Abstract
Background:
Some individuals living with obesity may be relatively metabolically healthy, whilst others suffer from multiple conditions that may be linked to adverse metabolic effects or other factors. The extent to which the adverse metabolic component of obesity contributes to disease compared to the non-metabolic components is often uncertain. We aimed to use Mendelian randomisation (MR) and specific genetic variants to separately test the causal roles of higher adiposity with and without its adverse metabolic effects on diseases.
Methods:
We selected 37 chronic diseases associated with obesity and genetic variants associated with different aspects of excess weight. These genetic variants included those associated with metabolically ‘favourable adiposity’ (FA) and ‘unfavourable adiposity’ (UFA) that are both associated with higher adiposity but with opposite effects on metabolic risk. We used these variants and two sample MR to test the effects on the chronic diseases.
Results:
MR identified two sets of diseases. First, 11 conditions where the metabolic effect of higher adiposity is the likely primary cause of the disease. Here, MR with the FA and UFA genetics showed opposing effects on risk of disease: coronary artery disease, peripheral artery disease, hypertension, stroke, type 2 diabetes, polycystic ovary syndrome, heart failure, atrial fibrillation, chronic kidney disease, renal cancer, and gout. Second, 9 conditions where the non-metabolic effects of excess weight (e.g. mechanical effect) are likely a cause. Here, MR with the FA genetics, despite leading to lower metabolic risk, and MR with the UFA genetics, both indicated higher disease risk: osteoarthritis, rheumatoid arthritis, osteoporosis, gastro-oesophageal reflux disease, gallstones, adult-onset asthma, psoriasis, deep vein thrombosis, and venous thromboembolism.
Conclusions:
Our results assist in understanding the consequences of higher adiposity uncoupled from its adverse metabolic effects, including the risks to individuals with high body mass index who may be relatively metabolically healthy.
Funding:
Diabetes UK, UK Medical Research Council, World Cancer Research Fund, National Cancer Institute.
Research organism: Human
Introduction
Obesity is associated with a higher risk of many diseases, notably metabolic conditions such as type 2 diabetes, but many individuals are often relatively metabolically healthy compared to others of similar body mass index (BMI). Whilst these metabolically healthier individuals may be at lower risk of some obesity-related conditions, they may be at risk of conditions that are linked to other aspects of obesity, such as the load-bearing effects. The burden of obesity on individuals and health-care systems is very large, and in the absence of a widely applicable, sustainable treatment or effective public health measures, it is important to understand the disease consequences of obesity, and how they may be best alleviated, in more detail.
To better understand the disease consequences of obesity, many previous studies have used the approach of Mendelian randomisation (MR) (Smith and Ebrahim, 2004). These studies used common genetic variants robustly associated with BMI as proxies for obesity to assess the causal effects of higher BMI on many diseases. MR studies have provided strong evidence that higher BMI leads to osteoarthritis (Tachmazidou et al., 2019), colorectal cancer (Thrift et al., 2015; Suzuki et al., 2021; Bull et al., 2020), and psoriasis (Budu-Aggrey et al., 2019), as well as metabolic conditions such as type 2 diabetes, cardiovascular disease (Hägg et al., 2015), and heart failure (Cheng et al., 2019; Corbin et al., 2016; Fall et al., 2013). Other MR studies indicate that higher BMI may lead to lower risk of some diseases, including postmenopausal breast cancer (Guo et al., 2016) and Parkinson’s disease (Noyce et al., 2017).
Obesity is heterogeneous – for example, for a given BMI, people vary widely in their amount of fat versus fat free mass, predominantly muscle, and their distribution of fat, predominantly subcutaneous versus ectopic and upper versus lower body fat. Even when there is strong evidence of causality, obesity may lead to disease through a variety of mechanisms. Despite many MR studies testing the role of higher BMI in disease, few have attempted to separate and test the different mechanisms that could lead from obesity to disease. Some MR studies have investigated the effects of fat distribution using genetic variants associated with waist-hip ratio (WHR) adjusted for BMI and shown that adverse fat distribution (more upper body, less lower body) leads to higher risk of metabolic disease (Emdin et al., 2017), some cancers (Cornish et al., 2020), and gastro-oesophageal reflux disease (Green et al., 2020).
Previous studies have identified genetic variants associated with more specific measures of adiposity. For example, several studies have characterised variants associated with ‘favourable adiposity’ (FA) or reduced adipose storage capacity using a variety of approaches (Ji et al., 2019; Lotta et al., 2017; Kilpeläinen et al., 2011; Huang et al., 2021). We recently identified 36 FA alleles which are collectively associated with a favourable metabolic profile, higher subcutaneous fat but lower ectopic liver fat (Ji et al., 2019; Martin et al., 2021), resembling a polygenic phenotype opposite to lipodystrophy (Semple et al., 2011). We also identified 38 unfavourable adiposity (UFA) alleles which are associated with higher fat in subcutaneous and visceral adipose tissue, and higher ectopic liver and pancreatic fat (Ji et al., 2019; Martin et al., 2021), resembling monogenic obesity (Supplementary file 1a). We performed MR studies and showed that FA and UFA have opposite causal effects on six metabolic conditions (Martin et al., 2021). While both FA and UFA were associated with higher adiposity, FA was causally associated with lower risk of type 2 diabetes, heart disease, hypertension, stroke, polycystic ovary syndrome, and non-alcoholic fatty liver disease. In contrast, as expected, UFA was associated with higher risk of these conditions. These results confirmed the ability of the two sets of adiposity variants to partially separate out the metabolic from the non-metabolic effects of higher adiposity.
In this study, we aimed to investigate the effects of separate components to higher adiposity on risk of additional metabolic diseases and many non-metabolic diseases. We used genetic variants associated with BMI, body fat percentage, FA, and UFA to understand the components of higher adiposity that are the predominant causes of disease risk. Our findings may give guidance on some obesity-related risks which are not dependent on metabolic consequences, thereby guiding appropriate medical care.
Methods
Study design
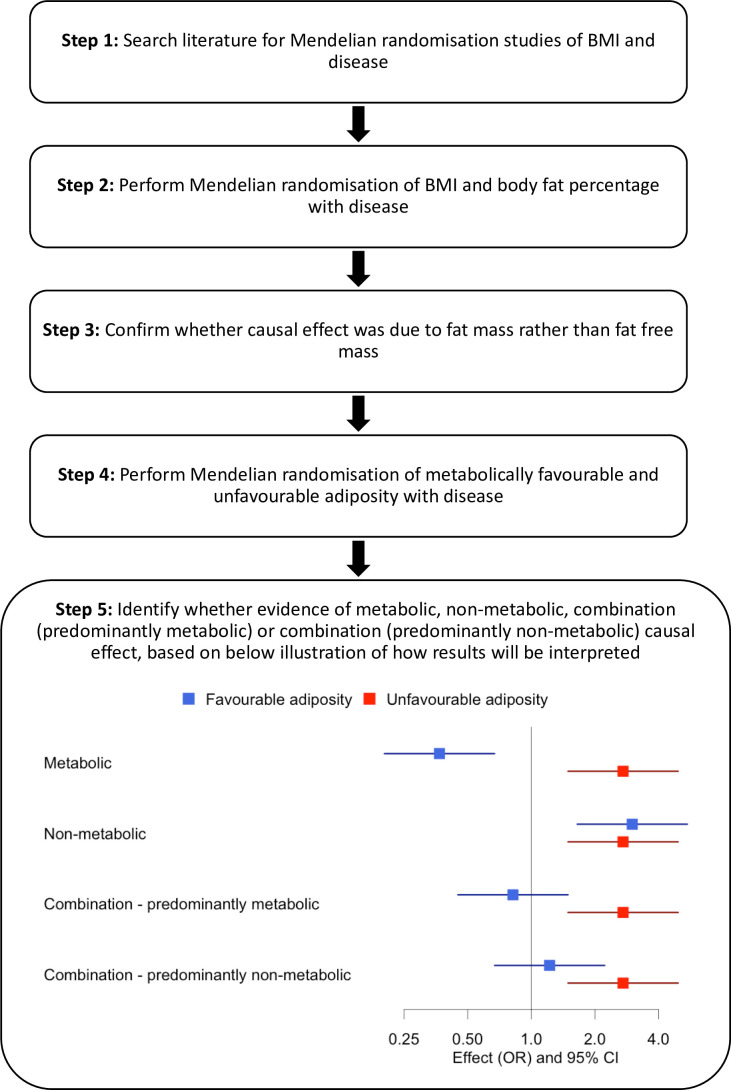
An overview of our approach is shown in Figure 1. First, we identified diseases by performing a literature search of studies that had used MR to assess the consequences of BMI on outcome phenotypes. We used the search terms ‘BMI and Mendelian randomisation’ and ‘BMI and Mendelian randomization’. We identified 37 diseases associated with BMI and for which MR studies had previously been performed (Supplementary file 1b). We included all diseases regardless of the MR result in the published study. Second, we reperformed MR studies using BMI as an exposure. Third, for those diseases where MR indicated higher BMI was causal, we tested the effects of body fat percentage to confirm that the causal effect was due to fat mass rather than fat-free mass. Fourth, for diseases where MR suggested the BMI effect was due to excess adiposity, we used genetic variants more specific to the metabolic and non-metabolic components of higher adiposity to help understand the extent to which these factors influence disease.
Figure 1. Study design.
Data sources
We used three data sources for disease outcomes: (i) published genome-wide association studies (GWAS; Okada et al., 2014; Nikpay et al., 2015; Jones et al., 2017; Michailidou et al., 2017; Phelan et al., 2017; Scelo et al., 2017; Tsoi et al., 2017; Day et al., 2018; Mahajan et al., 2018; Malik et al., 2018; O’Mara et al., 2018; Roselli et al., 2018; Schumacher et al., 2018; Wray et al., 2018; An et al., 2019; Ferreira et al., 2019; Huyghe et al., 2019; Jansen et al., 2019; Kunkle et al., 2019; Law et al., 2019; Lindström et al., 2019; Morris et al., 2019; Nalls et al., 2019; Shah et al., 2019; Tachmazidou et al., 2019; Tin et al., 2019; Wuttke et al., 2019; Huyghe et al., 2021) and (ii) FinnGen (FinnGen, 2021) as our main results, and (iii) UK Biobank (RRID:SCR_012815; Collins, 2012) as additional validation. FinnGen is a cohort of 176,899 individuals with linked medical records. UK Biobank is a population cohort of >500,000 individuals aged 37–73 years recruited between 2006 and 2010 from across the UK. For the 37 identified diseases, 25 had summary GWAS data available from both a published GWAS consortium and FinnGen, and 12 diseases had GWAS summary data available in FinnGen only. In addition, data from 31 of the 37 diseases were available in the UK Biobank. No GWAS data were available for Barrett’s oesophagus, but we included gastro-oesophageal reflux. The characteristics of the studies and measures, disease outcomes, and the definition of cases and controls are described in Supplementary file 1ci–iii.
GWAS of UK Biobank participants
For the GWAS of 31 diseases available in UK Biobank, we used a linear mixed model implemented in BOLT-LMM to account for population structure and relatedness (Loh et al., 2015). We used age, sex, genotyping platform, study centre, and the first five principal components as covariates in the model.
Genetic variants
We used four sets of genetic variants as proxies of four exposures (Supplementary file 1d).
Body mass index
In the broadest category, we used a set of 73 variants independently associated with BMI at genome-wide significance (p<5 × 10–8). These variants were identified in the GIANT consortium of up to 339,224 individuals of European ancestry (Locke et al., 2015).
Body fat percentage
We used 696 variants from a GWAS in the UK Biobank (Martin et al., 2021). We used bio-impedance measures of body fat % taken by the Tanita BC-418MA body composition analyser in 442,278 individuals of European ancestry.
The BMI and body fat percentage variants were partially overlapping (n = 5 variants), but we used exposure-trait-specific weights for each variant.
FA variants
There are 36 FA variants (Martin et al., 2021). These variants were identified in two steps. First, they were associated (at p<5 × 10–8) with body fat percentage and a composite metabolic phenotype consisting of body fat percentage, HDL-cholesterol, triglycerides, SHBG, alanine transaminase, and aspartate transaminase. Second, in a k-means clustering approach (a hard clustering approach) (Martin et al., 2021), they formed a cluster of variants that were collectively associated with higher HDL-cholesterol, higher SHBG, and lower triglycerides and liver enzymes – resembling a phenotype opposite to lipodystrophy.
UFA variants
There are 38 UFA variants (Martin et al., 2021). These variants were identified in two steps. First, they were associated (at p<5 × 10–8) with body fat percentage and a composite metabolic phenotype as detailed above. Second, in a k-means clustering approach (Martin et al., 2021), they formed a cluster of variants that were collectively associated with lower HDL-cholesterol, lower SHBG, and higher triglycerides and liver enzymes - resembling monogenic obesity.
Mendelian randomisation
We investigated the causal associations between the four exposures (BMI, body fat percentage, FA, and UFA) and 37 disease outcomes by performing two-sample MR analysis (Pierce and Burgess, 2013). We used the inverse-variance weighted (IVW) approach as our main analysis, and MR-Egger and weighted median as sensitivity analyses in order to detect and partially account for unidentified pleiotropy of our genetic instruments. For BMI, we used effect size estimates from the GWAS of BMI (Locke et al., 2015), and for body fat percentage, FA, and UFA, we used effect size estimates from the GWAS of body fat percentage (442,278 European ancestry individuals from the UK Biobank study) (Ji et al., 2019).
To estimate the effects of variants on our disease outcomes, we used two main sources of data: FinnGen GWAS summary results and published GWAS of the same diseases (Supplementary file 1ci–ii). We performed MR within each data source and then meta-analysed the results across the two datasets using a random-effects model with the R package metafor (RRID:SCR_003450; Viechtbauer, 2010), where the data was available in both. For one published GWAS (the GECCO consortium), we only had information for FA and UFA variants. To provide further MR evidence, we used a third source of disease data – disease status in the UK Biobank (Supplementary file 1ciii). We ran the same models but did not meta-analyse with published GWAS and FinnGen because most of the body fat percentage, FA, and UFA variants were identified in the UK Biobank.
We obtained heterogeneity Q statistics for each IVW MR and MR-Egger, and I2 statistics for each MR-Egger analysis using the MendelianRandomization R package (Yavorska and Burgess, 2017). All statistical analyses were conducted using R software (R Development Core Team, 2020). Given the number of tests performed, we used a Benjamini–Hochberg false discovery rate (FDR) procedure and an FDR of 0.1 to define meaningful results for each of the four exposures (Benjamini and Hochberg, 1995).
Results
We identified 37 diseases as associated with obesity and for which MR studies had previously been performed. Of these 37, 5 metabolic conditions were part of our previous study that validated the use of FA and UFA genetic variants as a way of partially separating the metabolic from non-metabolic components of higher adiposity (Martin et al., 2021). Once we had tested BMI and body fat percentage, we further characterised the likely causal component of higher adiposity using FA and UFA variants as follows (Figure 1, step 5): (i) diseases with evidence that the metabolic effect of higher adiposity is causal. Here, MR using the UFA genetic variants indicated that higher adiposity with its adverse metabolic consequences was causal to disease, whilst MR using the FA genetic variants indicated that higher adiposity with favourable metabolic effects was protective (at FDR 0.1). (ii) Diseases with evidence that there is a non-metabolic causal effect (e.g. mechanical effect, psychological/adverse social effect). Here, MR using the FA genetic variants indicated that higher adiposity without its adverse metabolic consequences was likely contributing to the disease, as well as the MR using the UFA genetic variants. (iii) Diseases with evidence that there is a combination of causal effects but with a predominantly metabolic component. Here, MR using the UFA genetic variants indicated that higher adiposity with its adverse metabolic consequences was causal to disease, and MR using the FA genetic variants was directionally consistent with higher adiposity with favourable metabolic effects being protective but FDR > 0.1. (iv) Diseases with evidence that there is a combination of causal effects but with a predominantly non-metabolic component. Here, MR using the UFA genetic variants indicated that higher adiposity without its adverse metabolic consequences was likely contributing to the disease, and MR of the FA genetic variants was directionally consistent with this but FDR > 0.1.
We grouped these disease outcomes into seven major categories – cardiovascular and metabolic conditions, musculoskeletal, gastrointestinal, nervous, integumentary and respiratory systems, and cancer. MR analysis of five conditions (coronary artery disease, hypertension, stroke, type 2 diabetes, and polycystic ovary syndrome) was part of our previous study (Martin et al., 2021). We focused on the MR of body fat percentage if a causal effect of BMI was indicated, and the MR of FA and UFA if a causal effect of BMI and body fat percentage was indicated, but have presented all results in Supplementary file 1e for completeness. Where random-effects meta-analyses were performed, the heterogeneity statistics are given in Supplementary file 1f.
(i) Diseases with evidence that the metabolic effect of higher adiposity is causal
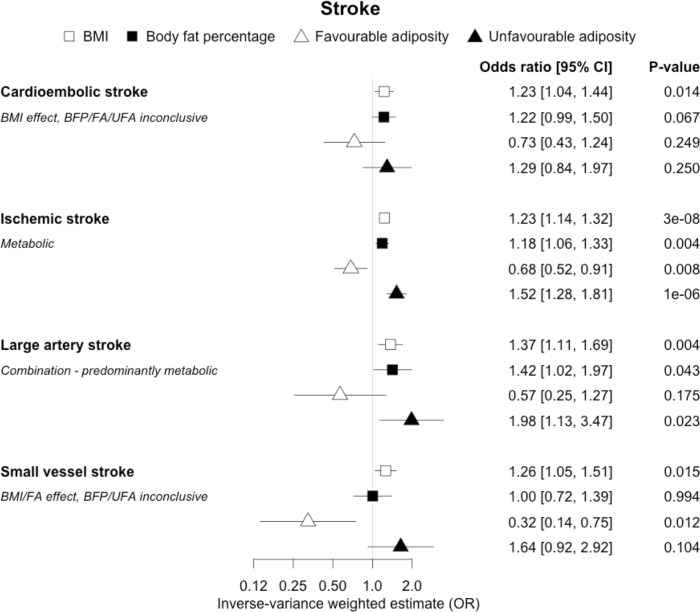
When comparing the MR analyses for FA and UFA, our results provided evidence that the metabolic effect of higher adiposity is contributing causally to coronary artery disease, peripheral artery disease, hypertension, stroke, type 2 diabetes, and gout (Figures 2—12, Supplementary file 1e). For stroke, our results were consistent when using sub-types of the condition (Figure 3—figure supplement 1, Supplementary file 1g). Our results also indicated that the metabolic effect of higher adiposity is causal to chronic kidney disease, although the results from BMI and body fat percentage were less conclusive (Figure 3).
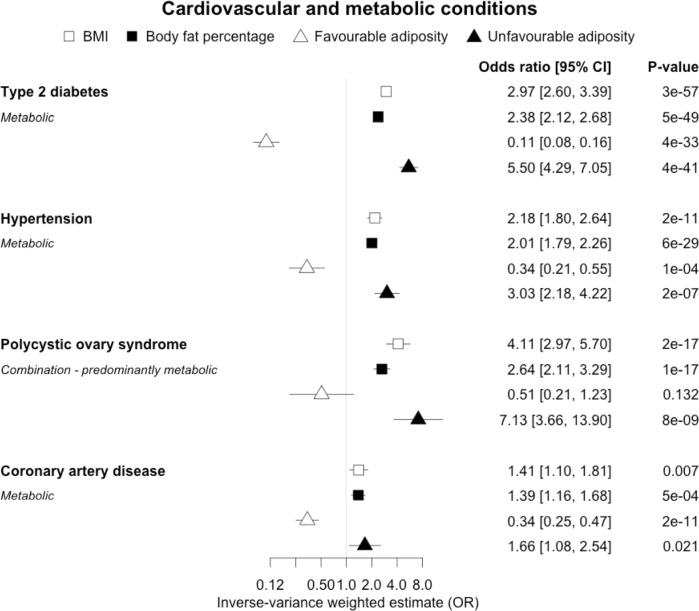
Figure 2. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on type 2 diabetes, hypertension, polycystic ovary syndrome and coronary artery disease.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
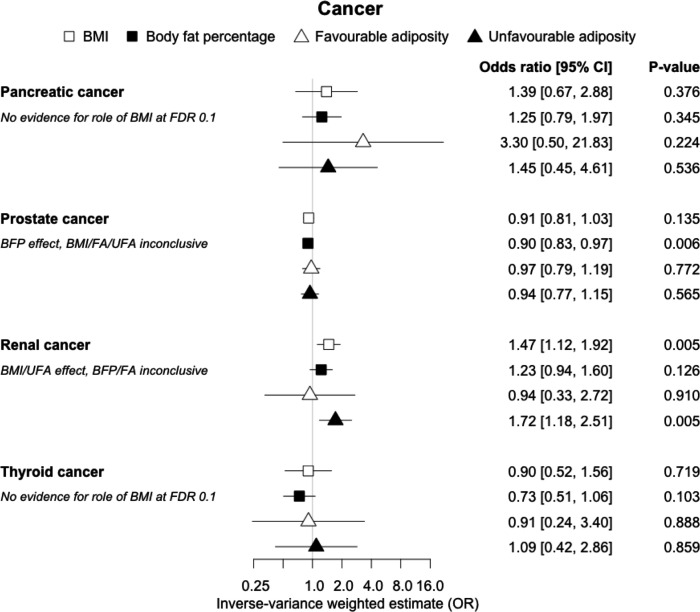
Figure 12. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on pancreatic, prostate, renal and thyroid cancer.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
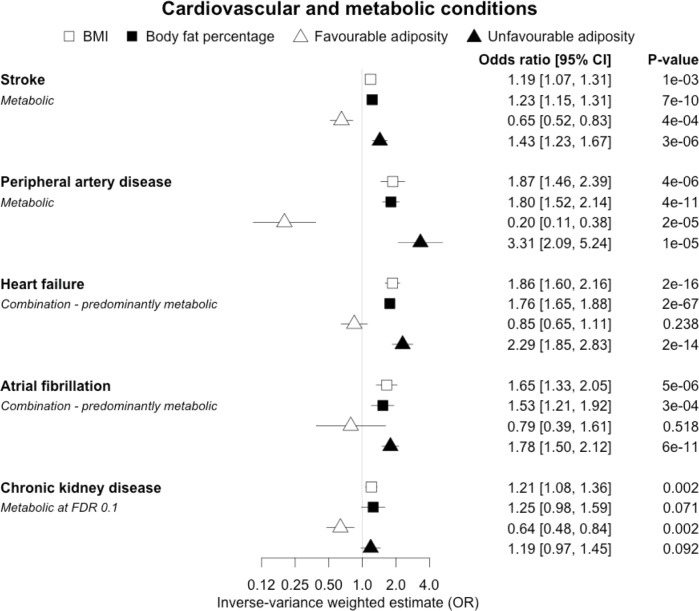
Figure 3. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on stroke, peripheral artery disease, heart failure, atrial fibrillation and chronic kidney disease.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
Figure 3—figure supplement 1. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on sub-types of stroke.
