Abstract
This study determined the proficiencies of laboratories measuring human immunodeficiency virus type 1 (HIV-1) viral loads and the accuracies of two assays used for HIV-1 viral load measurement in Australia and investigated the variability of the new versions of these assays. Quality assessment program panels containing (i) dilutions of HIV-1 subtype B, (ii) replicates of identical samples of HIV-1 subtype B, and (iii) samples of subtype E and B were tested by laboratories. Total variability (within and between laboratories) was tested with quality control samples. The coefficients of variation (CVs) for the Roche AMPLICOR HIV-1 MONITOR version (v) 1.0 and Chiron Quantiplex bDNA 2.0 assays ranged from 53 to 87% and 22 to 31%, respectively. The widespread occurrence of invalid runs with the AMPLICOR HIV-1 MONITOR 1.0 assay was identified. The CVs of the new versions of the assays were 82 to 86% for the AMPLICOR HIV-1 MONITOR v 1.5 assay and 16 to 23% for the Quantiplex bDNA 3.0 assay. For virus dilution samples, all but 5 of 19 laboratories obtained results within 2 standard deviations of the mean. The Quantiplex bDNA 2.0 assay reported values lower than those reported by the AMPLICOR HIV-1 MONITOR version 1.0 assay for samples containing HIV-1 subtype B, whereas the reverse was true for subtype E. Identification and resolution of the problem of invalid runs markedly improved the quality of HIV-1 viral load testing. The variability observed between laboratories and between assays, even the most recent versions, dictates that monitoring of viral load in an individual should always be by the same laboratory and by the same assay. Results for an individual which differ by less than 0.5 log10 HIV-1 RNA copy number/ml should not be considered clinically significant.
The quantification of human immunodeficiency virus (HIV) to establish the viral load or amount of measurable virus in the blood has become essential to the clinical management of HIV infection. It has been demonstrated that the viral load is the most reliable predictor of the course of infection if it is estimated once a steady state in viremia is achieved following seroconversion (8). Viral load increases with the development of resistance to antiretroviral therapy or with a detrimental alteration in clinical status (5). It has been advocated that the aim of current antiviral therapy for HIV infection should be to maintain the viral load below the level of detection of the available assays (3, 7).
The majority of commercial assays for the quantification of viral load use techniques involving amplification either of target nucleic acid (AMPLICOR HIV-1 MONITOR [Roche Molecular Systems, Pleasanton, Calif.]) or a hybridization signal (Quantiplex HIV RNA [Chiron Corporation, Emeryville, Calif., now Bayer Diagnostics], abbreviated in the text as Quantiplex bDNA). The majority of the data in this report were generated with version (v) 1.0 of the AMPLICOR HIV-1 MONITOR assay and the Quantiplex bDNA 2.0 assay. The report presents limited data from tests with the latest versions of these assays (v 1.5 and 3.0, respectively) over the period encompassed by the present study. The NucliSens HIV-1 QT assay (Organon Teknika, Boxtel, The Netherlands), which uses amplification of nucleic acid sequences, is not widely used for therapeutic monitoring in Australia.
It is a condition of registration of all HIV assays for use in Australia that laboratories that use the assay participate in the quality assurance (QA) programs run by the National Serology Reference Laboratory (NRL), Melbourne, Australia. This report presents the findings of the HIV type 1 (HIV-1) viral load QA program in Australia for the AMPLICOR HIV-1 MONITOR and Quantiplex bDNA assays. We investigated the proficiencies of laboratories measuring viral load across a range of dilutions (range of viral RNA copy number per milliliter), the reproducibility of determinations for replicate samples, and whether the performances of the assays were consistent between HIV-1 subtypes. The report also addresses the implication of reporting of data from invalid runs.
MATERIALS AND METHODS
Nineteen laboratories (testing sites) in Australia and one in New Zealand participate in the NRL's QA program for HIV viral load testing. The program was instituted in August 1996 and consists of two elements: a quality control (QC) program and a quality assessment program (QAP). All laboratories return QAP and QC program data to the NRL.
QC program.
A QC program provides a mechanism to monitor the run-to-run performance of an assay. For the HIV viral load QC program, NRL prepared and distributed the QC samples. The between-run, between-laboratory, and between-lot-number performances of the assays were monitored for laboratories returning data for QC samples to NRL.
