Abstract
The present report assesses the association between clonal groupings, disease, and the virulence fingerprint of 76 South African Helicobacter pylori cagA+ strains isolated from 57 Cape-colored subjects. Two methods, repetitive extragenic palindromic (REP)-PCR and random amplified polymorphic DNA (RAPD)-PCR, were used to generate DNA fingerprints, and computer-assisted analysis was used to derive clusters. The two PCR techniques were only partially complementary (48%). REP-PCR fingerprints identified a distinct pathological cluster consisting of strains from 63% of the patients and was strongly associated with both disease (P < 0.00001) and the vacuolating cytotoxin A (vacA) signal sequence type (P < 0.003). RAPD-PCR fingerprinting was not associated with disease and was less strongly associated with vacA (P < 0.05) than REP-PCR was. Hierarchical analysis indicated that isolates from patients with peptic ulcer disease tended to cluster differently than isolates from patients with gastritis alone or gastric adenocarcinoma. These relationships are consistent with a loosely clonal population structure associated with disease for H. pylori in the Cape-colored population in South Africa.
Genetic diversity between strains of Helicobacter pylori is more marked than for any other bacterial species (13). The main reason for this is that recombination between strains is also higher than has been described for any other bacterium (8, 13). Despite this high level of recombination, comparisons between strains based both on sequence analysis and on multilocus enzyme electrophoresis have shown that clonal population structure is not entirely destroyed (1, 9). It has been hypothesized that different clonal groupings may be associated with disease, but two studies using repetitive extragenic palindromic (REP)-PCR have reached different conclusions (7, 16).
In contrast to genome-based strategies such as REP-PCR, studies examining the association between specific virulence marker genes and disease have shown consistent associations. Virulence markers associated with disease include the presence of the gene cagA and the cag pathogenicity island (PAI), the s1/m1 and s1/m2 types of the vacuolating cytotoxin gene, vacA, and type 1 of the epithelial contact-induced gene, iceA. Another marker studied by us and others in populations in which cagA is ubiquitous is the length of the 3′ portion of cagA: larger fragments appear to be more closely associated with disease (10, 20). Many of these markers are usually (although not invariably) associated with each other; for example, strains possessing the cag PAI usually have vacA with an s1 signal type (4). One possible explanation for this would be the remnant of an original clonal structure.
The Cape-colored population of South Africa has a high rate of gastric adenocarcinoma (GCa), has a high prevalence of H. pylori infection, and harbors an interesting range of H. pylori strains (10, 12). Sequence analysis has suggested that the clonal population structure is stronger than for European and Asian strains, possibly because of interstrain recombination over a shorter time frame due to relatively recent mixing of racial groups (13). Strains are all cagA positive, but there is diversity in other virulence markers (10, 11). Access to this population offered us the opportunity to address several unresolved issues. We aimed to use the population to assess whether there was an association between clonal groupings and disease. To do this, we planned to use REP-PCR, as previously described, and also a second method, random amplified polymorphic DNA (RAPD)-PCR. If associations with disease were shown, we aimed to define whether specific virulence gene markers were associated with specific clonal groupings or whether these markers were independently associated with disease.
MATERIALS AND METHODS
H. pylori patients and strains.
A total of 76 clinical H. pylori strains from 57 Cape-colored patients (20 strains from 16 patients with gastritis alone, 34 strains from 26 patients with recurrent or existing peptic [duodenal] ulcer disease [PUD], and 22 strains from 15 patients with GCa) were examined. Thirty-nine patients had a single isolate from the antrum, while 17 patients had isolates from both the antrum and the body of the stomach and 1 patient had 3 isolates (two from the antrum [different sites] and one from the body). The study population included 38 males (67%), and the median age of the group was 51 years (range, 11 to 82 years). The signal sequence and the mid-region of the gene vacA, the presence of cagA and the length of the 3′ region of cagA, the cag PAI status, and the iceA status of isolates were previously determined (10; M. Kidd et al., unpublished data). H. pylori strain 26695, isolated from a patient in the United Kingdom with gastritis (15), and strain J99, isolated from a patient in the United States with duodenal ulcer disease (3) were used as controls.