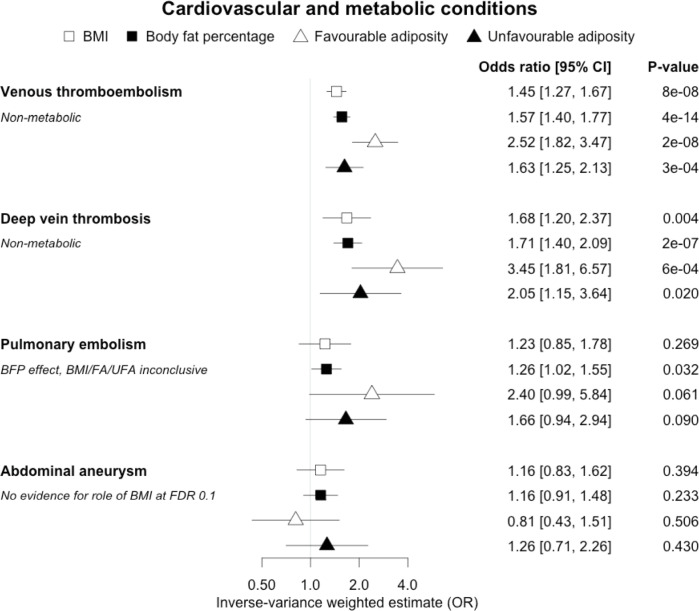
Figure 4. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on venous thromboembolism, deep vein thrombosis, pulmonary embolism and abdominal aneurysm.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
Figure 6. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on gallstones and gastro-oesophageal reflux disease.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
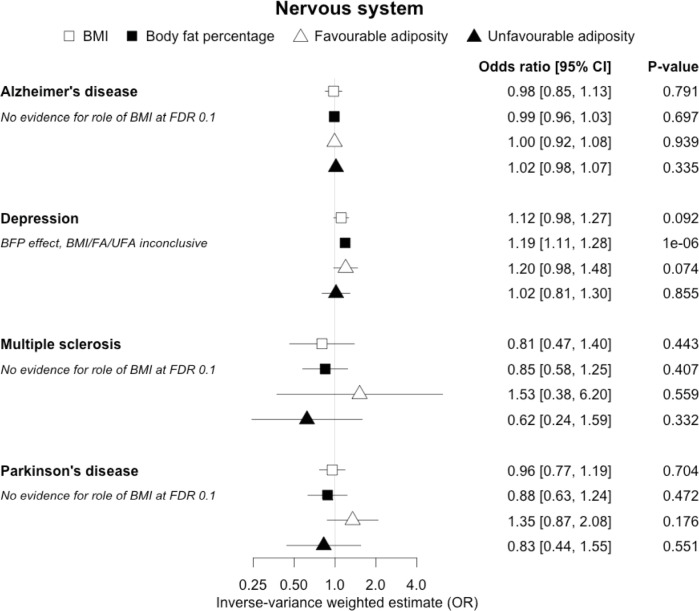
Figure 7. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on Alzheimer’s disease, depression, multiple sclerosis and Parkinson’s disease.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
Figure 8. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on psoriasis.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
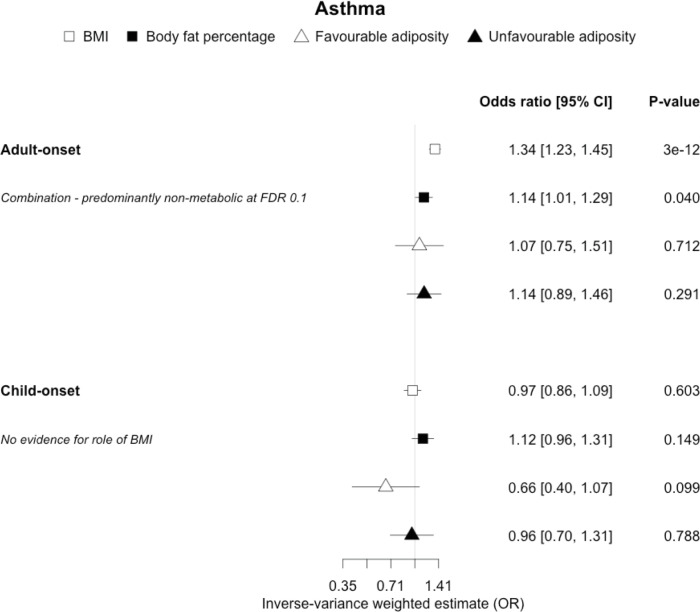
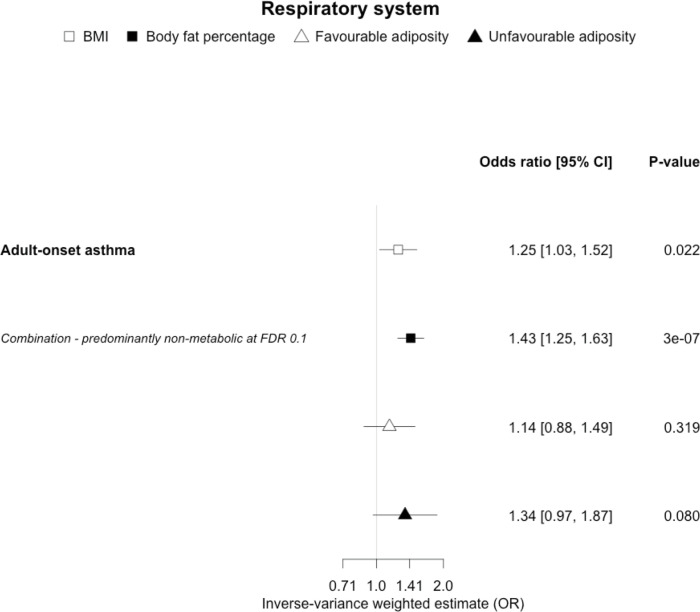
Figure 9. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on adult-onset asthma.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
Figure 9—figure supplement 1. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on sub-types of asthma.
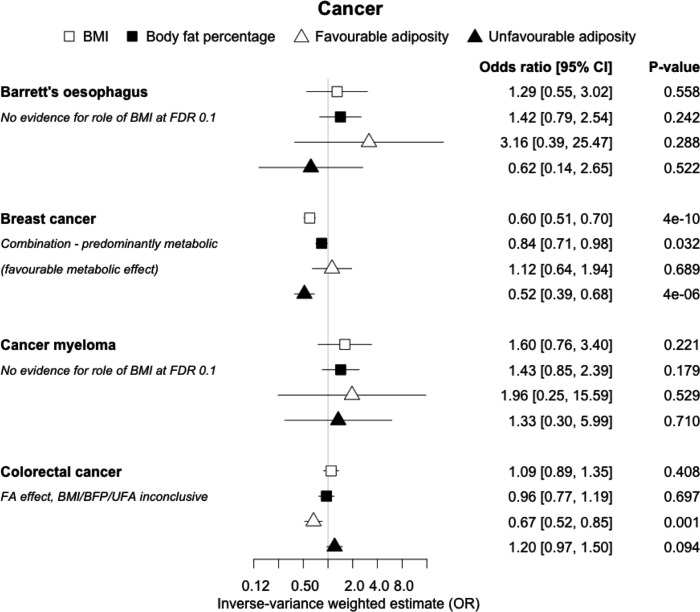
Figure 10. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on Barrett’s oesophagus, breast cancer, cancer myeloma and colorectal cancer.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
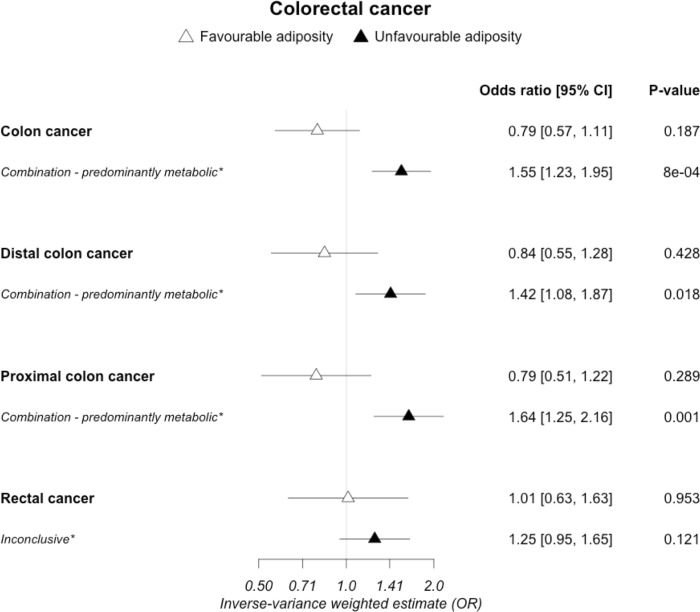
Figure 10—figure supplement 1. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on sub-types of colorectal cancer.
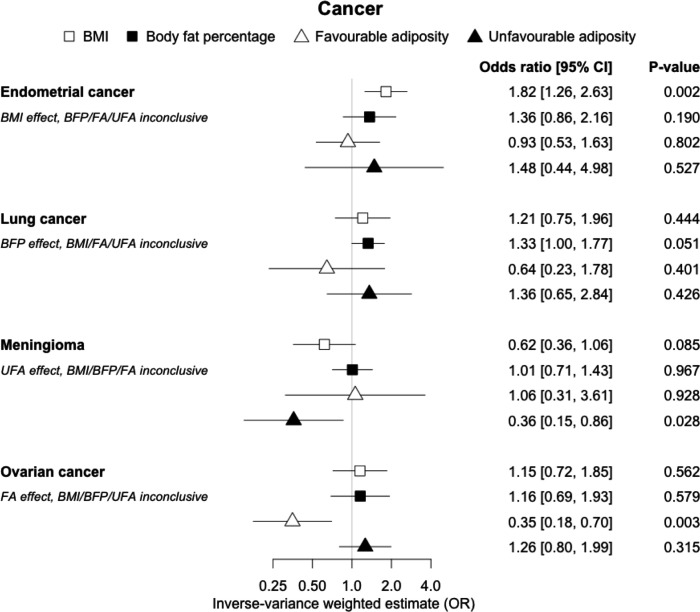
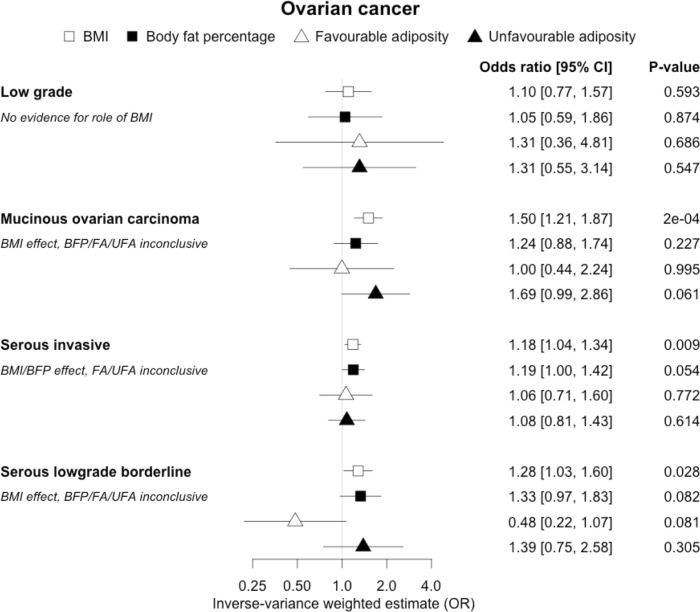
Figure 11. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on endometrial and lung cancer, meningioma and ovarian cancer.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
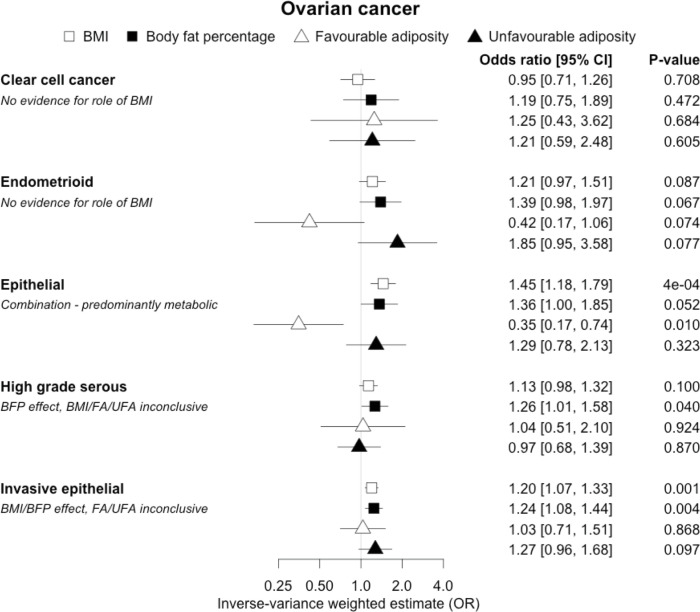
Figure 11—figure supplement 1. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on 5 sub-types of ovarian cancer.
Figure 11—figure supplement 2. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on 4 sub-types of ovarian cancer.
(ii) Diseases with evidence that there is a non-metabolic causal effect
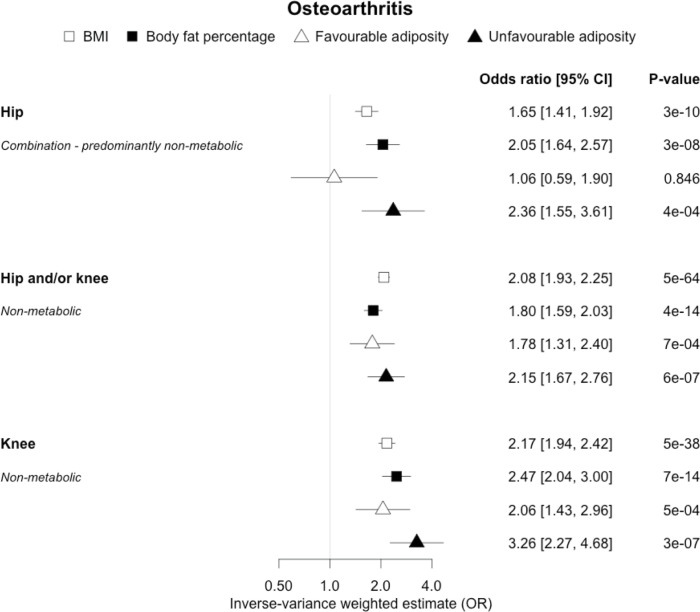
When comparing the MR analyses for FA and UFA, our results provided evidence that some non-metabolic effect of higher adiposity is contributing causally to venous thromboembolism, deep vein thrombosis, osteoarthritis, and rheumatoid arthritis (Figures 2—12, Supplementary file 1e). For osteoarthritis, our results were consistent when using sub-types of the condition (Figure 5—figure supplement 1, Supplementary file 1g).
(iii) Diseases with evidence that there is a combination of causal effects but with a predominantly metabolic component
When comparing the MR analyses for FA and UFA, our results provided evidence that the metabolic effect of higher adiposity is the predominate cause of the link between higher BMI and polycystic ovary syndrome, heart failure, and atrial fibrillation. Our results also provided evidence that the metabolic effect of higher adiposity is the predominate cause of the link between higher BMI and a reduced risk of breast cancer and higher risk of renal cancer, although the results from body fat percentage were less conclusive (Figures 2—12, Supplementary file 1e).
(iv) Diseases with evidence that there is a combination of causal effects but with a predominantly non-metabolic component
When comparing the MR analyses for FA and UFA, our results suggested that some non-metabolic effect of higher adiposity is the predominant cause of the link between higher BMI and gallstones, gastro-oesophageal reflux disease, adult-onset asthma, and psoriasis (Figures 2—12, Supplementary file 1e). Our results also indicated that some non-metabolic effect of higher adiposity is causal to osteoporosis, although the results from BMI were less conclusive (Figure 5). Our results found no evidence (at p<0.05) of an effect of BMI or adiposity on child-onset asthma (Figure 9—figure supplement 1, Supplementary file 1g).
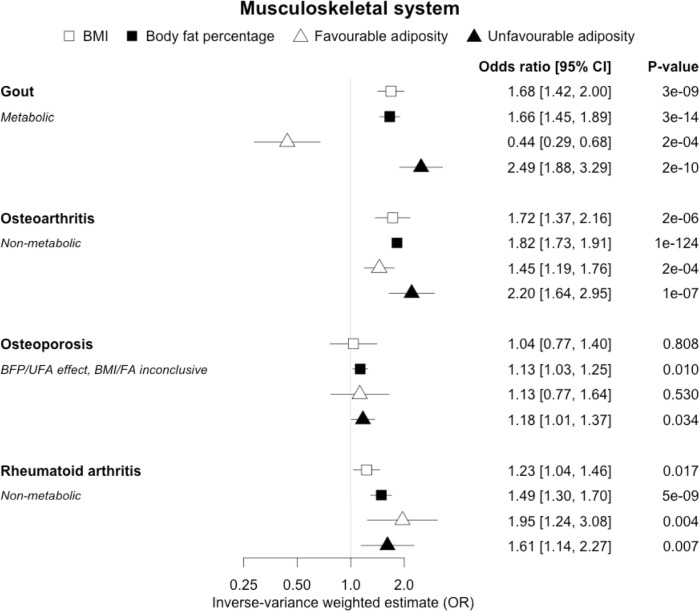
Figure 5. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on gout, osteoarthritis, osteoporosis and rheumatoid arthritis.
The error bars represent the 95% confidence intervals of the IVW estimates in odds ratio per standard deviation change in genetically determined BMI, body fat percentage, FA and UFA. Italics give our best interpretation of the data using the FDR 0.1 results.
Figure 5—figure supplement 1. The inverse-variance weighted (IVW) two-sample MR analysis/meta-analysis of the effects of body mass index (BMI), body fat percentage (BFP), “favourable adiposity” (FA) and “unfavourable adiposity” (UFA) on sub-types of osteoarthritis.
All other disease outcomes
Fifteen disease outcomes did not fit the criteria for definitions i–iv. For five of these conditions, our MR results indicated a causal effect of higher BMI or adiposity, but results from FA and UFA were inconclusive: pulmonary embolism, depression, endometrial cancer, lung cancer, and prostate cancer (Figures 2—12, Supplementary file 1e). Additionally, we identified some evidence of a metabolic effect of higher adiposity with colorectal and ovarian cancer, with the MR of FA indicating lower odds of colorectal (0.67 [0.52, 0.85]) and ovarian (0.35 [0.18, 0.70]) cancers, but MR of UFA was consistent with the null (p>0.05). For colorectal and ovarian cancer, our results were consistent when using sub-types of the conditions (Figure 10—figure supplement 1, Figure 11—figure supplements 1 and 2, Supplementary file 1g).
Sensitivity analyses
Out of 82 disease outcomes (including subtypes), weighted median MR results were directionally consistent with IVW analysis for 75 diseases for BMI and 73 for body fat percentage, with 33 and 47 of these having p<0.05, respectively. For FA and UFA, where sub-type colorectal cancer data was available, the total number of diseases was 87, and 76 were directionally consistent for both exposures, with 22 and 39 having p<0.05, respectively.
MR-Egger results were broadly consistent with the primary IVW MR results, indicating that pleiotropy (variants acting on the outcomes through more than one mechanism) appears to have had limited effect on our results. MR-Egger results were directionally consistent with IVW for 71 diseases for BMI and 70 for body fat percentage, with 25 and 38 of these having p<0.05, respectively. For FA and UFA, MR-Egger was directionally consistent for 60 and 67 diseases, with 6 and 15 having p<0.05, respectively (Supplementary file 1g). Of the 31 diseases available in the UK Biobank, the IVW analysis of these was directionally consistent with the FinnGen and/or published GWAS analysis for 28, 27, 24, and 27 traits for BMI, body fat percentage, FA, and UFA, respectively (Supplementary file 1h). Of these, 18, 21, 9, and 16 had p<0.05, respectively.
Discussion
We used a genetic approach to understand the role of higher adiposity uncoupled from its adverse metabolic effects in mechanisms linking obesity to higher risk of disease. We first used MR to provide evidence that higher BMI was causally associated with 21 diseases, broadly consistent with those from previous studies. For the majority (17) of these diseases, our results indicated that the BMI effect was predominantly due to excess adiposity rather than a non-fat mass component to BMI. We then used a more specific approach to test the separate roles of higher adiposity with and without its adverse metabolic effects. We provided genetic evidence that the adverse metabolic consequences of higher BMI lead to coronary artery disease, peripheral artery disease, hypertension, stroke, type 2 diabetes, polycystic ovary syndrome, heart failure, atrial fibrillation, chronic kidney disease, renal cancer, and gout, and the adverse non-metabolic consequences of higher BMI likely contribute to osteoarthritis, rheumatoid arthritis, osteoporosis, gastro-oesophageal reflux disease, gallstones, adult-onset asthma, psoriasis, deep vein thrombosis, and venous thromboembolism.
Understanding the reasons why obesity leads to disease is important in order to better advise health professionals and patients of health risks linked to obesity, whether or not they show metabolic derangements. Many previous studies have used an MR approach to support a causal role of higher BMI in disease, but here we attempted to systematically test many conditions and the role of separate components of higher BMI. We discuss some of the more notable, and potentially clinically important, results below.
Cardiometabolic diseases
Previous studies, including those using MR, have shown that higher BMI leads to many cardiometabolic diseases (Larsson et al., 2020; Riaz et al., 2018; Xu et al., 2020), but our results provide additional insight into the likely mechanisms. In addition to the previously established opposing effects of metabolically FA and UFA for coronary artery disease, stroke, hypertension, and type 2 diabetes (Martin et al., 2021), our results confirmed similarly strong metabolic components to peripheral artery disease and chronic kidney disease. These results are consistent with the well-established adverse metabolic effects of higher BMI on these diseases (contributing to atherosclerotic effects or linked to specific haemodynamic impacts) (Sattar and McGuire, 2018). For two further cardiovascular conditions, heart failure and atrial fibrillation, the results were less certain. For these two conditions, the evidence of a predominantly metabolic effect of higher BMI was very clear – with the MR of UFA consistent with effects at least as strong as those for coronary artery disease. However, in contrast to the results for coronary artery disease, the MR of FA was consistent with no effect. This comparison between the effects of FA and UFA may indicate that there is a partial mechanical, or other non-metabolic component, as well as metabolic effect, perhaps mediated by excess weight of any type placing extra strain on the heart.
In contrast to the results for most of the cardiometabolic diseases, our MR analyses provided evidence for a likely non-metabolic component mediating the effect of higher BMI on venous thromboembolism and deep vein thrombosis (two closely related conditions). This finding is clinically important as it suggests that treating metabolic risk factors associated with obesity without changing weight may not reduce the risk of deep vein thrombosis in individuals with obesity. Possible mechanisms could include higher intra‐abdominal pressure (due to excess fat) and slower blood circulation in the lower limbs (due to a more sedentary lifestyle secondary to obesity, or mechanical occlusion of veins) promoting clot initiation and formation (Lorenzet et al., 2012).
Musculoskeletal diseases
We observed clear differences for the role of higher BMI in different musculoskeletal diseases. For gout, opposing effects of FA and UFA clearly indicated a metabolic effect. Gout is a form of inflammatory arthritis caused by the deposition of urate crystals within the joints (Dalbeth et al., 2016). Weight loss from bariatric surgery is associated with lower serum uric acid and lower risk of gout (Maglio et al., 2017). A previous MR study showed that overall obesity, but not the central location of fat, increased the risk of gout (Larsson et al., 2018). The protective effect of FA could be due to improved insulin sensitivity leading to less insulin-enhanced reabsorption of organic anions such as urate (Choi et al., 2005). In contrast to gout, our MR analysis provided evidence that a non-metabolic effect of higher adiposity is a likely cause of osteoarthritis and rheumatoid arthritis – with both FA and UFA leading to disease. For osteoarthritis, the effect of UFA was stronger than that of FA, indicating both a metabolic and non-metabolic component. This is consistent with a causal association between higher adiposity and higher risk of osteoarthritis in non-weight-bearing joints including hands (Reyes et al., 2016). For rheumatoid arthritis, the effects of FA and UFA were similar, suggesting the non-metabolic effect accentuating, or more readily unmasking, the autoimmune background risk, as the key BMI-related factor, although the confidence intervals were wider than those for osteoarthritis. The UFA variants may potentially influence these conditions by load-bearing mechanisms, and tissue enrichment analysis for the FA and UFA variants previously found that FA and UFA loci are enriched for genes expressed in adipocytes and adipose tissue, and mesenchymal stem cells, respectively (Martin et al., 2021). For osteoporosis, we did not replicate the previous finding of a causal association between higher BMI and risk of osteoporosis (estimated by bone mineral density; Song et al., 2020); however, we observed a causal association between higher body fat percentage and a higher risk of osteoporosis with consistent risk increasing effects of both FA and UFA. This finding adds to the complex relationship between higher BMI and osteoporosis, where higher BMI at earlier ages may increase bone accrual, but in later years results in adverse effects.
Gastrointestinal diseases
We observed differences in the effects of BMI when comparing the two gastrointestinal diseases, although the results are less conclusive than those for the musculoskeletal conditions. Here, our results were consistent with a predominantly non-metabolic effect contributing to the association between higher BMI and higher risk of gallstones. Higher BMI has been shown to be causally associated with higher risk of gallstones (Yuan et al., 2021). There are several possible mechanisms that could explain how higher BMI without its adverse metabolic effects could increase the risk of gallstones. These could include a sedentary lifestyle and gallbladder hypomotility secondary to increased abdominal fat mass (Mathus-Vliegen et al., 2004). Metabolic mechanisms could include hepatic de novo cholesterol synthesis (Ståhlberg et al., 1997; Cruz-Monserrate et al., 2016). For gastro-oesophageal reflux, the consistent direction and effect sizes of higher FA and UFA indicate a non-metabolic component, an effect that may be mechanical and better explained by higher central adiposity rather than overall BMI (Green et al., 2020).
Other diseases
For most of the other diseases tested, it was difficult to draw firm conclusions about the role of metabolically FA and UFA. For some diseases, this was in part due to the lack of MR evidence for a role of any form of higher BMI. For example, our MR analyses provided no evidence for the role of higher BMI in the neurodegenerative diseases Alzheimer’s disease, multiple sclerosis, and Parkinson’s. These results are consistent with some but not all previous studies. For example, higher BMI is listed as a key risk factor for Alzheimer’s disease (Livingston et al., 2020), although with little evidence of causality, including MR studies that failed to show an effect (Larsson et al., 2017; Nordestgaard et al., 2017). In contrast to our results, recent MR studies have indicated that higher BMI is protective of Parkinson’s disease (Noyce et al., 2017) and causally associated with higher risk of multiple sclerosis (Mokry et al., 2016). For the inflammatory skin disorder psoriasis, our results indicated that both higher BMI and higher body fat percentage are causally associated with higher risk, but determining the underlying mechanism from the MR of FA and UFA was difficult. Higher BMI is a known cause of psoriasis (Budu-Aggrey et al., 2019; Iskandar et al., 2015) and weight loss is a recommended treatment (Iskandar et al., 2015). It is possible that both metabolic and non-metabolic pathways are driving the risk. The non-metabolic pathways could include inflammation which is one of the possible causal mechanisms (Sbidian et al., 2017; Dowlatshahi et al., 2013). Further work is required to understand if psoriasis could be effectively treated by targeting the metabolic factors alone, or whether only weight loss will benefit such patients. For cancers, our results do not provide any clear additional insight into the likely mechanisms, with potentially stronger effects for BMI and UFA compared to body fat percentage in some analyses hard to explain biologically. The reasons why higher BMI is associated with cancers is uncertain, although several MR studies indicate that the association with many is causal (Mariosa et al., 2019; Vincent and Yaghootkar, 2020), and that central adiposity may play a role (Jarvis et al., 2016). Exposure to higher insulin levels is a plausible mechanism, and some studies have used MR to test insulin directly (Nead et al., 2015; Shu et al., 2019; Carreras-Torres et al., 2017b; Carreras-Torres et al., 2017a; Johansson et al., 2019). Our MR analysis reproduced the previous finding between higher adiposity and higher risk of endometrial cancer (Painter et al., 2016) and renal cell carcinoma (Johansson et al., 2019), and lower risk of breast cancer (Guo et al., 2016; Shu et al., 2019). In contrast to previous MR studies showing a causal link between higher BMI and higher risk of prostate cancer (Kazmi et al., 2020; Davies et al., 2015), we identified a causal association between higher body fat percentage but lower risk of prostate cancer. The relationship between higher BMI and risk of breast cancer is complicated, with MR studies indicating that higher BMI is protective of postmenopausal breast cancer (Gao et al., 2016). This contrasts with the epidemiological associations but could be explained by effects of childhood BMI (Richardson et al., 2020).
