astrazeneca.com/media-centre/press-releases/2021/us-supply-agreement-for-additional-azd7442-doses.html. Accessed 16 Mar 2021.
- 108.Tang L, Yin Z, Hu Y, Mei H. Controlling cytokine storm is vital in COVID-19. Front Immunol. 2020;11:570993. doi: 10.3389/fimmu.2020.570993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Coperchini F, Chiovato L, Croce L, Magri F, Rotondi M. The cytokine storm in COVID-19: an overview of the involvement of the chemokine/chemokine-receptor system. Cytokine Growth Factor Rev. 2020;53:25–32. doi: 10.1016/j.cytogfr.2020.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Fajgenbaum DC, June CH. Cytokine storm. N Engl J Med. 2020;383(23):2255–2273. doi: 10.1056/NEJMra2026131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Gao YM, Xu G, Wang B, Liu BC. Cytokine storm syndrome in coronavirus disease 2019: a narrative review. J Intern Med. 2021;289(2):147–161. doi: 10.1111/joim.13144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Sinha P, Matthay MA, Calfee CS. Is a "Cytokine Storm" Relevant to COVID-19? JAMA Intern Med. 2020;180(9):1152–1154. doi: 10.1001/jamainternmed.2020.3313. [DOI] [PubMed] [Google Scholar]
- 113.Hunter CA, Jones SA. IL-6 as a keystone cytokine in health and disease. Nat Immunol. 2015;16(5):448–457. doi: 10.1038/ni.3153. [DOI] [PubMed] [Google Scholar]
- 114.Ruan Q, Yang K, Wang W, Jiang L, Song J. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan, China. Intensive Care Med. 2020;46(5):846–848. doi: 10.1007/s00134-020-05991-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Chen G, Wu D, Guo W, Cao Y, Huang D, Wang H, Wang T, Zhang X, Chen H, Yu H, Zhang X, Zhang M, Wu S, Song J, Chen T, Han M, Li S, Luo X, Zhao J, Ning Q. Clinical and immunological features of severe and moderate coronavirus disease 2019. J Clin Invest. 2020;130(5):2620–2629. doi: 10.1172/JCI137244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Wang J, Jiang M, Chen X, Montaner LJ. Cytokine storm and leukocyte changes in mild versus severe SARS-CoV-2 infection: review of 3939 COVID-19 patients in China and emerging pathogenesis and therapy concepts. J Leukoc Biol. 2020;108(1):17–41. doi: 10.1002/JLB.3COVR0520-272R. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Gritti G, Raimondi F, Bottazzi B, Ripamonti D, Riva I, Landi F, Alborghetti L, Frigeni M, Damiani M, Mico C, Fagiuoli S, Lorini FL, Gandini L, Novelli L, Morgan JP, Owens BMJ, Kanhai KJK, Reljanovic GT, Rizzi M, Di Marco F, Mantovani A, Rambaldi A. Siltuximab downregulates interleukin-8 and pentraxin 3 to improve ventilatory status and survival in severe COVID-19. Leukemia. 2021;35(9):2710–2714. doi: 10.1038/s41375-021-01299-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Lescure FX, Honda H, Fowler RA, Lazar JS, Shi G, Wung P, Patel N, Hagino O, Sarilumab C-GSG. Sarilumab in patients admitted to hospital with severe or critical COVID-19: a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Respir Med. 2021;9(5):522–532. doi: 10.1016/S2213-2600(21)00099-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Remap-Cap Investigators, Gordon AC, Mouncey PR, Al-Beidh F, Rowan KM, Nichol AD, Arabi YM, Annane D, Beane A, van Bentum-Puijk W, Berry LR, Bhimani Z, Bonten MJM, Bradbury CA, Brunkhorst FM, Buzgau A, Cheng AC, Detry MA, Duffy EJ, Estcourt LJ, Fitzgerald M, Goossens H, Haniffa R, Higgins AM, Hills TE, Horvat CM, Lamontagne F, Lawler PR, Leavis HL, Linstrum KM, Litton E, Lorenzi E, Marshall JC, Mayr FB, McAuley DF, McGlothlin A, McGuinness SP, McVerry BJ, Montgomery SK, Morpeth SC, Murthy S, Orr K, Parke RL, Parker JC, Patanwala AE, Pettila V, Rademaker E, Santos MS, Saunders CT, Seymour CW, Shankar-Hari M, Sligl WI, Turgeon AF, Turner AM, van de Veerdonk FL, Zarychanski R, Green C, Lewis RJ, Angus DC, McArthur CJ, Berry S, Webb SA, Derde LPG. Interleukin-6 receptor antagonists in critically ill patients with COVID-19. N Engl J Med. 2021; 384(16):1491–1502.
- 120.Gupta S, Wang W, Hayek SS, Chan L, Mathews KS, Melamed ML, Brenner SK, Leonberg-Yoo A, Schenck EJ, Radbel J, Reiser J, Bansal A, Srivastava A, Zhou Y, Finkel D, Green A, Mallappallil M, Faugno AJ, Zhang J, Velez JCQ, Shaefi S, Parikh CR, Charytan DM, Athavale AM, Friedman AN, Redfern RE, Short SAP, Correa S, Pokharel KK, Admon AJ, Donnelly JP, Gershengorn HB, Douin DJ, Semler MW, Hernan MA, Leaf DE, Investigators S-C. Association between early treatment with tocilizumab and mortality among critically ill patients with COVID-19. JAMA Intern Med. 2021;181(1):41–51. doi: 10.1001/jamainternmed.2020.6252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Stone JH, Frigault MJ, Serling-Boyd NJ, Fernandes AD, Harvey L, Foulkes AS, Horick NK, Healy BC, Shah R, Bensaci AM, Woolley AE, Nikiforow S, Lin N, Sagar M, Schrager H, Huckins DS, Axelrod M, Pincus MD, Fleisher J, Sacks CA, Dougan M, North CM, Halvorsen YD, Thurber TK, Dagher Z, Scherer A, Wallwork RS, Kim AY, Schoenfeld S, Sen P, Neilan TG, Perugino CA, Unizony SH, Collier DS, Matza MA, Yinh JM, Bowman KA, Meyerowitz E, Zafar A, Drobni ZD, Bolster MB, Kohler M, D'Silva KM, Dau J, Lockwood MM, Cubbison C, Weber BN, Mansour MK, Investigators BBTT. Efficacy of tocilizumab in patients hospitalized with COVID-19. N Engl J Med. 2020;383(24):2333–2344. doi: 10.1056/NEJMoa2028836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Della-Torre E, Campochiaro C, Cavalli G, De Luca G, Napolitano A, La Marca S, Boffini N, Da Prat V, Di Terlizzi G, Lanzillotta M, Rovere Querini P, Ruggeri A, Landoni G, Tresoldi M, Ciceri F, Zangrillo A, De Cobelli F, Dagna L, Groeup S-RS, Members S-RSG. Interleukin-6 blockade with sarilumab in severe COVID-19 pneumonia with systemic hyperinflammation: an open-label cohort study. Ann Rheum Dis. 2020;79(10):1277–1285. [DOI] [PMC free article] [PubMed]
- 123.Mullard A. Anti-IL-6Rs falter in COVID-19. Nat Rev Drug Discov. 2020;19(9):577. doi: 10.1038/d41573-020-00141-w. [DOI] [PubMed] [Google Scholar]
- 124.Rosas IO, Brau N, Waters M, Go RC, Hunter BD, Bhagani S, Skiest D, Aziz MS, Cooper N, Douglas IS, Savic S, Youngstein T, Del Sorbo L, Cubillo Gracian A, De La Zerda DJ, Ustianowski A, Bao M, Dimonaco S, Graham E, Matharu B, Spotswood H, Tsai L, Malhotra A. Tocilizumab in hospitalized patients with severe COVID-19 pneumonia. N Engl J Med. 2021;384(16):1503–1516. doi: 10.1056/NEJMoa2028700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Recovery Collaborative Group Tocilizumab in patients admitted to hospital with COVID-19 (RECOVERY): a randomised, controlled, open-label, platform trial. Lancet. 2021;397(10285):1637–1645. doi: 10.1016/S0140-6736(21)00676-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Salama C, Han J, Yau L, Reiss WG, Kramer B, Neidhart JD, Criner GJ, Kaplan-Lewis E, Baden R, Pandit L, Cameron ML, Garcia-Diaz J, Chavez V, Mekebeb-Reuter M, Limade Menezes F, Shah R, Gonzalez-Lara MF, Assman B, Freedman J, Mohan SV. Tocilizumab in patients hospitalized with COVID-19 pneumonia. N Engl J Med. 2021;384(1):20–30. doi: 10.1056/NEJMoa2030340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.WHO Rapid Evidence Appraisal for COVID-19 Therapies (REACT) Working Group. Shankar-Hari M, Vale CL, Godolphin PJ, Fisher D, Higgins JPT, Spiga F, Savovic J, Tierney J, Baron G, Benbenishty JS, Berry LR, Broman N, Cavalcanti AB, Colman R, De Buyser SL, Derde LPG, Domingo P, Omar SF, Fernandez-Cruz A, Feuth T, Garcia F, Garcia-Vicuna R, Gonzalez-Alvaro I, Gordon AC, Haynes R, Hermine O, Horby PW, Horick NK, Kumar K, Lambrecht BN, Landray MJ, Leal L, Lederer DJ, Lorenzi E, Mariette X, Merchante N, Misnan NA, Mohan SV, Nivens MC, Oksi J, Perez-Molina JA, Pizov R, Porcher R, Postma S, Rajasuriar R, Ramanan AV, Ravaud P, Reid PD, Rutgers A, Sancho-Lopez A, Seto TB, Sivapalasingam S, Soin AS, Staplin N, Stone JH, Strohbehn GW, Sunden-Cullberg J, Torre-Cisneros J, Tsai LW, van Hoogstraten H, van Meerten T, Veiga VC, Westerweel PE, Murthy S, Diaz JV, Marshall JC, Sterne JAC. Association between administration of IL-6 antagonists and mortality among patients hospitalized for COVID-19: a meta-analysis. JAMA. 2021;326(6):499–518. doi: 10.1001/jama.2021.11330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Charles P, Elliott MJ, Davis D, Potter A, Kalden JR, Antoni C, Breedveld FC, Smolen JS, Eberl G, deWoody K, Feldmann M, Maini RN. Regulation of cytokines, cytokine inhibitors, and acute-phase proteins following anti-TNF-alpha therapy in rheumatoid arthritis. J Immunol. 1999;163(3):1521–1528. [PubMed] [Google Scholar]