PCR amplification typing of H. pylori isolates.
For REP-PCR, the 18-mer degenerate primer pair of REP1R-Dt (17) and REP2-Dt (18) was used. Following initial denaturation at 95°C for 3 min, each reaction consisted of 35 cycles of denaturation at 95°C for 30 s, annealing and extension for 1 min, and a final extension at 72°C for 10 min. The annealing temperature was set at 45°C. Additional PCRs were performed with a single primer with 26695 and J99 to test the specificity of the REP product. Potential REP-PCR sites in strains 26695 and J99 were also identified using the National Center for Biotechnology Information (NCBI) BLAST server, and the expected number of band sizes was identified for each.
For RAPD-PCR, the specific H. pylori informative decanucleotide primer, 1254 (70% G+C), was used in a low-stringency PCR amplification (2). Briefly, 4 cycles of 94°C for 5 min, 36°C for 5 min, and 72°C for 5 min were followed by 30 cycles of 94°C for 1 min, 36°C for 1 min, and 72°C for 1 min, with a final extension of 10 min at 72°C. Potential RAPD-PCR sites in strains 26695 and J99 were identified using the NCBI BLAST server, and the expected number of band sizes was identified for each.
Twenty microliters of each PCR mixture was electrophoresed through a 1% agarose gel with a standard of 100 bp or a 1-kb DNA ladder (Roche Diagnostics, Johannesburg, South Africa). Variability in the intensity or shape of bands was not considered to represent differences.
Computer-assisted analysis.
The REP- and RAPD-PCR fingerprints of the H. pylori strains were analyzed with GelCompar software, Windows version 4.1 (Applied Math, Kortrijk, Belgium) (16). Both REP- and RAPD-PCR patterns were normalized using the 0.1-kbp molecular size standard. Comparison of the fingerprints was performed using the cluster analysis module. The calculation of a matrix of similarities was based on the Pearson product correlation coefficient and the final dendrogram was calculated by Ward's method (19). The similarity coefficient indicates the relatedness of the strains and was calculated using band positions (coefficient of Jaccard) per the GelCompar program. Briefly, for each couple of tracks, the coefficient of Jaccard [SJ]divides the number of corresponding bands by the total number of bands in both tracks using the formula SJ = nAB/[(nA + nB) − nAB], where nAB is the number of bands common for A and B, nA is the total number of bands in A, and nB is the total number of bands in B (16). A similarity coefficient of >70% was considered significant for analysis in this study.
For cluster analysis, the virulence data were summarized into two-way tables. Each table had 78 rows and columns for vacA genotype (s1 or s2 and m1 or m2), cagA 3′ fragment length (short, <600 bp; medium, 600 to 700 bp; long, >700 bp), cag PAI status (intact or partial), iceA status (iceA1+, iceA1+/2+, or iceA2+), REP fingerprint (REP1 or REP2), RAPD fingerprint (RAPD1 or RAPD2), and disease classification (gastritis alone, PUD, or GCa). The presence or absence of each character was binarily coded (present = 1, absent = 0). All analyses were performed with STATISTICA software (Gaithersburg, Md.)
Statistics.
Data were analyzed using the Wilcoxon rank sum test for independent samples, the chi-square test, or Fisher's exact test as appropriate. Probability levels of <0.05 were considered statistically significant.
RESULTS
Analysis of REP and RAPD sites in 26695 and J99.
Our preliminary studies using REP-PCR on clinical isolates yielded more numerous and smaller PCR amplicons than we expected. We therefore decided to assess whether REP-PCR products resulted purely from specific annealing to repetitive palindromic sequences or whether nonspecific annealing also occurred. To do this we assessed REP-PCR in the two previously completely sequenced strains, 26695 and J99. Firstly, we performed a BLAST analysis which revealed totals of 480 (26695) and 742 (J99) possible bands with sizes of >2 kb. The expected size range for the PCR protocol extends from 2 to 10 kb.
