Abstract
An unusual, highly diverged derivative of the Sabin type 2 oral poliovaccine (OPV) strain was recovered from environmental samples during routine screening for wild polioviruses. Virus was cultivated in L20B cells and then passaged on BGM cells at 40°C (RCT [reproductive capacity at supraoptimal temperature]-positive marker) to select against most OPV strains. All but 1 of 25 RCT-positive OPV-derived environmental isolates were antigenically and genetically (>99.5% VP1 sequence match) similar to the respective Sabin strains. However, isolate PV2/4568-1/ISR98 (referred to below as 4568-1) escaped neutralization with Sabin 2-specific monoclonal antibodies and cross-adsorbed sera, and had multiple nucleotide substitutions (220 of 2,646; 8.3%) in the P1 capsid region. Fourteen of the 44 associated amino acid substitutions in the capsid mapped to neutralizing antigenic sites. Neutralizing titers in the sera of 50 Israeli children 15 years old were significantly lower to 4568-1 (geometric mean titer [GMT], 47) than to Sabin 2 (GMT, 162) or to the prototype wild strain, PV2/MEF-1/EGY42 (GMT, 108). Two key attenuating sites had also reverted in 4568-1 (A481 to G in the 5′ untranslated region and the VP1 amino acid I143 to T), and the isolate was highly neurovirulent for transgenic mice expressing the poliovirus receptor (PVR-Tg21 mice). The extensive genetic divergence of 4568-1 from the parental Sabin 2 strain suggested that the virus had replicated in one or more people for ∼6 years. The presence in the environment of a highly evolved, neurovirulent OPV-derived poliovirus in the absence of polio cases has important implications for strategies for the cessation of immunization with OPV following global polio eradication.
Rapid evolution is characteristic of both wild and vaccine-derived polioviruses (4, 5, 11, 12, 15; A. Heim, A. Bellmunt, G. May, P. Pring-Akerblom, and W. Verhagen, Abstr. Eur. Soc. Clin. Virol. Prog. Clin. Virol. IV, abstr. 350, p. 72, 1998). For wild polioviruses, nucleotide substitutions accumulate at a rate of approximately 1% per year and consist primarily of changes at synonymous codon positions (10; L. De, J. Jorba, J. Boshell, R. Salas, and O. Kew, Abstr. 17th Annu. Meet. Am. Soc. Virol., abstr. W36-3, p. 123, 1998), i.e., do not result in amino acid changes at those loci. In contrast, the mutations initially appearing and fixed into the genomes of the Sabin vaccine strains upon administration of oral poliovaccine (OPV) are frequently associated with reversion of the attenuated phenotype and alteration of the neutralizing antigenic (NAg) sites of the OPV strains (1, 12). Reversion of the OPV strains to increased neurovirulence is one key factor for the occurrence of cases of vaccine-associated paralytic poliomyelitis (VAPP), which occur at a rate of ∼1 per 500,000 first doses of OPV in immunocompetent individuals (23) and at a ∼1,000-fold-higher rate for immunodeficient patients (24). Poliovirus replication is restricted to about 2 months in immunocompetent persons (1, 2) but may be prolonged up to 10 years in patients with deficiencies in antibody production (11). Because poliovirus genomes evolve rapidly, the duration of replication of an OPV-derived virus may be estimated by the extent of its nucleotide sequence divergence from its respective prototype OPV strain (11; De et al., Abstr. 17th Annu. Meet. Am. Soc. Virol.).
Routine environmental surveillance of wastewater for polioviruses was initiated in Israel and the Palestinian Authority following the last poliomyelitis outbreak in 1988 (14, 22). This approach has proven to be a powerful method to detect wild poliovirus circulation in communities where no poliomyelitis cases have been reported (14). Concentrated environmental samples are cultured using a double-selective cultivation technique (13) that selects against the growth of most OPV-derived strains by use of a supraoptimal temperature of incubation (40°C). Polioviruses that grow at 40°C are said to be positive for the RCT (reproductive capacity at supraoptimal temperature) marker. A small number (3 to 25) of RCT-positive isolates are recovered each year from the environment. Most of these isolates have been otherwise typical OPV-derived viruses that have lost their temperature-sensitive phenotypes; some have been imported wild polioviruses (14; L. M. Shulman, Y. Manor, R. Handsher, A. Vonsover, O. M. Kew, E. Mendelson, et al., Abstr. Xth Int. Cong. Virol., abstr. W54-6, p. 79, 1996); and one, PV2/4568-1/ISR98, isolated in 1998, proved to be a highly divergent, neurovirulent derivative of the Sabin type 2 OPV strain. In this report, we describe the molecular, antigenic, and neurovirulence properties of this unusual isolate. We estimate from the high degree of nucleotide divergence of 4568-1 from Sabin 2 that the initiating OPV dose was given approximately 6 years before recovery of the virus from the environment. The observations cannot distinguish between chronic infection of a single individual and person-to-person transmission of an OPV-derived poliovirus. In either event, the detection in the environment of a highly evolved derivative of Sabin 2 has important implications for development of the optimal strategy for cessation of immunization with OPV following the eradication of all wild poliovirus transmission (4).