- 129.Paleolog EM, Young S, Stark AC, McCloskey RV, Feldmann M, Maini RN. Modulation of angiogenic vascular endothelial growth factor by tumor necrosis factor alpha and interleukin-1 in rheumatoid arthritis. Arthritis Rheum. 1998;41(7):1258–1265. doi: 10.1002/1529-0131(199807)41:7<1258::AID-ART17>3.0.CO;2-1. [DOI] [PubMed] [Google Scholar]
- 130.Del Valle DM, Kim-Schulze S, Huang HH, Beckmann ND, Nirenberg S, Wang B, Lavin Y, Swartz TH, Madduri D, Stock A, Marron TU, Xie H, Patel M, Tuballes K, Van Oekelen O, Rahman A, Kovatch P, Aberg JA, Schadt E, Jagannath S, Mazumdar M, Charney AW, Firpo-Betancourt A, Mendu DR, Jhang J, Reich D, Sigel K, Cordon-Cardo C, Feldmann M, Parekh S, Merad M, Gnjatic S. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat Med. 2020;26(10):1636–1643. doi: 10.1038/s41591-020-1051-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Steeland S, Libert C, Vandenbroucke RE. A new venue of TNF targeting. Int J Mol Sci. 2018;19(5):1422. doi: 10.3390/ijms19051442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Stallmach A, Kortgen A, Gonnert F, Coldewey SM, Reuken P, Bauer M. Infliximab against severe COVID-19-induced cytokine storm syndrome with organ failure-a cautionary case series. Crit Care. 2020;24(1):444. doi: 10.1186/s13054-020-03158-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Mahase E. COVID-19: Anti-TNF drug adalimumab to be trialled for patients in the community. BMJ. 2020;371:m3847. doi: 10.1136/bmj.m3847. [DOI] [PubMed] [Google Scholar]
- 134.Rondeau JM, Ramage P, Zurini M, Gram H. The molecular mode of action and species specificity of canakinumab, a human monoclonal antibody neutralizing IL-1beta. MAbs. 2015;7(6):1151–1160. doi: 10.1080/19420862.2015.1081323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Dhimolea E. Canakinumab. MAbs. 2010;2(1):3–13. doi: 10.4161/mabs.2.1.10328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Landi L, Ravaglia C, Russo E, Cataleta P, Fusari M, Boschi A, Giannarelli D, Facondini F, Valentini I, Panzini I, Lazzari-Agli L, Bassi P, Marchionni E, Romagnoli R, De Giovanni R, Assirelli M, Baldazzi F, Pieraccini F, Rametta G, Rossi L, Santini L, Valenti I, Cappuzzo F. Blockage of interleukin-1beta with canakinumab in patients with COVID-19. Sci Rep. 2020;10(1):21775. doi: 10.1038/s41598-020-78492-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Ucciferri C, Auricchio A, Di Nicola M, Potere N, Abbate A, Cipollone F, Vecchiet J, Falasca K. Canakinumab in a subgroup of patients with COVID-19. Lancet Rheumatol. 2020;2(8):e457–ee458. doi: 10.1016/S2665-9913(20)30167-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Hamilton JA. GM-CSF-dependent inflammatory pathways. Front Immunol. 2019;10:2055. doi: 10.3389/fimmu.2019.02055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Minter RR, Cohen ES, Wang B, Liang M, Vainshtein I, Rees G, Eghobamien L, Harrison P, Sims DA, Matthews C, Wilkinson T, Monk P, Drinkwater C, Fabri L, Nash A, McCourt M, Jermutus L, Roskos L, Anderson IK, Sleeman MA. Protein engineering and preclinical development of a GM-CSF receptor antibody for the treatment of rheumatoid arthritis. Br J Pharmacol. 2013;168(1):200–211. doi: 10.1111/j.1476-5381.2012.02173.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Cremer PC, Abbate A, Hudock K, McWilliams C, Mehta J, Chang SY, Sheng CC, Van Tassell B, Bonaventura A, Vecchie A, Carey B, Wang Q, Wolski KE, Rajendram P, Duggal A, Wang TS, Paolini JF, Trapnell BC, Group M-CS. Mavrilimumab in patients with severe COVID-19 pneumonia and systemic hyperinflammation (MASH-COVID): an investigator initiated, multicentre, double-blind, randomised, placebo-controlled trial. Lancet Rheumatol. 2021;3(6):e410–e418. doi: 10.1016/S2665-9913(21)00070-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Temesgen Z, Assi M, Shweta FNU, Vergidis P, Rizza SA, Bauer PR, Pickering BW, Razonable RR, Libertin CR, Burger CD, Orenstein R, Vargas HE, Palraj R, Dababneh AS, Chappell G, Chappell D, Ahmed O, Sakemura R, Durrant C, Kenderian SS, Badley AD. GM-CSF neutralization with lenzilumab in severe COVID-19 pneumonia: a case-cohort study. Mayo Clin Proc. 2020;95(11):2382–2394. doi: 10.1016/j.mayocp.2020.08.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Barnum SR. Complement: a primer for the coming therapeutic revolution. Pharmacol Ther. 2017;172:63–72. doi: 10.1016/j.pharmthera.2016.11.014. [DOI] [PubMed] [Google Scholar]
- 143.Stahel PF, Barnum SR. Complement inhibition in coronavirus disease (COVID)-19: a neglected therapeutic option. Front Immunol. 2020;11:1661. doi: 10.3389/fimmu.2020.01661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Cugno M, Meroni PL, Gualtierotti R, Griffini S, Grovetti E, Torri A, Panigada M, Aliberti S, Blasi F, Tedesco F, Peyvandi F. Complement activation in patients with COVID-19: a novel therapeutic target. J Allergy Clin Immunol. 2020;146(1):215–217. doi: 10.1016/j.jaci.2020.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Carvelli J, Demaria O, Vely F, Batista L, Chouaki Benmansour N, Fares J, Carpentier S, Thibult ML, Morel A, Remark R, Andre P, Represa A, Piperoglou C, Explore C-IPHG, Explore C-MIG, Cordier PY, Le Dault E, Guervilly C, Simeone P, Gainnier M, Morel Y, Ebbo M, Schleinitz N, Vivier E. Association of COVID-19 inflammation with activation of the C5a–C5aR1 axis. Nature. 2020;588(7836):146–150. doi: 10.1038/s41586-020-2600-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Noris M, Benigni A, Remuzzi G. The case of complement activation in COVID-19 multiorgan impact. Kidney Int. 2020;98(2):314–322. doi: 10.1016/j.kint.2020.05.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Woodruff TM, Shukla AK. The complement C5a–C5aR1 GPCR axis in COVID-19 therapeutics. Trends Immunol. 2020;41(11):965–967. doi: 10.1016/j.it.2020.09.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Yu J, Yuan X, Chen H, Chaturvedi S, Braunstein EM, Brodsky RA. Direct activation of the alternative complement pathway by SARS-CoV-2 spike proteins is blocked by factor D inhibition. Blood. 2020;136(18):2080–2089. doi: 10.1182/blood.2020008248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Kang S, Yang M, He S, Wang Y, Chen X, Chen YQ, Hong Z, Liu J, Jiang G, Chen Q, Zhou Z, Zhou Z, Huang Z, Huang X, He H, Zheng W, Liao HX, Xiao F, Shan H, Chen S. A SARS-CoV-2 antibody curbs viral nucleocapsid protein-induced complement hyperactivation. Nat Commun. 2021;12(1):2697. doi: 10.1038/s41467-021-23036-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Annane D, Heming N, Grimaldi-Bensouda L, Fremeaux-Bacchi V, Vigan M, Roux AL, Marchal A, Michelon H, Rottman M, Moine P, Garches CCG. Eculizumab as an emergency treatment for adult patients with severe COVID-19 in the intensive care unit: a proof-of-concept study. EClinicalMedicine. 2020;28:100590. doi: 10.1016/j.eclinm.2020.100590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Vlaar APJ, de Bruin S, Busch M, Timmermans S, van Zeggeren IE, Koning R, Ter Horst L, Bulle EB, van Baarle F, van de Poll MCG, Kemper EM, van der Horst ICC, Schultz MJ, Horn J, Paulus F, Bos LD, Wiersinga WJ, Witzenrath M, Rueckinger S, Pilz K, Brouwer MC, Guo RF, Heunks L, van Paassen P, Riedemann NC, van de Beek D. Anti-C5a antibody IFX-1 (vilobelimab) treatment versus best supportive care for patients with severe COVID-19 (PANAMO): an exploratory, open-label, phase 2 randomised controlled trial. Lancet Rheumatol. 2020;2(12):e764–e773. doi: 10.1016/S2665-9913(20)30341-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Vandenberg O, Martiny D, Rochas O, van Belkum A, Kozlakidis Z. Considerations for diagnostic COVID-19 tests. Nat Rev Microbiol. 2021;19(3):171–183. doi: 10.1038/s41579-020-00461-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Beeching NJ, Fletcher TE, Beadsworth MBJ. COVID-19: testing times. BMJ. 2020;369:m1403. doi: 10.1136/bmj.m1403. [DOI] [PubMed] [Google Scholar]
- 154.Wolfel R, Corman VM, Guggemos W, Seilmaier M, Zange S, Muller MA, Niemeyer D, Jones TC, Vollmar P, Rothe C, Hoelscher M, Bleicker T, Brunink S, Schneider J, Ehmann R, Zwirglmaier K, Drosten C, Wendtner C. Virological assessment of hospitalized patients with COVID-2019. Nature. 2020;581(7809):465–469. doi: 10.1038/s41586-020-2196-x. [DOI] [PubMed] [Google Scholar]
- 155.Interim Guidelines for Collecting, Handling, and Testing Clinical Specimens from Persons for Coronavirus Disease 2019 (COVID-19). Centers for Disease Control and Prevention, USA. https://www.cdc.gov/coronavirus/2019-ncov/lab/guidelines-clinical-specimens.html. Accessed 25 Oct 2021.
