Summary
Robinow syndrome (RS) is a genetically heterogeneous disorder with six genes that converge on the WNT/planar cell polarity (PCP) signaling pathway implicated (DVL1, DVL3, FZD2, NXN, ROR2, and WNT5A). RS is characterized by skeletal dysplasia and distinctive facial and physical characteristics. To further explore the genetic heterogeneity, paralog contribution, and phenotypic variability of RS, we investigated a cohort of 22 individuals clinically diagnosed with RS from 18 unrelated families. Pathogenic or likely pathogenic variants in genes associated with RS or RS phenocopies were identified in all 22 individuals, including the first variant to be reported in DVL2. We retrospectively collected medical records of 16 individuals from this cohort and extracted clinical descriptions from 52 previously published cases. We performed Human Phenotype Ontology (HPO) based quantitative phenotypic analyses to dissect allele-specific phenotypic differences. Individuals with FZD2 variants clustered into two groups with demonstrable phenotypic differences between those with missense and truncating alleles. Probands with biallelic NXN variants clustered together with the majority of probands carrying DVL1, DVL2, and DVL3 variants, demonstrating no phenotypic distinction between the NXN-autosomal recessive and dominant forms of RS. While phenotypically similar diseases on the RS differential matched through HPO analysis, clustering using phenotype similarity score placed RS-associated phenotypes in a unique cluster containing WNT5A, FZD2, and ROR2 apart from non-RS-associated paralogs. Through human phenotype analyses of this RS cohort and OMIM clinical synopses of Mendelian disease, this study begins to tease apart specific biologic roles for non-canonical WNT-pathway proteins.
Keywords: genotype-phenotype correlation, skeletal dysplasia, HPO terms, quantitative phenotyping cluster heatmap, molecular diagnosis, traits and OMIM clinical synopsis, screw-tail breed dogs, DVL2, indel mutations
Robinow syndrome (RS) is a genetically heterogeneous disorder caused by WNT/PCP-signaling dysfunction. We identified pathogenic variants in RS individuals (n = 22), including the first DVL2 variant. Quantitative HPO-term-based analyses unveiled gene/allele-specific correlations to disease traits, providing a model for delineating the phenotypic consequences of variants affecting paralogs in the same pathway.
Introduction
Robinow syndrome (RS) is a genetically heterogeneous disorder first described by Robinow et al. in 1969.1,2 RS can be arbitrarily subdivided into autosomal recessive (AR) RS and autosomal dominant (AD) RS. There is significant locus heterogeneity, with pathogenic variants having been identified in six genes: DVL1 (MIM: 601365), DVL3 (MIM: 601368), FZD2 (MIM: 600667), NXN (MIM: 612895), ROR2 (MIM: 602337), and WNT5A (MIM: 164975). All RS-related genes encode proteins that play a role in the non-canonical WNT/planar cell polarity (PCP) signaling pathway.3
Recently, we have shown that RS is characterized by genetic and allelic heterogeneity,3,4 which poses a challenge to studying the disease as it may require use of multiple genomics approaches to investigate the underlying molecular causes in affected individuals. For instance, AR-RS is associated with biallelic loss-of-function (LoF) variants in the orphan tyrosine kinase receptor, ROR2 (RRS1 [MIM: 268310])5,6 and in the oxidative stress response protein, NXN (RRS2 [MIM: 618529]),7,8 both of which can be caused by single-nucleotide variants (SNVs) combined with large deletions.2,4 In contrast, insertions and deletions (indels) in two out of the three human orthologs of the Drosophila dishevelled (dvl) gene, DVL1 and DVL3, are the most frequent cause of sporadic and AD-RS (DRS2 [MIM: 616331], DRS3 [MIM: 616894])3,9, 10, 11 and recur independently in an estimated 33% of individuals with a diagnosis of RS.3 Disease-associated variant alleles found in DVL1 and DVL3 cluster within the penultimate and last exons of each gene, generating a −1 frameshift with a premature termination codon (PTC) in the last exon. All the indel variants are predicted to produce a highly basic, proline-rich C-terminal tail, suggesting those variants create a gain-of-function mutant protein. Notably, no variant affecting the DVL2 paralog has been observed in RS cohorts thus far, even though all three paralogs were shown to have overlapping function during development and share about 59%–67% of amino acid similarity.12, 13, 14 Screw-tail dog breeds (boxers and others) are homozygous for a 1-bp deletion in Dvl2 and the dogs' phenotype is similar to RS, further suggesting that, like DVL1 and DVL3, DVL2 could be yet another RS gene.15 Missense and truncating variants in FZD2, which encodes a highly conserved seven-pass transmembrane protein of the Frizzled family of membrane receptors, cause ∼14% of RS.3 Lastly, heterozygous missense variants, potentially leading to hypomorphic alleles of the extracellular soluble ligand of ROR2, WNT5A,3,16, 17, 18 are estimated to cause ∼10% of dominant RS (DRS1 [MIM: 180700]).3,16, 17, 18 A molecular understanding of RS is obfuscated by genetic heterogeneity, and phenotypic overlap between RS and other disorders, notably, Aarskog-Scott syndrome (AAS [MIM: 305400]).3,19
Clinically, RS is characterized by prominent forehead, hypertelorism, broad nasal bridge, anteverted nares, and midface hypoplasia, common features reported in both AR-RS and AD-RS.20,21 However, some phenotypic features are variable and differences in the frequency of manifestation might be considered as penetrance of the endophenotype. Hemivertebrae and scoliosis were reported in AR-RS, while umbilical hernias are almost exclusively observed in AD-RS.20 Thus far, there has been no comprehensive analysis of the genotype-phenotype correlations of RS caused by specific variants in all genes currently associated with RS; i.e., investigating allelic phenotypes of pathogenic variants, particularly for NXN and genes causative of AD-RS that were recently discovered.
Here, we identified RS DNA variants in 22 unpublished individuals with RS/RS-like phenotypes with a combination of gene-targeted Sanger sequencing, exome sequencing (ES), and genome sequencing (GS) and review the molecular and clinical published data on all previously published individuals carrying variants affecting NXN and AD-RS-associated genes (N = 68 individuals). Notably, we describe the first individual with RS with a likely pathogenic indel in DVL2, a gene that has not previously been associated with RS or any other human phenotype. We established genotype-phenotype correlations for all AD-RS genes, which revealed core as well as gene-specific clinical traits. Using Human Phenotype Ontology (HPO) terms, we performed a quantitative phenotypic similarity analysis of the entire cohort and for Online Mendelian Inheritance in Man (OMIM, https://omim.org/) diseases. Gene- and allele-specific phenotypic differences within RS were revealed. In addition, most of the differential diagnoses considered for RS share a high degree of phenotypic similarities with RS, and they together can be quantitatively distinguished from diseases caused by WNT, FZD, or ROR paralogs not associated with RS. These data support the hypothesis that the skeletal features in RS are caused by perturbation of the non-canonical WNT/PCP pathway rendered by alterations of specific paralogs during development.
Subjects and methods
Subjects
The RS cohort reported here consists of two groups: (1) a phenotype-driven cohort in which recruited individuals had a clinical diagnosis of RS; (2) a genotype-driven cohort for which individuals were referred for clinical laboratory diagnostic testing at the Baylor Genetics (BG) or GeneDx and were found to have likely pathogenic or pathogenic variants in AD-RS genes (NXN was not investigated in those diagnostic laboratories at the time of ascertainment) (Figure S1). After ascertainment of probands to the cohort using a genotype-driven approach, it was found that all probands were clinically diagnosed as having RS before molecular diagnosis, except individual BAB14592. This study focuses on AD-RS genes and the recently identified AR-RS gene NXN; individuals carrying ROR2 variants were not included in this study. In total, we collected 22 subjects from 18 unrelated families. RS was sporadic in the two cohorts except for three families (HOU5112, HOU4020, and HOU4115) with an affected father or an affected mother and grandfather. DNA was obtained after all relevant family members provided written, informed consent. The study was approved by the institutional review board (IRB) at BCM (IRB protocols nos. H-43246 and H-29697). BAB14964 was identified by the UK's 100,000 Genomes Project.22
Sanger sequencing
Samples from subjects clinically diagnosed as RS were investigated via Sanger sequencing of the penultimate and last exons of DVL1 and DVL3, in addition to the entire single-exon gene FZD2. Primers used are (1) DVL1 exon 14–15 forward (Fwd), GACACAGGTGCTGTCAGGAG; (2) DVL1 exon 14–15 reverse (Rev), GCACACGGTCAACAAGATCA; (3) DVL3 exon 14–15 Fwd, ACCACGGTCTCTCTCATCCA; (4) DVL3 exon 14–15 Rev, AAGACGGACGGATGGAGAGA; (5) FZD2 screen L, GCCGCGAGTAAAGTTTGCAA; (6) FZD2 screen R, ACAAACCTTGTGGGGGTGTT; (7) DVL2 exon15 F, TGGGCTACATAACATCCACAA; (8) DVL2 exon15 R, CCAGGGCTCCATCCCTAT. Primers (7) and (8) were used to confirm the variant found in BAB14964 via GS. The PCR reactions were performed using TaKaRa LA Taq DNA Polymerase (Takara Bio) with the protocol as follows: 98°C for 30 s (94°C for 60 s, 65°C for 20 s, 68°C for 20 min) repeated 32 times, 68°C for 10 min. PCR products were then sequenced by standard Sanger dideoxy capillary sequencing. All PCR products containing candidate indel alleles detected by Sanger sequencing were further confirmed by cloning both alleles into a TOPO TA cloning vector (Life Technologies). To determine specific allele sequences, the recombinant clones were transformed into chemically competent Escherichia coli strain One Shot TOP10 (Invitrogen). Single-colony purified clones were isolated and grown overnight in selective medium. Ten individual clonal colonies were then Sanger dideoxy capillary sequenced. Pedigrees and chromatograms of all variants are shown in Figure S2.
Variant detection
DNA samples from individuals lacking pathogenic variants in either DVL1, DVL3, or FZD2 and their unaffected relatives were submitted for ES through the Baylor-Hopkins Center for Mendelian Genomics initiative.23 ES was performed at the BCM Human Genome Sequencing Center (HGSC) using 0.5 μg of genomic DNA. An Illumina paired-end, pre-captured library was constructed according to the manufacturer's protocol with modifications described in the BCM-HGSC.23 Six to 10 pre-captured libraries were pooled and then hybridized in solution to the HGSC in-house-developed VCRome 2.1 design with custom spike-in24 according to the manufacturer's protocol (NimbleGen SeqCap EZ Exome Library SR User's Guide) with minor revisions. Illumina sequencing was performed, with a sequencing yield average of 11 Gb; 97.5% of targeted bases were covered to a depth of 20× or greater with an average depth of coverage of 118.6×. In parallel to the exome workflow, a single-nucleotide polymorphism (SNP) array was performed for a final quality assessment. This included orthogonal confirmation of sample identity and purity using the Error Rate In Sequencing (ERIS) pipeline developed at the HGSC. Using an e-GenoTyping approach, ERIS screens all sequence reads for exact matches to probe sequences defined by the variant and position of interest. A successfully sequenced sample must meet quality control metrics of ERIS SNP array concordance (>90%) and ERIS average contamination rate (<5%). To prevent bias in the filtering and parsing of rare variants caused by different annotation software, multiple annotation software tools that often provide distinct interpretations and varying levels of false-positive and false-negative findings were used. This has been shown to improve variant annotation, particularly for indels.25
Thus, two variant discovery methods were used in parallel, starting with the HGSC Mercury analysis pipeline.26,27 The Mercury pipeline moves data from initial sequence generation on the instrument to annotated variant call files (VCFs) via multiple analysis tools, including xAtlas, for variant calling.28 In addition, we used the Genome Analysis Toolkit (GATK) HaplotypeCaller to produce joint-called files with indel realignment and base recalibration for all families whose DNA was sequenced. We identified de novo mutations in silico by using read-depth information extracted from the BAM files of either parent or proband using the in-house developed software DNM-Finder.29 Candidate variants were filtered against exome data in publicly available databases, including the Genome Aggregation Database (gnomAD),30 the National Heart, Lung, and Blood Institute (NHLBI) Exome Sequencing Project (ESP) Exome Variant Server, the Atherosclerosis Risk in Communities Study (ARIC) database,31 and our internal Baylor-Hopkins Centers for Mendelian Genomics variant analyzer database of approximately 12,000 exomes. In parallel, we also took results from webtools that predict functional effects of candidate variants, such as Polymorphism Phenotyping v2 (PolyPhen-2),32 Sorting Intolerant From Tolerant (SIFT),33 and Combined Annotation Dependent Depletion (CADD).34 In addition, all affected individuals' exomes were screened for copy-number variants (CNVs) using XHMM35 and our in-house-developed algorithm for detecting homozygous exonic deletions, HMZDelFinder.36 ES data were also used to calculate B-allele frequency and delineate genomic intervals exhibiting absence of heterozygosity (AOH) consistent with identity-by-descent.37 Segregation studies were performed using standard PCR amplification of ES identified variants, and PCR products were purified with ExoSAP-IT (Affymetrix) and sequenced by dideoxy nucleotide Sanger sequencing. Pedigrees and chromatograms are shown (Figure S2).