QC samples were prepared in a class II biohazard cabinet by dilution of a stock HIV-1 subtype B (HIV-1 Ba-L) culture supernatant in pooled, defibrinated and delipidated normal human plasma (Basematrix; Boston Biomedica, Inc., West Bridgewater, Mass.). The dilutions were mixed in a sterile bottle, on a slow vortex, overnight at 4°C. Convenient single-use volumes (250 μl for the AMPLICOR HIV-1 MONITOR assays, 1,250 μl for the Quantiplex bDNA assay) were aliquoted into sterile screw-cap microcentrifuge tubes (Sarstedt, Nümbrecht, Germany), and the tubes and their contents were frozen at −70°C prior to dispatch to the laboratories on dry ice. Laboratories were required to include the QC samples in at least every fourth assay run and with a change of lot number or a change of operator. QC sample data were returned to NRL on a monthly basis and included not only the viral loads measured in the QC samples but also the numbers of invalid runs, the reasons for invalid runs, as well as the numbers of invalid patient results.
Since the institution of the HIV viral load QC program, three sets of coded QC samples have been circulated by the NRL. In the most recent set, samples QC105 and QC105C were designed to have loads that fall into the lower range (400 to 5,000 copies/ml), with QC105 to be used in the AMPLICOR HIV-1 MONITOR assay and QC105C to be used in the Quantiplex bDNA assay. Preparation of separate QC samples to monitor the lower ranges of the assays was necessary because the Quantiplex bDNA assay always returned lower values for the subtype B culture supernatant used for preparation of the QC samples than the AMPLICOR HIV-1 MONITOR assay did. Two samples with loads in the range of 103 copies/ml were prepared to ensure that the lower target value was achievable in both assays. The detection ranges stated by the manufacturers were 400 to 750,000 copies/ml for the AMPLICOR HIV-1 MONITOR v 1.0 assay and 500 to 500,000 copies/ml for the Quantiplex bDNA 2.0 assay. In the present report, only the results for the first set (QC101 and QC102; August 1996 to April 1997) and most recent set (QC105, QC105C, and QC106; February 1998 to June 1999) of QC samples are presented. The results obtained with the other QC set were similar to those obtained with the QC101 and QC102 set and have been omitted to avoid unnecessary repetition of results. Preliminary results obtained with a separate set of QC samples (108P and 109P) with low viral loads designed for use with the AMPLICOR HIV-1 MONITOR v 1.5 assay with Ultrasensitive sample preparation are also reported. These samples were prepared in normal human plasma because samples prepared in Basematrix proved unsuitable for ultracentrifugation. We have previously found that centrifugation in Basematrix led to inconsistent pelleting of the RNA and frequent loss of the pellet when the supernatant was removed.
Run validity.
Run validity was determined according to the manufacturer criteria. Two positive controls and one negative control are provided by the manufacturer for the AMPLICOR HIV-1 MONITOR assay, with a designated range into which the results for those controls must fall for an assay run to be valid. The Quantiplex bDNA assays include a standard curve with four standards, each of which is tested in duplicate in every run. The software accompanying the detection luminometer determines whether the linearities and variabilities of the values obtained for the standards and controls are acceptable for a valid run.
QAP.
HIV viral load QAP panels were distributed twice yearly to the 20 participating laboratories and consisted of up to eight samples, coded to disguise their identities. Laboratories were provided with a results sheet on which to record run information and results. Data were returned to the NRL for collation and analysis. To maintain confidentiality, each laboratory was assigned a unique code number under which results were reported (note that some laboratories may appear with more than one laboratory code in this study).
The first QAP panel, panel VLQA 97-1, contained five coded samples, four of which were members of a fivefold dilution series of HIV-1 Ba-L in Basematrix (subtype B), including one dilution presented in duplicate. The fifth sample contained diluent only.
The second QAP panel, panel VLQA 98-1, investigated whether sites achieved similar values within a run. Two samples (QC105 or QC105C and QC106) were coded and were included four times each in the panel. Panels containing QC105 or QC105C were distributed to laboratories that use the AMPLICOR HIV-1 MONITOR v 1.0 or Quantiplex bDNA 2.0 assay, respectively, for the reasons given above. In addition, laboratories were also requested to run the QC samples in the same run as the QAP samples, when practical.
The third panel, panel VLQA 98-2, contained five coded plasma samples from subjects infected with HIV-1 subtype B (one sample) or subtype E (four samples) and a control sample of anti-HIV-negative, pooled normal human plasma. These samples were estimated not to contain very high viral loads (<20,000 RNA copies/ml), but their loads were expected to fall comfortably within the linear range of the assays at the various testing sites. Again, when practical, laboratories were also requested to run QC samples QC105 or QC105C and QC106 (subtype B-spiked normal human plasma) in the same run as the QAP samples.
Data presentation and statistical analysis.
For the QC data, results for each QC sample are presented as the mean ± 1 standard deviation (SD) HIV-1 RNA copy number per milliliter. The coefficient of variation (CV) is given as a percentage and has been calculated as the SD divided by the mean. For the QAP panel data, the mean ± 2 SD was calculated from the raw data for the number of HIV-1 RNA copy number per milliliter. When the data are presented as residuals, values for copy number per milliliter were first transformed to log10 values and residuals were derived as the difference between the reported log10 copy number per milliliter and the mean log10 copy number per milliliter for each sample.