REP-PCR generated bands ranging in size from 0.4 to 5.6 kbp from the two strains. There were many small amplicons (<2,000 bp), and few amplicons of predicted sizes (20% [2 of 10] for 26695 and 13% [1 of 8] for J99, for bands of <10 kb) were demonstrated. The specificity of the REP products was then investigated by performing PCR with each primer separately. This generated nine bands with REP1R-Dt and five bands with REP2-Dt for 26695. Nine (64%) of the 14 bands were also found when both primers were used in a single reaction. For J99, nine bands with REP1R-Dt and six bands with REP2-Dt were identified. Three (20%) of the 15 bands were also found when both primers were used in a single reaction. Graphical analysis of the putative REP primer sites in both genomes demonstrated that both REP1R-Dt and REP2-Dt did not appear to be restricted to any specific region of the genome in either isolate. These data suggest that REP-PCR may be largely nonspecific and may instead possibly act as an arbitrarily primed protocol. Repeated REP-PCR (three different assays on different days) showed that the fingerprints from 26695 were stable.
We next examined the RAPD-PCR protocol. NCBI BLAST analysis revealed totals of 208 (26695) and 429 (J99) possible bands with sizes of >0.6 kb. The expected size range for the PCR protocol extends from 0.6 to 10 kb. RAPD-PCR was then performed on the two strains. RAPD-PCR generated bands ranging in size from 0.2 to 4.9 kbp. The results, which include a large number of small amplicons (<2,000 bp) as well as a low frequency of predicted amplicons (0% [0 of 7] for 26695 and 10% [1 of 10] for J99, for bands of <10 kb), confirm that RAPD-PCR is, as described, an arbitrarily primed protocol. Graphical analysis of the putative RAPD primer sites in both genomes demonstrated that these sites were normally distributed. Repeated RAPD-PCR (three different assays on different days) showed that the fingerprints from J99 were stable. Although both methodologies appear to be arbitrarily primed protocols, we decided to use both of them as described because of the previously described link between REP-PCR results and disease (7).