ES of BAB14592 was performed at the BG diagnostic laboratory,38 whereas ES of 8310037 was performed at GeneDx using a previously reported protocol.39 For 8310037, genomic DNA from the proband and parents was used; the exonic regions and flanking splice junctions of the genome were captured using the IDT xGen Exome Research Panel v1.0. Massively parallel (NextGen) sequencing was done on an Illumina system with 100-bp or greater paired-end reads. Reads were aligned to human genome build GRCh37/UCSC hg19 and analyzed for sequence variants using a custom-developed analysis tool. Additional sequencing technology and variant interpretation protocol have been described previously.39 The general assertion criteria for variant classification are publicly available on the GeneDx ClinVar submission page (http://www.ncbi.nlm.nih.gov/clinvar/submitters/26957/). For IFH, the variant was identified by a variety of testing methods and confirmed by Sanger at GeneDx.
GS of BAB14964 was performed by the UK's 100,000 Genomes Project (The National Genomics Research and Healthcare Knowledgebase v7, Genomics England. https://doi.org/10.6084/m9.figshare.4530893.v7. 2020).
Quantitative phenotypic analyses based on HPO terms
To quantify genotype-phenotype correlations within RS and across diseases included in the RS differential diagnosis, we retrospectively collected and re-analyzed the clinical phenotypes of 68 individuals (30 males and 38 females) carrying pathogenic variants in DVL1, DVL2, DVL3, FZD2, WNT5A, and NXN. This cohort consists of the 16 individuals described in this current study in addition to 52 individuals described previously in the literature. In total 23 DVL1, one DVL2, nine DVL3, 17 FZD2, 14 WNT5A, and four NXN individual clinical phenotypes were included (probands and affected relatives for whom phenotype was available) (Tables S1 and S2). Briefly, variant allele-associated phenotypes were annotated with HPO terms for each affected individual. All diseases (n = 7,977, including number symbol, plus sign, percentage sign, and no symbol in OMIM) and genes (n = 4,102, asterisk symbol in OMIM) that have been annotated with HPO terms by OMIM were downloaded from the HPO resource page (https://hpo.jax.org/app/download/annotation). Distribution plot of the number of HPO terms per proband was used to assess depth of phenotyping in this cohort. Individual similarity matrices were generated with the OntologyX suite of R packages using the Lin's semantic similarity score and the average method.40,41 Similarity matrices were then used to generate distance matrices of individual similarity. Hierarchical agglomerative clustering (HAC) was performed on distance matrices with the Ward's method42 with the number of clusters set based on visualization of the gap statistic curve. Individual similarity scores were visualized using the ComplexHeatmap package in R, and statistical analysis of individual groups was done using the OntologyX suite. Annotation grids were generated with the OntologyX suite of packages and then edited to exclude ancestral terms and to order columns by phenotype frequency.
To assess the phenotypic landscape of differential diagnoses of RS and compare it with WNT-pathway-associated diseases, a similar disease-to-gene and disease-to-disease HPO analysis was performed. Thirty-one Mendelian disorders (Table S3), including RS caused by perturbation of the WNT genes (DVL1, DVL3, FZD2, WNT5A, ROR2, and NXN), non-RS diseases related to WNT paralogs, and syndromes on the differential diagnosis for RS were identified to use in HPO analysis. These 31 diseases were separately assessed for phenotypic similarity to all (1) genes and (2) diseases with OMIM HPO annotation. HPO-annotated phenotypes for the 31 diseases were queried against all disease-associated genes (n = 4,102) or all diseases (n = 7,946) annotated with HPO terms by OMIM for phenotypic similarity. Lin semantic similarity scores between all pairs of the 31 diseases and all genes or diseases annotated with HPO terms were calculated. The top 10 phenotypically similar gene-associated or disease HPO term sets to each disease in the group of 31 diseases described above were parsed and duplicates removed. Every combination of two that includes one member from the group of 31 diseases and one from the top phenotypically similar gene-associated phenotype matches was taken, and p value calculated via comparison of the phenotypic similarity score between that group of two and 100,000 randomly selected groups of two from all OMIM HPO-annotated genes or diseases, respectively (p value cutoff <0.001). The gap statistic was calculated for cluster number k = 1 to 20 (gene analysis) or 15 (disease analysis), and the resultant curve was visualized to select optimal number of clusters to use. HAC analysis and visualization of phenotypic similarity and clustering was then performed as described above for RS proband phenotypes.
The same approach was used to perform an unbiased query for all HPO-annotated diseases (n = 7,971) to assess which clinically described diseases have significant (p < 0.003) similarity overlap with six HPO-annotated RS phenotypes associated with six non-canonical WNT/PCP pathway genes (DVL1, DVL3, WNT5A, FZD2, NXN, ROR2). A higher p value cutoff was used as p < 0.001 was too stringent for this analysis. p values here are used to define which set of diseases or genes in addition to the query set to include in HAC and heatmap visualization to further classify disease. This analysis was also done against HPO-annotated genes (data not shown). Lastly, the phenotypes of FZD2-related omodysplasia 2 (MIM: 164745), related mesomelic/rhizomelic dysplasias,43 and 16 FZD2 patients were compared with 7,963 OMIM disease phenotypes for similarity (p < 0.001). To help determine whether FZD2 proband phenotypes were more similar to RS than omodysplasias, a distribution of all pairwise scores between diseases and probands was visualized and average similarity score of FZD2 probands to both AD and AR omodysplasia and RS was performed (Figure S3).
Statistical analysis of gene-associated phenotypes
For each phenotype, the individuals were categorized into five genotype groups (Table S2). Fisher's exact test was done on each phenotype (considering all genotypes) by utilizing statistic package in R language. For every genotype, individuals were dichotomously categorized into either “have” or “not have” regarding pathogenic variants in the given RS-associated gene. If a particular phenotype is not correlated to a specific genotype, we should expect a similar prevalence of “have” and “not have” individuals distributed among all genotypes. The p values were adjusted using the p.adjust R function and Benjamini-Hochberg method (method = fdr). The significance threshold was set to 0.05. Patient prevalence of each genotype/phenotype was visualized by using the seaborn package in python language.
Results
Mutational patterns in AD-RS genes
Targeted gene sequencing (N = 7), ES (N = 14), and GS (N = 1) were performed to identify pathogenic variants that may cause RS in our cohort. As a result, pathogenic or likely pathogenic variants in genes/candidate genes associated with RS or Mendelian disorders with phenotypic overlap with RS were found in all 22 individuals (Figures S1 and S2). As shown in Table 1, four individuals (4 out of 22, 18%) have pathogenic or likely pathogenic variants in DVL1, including two de novo recurrent 1-bp deletion alleles and two novel variants for which inheritance could not be established. Notably, BAB13797 was found to have a previously unreported heterozygous splicing variant in DVL1, which is the first splicing variant reported in this gene thus far. Two individuals (2 out of 22, 9%) were found to have novel variants in DVL3. Five individuals (5 out of 22, 23%), including two from the same family, were found to have pathogenic or likely pathogenic variants in FZD2, including four recurrent missense and truncating variants, as well as one novel frameshift variant. Four individuals (4 out of 22, 18%), two of them from the same family, were found to have novel variants in WNT5A. Notably, one individual (1 out of 22, 5%) was found to have a de novo 1-bp duplication in exon 15 of DVL2 (MIM: 602151), which is the first RS-associated variant identified in DVL2.
Table 1.
Summary of variants detected in RS-related genes in this study
| Individual ID | Origin | Gene | Variant type | Inheritance | Zygosity | Variant | Effect | ACMG classification | ACMG criteria |
|---|---|---|---|---|---|---|---|---|---|
| BAB10168 | in-house cohort | DVL1 | −1 fs | de novo | Het | c.1519delT | p.Trp507Glyfs∗142 | pathogenic | PVS1, PS1, PS2, PM2, PP4 |
| BAB13793 | in-house cohort | DVL1 | −1 fs | unknown | Het | c.1592delC | p.Pro531Argfs∗118 | likely pathogenic | PVS1, PM2 |
| BAB9945 | in-house cohort | DVL1 | −1 fs | de novo | Het | c.1519delT | p.Trp507Glyfs∗142 | pathogenic | PVS1, PS1, PM2, PM6, PP4 |
| BAB13797 | in-house cohort | DVL1 | splicing | unknown | Het | c.1640-1C>T | NA | likely pathogenic | PVS1, PM2 |
| BAB14964 | 100,000 Genomes Project | DVL2 | +1 fs | de novo | Het | c.2105dupC | p.Pro703Serfs∗103 | likely pathogenic | PS2, PM2, PM4 |
| 8310037 | GeneDx | DVL3 | −1 fs | de novo | Het | c.1751_1754delATCG | p.Asp584Glyfs∗83 | pathogenic | PVS1, PS2, PM2, PP4 |
| IFH | GeneDx | DVL3 | splicing | unknown | Het | c.1715-2delA | NA | pathogenic | PVS1, PS1, PM2 |
| BAB10686 | in-house cohort | FZD2 | missense | unknown | Het | c.1300G>A | p.Gly434Ser | pathogenic | PS1, PM2, PM5, PP3, PP4 |
| BAB11377 | in-house cohort | FZD2 | nonsense | unknown | Het | c.1644G>A | p.Trp548∗ | pathogenic | PVS1, PS1, PM2, PP4 |
| BAB13798 | in-house cohort | FZD2 | nonsense | inherited | Het | c.1644G>A | p.Trp548∗ | pathogenic | PVS1, PS1, PM2, PP1, PP4 |
| Father of BAB13798 | in-house cohort | FZD2 | nonsense | unknown | Het | c.1644G>A | p.Trp548∗ | pathogenic | PVS1, PS1, PM2, PP4 |
| BAB14592 | BG | FZD2 | Fs | unknown | Het | c.367_388dup GCCCTCATGA ACAAGTTCGGTT |
p.Phe130Cysfs∗98 | likely pathogenic | PVS1, PM2 |
| BAB7986 | in-house cohort | WNT5A | missense | unknown | Het | c.248G>A | p.Cys83Tyr | likely pathogenic | PM2, PM5, PP3, PP4 |
| BAB11664 | in-house cohort | WNT5A | missense | inherited | Het | c.461G>T | p.Cys154Phe | uncertain significance | PM2, PP3, PP4 |
| BAB10972 Father of BAB11664 | in-house cohort | WNT5A | missense | unknown | Het | c.461G>T | p.Cys154Phe | uncertain significance | PM2, PP3, PP4 |
| BAB14600 | GeneDx | WNT5A | missense | de novo | Het | c.247T>G | p.Cys83Gly | likely pathogenic | PM2, PM5, PM6, PP3, PP4 |
Het, heterozygous; fs, frameshift; BG, Baylor Genetics Clinical Diagnostics Laboratory.
We found sixteen (16 out of 22, 73%) of the 22 studied subjects had pathogenic or likely pathogenic variants in 4 RS-related genes (DVL1, DVL3, FZD2, and WNT5A) or a candidate gene (DVL2), including nine novel (10 individuals) and three recurrent variants (six individuals). As shown in Figure 1, altogether, 18 unique variants from 24 unrelated subjects in DVL1,3,4,9, 11,44,45 10 variants in 10 DVL3-unrelated individuals,3,11,46,47 seven variants in 14 unrelated FZD2 individuals,3,48, 49, 50, 51 and 11 variants in WNT5A from 12 unrelated subjects3,16, 17, 18,52 were identified (Figure 1, Table S4).
Figure 1.
Map location of identified variants in RS-related genes resulting in AD-RS
Previously described variants (blue circles) and variants identified in this study (red circles) are shown. Larger circles represent identical variants in unrelated individuals; number of recurrent variants is proportional to size of circle. Chromosome and cytogenetic interval location of the canonical transcripts are provided: individual exons (black rectangles) on a horizontal line are drawn to represent gene structure. (A) Variants within the coding region of DVL1 resulting in AD-RS: variants are mostly small insertions or deletions, except for one splicing variant; all of them are predicted to lead to −1 frameshifting of the reading frame. Zoomed-in view shows the bases affected by variants in exon 14 (i.e., penultimate exon). PTC created by −1 frameshifting variants is displayed by an orange diamond. (B) Variant within the coding region of DVL2 resulting in AD-RS: BAB14964 was found to have a de novo 1-bp duplication at the position c.2105dupC; p.Pro703Serfs∗103. (C) Variants within the coding region of DVL3 resulting in AD-RS: the variants in DVL3 are mostly small insertions or deletions, except for four splicing variants. All mutations, analogous to DVL1, are predicted to lead to −1 frameshifting of the reading frame. PTC created by −1 frameshifting variants is displayed by an orange diamond. Zoomed-in view shows the bases affected by variants in exon 14 and 15. (D) Predicted protein translational effects of FZD2 variants resulting in AD-RS: pathogenic variants in FZD2 are missense or truncating variants affecting mostly the Frizzled domain. (E) Predicted protein translational effects of WNT5A variants resulting in AD-RS: pathogenic or likely pathogenic variants in WNT5A are mostly missense, except for two indels. The majority of WNT5A variants involve substitution or creation of a cysteine residue.