Strengths and limitations
Our study had a number of limitations. First, we do not know all of the potential effects of the FA and UFA genetic variants on intermediary mechanisms. For example, the inflammatory profile of the FA variants needs further characterisation. However, the consistent association of the FA genetic variants with lower risk of a wide range of metabolic conditions – from type 2 diabetes where insulin resistance predominates, to stroke where atherosclerotic and blood pressure mechanisms predominate – indicates that these variants collectively represent a profile of higher adiposity and favourable metabolic factors. Second, for some diseases, we may have not had sufficient power to detect an effect of BMI or to separate the effects, and this could explain some of the null findings, especially for conditions where we might have expected an effect, such as pulmonary embolism and aortic aneurysm, but there were smaller numbers of cases available. Third, in some situations it was harder to interpret the results from the MR FA and UFA analyses, especially when one appeared to show an effect and the other did not. One possibility is that some diseases are a combination of both non-metabolic and metabolic effects. Osteoarthritis was the best example of this potential scenario because both FA and UFA increased the risk of disease, but UFA to a greater extent. However, for other diseases, it could be hard to detect a combined effect because the MR with FA could be protective (if metabolic effects predominate), increase risk (if non-metabolic effects predominate), or null (if the two have similar effects). Finally, we used an FDR of 0.1 as a guide to discussing meaningful results. We observed 21 out of the 37 outcome diseases reaching an FDR of 0.1 (based on the Benjamini–Hochberg procedure) for BMI, and 19, 11, and 20 out of the 21 diseases causally associated with BMI reaching this FDR for body fat percentage, FA, and UFA, respectively. Equivalent numbers for an FDR of 0.05 were 21, 17, 11, and 17. Excluding the five metabolic conditions used in our previous study (which were all causally associated with BMI), these results are 16, 14, 7, and 15 for an FDR of 0.1, and 16, 12, 7, and 12 for an FDR of 0.05. In addition to correcting for multiple tests, we noted that 74 of the 37 × 4 MR tests reached a p-value of <0.05 when we would only expect 7 by chance, suggesting many of the tests that did not reach a strict Bonferroni p<0.05 were meaningful.
In summary, we have used a genetic approach to test the separate roles of higher adiposity with and without its adverse metabolic effects. These results emphasize that many people in the community who are of higher BMI are at risk of multiple chronic conditions that can severely impair their quality of life or cause morbidity or mortality, even if their metabolic parameters appear relatively normal.
Acknowledgements
This research has been conducted using the UK Biobank Resource under Application Number ‘9072’ and ‘9055’. We acknowledge the use of the University of Exeter High-Performance Computing (HPC) facility in carrying out this work. We acknowledge use of high-performance computing funded by an MRC Clinical Research Infrastructure award (MRC Grant: MR/M008924/1).
ASTERISK: We are very grateful to Dr. Bruno Buecher without whom this project would not have existed. We also thank all those who agreed to participate in this study, including the patients and the healthy control persons, as well as all the physicians, technicians and students.
CCFR: The Colon CFR graciously thanks the generous contributions of their study participants, dedication of study staff, and the financial support from the U.S. National Cancer Institute, without which this important registry would not exist. We would like to thank the study participants and staff of the Seattle Colon Cancer Family Registry and the Hormones and Colon Cancer study (CORE Studies).
CLUE II: We thank the participants of Clue II and appreciate the continued efforts of the staff at the Johns Hopkins George W. Comstock Center for Public Health Research and Prevention in the conduct of the Clue II Cohort Study.
COLON and NQplus: We would like to thank the COLON and NQplus investigators at Wageningen University & Research and the involved clinicians in the participating hospitals.
CORSA: We kindly thank all those who contributed to the screening project Burgenland against CRC. Furthermore, we are grateful to Doris Mejri and Monika Hunjadi for laboratory assistance.
CPS-II: We thank the CPS-II participants and Study Management Group for their invaluable contributions to this research. We would also like to acknowledge the contribution to this study from central cancer registries supported through the Centers for Disease Control and Prevention National Program of Cancer Registries, and cancer registries supported by the National Cancer Institute Surveillance Epidemiology and End Results program.
Czech Republic CCS: We are thankful to all clinicians in major hospitals in the Czech Republic, without whom the study would not be practicable. We are also sincerely grateful to all patients participating in this study.
DACHS: We thank all participants and cooperating clinicians, and Ute Handte-Daub, Utz Benscheid, Muhabbet Celik, and Ursula Eilber for excellent technical assistance.
EDRN: We acknowledge all the following contributors to the development of the resource: University of Pittsburgh School of Medicine, Department of Gastroenterology, Hepatology and Nutrition: Lynda Dzubinski; University of Pittsburgh School of Medicine, Department of Pathology: Michelle Bisceglia; and University of Pittsburgh School of Medicine, Department of Biomedical Informatics.
EPIC: Where authors are identified as personnel of the International Agency for Research on Cancer/World Health Organization, the authors alone are responsible for the views expressed in this article and they do not necessarily represent the decisions, policy or views of the International Agency for Research on Cancer/World Health Organization.
The EPIC-Norfolk study: we are grateful to all the participants who have been part of the project and to the many members of the study teams at the University of Cambridge who have enabled this research.
EPICOLON: We are sincerely grateful to all patients participating in this study who were recruited as part of the EPICOLON project. We acknowledge the Spanish National DNA Bank, Biobank of Hospital Clínic–IDIBAPS and Biobanco Vasco for the availability of the samples. The work was carried out (in part) at the Esther Koplowitz Centre, Barcelona.
Harvard cohorts (HPFS, NHS, PHS): The study protocol was approved by the institutional review boards of the Brigham and Women’s Hospital and Harvard T.H. Chan School of Public Health, and those of participating registries as required. We acknowledge Channing Division of Network Medicine, Department of Medicine, Brigham and Women’s Hospital as home of the NHS. We would like to thank the participants and staff of the HPFS, NHS and PHS for their valuable contributions as well as the following state cancer registries for their help: AL, AZ, AR, CA, CO, CT, DE, FL, GA, ID, IL, IN, IA, KY, LA, ME, MD, MA, MI, NE, NH, NJ, NY, NC, ND, OH, OK, OR, PA, RI, SC, TN, TX, VA, WA, WY. The authors assume full responsibility for analyses and interpretation of these data.
Interval: A complete list of the investigators and contributors to the INTERVAL trial is provided in reference (32Riaz et al., 2018). The academic coordinating centre would like to thank blood donor centre staff and blood donors for participating in the INTERVAL trial.
Kentucky: We would like to acknowledge the staff at the Kentucky Cancer Registry.
LCCS: We acknowledge the contributions of Jennifer Barrett, Robin Waxman, Gillian Smith and Emma Northwood in conducting this study.
NCCCS I & II: We would like to thank the study participants, and the NC Colorectal Cancer Study staff.
NSHDS investigators thank the Biobank Research Unit at Umeå University, the Västerbotten Intervention Programme, the Northern Sweden MONICA study and Region Västerbotten for providing data and samples and acknowledge the contribution from Biobank Sweden, supported by the Swedish Research Council (VR 2017-00650).
PLCO: We thank the PLCO Cancer Screening Trial screening center investigators and the staff from Information Management Services Inc and Westat Inc. Most importantly, we thank the study participants for their contributions that made this study possible.
SEARCH: We thank the SEARCH team.
SELECT: We thank the research and clinical staff at the sites that participated on SELECT study, without whom the trial would not have been successful. We are also grateful to the 35,533 dedicated men who participated in SELECT.
UK Biobank: We would like to thank the participants and researchers UK Biobank for their participation and acquisition of data.
WHI: We thank the WHI investigators and staff for their dedication, and the study participants for making the program possible. A full listing of WHI investigators can be found at: http://www.whi.org/researchers/Documents%20%20Write%20a%20Paper/WHI%20Investigator%20Short%20List.pdf.
HY is funded by Diabetes UK RD Lawrence fellowship (grant: 17/0005594). SM and TMF are funded by the MRC (MR/T002239/1). CJB is supported by the World Cancer Research Fund (WCRF UK), as part of the World Cancer Research Fund International grant programme (IIG_2019_2009). JAB works in a unit funded by the UK MRC (MC_UU_00011/1) and the University of Bristol. EEV is supported by Diabetes UK (17/0005587) and the World Cancer Research Fund (WCRF UK), as part of the World Cancer Research Fund International grant programme (IIG_2019_2009) and works within the CRUK Integrative Cancer Epidemiology Programme (C18281/A29019).
Genetics and Epidemiology of Colorectal Cancer Consortium (GECCO): National Cancer Institute, National Institutes of Health, U.S. Department of Health and Human Services (U01 CA164930, U01 CA137088, R01 CA059045, R21 CA191312, R01201407). Genotyping/Sequencing services were provided by the Center for Inherited Disease Research (CIDR) contract number HHSN268201700006I and HHSN268201200008I. This research was funded in part through the NIH/NCI Cancer Center Support Grant P30 CA015704. Scientific Computing Infrastructure at Fred Hutch funded by ORIP grant S10OD028685.
ASTERISK: a Hospital Clinical Research Program (PHRC-BRD09/C) from the University Hospital Center of Nantes (CHU de Nantes) and supported by the Regional Council of Pays de la Loire, the Groupement des Entreprises Françaises dans la Lutte contre le Cancer (GEFLUC), the Association Anne de Bretagne Génétique and the Ligue Régionale Contre le Cancer (LRCC).
The ATBC Study is supported by the Intramural Research Program of the U.S. National Cancer Institute, National Institutes of Health.
CLUE II funding was from the National Cancer Institute (U01 CA86308, Early Detection Research Network; P30 CA006973), National Institute on Aging (U01 AG18033), and the American Institute for Cancer Research. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US government.
Maryland Cancer Registry (MCR)
Cancer data was provided by the Maryland Cancer Registry, Center for Cancer Prevention and Control, Maryland Department of Health, with funding from the State of Maryland and the Maryland Cigarette Restitution Fund. The collection and availability of cancer registry data is also supported by the Cooperative Agreement NU58DP006333, funded by the Centers for Disease Control and Prevention. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the Centers for Disease Control and Prevention or the Department of Health and Human Services.
ColoCare: This work was supported by the National Institutes of Health (grant numbers R01 CA189184 [Li/Ulrich], U01 CA206110 [Ulrich/Li/Siegel/Figueireido/Colditz], 2P30CA015704-40 [Gilliland], R01 CA207371 [Ulrich/Li]), the Matthias Lackas-Foundation, the German Consortium for Translational Cancer Research, and the EU TRANSCAN initiative.
The Colon Cancer Family Registry (CCFR, https://www.coloncfr.org) is supported in part by funding from the National Cancer Institute (NCI), National Institutes of Health (NIH) (award U01 CA167551). Support for case ascertainment was provided in part from the Surveillance, Epidemiology, and End Results (SEER) Program and the following U.S. state cancer registries: AZ, CO, MN, NC, NH; and by the Victoria Cancer Registry (Australia) and Ontario Cancer Registry (Canada). The CCFR Set-1 (Illumina 1 M/1M-Duo) and Set-2 (Illumina Omni1-Quad) scans were supported by NIH awards U01 CA122839 and R01 CA143247 (to GC). The CCFR Set-3 (Affymetrix Axiom CORECT Set array) was supported by NIH award U19 CA148107 and R01 CA81488 (to SBG). The CCFR Set-4 (Illumina OncoArray 600K SNP array) was supported by NIH award U19 CA148107 (to SBG) and by the Center for Inherited Disease Research (CIDR), which is funded by the NIH to the Johns Hopkins University, contract number HHSN268201200008I. Additional funding for the OFCCR/ARCTIC was through award GL201-043 from the Ontario Research Fund (to BWZ), award 112746 from the Canadian Institutes of Health Research (to TJH), through a Cancer Risk Evaluation (CaRE) Program grant from the Canadian Cancer Society (to SG), and through generous support from the Ontario Ministry of Research and Innovation. The SFCCR Illumina HumanCytoSNP array was supported in part through NCI/NIH awards U01 CA074794 (to JDP) and /U24 CA074794 and R01 CA076366 (to PAN). The content of this manuscript does not necessarily reflect the views or policies of the NCI, NIH, or any of the collaborating centers in the Colon Cancer Family Registry (CCFR), nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government, any cancer registry, or the CCFR.
COLON: The COLON study is sponsored by Wereld Kanker Onderzoek Fonds, including funds from grant 2014/1179 as part of the World Cancer Research Fund International Regular Grant Programme, by Alpe d’Huzes and the Dutch Cancer Society (UM 2012-5653, UW 2013-5927, UW2015-7946), and by TRANSCAN (JTC2012-MetaboCCC, JTC2013-FOCUS). The Nqplus study is sponsored by a ZonMW investment grant (98-10030); by PREVIEW, the project PREVention of diabetes through lifestyle intervention and population studies in Europe and around the World (PREVIEW) project which received funding from the European Union Seventh Framework Programme (FP7/2007–2013) under grant no. 312057; by funds from TI Food and Nutrition (cardiovascular health theme), a public–private partnership on precompetitive research in food and nutrition; and by FOODBALL, the Food Biomarker Alliance, a project from JPI Healthy Diet for a Healthy Life.
Colorectal Cancer Transdisciplinary (CORECT) Study: The CORECT Study was supported by the National Cancer Institute, National Institutes of Health (NCI/NIH), U.S. Department of Health and Human Services (grant numbers U19 CA148107, R01 CA81488, P30 CA014089, R01 CA197350; P01 CA196569; R01 CA201407) and National Institutes of Environmental Health Sciences, National Institutes of Health (grant number T32 ES013678).
CORSA: “Österreichische Nationalbank Jubiläumsfondsprojekt” (12511) and Austrian Research Funding Agency (FFG) grant 829675.
CPS-II: The American Cancer Society funds the creation, maintenance, and updating of the Cancer Prevention Study-II (CPS-II) cohort. This study was conducted with Institutional Review Board approval.
CRCGEN: Colorectal Cancer Genetics & Genomics, Spanish study was supported by Instituto de Salud Carlos III, co-funded by FEDER funds – a way to build Europe – (grants PI14-613 and PI09-1286), Agency for Management of University and Research Grants (AGAUR) of the Catalan Government (grant 2017SGR723), and Junta de Castilla y León (grant LE22A10-2). Sample collection of this work was supported by the Xarxa de Bancs de Tumors de Catalunya sponsored by Pla Director d’Oncología de Catalunya (XBTC), Plataforma Biobancos PT13/0010/0013 and ICOBIOBANC, sponsored by the Catalan Institute of Oncology.
Czech Republic CCS: This work was supported by the Czech Science Foundation (20-03997 S) and by the Grant Agency of the Ministry of Health of the Czech Republic (grants NV18/03/00199 and NU21-07-00247).
DACHS: This work was supported by the German Research Council (BR 1704/6-1, BR 1704/6-3, BR 1704/6-4, CH 117/1-1, HO 5117/2-1, HE 5998/2-1, KL 2354/3-1, RO 2270/8-1, and BR 1704/17-1), the Interdisciplinary Research Program of the National Center for Tumor Diseases (NCT), Germany, and the German Federal Ministry of Education and Research (01KH0404, 01ER0814, 01ER0815, 01ER1505A, and 01ER1505B).
DALS: National Institutes of Health (R01 CA48998 to M.L. Slattery).
EDRN: This work is funded and supported by the NCI, EDRN Grant (U01 CA 84968-06).
EPIC: The coordination of EPIC is financially supported by the European Commission (DGSANCO) and the International Agency for Research on Cancer. The national cohorts are supported by Danish Cancer Society (Denmark); Ligue Contre le Cancer, Institut Gustave Roussy, Mutuelle Générale de l’Education Nationale, Institut National de la Santé et de la Recherche Médicale (INSERM) (France); German Cancer Aid, German Cancer Research Center (DKFZ), Federal Ministry of Education and Research (BMBF), Deutsche Krebshilfe, Deutsches Krebsforschungszentrum and Federal Ministry of Education and Research (Germany); the Hellenic Health Foundation (Greece); Associazione Italiana per la Ricerca sul Cancro-AIRCItaly and National Research Council (Italy); Dutch Ministry of Public Health, Welfare and Sports (VWS), Netherlands Cancer Registry (NKR), LK Research Funds, Dutch Prevention Funds, Dutch ZON (Zorg Onderzoek Nederland), World Cancer Research Fund (WCRF), Statistics Netherlands (The Netherlands); ERC-2009-AdG 232997 and Nordforsk, Nordic Centre of Excellence programme on Food, Nutrition and Health (Norway); Health Research Fund (FIS), PI13/00061 to Granada, PI13/01162 to EPIC-Murcia, Regional Governments of Andalucía, Asturias, Basque Country, Murcia and Navarra, ISCIII RETIC (RD06/0020) (Spain); Swedish Cancer Society, Swedish Research Council and County Councils of Skåne and Västerbotten (Sweden); Cancer Research UK (14136 to EPIC-Norfolk; C570/A16491 and C8221/A19170 to EPIC-Oxford), Medical Research Council (1000143 to EPIC-Norfolk, MR/M012190/1 to EPICOxford) (United Kingdom).
The EPIC-Norfolk study (https://doi.org/10.22025/2019.10.105.00004) has received funding from the Medical Research Council (MR/N003284/1 and MC-UU_12015/1) and Cancer Research UK (C864/A14136). The genetics work in the EPIC-Norfolk study was funded by the Medical Research Council (MC_PC_13048). Metabolite measurements in the EPIC-Norfolk study were supported by the MRC Cambridge Initiative in Metabolic Science (MR/L00002/1) and the Innovative Medicines Initiative Joint Undertaking under EMIF grant agreement no. 115372.
EPICOLON: This work was supported by grants from Fondo de Investigación Sanitaria/FEDER (PI08/0024, PI08/1276, PS09/02368, PI11/00219, PI11/00681, PI14/00173, PI14/00230, PI17/00509, 17/00878, PI20/00113, PI20/00226, Acción Transversal de Cáncer), Xunta de Galicia (PGIDIT07PXIB9101209PR), Ministerio de Economia y Competitividad (SAF07-64873, SAF 2010-19273, SAF2014-54453R), Fundación Científica de la Asociación Española contra el Cáncer (GCB13131592CAST), Beca Grupo de Trabajo “Oncología” AEG (Asociación Española de Gastroenterología), Fundación Privada Olga Torres, FP7 CHIBCHA Consortium, Agència de Gestió d’Ajuts Universitaris i de Recerca (AGAUR, Generalitat de Catalunya, 2014SGR135, 2014SGR255, 2017SGR21, 2017SGR653), Catalan Tumour Bank Network (Pla Director d’Oncologia, Generalitat de Catalunya), PERIS (SLT002/16/00398, Generalitat de Catalunya), CERCA Programme (Generalitat de Catalunya) and COST Actions BM1206 and CA17118. CIBERehd is funded by the Instituto de Salud Carlos III.
ESTHER/VERDI. This work was supported by grants from the Baden-Württemberg Ministry of Science, Research and Arts and the German Cancer Aid.
Harvard cohorts (HPFS, NHS, PHS): HPFS is supported by the National Institutes of Health (P01 CA055075, UM1 CA167552, U01 CA167552, R01 CA137178, R01 CA151993, R35 CA197735, K07 CA190673, and P50 CA127003), NHS by the National Institutes of Health (R01 CA137178, P01 CA087969, UM1 CA186107, R01 CA151993, R35 CA197735, K07CA190673, and P50 CA127003) and PHS by the National Institutes of Health (R01 CA042182).
Hawaii Adenoma Study: NCI grants R01 CA72520.
HCES-CRC: the Hwasun Cancer Epidemiology Study–Colon and Rectum Cancer (HCES-CRC; grants from Chonnam National University Hwasun Hospital, HCRI15011-1).
Kentucky: This work was supported by the following grant support: Clinical Investigator Award from Damon Runyon Cancer Research Foundation (CI-8); NCI R01CA136726.
LCCS: The Leeds Colorectal Cancer Study was funded by the Food Standards Agency and Cancer Research UK Programme Award (C588/A19167).
Melbourne Collaborative Cohort Study (MCCS) cohort recruitment was funded by VicHealth and Cancer Council Victoria. The MCCS was further augmented by Australian National Health and Medical Research Council grants 209057, 396414, and 1074383 and by infrastructure provided by Cancer Council Victoria. Cases and their vital status were ascertained through the Victorian Cancer Registry and the Australian Institute of Health and Welfare, including the National Death Index and the Australian Cancer Database.
Multiethnic Cohort (MEC) Study: National Institutes of Health (R37 CA54281, P01 CA033619, R01 CA063464, and U01 CA164973).
MECC: This work was supported by the National Institutes of Health, U.S. Department of Health and Human Services (R01 CA81488 to SBG and GR).
MSKCC: The work at Sloan Kettering in New York was supported by the Robert and Kate Niehaus Center for Inherited Cancer Genomics and the Romeo Milio Foundation. Moffitt: This work was supported by funding from the National Institutes of Health (grant numbers R01 CA189184, P30 CA076292), Florida Department of Health Bankhead-Coley Grant 09BN-13, and the University of South Florida Oehler Foundation. Moffitt contributions were supported in part by the Total Cancer Care Initiative, Collaborative Data Services Core, and Tissue Core at the H. Lee Moffitt Cancer Center & Research Institute, a National Cancer Institute-designated Comprehensive Cancer Center (grant number P30 CA076292).
NCCCS I & II: We acknowledge funding support for this project from the National Institutes of Health, R01 CA66635 and P30 DK034987.
NFCCR: This work was supported by an Interdisciplinary Health Research Team award from the Canadian Institutes of Health Research (CRT 43821); the National Institutes of Health, U.S. Department of Health and Human Serivces (U01 CA74783); and National Cancer Institute of Canada grants (18223 and 18226). We wish to acknowledge the contribution of Alexandre Belisle and the genotyping team of the McGill University and Génome Québec Innovation Centre, Montréal, Canada, for genotyping the Sequenom panel in the NFCCR samples. Funding was provided to Michael O. Woods by the Canadian Cancer Society Research Institute.
NSHDS: Swedish Research Council; Swedish Cancer Society; Cutting-Edge Research Grant and other grants from Region Västerbotten; Knut and Alice Wallenberg Foundation; Lion’s Cancer Research Foundation at Umeå University; the Cancer Research Foundation in Northern Sweden; and the Faculty of Medicine, Umeå University, Umeå, Sweden.
OSUMC: OCCPI funding was provided by Pelotonia and HNPCC funding was provided by the NCI (CA16058 and CA67941).
PLCO: Intramural Research Program of the Division of Cancer Epidemiology and Genetics and supported by contracts from the Division of Cancer Prevention, National Cancer Institute, NIH, DHHS. Funding was provided by National Institutes of Health (NIH), Genes, Environment and Health Initiative (GEI) Z01 CP 010200, NIH U01 HG004446, and NIH GEI U01 HG 004438.
SEARCH: The University of Cambridge has received salary support in respect of PDPP from the NHS in the East of England through the Clinical Academic Reserve. Cancer Research UK (C490/A16561); the UK National Institute for Health Research Biomedical Research Centres at the University of Cambridge.
SELECT: Research reported in this publication was supported in part by the National Cancer Institute of the National Institutes of Health under Award Numbers U10 CA37429 (CD Blanke), and UM1 CA182883 (CM Tangen/IM Thompson). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
SMS and REACH: This work was supported by the National Cancer Institute grant P01 CA074184 to JDP and PAN, grants R01 CA097325, R03 CA153323, and K05 CA152715 to PAN, and the National Center for Advancing Translational Sciences at the National Institutes of Health (grant KL2 TR000421 to ANB-H).
The Swedish Low-risk Colorectal Cancer Study: The study was supported by grants from the Swedish research council; K2015-55X-22674-01-4, K2008-55X-20157-03-3, K2006-72X-20157-01-2, and the Stockholm County Council (ALF project).
Swedish Mammography Cohort and Cohort of Swedish Men: This work is supported by the Swedish Research Council /Infrastructure grant, the Swedish Cancer Foundation, and the Karolinska Institute´s Distinguished Professor Award to Alicja Wolk.
UK Biobank: This research has been conducted using the UK Biobank Resource under Application Number 8,614
VITAL: National Institutes of Health (K05 CA154337).
WHI: The WHI program is funded by the National Heart, Lung, and Blood Institute, National Institutes of Health, U.S. Department of Health and Human Services through contracts HHSN268201100046C, HHSN268201100001C, HHSN268201100002C, HHSN268201100003C, HHSN268201100004C, and HHSN271201100004C.
Funding Statement
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Contributor Information
Timothy M Frayling, Email: t.m.frayling@exeter.ac.uk.
Hanieh Yaghootkar, Email: Hanieh.Yaghootkar@brunel.ac.uk.
Edward D Janus, University of Melbourne, Australia.
Matthias Barton, University of Zurich, Switzerland.
Funding Information
This paper was supported by the following grants:
Diabetes UK 17/0005594 to Hanieh Yaghootkar.
Medical Research Council MR/T002239/1 to Susan Martin, Timothy Frayling.
World Cancer Research Fund IIG_2019_2009 to Caroline Bull, Emma E Vincent.
Medical Research Council MC_UU_00011/1 to Joshua A Bell.
Diabetes UK 17/0005587 to Emma E Vincent.
Cancer Research UK C18281/A29019 to Emma E Vincent.
Additional information
Competing interests
No competing interests declared.
No competing interests declared.