- 156.Weissleder R, Lee H, Ko J, Pittet MJ. COVID-19 diagnostics in context. Sci Transl Med. 2020 doi: 10.1126/scitranslmed.abc1931. [DOI] [PubMed] [Google Scholar]
- 157.Li M, Wei R, Yang Y, He T, Shen Y, Qi T, Han T, Song Z, Zhu Z, Ma X, Lin Y, Yuan Y, Zhao K, Lu H, Zhou X. Comparing SARS-CoV-2 testing in anterior nasal vestibular swabs vs oropharyngeal swabs. Front Cell Infect Microbiol. 2021;11(598):653794. doi: 10.3389/fcimb.2021.653794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Kilic T, Weissleder R, Lee H. Molecular and immunological diagnostic tests of COVID-19: current status and challenges. iScience. 2020;23(8):101406. doi: 10.1016/j.isci.2020.101406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Li C, Zhao C, Bao J, Tang B, Wang Y, Gu B. Laboratory diagnosis of coronavirus disease-2019 (COVID-19) Clin Chim Acta. 2020;510:35–46. doi: 10.1016/j.cca.2020.06.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Long QX, Liu BZ, Deng HJ, Wu GC, Deng K, Chen YK, Liao P, Qiu JF, Lin Y, Cai XF, Wang DQ, Hu Y, Ren JH, Tang N, Xu YY, Yu LH, Mo Z, Gong F, Zhang XL, Tian WG, Hu L, Zhang XX, Xiang JL, Du HX, Liu HW, Lang CH, Luo XH, Wu SB, Cui XP, Zhou Z, Zhu MM, Wang J, Xue CJ, Li XF, Wang L, Li ZJ, Wang K, Niu CC, Yang QJ, Tang XJ, Zhang Y, Liu XM, Li JJ, Zhang DC, Zhang F, Liu P, Yuan J, Li Q, Hu JL, Chen J, Huang AL. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat Med. 2020;26(6):845–848. doi: 10.1038/s41591-020-0897-1. [DOI] [PubMed] [Google Scholar]
- 161.D'Cruz RJ, Currier AW, Sampson VB. Laboratory testing methods for novel severe acute respiratory syndrome-coronavirus-2 (SARS-CoV-2) Front Cell Dev Biol. 2020;8:468. doi: 10.3389/fcell.2020.00468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Hansen CH, Michlmayr D, Gubbels SM, Molbak K, Ethelberg S. Assessment of protection against reinfection with SARS-CoV-2 among 4 million PCR-tested individuals in Denmark in 2020: a population-level observational study. Lancet. 2021;397(10280):1204–1212. doi: 10.1016/S0140-6736(21)00575-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Darwish IA. Immunoassay methods and their applications in pharmaceutical analysis: basic methodology and recent advances. Int J Biomed Sci. 2006;2(3):217–235. [PMC free article] [PubMed] [Google Scholar]
- 164.Cong Y, Ulasli M, Schepers H, Mauthe M, V'Kovski P, Kriegenburg F, Thiel V, de Haan CAM, Reggiori F. Nucleocapsid protein recruitment to replication-transcription complexes plays a crucial role in coronaviral life cycle. J Virol. 2020;94(4):e01925–e11919. doi: 10.1128/JVI.01925-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Tai W, He L, Zhang X, Pu J, Voronin D, Jiang S, Zhou Y, Du L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol. 2020;17(6):613–620. doi: 10.1038/s41423-020-0400-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Vashist SK. In vitro diagnostic assays for COVID-19: recent advances and emerging trends. Diagnostics (Basel) 2020;10(4):202. doi: 10.3390/diagnostics10040202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Espejo AP, Akgun Y, Al Mana AF, Tjendra Y, Millan NC, Gomez-Fernandez C, Cray C. Review of current advances in serologic testing for COVID-19. Am J Clin Pathol. 2020;154(3):293–304. doi: 10.1093/ajcp/aqaa112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Amanat F, Stadlbauer D, Strohmeier S, Nguyen THO, Chromikova V, McMahon M, Jiang K, Arunkumar GA, Jurczyszak D, Polanco J, Bermudez-Gonzalez M, Kleiner G, Aydillo T, Miorin L, Fierer DS, Lugo LA, Kojic EM, Stoever J, Liu STH, Cunningham-Rundles C, Felgner PL, Moran T, Garcia-Sastre A, Caplivski D, Cheng AC, Kedzierska K, Vapalahti O, Hepojoki JM, Simon V, Krammer F. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat Med. 2020;26(7):1033–1036. doi: 10.1038/s41591-020-0913-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Guo JY, Liu IJ, Lin HT, Wang MJ, Chang YL, Lin SC, Liao MY, Hsu WC, Lin YL, Liao JC, Wu HC. Identification of COVID-19 B-cell epitopes with phage-displayed peptide library. J Biomed Sci. 2021;28(1):43. doi: 10.1186/s12929-021-00740-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.To KK, Tsang OT, Leung WS, Tam AR, Wu TC, Lung DC, Yip CC, Cai JP, Chan JM, Chik TS, Lau DP, Choi CY, Chen LL, Chan WM, Chan KH, Ip JD, Ng AC, Poon RW, Luo CT, Cheng VC, Chan JF, Hung IF, Chen Z, Chen H, Yuen KY. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis. 2020;20(5):565–574. doi: 10.1016/S1473-3099(20)30196-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Okba NMA, Muller MA, Li W, Wang C, GeurtsvanKessel CH, Corman VM, Lamers MM, Sikkema RS, de Bruin E, Chandler FD, Yazdanpanah Y, Le Hingrat Q, Descamps D, Houhou-Fidouh N, Reusken C, Bosch BJ, Drosten C, Koopmans MPG, Haagmans BL. Severe acute respiratory syndrome coronavirus 2-specific antibody responses in coronavirus disease patients. Emerg Infect Dis. 2020;26(7):1478–1488. doi: 10.3201/eid2607.200841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Kontou PI, Braliou GG, Dimou NL, Nikolopoulos G, Bagos PG. Antibody tests in detecting SARS-CoV-2 infection: a meta-analysis. Diagnostics (Basel) 2020;10(5):319. doi: 10.3390/diagnostics10050319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Suthar MS, Zimmerman MG, Kauffman RC, Mantus G, Linderman SL, Hudson WH, Vanderheiden A, Nyhoff L, Davis CW, Adekunle O, Affer M, Sherman M, Reynolds S, Verkerke HP, Alter DN, Guarner J, Bryksin J, Horwath MC, Arthur CM, Saakadze N, Smith GH, Edupuganti S, Scherer EM, Hellmeister K, Cheng A, Morales JA, Neish AS, Stowell SR, Frank F, Ortlund E, Anderson EJ, Menachery VD, Rouphael N, Mehta AK, Stephens DS, Ahmed R, Roback JD, Wrammert J. Rapid generation of neutralizing antibody responses in COVID-19 patients. Cell Rep Med. 2020;1(3):100040. doi: 10.1016/j.xcrm.2020.100040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Kohmer N, Westhaus S, Ruhl C, Ciesek S, Rabenau HF. Clinical performance of different SARS-CoV-2 IgG antibody tests. J Med Virol. 2020;92(10):2243–2247. doi: 10.1002/jmv.26145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Theel ES, Harring J, Hilgart H, Granger D. Performance characteristics of four high-throughput immunoassays for detection of IgG antibodies against SARS-CoV-2. J Clin Microbiol. 2020 doi: 10.1128/JCM.01243-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Zhu Y, Yu D, Han Y, Yan H, Chong H, Ren L, Wang J, Li T, He Y. Cross-reactive neutralization of SARS-CoV-2 by serum antibodies from recovered SARS patients and immunized animals. Sci Adv. 2020 doi: 10.1126/sciadv.abc9999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Aydin S. A short history, principles, and types of ELISA, and our laboratory experience with peptide/protein analyses using ELISA. Peptides. 2015;72:4–15. doi: 10.1016/j.peptides.2015.04.012. [DOI] [PubMed] [Google Scholar]
- 178.Beavis KG, Matushek SM, Abeleda APF, Bethel C, Hunt C, Gillen S, Moran A, Tesic V. Evaluation of the EUROIMMUN anti-SARS-CoV-2 ELISA assay for detection of IgA and IgG antibodies. J Clin Virol. 2020;129:104468. doi: 10.1016/j.jcv.2020.104468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Ghaffari A, Meurant R, Ardakani A. COVID-19 serological tests: how well do they actually perform? Diagnostics (Basel) 2020;10(7):453. doi: 10.3390/diagnostics10070453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Lisboa BM, Tavaziva G, Abidi SK, Campbell JR, Haraoui LP, Johnston JC, Lan Z, Law S, MacLean E, Trajman A, Menzies D, Benedetti A, Ahmad KF. Diagnostic accuracy of serological tests for COVID-19: systematic review and meta-analysis. BMJ. 2020;370:m2516. doi: 10.1136/bmj.m2516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Posthuma-Trumpie GA, Korf J, van Amerongen A. Lateral flow (immuno)assay: its strengths, weaknesses, opportunities and threats. A literature survey. Anal Bioanal Chem. 2009;393(2):569–582. doi: 10.1007/s00216-008-2287-2. [DOI] [PubMed] [Google Scholar]
- 182.Lu RM, Ko SH, Chen WY, Chang YL, Lin HT, Wu HC. Monoclonal antibodies against nucleocapsid protein of SARS-CoV-2 variants for detection of COVID-19. Int J Mol Sci. 2021;22(22):12412. doi: 10.3390/ijms222212412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Xiong X, Qu K, Ciazynska KA, Hosmillo M, Carter AP, Ebrahimi S, Ke Z, Scheres SHW, Bergamaschi L, Grice GL, Zhang Y, Collaboration C-NC-B, Nathan JA, Baker S, James LC, Baxendale HE, Goodfellow I, Doffinger R, Briggs JAG. A thermostable, closed SARS-CoV-2 spike protein trimer. Nat Struct Mol Biol. 2020;27(10):934–941. doi: 10.1038/s41594-020-0478-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.Ke Z, Oton J, Qu K, Cortese M, Zila V, McKeane L, Nakane T, Zivanov J, Neufeldt CJ, Cerikan B, Lu JM, Peukes J, Xiong X, Krausslich HG, Scheres SHW, Bartenschlager R, Briggs JAG. Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature. 2020;588(7838):498–502. doi: 10.1038/s41586-020-2665-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Xu C, Wang Y, Liu C, Zhang C, Han W, Hong X, Wang Y, Hong Q, Wang S, Zhao Q, Wang Y, Yang Y, Chen K, Zheng W, Kong L, Wang F, Zuo Q, Huang Z, Cong Y. Conformational dynamics of SARS-CoV-2 trimeric spike glycoprotein in complex with receptor ACE2 revealed by cryo-EM. Sci Adv. 2021;7(1):eabe5575. doi: 10.1126/sciadv.abe5575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Zhang J, Cai Y, Xiao T, Lu J, Peng H, Sterling SM, Walsh RM, Jr, Rits-Volloch S, Zhu H, Woosley AN, Yang W, Sliz P, Chen B. Structural impact on SARS-CoV-2 spike protein by D614G substitution. Science. 2021;372(6541):525–530. doi: 10.1126/science.abf2303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Cai Y, Zhang J, Xiao T, Peng H, Sterling SM, Walsh RM, Jr, Rawson S, Rits-Volloch S, Chen B. Distinct conformational states of SARS-CoV-2 spike protein. Science. 2020;369(6511):1586–1592. doi: 10.1126/science.abd4251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Lan J, Ge J, Yu J, Shan S, Zhou H, Fan S, Zhang Q, Shi X, Wang Q, Zhang L, Wang X. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. 2020;581(7807):215–220. doi: 10.1038/s41586-020-2180-5. [DOI] [PubMed] [Google Scholar]