MATERIALS AND METHODS
Reference poliovirus.
Nonattenuated Salk inactivated poliovaccine (IPV) reference strains of type 1 (PV1/Mahoney/USA42), type 2 (PV2/MEF-1/EGY42), and type 3 (PV3/Saukett/USA52) and attenuated Sabin OPV type 1 (LSc 2ab), type 2 (P712 ch 2ab), and type 3 (Leon 12 a1b) strains were obtained from Radu Crainic, Institut Pasteur, Paris, France.
Isolation of polioviruses from sewage samples.
Sampling and extraction of sewage samples were performed as described in detail previously (14), except that processed sewage extracts (15 to 24 ml) were first applied to monolayers of L20B cells (mouse L cells expressing the human poliovirus receptor [PVR]) (16, 20), kindly provided by David Wood, National Institute for Biological Standards and Control, London, England. Individual isolates have been named according to the following convention: PV (poliovirus) followed by a number denoting the type/the isolate number/the 3-letter country code followed by two digits indicating the year of isolation (e.g., PV2/4568-1/ISR98). In this report, the names have been shortened to the isolate number (e.g., PV2/4568-1/ISR98 is referred to as 4568-1).
Virus strain characterization.
Virus typing and intratypic differentiation (ITD) were carried out by standard methods (5, 13, 14). Further strain characterization was done by dot blot hybridization using RNA probes (6). The plasmids, containing the variable VP1 poliovirus strain-specific inserts or an enteroviral group probe (EV/5′UTR) were kindly provided by Lina De, Centers for Disease Control and Prevention, Atlanta, Ga. Preparation of digoxigenin (DIG)-labeled RNA transcripts from the linearized plasmid was performed according to the manufacturer's instructions (Boehringer GmbH, Mannheim, Germany). Supernatants (200 μl) from isolates grown to full cytopathic effect in tube cultures of HEp2 or L20B cells were spotted onto Hybond N+ nylon filters (Amersham Pharmacia Biotech, Little Chalfont, England) in quadruplicate, one spot for each Sabin-specific probe and one for the enteroviral-group-specific probe. Hybridization overnight at 65°C and detection by chemiluminescence were performed as described previously (6) except that Lumi Phos 530 was replaced by disodium 3-(4-methoxyspiro{1,2-dioxetane-3,2′-(5′-chloro)tricyclo[3.3.1.13,7]decan}4- yl)phenyl phosphate (CSPD; Boehringer).
Sequence analysis.
Sequence analysis of the capsid coding region (pP1, a 1,100-nucleotide [nt] reverse transcription-PCR [RT-PCR]-amplified segment which includes the entire VP1 gene and the VP1/2A junction) and 385 nt of the 5′ untranslated region (5′-UTR) was carried out as previously described (11, 22, 28). The RT-PCR product was purified directly from the RT-PCR mixture and sequenced using the ABI PRISM Dye Deoxy Terminator Cycle Sequencing Kit (Applied Biosystems, Foster City, Calif.). Reaction mixtures were analyzed on Applied Biosystems model 373 DNA Automatic Sequencing Systems. The University of Wisconsin Genetics Computer Group (GCG) gene analysis programs (version 9, 1994) were used for comparing nucleotide and amino acid sequences from isolates and the GenBank/EMBL database. Both strands of the entire VP1 gene and the 5′-UTR of the first five isolates were sequenced. All subsequent isolates were screened by partial sequencing (250 to 800 nt) starting from the 3′ end of VP1. The entire P1 region, coding for all capsid proteins, was determined for isolate 4568-1 using two additional primers, S0001 (sense polarity, matching genome positions 531 to 551, 5′-GCG GAA CCG ACT ACT TTG GGT-3′) and S0002 (sense polarity, matching genome positions 1249 to 1266, 5′-CAC TAT CTT GGG AGG GCT-3′). The deduced amino acid sequences of the capsid proteins from 4568-1 and sequences from other isolates were determined using either GCG programs or the DNA Strider program (Service de Biochemie et de Genetique, Institut de Recherche Fondamental, CEA, Saclay, France). Phylogenetic trees were constructed using the maximum-likelihood analysis programs PHYLIP (version 3.57c; University of Washington, Seattle [http://www.evolution.genetics.washington.edu/phylip.html]) and PUZZLE (version 4.0; Zoologisches Institut, Universität München, Munich, Germany, 1997 [http://www.zi.biologie.uni-muenchen.de/∼strimmer/puzzle.html]) and were visualized using the TREEVIEW program (Division of Environmental and Evolutionary Biology, Institute of Biomedical and Life Sciences, University of Glasgow, Glasgow, Scotland, 1998 [http://taxonomy.zoology.gla.ac.uk/rod/rod.html]).
Neurovirulence testing in transgenic mice.