Statistical analysis.
The results obtained for QC samples were compared between the two assay types by the t test. The incidence of invalid runs between different assay lot numbers was analyzed by the chi-square test. Between-sample data for the QAP panels were analyzed by two-way analysis of variance (ANOVA) to test variance between samples in the panel and between testing sites. When significance was observed across the ANOVA table, a t test was applied to locate significance within the data set.
RESULTS
QC program.
The results from two separate sets of QC samples (set QC101 and QC102 and set QC105, QC105C, and QC106) tested by previous assay versions and one set tested by the new versions of the AMPLICOR HIV-1 MONITOR and Quantiplex bDNA assays are presented in Table 1. Only results obtained from valid runs are included (specific data for invalid run rates in each assay appear below). The number of observations for the AMPLICOR HIV-1 MONITOR v 1.0 assay is substantially greater than the number of observations for the Quantiplex bDNA 2.0 assay due to the more widespread use of the former: 17 laboratories used the AMPLICOR HIV-1 MONITOR assay and between 2 and 6 laboratories used the Quantiplex bDNA assay (some laboratories used both assays for a period within the study). Despite the smaller numbers of observations, the variability of the data from the Quantiplex bDNA 2.0 assay was consistently lower than the variability of the data from the AMPLICOR HIV-1 MONITOR v 1.0 assay across both QC sample sets. The data obtained by the AMPLICOR HIV-1 MONITOR v 1.0 assay had a reduced variability with the more recent QC sample set (Table 1) compared to the data obtained a maximum of 2 years previously with the earlier QC101 and QC102 sample set (Table 1). Values for HIV-1 copy number per milliliter were consistently lower by the Quantiplex bDNA 2.0 assay than by the AMPLICOR HIV-1 MONITOR v 1.0 assay with these QC samples containing subtype B (P < 0.001).
TABLE 1.
HIV-1 RNA load measured by AMPLICOR HIV-1 MONITOR v 1.0 and v 1.5 assays and Quantiplex bDNA 2.0 and 3.0 assays in three separately distributed sets of QC samples
| Sample | AMPLICOR HIV-2 MONITOR v 1.0
|
Quantiplex bDNA 2.0
|
AMPLICOR HIV-1 MONITOR v 1.5
|
Quantiplex bDNA 3.0
|
AMPLICOR HIV-1 MONITOR v 1.5a
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| nb | Copy no./ml (103)c | CV (%) | n | Copy no./ml (103)c | CV (%) | n | Copy no./ml (103)c | CV (%) | n | Copy no./ml (103)c | CV (%) | n | Copy no./ml (103)c | CV (%) | |
| QC101 | 468 | 40.8 ± 24.6 | 60.3 | 77 | 22.9 ± 7.1 | 31.0 | |||||||||
| QC102 | 472 | 228.4 ± 198.4 | 86.9 | 77 | 129.4 ± 29.5 | 22.8 | |||||||||
| QC105 | 789 | 4.5 ± 2.4 | 53 | Not applicable | 18 | 5.3 ± 3.0 | 57 | Not applicable | |||||||
| QC105C | Not applicable | 158 | 2.7 ± 0.9 | 32 | Not applicable | 29 | 6.8 ± 1.1 | 16 | |||||||
| QC106 | 788 | 421.9 ± 249.2 | 59 | 159 | 125.4 ± 30.4 | 24 | 20 | 366.9 ± 126.0 | 34 | 28 | 242.8 ± 36.5 | 15 | |||
| QC108P | 11 | 0.87 ± 0.2 | 23 | 30 | 0.13 ± 0.10 | 82 | |||||||||
| QC109P | 11 | 11.7 ± 1.8 | 16 | 31 | 23.1 ± 19.7 | 86 | |||||||||
Ultrasensitive sample preparation.
n, number of samples.
Values are means ± SDs.
Data obtained to date with the new versions of each assay are shown in Table 1. The CV appears to be lower for the Quantiplex bDNA 3.0 assay than for the earlier version of the assay. This may be due to the fact that the later version is almost entirely automated. The CV does not appear to have changed for the AMPLICOR HIV-1 MONITOR assay between v 1.0 and v 1.5, but further data may be required to establish this with certainty.
Data from two laboratories that used Ultrasensitive sample preparation in conjunction with the AMPLICOR HIV-1 MONITOR v 1.5 assay are shown in Table 1. Data presented for the Quantiplex bDNA v 3.0 assay in Table 1 were generated in only one laboratory.