All variants in DVL1 and DVL3 are small indels or splicing variants (Figures 1A and 1C). In silico prediction of splicing variants by the Human Splicing Finder web tool53 (https://www.genomnis.com/access-hsf) indicates that cryptic splice acceptor sites downstream of the canonical site may be activated, resulting in the use of the −1 frame, which is consistent with other observed indels. Danyel et al. (2018) isolated RNA from blood and lymphoblastoid cell line samples of an individual who has a splicing variant at the position c.1715-2A>C in DVL3, and confirmed expression of both WT and expected −1 frameshifting mutant mRNAs.47 In summary, monoallelic −1 frameshift variants clustered within the penultimate and last exon of DVL1 or DVL3 cause AD-RS.
Recurrent monoallelic missense or truncating variants constitute the majority of the FZD2 variants associated with dominant RS (Figure 1D). Notably, six unrelated individuals have been found to have variants in FZD2 that alter the same amino acid Gly434 to serine or valine. The C to T transition rate at methylated CpG sites is ∼10-fold higher than those at unmethylated sites,54, 55, 56, 57 which may contribute to the high recurrence (two of the variants are C to T transitions). The high frequency of pathogenic variants at the G434 position may result in steric hindrance, altering the affinity or stability of the interaction between FZD2 and DVL158 as this conserved region is one of the motifs required for binding and stabilizing the interaction with DVL1.59 In addition, seven individuals present with truncating variants at p.Trp548∗ or p.Trp547∗. FZD2 is a single-exon gene that is likely not subject to nonsense-mediated mRNA decay (NMD), thus this truncating allele may produce a stable mRNA product but potentially prevent translation of a discontinuous motif across three intracellular loops in Frizzled protein that modulate the Fzd2–Dvl1 interaction (Figure 1D).48,59
Variants in WNT5A are mostly missense, except for two indels (one is an in-frame duplication and another is a homozygous frameshifting variant, with carrier parents unaffected), all localized to exon 3 and exon 4 of the gene (Figure 1E). A striking feature of WNT5A is that a majority (13 out of 14, 93%) of variants causing RS replace or introduce a cysteine, except for the biallelic frameshifting variant (p.Tyr188Metfs∗31).52 WNT5A is a cysteine-rich protein; there are 24 conserved cysteine residues across the entire protein. Disulfide bond formation is critical for tertiary structure, folding, and function of Wnt proteins.60 It is notable that four unrelated individuals were found to have variants altering the second cysteine of WNT5A, Cys83, to one of four different amino acids: glycine, serine, tryptophan, and tyrosine. A systematic analysis of cysteines and associated disulfide bonds in the mouse Wnt3a showed that mutation of any cysteine of Wnt3a leads to perturbation of disulfide bond formation, which either affects secretion/activity or leads to altered binding affinity with FZD, causing diminution of Wnt signaling.61 Based on these findings, we speculate that the removal or addition of cysteines disrupts disulfide bond formation in WNT5A, therefore altering the tertiary protein structure and affecting the function of WNT5A.
Biallelic LoF variants in NXN have been found to cause a new type of AR-RS.3,4 Notably, two related individuals were found to have compound heterozygous variants consisting of a maternally inherited 3-bp deletion and a paternally inherited intragenic 84-kb deletion that encompasses the entire first exon. This intragenic deletion was putatively formed by Alu-Alu-mediated rearrangement (AAMR). To evaluate the potential genomic instability AAMR risk for RS-associated genes, we used the AluAluCNVpredictor tool (http://alualucnvpredictor.research.bcm.edu:3838)62; the AAMR relative risk score for NXN is above the 0.6 cutoff for AAMR risk based on genome-wide Alu-Alu density and predicted relative risk susceptibility to genomic instability (MIM genes, 0.866; RefSeq genes, 0.843). This indicates that there is a higher probability of AAMR-mediated exonic deletion CNV formation at NXN compared with most other RefSeq genes.
Differential diagnosis of RS
We identified six individuals (6 out of 22, 27%) with pathogenic or likely pathogenic variants in other disease-associated genes (Table 2, Figures S1 and S2). Four individuals, three from the same family, were found to have novel hemizygous or heterozygous 1-bp deletions in FGD1. Missense and indel variants in FGD1 are associated with X-linked AAS,19 which has phenotypic overlap with AD-RS. Another individual, BAB9133, was found to have a variant of uncertain significance (VUS) in ARID1B (MIM: 614556) and a pathogenic copy-number loss spanning 17.5 kb of the gene TCF4 (MIM: 602272) (Figure S4), which is associated with Pitt-Hopkins syndrome (MIM: 610954).63 In BAB11295, a heterozygous de novo missense variant was identified in PPP1CB (MIM: 600590). This pathogenic variant has been reported to cause Noonan syndrome-like disorder with loose anagen hair 2 (MIM: 617506).64, 65, 66
Table 2.
Summary of variants detected in individuals with differential diagnoses of RS in our in-house cohort
| Individual ID | Origin | Gene | Variant type | Inheritance | Zygosity | Transcript | Variant | Effect | OMIM phenotype |
|---|---|---|---|---|---|---|---|---|---|
| BAB10066 | this report | FGD1 | Fs | unknown | Hemi | NM_004463.2 | c.1422delT | p.Phe474Leufs∗34 | Aarskog-Scott syndrome, 305400 |
|
BAB11334 Affected maternal grandfather of BAB11808 |
this report | FGD1 | Fs | unknown | Hemi | NM_004463.2 | c.367delC | p.Leu123∗ | Aarskog-Scott syndrome, 305400 |
|
BAB11333 Affected mother of BAB11808 |
this report | FGD1 | Fs | inherited | Het | NM_004463.2 | c.367delC | p.Leu123∗ | Aarskog-Scott syndrome, 305400 |
| BAB11808 | this report | FGD1 | Fs | inherited | Hemi | NM_004463.2 | c.367delC | p.Leu123∗ | Aarskog-Scott syndrome, 305400 |
| BAB8747 | White et al.3 2018 | FGD1 | Fs | de novo | Hemi | NM_004463.2 | c.892dupT | p.Cys298Leufs∗5 | Aarskog-Scott syndrome, 305400 |
| BAB8751 | White et al.3 2018 | FGD1 | Fs | de novo | Hemi | NM_004463.2 | c.527dupC | p.Leu177Thrfs∗40 | Aarskog-Scott syndrome, 305400 |
| BAB11295 | this report | PPP1CB | missense | de novo | Het | NM_206876.1 | c.146C>G | p.Pro49Arg | Noonan syndrome-like disorder with loose anagen hair 2, 617506 |
| BAB8743 | White et al.3 2018 | PTPN11 | missense | de novo | Het | NM_002834.4 | c.836A>G | p.Tyr279Cys | Noonan syndrome 1, 163950 |
| BAB8740 | White et al.3 2018 | RAC3 | missense | de novo | Het | NM_005052.2 | c.176C>G | p.Ala59Gly | neurodevelopmental disorder with structural brain anomalies and dysmorphic facies, 618577 |
| BAB8759 | White et al.3 2018 | SH3PXD2B | Fs | inherited | Hom | NM_001017995.2 | c.969delG | p.Arg324Glyfs∗19 | Frank-Ter Haar syndrome, 249420 |
| INPPL1 | missense | inherited | Hom | NM_001567.3 | c.1636G>A | p.Val546Ile | opsismodysplasia, 258480 | ||
| BAB8836 | White et al.3 2018 | – | translocation | de novo | Het/Hemi | – | 46,XY,der(X)t(X; 6)(p22.31;q25.3).arr[GRCh37] 6q25.3q27(157,870,814_170,881,475)x3, Xp22.33p22.31 (409,876_8,199,541)x0 | – | – |
| BAB9133 | this report | ARID1B | missense | unknown | Het | NM_017519.2 | c.2374T > C# | p.Tyr792His | Coffin-Siris syndrome 1, 135900 |
| TCF4 | deletion | unknown | Het | – | seq[GRCh37]del(18) (q21.2) chr18:g.52,940,259-52,957,780 | – | Pitt-Hopkins syndrome, 610954 |
Fs, frameshift; Hemi, hemizygous; Het, heterozygous; Hom, homozygous. #This variant is a VUS.
Clinical phenotyping in subjects with RS-associated variant alleles
A comprehensive genotype-phenotype table of both our in-house and published RS individuals was established by combining the individual patient phenotype information with the sequencing results (Table S2). The only subject carrying DVL2 variant was not included in this phenotyping analysis.
Based on the data collected in this study (Figure 2, Table S2), there are clinically recognizable phenotypic features that are present at the same rate in RS regardless of genotype. Some of the more common physical features among all genotypes (prevalence more than 60% in each genotype), potentially comprising a clinical synopsis, include midface hypoplasia (90%, N = 54 out of 60); hypertelorism (79%, N = 50 out of 63); prominent eyes (65%, N = 35 out of 54); anteverted nares (88%, N = 49 out of 56); wide, low nasal bridge (90%, N = 55 out of 61); gingival hyperplasia (81%, N = 42 out of 52); limb length anomaly, including mesomelia/micromelia/rhizomelia (91%, N = 60 out of 66); brachydactyly (89%, N = 55 out of 62); and broad first toe (67%, N = 33 out of 49). Additionally, hypoplastic genitalia in males (93%, N = 25 out of 27) was common among all RS subjects, irrespective of the genetic etiology, but prevalence of hypoplastic genitalia in females was variable in different genotypes.
Figure 2.
Phenotypic analysis of subjects with pathogenic variants in DVL1, DVL3, FZD2, WNT5A, and NXN
Prevalence (0–1.0) of phenotypes in subjects with pathogenic variants in DVL1, DVL3, FZD2, WNT5A, and NXN shown in Table S2 is displayed by heatmap. Within the heatmap, black indicates a higher prevalence, while light peach color indicates lower prevalence; light gray indicates these specific data are not available. The phenotypes are listed in order of overall decreasing prevalence. A key is provided on the right.
DVL1-associated RS probands were significantly more likely to have macrocephaly (95%, N = 21 out of 22), triangular mouth (82%, N = 14 out of 17), and increased bone density (82%, N = 9 out of 11). In addition, clinical findings suggest lower prevalence of upslanting palpebral fissures (26%, N = 5 out of 19) and short stature (38%, N = 8 out of 21), whereas an increased prevalence of high forehead (95%, N = 20 out of 21), bilobed tongue (80%, N = 16 out of 20), and umbilical hernia (59%, N = 10 out of 17) were associated with DVL1 variant alleles, although these observed trends do not reach statistical significance (Figure 2, Table S2). These findings further support the recently published association of cranial osteosclerosis in DVL1-associated RS.67 These data are not available for the majority of other genotypes. Exceptions include one or two individuals with other genotypes who were found to have normal bone density and, in a study by Shayota et al. (2020), an individual with WNT5A-associated RS and a second with ROR2-associated RS were found to have mild osteopenia of the non-cranial skeleton, or normal bone density measurements.67
Congenital heart defects were more common in RS subjects who have pathogenic variants in DVL3 (78%, N = 7 out of 9) (Figure 2, Table S2). Among the congenital heart defects observed across individuals with RS, atrial septal defect (ASD) and ventricular septal defect (VSD) were most common, while individuals carrying DVL3 pathogenic variants tended to have more complex defects, including hypoplastic right heart syndrome, hypoplastic left heart syndrome, pulmonary atresia, and aberrant subclavian artery. In addition, trends of increased prevalence of omphalocele (43%, N = 3 out of 7) and cleft lip and/or palate (67%, N = 6 out of 9) were also observed, as were decreased prevalence of abnormal ear shape and position (63%, N = 5 out of 8), female genital hypoplasia (25%, N = 1 out of 4), and hypoplastic phalanges (50%, N = 3 out of 6) among DVL3-associated RS.