The neurovirulence test was carried out as previously described (7) on PVR-Tg21 mice (kindly provided by A. Nomoto and T. Nomura). Ten mice (5 males and 5 females) were inoculated for each virus tested, except for 4568-1, for which 20 mice were used (10 females and 10 males). Each mouse received intraperitoneal injections of 0.33 × 108 to 1 × 108 PFU of virus suspended in 0.5 ml of Dulbecco modified Eagle's medium containing 0.1% fetal calf serum. The viral suspensions used for inoculation were back-titrated to confirm the quantity of virus injected per mouse. Mice were monitored daily for as long as 14 days after inoculation, and clinical symptoms (paresis, paralysis, or death) were recorded for each mouse. The mean healthy time (MHT) was determined for each virus by calculating the mean number of days before the appearance of any clinical symptom for individual mice inoculated with the corresponding virus.
Nucleotide sequence accession numbers.
The sequences of the RCT-positive isolates described in this article have been deposited in the EMBL/GenBank data library and have been assigned accession no. AJ237871 to AJ237885. Additional sequences referred to have the following accession numbers: PV2/Sabin 2, X00595; PV3/Sabin 3, X00925; and AF249260 to AF249265 and AF249260 to AF249265 for the type 2 clinical isolates as described in the legend to Fig. 2.
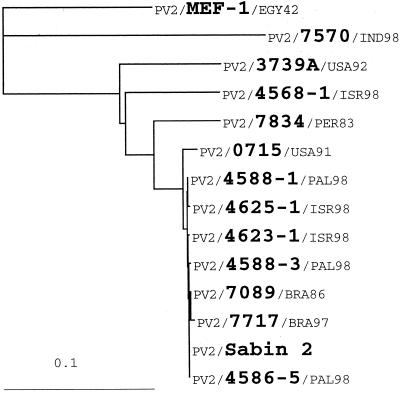
FIG. 2.
Phylogenetic tree of type 2 RCT-positive sewage isolates. VP1 sequences of 4568-1 were compared with those of Sabin 2, five other Sabin 2-derived RCT-positive sewage isolates from Israel (including the type 2 antigenic variant VP2/4625-1/ISR98), two isolates from immunodeficient VAPP patients in the United States (PV2/0715/USA91 and PV2/3739A/USA92), a divergent type 2 isolate from a patient in Peru (PV2/7834/PER83), two recent isolates from children with acute flaccid paralysis in Brazil (PV2/7089/BRA96 and PV2/7717/BRA97), the reference wild type 2 IPV strain, PV2/MEF-1/EGY42, and a wild type 2 isolate from a 1998 polio case in India (PV2/7570/IND98).
RESULTS
Properties of the poliovirus isolates from sewage.
Twenty-five RCT-positive sewage isolates were obtained in 1998, of which 20 were type 2 and 5 were type 3 (Table 1). Three of the isolates were found to be antigenically distinct from the parental vaccine strains: 4568-1 (type 2), which was not neutralized by Sabin 2-specific monoclonal antibodies and cross-adsorbed sera; 4625-1 (type 2), which was neutralized only by a Sabin 2-specific monoclonal antibody; and 4745-3 (type 3), which was neutralized only by Sabin 3-specific cross-adsorbed sera. Twenty of the 25 sewage isolates were partially sequenced, and 19 of these (including 4625-1 and 4745-3) were found to be very closely related (>99.5% nucleotide sequence similarity in the VP1 region) to their respective OPV strains (Table 1). One of the type 2 antigenic variants, 4568-1, differed from Sabin 2 at 8.6% (78 of 903) of VP1 nucleotides (Table 1) and did not form stable hybrids with the Sabin 2-specific RNA probe (Fig. 1).
TABLE 1.
Description of all RCT-positive isolates obtained from sewage in 1998
| Isolate | Date (mo yr) | Sampling site | Virus type | Neutralization by:
|
Nucleotide homology (%)c | Region(s) sequenced | No. of nucleotides analyzed | |
|---|---|---|---|---|---|---|---|---|
| Monoclonal antibodya | Polyclonal antiserab | |||||||
| PV3/4521-2/PAL98 | Mar. 98 | Schem | 3 | Yes | Yes | >99.5 | VP1/2A, partial VP1 | 900 |
| PV2/4543-1/PAL98 | Mar. 98 | Gaza (Beit-Lahiya) | 2 | Yes | ND | ND | ND | ND |
| PV3/4553-11/PAL98 | Apr. 98 | Qualquilya | 3 | Yes | ND | ND | ND | ND |
| PV2/4553-26/PAL98 | Apr. 98 | Qualquilya | 2 | Yes | Yes | ND | ND | ND |
| PV2/4553-27/PAL98 | Apr. 98 | Qualquilya | 2 | Yes | Yes | ND | ND | ND |
| PV2/4553-29/PAL98 | Apr. 98 | Qualquilya | 2 | Yes | Yes | ND | ND | ND |
| PV2/4568-1/ISR98 | May 98 | Shaf Dan | 2 | No | No | 90.6 | 5′-UTR, P1 | 3,301 |