Invalid assay runs. (i) AMPLICOR HIV-1 MONITOR v 1.0 assay.
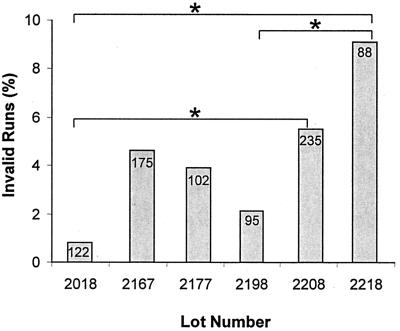
Early monitoring of HIV viral load tests revealed that approximately half the laboratories did not achieve consistently valid runs. Invalid runs were more likely to occur for particular lot numbers of AMPLICOR HIV-1 MONITOR v 1.0. Figure 1 shows the invalid run rate, according to the manufacturer's criteria, for six different lots supplied in 1998. The variability in the number of invalid runs between lots was not predictable from one lot to the next.
FIG. 1.
Incidence of invalid runs, by lot number, as proportion of total runs (shown by the number within each bar) of the AMPLICOR HIV-1 MONITOR assay in all laboratories performing HIV viral load testing in Australia. ∗, P < 0.05.
(ii) Quantiplex bDNA 2.0 assay.
During the reporting period for the first QC sample set for viral load (QC101 and QC102), the Quantiplex bDNA 2.0 assay had fewer invalid runs, with 5 of 77 runs being invalid. Results for a small percentage (<10%) of individual samples were rejected according to the manufacturer's criteria, because the CV between the mandatory two replicates per sample was >40% for negative samples (<500 HIV-1 RNA copies per ml) and >30% for positive samples. When repeated, these samples usually yielded valid results.
In the reporting period with QC105C and QC106, a low rate of invalid runs continued: 2 of 91 runs were invalid. There was a period of approximately 4 months in which one laboratory that used the Quantiplex bDNA 2.0 assay experienced an increase in the number of invalid runs (from August to November 1997) which was not seen in other laboratories using the assay. However, despite intensive investigation by the laboratory and the manufacturer, no clear cause of the invalid runs was established.
Insufficient data have been generated to permit analysis of the invalid run rate with the new versions of each assay.
QAP. (i) Panel VLQA 97-1: accuracy across a range of viral concentrations.
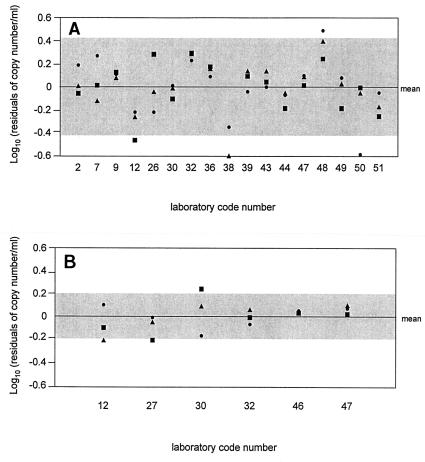
The results of testing of panel VLQA 97-1 by laboratories using the AMPLICOR HIV-1 MONITOR v 1.0 and Quantiplex bDNA 2.0 assays are shown in Fig. 2. Results were received from 19 laboratories: 13 used the AMPLICOR HIV-1 MONITOR v 1.0 assay only, 2 used the Quantiplex bDNA 2.0 assay only, and 4 used both assays. The data for the fivefold dilution series are presented as residuals (difference of the reported value of log10 HIV-1 RNA copy number per milliliter for each sample from the mean log10 HIV-1 RNA copy number per milliliter) and are given by testing site for each sample in the panel (dilution). The results for the 1:25 dilution were averaged, as this dilution was presented in duplicate in the panel. The panel also contained a negative sample with diluent only, which is not included on the graph, giving a total of three results for each testing site.
FIG. 2.
Results obtained by individual laboratories, identified by code number, for QAP panel VLQA 97-1, a panel of samples with a range of HIV-1 dilutions. The panel contained an undiluted sample (●), a 1/5 dilution of a sample (▴), and a 1/25 dilution of a sample in duplicate, shown as the average (■). Data are presented as residuals (difference of the reported value from the mean). The mean is indicated by the horizontal line, and 2 SDs above and below the mean are shown by the shaded region. (A) AMPLICOR HIV-1 MONITOR assay; (B) Quantiplex bDNA assay.