Individuals with RS caused by variants in FZD2 seem to be more mildly affected with significant lower prevalence of dental anomalies (58%, N = 7 out of 12) and hypoplastic phalanges (0%, N = 0 out of 3), as well as trends of decreased prevalence of hypertelorism (60%, N = 9 out of 15); wide, low nasal bridge (76%, N = 13 out of 17); brachydactyly (67%, N = 10 out of 15); clinodactyly (57%, N = 8 out of 14); broad thumb (25%, N = 3 out of 12); nail dysplasia (0%, N = 0 out of 8); bifid first and/or second phalanges (0%, N = 0 out of 3); hypoplastic phalanges (0%, N = 0 out of 3); and male genital hypoplasia (67%, N = 4 out of 6) compared with individuals with other genotypes (Figure 2, Table S2). The only exception is a trend of higher prevalence of long philtrum (92%, N = 12 out of 13). Interestingly, four females with FZD2-related RS, of 11 females studied, were found to have anomalies of the uterus: BAB7987 and BAB7988 have retroverted uterus, while BAB11377 and patient 1 reported by Warren et al.51 were found to have a bicornuate uterus.
For WNT5A-associated RS, a trend of decreased prevalence of abnormal ear shape and position (50%, N = 5 out of 10) was observed. In addition to core phenotypes, all individuals were observed to have dental anomalies (100%, N = 7 out of 7) and genital hypoplasia (100%, N = 5 out of 5 for male; N = 3 out of 3 for female) (Figure 2, Table S2). However, no individuals were found to have camptodactyly (0%, N = 0 out of 5), renal anomalies (0%, N = 0 out of 6), hearing loss (0%, N = 0 out of 5), or omphalocele (N = 0 out of 5). The prevalence of these phenotypic features is variable in other genotypes.
NXN is a novel AR-RS-related gene, with only four individuals identified, including two from the same family.3,4 Based on the phenotypic results (Figure 2, Table S2), in addition to core phenotypes, all NXN individuals have short stature (100%, N = 2 out of 2), high forehead (100%, N = 4 out of 4), long philtrum (100%, N = 4 out of 4), triangular mouth (100%, N = 4 out of 4), dental anomalies (100%, N = 2 out of 2), abnormal ear shape and position (100%, N = 2 out of 2), clinodactyly (100%, N = 4 out of 4), nail dysplasia (100%, N = 2 out of 2), hypoplastic phalanges (100%, N = 2 out of 2), and omphalocele (100%, N = 2 out of 2). NXN individuals were not found to have upslanting palpebral fissures (0%, N = 0 out of 3), bifid first and/or second phalanges (0%, N = 0 out of 2), scoliosis and/or kyphosis (0%, N = 0 out of 2), or pectus anomaly (0%, N = 0 out of 2). These phenotypic features are variably present in RS associated with other genes. Notably, the prevalence of short nose is greater than 90% in other genotypes, but only 50% in NXN individuals (N = 2 out of 4).
Quantitative assessment of RS clinical phenotypes
To quantify and visualize genotype-phenotype correlations, semantic similarity scores were calculated using an HPO-based analysis (Figure S5). The DVL2 subject was included in this analysis. The number of HPO terms per proband is shown in Figure S6. Phenotypic similarity scores between each RS proband and/or the annotated diseases were calculated and visualized in a cluster heatmap where number of clusters was determined by visualization of the gap statistic curve (Figures 3A, 3B, and S7).
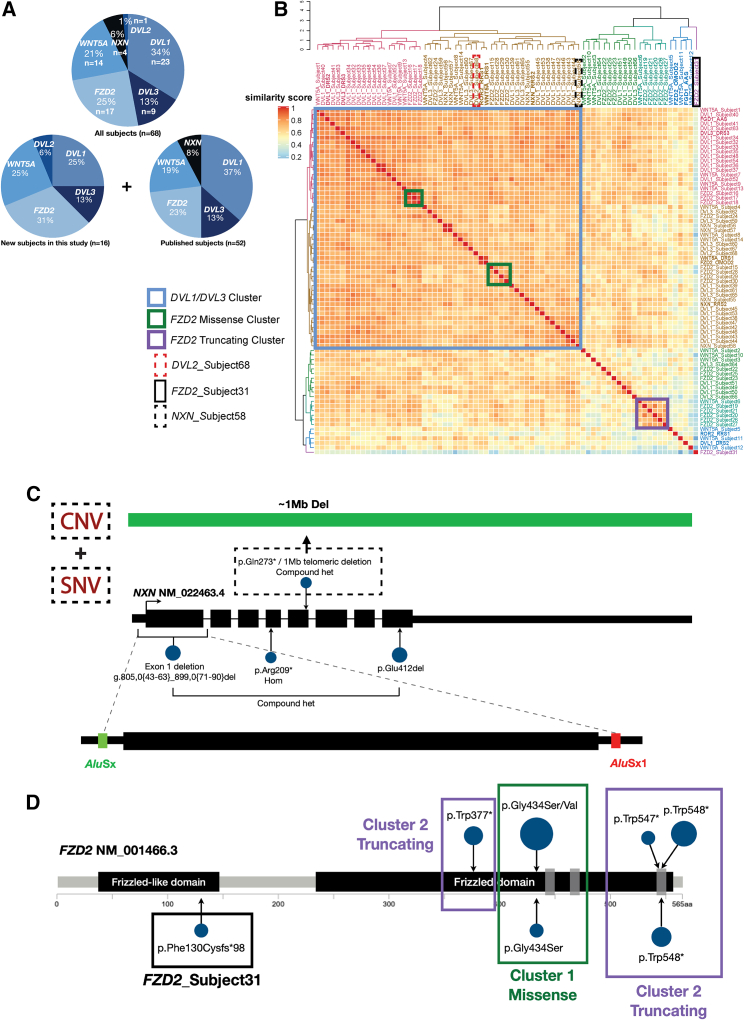
Figure 3.
RS cohort summary and genotype-phenotype characterization
(A) Cohort description. Clinical notes and molecular diagnoses of 68 subjects were collected, including 16 subjects in this study and 52 from previously published subjects. (B) HAC and visualization of quantitative phenotypic similarity allow refinement of genotype-phenotype correlations in RS. The dendrogram shown at the top and to the left of the heatmap is based on HAC analysis of the dissimilarity matrix produced from Lin semantic similarity scores and with k set to 6. Unique clusters are represented by different colors, and individual probands are labeled on top of and to the right of the heatmap. Within the heatmap, dark red indicates a higher similarity, while dark blue indicates lower similarity. A key is provided on the left. Blue rectangle: 84% of DVL1 and DVL3 individuals as well as DVL2 subject (highlighted by a red dashed rectangle) grouped into these two clusters in addition to the NXN individuals. Green rectangle: 62.5% (N = 5 out of 8) of FZD2 individuals grouped in this cluster have missense variants. Purple rectangle: 100% (N = 5 out of 5) of FZD2 individuals grouped in this cluster have truncating variants. Black rectangle: FZD2 outlier subject (FZD2_Subject31), whose clinical phenotypes are less similar to RS individuals. Black dashed rectangle: NXN_Subject58, who has a ∼1-Mb deletion. (C) Variants observed in NXN resulting in AR-RS: variants are displayed by blue circles; larger circles represent identical variants in unrelated individuals. The NXN subject marked as black dashed rectangle harbors a nonsense variant (SNV) in trans with an ∼1-Mb telomeric deletion (CNV), whereas the other three individuals all have biallelic LoF variants affecting NXN. Two of them from the same family were found to have compound heterozygous variants of an in-frame deletion and an Alu-Alu-mediated exonic deletion of the entire exon 1. (D) Patients with variants affecting FZD2 are separated in two distinct clusters based on phenotypic similarity score. Most of the individuals with missense variants affecting Gly434 in FZD2 were grouped into the missense cluster (green rectangle), whereas most of the individuals with truncating variants at C-terminal end of FZD2 were grouped into the truncating cluster (purple rectangle). Variants are displayed (blue circles) with larger circles representing identical variants in unrelated individuals. One outlier individual (FZD2_Subject31, black rectangle) was identified to have a truncating variant affecting the N terminus of the Frizzled-like domain.
Even within the highly similar phenotypic spectrum associated with RS variant alleles, subclusters of related phenotypes were computationally identifiable. As shown in Figure 3B, 84% of DVL1 and DVL3 probands (N = 27 out of 32), as well as the new DVL2 proband (red dashed rectangle), clustered into the two most similar groups (blue rectangle). This finding demonstrates that phenotypes related to variants in DVL1, DVL2, and DVL3 are similar and distinct from the “phenotype signature” and information content embodied within the specific clinical description of the trait for the given RS-associated genes. These results are supportive of a phenotypic diagnosis of RS in the individual with the DVL2 variant.
Only four individuals were found to have biallelic variants in NXN.3,4 Note that all the NXN individuals along with the OMIM-described NXN_RRS2 phenotype (MIM: 618529) were grouped into aforementioned DVL clusters (Figures 3B and 3C), blurring the phenotypic distinction between the AD and NXN-related AR forms of RS.
Probands with variants in FZD2 exhibited two phenotypic clusters (green boxes in DVL cluster and purple box, Figure 3B). Interestingly, most FZD2 individuals grouped into the DVL cluster have missense variants (N = 5 out of 8), whereas those in the purple cluster all have truncating variants (N = 5 out of 5). This suggests that RS probands with different variant allele types in FZD2 present with distinctive phenotypic signatures. In addition, for one FZD2 proband (FZD2_Subject 31; BAB14592), who has a 22-bp duplication (c.367_388dup), HPO similarity analysis indicates that this individual is an outlier whose phenotype is not similar to other RS individuals. As shown in Figure 3D, all other individuals have truncating variants affecting the C-terminal end of the Frizzled domain; the BAB14592 duplication variant allele affects the N-terminal end of the Frizzled-like domain. To ascertain which phenotypes differentiate these missense and truncating clusters, we performed genotype-phenotype analysis of FZD2 individuals with missense variants and individuals with truncating variants. The outlier individual was excluded. As shown in Figure S8 and Table S5, there are some core phenotypes (prevalence > 80% in both groups) including anteverted nares, long philtrum, short limbs, abnormal ear shape and position, broad forehead, and midface hypoplasia. In general, individuals having missense variants in FZD2 are overall more severely affected than those with truncating variants. All the individuals with missense variants have hypertelorism (N = 7 out of 7), upslanting palpebral fissures (N = 5 out of 5), prominent eyes (N = 5 out of 5), gingival hyperplasia (N = 5 out of 5), dental anomalies (N = 5 out of 5), and a sacral dimple (N = 4 out of 4); however, for individuals with truncating variants, frequencies for these features are only 29% (N = 2 out of 7), 17% (N = 1 out of 6), 43% (N = 3 out of 7), 33% (N = 1 out of 3), 33% (N = 2 out of 6), and 0% (N = 0 out of 3), respectively. Most of the individuals with missense variants exhibited a thin vermillion border (80%, N = 4 out of 5), bilobed tongue (67%, N = 46), brachydactyly (86%, N = 6 out of 7), and clinodactyly (83%, N = 5 out of 6). However, only 33% (N = 2 out of 6), 17% (N = 1 out of 6), 57% (N = 4 out of 7), and 43% (N = 3 out of 7), respectively, of individuals with truncating variants were observed to have those phenotypes. Exceptions are that high forehead was observed in 88% of individuals with truncating variants, but only half of individuals with missense variants (50%, N = 3 out of 6). All male subjects with a truncating variant have genital hypoplasia (N = 4 out of 4), but the only male with a missense variant does not.
Most WNT5A probands had quantitative phenotypes that grouped into five different clusters instead of grouping into any specific clusters (Figure 3B), which suggest that individuals with variants in WNT5A have more variable phenotypes. As WNT5A is an upstream ligand, this finding may potentially be due to the described crosstalk of WNT5A with pathways other than the non-canonical WNT/PCP pathway.68
Specificity of RS gene traits and clinical phenotyping
To visualize individual phenotypic feature overlap of individuals in this study compared with other selected Mendelian disorders (including different types of RS, FZD2-related omodysplasia, and the most frequent differential diagnosis, Aarskog-Scott syndrome) with phenotypes gleaned from OMIM, an annotation grid was generated to show HPO terms (i.e., clinical phenotypes) associated with each proband and disease (Figure S9). Individuals/different types of RS in DVL clusters and the FZD2 missense cluster had the highest similarity score, therefore reflecting the higher degree of overlap between phenotypic features. The outlier FZD2 individual (FZD2_Subject 31) only has six phenotypes in common with RS individuals and also presents with 12 distinct phenotypes (vertigo, knee flexion contracture, bowel incontinence, neurogenic bladder, fibular aplasia, bilateral talipes equinovarus, popliteal pterygium, myelomeningocele, sparse eyebrow, highly arched eyebrow, smooth philtrum, and curly hair), which suggests that frameshift variants at the N-terminal end of FZD2 may cause a different phenotype. In fact, this patient was clinically diagnosed with popliteal pterygium syndrome (MIM: 119500), although variants in IRF6 were not detected by clinical ES. The only individual (black dashed box) who has a nonsense variant in NXN in trans with a ∼1-Mb 17p13.3 telomeric deletion containing NXN4 and 14 additional genes (Figure 3B and 3C) presents with bladder stones and nephrolithiasis, suggesting that other genes deleted in this region, or mapping elsewhere in the genome, may have contributed to those phenotypes. Nine of these 14 deletion-mapped genes are known OMIM genes and three genes are related to AR disease traits, including VPS53 (MIM: 615850), GEMIN4 (MIM: 606969), and TIMM22 (MIM: 607251). In addition, a hemizygous missense VUS in a non-disease-associated gene, RPH3AL (MIM: 604881), was found in this region. Although none of these genes has been associated with disease in the heterozygous state, it is possible that haploinsufficiency of gene(s) within the region or deletion of regulatory regions cause additional phenotypic effects. In addition to overlapping phenotypic features, the disease most frequently on the differential diagnosis of RS, FGD1_AAS, exhibits distinct phenotypic features that may be useful in clinical differentiation.