- 189.Shang J, Ye G, Shi K, Wan Y, Luo C, Aihara H, Geng Q, Auerbach A, Li F. Structural basis of receptor recognition by SARS-CoV-2. Nature. 2020;581(7807):221–224. doi: 10.1038/s41586-020-2179-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Yan R, Zhang Y, Li Y, Xia L, Guo Y, Zhou Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science. 2020;367(6485):1444–1448. doi: 10.1126/science.abb2762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Wang Z, Schmidt F, Weisblum Y, Muecksch F, Barnes CO, Finkin S, Schaefer-Babajew D, Cipolla M, Gaebler C, Lieberman JA, Oliveira TY, Yang Z, Abernathy ME, Huey-Tubman KE, Hurley A, Turroja M, West KA, Gordon K, Millard KG, Ramos V, Da Silva J, Xu J, Colbert RA, Patel R, Dizon J, Unson-O'Brien C, Shimeliovich I, Gazumyan A, Caskey M, Bjorkman PJ, Casellas R, Hatziioannou T, Bieniasz PD, Nussenzweig MC. mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants. Nature. 2021;592(7855):616–622. doi: 10.1038/s41586-021-03324-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Li Q, Wu J, Nie J, Zhang L, Hao H, Liu S, Zhao C, Zhang Q, Liu H, Nie L, Qin H, Wang M, Lu Q, Li X, Sun Q, Liu J, Zhang L, Li X, Huang W, Wang Y. The impact of mutations in SARS-CoV-2 spike on viral infectivity and antigenicity. Cell. 2020;182(5):1284–1294 e1289. doi: 10.1016/j.cell.2020.07.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Liu Z, VanBlargan LA, Bloyet LM, Rothlauf PW, Chen RE, Stumpf S, Zhao H, Errico JM, Theel ES, Liebeskind MJ, Alford B, Buchser WJ, Ellebedy AH, Fremont DH, Diamond MS, Whelan SPJ. Identification of SARS-CoV-2 spike mutations that attenuate monoclonal and serum antibody neutralization. Cell Host Microbe. 2021;29(3):477–488 e474. doi: 10.1016/j.chom.2021.01.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Cao Y, Su B, Guo X, Sun W, Deng Y, Bao L, Zhu Q, Zhang X, Zheng Y, Geng C, Chai X, He R, Li X, Lv Q, Zhu H, Deng W, Xu Y, Wang Y, Qiao L, Tan Y, Song L, Wang G, Du X, Gao N, Liu J, Xiao J, Su XD, Du Z, Feng Y, Qin C, Qin C, Jin R, Xie XS. Potent neutralizing antibodies against SARS-CoV-2 identified by high-throughput single-cell sequencing of convalescent patients' B cells. Cell. 2020;182(1):73–84 e16. doi: 10.1016/j.cell.2020.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Yurkovetskiy L, Wang X, Pascal KE, Tomkins-Tinch C, Nyalile TP, Wang Y, Baum A, Diehl WE, Dauphin A, Carbone C, Veinotte K, Egri SB, Schaffner SF, Lemieux JE, Munro JB, Rafique A, Barve A, Sabeti PC, Kyratsous CA, Dudkina NV, Shen K, Luban J. Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell. 2020;183(3):739–751 e738. doi: 10.1016/j.cell.2020.09.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Pramanick I, Sengupta N, Mishra S, Pandey S, Girish N, Das A, Dutta S. Conformational flexibility and structural variability of SARS-CoV2 S protein. Structure. 2021;29(8):834–845 e835. doi: 10.1016/j.str.2021.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Barnes CO, Jette CA, Abernathy ME, Dam KA, Esswein SR, Gristick HB, Malyutin AG, Sharaf NG, Huey-Tubman KE, Lee YE, Robbiani DF, Nussenzweig MC, West AP, Jr, Bjorkman PJ. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature. 2020;588(7839):682–687. doi: 10.1038/s41586-020-2852-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Hodcroft EB, Zuber M, Nadeau S, Vaughan TG, Crawford KHD, Althaus CL, Reichmuth ML, Bowen JE, Walls AC, Corti D, Bloom JD, Veesler D, Mateo D, Hernando A, Comas I, Gonzalez-Candelas F, Seq C-SC, Stadler T, Neher RA. Spread of a SARS-CoV-2 variant through Europe in the summer of 2020. Nature. 2021;595(7869):707–712. doi: 10.1038/s41586-021-03677-y. [DOI] [PubMed] [Google Scholar]
- 199.Davies NG, Jarvis CI, Group CC-W, Edmunds WJ, Jewell NP, Diaz-Ordaz K, Keogh RH. Increased mortality in community-tested cases of SARS-CoV-2 lineage B.1.1.7. Nature. 2021;593(7858):270–274. doi: 10.1038/s41586-021-03426-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Challen R, Brooks-Pollock E, Read JM, Dyson L, Tsaneva-Atanasova K, Danon L. Risk of mortality in patients infected with SARS-CoV-2 variant of concern 202012/1: matched cohort study. BMJ. 2021;372:n579. doi: 10.1136/bmj.n579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Ratcliff J, Nguyen D, Fish M, Rynne J, Jennings A, Williams S, Al-Beidh F, Bonsall D, Evans A, Golubchik T, Gordon AC, Lamikanra A, Tsang P, Ciccone NA, Leuscher U, Slack W, Laing E, Mouncey PR, Ziyenge S, Oliveira M, Ploeg R, Rowan KM, Shankar-Hari M, Roberts DJ, Menon DK, Estcourt L, Simmonds P, Harvala H, Investigators R-CIDU. Virological characterization of critically ill patients with COVID-19 in the United Kingdom: interactions of viral load, antibody status, and B.1.1.7 infection. J Infect Dis. 2021;224(4):595–605. doi: 10.1093/infdis/jiab283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Leung K, Shum MH, Leung GM, Lam TT, Wu JT. Early transmissibility assessment of the N501Y mutant strains of SARS-CoV-2 in the United Kingdom, October to November 2020. Euro Surveill. 2021 doi: 10.2807/1560-7917.ES.2020.26.1.2002106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.Yi C, Sun X, Ye J, Ding L, Liu M, Yang Z, Lu X, Zhang Y, Ma L, Gu W, Qu A, Xu J, Shi Z, Ling Z, Sun B. Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies. Cell Mol Immunol. 2020;17(6):621–630. doi: 10.1038/s41423-020-0458-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Gu H, Chen Q, Yang G, He L, Fan H, Deng YQ, Wang Y, Teng Y, Zhao Z, Cui Y, Li Y, Li XF, Li J, Zhang NN, Yang X, Chen S, Guo Y, Zhao G, Wang X, Luo DY, Wang H, Yang X, Li Y, Han G, He Y, Zhou X, Geng S, Sheng X, Jiang S, Sun S, Qin CF, Zhou Y. Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy. Science. 2020;369(6511):1603–1607. doi: 10.1126/science.abc4730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 205.Chan KK, Tan TJC, Narayanan KK, Procko E. An engineered decoy receptor for SARS-CoV-2 broadly binds protein S sequence variants. Sci Adv. 2021 doi: 10.1126/sciadv.abf1738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Yuan M, Huang D, Lee CD, Wu NC, Jackson AM, Zhu X, Liu H, Peng L, van Gils MJ, Sanders RW, Burton DR, Reincke SM, Pruss H, Kreye J, Nemazee D, Ward AB, Wilson IA. Structural and functional ramifications of antigenic drift in recent SARS-CoV-2 variants. Science. 2021;373(6556):818–823. doi: 10.1126/science.abh1139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.McCarthy KR, Rennick LJ, Nambulli S, Robinson-McCarthy LR, Bain WG, Haidar G, Duprex WP. Recurrent deletions in the SARS-CoV-2 spike glycoprotein drive antibody escape. Science. 2021;371(6534):1139–1142. doi: 10.1126/science.abf6950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208.Kemp SA, Collier DA, Datir RP, Ferreira I, Gayed S, Jahun A, Hosmillo M, Rees-Spear C, Mlcochova P, Lumb IU, Roberts DJ, Chandra A, Temperton N, Collaboration C-NBC, Consortium C-GU, Sharrocks K, Blane E, Modis Y, Leigh KE, Briggs JAG, van Gils MJ, Smith KGC, Bradley JR, Smith C, Doffinger R, Ceron-Gutierrez L, Barcenas-Morales G, Pollock DD, Goldstein RA, Smielewska A, Skittrall JP, Gouliouris T, Goodfellow IG, Gkrania-Klotsas E, Illingworth CJR, McCoy LE, Gupta RK. SARS-CoV-2 evolution during treatment of chronic infection. Nature. 2021;592(7853):277–282. doi: 10.1038/s41586-021-03291-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Shen X, Tang H, McDanal C, Wagh K, Fischer W, Theiler J, Yoon H, Li D, Haynes BF, Sanders KO, Gnanakaran S, Hengartner N, Pajon R, Smith G, Glenn GM, Korber B, Montefiori DC. SARS-CoV-2 variant B.1.1.7 is susceptible to neutralizing antibodies elicited by ancestral spike vaccines. Cell Host Microbe. 2021;29(4):529–539 e523. doi: 10.1016/j.chom.2021.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Lubinski B, Tang T, Daniel S, Jaimes JA, Whittaker GR. Functional evaluation of proteolytic activation for the SARS-CoV-2 variant B.1.1.7: role of the P681H mutation. bioRxiv. 2021 doi: 10.2139/ssrn.3889709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Tegally H, Wilkinson E, Giovanetti M, Iranzadeh A, Fonseca V, Giandhari J, Doolabh D, Pillay S, San EJ, Msomi N, Mlisana K, von Gottberg A, Walaza S, Allam M, Ismail A, Mohale T, Glass AJ, Engelbrecht S, Van Zyl G, Preiser W, Petruccione F, Sigal A, Hardie D, Marais G, Hsiao NY, Korsman S, Davies MA, Tyers L, Mudau I, York D, Maslo C, Goedhals D, Abrahams S, Laguda-Akingba O, Alisoltani-Dehkordi A, Godzik A, Wibmer CK, Sewell BT, Lourenco J, Alcantara LCJ, Kosakovsky Pond SL, Weaver S, Martin D, Lessells RJ, Bhiman JN, Williamson C, de Oliveira T. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature. 2021;592(7854):438–443. doi: 10.1038/s41586-021-03402-9. [DOI] [PubMed] [Google Scholar]
- 212.Roquebert B, Trombert-Paolantoni S, Haim-Boukobza S, Lecorche E, Verdurme L, Foulongne V, Sofonea MT, Alizon S. The SARS-CoV-2 B1351 lineage (VOC beta) is outgrowing the B.1.1.7 lineage (VOC alpha) in some French regions in April 2021. Euro Surveill. 2021 doi: 10.2807/1560-7917.ES.2021.26.23.2100447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Callaway E. Delta coronavirus variant: scientists brace for impact. Nature. 2021;595(7865):17–18. doi: 10.1038/d41586-021-01696-3. [DOI] [PubMed] [Google Scholar]
- 214.Campbell F, Archer B, Laurenson-Schafer H, Jinnai Y, Konings F, Batra N, Pavlin B, Vandemaele K, Van Kerkhove MD, Jombart T, Morgan O, le Polain de Waroux O. Increased transmissibility and global spread of SARS-CoV-2 variants of concern as at June 2021. Euro Surveill. 2021 doi: 10.2807/1560-7917.ES.2021.26.24.2100509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Planas D, Veyer D, Baidaliuk A, Staropoli I, Guivel-Benhassine F, Rajah MM, Planchais C, Porrot F, Robillard N, Puech J, Prot M, Gallais F, Gantner P, Velay A, Le Guen J, Kassis-Chikhani N, Edriss D, Belec L, Seve A, Courtellemont L, Pere H, Hocqueloux L, Fafi-Kremer S, Prazuck T, Mouquet H, Bruel T, Simon-Loriere E, Rey FA, Schwartz O. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature. 2021;596(7871):276–280. doi: 10.1038/s41586-021-03777-9. [DOI] [PubMed] [Google Scholar]
- 216.Liu C, Ginn HM, Dejnirattisai W, Supasa P, Wang B, Tuekprakhon A, Nutalai R, Zhou D, Mentzer AJ, Zhao Y, Duyvesteyn HME, Lopez-Camacho C, Slon-Campos J, Walter TS, Skelly D, Johnson SA, Ritter TG, Mason C, Costa Clemens SA, Gomes Naveca F, Nascimento V, Nascimento F, Fernandes da Costa C, Resende PC, Pauvolid-Correa A, Siqueira MM, Dold C, Temperton N, Dong T, Pollard AJ, Knight JC, Crook D, Lambe T, Clutterbuck E, Bibi S, Flaxman A, Bittaye M, Belij-Rammerstorfer S, Gilbert SC, Malik T, Carroll MW, Klenerman P, Barnes E, Dunachie SJ, Baillie V, Serafin N, Ditse Z, Da Silva K, Paterson NG, Williams MA, Hall DR, Madhi S, Nunes MC, Goulder P, Fry EE, Mongkolsapaya J, Ren J, Stuart DI, Screaton GR. Reduced neutralization of SARS-CoV-2 B.1.617 by vaccine and convalescent serum. Cell. 2021;184(16):4220–42364213. doi: 10.1016/j.cell.2021.06.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Latif AA, Mullen JL, Alkuzweny M, Tsueng G, Cano M, Haag E, Zhou J, Zeller M, Hufbauer E, Matteson N, Wu C, Andersen KG, Su AI, Gangavarapu K, Hughes LD, Biology tCfVS. Lineage comparison. https://outbreak.info/compare-lineages?pango=Omicron&gene=S&threshold=0.2. Accessed 1 Dec 2021.