| PV2/4586-5/PAL98 | May 98 | Gaza (Sheik-Raduan) | 2 | Yes | Yes | >99.5 | 5′-UTR, VP1 | 1,301 |
| PV2/4588-1/PAL98 | May 98 | Gaza (Raffah) | 2 | Yes | Yes | >99.5 | 5′-UTR, VP1 | 1,301 |
| PV2/4588-3/PAL98 | May 98 | Gaza (Raffah) | 2 | Yes | Yes | >99.5 | 5′-UTR, VP1 | 1,301 |
| PV2/4623-1/ISR98 | Jun. 98 | Ofakim | 2 | Yes | Yes | >99.5 | 5′-UTR, VP1 | 1,301 |
| PV2/4625-1/ISR98 | Jun. 98 | Tiberia Elite | 2 | No | Yes | >99.5 | 5′-UTR, VP1 | 1,301 |
| PV2/4726-1/PAL98 | Oct. 98 | Jenin | 2 | ND | Yes | >99.5 | VP1/2A, VP1 | 1,057 |
| PV3/4745-1/PAL98 | Nov. 98 | Gaza (Sheik-Raduan) | 3 | ND | Yes | >99.5 | VP1/2A | 253 |
| PV2/4745-2/PAL98 | Nov. 98 | Gaza (Sheik-Raduan) | 2 | ND | Yes | >99.5 | VP1/2A, partial VP1 | 696 |
| PV3/4745-3/PAL98 | Nov. 98 | Gaza (Sheik-Raduan) | 3 | Yes | No | >99.5 | VP1/2A, VP1 | 1,050 |
| PV2/4746-1/PAL98 | Nov. 98 | Gaza (Beit-Lahiya) | 2 | ND | Yes | >99.5 | VP1/2A, partial VP1 | 696 |
| PV2/4746-2/PAL98 | Nov. 98 | Gaza (Beit-Lahiya) | 2 | ND | Yes | >99.5 | VP1/2A, partial VP1 | 696 |
| PV3/4747-1/PAL98 | Nov. 98 | Gaza (Raffah) | 3 | Yes | Yes | >99.5 | VP1/2A | 253 |
| PV2/4761-2/ISR98 | Nov. 98 | Quiryat-Gat | 2 | Yes | Yes | >99.5 | VP1/2A, partial VP1 | 696 |
| PV2/4772-1/PAL98 | Nov. 98 | Gaza (Beit-Lahiya) | 2 | Yes | Yes | >99.5 | VP1/2A, partial VP1 | 784 |
| PV2/4774-1/PAL98 | Nov. 98 | Jenin (N.) | 2 | Yes | Yes | >99.5 | VP1/2A, partial VP1 | 756 |
| PV2/4808-1/PAL98 | Dec. 98 | Gaza | 2 | Weak | ND | >99.5 | VP1/2A, partial VP1 | 837 |
| PV2/4809-1/PAL98 | Dec. 98 | Gaza | 2 | Yes | ND | >99.5 | VP1/2A, partial VP1 | 837 |
| PV2/4809-3/PAL98 | Dec. 98 | Gaza | 2 | Yes | ND | >99.5 | VP1/2A, partial VP1 | 836 |
Respective monoclonal anti-Sabin antibody. ND, test not done.
Respective cross-adsorbed, polyclonal anti-Sabin antisera.
Homology to the corresponding region of the respective Sabin strain.
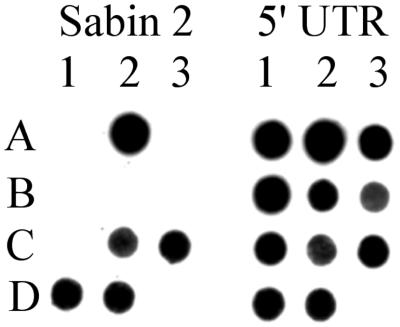
FIG. 1.
Intratypic differentiation of type 2, RCT-positive poliovirus isolates by blot hybridization. Viral RNA was bound to nylon filters and hybridized with a DIG-labeled Sabin 2-specific VP1 probe (left blot) or a panenteroviral 5′-UTR probe (right blot). Hybrids were detected by chemiluminescence (6). Controls: A1, Sabin 1; A2, Sabin 2; A3, Sabin 3; B1, Mahoney; B2, PV2/MEF-1/EGY42; B3, PV3/Saukett/USA52. RCT-positive isolates: C1, PV2/4568-1/ISR98; C2, PV2/4586-5/PAL98; C3, PV2/4588-3/PAL98; D1, PV2/4623-1/ISR98; D2, PV2/4625-1/ISR98.
Relationships of sewage isolates to other type 2 polioviruses.
To assess the genetic relationships of 4568-1 to other type 2 polioviruses, VP1 sequences of 4568-1 were compared with those of the Sabin 2 vaccine strain; the wild type 2 reference strain, MEF-1; a wild type 2 isolate (7570) from a 1998 polio case in India (the only country in which type 2 wild poliovirus was known to be endemic in 1998 [3]); and 10 Sabin 2-derived isolates. The other Sabin 2-derived isolates included five additional 1998 type 2 sewage isolates from Israel (4586-5, 4588-1, 4588-3, 4623-1, and the antigenic variant 4625-1), two isolates (3739A and 0715) from immunodeficient VAPP patients from the United States, a divergent poliovirus isolate (7834) from a polio patient in Peru, and two recent isolates (7089 and 7717) from children with acute flaccid paralysis in Brazil. The sequence relationships, summarized in a tree (Fig. 2) constructed using the DNA maximum-likelihood program of PHYLIP and rooted to the sequence of MEF-1, confirmed that 4568-1 was more closely related to the Sabin 2 group than to the wild viruses. Very similar relationships were obtained by quartet puzzling using the PUZZLE program (data not shown). Isolate 4568-1 was more divergent in VP1 sequences from Sabin 2 (8.6%) than were 7834 (6.0%) and 0715 (1.4%), but was less divergent than 3739A (10.2%). Isolate 4568-1 was unrelated to MEF-1 and 7570 (19.8 and 22.4% VP1 nucleotide differences, respectively) (Fig. 1).