Quantitative assessment of diseases on the RS differential and WNT signaling
To visualize similarity of RS and all of the diseases on the differential for RS-like phenotypes (Table 2), HPO analysis was performed similarly for 31 diseases (Figure S5; Table S3). Two distinct comparisons were performed: a disease-to-gene phenotypic comparison (Figures 4A and 4D) and a disease-to-disease phenotypic comparison (Figures 4B and 4E). We chose seven clusters for disease-to-gene phenotype comparison (Figure 4A) and seven clusters for disease-to-disease phenotypic comparison (Figure 4B) using visualization of the gap statistic curve along with phenotypic similarity by heatmap. All the abbreviations and identifier numbers for diseases included are shown in Table S3.
Figure 4.
Quantitative similarity analysis of RS phenotype supports high similarity of known differential diagnoses and indicate lack of similarity with diseases caused by variants affecting paralogous WNT-pathway genes
(A) Gap statistic curve for heatmap displayed in (D). Gap statistic for RS, RS-like, WNT paralog-associated diseases, and top disease-associated gene phenotype matches (p < 0.001) is displayed on the y axis and number of clusters tested on the x axis. The point on the curve where slope changed from a trend of higher to lower (i.e., additional numbers of clusters were not adding as much to the gap statistic) was used to choose optimal number of clusters (k = 7). (B) Gap statistic plot to determine number of clusters to display by heatmap in (E). Gap statistic for RS, RS-like, WNT paralog-associated diseases, and top disease phenotype matches (p < 0.001) is displayed on the y axis and number of clusters tested on the x axis. The gap statistic curve along with the heatmap representation in (C) were used to select the number of clusters that both represented the visually appreciable clusters in the heatmap and the greatest increase in gap statistic (k = 7). (C) Number of WNT, FZD, and ROR paralogs in human. (D and E) Semantic similarity heatmap results between disease-associated phenotypes of RS, RS phenocopies, WNT-associated disease phenotypes, and significantly similar known disease gene phenotypes (D) and OMIM-annotated disease phenotypes (E). Red rectangle: RS caused by DVL1, DVL3, WNT5A, ROR2, and NXN are clustered together with RS genes. Black arrow: the most frequent differential diagnosis of RS, Aarskog-Scott syndrome, has the highest similarity scores with RS. Green rectangle: most of the differential diagnoses observed in our in-house cohort are clustered together with RS and RS genes. Gray rectangle: diseases caused by WNTs, FZDs, or RORs not specific in WNT/PCP signaling, which can be distinguished from the RS cluster. Unique clusters are represented by different colors, and diseases/genes are labeled on top of and to the right of the heatmap. Within the heatmap, dark red indicates a higher similarity, while dark blue indicates lower similarity. A legend is provided on the left-hand side.
As expected, ROR2_RRS1, NXN_RRS2, WNT5A_DRS1, DVL1_DRS2, and DVL3_DRS3 are closely clustered together (red box in Figure 4D). In addition, most of the differential diagnoses observed in our in-house cohort (Table 2) are clustered together with RS and associated genes (green box). These are easily distinguished from diseases caused by WNTs, FZDs, or RORs paralogs (Figure 4C) not specific to WNT/PCP signaling (gray box). FGD1_AAS (black arrow) has the highest similarity scores with RS compared with other diseases on the differential diagnosis for RS, which is consistent with our observation that half of the individuals having variants associated with diseases on the RS differential rather than RS have Aarskog-Scott syndrome (Table 2).
The heatmap of disease-to-disease phenotypic similarity analysis is shown in Figure 4E. Similarly, RS caused by DVL1, DVL3, WNT5A, ROR2, and NXN are robustly clustered together (red rectangle). Most of the differential diagnoses of RS are also clustered together with RS (green rectangle), which indicates that phenotypes of those diseases are likely to substantially overlap with RS. Similar to the disease-to-gene comparison heatmap, diseases related to other WNTs, FZDs, or RORs were grouped into unique clusters (gray box) separate from RS and can be seen to have much larger dissimilarity than within the cohort-only comparison in Figure 3.
Discussion
In this study we investigate the molecular and clinical features of RS and conditions within the clinical differential diagnoses of RS. We present nine novel pathogenic or likely pathogenic variants in four RS genes and a new candidate gene, DVL2, which aids in the delineation of the molecular patterns of variants causative of AD-RS. HPO term-based quantitative phenotypic analyses of subjects and specific RS gene-associated disease traits (1) dissected gene- and allele-specific phenotypic differences between RS-associated variants, and (2) revealed that most of the phenotypically similar diseases on the RS differential cluster together with RS and can be quantitatively distinguished from diseases caused by non-RS-associated WNTs, FZDs, or RORs. These data support the hypothesis that specific WNT, FZD, and ROR paralogs are involved in RS.
We report the first RS-related likely pathogenic variant in DVL2, which is a 1-bp duplication and causes a +1 frameshift. Detailed phenotypic information, photographs, and radiographs of the DVL2 proband are shown in Figure 5 and Table S1. HPO analysis confirmed that this subject has similar phenotypes to subjects with DVL1/DVL3 variants (Figures 3B and S9). As shown in Figure S10, this +1 frameshifting variant in DVL2 and −1 frameshifting variants in DVL1/DVL3 are all predicted to generate a novel C-terminal tail of DVL proteins, whereas −1 variant in DVL2 and most of +1 variants in DVL1/DVL3 at the same location are predicted by conceptual translation to cause a much shorter C-terminal tail. This finding supports our hypothesis that frameshifting variants in DVL genes may act in a gain-of-function (GoF) manner instead of haploinsufficiency. Additional evidence to support a GoF role for the variant DVL C-terminus in bone development may come from a recent report that used whole-genome variant association across 100 screw-tail breed dogs. It suggested a similar underlying pathophysiology as it identified a homozygous −1 frameshift mutation in exon 15 of DVL2. The dogs exhibited corresponding phenotypes of RS, including brachycephaly, hypertelorism, short limbs, and vertebral malformations.15 Collectively, frameshift variants producing a variant C-terminus have been found in all three DVL gene paralogs in mammals, leading to skeletal dysplasia.
Figure 5.
Photographs and radiographs of BAB14964 who has a likely pathogenic variant in DVL2
(A) Facial characteristics include a tall forehead with high anterior hairline, hypertelorism, low nasal bridge, low-set ears, smooth philtrum, downturned corners of the mouth, and micro-retrognathia; standing photographs show micromelia with mesomelic predominance and genu valgus. (B–F) Selected radiographs from skeletal survey at 2 days old. (B) Chest radiograph and (C) magnified view: there are 11 rib pairs, with small accessory rib ossicles bilaterally, arising between the seventh and eighth ribs (white arrows). A resolving sagittal cleft is also shown in the T6 vertebral body (black arrow). There is a possible anterior rib fusion anomaly of the left first and second ribs. The T9 and T10 vertebral bodies are mildly hypoplastic, more easily seen on the lateral spine radiograph (D). Radiographs of upper limb (E) and lower limb (F) show diffuse osteosclerosis with some narrowing of the medullary spaces. There is mild mesomelic shortening of the upper limb, and mild rhizomelic shortening of the lower limb. (G and H) Radiographs of left arm (G) and left hand (H) at age 3 years. Mesomelic shortening is now more pronounced but remains mild. Diffuse osteosclerosis is no longer apparent. There is hypoplasia of the first metacarpal, second and fifth distal phalanges, and fifth middle phalanx, with associated clinodactyly of the little finger.
Most of the pathogenic variants in WNT5A are related to cysteine changes, whereas recurrent missense and truncating variants in FZD2 cause AD-RS. Probands with variants in FZD2 exhibited two phenotypic clusters, one of missense variants and another of truncating variants (Figure 3B). As shown in Figure 3D, all other individuals have truncating variants affecting the C-terminal end of the Frizzled domain, except for an outlier individual, who has a 22-bp duplication affecting the N-terminal end of the Frizzled-like domain. Whether the mutant mRNA escapes NMD cannot be predicted as FZD2 is a single-exon gene. This finding suggests that the frameshift variant at the N-terminal end of FZD2 may cause development of a unique set of phenotypic traits, and that the recurrent missense or truncating variants affecting the C-terminal Frizzled domain contribute to the development of AD-RS.
Although omodysplasia 2 (OMOD2) has been related to FZD2 variants in the OMIM database and several individuals,48, 49, 50, 51 a recent published review2 as well as our quantitative HPO analysis (Figure 3) indicate that it is unclear whether FZD2-related phenotypes should be classified as AD-RS, omodysplasia, or both. After the original description of the association of FZD2 with AD OMOD2 by Saal et al.,48 Nagasaki et al.50 also suggested that the phenotypic overlap between AD OMOD2 and RS, particularly AD-RS, was significant. The same study asserted that AD OMOD2 could be considered an RS-like syndrome. A review of OMOD2-related phenotypic traits, which were clinically described before as associated with FZD2 variation,69, 70, 71 revealed that many features not included in the OMIM clinical synopsis are indeed present in OMOD2 probands and represent features (i.e., hypertelorism, dental crowding, midface hypoplasia) that strengthen the phenotypic overlap between OMOD2 and RS. To address this question, an HPO analysis was performed for 16 FZD2 subjects (the outlier subject with the 22-bp duplication, FZD2_Subject31, is excluded) and mesomelic and rhizo-mesomelic dysplasias gleaned from the International Skeletal Dysplasia Society nosology and classification of genetic skeletal disorders (2019 revision).43 This analysis revealed that all 16 FZD2 subjects group with RS caused by NXN, DVL1, DVL3, ROR2, and WNT5A, and as well this FZD2 group, is distinct from the cluster of omodysplasias and mesomelic dysplasias (Figure 6). Taken together, these data objectively support the contention that OMOD2 is not a unique clinical entity distinguishable from RS and should instead be considered as FZD2-associated AD-RS.
Figure 6.
Quantitative similarity analysis of subjects carrying pathogenic variants affecting FZD2 reveal phenotypic clustering with RS
Phenotypic data from 16 subjects with pathogenic FZD2 variants were compiled and similarity analysis was performed against RS OMIM genes (n = 6) and other mesomelic/rhizo-mesomelic dysplasias (n = 9), including omodysplasia 1 and 2 as per Mortier et al.43 Left: gap statistic, a cutoff value of p < 0.001 for similarity was used to define the group of diseases used for gap statistic testing, which was subsequently used to choose the number of clusters (k = 2). Right: HAC and heatmap for mesomelic/rhizo-mesomelic dysplasias, FZD2 subjects, and significant disease matches.
It should be noted that there is an imbalance in the number of HPO terms among individuals from different RS gene groups (Figure S6), and this could represent either incomplete phenotypic evaluation of a particular individual or it could reflect true differences in phenotypic manifestations for particular genotypes. As discussed above, individuals with variants in FZD2, especially truncating variants, tend to be more mildly affected, as represented by the lower number of associated phenotypes. While individuals with WNT5A-RS have more variable numbers of associated phenotypes, this may be due to crosstalk of WNT5A in different WNT signaling pathways. One exception is that three individuals (WNT5A_Subject 5, 11, and 12) were not grouped with other RS subjects (Figure 3B). This exception may be primarily due to the lack of clinical data (those individuals all have fewer than 10 HPO terms; Figure S9), as most of the phenotypic information for these three individuals is not available. We anticipate that, with additional patient data collection and allelic series investigations, future studies of these genes could deepen our understanding of trait manifestations and potential limitations of quantitative phenotyping.