All but four results produced at any site by the AMPLICOR HIV-1 MONITOR v 1.0 assay were within 2 SDs of the mean (Fig. 2A). The four reported values which fell outside 2 SDs of the mean came from four separate testing sites and were for three different samples; i.e., it was not only one site or one sample concentration which provided the widely variant results. Only one value was outside 2 SDs by the Quantiplex bDNA 2.0 assay (Fig. 2B). Because fewer testing sites used the Quantiplex bDNA 2.0 assay, the SD was expected to have been greater than that seen with the AMPLICOR HIV-1 MONITOR assay. In spite of this, the log10 SD obtained by the Quantiplex bDNA 2.0 assay was approximately half that found by the AMPLICOR HIV-1 MONITOR v 1.0 assay for each dilution.
(ii) Panel VLQA 98-1: within-run reproducibility.
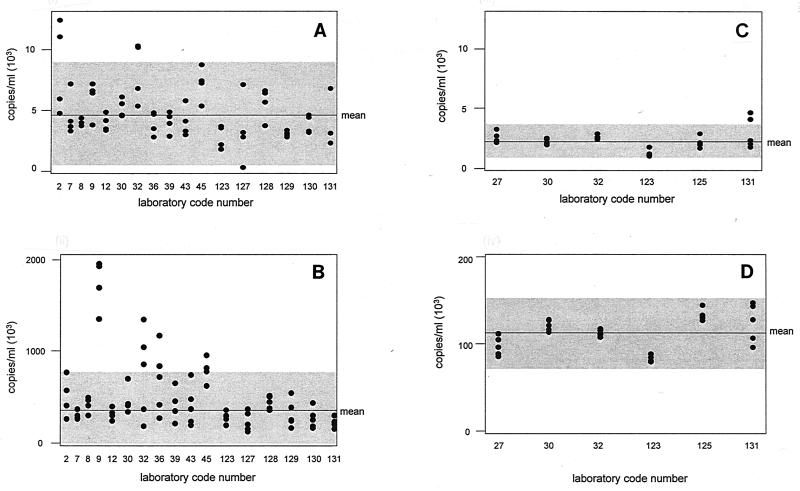
QAP panel VLQA 98-1 was decoded, and the values reported by each testing site for samples QC105, QC105C, and QC106 are shown in Fig. 3. When testing sites had included the matching samples for quality control, they were reported in the data set, giving five datum points for some testing sites. For sample QC106 tested by the AMPLICOR HIV-1 MONITOR v 1.0 assay, 14 of 81 results (17%) were greater than 7.5 × 105 copies/ml, the upper detection limit of the assay. To calculate the mean ± SD (Fig. 3B), these results have been included in the data set as 7.5 × 105 copies/ml, as it was not considered valid to extrapolate values beyond the upper limit of detection cited by the manufacturer. In practice, this is the value which would be reported clinically for a sample with a viral load which exceeded the upper limit of detection.
FIG. 3.
Results obtained by individual laboratories, identified by code number, for QAP panel VLQA 98-1, a panel of replicates of HIV-1 QC samples QC105, QC105C, and QC106. The mean is indicated by the horizontal line, and 2 SDs above and below the mean are shown by the shaded region. (A and B) Data for QC105 and QC106, respectively, obtained by the AMPLICOR HIV-1 MONITOR assay; (C and D) data for QC105C and QC106, respectively, obtained by the Quantiplex bDNA assay. Note the 10-fold difference in scale between panels B and D (results for QC106).
No significant difference was found between the results obtained for any of the samples in the panel by either the AMPLICOR HIV-1 MONITOR v 1.0 assay or the Quantiplex bDNA 2.0 assay. While most laboratories achieved clustered results within 2 SDs of the mean when testing replicate samples (Fig. 3) using the AMPLICOR HIV-1 MONITOR v 1.0 assay, results from four testing sites were spread over a wide range (a wide spread is defined as some of the results being greater than 2 SDs from the mean when other results from the same site fell on the mean or on the opposite side of the mean). These were testing sites 2 and 127, with the sample with a viral load in the lower range (QC105) (Fig. 3A), and sites 32 and 36, with the sample with a viral load in the upper range (QC106) (Fig. 3B). Testing sites 9 and 45 produced clustered results for QC106, all or some of which were greater than 2 SDs above the mean (Fig. 3B).
By the Quantiplex bDNA 2.0 assay, five of six laboratories reported results that were within 2 SDs of the mean for the sample with a viral load in the lower range (sample QC105C). One laboratory reported 2 of 5 results for QC105C greater than 2 SDs of the mean (Fig. 3C). Results reported from this site were spread over a wider range than those reported by other users for replicates of both QC105C and QC106 (Fig. 3C and D), although no site reported results greater than 2 SDs of the mean for sample QC106, which had a viral load in the upper range (Fig. 3D).
(iii) Panel VLQA 98-2: detection of HIV-1 subtypes.