REP-PCR fingerprinting in clinical isolates.
REP-PCR generated bands ranging in size from 0.1 to 6 kbp for all 76 clinical isolates. Each isolate was distinguished by one to nine distinct bands, with an average of four different bands per isolate. No single amplification band was common to all strains. Repeated REP-PCR (three different runs with 10 isolates) showed that fingerprints were stable. Of the 76 isolates, 58 different DNA fingerprints were seen; 2 pairs of isolates from the same patients and 6 pairs of isolates from different patients had DNA fingerprints with a similarity coefficient of >99%.
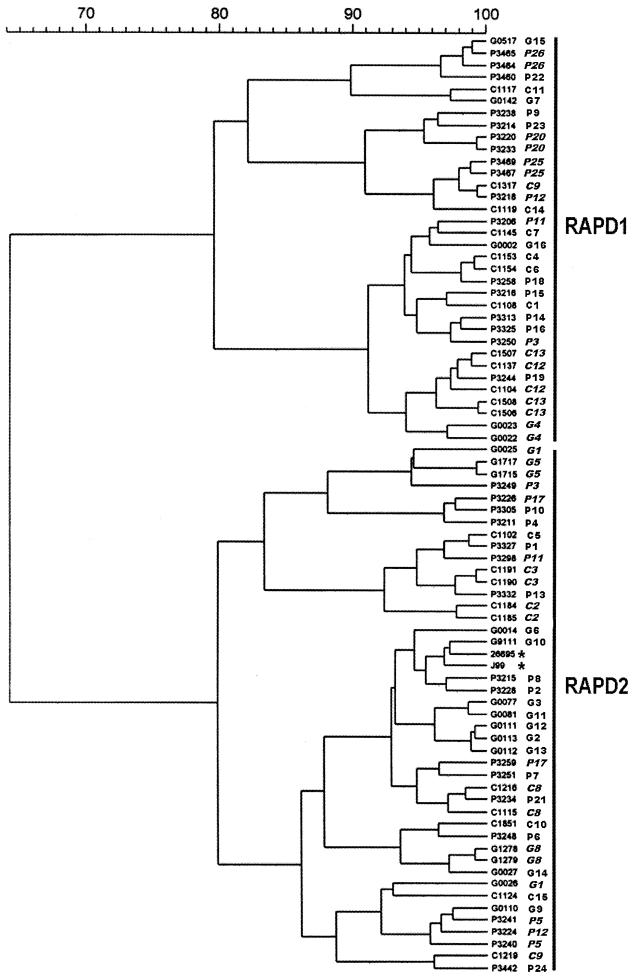
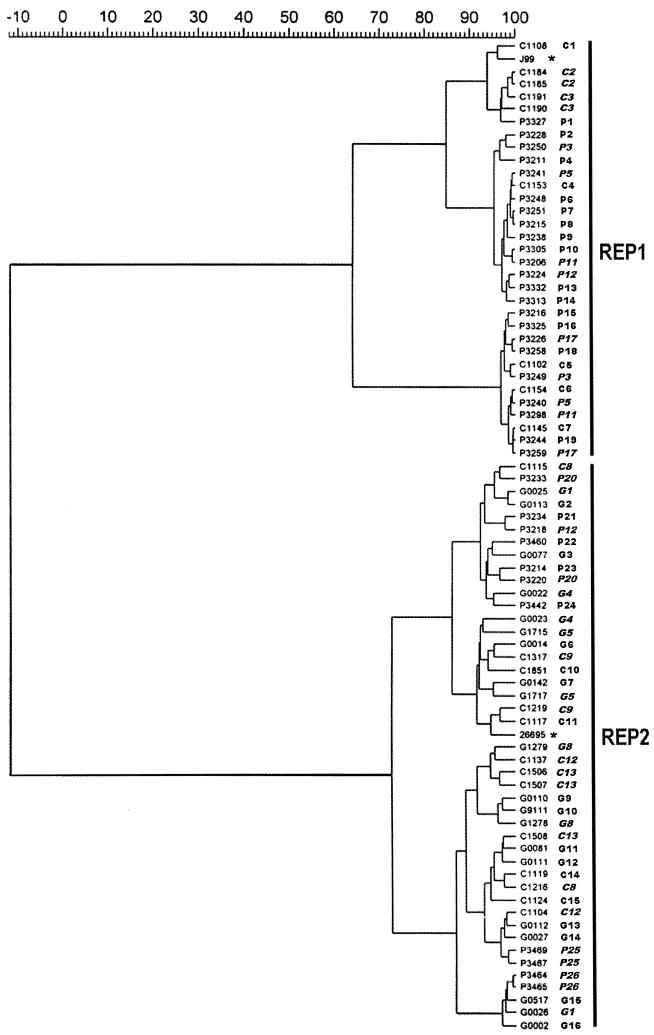
Cluster analysis revealed that there was a large degree of genetic heterogeneity among the H. pylori strains in this study (Fig. 1). Examination of the dendrogram demonstrated that REP-PCR divided isolates into two distinct and unrelated (similarity coefficient of 0%) clusters. Cluster 1 (REP1) consisted exclusively of 23 isolates from 19 patients with PUD, 9 isolates from 7 GCa patients, and J99, which had been isolated from a patient with duodenal ulcer disease. No isolates from patients with gastritis alone were present in this cluster. The second cluster (REP2) included 20 isolates from all 16 patients with gastritis alone, 13 isolates from 8 GCa patients, 11 isolates from 7 PUD patients, and 26695, which was isolated from a patient without clinically significant disease. Statistical analysis demonstrated a significant difference in the distribution of patients between these two clusters (χ2 = 20.14; P = 0.00004). More patients associated with clinically significant disease were present in REP1 (100%) than in REP2 (48%; P < 0.000001). Specifically, significantly more PUD patients (73%; P < 0.00001) and GCa patients (47%; P < 0.003) than patients with gastritis alone (0%) were present in REP1. While most patients with two isolates were present in the same cluster, one PUD patient had an isolate in REP1 (P3224) and one in REP2 (P3218).