Naveed Sattar has consulted for Afimmune, Amgen, AstraZeneca, Boehringer Ingelheim, Eli Lilly, Hanmi Pharmaceuticals, Merck Sharp & Dohme, Novartis, Novo Nordisk, Pfizer, and Sanofi; and received grant support paid to his University from AstraZeneca, Boehringer Ingelheim and Roche Diagnostics outside the submitted work.
Tim Frayling has consulted for Boehringer Ingelheim and Sanofi and has a student supported by GSK.
Author contributions
Data curation, Formal analysis, Investigation, Methodology, Software, Validation, Visualization, Writing – original draft, Writing – review and editing.
Conceptualization, Methodology, Project administration, Resources, Software, Supervision, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Software, Writing – review and editing.
Resources, Software, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Resources, Writing – review and editing.
Conceptualization, Funding acquisition, Investigation, Methodology, Project administration, Supervision, Validation, Writing – original draft, Writing – review and editing.
Conceptualization, Funding acquisition, Investigation, Methodology, Project administration, Supervision, Validation, Writing – original draft, Writing – review and editing.
Ethics
Human subjects: For the UK Biobank, all participants provided informed written consent and the National Research Ethics Service Committee North West-Haydock approved the study. All procedures in the UK Biobank study were conducted in accordance to the World Medical Association declaration of Helsinki ethical principles for medical research.
Additional files
(a) Characterisation of monogenic obesity, lipodystrophy, unfavourable adiposity (UFA), and favourable adiposity (FA) using body fat percentage and a selection of metabolic biomarkers. (b) Mendelian randomisation (MR) studies testing the role of obesity (usually as body mass index [BMI]) identified in literature search. (c) (i) Summary statistics of published genome-wide association studies (GWAS) used. Mean (standard deviation [SD] or range) are given for continuous study characteristics where available, mean ranges are given for meta-analyses unless otherwise specified. *Statistics represent only UK Biobank cohort of those included in meta-analysis. (c) (ii) Summary statistics of FinnGen studies used. Mean age of cases is given where available, BMI is not adjusted for, and UK Biobank is not included in these studies. ICD codes taken from hospital discharge register and/or causes of death register. (c) (iii) Summary statistics of UK Biobank studies used. Mean (SD) are given for continuous study characteristics of cases. Self-report code is from n_20002_* variable in UK Biobank. (d) The summary of 73 BMI and 696 body fat percentage genetic variants, the latter including 36 FA and 38 UFA genetic variants. Beta, SE, and p are from the GWAS of BMI and body fat percentage in UK Biobank, respectively. BMI variants were discovered using non-UK Biobank cohorts, and so some SNPs listed may have zero effect size in the UK Biobank GWAS of BMI. (e) The inverse-variance weighted two-sample MR analysis/meta-analysis of 37 identified diseases from published GWAS and/or FinnGen for BMI, body fat percentage, FA, and UFA clusters. Italicised results are those that were interpreted – including all BMI, body fat percentage if a causal effect of BMI was indicated, and FA/UFA if a causal effect of BMI and body fat percentage was indicated. (f) Heterogeneity statistics from random-effects meta-analysis of inverse-variance weighted MR of published GWAS and FinnGen studies. (g) (i) The inverse-variance weighted, weighted median, Egger, and penalised weighted median MR analyses for BMI using FinnGen and published GWAS. (g) (ii) The inverse-variance weighted, weighted median, Egger, and penalised weighted median MR analyses for body fat percentage using FinnGen and published GWAS. (g) (iii) The inverse-variance weighted, weighted median, Egger, and penalised weighted median MR analyses for FA using FinnGen and published GWAS. (g) (iv) The inverse-variance weighted, weighted median, Egger, and penalised weighted median MR analyses for UFA using FinnGen and published GWAS. (h) The inverse-variance weighted MR analysis of identified diseases from UK Biobank for BMI, body fat percentage, FA, and UFA clusters. PMID, PubMed ID; N, sample size; OPCS, operating procedure codes; SE, standard error; p, p-value; OR, odds ratio; 95% CI, 95% confidence interval; Q, Q-statistic; I2, I2-statistic; LCI, lower 95% confidence interval; UCI, upper 95% confidence interval; Intercept p, intercept p-value; I2 MR-Egger, I2-statistic MR-Egger.
Data availability
GWAS data from the outcome diseases studied is available from links published in the original studies (Supplementary File 1ci). FinnGen data is available at: https://finngen.gitbook.io/documentation/, and the list of disease outcomes used is in Supplementary File 1cii. Individual-level UK Biobank data cannot be provided, but it is available by application to the UK Biobank: https://www.ukbiobank.ac.uk, and a list of the traits used is in Supplementary File 1ciii. Code used to conduct this analysis will be made available on GitHub after removing any sensitive information (https://github.com/susiemartin/uncoupling-bmi, copy archived at swh:1:rev:f3472762ad6cb7f313656f684e07c14b8735efe5).
References
- An J, Gharahkhani P, Law MH, Ong J-S, Han X, Olsen CM, Neale RE, Lai J, Vaughan TL, Gockel I, Thieme R, Böhmer AC, Jankowski J, Fitzgerald RC, Schumacher J, Palles C, BEACON. 23andMe Research Team. Whiteman DC, MacGregor S. Gastroesophageal reflux GWAS identifies risk loci that also associate with subsequent severe esophageal diseases. Nature Communications. 2019;10:4219. doi: 10.1038/s41467-019-11968-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society. 1995;57:289–300. doi: 10.1111/j.2517-6161.1995.tb02031.x. [DOI] [Google Scholar]
- Budu-Aggrey A, Brumpton B, Tyrrell J, Watkins S, Modalsli EH, Celis-Morales C, Ferguson LD, Vie GÅ, Palmer T, Fritsche LG, Løset M, Nielsen JB, Zhou W, Tsoi LC, Wood AR, Jones SE, Beaumont R, Saunes M, Romundstad PR, Siebert S, McInnes IB, Elder JT, Davey Smith G, Frayling TM, Åsvold BO, Brown SJ, Sattar N, Paternoster L. Evidence of a causal relationship between body mass index and psoriasis: A mendelian randomization study. PLOS Medicine. 2019;16:e1002739. doi: 10.1371/journal.pmed.1002739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bull CJ, Bell JA, Murphy N, Sanderson E, Davey Smith G, Timpson NJ, Banbury BL, Albanes D, Berndt SI, Bézieau S, Bishop DT, Brenner H, Buchanan DD, Burnett-Hartman A, Casey G, Castellví-Bel S, Chan AT, Chang-Claude J, Cross AJ, de la Chapelle A, Figueiredo JC, Gallinger SJ, Gapstur SM, Giles GG, Gruber SB, Gsur A, Hampe J, Hampel H, Harrison TA, Hoffmeister M, Hsu L, Huang W-Y, Huyghe JR, Jenkins MA, Joshu CE, Keku TO, Kühn T, Kweon S-S, Le Marchand L, Li CI, Li L, Lindblom A, Martín V, May AM, Milne RL, Moreno V, Newcomb PA, Offit K, Ogino S, Phipps AI, Platz EA, Potter JD, Qu C, Quirós JR, Rennert G, Riboli E, Sakoda LC, Schafmayer C, Schoen RE, Slattery ML, Tangen CM, Tsilidis KK, Ulrich CM, van Duijnhoven FJB, van Guelpen B, Visvanathan K, Vodicka P, Vodickova L, Wang H, White E, Wolk A, Woods MO, Wu AH, Campbell PT, Zheng W, Peters U, Vincent EE, Gunter MJ. Adiposity, metabolites, and colorectal cancer risk: Mendelian randomization study. BMC Medicine. 2020;18:396. doi: 10.1186/s12916-020-01855-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carreras-Torres R, Johansson M, Gaborieau V, Haycock PC, Wade KH, Relton CL, Martin RM, Davey Smith G, Brennan P. The Role of Obesity, Type 2 Diabetes, and Metabolic Factors in Pancreatic Cancer: A Mendelian Randomization Study. Journal of the National Cancer Institute. 2017a;109:djx012. doi: 10.1093/jnci/djx012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carreras-Torres R, Johansson M, Haycock PC, Wade KH, Relton CL, Martin RM, Davey Smith G, Albanes D, Aldrich MC, Andrew A, Arnold SM, Bickeböller H, Bojesen SE, Brunnström H, Manjer J, Brüske I, Caporaso NE, Chen C, Christiani DC, Christian WJ, Doherty JA, Duell EJ, Field JK, Davies MPA, Marcus MW, Goodman GE, Grankvist K, Haugen A, Hong YC, Kiemeney LA, van der Heijden E, Kraft P, Johansson MB, Lam S, Landi MT, Lazarus P, Le Marchand L, Liu G, Melander O, Park SL, Rennert G, Risch A, Haura EB, Scelo G, Zaridze D, Mukeriya A, Savić M, Lissowska J, Swiatkowska B, Janout V, Holcatova I, Mates D, Schabath MB, Shen H, Tardon A, Teare MD, Woll P, Tsao MS, Wu X, Yuan JM, Hung RJ, Amos CI, McKay J, Brennan P. Obesity, metabolic factors and risk of different histological types of lung cancer: A Mendelian randomization study. PLOS ONE. 2017b;12:e0177875. doi: 10.1371/journal.pone.0177875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng L, Zhuang H, Ju H, Yang S, Han J, Tan R, Hu Y. Exposing the Causal Effect of Body Mass Index on the Risk of Type 2 Diabetes Mellitus: A Mendelian Randomization Study. Frontiers in Genetics. 2019;10:94. doi: 10.3389/fgene.2019.00094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi HK, Mount DB, Reginato AM. Pathogenesis of Gout. Annals of Internal Medicine. 2005;143:499. doi: 10.7326/0003-4819-143-7-200510040-00009. [DOI] [PubMed] [Google Scholar]
- Collins R. What makes UK Biobank special? Lancet. 2012;379:1173–1174. doi: 10.1016/S0140-6736(12)60404-8. [DOI] [PubMed] [Google Scholar]
- Corbin LJ, Richmond RC, Wade KH, Burgess S, Bowden J, Smith GD, Timpson NJ. BMI as a Modifiable Risk Factor for Type 2 Diabetes: Refining and Understanding Causal Estimates Using Mendelian Randomization. Diabetes. 2016;65:3002–3007. doi: 10.2337/db16-0418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cornish AJ, Law PJ, Timofeeva M, Palin K, Farrington SM, Palles C, Jenkins MA, Casey G, Brenner H, Chang-Claude J, Hoffmeister M, Kirac I, Maughan T, Brezina S, Gsur A, Cheadle JP, Aaltonen LA, Tomlinson I, Dunlop MG, Houlston RS. Modifiable pathways for colorectal cancer: a mendelian randomisation analysis. The Lancet. Gastroenterology & Hepatology. 2020;5:55–62. doi: 10.1016/S2468-1253(19)30294-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cruz-Monserrate Z, Conwell DL, Krishna SG. The Impact of Obesity on Gallstone Disease, Acute Pancreatitis, and Pancreatic Cancer. Gastroenterology Clinics of North America. 2016;45:625–637. doi: 10.1016/j.gtc.2016.07.010. [DOI] [PubMed] [Google Scholar]
- Dalbeth N, Merriman TR, Stamp LK. Gout. Lancet. 2016;388:2039–2052. doi: 10.1016/S0140-6736(16)00346-9. [DOI] [PubMed] [Google Scholar]
- Davies NM, Gaunt TR, Lewis SJ, Holly J, Donovan JL, Hamdy FC, Kemp JP, Eeles R, Easton D, Kote-Jarai Z, Al Olama AA, Benlloch S, Muir K, Giles GG, Wiklund F, Gronberg H, Haiman CA, Schleutker J, Nordestgaard BG, Travis RC, Neal D, Pashayan N, Khaw K-T, Stanford JL, Blot WJ, Thibodeau S, Maier C, Kibel AS, Cybulski C, Cannon-Albright L, Brenner H, Park J, Kaneva R, Batra J, Teixeira MR, Pandha H, PRACTICAL consortium. Lathrop M, Smith GD, Martin RM. The effects of height and BMI on prostate cancer incidence and mortality: a Mendelian randomization study in 20,848 cases and 20,214 controls from the PRACTICAL consortium. Cancer Causes & Control. 2015;26:1603–1616. doi: 10.1007/s10552-015-0654-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Day F, Karaderi T, Jones MR, Meun C, He C, Drong A, Kraft P, Lin N, Huang H, Broer L, Magi R, Saxena R, Laisk T, Urbanek M, Hayes MG, Thorleifsson G, Fernandez-Tajes J, Mahajan A, Mullin BH, Stuckey BGA, Spector TD, Wilson SG, Goodarzi MO, Davis L, Obermayer-Pietsch B, Uitterlinden AG, Anttila V, Neale BM, Jarvelin MR, Fauser B, Kowalska I, Visser JA, Andersen M, Ong K, Stener-Victorin E, Ehrmann D, Legro RS, Salumets A, McCarthy MI, Morin-Papunen L, Thorsteinsdottir U, Stefansson K, Styrkarsdottir U, Perry JRB, Dunaif A, Laven J, Franks S, Lindgren CM, Welt CK, 23andMe Research Team Large-scale genome-wide meta-analysis of polycystic ovary syndrome suggests shared genetic architecture for different diagnosis criteria. PLOS Genetics. 2018;14:e1007813. doi: 10.1371/journal.pgen.1007813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dowlatshahi EA, van der Voort EAM, Arends LR, Nijsten T. Markers of systemic inflammation in psoriasis: a systematic review and meta-analysis. The British Journal of Dermatology. 2013;169:266–282. doi: 10.1111/bjd.12355. [DOI] [PubMed] [Google Scholar]
- Emdin CA, Khera AV, Kathiresan S. Genetic Predisposition to Abdominal Obesity and Cardiometabolic Risk-Reply. JAMA. 2017;317:2334–2335. doi: 10.1001/jama.2017.5044. [DOI] [PubMed] [Google Scholar]
- Fall T, Hägg S, Mägi R, Ploner A, Fischer K, Horikoshi M, Sarin A-P, Thorleifsson G, Ladenvall C, Kals M, Kuningas M, Draisma HHM, Ried JS, van Zuydam NR, Huikari V, Mangino M, Sonestedt E, Benyamin B, Nelson CP, Rivera NV, Kristiansson K, Shen H-Y, Havulinna AS, Dehghan A, Donnelly LA, Kaakinen M, Nuotio M-L, Robertson N, de Bruijn RFAG, Ikram MA, Amin N, Balmforth AJ, Braund PS, Doney ASF, Döring A, Elliott P, Esko T, Franco OH, Gretarsdottir S, Hartikainen A-L, Heikkilä K, Herzig K-H, Holm H, Hottenga JJ, Hyppönen E, Illig T, Isaacs A, Isomaa B, Karssen LC, Kettunen J, Koenig W, Kuulasmaa K, Laatikainen T, Laitinen J, Lindgren C, Lyssenko V, Läärä E, Rayner NW, Männistö S, Pouta A, Rathmann W, Rivadeneira F, Ruokonen A, Savolainen MJ, Sijbrands EJG, Small KS, Smit JH, Steinthorsdottir V, Syvänen A-C, Taanila A, Tobin MD, Uitterlinden AG, Willems SM, Willemsen G, Witteman J, Perola M, Evans A, Ferrières J, Virtamo J, Kee F, Tregouet D-A, Arveiler D, Amouyel P, Ferrario MM, Brambilla P, Hall AS, Heath AC, Madden PAF, Martin NG, Montgomery GW, Whitfield JB, Jula A, Knekt P, Oostra B, van Duijn CM, Penninx BWJH, Smith GD, Kaprio J, Samani NJ, Gieger C, Peters A, Wichmann HE, Boomsma DI, de Geus EJC, Tuomi T, Power C, Hammond CJ, Spector TD, Lind L, Orho-Melander M, Palmer CNA, Morris AD, Groop L, Järvelin M-R, Salomaa V, Vartiainen E, Hofman A, Ripatti S, Metspalu A, Thorsteinsdottir U, Stefansson K, Pedersen NL, McCarthy MI, Ingelsson E, Prokopenko I, European Network for Genetic and Genomic Epidemiology (ENGAGE) consortium The role of adiposity in cardiometabolic traits: a Mendelian randomization analysis. PLOS Medicine. 2013;10:e1001474. doi: 10.1371/journal.pmed.1001474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferreira MAR, Mathur R, Vonk JM, Szwajda A, Brumpton B, Granell R, Brew BK, Ullemar V, Lu Y, Jiang Y, 23andMe Research Team. eQTLGen Consortium. BIOS Consortium. Magnusson PKE, Karlsson R, Hinds DA, Paternoster L, Koppelman GH, Almqvist C. Genetic Architectures of Childhood- and Adult-Onset Asthma Are Partly Distinct. American Journal of Human Genetics. 2019;104:665–684. doi: 10.1016/j.ajhg.2019.02.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- FinnGen FinnGen Documentation of R4 release. 2021. [April 21, 2021]. https://finngen.gitbook.io/documentation
- Gao C, Patel CJ, Michailidou K, Peters U, Gong J, Schildkraut J, Schumacher FR, Zheng W, Boffetta P, Stucker I, Willett W, Gruber S, Easton DF, Hunter DJ, Sellers TA, Haiman C, Henderson BE, Hung RJ, Amos C, Pierce BL, Lindström S, Kraft P, the Colorectal Transdisciplinary Study (CORECT); Discovery, Biology and Risk of Inherited Variants in Breast Cancer (DRIVE); Elucidating Loci Involved in Prostate Cancer Susceptibility (ELLIPSE); Follow-up of Ovarian Cancer Genetic Association and Interaction Studies (FOCI); and Transdisciplinary Research in Cancer of the Lung (TRICL) Mendelian randomization study of adiposity-related traits and risk of breast, ovarian, prostate, lung and colorectal cancer. International Journal of Epidemiology. 2016;45:896–908. doi: 10.1093/ije/dyw129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green HD, Beaumont RN, Wood AR, Hamilton B, Jones SE, Goodhand JR, Kennedy NA, Ahmad T, Yaghootkar H, Weedon MN, Frayling TM, Tyrrell J. Genetic evidence that higher central adiposity causes gastro-oesophageal reflux disease: a Mendelian randomization study. International Journal of Epidemiology. 2020;49:1270–1281. doi: 10.1093/ije/dyaa082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo Y, Warren Andersen S, Shu X-O, Michailidou K, Bolla MK, Wang Q, Garcia-Closas M, Milne RL, Schmidt MK, Chang-Claude J, Dunning A, Bojesen SE, Ahsan H, Aittomäki K, Andrulis IL, Anton-Culver H, Arndt V, Beckmann MW, Beeghly-Fadiel A, Benitez J, Bogdanova NV, Bonanni B, Børresen-Dale A-L, Brand J, Brauch H, Brenner H, Brüning T, Burwinkel B, Casey G, Chenevix-Trench G, Couch FJ, Cox A, Cross SS, Czene K, Devilee P, Dörk T, Dumont M, Fasching PA, Figueroa J, Flesch-Janys D, Fletcher O, Flyger H, Fostira F, Gammon M, Giles GG, Guénel P, Haiman CA, Hamann U, Hooning MJ, Hopper JL, Jakubowska A, Jasmine F, Jenkins M, John EM, Johnson N, Jones ME, Kabisch M, Kibriya M, Knight JA, Koppert LB, Kosma V-M, Kristensen V, Le Marchand L, Lee E, Li J, Lindblom A, Luben R, Lubinski J, Malone KE, Mannermaa A, Margolin S, Marme F, McLean C, Meijers-Heijboer H, Meindl A, Neuhausen SL, Nevanlinna H, Neven P, Olson JE, Perez JIA, Perkins B, Peterlongo P, Phillips K-A, Pylkäs K, Rudolph A, Santella R, Sawyer EJ, Schmutzler RK, Seynaeve C, Shah M, Shrubsole MJ, Southey MC, Swerdlow AJ, Toland AE, Tomlinson I, Torres D, Truong T, Ursin G, Van Der Luijt RB, Verhoef S, Whittemore AS, Winqvist R, Zhao H, Zhao S, Hall P, Simard J, Kraft P, Pharoah P, Hunter D, Easton DF, Zheng W. Genetically Predicted Body Mass Index and Breast Cancer Risk: Mendelian Randomization Analyses of Data from 145,000 Women of European Descent. PLOS Medicine. 2016;13:e1002105. doi: 10.1371/journal.pmed.1002105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hägg S, Fall T, Ploner A, Mägi R, Fischer K, Draisma HHM, Kals M, de Vries PS, Dehghan A, Willems SM, Sarin A-P, Kristiansson K, Nuotio M-L, Havulinna AS, de Bruijn RFAG, Ikram MA, Kuningas M, Stricker BH, Franco OH, Benyamin B, Gieger C, Hall AS, Huikari V, Jula A, Järvelin M-R, Kaakinen M, Kaprio J, Kobl M, Mangino M, Nelson CP, Palotie A, Samani NJ, Spector TD, Strachan DP, Tobin MD, Whitfield JB, Uitterlinden AG, Salomaa V, Syvänen A-C, Kuulasmaa K, Magnusson PK, Esko T, Hofman A, de Geus EJC, Lind L, Giedraitis V, Perola M, Evans A, Ferrières J, Virtamo J, Kee F, Tregouet D-A, Arveiler D, Amouyel P, Gianfagna F, Brambilla P, Ripatti S, van Duijn CM, Metspalu A, Prokopenko I, McCarthy MI, Pedersen NL, Ingelsson E, European Network for Genetic and Genomic Epidemiology Consortium Adiposity as a cause of cardiovascular disease: a Mendelian randomization study. International Journal of Epidemiology. 2015;44:578–586. doi: 10.1093/ije/dyv094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang LO, Rauch A, Mazzaferro E, Preuss M, Carobbio S, Bayrak CS, Chami N, Wang Z, Schick UM, Yang N, Itan Y, Vidal-Puig A, den Hoed M, Mandrup S, Kilpeläinen TO, Loos RJF. Genome-wide discovery of genetic loci that uncouple excess adiposity from its comorbidities. Nature Metabolism. 2021;3:228–243. doi: 10.1038/s42255-021-00346-2. [DOI] [PubMed] [Google Scholar]