The results obtained from testing sites for panel VLQA 98-2 are presented in Table 2. Results were received from 19 laboratories. Only one laboratory used both the AMPLICOR HIV-1 MONITOR v 1.0 and the Quantiplex bDNA 2.0 assays. Two laboratories used the Quantiplex bDNA 2.0 assay only, but in one of those laboratories, the run containing the QAP samples was invalid. Therefore, those results were excluded from the analysis. The remaining 16 laboratories used the AMPLICOR HIV-1 MONITOR v 1.0 assay only. All testing sites except site 128 included samples QC105 or QC105C and QC106 in the run. Site 39 included only QC106. Data for the HIV-1-negative sample were <400 copies/ml for the AMPLICOR HIV-1 MONITOR v 1.0 assay and <500 copies/ml for the Quantiplex bDNA 2.0 assay (data not shown). For the same sample, the averaged results obtained for the samples with HIV-1 subtype by the AMPLICOR HIV-1 MONITOR v 1.0 assay were consistently lower than those obtained from either laboratory by the Quantiplex bDNA 2.0 assay. In contrast, for the samples with HIV-1 subtype B, the AMPLICOR HIV-1 MONITOR v 1.0 assay viral load results were higher than the Quantiplex bDNA 2.0 assay results. This was true for both the subtype B-containing sample in the panel and QC106. Although the viral loads in QC105 and QC105C fell within the same range, they are different samples, one being used in each assay. Thus, it is not valid to compare the values for the latter samples.
TABLE 2.
HIV-1 RNA levels in samples with different HIV-1 subtypes (QAP panel VLQA 98-2 and QC samples) measured by AMPLICOR HIV-1 MONITOR and Quantiplex bDNA assays
| Assay | HIV-1 RNA load (no. of copies/ml)
|
|||||||
|---|---|---|---|---|---|---|---|---|
| Subtype E-containing samples
|
Subtype B-containing samples
|
|||||||
| VLQA 98-2/1 | VLQA 98-2/2 | VLQA 98-2/4 | VLQA 98-2/5 | VLQA 98-2/6 | QC105 | QC105C | QC106 | |
| AMPLICOR HIV-1 MONITOR (mean ± SD) | 6,977 ± 5,673 | 3,166 ± 1,833 | 1,809 ± 1,543 | 5,703 ± 3,192 | 4,276 ± 2,327 | 4,085 ± 1,835 | Not applicable | 327,151 ± 132,791 |
| Quantiplex bDNA (site 27/site 131) | 14,300/10,510 | 7,800/5,652 | 4,400/2,494 | 9,500/6,450 | 1,200/980 | Not applicable | 2,300/2,596 | 128,900/123,900 |
DISCUSSION
The use of viral load assessments is relatively new in Australia compared to the United States, where clinical trials (American Clinical Trials Group) necessitated the establishment of a QA program at an earlier date (13). Both the AMPLICOR HIV-1 MONITOR and Quantiplex bDNA assays were first registered in Australia in 1996 and, together with the Organon Teknika NucliSens HIV-1 QT, are the only assays currently licensed for diagnostic viral load testing. In contrast, a larger variety of assays is used in the United States, including in-house assays set up within individual laboratories. There is abundant evidence to show that without quality assurance, the performance parameters of such assays are poor (3).
The institution of monitoring of HIV viral load tests in Australia found that approximately half the laboratories performing the AMPLICOR HIV-1 MONITOR v 1.0 assay had intermittent or continuing difficulty achieving assay runs which were valid according to the manufacturer's criteria. Consistently, the reason for invalidity was a failure to achieve a value for the high-positive kit control which was within the specified range. With cooperation between the manufacturer and personnel in several laboratories, very close attention to equipment calibration, and strict adherence to the assay protocol, the proportion of invalid runs by the AMPLICOR HIV-1 MONITOR v 1.0 assay was reduced from 16 to 3% over a period of 6 months across all laboratories. However, although laboratories demonstrated this improved performance, increased invalid run rates continue to occur periodically and can be correlated with the supply of different assay lots. A combination of factors, including minor manufacturing and laboratory variations, probably contribute to an invalid run. Prior to the availability of the NRL's program, results from invalid runs were regularly reported to clinicians in the belief that a single invalid control result did not necessarily invalidate the entire run. This practice was justified on the basis of the nature of the test and its cost rather than on the criterion of accuracy. The NRL's QA program has shown that, when analyzed overall, the numerical results for QC samples do not change significantly in invalid runs (data not shown). However, in multiple instances the high-positive control, provided by the manufacturer, returned values greater than the upper limit of detection and the value for the higher-load NRL QC sample was greater than 2 SDs from the mean. Thus, while values reported from invalid runs may not affect the collective interpretation of a large amount of data, this argument cannot be used to defend reporting of values from invalid runs, as the error in an individual run may be substantial.