FIG. 1.
Dendrogram of H. pylori REP-PCR DNA fingerprints. Similarity coefficients are included in the top bar. G, gastritis alone; P, PUD; C, GCa. J99 and 26695 are indicated by asterisks. Strain numbers (immediately adjacent to the dendrogram) and patient numbers are indicated, and numbers from patients with more than one isolate are italicized. Two separate clusters are indicated.
RAPD-PCR fingerprinting.
RAPD-PCR resulted in one to seven distinct bands, with an average of four different bands per isolate. Repeated RAPD-PCR (n = 3) showed that fingerprints were stable. The band sizes ranged in size from 0.1 to 4.6 kbp, and there was no amplification band common to all strains. In RAPD-PCR of the 76 isolates, 60 different DNA fingerprints were seen; 5 pairs of isolates from the same patients and 3 pairs of isolates from different patients had DNA fingerprints with similarity coefficients of >99%.
Cluster analysis revealed that there was a large degree of genetic heterogeneity among the H. pylori strains in this study (Fig. 2). Examination of the dendrogram demonstrated two readily distinguishable clusters of H. pylori strains at a similarity coefficient of 64.2% ± 4.3%. Cluster 1 (RAPD1) included 5 isolates from 4 patients with gastritis alone, 12 isolates from 9 patients with GCa, and 17 isolates from 14 PUD patients. The second cluster (RAPD2) consisted of 17 isolates from 15 patients with PUD, 10 isolates from 7 GCa patients, and 15 isolates from 12 patients with gastritis alone. Both type strains were also present in this cluster. Isolates associated with specific disease subtypes did not appear to be differently distributed between the two clusters (χ2 = 3.53; P = 0.17). In addition, subgroup analysis did not suggest that patients with clinically significant disease clustered separately (P < 0.07). Interestingly, isolates from 4 of the 17 patients with multiple strains were present in both RAPD1 and RAPD2.
FIG. 2.
Dendrogram of H. pylori RAPD-PCR DNA fingerprints. Similarity coefficients are included in the top bar. G, gastritis alone; P, PUD; C, GCa. The strain and patient numbers are indicated as described for Fig. 1. J99 and 26695 are indicated by asterisks. The two clusters are indicated.
Relationship between REP-DNA fingerprinting and virulence status.
The dendrogram based on REP-DNA contains two clusters that segregate with pathology. We have previously identified that specific virulence gene markers also segregate with pathology. We therefore next determined whether these specific virulence factors were associated with the REP clusters. Analysis of vacA signal sequences demonstrated that vacA type s1 occurred more frequently in the pathology-associated REP1 cluster (100%) than in cluster 2 (77%; P < 0.003). There were also differences in the segregation of vacA mid-regions between the two groups (88 versus 61%; P = 0.01). No specific segregation of cagA 3′ fragment length or cag PAI status (intact or partial) was evident. iceA genotypes were, however, differentially segregated (χ2 = 7.43; P < 0.03). Specifically, iceA1+/2+ occurred more often in REP2 (62 versus 33%; P < 0.02), while iceA2 occurred more often in REP1 (42 versus 18%; P < 0.02). These results suggest that there may also be a strong association between the REP fingerprint and the vacA and iceA allelic types.
Next, to define whether specific virulence factors were associated with disease independent of REP grouping, we examined their association with disease within REP clusters. Because all strains in the REP1 cluster were disease associated, this cluster was not suitable for analysis. However, within REP2, significantly more PUD isolates (100%; P < 0.004) and GCa isolates (100%; P < 0.002) were vacA s1 positive than gastritis-alone isolates (50%). There were no significant differences in the segregation of vacA mid-region alleles or in iceA genotypes. These results suggest that at least within the REP2 cluster (which is an amalgam of strains from patients with and without gastroduodenal disease), virulent type s1 vacA is independently associated with disease. The disease homogeneity in the REP1 cluster precludes meaningful analysis for REP1 strains.