- 218.Greaney AJ, Starr TN, Gilchuk P, Zost SJ, Binshtein E, Loes AN, Hilton SK, Huddleston J, Eguia R, Crawford KHD, Dingens AS, Nargi RS, Sutton RE, Suryadevara N, Rothlauf PW, Liu Z, Whelan SPJ, Carnahan RH, Crowe JE, Jr, Bloom JD. Complete mapping of mutations to the SARS-CoV-2 spike receptor-binding domain that escape antibody recognition. Cell Host Microbe. 2021;29(1):44–57 e49. doi: 10.1016/j.chom.2020.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.Muecksch F, Weisblum Y, Barnes CO, Schmidt F, Schaefer-Babajew D, Wang Z, Lorenzi JCC, Flyak AI, DeLaitsch AT, Huey-Tubman KE, Hou S, Schiffer CA, Gaebler C, Da Silva J, Poston D, Finkin S, Cho A, Cipolla M, Oliveira TY, Millard KG, Ramos V, Gazumyan A, Rutkowska M, Caskey M, Nussenzweig MC, Bjorkman PJ, Hatziioannou T, Bieniasz PD. Affinity maturation of SARS-CoV-2 neutralizing antibodies confers potency, breadth, and resilience to viral escape mutations. Immunity. 2021;54(8):1853–1868 e1857. doi: 10.1016/j.immuni.2021.07.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Gaebler C, Wang Z, Lorenzi JCC, Muecksch F, Finkin S, Tokuyama M, Cho A, Jankovic M, Schaefer-Babajew D, Oliveira TY, Cipolla M, Viant C, Barnes CO, Bram Y, Breton G, Hagglof T, Mendoza P, Hurley A, Turroja M, Gordon K, Millard KG, Ramos V, Schmidt F, Weisblum Y, Jha D, Tankelevich M, Martinez-Delgado G, Yee J, Patel R, Dizon J, Unson-O'Brien C, Shimeliovich I, Robbiani DF, Zhao Z, Gazumyan A, Schwartz RE, Hatziioannou T, Bjorkman PJ, Mehandru S, Bieniasz PD, Caskey M, Nussenzweig MC. Evolution of antibody immunity to SARS-CoV-2. Nature. 2021;591(7851):639–644. doi: 10.1038/s41586-021-03207-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221.Sakharkar M, Rappazzo CG, Wieland-Alter WF, Hsieh CL, Wrapp D, Esterman ES, Kaku CI, Wec AZ, Geoghegan JC, McLellan JS, Connor RI, Wright PF, Walker LM. Prolonged evolution of the human B cell response to SARS-CoV-2 infection. Sci Immunol. 2021 doi: 10.1126/sciimmunol.abg6916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Sokal A, Chappert P, Barba-Spaeth G, Roeser A, Fourati S, Azzaoui I, Vandenberghe A, Fernandez I, Meola A, Bouvier-Alias M, Crickx E, Beldi-Ferchiou A, Hue S, Languille L, Michel M, Baloul S, Noizat-Pirenne F, Luka M, Megret J, Menager M, Pawlotsky JM, Fillatreau S, Rey FA, Weill JC, Reynaud CA, Mahevas M. Maturation and persistence of the anti-SARS-CoV-2 memory B cell response. Cell. 2021;184(5):1201–1213 e1214. doi: 10.1016/j.cell.2021.01.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.COVID-19 Dashboard. World health organization. https://covid19.who.int/. Accessed 28 Dec 2021.
- 224.Laboratory-Confirmed COVID-19-Associated Hospitalizations. Centers for Disease Control and Prevention, USA. https://gis.cdc.gov/grasp/covidnet/covid19_3.html. Accessed 18 Feb 2021.
- 225.Pak A, Adegboye OA, Adekunle AI, Rahman KM, McBryde ES, Eisen DP. Economic consequences of the COVID-19 outbreak: the need for epidemic preparedness. Front Public Health. 2020;8:241. doi: 10.3389/fpubh.2020.00241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Sharma O, Sultan AA, Ding H, Triggle CR. A review of the progress and challenges of developing a vaccine for COVID-19. Front Immunol. 2020;11:585354. doi: 10.3389/fimmu.2020.585354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227.Hacisuleyman E, Hale C, Saito Y, Blachere NE, Bergh M, Conlon EG, Schaefer-Babajew DJ, DaSilva J, Muecksch F, Gaebler C, Lifton R, Nussenzweig MC, Hatziioannou T, Bieniasz PD, Darnell RB. Vaccine breakthrough infections with SARS-CoV-2 variants. N Engl J Med. 2021;384(23):2212–2218. doi: 10.1056/NEJMoa2105000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Brosh-Nissimov T, Orenbuch-Harroch E, Chowers M, Elbaz M, Nesher L, Stein M, Maor Y, Cohen R, Hussein K, Weinberger M, Zimhony O, Chazan B, Najjar R, Zayyad H, Rahav G, Wiener-Well Y. BNT162b2 vaccine breakthrough: clinical characteristics of 152 fully vaccinated hospitalized COVID-19 patients in Israel. Clin Microbiol Infect. 2021 doi: 10.1016/j.cmi.2021.06.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229.Lange B, Gerigk M, Tenenbaum T. Breakthrough infections in BNT162b2-vaccinated health care workers. N Engl J Med. 2021;385(12):1145–1146. doi: 10.1056/NEJMc2108076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Callaway E. Heavily mutated Omicron variant puts scientists on alert. Nature. 2021 doi: 10.1038/d41586-021-03552-w. [DOI] [PubMed] [Google Scholar]
- 231.Philippidis A. Top 7 Best Selling COVID-19 Vaccines and Drugs of 2020. In Genetic Engineering & Biotechnology News. Mary Ann Liebert, Inc. https://www.genengnews.com/a-lists/top-7-best-selling-covid-19-vaccines-and-drugs-of-2020/. Accessed 25 Feb 2021.
- 232.Regeneron Reports Third Quarter 2021 Financial and Operating Results. PR Newswire Association LLC. https://www.prnewswire.com/news-releases/regeneron-reports-third-quarter-2021-financial-and-operating-results-301416042.html. Accessed 4 Nov 2021.
- 233.Tea F, Ospina Stella A, Aggarwal A, Ross Darley D, Pilli D, Vitale D, Merheb V, Lee FXZ, Cunningham P, Walker GJ, Fichter C, Brown DA, Rawlinson WD, Isaacs SR, Mathivanan V, Hoffmann M, Pohlman S, Mazigi O, Christ D, Dwyer DE, Rockett RJ, Sintchenko V, Hoad VC, Irving DO, Dore GJ, Gosbell IB, Kelleher AD, Matthews GV, Brilot F, Turville SG. SARS-CoV-2 neutralizing antibodies: longevity, breadth, and evasion by emerging viral variants. PLoS Med. 2021;18(7):e1003656. doi: 10.1371/journal.pmed.1003656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.McAndrews KM, Dowlatshahi DP, Dai J, Becker LM, Hensel J, Snowden LM, Leveille JM, Brunner MR, Holden KW, Hopkins NS, Harris AM, Kumpati J, Whitt MA, Lee JJ, Ostrosky-Zeichner LL, Papanna R, LeBleu VS, Allison JP, Kalluri R. Heterogeneous antibodies against SARS-CoV-2 spike receptor binding domain and nucleocapsid with implications for COVID-19 immunity. JCI Insight. 2020;5(18):e142386. doi: 10.1172/jci.insight.142386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235.Thomson EC, Rosen LE, Shepherd JG, Spreafico R, da Silva Filipe A, Wojcechowskyj JA, Davis C, Piccoli L, Pascall DJ, Dillen J, Lytras S, Czudnochowski N, Shah R, Meury M, Jesudason N, De Marco A, Li K, Bassi J, O'Toole A, Pinto D, Colquhoun RM, Culap K, Jackson B, Zatta F, Rambaut A, Jaconi S, Sreenu VB, Nix J, Zhang I, Jarrett RF, Glass WG, Beltramello M, Nomikou K, Pizzuto M, Tong L, Cameroni E, Croll TI, Johnson N, Di Iulio J, Wickenhagen A, Ceschi A, Harbison AM, Mair D, Ferrari P, Smollett K, Sallusto F, Carmichael S, Garzoni C, Nichols J, Galli M, Hughes J, Riva A, Ho A, Schiuma M, Semple MG, Openshaw PJM, Fadda E, Baillie JK, Chodera JD, Investigators IC, Consortium CGU, Rihn SJ, Lycett SJ, Virgin HW, Telenti A, Corti D, Robertson DL, Snell G. Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity. Cell. 2021;184(5):1171–1187 e1120. doi: 10.1016/j.cell.2021.01.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 236.Ku Z, Xie X, Davidson E, Ye X, Su H, Menachery VD, Li Y, Yuan Z, Zhang X, Muruato AE, Escuer AGI, Tyrell B, Doolan K, Doranz BJ, Wrapp D, Bates PF, McLellan JS, Weiss SR, Zhang N, Shi PY, An Z. Molecular determinants and mechanism for antibody cocktail preventing SARS-CoV-2 escape. Nat Commun. 2021;12(1):469. doi: 10.1038/s41467-020-20789-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 237.Guglielmi G. Rapid coronavirus tests: a guide for the perplexed. Nature. 2021;590(7845):202–205. doi: 10.1038/d41586-021-00332-4. [DOI] [PubMed] [Google Scholar]
- 238.Oyewole AO, Barrass L, Robertson EG, Woltmann J, O'Keefe H, Sarpal H, Dangova K, Richmond C, Craig D. COVID-19 impact on diagnostic innovations: emerging trends and implications. Diagnostics (Basel). 2021 doi: 10.3390/diagnostics11020182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 239.Neilan AM, Losina E, Bangs AC, Flanagan C, Panella C, Eskibozkurt GE, Mohareb A, Hyle EP, Scott JA, Weinstein MC, Siedner MJ, Reddy KP, Harling G, Freedberg KA, Shebl FM, Kazemian P, Ciaranello AL. Clinical impact, costs, and cost-effectiveness of expanded severe acute respiratory syndrome coronavirus 2 testing in Massachusetts. Clin Infect Dis. 2021;73(9):e2908–e2917. [DOI] [PMC free article] [PubMed]
- 240.Ahmad FB, Anderson RN. The leading causes of death in the US for 2020. JAMA. 2021;325(18):1829–1830. doi: 10.1001/jama.2021.5469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Forman R, Shah S, Jeurissen P, Jit M, Mossialos E. COVID-19 vaccine challenges: what have we learned so far and what remains to be done? Health Policy. 2021;125(5):553–567. doi: 10.1016/j.healthpol.2021.03.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242.Lamb YN. BNT162b2 mRNA COVID-19 vaccine: first approval. Drugs. 2021;81(4):495–501. doi: 10.1007/s40265-021-01480-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 243.Martonosi SE, Behzad B, Cummings K. Pricing the COVID-19 vaccine: a mathematical approach. Omega. 2021;103:102451. doi: 10.1016/j.omega.2021.102451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 244.Wouters OJ, Shadlen KC, Salcher-Konrad M, Pollard AJ, Larson HJ, Teerawattananon Y, Jit M. Challenges in ensuring global access to COVID-19 vaccines: production, affordability, allocation, and deployment. Lancet. 2021;397(10278):1023–1034. doi: 10.1016/S0140-6736(21)00306-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 245.Bowie A. Regeneron announces U.S. Government agreement to purchase additional covid-19 antibody cocktail doses. Regeneron Pharmaceuticals Inc., https://investor.regeneron.com/index.php/news-releases/news-release-details/regeneron-announces-us-government-agreement-purchase-additional. Accessed 12 Jan 2021.