Familiarity with the assays and attention to detail have also served to reduce the between-run variability both within and between laboratories, reflected by the reduction in CVs seen between the data generated by the AMPLICOR HIV-1 MONITOR v 1.0 assay for the first set of QC samples (samples QC101 and QC102) and the more recent set of samples (samples QC105 and QC106). In the new version of the AMPLICOR HIV-1 MONITOR assay, v 1.5, results collected to date suggest that there is no difference in the CV from the previous version of the same assay except when Ultrasensitive sample preparation is used. The high CV observed when the modified sample preparation is used may reflect the inexperience of the operators with the new technique but may also highlight a difficulty obtaining precise results for samples with low viral loads by the AMPLICOR HIV-1 MONITOR assay in conjunction with Ultrasensitive sample preparation. As the new version does not differ in practical terms from the previous version, it is not anticipated that the CV will decrease with use of the assay but reflects inherent assay variability. It should be noted that the variability observed with the Quantiplex bDNA 2.0 assay, and now with the Quantiplex bDNA 3.0 assay, has remained consistently less than 35% throughout the program.
Each QAP is designed to ask a question specifically of the quality of HIV viral load testing. In the present study, the questions were, “Can laboratories consistently distinguish a 5-fold difference in viral load?,” “What is the within-run reproducibility of the same sample?,” and “Are assays equally effective for detection of different HIV-1 subtypes?” Only 5 of 19 laboratories, using either assay, produced one result for one of the dilution samples (a different one in each case) which fell outside the range of acceptable variability. The reproducibility of the results obtained with replicate samples demonstrated greater variability, particularly in laboratories using the AMPLICOR HIV-1 MONITOR assay, which may be expected for the PCR technique. Of 17 laboratories, 3 had widely divergent results for replicate samples with high and low viral loads. Only one of these laboratories had problems with reproducibility for samples with both high and low viral loads. The overall results indicate that while variability is important, there are no systematic testing problems with any laboratory in the detection of virus dilutions.
It has previously been recognized that quantification of viral load by the different assays varies. A lower copy number by the Quantiplex bDNA 2.0 assay has consistently been reported (1, 11, 12) and is also supported by the present study. Of greater significance, however, is the fact that the genetic variability of HIV-1 can influence viral load quantification by the various assays. The AMPLICOR HIV-1 MONITOR v 1.0 assay demonstrated poorer sensitivity for some HIV-1 non-B subtypes, with the Quantiplex bDNA 2.0 assay more efficiently detecting subtypes A and E (1, 4, 9, 11). In the present study the values obtained by the AMPLICOR HIV-1 MONITOR v 1.0 assay for subtype E were lower than those obtained for subtype B. The predominance of HIV-1 subtype B in Australia reduced the impact of this finding. However, with the prevalence of subtype E in Southeast Asia (2), the incidence of this subtype in Australia might reasonably be expected to change. The updated version of the AMPLICOR HIV-1 MONITOR assay has been designed to address the deficiency in detection of HIV-1 non-B subtypes seen in v 1.0 by the use of modified primers for nucleic acid amplification. When alternative primers have been used, improved detection of subtype A has been observed (9). Furthermore, in a recent direct comparison, the updated versions of the AMPLICOR HIV-1 MONITOR and Quantiplex bDNA assays demonstrated very close correlations in the number of copies per milliliter, unlike the earlier versions of the same assays (6). Insufficient data have been obtained through the NRL QA program to establish whether improved sensitivity to HIV-1 non-B subtypes has been achieved or whether a better correlation of results is demonstrated by the newer versions of the assays.
In conclusion, the results obtained from the NRL QA program support the recommendation that testing for HIV-1 viral load in an individual patient should always be conducted by the same test and by the same laboratory. The continued variability observed with new versions of the most commonly used assays suggests that this recommendation should remain in place. As long as the same test is consistently used for within-patient measures, the relative values obtained will indicate the efficacy of antiretroviral treatment. The new versions of the viral load assays will probably redress some of the problems associated with the sensitivities of the assays for various HIV-1 subtypes. However highly divergent subtypes may remain undetectable (10). In addition, despite the increased levels of sensitivity claimed by each manufacturer, problems of variability in the assays continue with low viral loads, particularly in the Roche assay.
ACKNOWLEDGMENTS
NRL's HIV viral load QA program was partially funded by a grant from the Diagnostics and Technology Branch of the Department of Health and Aged Care (DHAC) of the Australian Government. NRL is funded by an operational grant from the Population Health Division of DHAC.
We are grateful to A. Dunne and Professor S. Crowe of the Macfarlane Burnet Centre for Medical Research for the generous supply of HIV-1 subtype B supernatants used in the preparation of the QC samples and to S. Walker for assistance with data collection. In addition, we acknowledge the cooperation of the participating laboratories.