Relationship between RAPD-DNA fingerprinting and virulence status.
The dendrogram based on RAPD-PCR analysis contains two clusters that do not appear to correlate with pathology. We therefore tested whether there was an association between virulence factors and the RAPD fingerprint. Analysis of signal sequences demonstrated that the vacA s1 type occurred more frequently in RAPD1 (100%) than in RAPD2 (77%; P < 0.002), while vacA s2 alleles were found exclusively in the second cluster. There were no differences in the segregation of vacA mid-regions between the two clusters (74% m1 versus 70% m2; P = 0.5). No specific segregation of cagA 3′ fragment lengths was evident. The cag PAI status was, however, differentially segregated (P < 0.002). Specifically, an intact PAI was present more often for RAPD1 isolates (85%) than for RAPD2 isolates (52%). While iceA genotypes did not appear to be differentially segregated between the two clusters (χ2 = 5.4; P = 0.07), iceA2 genotypes were present significantly more often (P < 0.02) in RAPD2 (63%) than in RAPD1 (15%). These results suggest that there may be an association between the RAPD fingerprint and the vacA and cag PAI status.
Next, to define whether specific virulence factors were associated with disease independent of RAPD grouping, we examined their association with disease within each cluster. Since all RAPD1 isolates were vacA s1 and m1 positive, this cluster was not suitable for further analysis. However, within RAPD2 significantly more PUD isolates were vacA s1 positive (100%; P < 0.00005) and vacA m1 positive (94%; P < 0.0005) than gastritis-alone isolates (s1, 33%; m1, 33%). This was similar for GCa isolates (vacA s1, 100%, P < 0.001; vacA m1, 80%, P < 0.03). No specific segregation of iceA genotypes was evident. The cag PAI status was, however, differentially segregated between disease groups in RAPD2 (χ2 = 12.13; P < 0.003). Specifically, an intact PAI was present more often in PUD isolates (78%; P < 0.001) and GCa isolates (60%; P < 0.05) than in gastritis-alone isolates (23%).
Overall, these results suggest that RAPD fingerprints are not clearly associated with disease potential and are unable to identify genomic correlations with disease. The virulence factors considered are more strongly associated with disease independently of RAPD fingerprint group.
Cluster analysis of virulence and fingerprint data.
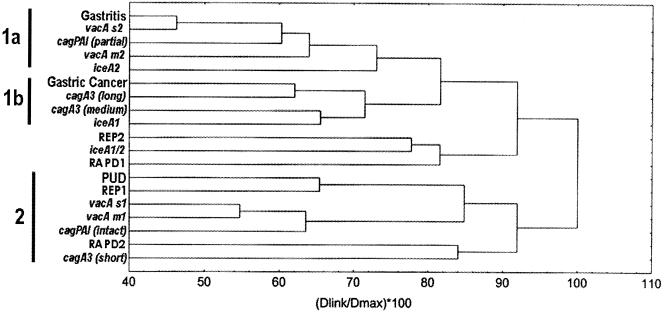
The results thus far suggest that REP-PCR generates two clusters, one of which is strongly associated with both pathology and virulence fingerprint, while RAPD-PCR generates two clusters which are more heterogeneous and are less strongly related to these parameters. In order to investigate further the interrelationship of these clusters, specific virulence factors, and disease, we performed a hierarchical (cluster) analysis using unweighted pair-group average and Euclidean distances for all the variables (pathology, REP and RAPD fingerprints, and virulence fingerprint). This analysis resulted in two distinct clusters (Fig. 3) which were clearly delineated. Cluster 1 was defined by the presence of isolates from patients with gastritis alone and GCa, vacA s2 and m2, a partial cag PAI, long cagA 3′ fragment lengths, the iceA alleles, and the RAPD1 and REP2 fingerprints. Further analysis of cluster 1, however, suggested the presence of two subclusters (1a and 1b). Cluster 1a demonstrated that gastritis, vacA s2, a partial cag PAI, and the iceA2 genotype were related, while cluster 1b was characterized by GCa isolates, the medium and long cagA 3′ fragments, and iceA1. The RAPD1 and REP2 fingerprints occurred outside these clusters. Cluster 2 included isolates from patients with PUD, the REP1 fingerprint, the virulence-associated vacA s1 and m1 alleles, an intact cag PAI, the RAPD2 fingerprint, and a short cagA 3′ fragment length. This suggests that PUD was associated with specific virulence genes as well as specific genomic fingerprints. Analysis using the k-means test confirmed the presence of two clusters.