- 246.Haagmans BL, Noack D, Okba NMA, Li W, Wang C, Bestebroer T, de Vries R, Herfst S, de Meulder D, Verveer E, van Run P, Lamers MM, Rijnders B, Rokx C, van Kuppeveld F, Grosveld F, Drabek D, Geurts van Kessel C, Koopmans M, Bosch BJ, Kuiken T, Rockx B. SARS-CoV-2 neutralizing human antibodies protect against lower respiratory tract disease in a hamster model. J Infect Dis. 2021;223(12):2020–2028. doi: 10.1093/infdis/jiab289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Alsoussi WB, Turner JS, Case JB, Zhao H, Schmitz AJ, Zhou JQ, Chen RE, Lei T, Rizk AA, McIntire KM, Winkler ES, Fox JM, Kafai NM, Thackray LB, Hassan AO, Amanat F, Krammer F, Watson CT, Kleinstein SH, Fremont DH, Diamond MS, Ellebedy AH. A potently neutralizing antibody protects mice against SARS-CoV-2 Infection. J Immunol. 2020;205(4):915–922. doi: 10.4049/jimmunol.2000583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 248.Maisonnasse P, Aldon Y, Marc A, Marlin R, Dereuddre-Bosquet N, Kuzmina NA, Freyn AW, Snitselaar JL, Gonçalves A, Caniels TG, Burger JA, Poniman M, Bontjer I, Chesnais V, Diry S, Iershov A, Ronk AJ, Jangra S, Rathnasinghe R, Brouwer PJM, Bijl TPL, van Schooten J, Brinkkemper M, Liu H, Yuan M, Mire CE, van Breemen MJ, Contreras V, Naninck T, Lemaître J, Kahlaoui N, Relouzat F, Chapon C, Ho Tsong Fang R, McDanal C, Osei-Twum M, St-Amant N, Gagnon L, Montefiori DC, Wilson IA, Ginoux E, de Bree GJ, García-Sastre A, Schotsaert M, Coughlan L, Bukreyev A, van der Werf S, Guedj J, Sanders RW, van Gils MJ, Le Grand R. COVA1–18 neutralizing antibody protects against SARS-CoV-2 in three preclinical models. Nat Commun. 2021;12(1):6097. doi: 10.1038/s41467-021-26354-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 249.Seydoux E, Homad LJ, MacCamy AJ, Parks KR, Hurlburt NK, Jennewein MF, Akins NR, Stuart AB, Wan YH, Feng J, Whaley RE, Singh S, Boeckh M, Cohen KW, McElrath MJ, Englund JA, Chu HY, Pancera M, McGuire AT, Stamatatos L. Analysis of a SARS-CoV-2-infected individual reveals development of potent neutralizing antibodies with limited somatic mutation. Immunity. 2020;53(1):98–105 e105. doi: 10.1016/j.immuni.2020.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 250.Ju B, Zhang Q, Ge J, Wang R, Sun J, Ge X, Yu J, Shan S, Zhou B, Song S, Tang X, Yu J, Lan J, Yuan J, Wang H, Zhao J, Zhang S, Wang Y, Shi X, Liu L, Zhao J, Wang X, Zhang Z, Zhang L. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature. 2020;584(7819):115–119. doi: 10.1038/s41586-020-2380-z. [DOI] [PubMed] [Google Scholar]
- 251.Robbiani DF, Gaebler C, Muecksch F, Lorenzi JCC, Wang Z, Cho A, Agudelo M, Barnes CO, Gazumyan A, Finkin S, Hagglof T, Oliveira TY, Viant C, Hurley A, Hoffmann HH, Millard KG, Kost RG, Cipolla M, Gordon K, Bianchini F, Chen ST, Ramos V, Patel R, Dizon J, Shimeliovich I, Mendoza P, Hartweger H, Nogueira L, Pack M, Horowitz J, Schmidt F, Weisblum Y, Michailidis E, Ashbrook AW, Waltari E, Pak JE, Huey-Tubman KE, Koranda N, Hoffman PR, West AP, Jr, Rice CM, Hatziioannou T, Bjorkman PJ, Bieniasz PD, Caskey M, Nussenzweig MC. Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature. 2020;584(7821):437–442. doi: 10.1038/s41586-020-2456-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 252.Zeng X, Li L, Lin J, Li X, Liu B, Kong Y, Zeng S, Du J, Xiao H, Zhang T, Zhang S, Liu J. Isolation of a human monoclonal antibody specific for the receptor binding domain of SARS-CoV-2 using a competitive phage biopanning strategy. Antib Ther. 2020;3(2):95–100. doi: 10.1093/abt/tbaa008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 253.Kreer C, Zehner M, Weber T, Ercanoglu MS, Gieselmann L, Rohde C, Halwe S, Korenkov M, Schommers P, Vanshylla K, Di Cristanziano V, Janicki H, Brinker R, Ashurov A, Krahling V, Kupke A, Cohen-Dvashi H, Koch M, Eckert JM, Lederer S, Pfeifer N, Wolf T, Vehreschild M, Wendtner C, Diskin R, Gruell H, Becker S, Klein F. Longitudinal isolation of potent near-germline SARS-CoV-2-neutralizing antibodies from COVID-19 patients. Cell. 2020;182(4):843–854 e812. doi: 10.1016/j.cell.2020.06.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 254.Rapp M, Guo Y, Reddem ER, Yu J, Liu L, Wang P, Cerutti G, Katsamba P, Bimela JS, Bahna FA, Mannepalli SM, Zhang B, Kwong PD, Huang Y, Ho DD, Shapiro L, Sheng Z. Modular basis for potent SARS-CoV-2 neutralization by a prevalent VH1–2-derived antibody class. Cell Rep. 2021;35(1):108950. doi: 10.1016/j.celrep.2021.108950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255.Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D. Mapping neutralizing and immunodominant sites on the SARS-CoV-2 spike receptor-binding domain by structure-guided high-resolution serology. Cell. 2020;183(4):1024–1042 e1021. doi: 10.1016/j.cell.2020.09.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Tortorici MA, Beltramello M, Lempp FA, Pinto D, Dang HV, Rosen LE, McCallum M, Bowen J, Minola A, Jaconi S, Zatta F, De Marco A, Guarino B, Bianchi S, Lauron EJ, Tucker H, Zhou J, Peter A, Havenar-Daughton C, Wojcechowskyj JA, Case JB, Chen RE, Kaiser H, Montiel-Ruiz M, Meury M, Czudnochowski N, Spreafico R, Dillen J, Ng C, Sprugasci N, Culap K, Benigni F, Abdelnabi R, Foo SC, Schmid MA, Cameroni E, Riva A, Gabrieli A, Galli M, Pizzuto MS, Neyts J, Diamond MS, Virgin HW, Snell G, Corti D, Fink K, Veesler D. Ultrapotent human antibodies protect against SARS-CoV-2 challenge via multiple mechanisms. Science. 2020;370(6519):950–957. doi: 10.1126/science.abe3354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 257.Kreye J, Reincke SM, Kornau HC, Sanchez-Sendin E, Corman VM, Liu H, Yuan M, Wu NC, Zhu X, Lee CD, Trimpert J, Holtje M, Dietert K, Stoffler L, von Wardenburg N, van Hoof S, Homeyer MA, Hoffmann J, Abdelgawad A, Gruber AD, Bertzbach LD, Vladimirova D, Li LY, Barthel PC, Skriner K, Hocke AC, Hippenstiel S, Witzenrath M, Suttorp N, Kurth F, Franke C, Endres M, Schmitz D, Jeworowski LM, Richter A, Schmidt ML, Schwarz T, Muller MA, Drosten C, Wendisch D, Sander LE, Osterrieder N, Wilson IA, Pruss H. A therapeutic non-self-reactive SARS-CoV-2 antibody protects from lung pathology in a COVID-19 Hamster Model. Cell. 2020;183(4):1058–1069 e1019. doi: 10.1016/j.cell.2020.09.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 258.Rosa A, Pye VE, Graham C, Muir L, Seow J, Ng KW, Cook NJ, Rees-Spear C, Parker E, Dos Santos MS, Rosadas C, Susana A, Rhys H, Nans A, Masino L, Roustan C, Christodoulou E, Ulferts R, Wrobel AG, Short CE, Fertleman M, Sanders RW, Heaney J, Spyer M, Kjaer S, Riddell A, Malim MH, Beale R, MacRae JI, Taylor GP, Nastouli E, van Gils MJ, Rosenthal PB, Pizzato M, McClure MO, Tedder RS, Kassiotis G, McCoy LE, Doores KJ, Cherepanov P. SARS-CoV-2 can recruit a heme metabolite to evade antibody immunity. Sci Adv. 2021;7(22):eabg7607. doi: 10.1126/sciadv.abg7607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259.Guo Y, Kawaguchi A, Takeshita M, Sekiya T, Hirohama M, Yamashita A, Siomi H, Murano K. Potent mouse monoclonal antibodies that block SARS-CoV-2 infection. J Biol Chem. 2021;296:100346. doi: 10.1016/j.jbc.2021.100346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Wang Z, Lorenzi JCC, Muecksch F, Finkin S, Viant C, Gaebler C, Cipolla M, Hoffmann HH, Oliveira TY, Oren DA, Ramos V, Nogueira L, Michailidis E, Robbiani DF, Gazumyan A, Rice CM, Hatziioannou T, Bieniasz PD, Caskey M, Nussenzweig MC. Enhanced SARS-CoV-2 neutralization by dimeric IgA. Sci Transl Med. 2021 doi: 10.1126/scitranslmed.abf1555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 261.Zhou Y, Liu Z, Li S, Xu W, Zhang Q, Silva IT, Li C, Wu Y, Jiang Q, Liu Z, Wang Q, Guo Y, Wu J, Gu C, Cai X, Qu D, Mayer CT, Wang X, Jiang S, Ying T, Yuan Z, Xie Y, Wen Y, Lu L, Wang Q. Enhancement versus neutralization by SARS-CoV-2 antibodies from a convalescent donor associates with distinct epitopes on the RBD. Cell Rep. 2021;34(5):108699. doi: 10.1016/j.celrep.2021.108699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 262.Voss WN, Hou YJ, Johnson NV, Delidakis G, Kim JE, Javanmardi K, Horton AP, Bartzoka F, Paresi CJ, Tanno Y, Chou CW, Abbasi SA, Pickens W, George K, Boutz DR, Towers DM, McDaniel JR, Billick D, Goike J, Rowe L, Batra D, Pohl J, Lee J, Gangappa S, Sambhara S, Gadush M, Wang N, Person MD, Iverson BL, Gollihar JD, Dye JM, Herbert AS, Finkelstein IJ, Baric RS, McLellan JS, Georgiou G, Lavinder JJ, Ippolito GC. Prevalent, protective, and convergent IgG recognition of SARS-CoV-2 non-RBD spike epitopes. Science. 2021;372(6546):1108–1112. doi: 10.1126/science.abg5268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 263.Bell BN, Powell AE, Rodriguez C, Cochran JR, Kim PS. Neutralizing antibodies targeting the SARS-CoV-2 receptor binding domain isolated from a naive human antibody library. Protein Sci. 2021;30(4):716–727. doi: 10.1002/pro.4044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 264.Bullen G, Galson JD, Hall G, Villar P, Moreels L, Ledsgaard L, Mattiuzzo G, Bentley EM, Masters EW, Tang D, Millett S, Tongue D, Brown R, Diamantopoulos I, Parthiban K, Tebbutt C, Leah R, Chaitanya K, Ergueta-Carballo S, Pazeraitis D, Surade SB, Ashiru O, Crippa L, Cowan R, Bowler MW, Campbell JI, Lee WJ, Carr MD, Matthews D, Pfeffer P, Hufton SE, Sawmynaden K, Osbourn J, McCafferty J, Karatt-Vellatt A. Cross-reactive SARS-CoV-2 neutralizing antibodies from deep mining of early patient responses. Front Immunol. 