An additional measure of genetic relatedness is the proportion of substitutions that are transversions (substitutions between a purine and a pyrimidine). The proportion of transversional differences between closely related polioviruses is ∼5 to 15% but rises to >30% with increased genetic distance (10). The proportion of transversional differences between 4568-1 and Sabin 2 (15.5%) was significantly lower than the proportion of transversional differences between 4568-1 and MEF-1 (27.5%) or 7570 (45.9%).
Sequence properties of isolate 4568-1.
Analysis of the VP1 nucleotide sequence of isolate 4568-1 was extended to include the complete P1 region of the genome (2,646 nt) encoding the four capsid proteins (VP1 to VP4). The P1 capsid region differed from that of Sabin 2 by 8.3% (220 of 2,646) of nucleotides. Nucleotide substitutions were widely distributed throughout the P1 region. Most (166 of 220; 75.5%) substitutions occurred in the third codon position. In 196 codons with single nucleotide substitutions, 99.5% (155 of 156) of the third-position nucleotide changes generated synonymous codons (i.e., did not encode an amino acid substitution). Of the 44 deduced amino acid substitutions into the capsid protein, 14 clustered in virion surface residues forming the NAg sites (9, 17, 18) (Fig. 3). Six additional amino acid substitutions occurred in the variable interval (residues 1 to 32) (26) near the amino terminus of VP1. Twelve codons had double nucleotide substitutions. Three of these codons were still synonymous, whereas the remaining nine encoded amino acid substitutions. The pattern of double nucleotide substitution into three of these nine codons was consistent with two successive amino acid substitutions at the same position in the capsid polypeptide.
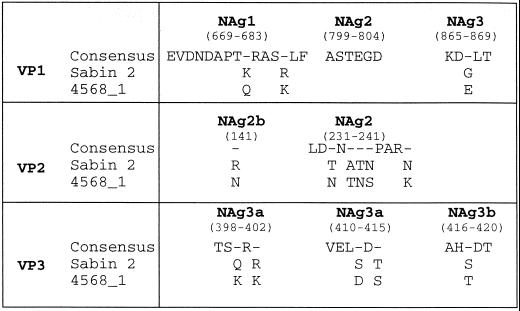
FIG. 3.
Amino acid substitutions in NAg sites of 4568-1. Nonhomologous amino acids are indicated below the consensus sequence for each NAg site. Amino acid positions are numbered according to that for Sabin 2 (26).
Two key mutations (a G→A substitution at nucleotide position 481 in the 5′-UTR and an isoleucine→threonine substitution at amino acid position 143 of VP1) have been found to be associated with reversion of the attenuated phenotype of the Sabin 2 OPV strain (12). Both of these substitutions were found in the genome of 4568-1.
Antigenic drift of 4568-1.
Because isolate 4568-1 was initially recognized as an antigenic variant when tested with highly specific animal sera, we sought to determine whether the antigenic differences between 4568-1 and Sabin 2 would also be evident when they were tested against human immune sera. Fifty serum samples, collected from a cohort of 15-year-old high school students who had received at least three OPV doses in early childhood and a booster dose during the 1988 Israeli outbreak (22), were tested for titers of neutralizing antibodies to 4568-1, Sabin 2, and MEF-1 (the type 2 IPV strain). While all 50 serum samples contained protective levels of neutralizing antibody to all three type 2 polioviruses, the geometric mean titers (GMT) of antibody to 4568-1 were significantly lower (GMT, 47) than those to Sabin 2 (GMT, 162) or MEF-1 (GMT, 108).
Neurovirulence of 4568-1.
The neurovirulence of 4568-1 was examined in PVR-Tg21 transgenic mice (7, 21) in comparison with Sabin 2 and the nonattenuated reference strains MEF-1 and Mahoney (type 1). Under our assay conditions, inoculation with 108 PFU/mouse and follow-up for 14 days after challenge with virus, the original Sabin 2 strain (n = 10 mice tested) showed maximum attenuation, with an MHT of 14.0 ± 0.0 days (mean ± standard error of the mean). The equivalent challenge dose of the nonattenuated polioviruses Mahoney (n = 10) and MEF-1 (n = 10) gave MHTs of 4.7 ± 1.14 and 9.6 ± 1.49 days, respectively. The Sabin 2-derived field isolate 4568-1 (n = 20) had clearly lost the attenuated phenotype, since it had a shorter MHT, 3.1 ± 0.07 days, than either of the two reference nonattenuated polioviruses.