- Huyghe JR, Bien SA, Harrison TA, Kang HM, Chen S, Schmit SL, Conti DV, Qu C, Jeon J, Edlund CK, Greenside P, Wainberg M, Schumacher FR, Smith JD, Levine DM, Nelson SC, Sinnott-Armstrong NA, Albanes D, Alonso MH, Anderson K, Arnau-Collell C, Arndt V, Bamia C, Banbury BL, Baron JA, Berndt SI, Bézieau S, Bishop DT, Boehm J, Boeing H, Brenner H, Brezina S, Buch S, Buchanan DD, Burnett-Hartman A, Butterbach K, Caan BJ, Campbell PT, Carlson CS, Castellví-Bel S, Chan AT, Chang-Claude J, Chanock SJ, Chirlaque M-D, Cho SH, Connolly CM, Cross AJ, Cuk K, Curtis KR, de la Chapelle A, Doheny KF, Duggan D, Easton DF, Elias SG, Elliott F, English DR, Feskens EJM, Figueiredo JC, Fischer R, FitzGerald LM, Forman D, Gala M, Gallinger S, Gauderman WJ, Giles GG, Gillanders E, Gong J, Goodman PJ, Grady WM, Grove JS, Gsur A, Gunter MJ, Haile RW, Hampe J, Hampel H, Harlid S, Hayes RB, Hofer P, Hoffmeister M, Hopper JL, Hsu W-L, Huang W-Y, Hudson TJ, Hunter DJ, Ibañez-Sanz G, Idos GE, Ingersoll R, Jackson RD, Jacobs EJ, Jenkins MA, Joshi AD, Joshu CE, Keku TO, Key TJ, Kim HR, Kobayashi E, Kolonel LN, Kooperberg C, Kühn T, Küry S, Kweon S-S, Larsson SC, Laurie CA, Le Marchand L, Leal SM, Lee SC, Lejbkowicz F, Lemire M, Li CI, Li L, Lieb W, Lin Y, Lindblom A, Lindor NM, Ling H, Louie TL, Männistö S, Markowitz SD, Martín V, Masala G, McNeil CE, Melas M, Milne RL, Moreno L, Murphy N, Myte R, Naccarati A, Newcomb PA, Offit K, Ogino S, Onland-Moret NC, Pardini B, Parfrey PS, Pearlman R, Perduca V, Pharoah PDP, Pinchev M, Platz EA, Prentice RL, Pugh E, Raskin L, Rennert G, Rennert HS, Riboli E, Rodríguez-Barranco M, Romm J, Sakoda LC, Schafmayer C, Schoen RE, Seminara D, Shah M, Shelford T, Shin M-H, Shulman K, Sieri S, Slattery ML, Southey MC, Stadler ZK, Stegmaier C, Su Y-R, Tangen CM, Thibodeau SN, Thomas DC, Thomas SS, Toland AE, Trichopoulou A, Ulrich CM, Van Den Berg DJ, van Duijnhoven FJB, Van Guelpen B, van Kranen H, Vijai J, Visvanathan K, Vodicka P, Vodickova L, Vymetalkova V, Weigl K, Weinstein SJ, White E, Win AK, Wolf CR, Wolk A, Woods MO, Wu AH, Zaidi SH, Zanke BW, Zhang Q, Zheng W, Scacheri PC, Potter JD, Bassik MC, Kundaje A, Casey G, Moreno V, Abecasis GR, Nickerson DA, Gruber SB, Hsu L, Peters U. Discovery of common and rare genetic risk variants for colorectal cancer. Nature Genetics. 2019;51:76–87. doi: 10.1038/s41588-018-0286-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huyghe JR, Harrison TA, Bien SA, Hampel H, Figueiredo JC, Schmit SL, Conti DV, Chen S, Qu C, Lin Y, Barfield R, Baron JA, Cross AJ, Diergaarde B, Duggan D, Harlid S, Imaz L, Kang HM, Levine DM, Perduca V, Perez-Cornago A, Sakoda LC, Schumacher FR, Slattery ML, Toland AE, van Duijnhoven FJB, Van Guelpen B, Agudo A, Albanes D, Alonso MH, Anderson K, Arnau-Collell C, Arndt V, Banbury BL, Bassik MC, Berndt SI, Bézieau S, Bishop DT, Boehm J, Boeing H, Boutron-Ruault M-C, Brenner H, Brezina S, Buch S, Buchanan DD, Burnett-Hartman A, Caan BJ, Campbell PT, Carr PR, Castells A, Castellví-Bel S, Chan AT, Chang-Claude J, Chanock SJ, Curtis KR, de la Chapelle A, Easton DF, English DR, Feskens EJM, Gala M, Gallinger SJ, Gauderman WJ, Giles GG, Goodman PJ, Grady WM, Grove JS, Gsur A, Gunter MJ, Haile RW, Hampe J, Hoffmeister M, Hopper JL, Hsu W-L, Huang W-Y, Hudson TJ, Jenab M, Jenkins MA, Joshi AD, Keku TO, Kooperberg C, Kühn T, Küry S, Le Marchand L, Lejbkowicz F, Li CI, Li L, Lieb W, Lindblom A, Lindor NM, Männistö S, Markowitz SD, Milne RL, Moreno L, Murphy N, Nassir R, Offit K, Ogino S, Panico S, Parfrey PS, Pearlman R, Pharoah PDP, Phipps AI, Platz EA, Potter JD, Prentice RL, Qi L, Raskin L, Rennert G, Rennert HS, Riboli E, Schafmayer C, Schoen RE, Seminara D, Song M, Su Y-R, Tangen CM, Thibodeau SN, Thomas DC, Trichopoulou A, Ulrich CM, Visvanathan K, Vodicka P, Vodickova L, Vymetalkova V, Weigl K, Weinstein SJ, White E, Wolk A, Woods MO, Wu AH, Abecasis GR, Nickerson DA, Scacheri PC, Kundaje A, Casey G, Gruber SB, Hsu L, Moreno V, Hayes RB, Newcomb PA, Peters U. Genetic architectures of proximal and distal colorectal cancer are partly distinct. Gut. 2021;70:1325–1334. doi: 10.1136/gutjnl-2020-321534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iskandar IYK, Ashcroft DM, Warren RB, Yiu ZZN, McElhone K, Lunt M, Barker JNWN, Burden AD, Ormerod AD, Reynolds NJ, Smith CH, Griffiths CEM. Demographics and disease characteristics of patients with psoriasis enrolled in the British Association of Dermatologists Biologic Interventions Register. The British Journal of Dermatology. 2015;173:510–518. doi: 10.1111/bjd.13908. [DOI] [PubMed] [Google Scholar]
- Jansen IE, Savage JE, Watanabe K, Bryois J, Williams DM, Steinberg S, Sealock J, Karlsson IK, Hägg S, Athanasiu L, Voyle N, Proitsi P, Witoelar A, Stringer S, Aarsland D, Almdahl IS, Andersen F, Bergh S, Bettella F, Bjornsson S, Brækhus A, Bråthen G, de Leeuw C, Desikan RS, Djurovic S, Dumitrescu L, Fladby T, Hohman TJ, Jonsson PV, Kiddle SJ, Rongve A, Saltvedt I, Sando SB, Selbæk G, Shoai M, Skene NG, Snaedal J, Stordal E, Ulstein ID, Wang Y, White LR, Hardy J, Hjerling-Leffler J, Sullivan PF, van der Flier WM, Dobson R, Davis LK, Stefansson H, Stefansson K, Pedersen NL, Ripke S, Andreassen OA, Posthuma D. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nature Genetics. 2019;51:404–413. doi: 10.1038/s41588-018-0311-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarvis D, Mitchell JS, Law PJ, Palin K, Tuupanen S, Gylfe A, Hänninen UA, Cajuso T, Tanskanen T, Kondelin J, Kaasinen E, Sarin A-P, Kaprio J, Eriksson JG, Rissanen H, Knekt P, Pukkala E, Jousilahti P, Salomaa V, Ripatti S, Palotie A, Järvinen H, Renkonen-Sinisalo L, Lepistö A, Böhm J, Meklin J-P, Al-Tassan NA, Palles C, Martin L, Barclay E, Farrington SM, Timofeeva MN, Meyer BF, Wakil SM, Campbell H, Smith CG, Idziaszczyk S, Maughan TS, Kaplan R, Kerr R, Kerr D, Buchanan DD, Win AK, Hopper JL, Jenkins MA, Lindor NM, Newcomb PA, Gallinger S, Conti D, Schumacher F, Casey G, Taipale J, Aaltonen LA, Cheadle JP, Dunlop MG, Tomlinson IP, Houlston RS. Mendelian randomisation analysis strongly implicates adiposity with risk of developing colorectal cancer. British Journal of Cancer. 2016;115:266–272. doi: 10.1038/bjc.2016.188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji Y, Yiorkas AM, Frau F, Mook-Kanamori D, Staiger H, Thomas EL, Atabaki-Pasdar N, Campbell A, Tyrrell J, Jones SE, Beaumont RN, Wood AR, Tuke MA, Ruth KS, Mahajan A, Murray A, Freathy RM, Weedon MN, Hattersley AT, Hayward C, Machann J, Häring H-U, Franks P, de Mutsert R, Pearson E, Stefan N, Frayling TM, Allebrandt KV, Bell JD, Blakemore AI, Yaghootkar H. Genome-Wide and Abdominal MRI Data Provide Evidence That a Genetically Determined Favorable Adiposity Phenotype Is Characterized by Lower Ectopic Liver Fat and Lower Risk of Type 2 Diabetes, Heart Disease, and Hypertension. Diabetes. 2019;68:207–219. doi: 10.2337/db18-0708. [DOI] [PubMed] [Google Scholar]
- Johansson M, Carreras-Torres R, Scelo G, Purdue MP, Mariosa D, Muller DC, Timpson NJ, Haycock PC, Brown KM, Wang Z, Ye Y, Hofmann JN, Foll M, Gaborieau V, Machiela MJ, Colli LM, Li P, Garnier J-G, Blanche H, Boland A, Burdette L, Prokhortchouk E, Skryabin KG, Yeager M, Radojevic-Skodric S, Ognjanovic S, Foretova L, Holcatova I, Janout V, Mates D, Mukeriya A, Rascu S, Zaridze D, Bencko V, Cybulski C, Fabianova E, Jinga V, Lissowska J, Lubinski J, Navratilova M, Rudnai P, Benhamou S, Cancel-Tassin G, Cussenot O, Weiderpass E, Ljungberg B, Tumkur Sitaram R, Häggström C, Bruinsma F, Jordan SJ, Severi G, Winship I, Hveem K, Vatten LJ, Fletcher T, Larsson SC, Wolk A, Banks RE, Selby PJ, Easton DF, Andreotti G, Beane Freeman LE, Koutros S, Männistö S, Weinstein S, Clark PE, Edwards TL, Lipworth L, Gapstur SM, Stevens VL, Carol H, Freedman ML, Pomerantz MM, Cho E, Wilson KM, Gaziano JM, Sesso HD, Freedman ND, Parker AS, Eckel-Passow JE, Huang W-Y, Kahnoski RJ, Lane BR, Noyes SL, Petillo D, Teh BT, Peters U, White E, Anderson GL, Johnson L, Luo J, Buring J, Lee I-M, Chow W-H, Moore LE, Eisen T, Henrion M, Larkin J, Barman P, Leibovich BC, Choueiri TK, Lathrop GM, Deleuze J-F, Gunter M, McKay JD, Wu X, Houlston RS, Chanock SJ, Relton C, Richards JB, Martin RM, Davey Smith G, Brennan P. The influence of obesity-related factors in the etiology of renal cell carcinoma-A mendelian randomization study. PLOS Medicine. 2019;16:e1002724. doi: 10.1371/journal.pmed.1002724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones GT, Tromp G, Kuivaniemi H, Gretarsdottir S, Baas AF, Giusti B, Strauss E, Van’t Hof FNG, Webb TR, Erdman R, Ritchie MD, Elmore JR, Verma A, Pendergrass S, Kullo IJ, Ye Z, Peissig PL, Gottesman O, Verma SS, Malinowski J, Rasmussen-Torvik LJ, Borthwick KM, Smelser DT, Crosslin DR, de Andrade M, Ryer EJ, McCarty CA, Böttinger EP, Pacheco JA, Crawford DC, Carrell DS, Gerhard GS, Franklin DP, Carey DJ, Phillips VL, Williams MJA, Wei W, Blair R, Hill AA, Vasudevan TM, Lewis DR, Thomson IA, Krysa J, Hill GB, Roake J, Merriman TR, Oszkinis G, Galora S, Saracini C, Abbate R, Pulli R, Pratesi C, Saratzis A, Verissimo AR, Bumpstead S, Badger SA, Clough RE, Cockerill G, Hafez H, Scott DJA, Futers TS, Romaine SPR, Bridge K, Griffin KJ, Bailey MA, Smith A, Thompson MM, van Bockxmeer FM, Matthiasson SE, Thorleifsson G, Thorsteinsdottir U, Blankensteijn JD, Teijink JAW, Wijmenga C, de Graaf J, Kiemeney LA, Lindholt JS, Hughes A, Bradley DT, Stirrups K, Golledge J, Norman PE, Powell JT, Humphries SE, Hamby SE, Goodall AH, Nelson CP, Sakalihasan N, Courtois A, Ferrell RE, Eriksson P, Folkersen L, Franco-Cereceda A, Eicher JD, Johnson AD, Betsholtz C, Ruusalepp A, Franzén O, Schadt EE, Björkegren JLM, Lipovich L, Drolet AM, Verhoeven EL, Zeebregts CJ, Geelkerken RH, van Sambeek MR, van Sterkenburg SM, de Vries J-P, Stefansson K, Thompson JR, de Bakker PIW, Deloukas P, Sayers RD, Harrison SC, van Rij AM, Samani NJ, Bown MJ. Meta-Analysis of Genome-Wide Association Studies for Abdominal Aortic Aneurysm Identifies Four New Disease-Specific Risk Loci. Circulation Research. 2017;120:341–353. doi: 10.1161/CIRCRESAHA.116.308765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kazmi N, Haycock P, Tsilidis K, Lynch BM, Truong T, PRACTICAL Consortium, CRUK, BPC3, CAPS, PEGASUS. Martin RM, Lewis SJ. Appraising causal relationships of dietary, nutritional and physical-activity exposures with overall and aggressive prostate cancer: two-sample Mendelian-randomization study based on 79 148 prostate-cancer cases and 61 106 controls. International Journal of Epidemiology. 2020;49:587–596. doi: 10.1093/ije/dyz235. [DOI] [PubMed] [Google Scholar]
- Kilpeläinen TO, Zillikens MC, Stančákova A, Finucane FM, Ried JS, Langenberg C, Zhang W, Beckmann JS, Luan J, Vandenput L, Styrkarsdottir U, Zhou Y, Smith AV, Zhao J-H, Amin N, Vedantam S, Shin S-Y, Haritunians T, Fu M, Feitosa MF, Kumari M, Halldorsson BV, Tikkanen E, Mangino M, Hayward C, Song C, Arnold AM, Aulchenko YS, Oostra BA, Campbell H, Cupples LA, Davis KE, Döring A, Eiriksdottir G, Estrada K, Fernández-Real JM, Garcia M, Gieger C, Glazer NL, Guiducci C, Hofman A, Humphries SE, Isomaa B, Jacobs LC, Jula A, Karasik D, Karlsson MK, Khaw K-T, Kim LJ, Kivimäki M, Klopp N, Kühnel B, Kuusisto J, Liu Y, Ljunggren O, Lorentzon M, Luben RN, McKnight B, Mellström D, Mitchell BD, Mooser V, Moreno JM, Männistö S, O’Connell JR, Pascoe L, Peltonen L, Peral B, Perola M, Psaty BM, Salomaa V, Savage DB, Semple RK, Skaric-Juric T, Sigurdsson G, Song KS, Spector TD, Syvänen A-C, Talmud PJ, Thorleifsson G, Thorsteinsdottir U, Uitterlinden AG, van Duijn CM, Vidal-Puig A, Wild SH, Wright AF, Clegg DJ, Schadt E, Wilson JF, Rudan I, Ripatti S, Borecki IB, Shuldiner AR, Ingelsson E, Jansson J-O, Kaplan RC, Gudnason V, Harris TB, Groop L, Kiel DP, Rivadeneira F, Walker M, Barroso I, Vollenweider P, Waeber G, Chambers JC, Kooner JS, Soranzo N, Hirschhorn JN, Stefansson K, Wichmann H-E, Ohlsson C, O’Rahilly S, Wareham NJ, Speliotes EK, Fox CS, Laakso M, Loos RJF. Genetic variation near IRS1 associates with reduced adiposity and an impaired metabolic profile. Nature Genetics. 2011;43:753–760. doi: 10.1038/ng.866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkle BW, Grenier-Boley B, Sims R, Bis JC, Damotte V, Naj AC, Boland A, Vronskaya M, van der Lee SJ, Amlie-Wolf A, Bellenguez C, Frizatti A, Chouraki V, Martin ER, Sleegers K, Badarinarayan N, Jakobsdottir J, Hamilton-Nelson KL, Moreno-Grau S, Olaso R, Raybould R, Chen Y, Kuzma AB, Hiltunen M, Morgan T, Ahmad S, Vardarajan BN, Epelbaum J, Hoffmann P, Boada M, Beecham GW, Garnier J-G, Harold D, Fitzpatrick AL, Valladares O, Moutet M-L, Gerrish A, Smith AV, Qu L, Bacq D, Denning N, Jian X, Zhao Y, Del Zompo M, Fox NC, Choi S-H, Mateo I, Hughes JT, Adams HH, Malamon J, Sanchez-Garcia F, Patel Y, Brody JA, Dombroski BA, Naranjo MCD, Daniilidou M, Eiriksdottir G, Mukherjee S, Wallon D, Uphill J, Aspelund T, Cantwell LB, Garzia F, Galimberti D, Hofer E, Butkiewicz M, Fin B, Scarpini E, Sarnowski C, Bush WS, Meslage S, Kornhuber J, White CC, Song Y, Barber RC, Engelborghs S, Sordon S, Voijnovic D, Adams PM, Vandenberghe R, Mayhaus M, Cupples LA, Albert MS, De Deyn PP, Gu W, Himali JJ, Beekly D, Squassina A, Hartmann AM, Orellana A, Blacker D, Rodriguez-Rodriguez E, Lovestone S, Garcia ME, Doody RS, Munoz-Fernadez C, Sussams R, Lin H, Fairchild TJ, Benito YA, Holmes C, Karamujić-Čomić H, Frosch MP, Thonberg H, Maier W, Roshchupkin G, Ghetti B, Giedraitis V, Kawalia A, Li S, Huebinger RM, Kilander L, Moebus S, Hernández I, Kamboh MI, Brundin R, Turton J, Yang Q, Katz MJ, Concari L, Lord J, Beiser AS, Keene CD, Helisalmi S, Kloszewska I, Kukull WA, Koivisto AM, Lynch A, Tarraga L, Larson EB, Haapasalo A, Lawlor B, Mosley TH, Lipton RB, Solfrizzi V, Gill M, Longstreth WT, Montine TJ, Frisardi V, Diez-Fairen M, Rivadeneira F, Petersen RC, Deramecourt V, Alvarez I, Salani F, Ciaramella A, Boerwinkle E, Reiman EM, Fievet N, Rotter JI, Reisch JS, Hanon O, Cupidi C, Andre Uitterlinden AG, Royall DR, Dufouil C, Maletta RG, de Rojas I, Sano M, Brice A, Cecchetti R, George-Hyslop PS, Ritchie K, Tsolaki M, Tsuang DW, Dubois B, Craig D, Wu C-K, Soininen H, Avramidou D, Albin RL, Fratiglioni L, Germanou A, Apostolova LG, Keller L, Koutroumani M, Arnold SE, Panza F, Gkatzima O, Asthana S, Hannequin D, Whitehead P, Atwood CS, Caffarra P, Hampel H, Quintela I, Carracedo Á, Lannfelt L, Rubinsztein DC, Barnes LL, Pasquier F, Frölich L, Barral S, McGuinness B, Beach TG, Johnston JA, Becker JT, Passmore P, Bigio EH, Schott JM, Bird TD, Warren JD, Boeve BF, Lupton MK, Bowen JD, Proitsi P, Boxer A, Powell JF, Burke JR, Kauwe JSK, Burns JM, Mancuso M, Buxbaum JD, Bonuccelli U, Cairns NJ, McQuillin A, Cao C, Livingston G, Carlson CS, Bass NJ, Carlsson CM, Hardy J, Carney RM, Bras J, Carrasquillo MM, Guerreiro R, Allen M, Chui HC, Fisher E, Masullo C, Crocco EA, DeCarli C, Bisceglio G, Dick M, Ma L, Duara R, Graff-Radford NR, Evans DA, Hodges A, Faber KM, Scherer M, Fallon KB, Riemenschneider M, Fardo DW, Heun R, Farlow MR, Kölsch H, Ferris S, Leber M, Foroud TM, Heuser I, Galasko DR, Giegling I, Gearing M, Hüll M, Geschwind DH, Gilbert JR, Morris J, Green RC, Mayo K, Growdon JH, Feulner T, Hamilton RL, Harrell LE, Drichel D, Honig LS, Cushion TD, Huentelman MJ, Hollingworth P, Hulette CM, Hyman BT, Marshall R, Jarvik GP, Meggy A, Abner E, Menzies GE, Jin L-W, Leonenko G, Real LM, Jun GR, Baldwin CT, Grozeva D, Karydas A, Russo G, Kaye JA, Kim R, Jessen F, Kowall NW, Vellas B, Kramer JH, Vardy E, LaFerla FM, Jöckel K-H, Lah JJ, Dichgans M, Leverenz JB, Mann D, Levey AI, Pickering-Brown S, Lieberman AP, Klopp N, Lunetta KL, Wichmann H-E, Lyketsos CG, Morgan K, Marson DC, Brown K, Martiniuk F, Medway C, Mash DC, Nöthen MM, Masliah E, Hooper NM, McCormick WC, Daniele A, McCurry SM, Bayer A, McDavid AN, Gallacher J, McKee AC, van den Bussche H, Mesulam M, Brayne C, Miller BL, Riedel-Heller S, Miller CA, Miller JW, Al-Chalabi A, Morris JC, Shaw CE, Myers AJ, Wiltfang J, O’Bryant S, Olichney JM, Alvarez V, Parisi JE, Singleton AB, Paulson HL, Collinge J, Perry WR, Mead S, Peskind E, Cribbs DH, Rossor M, Pierce A, Ryan NS, Poon WW, Nacmias B, Potter H, Sorbi S, Quinn JF, Sacchinelli E, Raj A, Spalletta G, Raskind M, Caltagirone C, Bossù P, Orfei MD, Reisberg B, Clarke R, Reitz C, Smith AD, Ringman JM, Warden D, Roberson ED, Wilcock G, Rogaeva E, Bruni AC, Rosen HJ, Gallo M, Rosenberg RN, Ben-Shlomo Y, Sager MA, Mecocci P, Saykin AJ, Pastor P, Cuccaro ML, Vance JM, Schneider JA, Schneider LS, Slifer S, Seeley WW, Smith AG, Sonnen JA, Spina S, Stern RA, Swerdlow RH, Tang M, Tanzi RE, Trojanowski JQ, Troncoso JC, Van Deerlin VM, Van Eldik LJ, Vinters HV, Vonsattel JP, Weintraub S, Welsh-Bohmer KA, Wilhelmsen KC, Williamson J, Wingo TS, Woltjer RL, Wright CB, Yu C-E, Yu L, Saba Y, Pilotto A, Bullido MJ, Peters O, Crane PK, Bennett D, Bosco P, Coto E, Boccardi V, De Jager PL, Lleo A, Warner N, Lopez OL, Ingelsson M, Deloukas P, Cruchaga C, Graff C, Gwilliam R, Fornage M, Goate AM, Sanchez-Juan P, Kehoe PG, Amin N, Ertekin-Taner N, Berr C, Debette S, Love S, Launer LJ, Younkin SG, Dartigues J-F, Corcoran C, Ikram MA, Dickson DW, Nicolas G, Campion D, Tschanz J, Schmidt H, Hakonarson H, Clarimon J, Munger R, Schmidt R, Farrer LA, Van Broeckhoven C, C O’Donovan M, DeStefano AL, Jones L, Haines JL, Deleuze J-F, Owen MJ, Gudnason V, Mayeux R, Escott-Price V, Psaty BM, Ramirez A, Wang L-S, Ruiz A, van Duijn CM, Holmans PA, Seshadri S, Williams J, Amouyel P, Schellenberg GD, Lambert J-C, Pericak-Vance MA, Alzheimer Disease Genetics Consortium (ADGC) European Alzheimer’s Disease Initiative (EADI) Cohorts for Heart and Aging Research in Genomic Epidemiology Consortium (CHARGE) Genetic and Environmental Risk in AD/Defining Genetic, Polygenic and Environmental Risk for Alzheimer’s Disease Consortium (GERAD/PERADES) Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nature Genetics. 2019;51:414–430. doi: 10.1038/s41588-019-0358-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsson SC, Traylor M, Malik R, Dichgans M, Burgess S, Markus HS, CoSTREAM Consortium, on behalf of the International Genomics of Alzheimer’s Project Modifiable pathways in Alzheimer’s disease: Mendelian randomisation analysis. BMJ. 2017;359:j5375. doi: 10.1136/bmj.j5375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsson SC, Burgess S, Michaëlsson K. Genetic association between adiposity and gout: a Mendelian randomization study. Rheumatology. 2018;57:2145–2148. doi: 10.1093/rheumatology/key229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsson SC, Bäck M, Rees JMB, Mason AM, Burgess S. Body mass index and body composition in relation to 14 cardiovascular conditions in UK Biobank: a Mendelian randomization study. European Heart Journal. 2020;41:221–226. doi: 10.1093/eurheartj/ehz388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Law PJ, Timofeeva M, Fernandez-Rozadilla C, Broderick P, Studd J, Fernandez-Tajes J, Farrington S, Svinti V, Palles C, Orlando G, Sud A, Holroyd A, Penegar S, Theodoratou E, Vaughan-Shaw P, Campbell H, Zgaga L, Hayward C, Campbell A, Harris S, Deary IJ, Starr J, Gatcombe L, Pinna M, Briggs S, Martin L, Jaeger E, Sharma-Oates A, East J, Leedham S, Arnold R, Johnstone E, Wang H, Kerr D, Kerr R, Maughan T, Kaplan R, Al-Tassan N, Palin K, Hänninen UA, Cajuso T, Tanskanen T, Kondelin J, Kaasinen E, Sarin A-P, Eriksson JG, Rissanen H, Knekt P, Pukkala E, Jousilahti P, Salomaa V, Ripatti S, Palotie A, Renkonen-Sinisalo L, Lepistö A, Böhm J, Mecklin J-P, Buchanan DD, Win A-K, Hopper J, Jenkins ME, Lindor NM, Newcomb PA, Gallinger S, Duggan D, Casey G, Hoffmann P, Nöthen MM, Jöckel K-H, Easton DF, Pharoah PDP, Peto J, Canzian F, Swerdlow A, Eeles RA, Kote-Jarai Z, Muir K, Pashayan N, PRACTICAL consortium. Harkin A, Allan K, McQueen J, Paul J, Iveson T, Saunders M, Butterbach K, Chang-Claude J, Hoffmeister M, Brenner H, Kirac I, Matošević P, Hofer P, Brezina S, Gsur A, Cheadle JP, Aaltonen LA, Tomlinson I, Houlston RS, Dunlop MG. Association analyses identify 31 new risk loci for colorectal cancer susceptibility. Nature Communications. 