2021;12:678570. doi: 10.3389/fimmu.2021.678570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 265.Wang N, Sun Y, Feng R, Wang Y, Guo Y, Zhang L, Deng YQ, Wang L, Cui Z, Cao L, Zhang YJ, Li W, Zhu FC, Qin CF, Wang X. Structure-based development of human antibody cocktails against SARS-CoV-2. Cell Res. 2021;31(1):101–103. doi: 10.1038/s41422-020-00446-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 266.Yao H, Sun Y, Deng YQ, Wang N, Tan Y, Zhang NN, Li XF, Kong C, Xu YP, Chen Q, Cao TS, Zhao H, Yan X, Cao L, Lv Z, Zhu D, Feng R, Wu N, Zhang W, Hu Y, Chen K, Zhang RR, Lv Q, Sun S, Zhou Y, Yan R, Yang G, Sun X, Liu C, Lu X, Cheng L, Qiu H, Huang XY, Weng T, Shi D, Jiang W, Shao J, Wang L, Zhang J, Jiang T, Lang G, Qin CF, Li L, Wang X. Rational development of a human antibody cocktail that deploys multiple functions to confer Pan-SARS-CoVs protection. Cell Res. 2021;31(1):25–36. doi: 10.1038/s41422-020-00444-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 267.Zhu L, Deng YQ, Zhang RR, Cui Z, Sun CY, Fan CF, Xing X, Huang W, Chen Q, Zhang NN, Ye Q, Cao TS, Wang N, Wang L, Cao L, Wang H, Kong D, Ma J, Luo C, Zhang Y, Nie J, Sun Y, Lv Z, Shaw N, Li Q, Li XF, Hu J, Xie L, Rao Z, Wang Y, Wang X, Qin CF. Double lock of a potent human therapeutic monoclonal antibody against SARS-CoV-2. Natl Sci Rev. 2021;8(3):nwaa297. doi: 10.1093/nsr/nwaa297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 268.Cheng MH, Porritt RA, Rivas MN, Krieger JM, Ozdemir AB, Garcia G, Jr, Arumugaswami V, Fries BC, Arditi M, Bahar I. A monoclonal antibody against staphylococcal enterotoxin B superantigen inhibits SARS-CoV-2 entry in vitro. Structure. 2021;29(9):951–962 e953. doi: 10.1016/j.str.2021.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 269.Yan R, Wang R, Ju B, Yu J, Zhang Y, Liu N, Wang J, Zhang Q, Chen P, Zhou B, Li Y, Shen Y, Zhang S, Tian L, Guo Y, Xia L, Zhong X, Cheng L, Ge X, Zhao J, Wang HW, Wang X, Zhang Z, Zhang L, Zhou Q. Structural basis for bivalent binding and inhibition of SARS-CoV-2 infection by human potent neutralizing antibodies. Cell Res. 2021;31(5):517–525. doi: 10.1038/s41422-021-00487-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 270.Mor M, Werbner M, Alter J, Safra M, Chomsky E, Lee JC, Hada-Neeman S, Polonsky K, Nowell CJ, Clark AE, Roitburd-Berman A, Ben-Shalom N, Navon M, Rafael D, Sharim H, Kiner E, Griffis ER, Gershoni JM, Kobiler O, Leibel SL, Zimhony O, Carlin AF, Yaari G, Dessau M, Gal-Tanamy M, Hagin D, Croker BA, Freund NT. Multi-clonal SARS-CoV-2 neutralization by antibodies isolated from severe COVID-19 convalescent donors. PLoS Pathog. 2021;17(2):e1009165. doi: 10.1371/journal.ppat.1009165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 271.Han X, Wang Y, Li S, Hu C, Li T, Gu C, Wang K, Shen M, Wang J, Hu J, Wu R, Mu S, Gong F, Chen Q, Gao F, Huang J, Long Y, Luo F, Song S, Long S, Hao Y, Li L, Wu Y, Xu W, Cai X, Gao Q, Zhang G, He C, Deng K, Du L, Nai Y, Wang W, Xie Y, Qu D, Huang A, Tang N, Jin A. A rapid and efficient screening system for neutralizing antibodies and its application for SARS-CoV-2. Front Immunol. 2021;12(837):653189. doi: 10.3389/fimmu.2021.653189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 272.Du S, Cao Y, Zhu Q, Yu P, Qi F, Wang G, Du X, Bao L, Deng W, Zhu H, Liu J, Nie J, Zheng Y, Liang H, Liu R, Gong S, Xu H, Yisimayi A, Lv Q, Wang B, He R, Han Y, Zhao W, Bai Y, Qu Y, Gao X, Ji C, Wang Q, Gao N, Huang W, Wang Y, Xie XS, Su XD, Xiao J, Qin C. Structurally resolved SARS-CoV-2 antibody shows high efficacy in severely infected hamsters and provides a potent cocktail pairing strategy. Cell. 2020;183(4):1013–10231013. doi: 10.1016/j.cell.2020.09.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 273.Wan J, Xing S, Ding L, Wang Y, Gu C, Wu Y, Rong B, Li C, Wang S, Chen K, He C, Zhu D, Yuan S, Qiu C, Zhao C, Nie L, Gao Z, Jiao J, Zhang X, Wang X, Ying T, Wang H, Xie Y, Lu Y, Xu J, Lan F. Human-IgG-neutralizing monoclonal antibodies block the SARS-CoV-2 infection. Cell Rep. 2020;32(3):107918. doi: 10.1016/j.celrep.2020.107918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 274.Yuan M, Liu H, Wu NC, Lee CD, Zhu X, Zhao F, Huang D, Yu W, Hua Y, Tien H, Rogers TF, Landais E, Sok D, Jardine JG, Burton DR, Wilson IA. Structural basis of a shared antibody response to SARS-CoV-2. Science. 2020;369(6507):1119–1123. doi: 10.1126/science.abd2321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 275.Song D, Wang W, Dong C, Ning Z, Liu X, Liu C, Du G, Sha C, Wang K, Lu J, Sun B, Zhao Y, Wang Q, Xu H, Li Y, Shen Z, Jiao J, Wang R, Tian J, Liu W, Wang L, Deng YQ, Dou C. Structure and function analysis of a potent human neutralizing antibody CA521(FALA) against SARS-CoV-2. Commun Biol. 2021;4(1):500. doi: 10.1038/s42003-021-02029-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 276.Scheid JF, Barnes CO, Eraslan B, Hudak A, Keeffe JR, Cosimi LA, Brown EM, Muecksch F, Weisblum Y, Zhang S, Delorey T, Woolley AE, Ghantous F, Park SM, Phillips D, Tusi B, Huey-Tubman KE, Cohen AA, Gnanapragasam PNP, Rzasa K, Hatziioanno T, Durney MA, Gu X, Tada T, Landau NR, West AP, Jr, Rozenblatt-Rosen O, Seaman MS, Baden LR, Graham DB, Deguine J, Bieniasz PD, Regev A, Hung D, Bjorkman PJ, Xavier RJ. B cell genomics behind cross-neutralization of SARS-CoV-2 variants and SARS-CoV. Cell. 2021;184(12):3205–3221 e3224. doi: 10.1016/j.cell.2021.04.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 277.Schmitz AJ, Turner JS, Liu Z, Zhou JQ, Aziati ID, Chen RE, Joshi A, Bricker TL, Darling TL, Adelsberg DC, Altomare CG, Alsoussi WB, Case JB, VanBlargan LA, Lei T, Thapa M, Amanat F, Jeevan T, Fabrizio T, O’Halloran JA, Shi P-Y, Presti RM, Webby RJ, Krammer F, Whelan SPJ, Bajic G, Diamond MS, Boon ACM, Ellebedy AH. A vaccine-induced public antibody protects against SARS-CoV-2 and emerging variants. Immunity. 2021;54(9):2159–2166.e2156. doi: 10.1016/j.immuni.2021.08.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 278.Wang L, Zhou T, Zhang Y, Yang ES, Schramm CA, Shi W, Pegu A, Oloniniyi OK, Henry AR, Darko S, Narpala SR, Hatcher C, Martinez DR, Tsybovsky Y, Phung E, Abiona OM, Antia A, Cale EM, Chang LA, Choe M, Corbett KS, Davis RL, DiPiazza AT, Gordon IJ, Hait SH, Hermanus T, Kgagudi P, Laboune F, Leung K, Liu T, Mason RD, Nazzari AF, Novik L, O'Connell S, O'Dell S, Olia AS, Schmidt SD, Stephens T, Stringham CD, Talana CA, Teng IT, Wagner DA, Widge AT, Zhang B, Roederer M, Ledgerwood JE, Ruckwardt TJ, Gaudinski MR, Moore PL, Doria-Rose NA, Baric RS, Graham BS, McDermott AB, Douek DC, Kwong PD, Mascola JR, Sullivan NJ, Misasi J. Ultrapotent antibodies against diverse and highly transmissible SARS-CoV-2 variants. Science. 2021 doi: 10.1126/science.abh1766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 279.Tan TJC, Yuan M, Kuzelka K, Padron GC, Beal JR, Chen X, Wang Y, Rivera-Cardona J, Zhu X, Stadtmueller BM, Brooke CB, Wilson IA, Wu NC. Sequence signatures of two public antibody clonotypes that bind SARS-CoV-2 receptor binding domain. Nat Commun. 2021;12(1):3815. doi: 10.1038/s41467-021-24123-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 280.Banach BB, Cerutti G, Fahad AS, Shen CH, Oliveira De Souza M, Katsamba PS, Tsybovsky Y, Wang P, Nair MS, Huang Y, Francino-Urdaniz IM, Steiner PJ, Gutierrez-Gonzalez M, Liu L, Lopez Acevedo SN, Nazzari AF, Wolfe JR, Luo Y, Olia AS, Teng IT, Yu J, Zhou T, Reddem ER, Bimela J, Pan X, Madan B, Laflin AD, Nimrania R, Yuen KY, Whitehead TA, Ho DD, Kwong PD, Shapiro L, DeKosky BJ. Paired heavy- and light-chain signatures contribute to potent SARS-CoV-2 neutralization in public antibody responses. Cell Rep. 2021;37(1):109771. doi: 10.1016/j.celrep.2021.109771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 281.Li D, Edwards RJ, Manne K, Martinez DR, Schafer A, Alam SM, Wiehe K, Lu X, Parks R, Sutherland LL, Oguin TH, 3rd, McDanal C, Perez LG, Mansouri K, Gobeil SMC, Janowska K, Stalls V, Kopp M, Cai F, Lee E, Foulger A, Hernandez GE, Sanzone A, Tilahun K, Jiang C, Tse LV, Bock KW, Minai M, Nagata BM, Cronin K, Gee-Lai V, Deyton M, Barr M, Von Holle T, Macintyre AN, Stover E, Feldman J, Hauser BM, Caradonna TM, Scobey TD, Rountree W, Wang Y, Moody MA, Cain DW, DeMarco CT, Denny TN, Woods CW, Petzold EW, Schmidt AG, Teng IT, Zhou T, Kwong PD, Mascola JR, Graham BS, Moore IN, Seder R, Andersen H, Lewis MG, Montefiori DC, Sempowski GD, Baric RS, Acharya P, Haynes BF, Saunders KO. In vitro and in vivo functions of SARS-CoV-2 infection-enhancing and neutralizing antibodies. Cell. 2021;184(16):4203–4219 e4232. doi: 10.1016/j.cell.2021.06.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 282.Clark SA, Clark LE, Pan J, Coscia A, McKay LGA, Shankar S, Johnson RI, Brusic V, Choudhary MC, Regan J, Li JZ, Griffiths A, Abraham J. SARS-CoV-2 evolution in an immunocompromised host reveals shared neutralization escape mechanisms. Cell. 2021;184(10):2605–2617 e2618. doi: 10.1016/j.cell.2021.03.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 283.Zhou X, Ma F, Xie J, Yuan M, Li Y, Shaabani N, Zhao F, Huang D, Wu NC, Lee CD, Liu H, Li J, Chen Z, Hong Y, Liu WH, Xiao N, Burton DR, Tu H, Li H, Chen X, Teijaro JR, Wilson IA, Xiao C, Huang Z. Diverse immunoglobulin gene usage and convergent epitope targeting in neutralizing antibody responses to SARS-CoV-2. Cell Rep. 2021;35(6):109109. doi: 10.1016/j.celrep.2021.109109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 284.Pinto D, Sauer MM, Czudnochowski N, Low JS, Tortorici MA, Housley MP, Noack J, Walls AC, Bowen JE, Guarino B, Rosen LE, di Iulio J, Jerak J, Kaiser H, Islam S, Jaconi S, Sprugasci N, Culap K, Abdelnabi R, Foo C, Coelmont L, Bartha I, Bianchi S, Silacci-Fregni C, Bassi J, Marzi R, Vetti E, Cassotta A, Ceschi A, Ferrari P, Cippa PE, Giannini O, Ceruti S, Garzoni C, Riva A, Benigni F, Cameroni E, Piccoli L, Pizzuto MS, Smithey M, Hong D, Telenti A, Lempp FA, Neyts J, Havenar-Daughton C, Lanzavecchia A, Sallusto F, Snell G, Virgin HW, Beltramello M, Corti D, Veesler D. Broad betacoronavirus neutralization by a stem helix-specific human antibody. Science. 