Estimation of the duration of replication of the 4568-1 lineage.
The duration of replication of an OPV-derived isolate after the initiating vaccine dose can be estimated from the degree of its sequence divergence from its parental Sabin strain. The VP1 evolution rate for type 1 poliovirus, whether a vaccine-derived strain replicating in an immunodeficient patient (11) or a wild poliovirus circulating over a period of 10 years (De et al., Abstr. 17th Annu. Meet. Am. Soc. Virol.) or 1 year (22), is ∼3% third-codon-position substitutions/year. Similar values have been obtained for the entire P1 capsid region (4) and for a Sabin 3-derived poliovirus (15). The observed number of third-codon-position differences between 4568-1 and Sabin 2 over the entire P1 capsid region was 166 of 882 codons (18.8%). By assuming that the evolution rate for the P1 region of 4568-1 was constant throughout the entire period of replication and similar to the rates observed for other polioviruses (3% third-position substitutions/year), and without correcting for the small effects of multiple substitutions at a site, we estimate that the total period of replication was about 6 years.
DISCUSSION
Environmental sampling has been a powerful tool for poliovirus surveillance in Israel and the Palestinian Authority (14). Several independent introductions of wild polioviruses (type 1 in 1991, 1994 to 1995, and 1996; type 3 in 1990) have been detected since the appearance of the last polio cases in Israel, Gaza, and the West Bank in 1987 to 1988 (14, 22; Shulman et al., Abstr. Xth Int. Cong. Virol.). Because OPV is widely used (in combination with IPV [25, 27] by both Israeli and Palestinian health authorities (14), OPV-derived viruses are prevalent in local wastewater. The key to detecting polioviruses that have properties distinct from most OPV-derived strains is the use of a double-selective culture technique (13) that favors the growth of RCT-positive isolates. The yield of RCT-positive isolates has increased since the replacement of BGM cells in the culture system with L20B cells that express the human gene for the PVR (16, 20). All RCT-positive isolates are subsequently characterized by molecular and antigenic methods.
Isolate 4568-1 was unusual because it was highly divergent from the prototype Sabin 2 OPV strain. A nonvaccine origin for 4568-1 could be ruled out because it is genetically much closer to Sabin 2 than to any of the wild polioviruses found in the Middle East or elsewhere since 1980 (10), and because the only known remaining foci of endemicity for wild type 2 poliovirus in 1998 were in northern India (3), and those wild viruses were unrelated to 4568-1. On the other hand, 4568-1 was similar to, but not derived from, highly divergent Sabin 2-derived polioviruses isolated from immunodeficient patients in the United States.
The extent of sequence divergence of 4568-1 from Sabin 2 suggests that the virus had replicated in one or more people for about 6 years since the administration of the initiating OPV dose. This estimate of the duration of replication is only approximate and is based upon the assumptions that rate of VP1 evolution for poliovirus type 2 is essentially constant over the period of replication and similar to the rates observed for types 1 and 3 (11, 15, 22; De et al., Abstr. 17th Annu. Meet. Am. Soc. Virol.; Heim et al., Abstr. Eur. Soc. Clin. Virol.; C.-F. Yang, S.-J. Yang, M. A. Pallansch, and O. M. Kew, Abstr. 15th Annu. Meet. Am. Soc. Virol., 1996).
We cannot distinguish from the existing evidence whether virus replication was restricted to a single chronically infected individual (such as an immunodeficient patient) or whether a succession of people were infected through continuous transmission. The largely urban communities sampled at the wastewater collection site for isolate 4568-1 have a combined population of 1.3 million and include both long-term resident and recent immigrant populations. Additional sampling sites have been selected within this community to better localize the source of the unusual type 2 virus. Although it is possible that we have detected a unique infection, it appears more likely that viruses related to 4568-1 were more widely distributed in the community (8). Moreover, we do not know whether the initial infection occurred within Israel or came from some external source. One factor which might potentially permit the spread of the unusual vaccine strain derivative is the reduced titers of neutralizing antibodies to 4568-1 relative to Sabin 2 and MEF-1 in the sera of children immunized with OPV. This factor may be negligible in communities with high OPV coverage but could be important in communities with lower OPV coverage and where the immunogenicity of OPV is reduced (25, 27).
It appears unlikely that the presence of a neurovirulent, highly diverged Sabin 2 derivative currently presents any significantly increased risk to a well-immunized community over that already presented by the frequent excretion of neurovirulent OPV-derived strains by normal OPV recipients (1, 7, 23). No paralytic case in Israel has been found to be associated with any virus related to isolate 4568-1 or to any other highly diverged Sabin strain derivative (14). However, under the current conditions of high overall OPV coverage in Israel, cases associated with neurovirulent type 2 polioviruses would presumably be quite rare. In the prevaccine era, type 2 poliovirus infections were found to have the lowest paralytic attack rates (∼1 case per 2,500 infections) of the three poliovirus serotypes (19), and widespread immunization with OPV has reduced type 2 attack rates to immeasurably low levels. Thus, it is possible that type 2 circulation could occur in the absence of paralytic cases within a localized community with suboptimal OPV coverage, as apparently occurred with wild type 1 polioviruses in Gaza in 1996 (14; Shulman et al., Abstr. Xth Int. Cong. Virol.).