2019;10:2154. doi: 10.1038/s41467-019-09775-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindström S, Wang L, Smith EN, Gordon W, van Hylckama Vlieg A, de Andrade M, Brody JA, Pattee JW, Haessler J, Brumpton BM, Chasman DI, Suchon P, Chen M-H, Turman C, Germain M, Wiggins KL, MacDonald J, Braekkan SK, Armasu SM, Pankratz N, Jackson RD, Nielsen JB, Giulianini F, Puurunen MK, Ibrahim M, Heckbert SR, Damrauer SM, Natarajan P, Klarin D, Million Veteran Program. de Vries PS, Sabater-Lleal M, Huffman JE, CHARGE Hemostasis Working Group. Bammler TK, Frazer KA, McCauley BM, Taylor K, Pankow JS, Reiner AP, Gabrielsen ME, Deleuze J-F, O’Donnell CJ, Kim J, McKnight B, Kraft P, Hansen J-B, Rosendaal FR, Heit JA, Psaty BM, Tang W, Kooperberg C, Hveem K, Ridker PM, Morange P-E, Johnson AD, Kabrhel C, Trégouët D-A, Smith NL. Genomic and transcriptomic association studies identify 16 novel susceptibility loci for venous thromboembolism. Blood. 2019;134:1645–1657. doi: 10.1182/blood.2019000435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Livingston G, Huntley J, Sommerlad A, Ames D, Ballard C, Banerjee S, Brayne C, Burns A, Cohen-Mansfield J, Cooper C, Costafreda SG, Dias A, Fox N, Gitlin LN, Howard R, Kales HC, Kivimäki M, Larson EB, Ogunniyi A, Orgeta V, Ritchie K, Rockwood K, Sampson EL, Samus Q, Schneider LS, Selbæk G, Teri L, Mukadam N. Dementia prevention, intervention, and care: 2020 report of the Lancet Commission. Lancet. 2020;396:413–446. doi: 10.1016/S0140-6736(20)30367-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Locke AE, Kahali B, Berndt SI, Justice AE, Pers TH, Day FR, Powell C, Vedantam S, Buchkovich ML, Yang J, Croteau-Chonka DC, Esko T, Fall T, Ferreira T, Gustafsson S, Kutalik Z, Luan J, Mägi R, Randall JC, Winkler TW, Wood AR, Workalemahu T, Faul JD, Smith JA, Zhao JH, Zhao W, Chen J, Fehrmann R, Hedman ÅK, Karjalainen J, Schmidt EM, Absher D, Amin N, Anderson D, Beekman M, Bolton JL, Bragg-Gresham JL, Buyske S, Demirkan A, Deng G, Ehret GB, Feenstra B, Feitosa MF, Fischer K, Goel A, Gong J, Jackson AU, Kanoni S, Kleber ME, Kristiansson K, Lim U, Lotay V, Mangino M, Leach IM, Medina-Gomez C, Medland SE, Nalls MA, Palmer CD, Pasko D, Pechlivanis S, Peters MJ, Prokopenko I, Shungin D, Stančáková A, Strawbridge RJ, Sung YJ, Tanaka T, Teumer A, Trompet S, van der Laan SW, van Setten J, Van Vliet-Ostaptchouk JV, Wang Z, Yengo L, Zhang W, Isaacs A, Albrecht E, Ärnlöv J, Arscott GM, Attwood AP, Bandinelli S, Barrett A, Bas IN, Bellis C, Bennett AJ, Berne C, Blagieva R, Blüher M, Böhringer S, Bonnycastle LL, Böttcher Y, Boyd HA, Bruinenberg M, Caspersen IH, Chen Y-DI, Clarke R, Daw EW, de Craen AJM, Delgado G, Dimitriou M, Doney ASF, Eklund N, Estrada K, Eury E, Folkersen L, Fraser RM, Garcia ME, Geller F, Giedraitis V, Gigante B, Go AS, Golay A, Goodall AH, Gordon SD, Gorski M, Grabe H-J, Grallert H, Grammer TB, Gräßler J, Grönberg H, Groves CJ, Gusto G, Haessler J, Hall P, Haller T, Hallmans G, Hartman CA, Hassinen M, Hayward C, Heard-Costa NL, Helmer Q, Hengstenberg C, Holmen O, Hottenga J-J, James AL, Jeff JM, Johansson Å, Jolley J, Juliusdottir T, Kinnunen L, Koenig W, Koskenvuo M, Kratzer W, Laitinen J, Lamina C, Leander K, Lee NR, Lichtner P, Lind L, Lindström J, Lo KS, Lobbens S, Lorbeer R, Lu Y, Mach F, Magnusson PKE, Mahajan A, McArdle WL, McLachlan S, Menni C, Merger S, Mihailov E, Milani L, Moayyeri A, Monda KL, Morken MA, Mulas A, Müller G, Müller-Nurasyid M, Musk AW, Nagaraja R, Nöthen MM, Nolte IM, Pilz S, Rayner NW, Renstrom F, Rettig R, Ried JS, Ripke S, Robertson NR, Rose LM, Sanna S, Scharnagl H, Scholtens S, Schumacher FR, Scott WR, Seufferlein T, Shi J, Smith AV, Smolonska J, Stanton AV, Steinthorsdottir V, Stirrups K, Stringham HM, Sundström J, Swertz MA, Swift AJ, Syvänen A-C, Tan S-T, Tayo BO, Thorand B, Thorleifsson G, Tyrer JP, Uh H-W, Vandenput L, Verhulst FC, Vermeulen SH, Verweij N, Vonk JM, Waite LL, Warren HR, Waterworth D, Weedon MN, Wilkens LR, Willenborg C, Wilsgaard T, Wojczynski MK, Wong A, Wright AF, Zhang Q, LifeLines Cohort Study. Brennan EP, Choi M, Dastani Z, Drong AW, Eriksson P, Franco-Cereceda A, Gådin JR, Gharavi AG, Goddard ME, Handsaker RE, Huang J, Karpe F, Kathiresan S, Keildson S, Kiryluk K, Kubo M, Lee J-Y, Liang L, Lifton RP, Ma B, McCarroll SA, McKnight AJ, Min JL, Moffatt MF, Montgomery GW, Murabito JM, Nicholson G, Nyholt DR, Okada Y, Perry JRB, Dorajoo R, Reinmaa E, Salem RM, Sandholm N, Scott RA, Stolk L, Takahashi A, Tanaka T, van ’t Hooft FM, Vinkhuyzen AAE, Westra H-J, Zheng W, Zondervan KT, ADIPOGen Consortium. AGEN-BMI Working Group. CARDIOGRAMplusC4D Consortium. CKDGen Consortium. GLGC. ICBP. MAGIC Investigators. MuTHER Consortium. MIGen Consortium. PAGE Consortium. ReproGen Consortium. GENIE Consortium. International Endogene Consortium. Heath AC, Arveiler D, Bakker SJL, Beilby J, Bergman RN, Blangero J, Bovet P, Campbell H, Caulfield MJ, Cesana G, Chakravarti A, Chasman DI, Chines PS, Collins FS, Crawford DC, Cupples LA, Cusi D, Danesh J, de Faire U, den Ruijter HM, Dominiczak AF, Erbel R, Erdmann J, Eriksson JG, Farrall M, Felix SB, Ferrannini E, Ferrières J, Ford I, Forouhi NG, Forrester T, Franco OH, Gansevoort RT, Gejman PV, Gieger C, Gottesman O, Gudnason V, Gyllensten U, Hall AS, Harris TB, Hattersley AT, Hicks AA, Hindorff LA, Hingorani AD, Hofman A, Homuth G, Hovingh GK, Humphries SE, Hunt SC, Hyppönen E, Illig T, Jacobs KB, Jarvelin M-R, Jöckel K-H, Johansen B, Jousilahti P, Jukema JW, Jula AM, Kaprio J, Kastelein JJP, Keinanen-Kiukaanniemi SM, Kiemeney LA, Knekt P, Kooner JS, Kooperberg C, Kovacs P, Kraja AT, Kumari M, Kuusisto J, Lakka TA, Langenberg C, Marchand LL, Lehtimäki T, Lyssenko V, Männistö S, Marette A, Matise TC, McKenzie CA, McKnight B, Moll FL, Morris AD, Morris AP, Murray JC, Nelis M, Ohlsson C, Oldehinkel AJ, Ong KK, Madden PAF, Pasterkamp G, Peden JF, Peters A, Postma DS, Pramstaller PP, Price JF, Qi L, Raitakari OT, Rankinen T, Rao DC, Rice TK, Ridker PM, Rioux JD, Ritchie MD, Rudan I, Salomaa V, Samani NJ, Saramies J, Sarzynski MA, Schunkert H, Schwarz PEH, Sever P, Shuldiner AR, Sinisalo J, Stolk RP, Strauch K, Tönjes A, Trégouët D-A, Tremblay A, Tremoli E, Virtamo J, Vohl M-C, Völker U, Waeber G, Willemsen G, Witteman JC, Zillikens MC, Adair LS, Amouyel P, Asselbergs FW, Assimes TL, Bochud M, Boehm BO, Boerwinkle E, Bornstein SR, Bottinger EP, Bouchard C, Cauchi S, Chambers JC, Chanock SJ, Cooper RS, de Bakker PIW, Dedoussis G, Ferrucci L, Franks PW, Froguel P, Groop LC, Haiman CA, Hamsten A, Hui J, Hunter DJ, Hveem K, Kaplan RC, Kivimaki M, Kuh D, Laakso M, Liu Y, Martin NG, März W, Melbye M, Metspalu A, Moebus S, Munroe PB, Njølstad I, Oostra BA, Palmer CNA, Pedersen NL, Perola M, Pérusse L, Peters U, Power C, Quertermous T, Rauramaa R, Rivadeneira F, Saaristo TE, Saleheen D, Sattar N, Schadt EE, Schlessinger D, Slagboom PE, Snieder H, Spector TD, Thorsteinsdottir U, Stumvoll M, Tuomilehto J, Uitterlinden AG, Uusitupa M, van der Harst P, Walker M, Wallaschofski H, Wareham NJ, Watkins H, Weir DR, Wichmann H-E, Wilson JF, Zanen P, Borecki IB, Deloukas P, Fox CS, Heid IM, O’Connell JR, Strachan DP, Stefansson K, van Duijn CM, Abecasis GR, Franke L, Frayling TM, McCarthy MI, Visscher PM, Scherag A, Willer CJ, Boehnke M, Mohlke KL, Lindgren CM, Beckmann JS, Barroso I, North KE, Ingelsson E, Hirschhorn JN, Loos RJF, Speliotes EK. Genetic studies of body mass index yield new insights for obesity biology. Nature. 2015;518:197–206. doi: 10.1038/nature14177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loh P-R, Tucker G, Bulik-Sullivan BK, Vilhjálmsson BJ, Finucane HK, Salem RM, Chasman DI, Ridker PM, Neale BM, Berger B, Patterson N, Price AL. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nature Genetics. 2015;47:284–290. doi: 10.1038/ng.3190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorenzet R, Napoleone E, Cutrone A, Donati MB. Thrombosis and obesity: cellular bases. Thrombosis Research. 2012;129:285–289. doi: 10.1016/j.thromres.2011.10.021. [DOI] [PubMed] [Google Scholar]
- Lotta LA, Gulati P, Day FR, Payne F, Ongen H, van de Bunt M, Gaulton KJ, Eicher JD, Sharp SJ, Luan J, De Lucia Rolfe E, Stewart ID, Wheeler E, Willems SM, Adams C, Yaghootkar H, EPIC-InterAct Consortium. Cambridge FPLD1 Consortium. Forouhi NG, Khaw K-T, Johnson AD, Semple RK, Frayling T, Perry JRB, Dermitzakis E, McCarthy MI, Barroso I, Wareham NJ, Savage DB, Langenberg C, O’Rahilly S, Scott RA. Integrative genomic analysis implicates limited peripheral adipose storage capacity in the pathogenesis of human insulin resistance. Nature Genetics. 2017;49:17–26. doi: 10.1038/ng.3714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maglio C, Peltonen M, Neovius M, Jacobson P, Jacobsson L, Rudin A, Carlsson LMS. Effects of bariatric surgery on gout incidence in the Swedish Obese Subjects study: a non-randomised, prospective, controlled intervention trial. Annals of the Rheumatic Diseases. 2017;76:688–693. doi: 10.1136/annrheumdis-2016-209958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mahajan A, Taliun D, Thurner M, Robertson NR, Torres JM, Rayner NW, Payne AJ, Steinthorsdottir V, Scott RA, Grarup N, Cook JP, Schmidt EM, Wuttke M, Sarnowski C, Mägi R, Nano J, Gieger C, Trompet S, Lecoeur C, Preuss MH, Prins BP, Guo X, Bielak LF, Below JE, Bowden DW, Chambers JC, Kim YJ, Ng MCY, Petty LE, Sim X, Zhang W, Bennett AJ, Bork-Jensen J, Brummett CM, Canouil M, Ec Kardt K-U, Fischer K, Kardia SLR, Kronenberg F, Läll K, Liu C-T, Locke AE, Luan J, Ntalla I, Nylander V, Schönherr S, Schurmann C, Yengo L, Bottinger EP, Brandslund I, Christensen C, Dedoussis G, Florez JC, Ford I, Franco OH, Frayling TM, Giedraitis V, Hackinger S, Hattersley AT, Herder C, Ikram MA, Ingelsson M, Jørgensen ME, Jørgensen T, Kriebel J, Kuusisto J, Ligthart S, Lindgren CM, Linneberg A, Lyssenko V, Mamakou V, Meitinger T, Mohlke KL, Morris AD, Nadkarni G, Pankow JS, Peters A, Sattar N, Stančáková A, Strauch K, Taylor KD, Thorand B, Thorleifsson G, Thorsteinsdottir U, Tuomilehto J, Witte DR, Dupuis J, Peyser PA, Zeggini E, Loos RJF, Froguel P, Ingelsson E, Lind L, Groop L, Laakso M, Collins FS, Jukema JW, Palmer CNA, Grallert H, Metspalu A, Dehghan A, Köttgen A, Abecasis GR, Meigs JB, Rotter JI, Marchini J, Pedersen O, Hansen T, Langenberg C, Wareham NJ, Stefansson K, Gloyn AL, Morris AP, Boehnke M, McCarthy MI. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nature Genetics. 2018;50:1505–1513. doi: 10.1038/s41588-018-0241-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malik R, Chauhan G, Traylor M, Sargurupremraj M, Okada Y, Mishra A, Rutten-Jacobs L, Giese A-K, van der Laan SW, Gretarsdottir S, Anderson CD, Chong M, Adams HHH, Ago T, Almgren P, Amouyel P, Ay H, Bartz TM, Benavente OR, Bevan S, Boncoraglio GB, Brown RD, Butterworth AS, Carrera C, Carty CL, Chasman DI, Chen W-M, Cole JW, Correa A, Cotlarciuc I, Cruchaga C, Danesh J, de Bakker PIW, DeStefano AL, den Hoed M, Duan Q, Engelter ST, Falcone GJ, Gottesman RF, Grewal RP, Gudnason V, Gustafsson S, Haessler J, Harris TB, Hassan A, Havulinna AS, Heckbert SR, Holliday EG, Howard G, Hsu F-C, Hyacinth HI, Ikram MA, Ingelsson E, Irvin MR, Jian X, Jiménez-Conde J, Johnson JA, Jukema JW, Kanai M, Keene KL, Kissela BM, Kleindorfer DO, Kooperberg C, Kubo M, Lange LA, Langefeld CD, Langenberg C, Launer LJ, Lee J-M, Lemmens R, Leys D, Lewis CM, Lin W-Y, Lindgren AG, Lorentzen E, Magnusson PK, Maguire J, Manichaikul A, McArdle PF, Meschia JF, Mitchell BD, Mosley TH, Nalls MA, Ninomiya T, O’Donnell MJ, Psaty BM, Pulit SL, Rannikmäe K, Reiner AP, Rexrode KM, Rice K, Rich SS, Ridker PM, Rost NS, Rothwell PM, Rotter JI, Rundek T, Sacco RL, Sakaue S, Sale MM, Salomaa V, Sapkota BR, Schmidt R, Schmidt CO, Schminke U, Sharma P, Slowik A, Sudlow CLM, Tanislav C, Tatlisumak T, Taylor KD, Thijs VNS, Thorleifsson G, Thorsteinsdottir U, Tiedt S, Trompet S, Tzourio C, van Duijn CM, Walters M, Wareham NJ, Wassertheil-Smoller S, Wilson JG, Wiggins KL, Yang Q, Yusuf S, AFGen Consortium. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium. International Genomics of Blood Pressure (iGEN-BP) Consortium. INVENT Consortium. STARNET. Bis JC, Pastinen T, Ruusalepp A, Schadt EE, Koplev S, Björkegren JLM, Codoni V, Civelek M, Smith NL, Trégouët DA, Christophersen IE, Roselli C, Lubitz SA, Ellinor PT, Tai ES, Kooner JS, Kato N, He J, van der Harst P, Elliott P, Chambers JC, Takeuchi F, Johnson AD, BioBank Japan Cooperative Hospital Group. COMPASS Consortium. EPIC-CVD Consortium. EPIC-InterAct Consortium. International Stroke Genetics Consortium (ISGC) METASTROKE Consortium. Neurology Working Group of the CHARGE Consortium. NINDS Stroke Genetics Network (SiGN) UK Young Lacunar DNA Study. MEGASTROKE Consortium. Sanghera DK, Melander O, Jern C, Strbian D, Fernandez-Cadenas I, Longstreth WT, Rolfs A, Hata J, Woo D, Rosand J, Pare G, Hopewell JC, Saleheen D, Stefansson K, Worrall BB, Kittner SJ, Seshadri S, Fornage M, Markus HS, Howson JMM, Kamatani Y, Debette S, Dichgans M. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nature Genetics. 2018;50:524–537. doi: 10.1038/s41588-018-0058-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mariosa D, Carreras-Torres R, Martin RM, Johansson M, Brennan P. Commentary: What can Mendelian randomization tell us about causes of cancer? International Journal of Epidemiology. 2019;48:816–821. doi: 10.1093/ije/dyz151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin S, Cule M, Basty N, Tyrrell J, Beaumont RN, Wood AR, Frayling TM, Sorokin E, Whitcher B, Liu Y, Bell JD, Thomas EL, Yaghootkar H. Genetic Evidence for Different Adiposity Phenotypes and Their Opposing Influences on Ectopic Fat and Risk of Cardiometabolic Disease. Diabetes. 2021;70:1843–1856. doi: 10.2337/db21-0129. [DOI] [PubMed] [Google Scholar]
- Mathus-Vliegen EMH, Van Ierland-Van Leeuwen ML, Terpstra A. Determinants of gallbladder kinetics in obesity. Digestive Diseases and Sciences. 2004;49:9–16. doi: 10.1023/b:ddas.0000011595.39555.c0. [DOI] [PubMed] [Google Scholar]
- Michailidou K, Lindström S, Dennis J, Beesley J, Hui S, Kar S, Lemaçon A, Soucy P, Glubb D, Rostamianfar A, Bolla MK, Wang Q, Tyrer J, Dicks E, Lee A, Wang Z, Allen J, Keeman R, Eilber U, French JD, Qing Chen X, Fachal L, McCue K, McCart Reed AE, Ghoussaini M, Carroll JS, Jiang X, Finucane H, Adams M, Adank MA, Ahsan H, Aittomäki K, Anton-Culver H, Antonenkova NN, Arndt V, Aronson KJ, Arun B, Auer PL, Bacot F, Barrdahl M, Baynes C, Beckmann MW, Behrens S, Benitez J, Bermisheva M, Bernstein L, Blomqvist C, Bogdanova NV, Bojesen SE, Bonanni B, Børresen-Dale A-L, Brand JS, Brauch H, Brennan P, Brenner H, Brinton L, Broberg P, Brock IW, Broeks A, Brooks-Wilson A, Brucker SY, Brüning T, Burwinkel B, Butterbach K, Cai Q, Cai H, Caldés T, Canzian F, Carracedo A, Carter BD, Castelao JE, Chan TL, David Cheng T-Y, Seng Chia K, Choi J-Y, Christiansen H, Clarke CL, NBCS Collaborators. Collée M, Conroy DM, Cordina-Duverger E, Cornelissen S, Cox DG, Cox A, Cross SS, Cunningham JM, Czene K, Daly MB, Devilee P, Doheny KF, Dörk T, Dos-Santos-Silva I, Dumont M, Durcan L, Dwek M, Eccles DM, Ekici AB, Eliassen AH, Ellberg C, Elvira M, Engel C, Eriksson M, Fasching PA, Figueroa J, Flesch-Janys D, Fletcher O, Flyger H, Fritschi L, Gaborieau V, Gabrielson M, Gago-Dominguez M, Gao Y-T, Gapstur SM, García-Sáenz JA, Gaudet MM, Georgoulias V, Giles GG, Glendon G, Goldberg MS, Goldgar DE, González-Neira A, Grenaker Alnæs GI, Grip M, Gronwald J, Grundy A, Guénel P, Haeberle L, Hahnen E, Haiman CA, Håkansson N, Hamann U, Hamel N, Hankinson S, Harrington P, Hart SN, Hartikainen JM, Hartman M, Hein A, Heyworth J, Hicks B, Hillemanns P, Ho DN, Hollestelle A, Hooning MJ, Hoover RN, Hopper JL, Hou M-F, Hsiung C-N, Huang G, Humphreys K, Ishiguro J, Ito H, Iwasaki M, Iwata H, Jakubowska A, Janni W, John EM, Johnson N, Jones K, Jones M, Jukkola-Vuorinen A, Kaaks R, Kabisch M, Kaczmarek K, Kang D, Kasuga Y, Kerin MJ, Khan S, Khusnutdinova E, Kiiski JI, Kim S-W, Knight JA, Kosma V-M, Kristensen VN, Krüger U, Kwong A, Lambrechts D, Le Marchand L, Lee E, Lee MH, Lee JW, Neng Lee C, Lejbkowicz F, Li J, Lilyquist J, Lindblom A, Lissowska J, Lo W-Y, Loibl S, Long J, Lophatananon A, Lubinski J, Luccarini C, Lux MP, Ma ESK, MacInnis RJ, Maishman T, Makalic E, Malone KE, Kostovska IM, Mannermaa A, Manoukian S, Manson JE, Margolin S, Mariapun S, Martinez ME, Matsuo K, Mavroudis D, McKay J, McLean C, Meijers-Heijboer H, Meindl A, Menéndez P, Menon U, Meyer J, Miao H, Miller N, Taib NAM, Muir K, Mulligan AM, Mulot C, Neuhausen SL, Nevanlinna H, Neven P, Nielsen SF, Noh D-Y, Nordestgaard BG, Norman A, Olopade OI, Olson JE, Olsson H, Olswold C, Orr N, Pankratz VS, Park SK, Park-Simon T-W, Lloyd R, Perez JIA, Peterlongo P, Peto J, Phillips K-A, Pinchev M, Plaseska-Karanfilska D, Prentice R, Presneau N, Prokofyeva D, Pugh E, Pylkäs K, Rack B, Radice P, Rahman N, Rennert G, Rennert HS, Rhenius V, Romero A, Romm J, Ruddy KJ, Rüdiger T, Rudolph A, Ruebner M, Rutgers EJT, Saloustros E, Sandler DP, Sangrajrang S, Sawyer EJ, Schmidt DF, Schmutzler RK, Schneeweiss A, Schoemaker MJ, Schumacher F, Schürmann P, Scott RJ, Scott C, Seal S, Seynaeve C, Shah M, Sharma P, Shen C-Y, Sheng G, Sherman ME, Shrubsole MJ, Shu X-O, Smeets A, Sohn C, Southey MC, Spinelli JJ, Stegmaier C, Stewart-Brown S, Stone J, Stram DO, Surowy H, Swerdlow A, Tamimi R, Taylor JA, Tengström M, Teo SH, Beth Terry M, Tessier DC, Thanasitthichai S, Thöne K, Tollenaar RAEM, Tomlinson I, Tong L, Torres D, Truong T, Tseng C-C, Tsugane S, Ulmer H-U, Ursin G, Untch M, Vachon C, van Asperen CJ, Van Den Berg D, van den Ouweland AMW, van der Kolk L, van der Luijt RB, Vincent D, Vollenweider J, Waisfisz Q, Wang-Gohrke S, Weinberg CR, Wendt C, Whittemore AS, Wildiers H, Willett W, Winqvist R, Wolk A, Wu AH, Xia L, Yamaji T, Yang XR, Har Yip C, Yoo K-Y, Yu J-C, Zheng W, Zheng Y, Zhu B, Ziogas A, Ziv E, ABCTB Investigators. ConFab/AOCS Investigators. Lakhani SR, Antoniou AC, Droit A, Andrulis IL, Amos CI, Couch FJ, Pharoah PDP, Chang-Claude J, Hall P, Hunter DJ, Milne RL, García-Closas M, Schmidt MK, Chanock SJ, Dunning AM, Edwards SL, Bader GD, Chenevix-Trench G, Simard J, Kraft P, Easton DF. Association analysis identifies 65 new breast cancer risk loci. Nature. 2017;551:92–94. doi: 10.1038/nature24284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mokry LE, Ross S, Timpson NJ, Sawcer S, Davey Smith G, Richards JB. Obesity and Multiple Sclerosis: A Mendelian Randomization Study. PLOS Medicine. 2016;13:e1002053. doi: 10.1371/journal.pmed.1002053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morris JA, Kemp JP, Youlten SE, Laurent L, Logan JG, Chai RC, Vulpescu NA, Forgetta V, Kleinman A, Mohanty ST, Sergio CM, Quinn J, Nguyen-Yamamoto L, Luco A-L, Vijay J, Simon M-M, Pramatarova A, Medina-Gomez C, Trajanoska K, Ghirardello EJ, Butterfield NC, Curry KF, Leitch VD, Sparkes PC, Adoum A-T, Mannan NS, Komla-Ebri DSK, Pollard AS, Dewhurst HF, Hassall TAD, Beltejar M-JG, 23andMe Research Team. Adams DJ, Vaillancourt SM, Kaptoge S, Baldock P, Cooper C, Reeve J, Ntzani EE, Evangelou E, Ohlsson C, Karasik D, Rivadeneira F, Kiel DP, Tobias JH, Gregson CL, Harvey NC, Grundberg E, Goltzman D, Adams DJ, Lelliott CJ, Hinds DA, Ackert-Bicknell CL, Hsu Y-H, Maurano MT, Croucher PI, Williams GR, Bassett JHD, Evans DM, Richards JB. An atlas of genetic influences on osteoporosis in humans and mice. Nature Genetics. 