2021;373(6559):1109–1116. doi: 10.1126/science.abj3321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 285.Zhang Q, Ju B, Ge J, Chan JF, Cheng L, Wang R, Huang W, Fang M, Chen P, Zhou B, Song S, Shan S, Yan B, Zhang S, Ge X, Yu J, Zhao J, Wang H, Liu L, Lv Q, Fu L, Shi X, Yuen KY, Liu L, Wang Y, Chen Z, Zhang L, Wang X, Zhang Z. Potent and protective IGHV3-53/3-66 public antibodies and their shared escape mutant on the spike of SARS-CoV-2. Nat Commun. 2021;12(1):4210. doi: 10.1038/s41467-021-24514-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 286.Asarnow D, Wang B, Lee WH, Hu Y, Huang CW, Faust B, Ng PML, Ngoh EZX, Bohn M, Bulkley D, Pizzorno A, Ary B, Tan HC, Lee CY, Minhat RA, Terrier O, Soh MK, Teo FJ, Yeap YYC, Seah SGK, Chan CEZ, Connelly E, Young NJ, Maurer-Stroh S, Renia L, Hanson BJ, Rosa-Calatrava M, Manglik A, Cheng Y, Craik CS, Wang CI. Structural insight into SARS-CoV-2 neutralizing antibodies and modulation of syncytia. Cell. 2021;184(12):3192–3204 e3116. doi: 10.1016/j.cell.2021.04.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 287.Noy-Porat T, Mechaly A, Levy Y, Makdasi E, Alcalay R, Gur D, Aftalion M, Falach R, Leviatan Ben-Arye S, Lazar S, Zauberman A, Epstein E, Chitlaru T, Weiss S, Achdout H, Edgeworth JD, Kikkeri R, Yu H, Chen X, Yitzhaki S, Shapira SC, Padler-Karavani V, Mazor O, Rosenfeld R. Therapeutic antibodies, targeting the SARS-CoV-2 spike N-terminal domain, protect lethally infected K18-hACE2 mice. Science. 2021;24(5):102479. doi: 10.1016/j.isci.2021.102479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 288.Cao Y, Yisimayi A, Bai Y, Huang W, Li X, Zhang Z, Yuan T, An R, Wang J, Xiao T, Du S, Ma W, Song L, Li Y, Li X, Song W, Wu J, Liu S, Li X, Zhang Y, Su B, Guo X, Wei Y, Gao C, Zhang N, Zhang Y, Dou Y, Xu X, Shi R, Lu B, Jin R, Ma Y, Qin C, Wang Y, Feng Y, Xiao J, Xie XS. Humoral immune response to circulating SARS-CoV-2 variants elicited by inactivated and RBD-subunit vaccines. Cell Res. 2021;31(7):732–741. doi: 10.1038/s41422-021-00514-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 289.Pan Y, Du J, Liu J, Wu H, Gui F, Zhang N, Deng X, Song G, Li Y, Lu J, Wu X, Zhan S, Jing Z, Wang J, Yang Y, Liu J, Chen Y, Chen Q, Zhang H, Hu H, Duan K, Wang M, Wang Q, Yang X. Screening of potent neutralizing antibodies against SARS-CoV-2 using convalescent patients-derived phage-display libraries. Cell Discov. 2021;7(1):57. doi: 10.1038/s41421-021-00295-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 290.VanBlargan LA, Adams LJ, Liu Z, Chen RE, Gilchuk P, Raju S, Smith BK, Zhao H, Case JB, Winkler ES, Whitener BM, Droit L, Aziati ID, Bricker TL, Joshi A, Shi PY, Creanga A, Pegu A, Handley SA, Wang D, Boon ACM, Crowe JE, Jr, Whelan SPJ, Fremont DH, Diamond MS. A potently neutralizing SARS-CoV-2 antibody inhibits variants of concern by utilizing unique binding residues in a highly conserved epitope. Immunity. 2021;54(10):2399–2416 e2396. doi: 10.1016/j.immuni.2021.08.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 291.Kramer KJ, Johnson NV, Shiakolas AR, Suryadevara N, Periasamy S, Raju N, Williams JK, Wrapp D, Zost SJ, Walker LM, Wall SC, Holt CM, Hsieh CL, Sutton RE, Paulo A, Nargi RS, Davidson E, Doranz BJ, Crowe JE, Jr, Bukreyev A, Carnahan RH, McLellan JS, Georgiev IS. Potent neutralization of SARS-CoV-2 variants of concern by an antibody with an uncommon genetic signature and structural mode of spike recognition. Cell Rep. 2021;37(1):109784. doi: 10.1016/j.celrep.2021.109784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 292.Jia L, Liu YP, Tian LF, Xiong C, Xu X, Qu H, Xiong W, Zhou D, Wang F, Liu Z, Yan XX, Xu W, Tang L. Potent neutralizing RBD-specific antibody cocktail against SARS-CoV-2 and its mutant. MedComm (2020). 2021 doi: 10.1002/mco2.79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 293.Tong P, Gautam A, Windsor IW, Travers M, Chen Y, Garcia N, Whiteman NB, McKay LGA, Storm N, Malsick LE, Honko AN, Lelis FJN, Habibi S, Jenni S, Cai Y, Rennick LJ, Duprex WP, McCarthy KR, Lavine CL, Zuo T, Lin J, Zuiani A, Feldman J, MacDonald EA, Hauser BM, Griffths A, Seaman MS, Schmidt AG, Chen B, Neuberg D, Bajic G, Harrison SC, Wesemann DR. Memory B cell repertoire for recognition of evolving SARS-CoV-2 spike. Cell. 2021;184(19):4969–4980 e4915. doi: 10.1016/j.cell.2021.07.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 294.Peter AS, Roth E, Schulz SR, Fraedrich K, Steinmetz T, Damm D, Hauke M, Richel E, Mueller-Schmucker S, Habenicht K, Eberlein V, Issmail L, Uhlig N, Dolles S, Gruner E, Peterhoff D, Ciesek S, Hoffmann M, Pohlmann S, McKay PF, Shattock RJ, Wolfel R, Socher E, Wagner R, Eichler J, Sticht H, Schuh W, Neipel F, Ensser A, Mielenz D, Tenbusch M, Winkler TH, Grunwald T, Uberla K, Jack HM. A pair of noncompeting neutralizing human monoclonal antibodies protecting from disease in a SARS-CoV-2 infection model. Eur J Immunol. 2021 doi: 10.1002/eji.202149374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 295.Wang Z, Muecksch F, Schaefer-Babajew D, Finkin S, Viant C, Gaebler C, Hoffmann HH, Barnes CO, Cipolla M, Ramos V, Oliveira TY, Cho A, Schmidt F, Da Silva J, Bednarski E, Aguado L, Yee J, Daga M, Turroja M, Millard KG, Jankovic M, Gazumyan A, Zhao Z, Rice CM, Bieniasz PD, Caskey M, Hatziioannou T, Nussenzweig MC. Naturally enhanced neutralizing breadth against SARS-CoV-2 one year after infection. Nature. 2021;595(7867):426–431. doi: 10.1038/s41586-021-03696-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 296.Onodera T, Kita S, Adachi Y, Moriyama S, Sato A, Nomura T, Sakakibara S, Inoue T, Tadokoro T, Anraku Y, Yumoto K, Tian C, Fukuhara H, Sasaki M, Orba Y, Shiwa N, Iwata N, Nagata N, Suzuki T, Sasaki J, Sekizuka T, Tonouchi K, Sun L, Fukushi S, Satofuka H, Kazuki Y, Oshimura M, Kurosaki T, Kuroda M, Matsuura Y, Suzuki T, Sawa H, Hashiguchi T, Maenaka K, Takahashi Y. A SARS-CoV-2 antibody broadly neutralizes SARS-related coronaviruses and variants by coordinated recognition of a virus-vulnerable site. Immunity. 2021;54(10):2385–2398 e2310. doi: 10.1016/j.immuni.2021.08.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 297.Nie J, Xie J, Liu S, Wu J, Liu C, Li J, Liu Y, Wang M, Zhao H, Zhang Y, Yao J, Chen L, Shen Y, Yang Y, Wang HW, Wang Y, Huang W. Three epitope-distinct human antibodies from RenMab mice neutralize SARS-CoV-2 and cooperatively minimize the escape of mutants. Cell Discov. 2021;7(1):53. doi: 10.1038/s41421-021-00292-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 298.Song G, He W-T, Callaghan S, Anzanello F, Huang D, Ricketts J, Torres JL, Beutler N, Peng L, Vargas S, Cassell J, Parren M, Yang L, Ignacio C, Smith DM, Voss JE, Nemazee D, Ward AB, Rogers T, Burton DR, Andrabi R. Cross-reactive serum and memory B-cell responses to spike protein in SARS-CoV-2 and endemic coronavirus infection. Nat Commun. 2021;12(1):2938. doi: 10.1038/s41467-021-23074-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 299.Rujas E, Kucharska I, Tan YZ, Benlekbir S, Cui H, Zhao T, Wasney GA, Budylowski P, Guvenc F, Newton JC, Sicard T, Semesi A, Muthuraman K, Nouanesengsy A, Aschner CB, Prieto K, Bueler SA, Youssef S, Liao-Chan S, Glanville J, Christie-Holmes N, Mubareka S, Gray-Owen SD, Rubinstein JL, Treanor B, Julien JP. Multivalency transforms SARS-CoV-2 antibodies into ultrapotent neutralizers. Nat Commun. 2021;12(1):3661. doi: 10.1038/s41467-021-23825-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 300.Fenwick C, Turelli P, Perez L, Pellaton C, Esteves-Leuenberger L, Farina A, Campos J, Lana E, Fiscalini F, Raclot C, Pojer F, Lau K, Demurtas D, Descatoire M, Joo VS, Foglierini M, Noto A, Abdelnabi R, Foo CS, Vangeel L, Neyts J, Du W, Bosch BJ, Veldman G, Leyssen P, Thiel V, LeGrand R, Levy Y, Trono D, Pantaleo G. A highly potent antibody effective against SARS-CoV-2 variants of concern. Cell Rep. 2021;37(2):109814. doi: 10.1016/j.celrep.2021.109814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 301.Wheatley AK, Pymm P, Esterbauer R, Dietrich MH, Lee WS, Drew D, Kelly HG, Chan L-J, Mordant FL, Black KA, Adair A, Tan H-X, Juno JA, Wragg KM, Amarasena T, Lopez E, Selva KJ, Haycroft ER, Cooney JP, Venugopal H, Tan LL, Neill MTO, Allison CC, Cromer D, Davenport MP, Bowen RA, Chung AW, Pellegrini M, Liddament MT, Glukhova A, Subbarao K, Kent SJ, Tham W-H. Landscape of human antibody recognition of the SARS-CoV-2 receptor binding domain. Cell Rep. 2021;37(2):109822. doi: 10.1016/j.celrep.2021.109822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 302.Cho A, Muecksch F, Schaefer-Babajew D, Wang Z, Finkin S, Gaebler C, Ramos V, Cipolla M, Mendoza P, Agudelo M, Bednarski E, DaSilva J, Shimeliovich I, Dizon J, Daga M, Millard KG, Turroja M, Schmidt F, Zhang F, Tanfous TB, Jankovic M, Oliveria TY, Gazumyan A, Caskey M, Bieniasz PD, Hatziioannou T, Nussenzweig MC. Anti-SARS-CoV-2 receptor-binding domain antibody evolution after mRNA vaccination. Nature. 2021 doi: 10.1038/s41586-021-04060-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 303.Zost SJ, Gilchuk P, Chen RE, Case JB, Reidy JX, Trivette A, Nargi RS, Sutton RE, Suryadevara N, Chen EC, Binshtein E, Shrihari S, Ostrowski M, Chu HY, Didier JE, MacRenaris KW, Jones T, Day S, Myers L, Eun-Hyung Lee F, Nguyen DC, Sanz I, Martinez DR, Rothlauf PW, Bloyet LM, Whelan SPJ, Baric RS, Thackray LB, Diamond MS, Carnahan RH, Crowe JE., Jr Rapid isolation and profiling of a diverse panel of human monoclonal antibodies targeting the SARS-CoV-2 spike protein. Nat Med. 2020;26(9):1422–1427. doi: 10.1038/s41591-020-0998-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All the data and materials supporting the conclusions were included in the main paper.