Neurovirulent OPV-derived polioviruses could present a potentially serious global health risk if they were present in any community after the cessation of immunization with OPV following certification of the interruption of all wild poliovirus transmission (4). Little information is currently available on the environmental prevalence of highly divergent OPV-derived polioviruses. Environmental studies in most countries have been impeded by the continuous presence of typical OPV-derived strains in wastewaters. However, the availability of efficient methods to select for atypical OPV-derived environmental isolates (13) opens the way to assess the possible broader significance of the findings reported here.
ACKNOWLEDGMENTS
This study was partially supported by The Israel-U.S. Binational Science Foundation grant 96-00159.
We also acknowledge the District Health Departments in Israel and the Palestinian Authority who routinely supply the sewage samples. Jagadish Deshpande, Enterovirus Research Institute, Mumbai, India, contributed the Indian PV2 isolate, and Edson da Silva, Instituto Oswaldo Cruz, Rio de Janeiro, Brazil provided the Brazilian PV2 isolates. We also acknowledge Cara Burns, Lina De, Jaume Jorba, David R. Kilkpatrick, Chen-Fu Yang, and Su-Ju Yang from the Centers for Disease Control and Prevention for their assistance.
REFERENCES
- 1.Abraham R, Minor P, Dunn G, Modlin J F, Ogra P L. Shedding of virulent poliovirus revertants during immunization with oral poliovirus vaccine after prior immunization with inactivated polio vaccine. J Infect Dis. 1993;168:1105–1109. doi: 10.1093/infdis/168.5.1105. [DOI] [PubMed] [Google Scholar]
- 2.Alexander J P, Jr, Gary H E, Jr, Pallansch M A. Duration of poliovirus excretion and its implications for acute flaccid paralysis surveillance: a review of the literature. J Infect Dis. 1997;175(Suppl. 1):S176–S182. doi: 10.1093/infdis/175.supplement_1.s176. [DOI] [PubMed] [Google Scholar]
- 3.Centers for Disease Control and Prevention. Progress toward the global interruption of wild poliovirus type 2 transmission, 1999. Morb Mortal Wkly Rep. 1999;48:736–738. [PubMed] [Google Scholar]
- 4.Cochi S L, Sutter R W, Kew O M, Pallansch M A, Dowdle W R. A decision tree for stopping polio immunization. Technical Consultation on Global Eradication of Poliomyelitis. EPI/POLIO/TECH.97/WP18. Geneva, Switzerland: World Health Organization; 1997. [Google Scholar]
- 5.Crainic R, Couillin P, Blondel B, Cabau N, Boue A, Horodniceanu F. Natural variation of poliovirus neutralization epitopes. Infect Immun. 1983;41:1217–1225. doi: 10.1128/iai.41.3.1217-1225.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.De L, Nottay B, Yang C-F, Holloway B P, Pallansch M, Kew O. Identification of vaccine-related poliovirus by hybridization with specific RNA probes. J Clin Microbiol. 1995;33:562–571. doi: 10.1128/jcm.33.3.562-571.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Georgescu M-M, Delpeyroux F, Tardy-Panit M, Balanant J, Combiescu M, Combiescu A A, Guillot S, Crainic R. High diversity of poliovirus strains isolated from the central nervous system from patients with vaccine-associated paralytic poliomyelitis. J Virol. 1994;68:8089–8101. doi: 10.1128/jvi.68.12.8089-8101.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hovi T. Review of environmental surveillance for wild polioviruses. Global Polio Laboratory Network Meeting. WHO/EPPI/Polio/LAB98/WP. Geneva, Switzerland: World Health Organization; 1998. Potential role of environmental surveillance for wild polioviruses in the global eradication programme. [Google Scholar]
- 9.Huovilainen A, Kinnunen L, Ferguson M, Hovi T. Antigenic variation among 173 strains of type 3 poliovirus isolated in Finland during the 1984 to 1985 outbreak. J Gen Virol. 1988;69:1941–1948. doi: 10.1099/0022-1317-69-8-1941. [DOI] [PubMed] [Google Scholar]
- 10.Kew O M, Mulders M N, Lipskaya G Y, da Silva E E, Pallansch M A. Molecular epidemiology of polioviruses. Semin Virol. 1995;6:401–414. [Google Scholar]
- 11.Kew O M, Sutter R W, Nottay B K, McDonough M J, Prevots D R, Quick L, Pallansch M A. Prolonged replication of a type 1 vaccine-derived poliovirus in an immunodeficient patient. J Clin Microbiol. 1998;36:2893–2899. doi: 10.1128/jcm.36.10.2893-2899.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Macadam A J, Pollard S R, Ferguson G, Skuce R, Wood D, Almond J W, Minor P D. Genetic basis of attenuation of the Sabin type 2 vaccine strain of poliovirus in primates. Virology. 1993;192:18–26. doi: 10.1006/viro.1993.1003. [DOI] [PubMed] [Google Scholar]