2019;51:258–266. doi: 10.1038/s41588-018-0302-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nalls MA, Blauwendraat C, Vallerga CL, Heilbron K, Bandres-Ciga S, Chang D, Tan M, Kia DA, Noyce AJ, Xue A, Bras J, Young E, von Coelln R, Simón-Sánchez J, Schulte C, Sharma M, Krohn L, Pihlstrøm L, Siitonen A, Iwaki H, Leonard H, Faghri F, Gibbs JR, Hernandez DG, Scholz SW, Botia JA, Martinez M, Corvol J-C, Lesage S, Jankovic J, Shulman LM, Sutherland M, Tienari P, Majamaa K, Toft M, Andreassen OA, Bangale T, Brice A, Yang J, Gan-Or Z, Gasser T, Heutink P, Shulman JM, Wood NW, Hinds DA, Hardy JA, Morris HR, Gratten J, Visscher PM, Graham RR, Singleton AB, 23andMe Research Team Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. The Lancet. Neurology. 2019;18:1091–1102. doi: 10.1016/S1474-4422(19)30320-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nead KT, Sharp SJ, Thompson DJ, Painter JN, Savage DB, Semple RK, Barker A, Australian National Endometrial Cancer Study Group (ANECS) Perry JRB, Attia J, Dunning AM, Easton DF, Holliday E, Lotta LA, O’Mara T, McEvoy M, Pharoah PDP, Scott RJ, Spurdle AB, Langenberg C, Wareham NJ, Scott RA. Evidence of a Causal Association Between Insulinemia and Endometrial Cancer: A Mendelian Randomization Analysis. Journal of the National Cancer Institute. 2015;107:djv178. doi: 10.1093/jnci/djv178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikpay M, Goel A, Won H-H, Hall LM, Willenborg C, Kanoni S, Saleheen D, Kyriakou T, Nelson CP, Hopewell JC, Webb TR, Zeng L, Dehghan A, Alver M, Armasu SM, Auro K, Bjonnes A, Chasman DI, Chen S, Ford I, Franceschini N, Gieger C, Grace C, Gustafsson S, Huang J, Hwang S-J, Kim YK, Kleber ME, Lau KW, Lu X, Lu Y, Lyytikäinen L-P, Mihailov E, Morrison AC, Pervjakova N, Qu L, Rose LM, Salfati E, Saxena R, Scholz M, Smith AV, Tikkanen E, Uitterlinden A, Yang X, Zhang W, Zhao W, de Andrade M, de Vries PS, van Zuydam NR, Anand SS, Bertram L, Beutner F, Dedoussis G, Frossard P, Gauguier D, Goodall AH, Gottesman O, Haber M, Han B-G, Huang J, Jalilzadeh S, Kessler T, König IR, Lannfelt L, Lieb W, Lind L, Lindgren CM, Lokki M-L, Magnusson PK, Mallick NH, Mehra N, Meitinger T, Memon F-U-R, Morris AP, Nieminen MS, Pedersen NL, Peters A, Rallidis LS, Rasheed A, Samuel M, Shah SH, Sinisalo J, Stirrups KE, Trompet S, Wang L, Zaman KS, Ardissino D, Boerwinkle E, Borecki IB, Bottinger EP, Buring JE, Chambers JC, Collins R, Cupples LA, Danesh J, Demuth I, Elosua R, Epstein SE, Esko T, Feitosa MF, Franco OH, Franzosi MG, Granger CB, Gu D, Gudnason V, Hall AS, Hamsten A, Harris TB, Hazen SL, Hengstenberg C, Hofman A, Ingelsson E, Iribarren C, Jukema JW, Karhunen PJ, Kim B-J, Kooner JS, Kullo IJ, Lehtimäki T, Loos RJF, Melander O, Metspalu A, März W, Palmer CN, Perola M, Quertermous T, Rader DJ, Ridker PM, Ripatti S, Roberts R, Salomaa V, Sanghera DK, Schwartz SM, Seedorf U, Stewart AF, Stott DJ, Thiery J, Zalloua PA, O’Donnell CJ, Reilly MP, Assimes TL, Thompson JR, Erdmann J, Clarke R, Watkins H, Kathiresan S, McPherson R, Deloukas P, Schunkert H, Samani NJ, Farrall M. A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nature Genetics. 2015;47:1121–1130. doi: 10.1038/ng.3396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nordestgaard LT, Tybjærg-Hansen A, Nordestgaard BG, Frikke-Schmidt R. Body Mass Index and Risk of Alzheimer’s Disease: A Mendelian Randomization Study of 399,536 Individuals. The Journal of Clinical Endocrinology and Metabolism. 2017;102:2310–2320. doi: 10.1210/jc.2017-00195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noyce AJ, Kia DA, Hemani G, Nicolas A, Price TR, De Pablo-Fernandez E, Haycock PC, Lewis PA, Foltynie T, Davey Smith G, International Parkinson Disease Genomics Consortium. Schrag A, Lees AJ, Hardy J, Singleton A, Nalls MA, Pearce N, Lawlor DA, Wood NW. Estimating the causal influence of body mass index on risk of Parkinson disease: A Mendelian randomisation study. PLOS Medicine. 2017;14:e1002314. doi: 10.1371/journal.pmed.1002314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okada Y, Wu D, Trynka G, Raj T, Terao C, Ikari K, Kochi Y, Ohmura K, Suzuki A, Yoshida S, Graham RR, Manoharan A, Ortmann W, Bhangale T, Denny JC, Carroll RJ, Eyler AE, Greenberg JD, Kremer JM, Pappas DA, Jiang L, Yin J, Ye L, Su D-F, Yang J, Xie G, Keystone E, Westra H-J, Esko T, Metspalu A, Zhou X, Gupta N, Mirel D, Stahl EA, Diogo D, Cui J, Liao K, Guo MH, Myouzen K, Kawaguchi T, Coenen MJH, van Riel PLCM, van de Laar MAFJ, Guchelaar H-J, Huizinga TWJ, Dieudé P, Mariette X, Bridges SL, Zhernakova A, Toes REM, Tak PP, Miceli-Richard C, Bang S-Y, Lee H-S, Martin J, Gonzalez-Gay MA, Rodriguez-Rodriguez L, Rantapää-Dahlqvist S, Arlestig L, Choi HK, Kamatani Y, Galan P, Lathrop M, RACI consortium. GARNET consortium. Eyre S, Bowes J, Barton A, de Vries N, Moreland LW, Criswell LA, Karlson EW, Taniguchi A, Yamada R, Kubo M, Liu JS, Bae S-C, Worthington J, Padyukov L, Klareskog L, Gregersen PK, Raychaudhuri S, Stranger BE, De Jager PL, Franke L, Visscher PM, Brown MA, Yamanaka H, Mimori T, Takahashi A, Xu H, Behrens TW, Siminovitch KA, Momohara S, Matsuda F, Yamamoto K, Plenge RM. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature. 2014;506:376–381. doi: 10.1038/nature12873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Mara TA, Glubb DM, Amant F, Annibali D, Ashton K, Attia J, Auer PL, Beckmann MW, Black A, Bolla MK, Brauch H, Brenner H, Brinton L, Buchanan DD, Burwinkel B, Chang-Claude J, Chanock SJ, Chen C, Chen MM, Cheng THT, Clarke CL, Clendenning M, Cook LS, Couch FJ, Cox A, Crous-Bous M, Czene K, Day F, Dennis J, Depreeuw J, Doherty JA, Dörk T, Dowdy SC, Dürst M, Ekici AB, Fasching PA, Fridley BL, Friedenreich CM, Fritschi L, Fung J, García-Closas M, Gaudet MM, Giles GG, Goode EL, Gorman M, Haiman CA, Hall P, Hankison SE, Healey CS, Hein A, Hillemanns P, Hodgson S, Hoivik EA, Holliday EG, Hopper JL, Hunter DJ, Jones A, Krakstad C, Kristensen VN, Lambrechts D, Marchand LL, Liang X, Lindblom A, Lissowska J, Long J, Lu L, Magliocco AM, Martin L, McEvoy M, Meindl A, Michailidou K, Milne RL, Mints M, Montgomery GW, Nassir R, Olsson H, Orlow I, Otton G, Palles C, Perry JRB, Peto J, Pooler L, Prescott J, Proietto T, Rebbeck TR, Risch HA, Rogers PAW, Rübner M, Runnebaum I, Sacerdote C, Sarto GE, Schumacher F, Scott RJ, Setiawan VW, Shah M, Sheng X, Shu X-O, Southey MC, Swerdlow AJ, Tham E, Trovik J, Turman C, Tyrer JP, Vachon C, VanDen Berg D, Vanderstichele A, Wang Z, Webb PM, Wentzensen N, Werner HMJ, Winham SJ, Wolk A, Xia L, Xiang Y-B, Yang HP, Yu H, Zheng W, Pharoah PDP, Dunning AM, Kraft P, De Vivo I, Tomlinson I, Easton DF, Spurdle AB, Thompson DJ. Identification of nine new susceptibility loci for endometrial cancer. Nature Communications. 2018;9:3166. doi: 10.1038/s41467-018-05427-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Painter JN, O’Mara TA, Marquart L, Webb PM, Attia J, Medland SE, Cheng T, Dennis J, Holliday EG, McEvoy M, Scott RJ, Ahmed S, Healey CS, Shah M, Gorman M, Martin L, Hodgson SV, Beckmann MW, Ekici AB, Fasching PA, Hein A, Rübner M, Czene K, Darabi H, Hall P, Li J, Dörk T, Dürst M, Hillemanns P, Runnebaum IB, Amant F, Annibali D, Depreeuw J, Lambrechts D, Neven P, Cunningham JM, Dowdy SC, Goode EL, Fridley BL, Winham SJ, Njølstad TS, Salvesen HB, Trovik J, Werner HMJ, Ashton KA, Otton G, Proietto A, Mints M, Tham E, Bolla MK, Michailidou K, Wang Q, Tyrer JP, Hopper JL, Peto J, Swerdlow AJ, Burwinkel B, Brenner H, Meindl A, Brauch H, Lindblom A, Chang-Claude J, Couch FJ, Giles GG, Kristensen VN, Cox A, Pharoah PDP, Tomlinson I, Dunning AM, Easton DF, Thompson DJ, Spurdle AB, AOCS Group. for RENDOCAS. National Study of Endometrial Cancer Genetics Group (NSECG) Australian National Endometrial Cancer Study Group (ANECS) Genetic Risk Score Mendelian Randomization Shows that Obesity Measured as Body Mass Index, but not Waist:Hip Ratio, Is Causal for Endometrial Cancer. Cancer Epidemiology, Biomarkers & Prevention. 2016;25:1503–1510. doi: 10.1158/1055-9965.EPI-16-0147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phelan CM, Kuchenbaecker KB, Tyrer JP, Kar SP, Lawrenson K, Winham SJ, Dennis J, Pirie A, Riggan MJ, Chornokur G, Earp MA, Lyra PC, Lee JM, Coetzee S, Beesley J, McGuffog L, Soucy P, Dicks E, Lee A, Barrowdale D, Lecarpentier J, Leslie G, Aalfs CM, Aben KKH, Adams M, Adlard J, Andrulis IL, Anton-Culver H, Antonenkova N, AOCS study group. Aravantinos G, Arnold N, Arun BK, Arver B, Azzollini J, Balmaña J, Banerjee SN, Barjhoux L, Barkardottir RB, Bean Y, Beckmann MW, Beeghly-Fadiel A, Benitez J, Bermisheva M, Bernardini MQ, Birrer MJ, Bjorge L, Black A, Blankstein K, Blok MJ, Bodelon C, Bogdanova N, Bojesen A, Bonanni B, Borg Å, Bradbury AR, Brenton JD, Brewer C, Brinton L, Broberg P, Brooks-Wilson A, Bruinsma F, Brunet J, Buecher B, Butzow R, Buys SS, Caldes T, Caligo MA, Campbell I, Cannioto R, Carney ME, Cescon T, Chan SB, Chang-Claude J, Chanock S, Chen XQ, Chiew Y-E, Chiquette J, Chung WK, Claes KBM, Conner T, Cook LS, Cook J, Cramer DW, Cunningham JM, D’Aloisio AA, Daly MB, Damiola F, Damirovna SD, Dansonka-Mieszkowska A, Dao F, Davidson R, DeFazio A, Delnatte C, Doheny KF, Diez O, Ding YC, Doherty JA, Domchek SM, Dorfling CM, Dörk T, Dossus L, Duran M, Dürst M, Dworniczak B, Eccles D, Edwards T, Eeles R, Eilber U, Ejlertsen B, Ekici AB, Ellis S, Elvira M, EMBRACE Study. Eng KH, Engel C, Evans DG, Fasching PA, Ferguson S, Ferrer SF, Flanagan JM, Fogarty ZC, Fortner RT, Fostira F, Foulkes WD, Fountzilas G, Fridley BL, Friebel TM, Friedman E, Frost D, Ganz PA, Garber J, García MJ, Garcia-Barberan V, Gehrig A, GEMO Study Collaborators. Gentry-Maharaj A, Gerdes A-M, Giles GG, Glasspool R, Glendon G, Godwin AK, Goldgar DE, Goranova T, Gore M, Greene MH, Gronwald J, Gruber S, Hahnen E, Haiman CA, Håkansson N, Hamann U, Hansen TVO, Harrington PA, Harris HR, Hauke J, HEBON Study. Hein A, Henderson A, Hildebrandt MAT, Hillemanns P, Hodgson S, Høgdall CK, Høgdall E, Hogervorst FBL, Holland H, Hooning MJ, Hosking K, Huang R-Y, Hulick PJ, Hung J, Hunter DJ, Huntsman DG, Huzarski T, Imyanitov EN, Isaacs C, Iversen ES, Izatt L, Izquierdo A, Jakubowska A, James P, Janavicius R, Jernetz M, Jensen A, Jensen UB, John EM, Johnatty S, Jones ME, Kannisto P, Karlan BY, Karnezis A, Kast K, KConFab Investigators. Kennedy CJ, Khusnutdinova E, Kiemeney LA, Kiiski JI, Kim S-W, Kjaer SK, Köbel M, Kopperud RK, Kruse TA, Kupryjanczyk J, Kwong A, Laitman Y, Lambrechts D, Larrañaga N, Larson MC, Lazaro C, Le ND, Le Marchand L, Lee JW, Lele SB, Leminen A, Leroux D, Lester J, Lesueur F, Levine DA, Liang D, Liebrich C, Lilyquist J, Lipworth L, Lissowska J, Lu KH, Lubinński J, Luccarini C, Lundvall L, Mai PL, Mendoza-Fandiño G, Manoukian S, Massuger LFAG, May T, Mazoyer S, McAlpine JN, McGuire V, McLaughlin JR, McNeish I, Meijers-Heijboer H, Meindl A, Menon U, Mensenkamp AR, Merritt MA, Milne RL, Mitchell G, Modugno F, Moes-Sosnowska J, Moffitt M, Montagna M, Moysich KB, Mulligan AM, Musinsky J, Nathanson KL, Nedergaard L, Ness RB, Neuhausen SL, Nevanlinna H, Niederacher D, Nussbaum RL, Odunsi K, Olah E, Olopade OI, Olsson H, Olswold C, O’Malley DM, Ong K-R, Onland-Moret NC, OPAL study group. Orr N, Orsulic S, Osorio A, Palli D, Papi L, Park-Simon T-W, Paul J, Pearce CL, Pedersen IS, Peeters PHM, Peissel B, Peixoto A, Pejovic T, Pelttari LM, Permuth JB, Peterlongo P, Pezzani L, Pfeiler G, Phillips K-A, Piedmonte M, Pike MC, Piskorz AM, Poblete SR, Pocza T, Poole EM, Poppe B, Porteous ME, Prieur F, Prokofyeva D, Pugh E, Pujana MA, Pujol P, Radice P, Rantala J, Rappaport-Fuerhauser C, Rennert G, Rhiem K, Rice P, Richardson A, Robson M, Rodriguez GC, Rodríguez-Antona C, Romm J, Rookus MA, Rossing MA, Rothstein JH, Rudolph A, Runnebaum IB, Salvesen HB, Sandler DP, Schoemaker MJ, Senter L, Setiawan VW, Severi G, Sharma P, Shelford T, Siddiqui N, Side LE, Sieh W, Singer CF, Sobol H, Song H, Southey MC, Spurdle AB, Stadler Z, Steinemann D, Stoppa-Lyonnet D, Sucheston-Campbell LE, Sukiennicki G, Sutphen R, Sutter C, Swerdlow AJ, Szabo CI, Szafron L, Tan YY, Taylor JA, Tea M-K, Teixeira MR, Teo S-H, Terry KL, Thompson PJ, Thomsen LCV, Thull DL, Tihomirova L, Tinker AV, Tischkowitz M, Tognazzo S, Toland AE, Tone A, Trabert B, Travis RC, Trichopoulou A, Tung N, Tworoger SS, van Altena AM, Van Den Berg D, van der Hout AH, van der Luijt RB, Van Heetvelde M, Van Nieuwenhuysen E, van Rensburg EJ, Vanderstichele A, Varon-Mateeva R, Vega A, Edwards DV, Vergote I, Vierkant RA, Vijai J, Vratimos A, Walker L, Walsh C, Wand D, Wang-Gohrke S, Wappenschmidt B, Webb PM, Weinberg CR, Weitzel JN, Wentzensen N, Whittemore AS, Wijnen JT, Wilkens LR, Wolk A, Woo M, Wu X, Wu AH, Yang H, Yannoukakos D, Ziogas A, Zorn KK, Narod SA, Easton DF, Amos CI, Schildkraut JM, Ramus SJ, Ottini L, Goodman MT, Park SK, Kelemen LE, Risch HA, Thomassen M, Offit K, Simard J, Schmutzler RK, Hazelett D, Monteiro AN, Couch FJ, Berchuck A, Chenevix-Trench G, Goode EL, Sellers TA, Gayther SA, Antoniou AC, Pharoah PDP. Identification of 12 new susceptibility loci for different histotypes of epithelial ovarian cancer. Nature Genetics. 2017;49:680–691. doi: 10.1038/ng.3826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pierce BL, Burgess S. Efficient design for Mendelian randomization studies: subsample and 2-sample instrumental variable estimators. American Journal of Epidemiology. 2013;178:1177–1184. doi: 10.1093/aje/kwt084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- R Development Core Team . Vienna, Austria: R Foundation for Statistical Computing; 2020. http://www.r-project.org [Google Scholar]
- Reyes C, Leyland KM, Peat G, Cooper C, Arden NK, Prieto-Alhambra D. Association Between Overweight and Obesity and Risk of Clinically Diagnosed Knee, Hip, and Hand Osteoarthritis: A Population-Based Cohort Study. Arthritis & Rheumatology. 2016;68:1869–1875. doi: 10.1002/art.39707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riaz H, Khan MS, Siddiqi TJ, Usman MS, Shah N, Goyal A, Khan SS, Mookadam F, Krasuski RA, Ahmed H. Association Between Obesity and Cardiovascular Outcomes: A Systematic Review and Meta-analysis of Mendelian Randomization Studies. JAMA Network Open. 2018;1:e183788. doi: 10.1001/jamanetworkopen.2018.3788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richardson TG, Sanderson E, Elsworth B, Tilling K, Davey Smith G. Use of genetic variation to separate the effects of early and later life adiposity on disease risk: mendelian randomisation study. BMJ. 2020;369:m1203. doi: 10.1136/bmj.m1203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roselli C, Chaffin MD, Weng L-C, Aeschbacher S, Ahlberg G, Albert CM, Almgren P, Alonso A, Anderson CD, Aragam KG, Arking DE, Barnard J, Bartz TM, Benjamin EJ, Bihlmeyer NA, Bis JC, Bloom HL, Boerwinkle E, Bottinger EB, Brody JA, Calkins H, Campbell A, Cappola TP, Carlquist J, Chasman DI, Chen LY, Chen Y-DI, Choi E-K, Choi SH, Christophersen IE, Chung MK, Cole JW, Conen D, Cook J, Crijns HJ, Cutler MJ, Damrauer SM, Daniels BR, Darbar D, Delgado G, Denny JC, Dichgans M, Dörr M, Dudink EA, Dudley SC, Esa N, Esko T, Eskola M, Fatkin D, Felix SB, Ford I, Franco OH, Geelhoed B, Grewal RP, Gudnason V, Guo X, Gupta N, Gustafsson S, Gutmann R, Hamsten A, Harris TB, Hayward C, Heckbert SR, Hernesniemi J, Hocking LJ, Hofman A, Horimoto ARVR, Huang J, Huang PL, Huffman J, Ingelsson E, Ipek EG, Ito K, Jimenez-Conde J, Johnson R, Jukema JW, Kääb S, Kähönen M, Kamatani Y, Kane JP, Kastrati A, Kathiresan S, Katschnig-Winter P, Kavousi M, Kessler T, Kietselaer BL, Kirchhof P, Kleber ME, Knight S, Krieger JE, Kubo M, Launer LJ, Laurikka J, Lehtimäki T, Leineweber K, Lemaitre RN, Li M, Lim HE, Lin HJ, Lin H, Lind L, Lindgren CM, Lokki M-L, London B, Loos RJF, Low S-K, Lu Y, Lyytikäinen L-P, Macfarlane PW, Magnusson PK, Mahajan A, Malik R, Mansur AJ, Marcus GM, Margolin L, Margulies KB, März W, McManus DD, Melander O, Mohanty S, Montgomery JA, Morley MP, Morris AP, Müller-Nurasyid M, Natale A, Nazarian S, Neumann B, Newton-Cheh C, Niemeijer MN, Nikus K, Nilsson P, Noordam R, Oellers H, Olesen MS, Orho-Melander M, Padmanabhan S, Pak H-N, Paré G, Pedersen NL, Pera J, Pereira A, Porteous D, Psaty BM, Pulit SL, Pullinger CR, Rader DJ, Refsgaard L, Ribasés M, Ridker PM, Rienstra M, Risch L, Roden DM, Rosand J, Rosenberg MA, Rost N, Rotter JI, Saba S, Sandhu RK, Schnabel RB, Schramm K, Schunkert H, Schurman C, Scott SA, Seppälä I, Shaffer C, Shah S, Shalaby AA, Shim J, Shoemaker MB, Siland JE, Sinisalo J, Sinner MF, Slowik A, Smith AV, Smith BH, Smith JG, Smith JD, Smith NL, Soliman EZ, Sotoodehnia N, Stricker BH, Sun A, Sun H, Svendsen JH, Tanaka T, Tanriverdi K, Taylor KD, Teder-Laving M, Teumer A, Thériault S, Trompet S, Tucker NR, Tveit A, Uitterlinden AG, Van Der Harst P, Van Gelder IC, Van Wagoner DR, Verweij N, Vlachopoulou E, Völker U, Wang B, Weeke PE, Weijs B, Weiss R, Weiss S, Wells QS, Wiggins KL, Wong JA, Woo D, Worrall BB, Yang P-S, Yao J, Yoneda ZT, Zeller T, Zeng L, Lubitz SA, Lunetta KL, Ellinor PT. Multi-ethnic genome-wide association study for atrial fibrillation. Nature Genetics. 2018;50:1225–1233. doi: 10.1038/s41588-018-0133-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sattar N, McGuire DK. Pathways to Cardiorenal Complications in Type 2 Diabetes Mellitus: A Need to Rethink. Circulation. 2018;138:7–9. doi: 10.1161/CIRCULATIONAHA.118.035083. [DOI] [PubMed] [Google Scholar]