As shown in Table 2, 12 individuals reported here and from the literature were found to have pathogenic variants in genes related to diseases with phenotypic overlap with RS in our cohort, and most of the differential diagnoses are grouped together with RS based on our HPO analysis (Figures 4D and 4E). We suggest adding FGD1 to RS gene panels for clinical studies as (1) pathogenic variants in FGD1 are the most frequent molecular diagnostic finding in clinically diagnosed RS with otherwise negative testing, and (2) FGD1_AAS has the highest similarity score with RS in our HPO analysis.
To specifically investigate if there are any other phenocopies with RS, RS caused by DVL1, DVL3, WNT5A, ROR2, and NXN were used to query for similarity to OMIM-annotated diseases (n = 7,946), diseases with the highest similarity score were selected and the results were visualized by a cluster heatmap. As shown in Figure S11, RS caused by different genes are strongly clustered together (red box). FGD_AAS1 grouped with 28 skeletal dysplasias in the cluster with the highest similarity to the RS cluster (blue box). FZD2_OMOD2 was clustered with omodysplasia 1 (MIM: 258315) (green box), with lower similarity scores than diseases in blue cluster. The identified diseases observed to be phenotypically similar to RS may help identify molecular diagnoses in unsolved cases clinically diagnosed as RS.
We have previously suggested RAC3 (MIM: 602050) as a potential candidate AD-RS gene based on one clinically diagnosed subject with apparent RS and a de novo missense variant allele in RAC3.3 However, recent studies have defined a neurodevelopmental clinical phenotype distinct from RS associated with RAC3 variations.72,73 Besides dysmorphic facial features, the individual described in White et al.3 also had developmental delay, seizures, abnormal electroencephalogram (EEG), and a thin corpus callosum. A clinical reevaluation of this individual is more consistent with a distinct RAC3-related neurodevelopmental disorder (MIM: 618577)72, 73, 74 than with RS.
Previously, the most important clinical signs for the characterization of RS were defined as hypertelorism, nasal features, midface hypoplasia, mesomelic limb shortening, brachydactyly, clinodactyly, micropenis, and short stature, which were present in more than 75% of individuals with either AD-RS or AR-RS.20 In our study (Table S2, Figure 2), midface hypoplasia, hypertelorism, prominent eyes, anteverted nares, wide, low nasal bridge, gingival hyperplasia, limb length anomaly, brachydactyly, broad first toe, and male hypoplastic genitalia were observed in more than 60% of individuals in each genotype. However, only 38% (N = 8 out of 21) of DVL1 individuals have short stature, which contrasts with the prevalence of short stature in all other RS types (69%–100%), although this trend did not reach statistical significance (Table S2). Moreover, scoliosis was present in more than 75% of individuals with the recessive form (ROR2 mediated), but less than 25% of individuals with the dominant form based on a previous report.20 In this study, only AR-RS individuals with variants in NXN were studied but none of them had scoliosis. In AD-RS groups, there was a lower than 50% prevalence of scoliosis found for each genotype, consistent with previous observation that this is not a common phenotype in AD-RS. It has been suggested that dental crowding and hypodontia were more severe and frequently observed in individuals with AD-RS.21 We observed that the prevalence of dental anomalies reaches >85% in all genotype groups except in individuals with FZD2-RS (58%).
A group of papers published recently reported some genotype-phenotype correlations among 13 individuals recruited at an RS clinical conference in 2018, and most of those patients are also included in this current analysis. Bifid tongue was associated with individuals with pathogenic variants in the DVL1, ROR2, GPC4, or NXN genes but was not observed in individuals with WNT5A-RS.75 The prevalence of bifid tongue is 38% (N = 3 out of 8) in our cohort of individuals with pathogenic WNT5A variation, comparable with other genotypes. Mesomelia was seen in individuals encompassing all genetic variants except one NXN individual based on previous report.76 However, the NXN individuals reported here were observed to have mesomelia at a prevalence of 50% (N = 2 out of 4). These differences demonstrate the need for characterization of additional NXN individuals to more precisely define an NXN-associated phenotypic spectrum.
The genotype-phenotype correlations identified in this study may also help characterize the phenotypic spectrum of RS caused by variants in different genes and allelic types (e.g., missense versus frameshift) within those genes, helping to understand the molecular functions of RS genes. Almost all cases of DVL1-RS have macrocephaly, and increased bone density (skull), which are phenotypes suggestive of increased WNT canonical signaling levels.77 It is possible that not only perturbation of WNT/PCP signaling but also a disturbed balance between the levels of WNT/β-catenin and WNT/PCP signaling may underlie RS pathobiology. In DVL3-related RS, a significantly higher prevalence of heart defects was found. Notably, Dvl3 knockout mice show cardiovascular outflow tract defects, but Dvl1 knockout mice did not show any cardiac defects,13,78 which implicates DVL3 as a key factor in heart development in both humans and mice. The complexity of the WNT signaling mechanisms in humans is primarily due to an abundance of gene paralogs encoding for 19 different WNTs, 10 different FZDs, three different DVLs, and two RORs.79 As demonstrated in Figure 4, diseases caused by other WNTs, FZDs, and RORs are not similar to RS and formed distinct clusters. Thus, our analysis suggests that phenotypes of diseases caused by other WNTs, FZDs, or RORs are distinct from RS and further supports that RS is caused by perturbation of the specific pathway activated by WNT5A, FZD2, and ROR2 instead of other genes in or branches of WNT signaling.
We identified previously unreported pathogenic variants in known RS genes as well as genes associated with other conditions on the RS differential. These findings deepen our understanding of the molecular and clinical etiology of RS. In 2018, White et al.3 reported molecular diagnoses of a cohort of 21 individuals clinically diagnosed with RS. In that cohort, 61.5% of individuals were found to have pathogenic variants in known, previously published RS-associated genes, 24% of individuals were found to have variants in genes associated with diseases considered to be on the AD-RS differential, and 14.5% of individuals were unsolved or solved by candidate RS-associated genes that had not yet been observed in a sufficient number of probands to establish a firm association. If we re-examine those data in our current in-house database (as of October 2021, 82 individuals in total), those numbers are 75.6%, 14.6%, and 9.8%, respectively. Based on these numbers, it can be concluded that the major genes underlying the molecular etiology of RS have been identified, although the discovery of novel candidate genes only observed in a single individual (i.e., DVL2) or in a few individuals (i.e., NXN) requires additional investigation of individuals with RS.
ES is becoming a first-line test for molecular diagnosis of congenital disease,80 and the dissection of gene and allele-specific phenotypic spectrums using HPO-based phenotypic similarity analyses may help to provide more specific prognosis and counseling based on variant or gene implicated, and, ultimately, provide genotype-guided management. Moreover, ES can be particularly enlightening in (1) conditions with locus heterogeneity, (2) multi-locus pathogenic variation (MPV), and (3) genocopies as exemplified by RS and Aarskog syndrome.
In summary, our study: (1) identifies previously undescribed pathogenic variant alleles, (2) provides evidence for a new RS disease gene DVL2, (3) examines paralog specificity of trait manifestations, (4) delineates allele-specific phenotypic differences, (5) provides biological insights into a genetically heterogeneous disorder, (6) indicates that missense or truncating variants affecting the C-terminal end of the Frizzled domain in FZD2 lead to a clinical phenotype consistent with RS, and (7) reveals the power of deep phenotypic analyses to tease apart genetically heterogeneous disorders.
Data and code availability
The code supporting the current study has not been deposited in a public repository because it is being updated locally to accommodate further analysis, but it is available from the corresponding author on request. The accession numbers for the identified variants in this paper are ClinVar: SCV000256688, SCV001438002, SCV001438003, SCV001438004, SCV001438005, SCV001438006, SCV001438007, SCV001438008, SCV001438009, SCV001438010, SCV001438011, SCV001438012, SCV001438013, SCV001438014, and SCV00143805 and the preliminary identifier SUB9931810. The dbGAP accession number for all exome sequences reported in this paper and with consent for data sharing in controlled-access databases is dbGAP:phs000711.v5.p1.
Acknowledgments
The authors would like to thank the individuals and their families who contributed to this study, especially Kim Kremeier and the Robinow Syndrome Foundation for facilitating this collaboration. We would also like to thank Dr. Chad A. Shaw for helpful discussions about the quantitative methodology applied here to study HPO terms. This work was supported in part by the US National Human Genome Research Institute (NHGRI)/NHLBI grant number UM1HG006542 to the Baylor-Hopkins Center for Mendelian Genomics (BHCMG); the National Institute of Neurological Disorders and Stroke (NINDS) R35 NS105078 (J.R.L.); the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD) R03 HD092569 (C.M.B.C and V.R.S.); the National Institute of General Medical Sciences (NIGMS) R01 GM132589 (C.M.B.C); NHGRI K08 HG008986 (J.E.P.); the JPB Foundation (W.K.C.); the BCM Medical Scientist Training Program and the Department of Genetics and Genomics (A.J.); the NIH T32 (GM07526-43) Medical Genetics Research Fellowship Program (B.J.S.); and the American College of Medical Genetics and Genomics Foundation for Genetics and Genomic Medicine (B.J.S.).
This research was made possible through access to the data and findings generated by the 100,000 Genomes Project. The 100,000 Genomes Project is managed by Genomics England Limited (a wholly owned company of the Department of Health and Social Care). The 100,000 Genomes Project is funded by the National Institute for Health Research and NHS England. The Wellcome Trust, Cancer Research UK, and the Medical Research Council have also funded research infrastructure. The 100,000 Genomes Project uses data provided by patients and collected by the National Health Service as part of their care and support.
Declaration of interests
BCM and Miraca Holdings have formed a joint venture with shared ownership and governance of BG, which performs clinical microarray analysis (CMA), clinical ES (cES), and clinical biochemical studies. V.R.S. and P.L. receive professional services compensation from BG and J.R.L. serves on the Scientific Advisory Board of the BG. J.R.L. has stock ownership in 23andMe, is a paid consultant for the Regeneron Genetics Center, and is a coinventor on multiple United States and European patents related to molecular diagnostics for inherited neuropathies, eye diseases, and bacterial genomic fingerprinting. W.K.C. is a is a paid consultant for the Regeneron Genetics Center. P.C.S. and S.M. are employees of GeneDx. The other authors declare no competing interests.
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.xhgg.2021.100074.
Web resources
The URLs for data presented herein are as follows:
ARIC database, http://www2.cscc.unc.edu/aric/
BCM-HGSC Mercury analysis pipeline, https://www.hgsc.bcm.edu/software/mercury.
BCM-HGSC ERIS pipeline, https://www.hgsc.bcm.edu/software/eris.
GATK HaplotypeCaller, https://gatk.broadinstitute.org/hc/en-us/articles/360037225632-HaplotypeCaller.
gnomAD, https://gnomad.broadinstitute.org/
PolyPhen-2, http://genetics.bwh.harvard.edu/pph2/
SIFT, https://sift.bii.a-star.edu.sg/
CADD, https://cadd.gs.washington.edu/
AluAluCNVpredictor tool, http://alualucnvpredictor.research.bcm.edu:3838.
HPO resource page, https://hpo.jax.org/app/download/annotation.
Human Splicing Finder web tool, https://www.genomnis.com/access-hsf
dbGaP, http://www.ncbi.nlm.nih.gov/gap.