- 13.Manor Y, Handsher R, Halmut T, Neuman M, Abramovitz B, Mates A, Mendelson E. A double-selective tissue culture system for isolation of wild-type poliovirus from sewage applied in a long-term environmental surveillance. Appl Environ Microbiol. 1999;65:1794–1797. doi: 10.1128/aem.65.4.1794-1797.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Manor Y, Handsher R, Halmut T, Neuman M, Bobrov A, Rudich H, Vonsover A, Shulman L, Kew O, Mendelson E. Detection of poliovirus circulation by environmental surveillance in the absence of clinical cases in Israel and the Palestinian Authority. J Clin Microbiol. 1999;37:1670–1675. doi: 10.1128/jcm.37.6.1670-1675.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Martin J, Dunn G, Hull R, Patel V, Minor P D. Evolution of the Sabin strain of type 3 poliovirus in an immunodeficient patient during the entire 637-day period of virus excretion. J Virol. 2000;74:3001–3010. doi: 10.1128/jvi.74.7.3001-3010.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mendelsohn C D, Wimmer E, Racaniello V R. Cellular receptor for poliovirus: molecular cloning, nucleotide sequencing, and expression of a new member of the immunoglobulin superfamily. Cell. 1989;56:855–865. doi: 10.1016/0092-8674(89)90690-9. [DOI] [PubMed] [Google Scholar]
- 17.Minor P D. Antigenic structure of picornavirus. Curr Top Microbiol Immunol. 1990;161:122–151. doi: 10.1007/978-3-642-75602-3_5. [DOI] [PubMed] [Google Scholar]
- 18.Minor P D, Ferguson M, Evans D M A, Almond J W, Icenogle J P. Antigenic structure of polioviruses of serotypes 1, 2, and 3. J Gen Virol. 1986;67:1283–1291. doi: 10.1099/0022-1317-67-7-1283. [DOI] [PubMed] [Google Scholar]
- 19.Nathanson N, Martin J R. The epidemiology of poliomyelitis: enigmas surrounding its appearance, epidemicity, and disappearance. Am J Epidemiol. 1979;110:672–692. doi: 10.1093/oxfordjournals.aje.a112848. [DOI] [PubMed] [Google Scholar]
- 20.Pipkin P A, Wood D J, Racaniello V R, Minor P D. Characterization of L cells expressing the human poliovirus receptor for the specific detection of polioviruses in vitro. J Virol Methods. 1993;41:333–340. doi: 10.1016/0166-0934(93)90022-j. [DOI] [PubMed] [Google Scholar]
- 21.Ren R, Racaniello V R. Human poliovirus receptor gene expression and poliovirus tissue tropism in transgenic mice. J Virol. 1992;66:296–304. doi: 10.1128/jvi.66.1.296-304.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shulman L M, Handsher R, Yang C-F, Yang S-J, Manor J, Vonsover A, Grossman Z, Pallansch M, Mendelson E, Kew O. Resolution of the pathways of poliovirus type 1 transmission during an outbreak. J Clin Microbiol. 2000;38:945–952. doi: 10.1128/jcm.38.3.945-952.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Strebel P M, Sutter R W, Cochi S L, Biellik R J, Brink E W, Kew O M, Pallansch M A, Orenstein W A, Hinman A R. Epidemiology of poliomyelitis in the United States one decade after the last reported case of indigenous wild virus-associated disease. Clin Infect Dis. 1992;14:568–579. doi: 10.1093/clinids/14.2.568. [DOI] [PubMed] [Google Scholar]
- 24.Sutter R W, Prevots R. Vaccine-associated paralytic poliomyelitis among immunodeficient persons. Infect Med. 1994;11:426–438. doi: 10.1086/379791. [DOI] [PubMed] [Google Scholar]
- 25.Swartz T A, Handsher R, Manor Y, Stoeckel P, Barkay A, Mendelson E, Leventhal A. Immune response to an intercalated enhanced inactivated polio vaccine/oral polio vaccine programme in Israel: impact on the control of poliomyelitis. Vaccine. 1998;16:2090–2095. doi: 10.1016/s0264-410x(98)00071-1. [DOI] [PubMed] [Google Scholar]
- 26.Toyoda H, Kohara M, Kataoka Y, Suganuma T, Omata T, Imura N, Nomoto A. Complete nucleotide sequences of all three poliovirus serotype genomes. J Mol Biol. 1984;174:561–585. doi: 10.1016/0022-2836(84)90084-6. [DOI] [PubMed] [Google Scholar]
- 27.Tulchinsky T, Abed Y, Handsher R, Toubassi N, Acker C, Melnick J. Successful control of poliomyelitis by a combined OPV/IPV polio vaccine program in the West Bank and Gaza, 1978–93. Isr J Med Sci. 1994;30:489–494. [PubMed] [Google Scholar]
- 28.Yang C-F, De L, Holloway B P, Pallansch M A, Kew O M. Detection and identification of vaccine-related polioviruses by the polymerase chain reaction. Virus Res. 1991;20:159–179. doi: 10.1016/0168-1702(91)90107-7. [DOI] [PubMed] [Google Scholar]