- Sbidian E, Chaimani A, Hua C, Mazaud C, Droitcourt C, Hughes C, Ingram JR, Naldi L, Chosidow O, Le Cleach L. Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis. The Cochrane Database of Systematic Reviews. 2017;12:CD011535. doi: 10.1002/14651858.CD011535.pub2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scelo G, Purdue MP, Brown KM, Johansson M, Wang Z, Eckel-Passow JE, Ye Y, Hofmann JN, Choi J, Foll M, Gaborieau V, Machiela MJ, Colli LM, Li P, Sampson JN, Abedi-Ardekani B, Besse C, Blanche H, Boland A, Burdette L, Chabrier A, Durand G, Le Calvez-Kelm F, Prokhortchouk E, Robinot N, Skryabin KG, Wozniak MB, Yeager M, Basta-Jovanovic G, Dzamic Z, Foretova L, Holcatova I, Janout V, Mates D, Mukeriya A, Rascu S, Zaridze D, Bencko V, Cybulski C, Fabianova E, Jinga V, Lissowska J, Lubinski J, Navratilova M, Rudnai P, Szeszenia-Dabrowska N, Benhamou S, Cancel-Tassin G, Cussenot O, Baglietto L, Boeing H, Khaw K-T, Weiderpass E, Ljungberg B, Sitaram RT, Bruinsma F, Jordan SJ, Severi G, Winship I, Hveem K, Vatten LJ, Fletcher T, Koppova K, Larsson SC, Wolk A, Banks RE, Selby PJ, Easton DF, Pharoah P, Andreotti G, Freeman LEB, Koutros S, Albanes D, Männistö S, Weinstein S, Clark PE, Edwards TL, Lipworth L, Gapstur SM, Stevens VL, Carol H, Freedman ML, Pomerantz MM, Cho E, Kraft P, Preston MA, Wilson KM, Michael Gaziano J, Sesso HD, Black A, Freedman ND, Huang W-Y, Anema JG, Kahnoski RJ, Lane BR, Noyes SL, Petillo D, Teh BT, Peters U, White E, Anderson GL, Johnson L, Luo J, Buring J, Lee I-M, Chow W-H, Moore LE, Wood C, Eisen T, Henrion M, Larkin J, Barman P, Leibovich BC, Choueiri TK, Mark Lathrop G, Rothman N, Deleuze J-F, McKay JD, Parker AS, Wu X, Houlston RS, Brennan P, Chanock SJ. Genome-wide association study identifies multiple risk loci for renal cell carcinoma. Nature Communications. 2017;8:15724. doi: 10.1038/ncomms15724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schumacher FR, Al Olama AA, Berndt SI, Benlloch S, Ahmed M, Saunders EJ, Dadaev T, Leongamornlert D, Anokian E, Cieza-Borrella C, Goh C, Brook MN, Sheng X, Fachal L, Dennis J, Tyrer J, Muir K, Lophatananon A, Stevens VL, Gapstur SM, Carter BD, Tangen CM, Goodman PJ, Thompson IM, Batra J, Chambers S, Moya L, Clements J, Horvath L, Tilley W, Risbridger GP, Gronberg H, Aly M, Nordström T, Pharoah P, Pashayan N, Schleutker J, Tammela TLJ, Sipeky C, Auvinen A, Albanes D, Weinstein S, Wolk A, Håkansson N, West CML, Dunning AM, Burnet N, Mucci LA, Giovannucci E, Andriole GL, Cussenot O, Cancel-Tassin G, Koutros S, Beane Freeman LE, Sorensen KD, Orntoft TF, Borre M, Maehle L, Grindedal EM, Neal DE, Donovan JL, Hamdy FC, Martin RM, Travis RC, Key TJ, Hamilton RJ, Fleshner NE, Finelli A, Ingles SA, Stern MC, Rosenstein BS, Kerns SL, Ostrer H, Lu Y-J, Zhang H-W, Feng N, Mao X, Guo X, Wang G, Sun Z, Giles GG, Southey MC, MacInnis RJ, FitzGerald LM, Kibel AS, Drake BF, Vega A, Gómez-Caamaño A, Szulkin R, Eklund M, Kogevinas M, Llorca J, Castaño-Vinyals G, Penney KL, Stampfer M, Park JY, Sellers TA, Lin H-Y, Stanford JL, Cybulski C, Wokolorczyk D, Lubinski J, Ostrander EA, Geybels MS, Nordestgaard BG, Nielsen SF, Weischer M, Bisbjerg R, Røder MA, Iversen P, Brenner H, Cuk K, Holleczek B, Maier C, Luedeke M, Schnoeller T, Kim J, Logothetis CJ, John EM, Teixeira MR, Paulo P, Cardoso M, Neuhausen SL, Steele L, Ding YC, De Ruyck K, De Meerleer G, Ost P, Razack A, Lim J, Teo S-H, Lin DW, Newcomb LF, Lessel D, Gamulin M, Kulis T, Kaneva R, Usmani N, Singhal S, Slavov C, Mitev V, Parliament M, Claessens F, Joniau S, Van den Broeck T, Larkin S, Townsend PA, Aukim-Hastie C, Gago-Dominguez M, Castelao JE, Martinez ME, Roobol MJ, Jenster G, van Schaik RHN, Menegaux F, Truong T, Koudou YA, Xu J, Khaw K-T, Cannon-Albright L, Pandha H, Michael A, Thibodeau SN, McDonnell SK, Schaid DJ, Lindstrom S, Turman C, Ma J, Hunter DJ, Riboli E, Siddiq A, Canzian F, Kolonel LN, Le Marchand L, Hoover RN, Machiela MJ, Cui Z, Kraft P, Amos CI, Conti DV, Easton DF, Wiklund F, Chanock SJ, Henderson BE, Kote-Jarai Z, Haiman CA, Eeles RA, Profile Study. Australian Prostate Cancer BioResource (APCB) IMPACT Study. Canary PASS Investigators. Breast and Prostate Cancer Cohort Consortium (BPC3) PRACTICAL (Prostate Cancer Association Group to Investigate Cancer-Associated Alterations in the Genome) Consortium. Cancer of the Prostate in Sweden (CAPS) Prostate Cancer Genome-wide Association Study of Uncommon Susceptibility Loci (PEGASUS) Genetic Associations and Mechanisms in Oncology (GAME-ON)/Elucidating Loci Involved in Prostate Cancer Susceptibility (ELLIPSE) Consortium Association analyses of more than 140,000 men identify 63 new prostate cancer susceptibility loci. Nature Genetics. 2018;50:928–936. doi: 10.1038/s41588-018-0142-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Semple RK, Savage DB, Cochran EK, Gorden P, O’Rahilly S. Genetic syndromes of severe insulin resistance. Endocrine Reviews. 2011;32:498–514. doi: 10.1210/er.2010-0020. [DOI] [PubMed] [Google Scholar]
- Shah S, Henry A, Roselli C, Lin H, Sveinbjörnsson G, Fatemifar G, Hedman AK, Wilk JB, Morley MP, Chaffin MD, Helgadottir A, Verwei N, Dehghan A, Almgren P, Andersson C, Aragam KG, Ärnlöv J, Backman JD, Biggs ML, Bloom HL, Brandimarto J, Brown MR, Leonard B, Buckbinder L, Carey DJ, Chasman DI, Chen X, Chen X, Chung J, Chutkow W, Cook JP, Delgado GE, Denaxas S, Doney AS, Dörr M, Dudley SC, Dunn ME, Engström G, Esko T, Felix SB, Finan S, Ford I, Ghanbari M, Ghasemi S, Giedraitis V, Giulianini F, Gottdiener JS, Gross S, Guðbjartsson DF, Gutmann R, Haggerty CM, Harst P, Hyde CL, Ingelsson E, Wouter Jukema J, Kavousi M, Khaw KT, Kleber ME, Køber L, Koekemoer A, Langenberg C, Lind L, Lindgren CM, London B, Lotta LA, Lovering RA, Luan J, Magnusson P, Mahajan A, Margulies KB, März W, Melander O, Mordi IR, Morgan T, Morris AD, Morris AP, Morrison AC, Nagle MW, Nelson CP, Niessner A, Niiranen T, O’Donoghue ML, Owens AT, Palmer CNA, Parry HM, Perola M, Portilla-Fernandez E, Psaty BM, Rice KM, Ridker PM, Romaine SPR, Rotter JI, Salo P, Salomaa V, Setten J, Shalaby AA, Smelser DT, Smith NL, Stender S, Stott DJ, Svensson P, Tammesoo ML, Taylor KD, Teder-Laving M, Teumer A, Thorgeirsson G, Thorsteinsdottir U, Torp-Pedersen C, Trompet S, Tyl B, Uitterlinden AG, Veluchamy A, Völker U, Voors AA, Wang X, Wareham NJ, Waterworth D, Weeke PE, Weiss R, Wiggins KL, Xing H, Yerges-Armstrong LM, Yu B, Zannad F, Hua Zhao J, Hemingway H, Samani NJ, McMurray JJV, Yang J, Visscher PM, Newton-Cheh C, Malarstig A, Holm H, Lubitz SA, Sattar N, Holmes MV, Cappola TP, Asselbergs F, Hingorani AD, Kuchenbaecker K, Ellinor PT, Lang CC, Stefansson K, Gustav Smith J, Vasan RS, Swerdlow DI, Thomas Lumbers R, EchoGen Consortium. Broad AF Investigators. Regeneron Genetics Center Genome-Wide Association Study Provides New Insights into the Genetic Architecture and Pathogenesis of Heart Failure. bioRxiv. 2019 doi: 10.1101/682013. [DOI]
- Shu X, Wu L, Khankari NK, Shu X-O, Wang TJ, Michailidou K, Bolla MK, Wang Q, Dennis J, Milne RL, Schmidt MK, Pharoah PDP, Andrulis IL, Hunter DJ, Simard J, Easton DF, Zheng W, Breast Cancer Association Consortium Associations of obesity and circulating insulin and glucose with breast cancer risk: a Mendelian randomization analysis. International Journal of Epidemiology. 2019;48:795–806. doi: 10.1093/ije/dyy201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith GD, Ebrahim S. Mendelian randomization: prospects, potentials, and limitations. International Journal of Epidemiology. 2004;33:30–42. doi: 10.1093/ije/dyh132. [DOI] [PubMed] [Google Scholar]
- Song J, Zhang R, Lv L, Liang J, Wang W, Liu R, Dang X. The Relationship Between Body Mass Index and Bone Mineral Density: A Mendelian Randomization Study. Calcified Tissue International. 2020;107:440–445. doi: 10.1007/s00223-020-00736-w. [DOI] [PubMed] [Google Scholar]
- Ståhlberg D, Rudling M, Angelin B, Björkhem I, Forsell P, Nilsell K, Einarsson K. Hepatic cholesterol metabolism in human obesity. Hepatology. 1997;25:1447–1450. doi: 10.1002/hep.510250623. [DOI] [PubMed] [Google Scholar]
- Suzuki S, Goto A, Nakatochi M, Narita A, Yamaji T, Sawada N, Katagiri R, Iwagami M, Hanyuda A, Hachiya T, Sutoh Y, Oze I, Koyanagi YN, Kasugai Y, Taniyama Y, Ito H, Ikezaki H, Nishida Y, Tamura T, Mikami H, Takezaki T, Suzuki S, Ozaki E, Kuriki K, Takashima N, Arisawa K, Takeuchi K, Tanno K, Shimizu A, Tamiya G, Hozawa A, Kinoshita K, Wakai K, Sasaki M, Yamamoto M, Matsuo K, Tsugane S, Iwasaki M. Body mass index and colorectal cancer risk: A Mendelian randomization study. Cancer Science. 2021;112:1579–1588. doi: 10.1111/cas.14824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tachmazidou I, Hatzikotoulas K, Southam L, Esparza-Gordillo J, Haberland V, Zheng J, Johnson T, Koprulu M, Zengini E, Steinberg J, Wilkinson JM, Bhatnagar S, Hoffman JD, Buchan N, Süveges D, arcOGEN Consortium. Yerges-Armstrong L, Smith GD, Gaunt TR, Scott RA, McCarthy LC, Zeggini E. Identification of new therapeutic targets for osteoarthritis through genome-wide analyses of UK Biobank data. Nature Genetics. 2019;51:230–236. doi: 10.1038/s41588-018-0327-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thrift AP, Gong J, Peters U, Chang-Claude J, Rudolph A, Slattery ML, Chan AT, Locke AE, Kahali B, Justice AE, Pers TH, Gallinger S, Hayes RB, Baron JA, Caan BJ, Ogino S, Berndt SI, Chanock SJ, Casey G, Haile RW, Du M, Harrison TA, Thornquist M, Duggan DJ, Le Marchand L, Lindor NM, Seminara D, Song M, Wu K, Thibodeau SN, Cotterchio M, Win AK, Jenkins MA, Hopper JL, Ulrich CM, Potter JD, Newcomb PA, Hoffmeister M, Brenner H, White E, Hsu L, Campbell PT. Mendelian Randomization Study of Body Mass Index and Colorectal Cancer Risk. Cancer Epidemiology, Biomarkers & Prevention. 2015;24:1024–1031. doi: 10.1158/1055-9965.EPI-14-1309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tin A, Marten J, Halperin Kuhns VL, Li Y, Wuttke M, Kirsten H, Sieber KB, Qiu C, Gorski M, Yu Z, Giri A, Sveinbjornsson G, Li M, Chu AY, Hoppmann A, O’Connor LJ, Prins B, Nutile T, Noce D, Akiyama M, Cocca M, Ghasemi S, van der Most PJ, Horn K, Xu Y, Fuchsberger C, Sedaghat S, Afaq S, Amin N, Ärnlöv J, Bakker SJL, Bansal N, Baptista D, Bergmann S, Biggs ML, Biino G, Boerwinkle E, Bottinger EP, Boutin TS, Brumat M, Burkhardt R, Campana E, Campbell A, Campbell H, Carroll RJ, Catamo E, Chambers JC, Ciullo M, Concas MP, Coresh J, Corre T, Cusi D, Felicita SC, de Borst MH, De Grandi A, de Mutsert R, de Vries APJ, Delgado G, Demirkan A, Devuyst O, Dittrich K, Eckardt K-U, Ehret G, Endlich K, Evans MK, Gansevoort RT, Gasparini P, Giedraitis V, Gieger C, Girotto G, Gögele M, Gordon SD, Gudbjartsson DF, Gudnason V, German Chronic Kidney Disease Study. Haller T, Hamet P, Harris TB, Hayward C, Hicks AA, Hofer E, Holm H, Huang W, Hutri-Kähönen N, Hwang S-J, Ikram MA, Lewis RM, Ingelsson E, Jakobsdottir J, Jonsdottir I, Jonsson H, Joshi PK, Josyula NS, Jung B, Kähönen M, Kamatani Y, Kanai M, Kerr SM, Kiess W, Kleber ME, Koenig W, Kooner JS, Körner A, Kovacs P, Krämer BK, Kronenberg F, Kubo M, Kühnel B, La Bianca M, Lange LA, Lehne B, Lehtimäki T, Lifelines Cohort Study. Liu J, Loeffler M, Loos RJF, Lyytikäinen L-P, Magi R, Mahajan A, Martin NG, März W, Mascalzoni D, Matsuda K, Meisinger C, Meitinger T, Metspalu A, Milaneschi Y, V. A. Million Veteran Program. O’Donnell CJ, Wilson OD, Gaziano JM, Mishra PP, Mohlke KL, Mononen N, Montgomery GW, Mook-Kanamori DO, Müller-Nurasyid M, Nadkarni GN, Nalls MA, Nauck M, Nikus K, Ning B, Nolte IM, Noordam R, O’Connell JR, Olafsson I, Padmanabhan S, Penninx BWJH, Perls T, Peters A, Pirastu M, Pirastu N, Pistis G, Polasek O, Ponte B, Porteous DJ, Poulain T, Preuss MH, Rabelink TJ, Raffield LM, Raitakari OT, Rettig R, Rheinberger M, Rice KM, Rizzi F, Robino A, Rudan I, Krajcoviechova A, Cifkova R, Rueedi R, Ruggiero D, Ryan KA, Saba Y, Salvi E, Schmidt H, Schmidt R, Shaffer CM, Smith AV, Smith BH, Spracklen CN, Strauch K, Stumvoll M, Sulem P, Tajuddin SM, Teren A, Thiery J, Thio CHL, Thorsteinsdottir U, Toniolo D, Tönjes A, Tremblay J, Uitterlinden AG, Vaccargiu S, van der Harst P, van Duijn CM, Verweij N, Völker U, Vollenweider P, Waeber G, Waldenberger M, Whitfield JB, Wild SH, Wilson JF, Yang Q, Zhang W, Zonderman AB, Bochud M, Wilson JG, Pendergrass SA, Ho K, Parsa A, Pramstaller PP, Psaty BM, Böger CA, Snieder H, Butterworth AS, Okada Y, Edwards TL, Stefansson K, Susztak K, Scholz M, Heid IM, Hung AM, Teumer A, Pattaro C, Woodward OM, Vitart V, Köttgen A. Target genes, variants, tissues and transcriptional pathways influencing human serum urate levels. Nature Genetics. 2019;51:1459–1474. doi: 10.1038/s41588-019-0504-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsoi LC, Stuart PE, Tian C, Gudjonsson JE, Das S, Zawistowski M, Ellinghaus E, Barker JN, Chandran V, Dand N, Duffin KC, Enerbäck C, Esko T, Franke A, Gladman DD, Hoffmann P, Kingo K, Kõks S, Krueger GG, Lim HW, Metspalu A, Mrowietz U, Mucha S, Rahman P, Reis A, Tejasvi T, Trembath R, Voorhees JJ, Weidinger S, Weichenthal M, Wen X, Eriksson N, Kang HM, Hinds DA, Nair RP, Abecasis GR, Elder JT. Large scale meta-analysis characterizes genetic architecture for common psoriasis associated variants. Nature Communications. 2017;8:15382. doi: 10.1038/ncomms15382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Viechtbauer W. Conducting meta-analyses in R with the metafor package. Journal of Statistical Software. 2010;36:1–48. doi: 10.18637/jss.v036.i03. [DOI] [Google Scholar]
- Vincent EE, Yaghootkar H. Using genetics to decipher the link between type 2 diabetes and cancer: shared aetiology or downstream consequence? Diabetologia. 2020;63:1706–1717. doi: 10.1007/s00125-020-05228-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, Adams MJ, Agerbo E, Air TM, Andlauer TMF, Bacanu S-A, Bækvad-Hansen M, Beekman AFT, Bigdeli TB, Binder EB, Blackwood DRH, Bryois J, Buttenschøn HN, Bybjerg-Grauholm J, Cai N, Castelao E, Christensen JH, Clarke T-K, Coleman JIR, Colodro-Conde L, Couvy-Duchesne B, Craddock N, Crawford GE, Crowley CA, Dashti HS, Davies G, Deary IJ, Degenhardt F, Derks EM, Direk N, Dolan CV, Dunn EC, Eley TC, Eriksson N, Escott-Price V, Kiadeh FHF, Finucane HK, Forstner AJ, Frank J, Gaspar HA, Gill M, Giusti-Rodríguez P, Goes FS, Gordon SD, Grove J, Hall LS, Hannon E, Hansen CS, Hansen TF, Herms S, Hickie IB, Hoffmann P, Homuth G, Horn C, Hottenga J-J, Hougaard DM, Hu M, Hyde CL, Ising M, Jansen R, Jin F, Jorgenson E, Knowles JA, Kohane IS, Kraft J, Kretzschmar WW, Krogh J, Kutalik Z, Lane JM, Li Y, Li Y, Lind PA, Liu X, Lu L, MacIntyre DJ, MacKinnon DF, Maier RM, Maier W, Marchini J, Mbarek H, McGrath P, McGuffin P, Medland SE, Mehta D, Middeldorp CM, Mihailov E, Milaneschi Y, Milani L, Mill J, Mondimore FM, Montgomery GW, Mostafavi S, Mullins N, Nauck M, Ng B, Nivard MG, Nyholt DR, O’Reilly PF, Oskarsson H, Owen MJ, Painter JN, Pedersen CB, Pedersen MG, Peterson RE, Pettersson E, Peyrot WJ, Pistis G, Posthuma D, Purcell SM, Quiroz JA, Qvist P, Rice JP, Riley BP, Rivera M, Saeed Mirza S, Saxena R, Schoevers R, Schulte EC, Shen L, Shi J, Shyn SI, Sigurdsson E, Sinnamon GBC, Smit JH, Smith DJ, Stefansson H, Steinberg S, Stockmeier CA, Streit F, Strohmaier J, Tansey KE, Teismann H, Teumer A, Thompson W, Thomson PA, Thorgeirsson TE, Tian C, Traylor M, Treutlein J, Trubetskoy V, Uitterlinden AG, Umbricht D, Van der Auwera S, van Hemert AM, Viktorin A, Visscher PM, Wang Y, Webb BT, Weinsheimer SM, Wellmann J, Willemsen G, Witt SH, Wu Y, Xi HS, Yang J, Zhang F, eQTLGen. 23andMe. Arolt V, Baune BT, Berger K, Boomsma DI, Cichon S, Dannlowski U, de Geus ECJ, DePaulo JR, Domenici E, Domschke K, Esko T, Grabe HJ, Hamilton SP, Hayward C, Heath AC, Hinds DA, Kendler KS, Kloiber S, Lewis G, Li QS, Lucae S, Madden PFA, Magnusson PK, Martin NG, McIntosh AM, Metspalu A, Mors O, Mortensen PB, Müller-Myhsok B, Nordentoft M, Nöthen MM, O’Donovan MC, Paciga SA, Pedersen NL, Penninx BWJH, Perlis RH, Porteous DJ, Potash JB, Preisig M, Rietschel M, Schaefer C, Schulze TG, Smoller JW, Stefansson K, Tiemeier H, Uher R, Völzke H, Weissman MM, Werge T, Winslow AR, Lewis CM, Levinson DF, Breen G, Børglum AD, Sullivan PF, Major Depressive Disorder Working Group of the Psychiatric Genomics Consortium Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nature Genetics. 2018;50:668–681. doi: 10.1038/s41588-018-0090-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wuttke M, Li Y, Li M, Sieber KB, Feitosa MF, Gorski M, Tin A, Wang L, Chu AY, Hoppmann A, Kirsten H, Giri A, Chai J-F, Sveinbjornsson G, Tayo BO, Nutile T, Fuchsberger C, Marten J, Cocca M, Ghasemi S, Xu Y, Horn K, Noce D, van der Most PJ, Sedaghat S, Yu Z, Akiyama M, Afaq S, Ahluwalia TS, Almgren P, Amin N, Ärnlöv J, Bakker SJL, Bansal N, Baptista D, Bergmann S, Biggs ML, Biino G, Boehnke M, Boerwinkle E, Boissel M, Bottinger EP, Boutin TS, Brenner H, Brumat M, Burkhardt R, Butterworth AS, Campana E, Campbell A, Campbell H, Canouil M, Carroll RJ, Catamo E, Chambers JC, Chee M-L, Chee M-L, Chen X, Cheng C-Y, Cheng Y, Christensen K, Cifkova R, Ciullo M, Concas MP, Cook JP, Coresh J, Corre T, Sala CF, Cusi D, Danesh J, Daw EW, de Borst MH, De Grandi A, de Mutsert R, de Vries APJ, Degenhardt F, Delgado G, Demirkan A, Di Angelantonio E, Dittrich K, Divers J, Dorajoo R, Eckardt K-U, Ehret G, Elliott P, Endlich K, Evans MK, Felix JF, Foo VHX, Franco OH, Franke A, Freedman BI, Freitag-Wolf S, Friedlander Y, Froguel P, Gansevoort RT, Gao H, Gasparini P, Gaziano JM, Giedraitis V, Gieger C, Girotto G, Giulianini F, Gögele M, Gordon SD, Gudbjartsson DF, Gudnason V, Haller T, Hamet P, Harris TB, Hartman CA, Hayward C, Hellwege JN, Heng C-K, Hicks AA, Hofer E, Huang W, Hutri-Kähönen N, Hwang S-J, Ikram MA, Indridason OS, Ingelsson E, Ising M, Jaddoe VWV, Jakobsdottir J, Jonas JB, Joshi PK, Josyula NS, Jung B, Kähönen M, Kamatani Y, Kammerer CM, Kanai M, Kastarinen M, Kerr SM, Khor C-C, Kiess W, Kleber ME, Koenig W, Kooner JS, Körner A, Kovacs P, Kraja AT, Krajcoviechova A, Kramer H, Krämer BK, Kronenberg F, Kubo M, Kühnel B, Kuokkanen M, Kuusisto J, La Bianca M, Laakso M, Lange LA, Langefeld CD, Lee JJ-M, Lehne B, Lehtimäki T, Lieb W, Lifelines Cohort Study. Lim S-C, Lind L, Lindgren CM, Liu J, Liu J, Loeffler M, Loos RJF, Lucae S, Lukas MA, Lyytikäinen L-P, Mägi R, Magnusson PKE, Mahajan A, Martin NG, Martins J, März W, Mascalzoni D, Matsuda K, Meisinger C, Meitinger T, Melander O, Metspalu A, Mikaelsdottir EK, Milaneschi Y, Miliku K, Mishra PP, V. A. Million Veteran Program. Mohlke KL, Mononen N, Montgomery GW, Mook-Kanamori DO, Mychaleckyj JC, Nadkarni GN, Nalls MA, Nauck M, Nikus K, Ning B, Nolte IM, Noordam R, O’ J, Olafsson I, Oldehinkel AJ, Orho-Melander M, Ouwehand WH, Padmanabhan S, Palmer ND, Palsson R, Penninx BWJH, Perls T, Perola M, Pirastu M, Pirastu N, Pistis G, Podgornaia AI, Polasek O, Ponte B, Porteous DJ, Poulain T, Pramstaller PP, Preuss MH, Prins BP, Province MA, Rabelink TJ, Raffield LM, Raitakari OT, Reilly DF, Rettig R, Rheinberger M, Rice KM, Ridker PM, Rivadeneira F, Rizzi F, Roberts DJ, Robino A, Rossing P, Rudan I, Rueedi R, Ruggiero D, Ryan KA, Saba Y, Sabanayagam C, Salomaa V, Salvi E, Saum K-U, Schmidt H, Schmidt R, Schöttker B, Schulz C-A, Schupf N, Shaffer CM, Shi Y, Smith AV, Smith BH, Soranzo N, Spracklen CN, Strauch K, Stringham HM, Stumvoll M, Svensson PO, Szymczak S, Tai E-S, Tajuddin SM, Tan NYQ, Taylor KD, Teren A, Tham Y-C, Thiery J, Thio CHL, Thomsen H, Thorleifsson G, Toniolo D, Tönjes A, Tremblay J, Tzoulaki I, Uitterlinden AG, Vaccargiu S, van Dam RM, van der Harst P, van Duijn CM, Velez Edward DR, Verweij N, Vogelezang S, Völker U, Vollenweider P, Waeber G, Waldenberger M, Wallentin L, Wang YX, Wang C, Waterworth DM, Bin Wei W, White H, Whitfield JB, Wild SH, Wilson JF, Wojczynski MK, Wong C, Wong T-Y, Xu L, Yang Q, Yasuda M, Yerges-Armstrong LM, Zhang W, Zonderman AB, Rotter JI, Bochud M, Psaty BM, Vitart V, Wilson JG, Dehghan A, Parsa A, Chasman DI, Ho K, Morris AP, Devuyst O, Akilesh S, Pendergrass SA, Sim X, Böger CA, Okada Y, Edwards TL, Snieder H, Stefansson K, Hung AM, Heid IM, Scholz M, Teumer A, Köttgen A, Pattaro C. A catalog of genetic loci associated with kidney function from analyses of a million individuals. Nature Genetics. 2019;51:957–972. doi: 10.1038/s41588-019-0407-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu X, Eales JM, Jiang X, Sanderson E, Scannali D, Morris AP. Obesity as a Cause of Kidney Disease – Insights from Mendelian Randomisation Studies. medRxiv. 2020 doi: 10.1101/2020.09.13.20155234. [DOI]
- Yavorska OO, Burgess S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. International Journal of Epidemiology. 2017;46:1734–1739. doi: 10.1093/ije/dyx034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan S, Gill D, Giovannucci EL, Larsson SC. Obesity, Type 2 Diabetes, Lifestyle Factors, and Risk of Gallstone Disease: A Mendelian Randomization Investigation. Clinical Gastroenterology and Hepatology. 2021;6:S1542-3565(21)00001-X. doi: 10.1016/j.cgh.2020.12.034. [DOI] [PubMed] [Google Scholar]