NHLBI ESP Exome Variant Server, http://evs.gs.washington.edu/EVS/
OMIM, http://www.omim.org/
Supplemental information
References
- 1.Robinow M., Silverman F.N., Smith H.D. A newly recognized dwarfing syndrome. Am. J. Dis. Child. 1969;117:645–651. doi: 10.1001/archpedi.1969.02100030647005. [DOI] [PubMed] [Google Scholar]
- 2.Mazzeu J.F., Brunner H.G. 50 years of Robinow syndrome. Am. J. Med. Genet. A. 2020;182:2005–2007. doi: 10.1002/ajmg.a.61756. [DOI] [PubMed] [Google Scholar]
- 3.White J.J., Mazzeu J.F., Coban-Akdemir Z., Bayram Y., Bahrambeigi V., Hoischen A., van Bon B.W.M., Gezdirici A., Gulec E.Y., Ramond F., et al. WNT signaling perturbations underlie the genetic heterogeneity of Robinow syndrome. Am. J. Hum. Genet. 2018;102:27–43. doi: 10.1016/j.ajhg.2017.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zhang C., Mazzeu J.F., Eisfeldt J., Grochowski C.M., White J., Akdemir Z.C., Jhangiani S.N., Muzny D.M., Gibbs R.A., Lindstrand A., et al. Novel pathogenic genomic variants leading to autosomal dominant and recessive Robinow syndrome. Am. J. Med. Genet. A. 2021;185:3593–3600. doi: 10.1002/ajmg.a.61908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Oishi I., S.H., Onishi N., Takada R., Kani S., Ohkawara B., Koshida I., Suzuki K., Yamada G., Schwabe G.C., et al. The receptor tyrosine kinase Ror2 is involved in non-canonical Wnt5a/JNK signalling pathway. Genes Cells. 2003;8:645–654. doi: 10.1046/j.1365-2443.2003.00662.x. [DOI] [PubMed] [Google Scholar]
- 6.Minami Y., Oishi I., Endo M., Nishita M. Ror-family receptor tyrosine kinases in noncanonical Wnt signaling: their implications in developmental morphogenesis and human diseases. Dev. Dyn. 2010;239:1–15. doi: 10.1002/dvdy.21991. [DOI] [PubMed] [Google Scholar]
- 7.Funato Y., Michiue T., Asashima M., Miki H. The thioredoxin-related redox-regulating protein nucleoredoxin inhibits Wnt-beta-catenin signalling through dishevelled. Nat. Cell. Biol. 2006;8:501–508. doi: 10.1038/ncb1405. [DOI] [PubMed] [Google Scholar]
- 8.Funato Y., Michiue T., Terabayashi T., Yukita A., Danno H., Asashima M., Miki H. Nucleoredoxin regulates the Wnt/planar cell polarity pathway in Xenopus. Genes Cells. 2008;13:965–975. doi: 10.1111/j.1365-2443.2008.01220.x. [DOI] [PubMed] [Google Scholar]
- 9.Bunn K.J., Daniel P., Rosken H.S., O'Neill A.C., Cameron-Christie S.R., Morgan T., Brunner H.G., Lai A., Kunst H.P., Markie D.M., et al. Mutations in DVL1 cause an osteosclerotic form of Robinow syndrome. Am. J. Hum. Genet. 2015;96:623–630. doi: 10.1016/j.ajhg.2015.02.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.White J., Mazzeu J.F., Hoischen A., Jhangiani S.N., Gambin T., Alcino M.C., Penney S., Saraiva J.M., Hove H., Skovby F., et al. DVL1 frameshift mutations clustering in the penultimate exon cause autosomal-dominant Robinow syndrome. Am. J. Hum. Genet. 2015;96:612–622. doi: 10.1016/j.ajhg.2015.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.White J.J., Mazzeu J.F., Hoischen A., Bayram Y., Withers M., Gezdirici A., Kimonis V., Steehouwer M., Jhangiani S.N., Muzny D.M., et al. DVL3 alleles resulting in a −1 frameshift of the last exon mediate autosomal-dominant Robinow syndrome. Am. J. Hum. Genet. 2016;98:553–561. doi: 10.1016/j.ajhg.2016.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Semenov M.V., Snyder M. Human dishevelled genes constitute a DHR-containing multigene family. Genomics. 1997;42:302–310. doi: 10.1006/geno.1997.4713. [DOI] [PubMed] [Google Scholar]
- 13.Etheridge S.L., Ray S., Li S., Hamblet N.S., Lijam N., Tsang M., Greer J., Kardos N., Wang J., Sussman D.J., et al. Murine dishevelled 3 functions in redundant pathways with dishevelled 1 and 2 in normal cardiac outflow tract, cochlea, and neural tube development. PLoS Genet. 2008;4:e1000259. doi: 10.1371/journal.pgen.1000259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hamblet N.S., Lijam N., Ruiz-Lozano P., Wang J., Yang Y., Luo Z., Mei L., Chien K.R., Sussman D.J., Wynshaw-Boris A. Dishevelled 2 is essential for cardiac outflow tract development, somite segmentation and neural tube closure. Development. 2002;129:5827–5838. doi: 10.1242/dev.00164. [DOI] [PubMed] [Google Scholar]
- 15.Mansour T.A., Lucot K., Konopelski S.E., Dickinson P.J., Sturges B.K., Vernau K.L., Choi S., Stern J.A., Thomasy S.M., Doring S., et al. Whole genome variant association across 100 dogs identifies a frame shift mutation in DISHEVELLED 2 which contributes to Robinow-like syndrome in Bulldogs and related screw tail dog breeds. PLoS Genet. 2018;14:e1007850. doi: 10.1371/journal.pgen.1007850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Person A.D., Beiraghi S., Sieben C.M., Hermanson S., Neumann A.N., Robu M.E., Schleiffarth J.R., Billington C.J., Jr., van Bokhoven H., Hoogeboom J.M., et al. WNT5A mutations in patients with autosomal dominant Robinow syndrome. Dev. Dyn. 2010;239:327–337. doi: 10.1002/dvdy.22156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Roifman M., Marcelis C.L., Paton T., Marshall C., Silver R., Lohr J.L., Yntema H.G., Venselaar H., Kayserili H., van Bon B., et al. De novo WNT5A-associated autosomal dominant Robinow syndrome suggests specificity of genotype and phenotype. Clin. Genet. 2015;87:34–41. doi: 10.1111/cge.12401. [DOI] [PubMed] [Google Scholar]
- 18.Xiong S., Chitayat D., Wei X., Zhu J., Lu W., Sun L.M., Chopra M. A novel de-novo WNT5A mutation in a Chinese patient with Robinow syndrome. Clin. Dysmorphol. 2016;25:186–189. doi: 10.1097/MCD.0000000000000130. [DOI] [PubMed] [Google Scholar]
- 19.Orrico A., Galli L., Faivre L., Clayton-Smith J., Azzarello-Burri S.M., Hertz J.M., Jacquemont S., Taurisano R., Arroyo Carrera I., Tarantino E., et al. Aarskog-Scott syndrome: clinical update and report of nine novel mutations of the FGD1 gene. Am. J. Med. Genet. A. 2010;152a:313–318. doi: 10.1002/ajmg.a.33199. [DOI] [PubMed] [Google Scholar]
- 20.Mazzeu J.F., Pardono E., Vianna-Morgante A.M., Richieri-Costa A., Ae Kim C., Brunoni D., Martelli L., de Andrade C.E., Colin G., Otto P.A. Clinical characterization of autosomal dominant and recessive variants of Robinow syndrome. Am. J. Med. Genet. A. 2007;143:320–325. doi: 10.1002/ajmg.a.31592. [DOI] [PubMed] [Google Scholar]
- 21.Beiraghi S., Leon-Salazar V., Larson B.E., John M.T., Cunningham M.L., Petryk A., Lohr J.L. Craniofacial and intraoral phenotype of Robinow syndrome forms. Clin. Genet. 2011;80:15–24. doi: 10.1111/j.1399-0004.2011.01683.x. [DOI] [PubMed] [Google Scholar]
- 22.Samuel G.N., Farsides B. The UK's 100,000 Genomes Project: manifesting policymakers' expectations. New Genet. Soc. 2017;36:336–353. doi: 10.1080/14636778.2017.1370671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Posey J.E., O'Donnell-Luria A.H., Chong J.X., Harel T., Jhangiani S.N., Coban Akdemir Z.H., Buyske S., Pehlivan D., Carvalho C.M.B., Baxter S., et al. Insights into genetics, human biology and disease gleaned from family based genomic studies. Genet. Med. 2019;21:798–812. doi: 10.1038/s41436-018-0408-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bainbridge M.N., Wang M., Wu Y., Newsham I., Muzny D.M., Jefferies J.L., Albert T.J., Burgess D.L., Gibbs R.A. Targeted enrichment beyond the consensus coding DNA sequence exome reveals exons with higher variant densities. Genome Biol. 2011;12:R68. doi: 10.1186/gb-2011-12-7-r68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.McCarthy D.J., Humburg P., Kanapin A., Rivas M.A., Gaulton K., Cazier J.B., Donnelly P. Choice of transcripts and software has a large effect on variant annotation. Genome Med. 2014;6:26. doi: 10.1186/gm543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Challis D., Yu J., Evani U.S., Jackson A.R., Paithankar S., Coarfa C., Milosavljevic A., Gibbs R.A., Yu F. An integrative variant analysis suite for whole exome next-generation sequencing data. BMC Bioinformatics. 2012;13:8. doi: 10.1186/1471-2105-13-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Reid J.G., Carroll A., Veeraraghavan N., Dahdouli M., Sundquist A., English A., Bainbridge M., White S., Salerno W., Buhay C., et al. Launching genomics into the cloud: deployment of Mercury, a next generation sequence analysis pipeline. BMC Bioinformatics. 2014;15:30. doi: 10.1186/1471-2105-15-30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Farek J., Hughes D., Mansfield A., Krasheninina O., Nasser W., Sedlazeck F.J., Khan Z., Venner E., Metcalf G., Boerwinkle E., et al. xAtlas: scalable small variant calling across heterogeneous next-generation sequencing experiments. 2018. https://doiorg/101101/295071 [DOI] [PMC free article] [PubMed]
- 29.Eldomery M.K., Coban-Akdemir Z., Harel T., Rosenfeld J.A., Gambin T., Stray-Pedersen A., Kury S., Mercier S., Lessel D., Denecke J., et al. Lessons learned from additional research analyses of unsolved clinical exome cases. Genome Med. 2017;9:26. doi: 10.1186/s13073-017-0412-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Karczewski K.J., Francioli L.C., Tiao G., Cummings B.B., Alfoldi J., Wang Q., Collins R.L., Laricchia K.M., Ganna A., Birnbaum D.P., et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–443. doi: 10.1038/s41586-020-2308-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Investigators T.A. The Atherosclerosis Risk in Communities (ARIC) study: design and objectives. Am. J. Epidemiol. 1989;129:687–702. [PubMed] [Google Scholar]
- 32.Adzhubei I.A., Schmidt S., Peshkin L., Ramensky V.E., Gerasimova A., Bork P., Kondrashov A.S., Sunyaev S.R. A method and server for predicting damaging missense mutations. Nat. Methods. 2010;7:248–249. doi: 10.1038/nmeth0410-248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Vaser R., Adusumalli S., Leng S.N., Sikic M., Ng P.C. SIFT missense predictions for genomes. Nat. Protoc. 2016;11:1–9. doi: 10.1038/nprot.2015.123. [DOI] [PubMed] [Google Scholar]
- 34.Kircher M., Witten D.M., Jain P., O'Roak B.J., Cooper G.M., Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 2014;46:310–315. doi: 10.1038/ng.2892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Fromer M., Moran J.L., Chambert K., Banks E., Bergen S.E., Ruderfer D.M., Handsaker R.E., McCarroll S.A., O'Donovan M.C., Owen M.J., et al. Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth. Am. J. Hum. Genet. 2012;91:597–607. doi: 10.1016/j.ajhg.2012.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Gambin T., Akdemir Z.C., Yuan B., Gu S., Chiang T., Carvalho C.M.B., Shaw C., Jhangiani S., Boone P.M., Eldomery M.K., et al. Homozygous and hemizygous CNV detection from exome sequencing data in a Mendelian disease cohort. Nucleic Acids Res. 2017;45:1633–1648. doi: 10.1093/nar/gkw1237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Karaca E., Posey J.E., Coban Akdemir Z., Pehlivan D., Harel T., Jhangiani S.N., Bayram Y., Song X., Bahrambeigi V., Yuregir O.O., et al. Phenotypic expansion illuminates multilocus pathogenic variation. Genet. Med. 2018;20:1528–1537. doi: 10.1038/gim.2018.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yang Y., Muzny D.M., Reid J.G., Bainbridge M.N., Willis A., Ward P.A., Braxton A., Beuten J., Xia F., Niu Z., et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N. Engl. J. Med. 2013;369:1502–1511. doi: 10.1056/NEJMoa1306555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Retterer K., Juusola J., Cho M.T., Vitazka P., Millan F., Gibellini F., Vertino-Bell A., Smaoui N., Neidich J., Monaghan K.G., et al. Clinical application of whole-exome sequencing across clinical indications. Genet. Med. 2016;18:696–704. doi: 10.1038/gim.2015.148. [DOI] [PubMed] [Google Scholar]
- 40.Lin D. An information-theoretic definition of similarity. ICML. 1998;98:296–304. [Google Scholar]