FIG. 3.
Cluster diagram based on hierarchical analysis using genomic analysis (REP and RAPD fingerprints) and virulence data (vacA, cagA and cag PAI, and iceA genotypes). The two major clusters are evident, as are the subclusters of cluster 1. (Dlink/Dmax) × 100, similarity coefficient.
DISCUSSION
The DNA fingerprints of 76 cagA+ H. pylori isolates from 57 Cape-colored individuals with a spectrum of gastroduodenal diseases were analyzed by two genomic DNA-based PCR techniques, REP-PCR and RAPD-PCR. This study indicates that REP-PCR probably acts as an arbitrarily primed protocol with some site-directed genome analysis. Because of differences in primers and annealing temperatures, these two techniques, however, probe different portions of the genome that may include virulence-specific areas and generate different fingerprints. Both the REP- and RAPD-PCR fingerprints were stable and reproducible for each isolate. It appears, however, that REP-PCR may be more useful than RAPD-PCR for generating DNA fingerprints that can classify pathogenic South African strains for this study population.
REP-PCR typing has generated stable, reproducible DNA fingerprints that have been successfully used to discriminate H. pylori strains associated with PUD in a U.S. study (7), but this was not confirmed in a European study (16). The results from the present study suggest that REP fingerprints may be useful for defining disease-associated strains in a recent bacterial population derived from mixed population groups (European, Southeast Asian, and African). In particular, REP-PCR clearly discriminates a pathogenic cluster that accounts for isolates from ∼75% of PUD patients and ∼50% of GCa patients in our study group. This cluster was also distinguished by the almost universal presence of the vacA s1/m1 allelic type as well as by the inclusion of the type strain (J99; vacA s1/m1) isolated from a patient with duodenal ulcer disease. The vacA s2 allele (with low or noncytotoxic activity and not associated with peptic ulceration in the United States [5]) was completely absent from this cluster. The second REP cluster was composed chiefly of gastritis-alone isolates (61%) and consisted of 52% of the study population. In addition, vacA s2 alleles were found exclusively in this cluster, while the vacA m2 allele, which has been less strongly correlated with epithelial injury (epithelial degeneration, mucus depletion, and microscopic erosions) than mid-region m1 (6), tended to segregate into this cluster (81%). Interestingly, a fair percentage of GCa isolates (59%) was also found in cluster 2. These results suggest that carcinogenic potential, unlike PUD potential, may not be associated with any particular REP cluster.
RAPD-PCR uses an oligonucleotide of arbitrarily chosen sequence to prime DNA synthesis from pairs of sites to which it is matched or partially matched. It results in strain-specific arrays of DNA products and has been successfully used to analyze DNA diversity among clinical isolates of H. pylori (2, 14). The RAPD results from this study using a high (70%)-GC-content primer show two clusters that are quite heterogeneous. In contrast to REP fingerprinting, which distinguished peptic ulcer isolates in a separate cluster, RAPD-PCR may not be associated with PUD potential, because isolates from PUD patients were present in similar numbers in both RAPD1 and RAPD2. Similar to REP fingerprinting, there was a correlation between RAPD-PCR and vacA polymorphism. Strains with the vacA s2 allele were exclusively present in isolates from RAPD2, while RAPD1 contained only strains with vacA s1 alleles. As noted for REP analysis, the iceA2 genotype segregated with RAPD fingerprints. Interestingly, an intact cag PAI also appeared to segregate with the RAPD1 fingerprint. Sixty-four percent of strains identified in the virulence-associated REP1 cluster were also present in the heterogeneous RAPD2 cluster (but made up only 48% of the isolates identified in this cluster), suggesting that the RAPD and REP techniques may be only partially complementary.