REFERENCES
- 1.Berger A, Braner J, Doerr H W, Weber B. Quantification of viral load: clinical relevance for human immunodeficiency virus, hepatitis B virus and hepatitis C virus infection. Intervirology. 1998;41:24–34. doi: 10.1159/000024912. [DOI] [PubMed] [Google Scholar]
- 2.Brown A E, McNeil J G. HIV vaccine development: a subtype E-specific strategy. Southeast Asian J Trop Med Public Health. 1998;29:377–382. [PubMed] [Google Scholar]
- 3.Carpenter C C, Fischl M A, Hammer S M, Hirsch M S, Jacobsen D M, Katzenstein D A, Montaner J S, Richman D D, Saag M S, Schooley R T, Thompson M A, Vella S, Yeni P G, Volberding P A. Antiretroviral therapy for HIV infection in 1998. JAMA. 1998;280:78–86. doi: 10.1001/jama.280.1.78. [DOI] [PubMed] [Google Scholar]
- 4.Dunne A L, Crowe S M. Comparison of branched DNA and reverse transcriptase polymerase chain reaction for quantifying six different HIV-1 subtypes in plasma. AIDS. 1997;11:126–127. [PubMed] [Google Scholar]
- 5.Henrard D R, Daar E, Farzadegan H, Clark S J, Phillips J, Shaw G M, Busch M P. Virologic and immunologic characterization of symptomatic and asymptomatic primary HIV-1 infection. J Acquir Immune Defic Syndr Hum Retrovirol. 1995;9:305–310. [PubMed] [Google Scholar]
- 6.Highbarger H C, Alvord W G, Jlang M K, Shah A S, Metcalf J A, Lane H C, Dewar R L. Comparison of the Quantiplex version 3.0 assay and a sensitized Amplicor Monitor assay for measurement of human immunodeficiency virus type 1 RNA levels in plasma samples. J Clin Microbiol. 1999;37:3612–3614. doi: 10.1128/jcm.37.11.3612-3614.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ho D D, Neumann A U, Perelson A S, Chen W, Leonard J M, Markowitz M. Rapid turnover of plasma virions and CD4 lymphocytes in HIV-1 infection. Nature. 1995;373:123–126. doi: 10.1038/373123a0. [DOI] [PubMed] [Google Scholar]
- 8.Mellors J W, Rinaldo C R, Gupta P, White R M, Todd J A, Kinsley L A. Prognosis in HIV-1 infection predicted by the quantity of virus in plasma. Science. 1996;272:1167–1170. doi: 10.1126/science.272.5265.1167. [DOI] [PubMed] [Google Scholar]
- 9.Nkengasong J N, Kalou M, Maurice C, Bile C, Borget M Y, Koblari S, Boateng E, Sassan-Morokro M, Anatole-Ehounou E, Ghys P, Greenberg A E, Wiktor S Z. Comparison of NucliSens and Amplicor Monitor assays for quantification of human immunodeficiency virus type 1 (HIV-1) RNA in plasma of persons with HIV-1 subtype A infection in Abidjan, Côte d'Ivoire. J Clin Microbiol. 1998;36:2495–2498. doi: 10.1128/jcm.36.9.2495-2498.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Oelrichs R B, Workman C. Beside the limit—how HIV-1 can mask its true viral load. Victorian Infect Dis Bull. 1999;2:33. [Google Scholar]
- 11.Parekh B, Phillips S, Granade T C, Baggs J, Hu D J, Respess R. Impact of HIV type 1 subtype variation on viral RNA quantitation. AIDS Res Hum Retrovir. 1999;15:133–142. doi: 10.1089/088922299311556. [DOI] [PubMed] [Google Scholar]
- 12.Prud'homme I T, Kim J E, Pilon R G, Minkus T, Hawley-Foss N, Cameron W, Rud E W. Amplicor HIV monitor, NASBA HIV-1 RNA QT and Quantiplex HIV RNA version 2.0 viral load assays: a Canadian evaluation. J Clin Virol. 1998;11:189–202. doi: 10.1016/s0928-0197(98)00058-0. [DOI] [PubMed] [Google Scholar]
- 13.Yen-Lieberman B, Brambilla D, Jackson B, Bremner J, Coombs R, Cronin M, Herman S, Katzenstein D, Leung S, Lin H J, Palumbo P, Rasheed S, Todd J, Vahey M, Reichelderfer P. Evaluation of a quality assurance program for quantitation of human immunodeficiency virus type 1 RNA in plasma by the AIDS Clinical Trials Group virology laboratories. J Clin Microbiol. 1996;34:2695–2701. doi: 10.1128/jcm.34.11.2695-2701.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]