- 41.Liu P., Meng L., Normand E.A., Xia F., Song X., Ghazi A., Rosenfeld J., Magoulas P.L., Braxton A., Ward P., et al. Reanalysis of clinical exome sequencing data. N. Engl. J. Med. 2019;380:2478–2480. doi: 10.1056/NEJMc1812033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ward J.H. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 1963;58:236–244. [Google Scholar]
- 43.Mortier G.R., Cohn D.H., Cormier-Daire V., Hall C., Krakow D., Mundlos S., Nishimura G., Robertson S., Sangiorgi L., Savarirayan R., et al. Nosology and classification of genetic skeletal disorders: 2019 revision. Am. J. Med. Genet. A. 2019;179:2393–2419. doi: 10.1002/ajmg.a.61366. [DOI] [PubMed] [Google Scholar]
- 44.Mishra R., Jain V., Gupta D., Saxena R., Kulshreshtha S., Ramprasad V.L., Verma I.C., Dua Puri R. Robinow syndrome and brachydactyly: an interplay of high-throughput sequencing and deep phenotyping in a kindred. Mol. Syndromol. 2020;11:43–49. doi: 10.1159/000505506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Rai A., Patil S.J., Srivastava P., Gaurishankar K., Phadke S.R. Clinical and molecular characterization of four patients with Robinow syndrome from different families. Am. J. Med. Genet. A. 2021;185:1105–1112. doi: 10.1002/ajmg.a.62082. [DOI] [PubMed] [Google Scholar]
- 46.Murali C.N., Keena B., Zackai E.H. Robinow syndrome: a diagnosis at the fingertips. Clin. Dysmorphol. 2018;27:135–137. doi: 10.1097/MCD.0000000000000230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Danyel M., Kortum F., Dathe K., Kutsche K., Horn D. Autosomal dominant Robinow syndrome associated with a novel DVL3 splice mutation. Am. J. Med. Genet. A. 2018;176:992–996. doi: 10.1002/ajmg.a.38635. [DOI] [PubMed] [Google Scholar]
- 48.Saal H.M., Prows C.A., Guerreiro I., Donlin M., Knudson L., Sund K.L., Chang C.F., Brugmann S.A., Stottmann R.W. A mutation in FRIZZLED2 impairs Wnt signaling and causes autosomal dominant omodysplasia. Hum. Mol. Genet. 2015;24:3399–3409. doi: 10.1093/hmg/ddv088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Turkmen S., Spielmann M., Gunes N., Knaus A., Flottmann R., Mundlos S., Tuysuz B. A novel de novo FZD2 mutation in a patient with autosomal dominant omodysplasia. Mol. Syndromol. 2017;8:318–324. doi: 10.1159/000479721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Nagasaki K., Nishimura G., Kikuchi T., Nyuzuki H., Sasaki S., Ogawa Y., Saitoh A. Nonsense mutations in FZD2 cause autosomal-dominant omodysplasia: Robinow syndrome-like phenotypes. Am. J. Med. Genet. A. 2018;176:739–742. doi: 10.1002/ajmg.a.38623. [DOI] [PubMed] [Google Scholar]
- 51.Warren H.E., Louie R.J., Friez M.J., Frias J.L., Leroy J.G., Spranger J.W., Skinner S.A., Champaigne N.L. Two unrelated patients with autosomal dominant omodysplasia and FRIZZLED2 mutations. Clin. Case Rep. 2018;6:2252–2255. doi: 10.1002/ccr3.1818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Birgmeier J., Esplin E.D., Jagadeesh K.A., Guturu H., Wenger A.M., Chaib H., Buckingham J.A., Bejerano G., Bernstein J.A. Biallelic loss-of-function WNT5A mutations in an infant with severe and atypical manifestations of Robinow syndrome. Am. J. Med. Genet. A. 2018;176:1030–1036. doi: 10.1002/ajmg.a.38636. [DOI] [PubMed] [Google Scholar]
- 53.Desmet F.O., Hamroun D., Lalande M., Collod-Beroud G., Claustres M., Beroud C. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009;37:e67. doi: 10.1093/nar/gkp215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Hwang D.G., Green P. Bayesian Markov chain Monte Carlo sequence analysis reveals varying neutral substitution patterns in mammalian evolution. Proc. Natl. Acad. Sci. U S A. 2004;101:13994–14001. doi: 10.1073/pnas.0404142101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Walsh C.P., Xu G.L. Cytosine methylation and DNA repair. Curr. Top. Microbiol. Immunol. 2006;301:283–315. doi: 10.1007/3-540-31390-7_11. [DOI] [PubMed] [Google Scholar]
- 56.Arnheim N., Calabrese P. Understanding what determines the frequency and pattern of human germline mutations. Nat. Rev. Genet. 2009;10:478–488. doi: 10.1038/nrg2529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Segurel L., Wyman M.J., Przeworski M. Determinants of mutation rate variation in the human germline. Annu. Rev. Genomics Hum. Genet. 2014;15:47–70. doi: 10.1146/annurev-genom-031714-125740. [DOI] [PubMed] [Google Scholar]
- 58.Punchihewa C., Ferreira A.M., Cassell R., Rodrigues P., Fujii N. Sequence requirement and subtype specificity in the high-affinity interaction between human frizzled and dishevelled proteins. Protein Sci. 2009;18:994–1002. doi: 10.1002/pro.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Tauriello D.V., Jordens I., Kirchner K., Slootstra J.W., Kruitwagen T., Bouwman B.A., Noutsou M., Rudiger S.G., Schwamborn K., Schambony A., et al. Wnt/beta-catenin signaling requires interaction of the Dishevelled DEP domain and C terminus with a discontinuous motif in Frizzled. Proc. Natl. Acad. Sci. U S A. 2012;109:E812–E820. doi: 10.1073/pnas.1114802109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Janda C.Y., Waghray D., Levin A.M., Thomas C., Garcia K.C. Structural basis of Wnt recognition by frizzled. Science. 2012;337:59–64. doi: 10.1126/science.1222879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.MacDonald B.T., Hien A., Zhang X., Iranloye O., Virshup D.M., Waterman M.L., He X. Disulfide bond requirements for active Wnt ligands. J. Biol. Chem. 2014;289:18122–18136. doi: 10.1074/jbc.M114.575027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Song X., Beck C.R., Du R., Campbell I.M., Coban-Akdemir Z., Gu S., Breman A.M., Stankiewicz P., Ira G., Shaw C.A., et al. Predicting human genes susceptible to genomic instability associated with Alu/Alu-mediated rearrangements. Genome Res. 2018;28:1228–1242. doi: 10.1101/gr.229401.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Rosenfeld J.A., Leppig K., Ballif B.C., Thiese H., Erdie-Lalena C., Bawle E., Sastry S., Spence J.E., Bandholz A., Surti U., et al. Genotype-phenotype analysis of TCF4 mutations causing Pitt-Hopkins syndrome shows increased seizure activity with missense mutations. Genet. Med. 2009;11:797–805. doi: 10.1097/GIM.0b013e3181bd38a9. [DOI] [PubMed] [Google Scholar]
- 64.Gripp K.W., Aldinger K.A., Bennett J.T., Baker L., Tusi J., Powell-Hamilton N., Stabley D., Sol-Church K., Timms A.E., Dobyns W.B. A novel rasopathy caused by recurrent de novo missense mutations in PPP1CB closely resembles Noonan syndrome with loose anagen hair. Am. J. Med. Genet. A. 2016;170:2237–2247. doi: 10.1002/ajmg.a.37781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Bertola D., Yamamoto G., Buscarilli M., Jorge A., Passos-Bueno M.R., Kim C. The recurrent PPP1CB mutation p.Pro49Arg in an additional Noonan-like syndrome individual: broadening the clinical phenotype. Am. J. Med. Genet. A. 2017;173:824–828. doi: 10.1002/ajmg.a.38070. [DOI] [PubMed] [Google Scholar]
- 66.Zambrano R.M., Marble M., Chalew S.A., Lilje C., Vargas A., Lacassie Y. Further evidence that variants in PPP1CB cause a rasopathy similar to Noonan syndrome with loose anagen hair. Am. J. Med. Genet. A. 2017;173:565–567. doi: 10.1002/ajmg.a.38056. [DOI] [PubMed] [Google Scholar]
- 67.Shayota B.J., Zhang C., Shypailo R.J., Mazzeu J.F., Carvalho C.M.B., Sutton V.R. Characterization of the Robinow syndrome skeletal phenotype, bone micro-architecture, and genotype-phenotype correlations with the osteosclerotic form. Am. J. Med. Genet. A. 2020;182:2632–2640. doi: 10.1002/ajmg.a.61843. [DOI] [PubMed] [Google Scholar]
- 68.Ishitani T., Kishida S., Hyodo-Miura J., Ueno N., Yasuda J., Waterman M., Shibuya H., Moon R.T., Ninomiya-Tsuji J., Matsumoto K. The TAK1-NLK mitogen-activated protein kinase cascade functions in the Wnt-5a/Ca(2+) pathway to antagonize Wnt/beta-catenin signaling. Mol. Cell. Biol. 2003;23:131–139. doi: 10.1128/MCB.23.1.131-139.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Gordon B.L., Champaigne N.L., Rogers R.C., Frias J.L., Leroy J.G. Long-term observation of a patient with dominant omodysplasia. Am. J. Med. Genet. A. 2014;164A:1234–1238. doi: 10.1002/ajmg.a.36408. [DOI] [PubMed] [Google Scholar]
- 70.Maroteaux P., Sauvegrain J., Chrispin A., Farriaux J.P. Omodysplasia. Am. J. Med. Genet. 1989;32:371–375. doi: 10.1002/ajmg.1320320321. [DOI] [PubMed] [Google Scholar]
- 71.Venditti C.P., Farmer J., Russell K.L., Friedrich C.A., Alter C., Canning D., Whitaker L., Mennuti M.T., Driscoll D.A., Zackai E.H. Omodysplasia: an affected mother and son. Am. J. Med. Genet. 2002;111:169–177. doi: 10.1002/ajmg.10555. [DOI] [PubMed] [Google Scholar]
- 72.Hiraide T., Kaba Yasui H., Kato M., Nakashima M., Saitsu H. A de novo variant in RAC3 causes severe global developmental delay and a middle interhemispheric variant of holoprosencephaly. J. Hum. Genet. 2019;64:1127–1132. doi: 10.1038/s10038-019-0656-7. [DOI] [PubMed] [Google Scholar]
- 73.Costain G., Callewaert B., Gabriel H., Tan T.Y., Walker S., Christodoulou J., Lazar T., Menten B., Orkin J., Sadedin S., et al. De novo missense variants in RAC3 cause a novel neurodevelopmental syndrome. Genet. Med. 2019;21:1021–1026. doi: 10.1038/s41436-018-0323-y. [DOI] [PubMed] [Google Scholar]
- 74.de Curtis I. The Rac3 GTPase in neuronal development, neurodevelopmental disorders, and cancer. Cells. 2019;8:1063. doi: 10.3390/cells8091063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Conlon C.J., Abu-Ghname A., Raghuram A.C., Davis M.J., Guillen D.E., Sutton V.R., Carvalho C.M.B., Maricevich R.S. Craniofacial phenotypes associated with Robinow syndrome. Am. J. Med. Genet. A. 2021;185:3606–3612. doi: 10.1002/ajmg.a.61986. [DOI] [PubMed] [Google Scholar]
- 76.Abu-Ghname A., Trost J., Davis M.J., Sutton V.R., Zhang C., Guillen D.E., Carvalho C.M.B., Maricevich R.S. Extremity anomalies associated with Robinow syndrome. Am. J. Med. Genet. A. 2021;185:3584–3592. doi: 10.1002/ajmg.a.61884. [DOI] [PubMed] [Google Scholar]
- 77.Jenkins Z.A., van Kogelenberg M., Morgan T., Jeffs A., Fukuzawa R., Pearl E., Thaller C., Hing A.V., Porteous M.E., Garcia-Minaur S., et al. Germline mutations in WTX cause a sclerosing skeletal dysplasia but do not predispose to tumorigenesis. Nat. Genet. 2009;41:95–100. doi: 10.1038/ng.270. [DOI] [PubMed] [Google Scholar]
- 78.Gao C., Chen Y.G. Dishevelled: the hub of Wnt signaling. Cell. Signal. 2010;22:717–727. doi: 10.1016/j.cellsig.2009.11.021. [DOI] [PubMed] [Google Scholar]
- 79.Lerner U.H., Ohlsson C. The WNT system: background and its role in bone. J. Intern. Med. 2015;277:630–649. doi: 10.1111/joim.12368. [DOI] [PubMed] [Google Scholar]
- 80.Manickam K., McClain M.R., Demmer L.A., Biswas S., Kearney H.M., Malinowski J., et al. Exome and genome sequencing for pediatric patients with congenital anomalies or intellectual disability: an evidence-based clinical guideline of the American College of Medical Genetics and Genomics (ACMG) Genet. Med. 2021;11:2029–2037. doi: 10.1038/s41436-021-01242-6. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The code supporting the current study has not been deposited in a public repository because it is being updated locally to accommodate further analysis, but it is available from the corresponding author on request. The accession numbers for the identified variants in this paper are ClinVar: SCV000256688, SCV001438002, SCV001438003, SCV001438004, SCV001438005, SCV001438006, SCV001438007, SCV001438008, SCV001438009, SCV001438010, SCV001438011, SCV001438012, SCV001438013, SCV001438014, and SCV00143805 and the preliminary identifier SUB9931810. The dbGAP accession number for all exome sequences reported in this paper and with consent for data sharing in controlled-access databases is dbGAP:phs000711.v5.p1.