Using a different computer algorithm revealed a complementary interpretation of the results. Binary coding and hierarchical analysis using unweighted pair-group average and k-means clustering with the STATISTICA program demonstrated a clear clustering of PUD, virulence-associated H. pylori alleles (vacA s1/m1 and an intact cag PAI), and the REP1 and RAPD2 fingerprints. This was distinct from a second cluster containing the remainder of the variables. The relationships noted in this analysis and analysis of the REP and RAPD groups support the suggestion that H. pylori in the Cape-colored population may be more clonal than European strains (13).
In conclusion, this analysis of genome relatedness demonstrated by computer-assisted analysis that clustering of REP-PCR fingerprints was strongly associated with disease (particularly PUD) and with vacA signal sequence type in South African isolates from Cape Town. RAPD-PCR fingerprints were also associated with these parameters, although less strongly. Isolates from peptic ulcer patients and patients without ulcers tended to cluster differently, but strains from patients with GCa showed no definite genomic clustering. The later observation may be of some importance given the often controversial causal relationship between infection with the organism and this disease. Some specific virulence factors, such as the s1 type of vacA, were independently associated with disease within genomic clusters. Additional studies are now needed of other mixed populations to confirm and extend these results.
ACKNOWLEDGMENTS
Part of this project was sponsored by the Freda and David Bekker Award (to J.A.L.) and the ABBOTT-SAGES Award (to M.K.). M.K. is a recipient of a Claude Harris Leon Foundation Fellowship.
We thank Mark Achtman of the Max-Planck Institut fur Infektionsbiologie for critically reviewing the manuscript.
REFERENCES
- 1.Achtman M, Azuma T, Berg D, Ito Y, Morelli G, Pan Z-J, Suerbaum S, Thompson S, van der Ende A, van Doorn L-J. Recombination and clonal groupings within Helicobacter pylori from different geographical regions. Mol Microbiol. 1999;32:459–470. doi: 10.1046/j.1365-2958.1999.01382.x. [DOI] [PubMed] [Google Scholar]
- 2.Akopyanz N, Bukanov N, Westblom T, Kresovich S, Berg D. DNA diversity among clinical isolates of Helicobacter pylori detected by PCR-based RAPD fingerprinting. Nucleic Acids Res. 1992;20:5137–5142. doi: 10.1093/nar/20.19.5137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Alm R, Ling L-S, Moir D, King B, Brown E, Doig P, Smith D, Noonan B, Guild B, deJonge B, Carmel G, Tummino P, Caruso A, Uria-Nickelsen M, Mills D, Ives C, Gibson R, Merberg D, Mills S, Jiang Q, Taylor D, Vovis G, Trust T. Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature. 1999;397:176–180. doi: 10.1038/16495. [DOI] [PubMed] [Google Scholar]
- 4.Atherton J, Cao P, Peek R, Tummuru M, Blaser M, Cover T. Mosaicism in vacuolating cytotoxin alleles of Helicobacter pylori. J Biol Chem. 1995;270:17771–17777. doi: 10.1074/jbc.270.30.17771. [DOI] [PubMed] [Google Scholar]
- 5.Atherton J, Peek R, Tham K, Cover T, Blaser M. Clinical and pathological importance of heterogeneity in vacA, the vacuolating cytotoxin gene of Helicobacter pylori. Gastroenterology. 1997;112:92–99. doi: 10.1016/s0016-5085(97)70223-3. [DOI] [PubMed] [Google Scholar]
- 6.Atherton J. The clinical relevance of strain types of Helicobacter pylori. Gut. 1997;40:701–703. doi: 10.1136/gut.40.6.701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Go M, Chan K, Versalovic J, Koeuth T, Graham D, Lupski J. Cluster analysis of Helicobacter pylori genomic DNA fingerprints suggests gastroduodenal disease-specific associations. Scand J Gastroenterol. 1995;30:640–646. doi: 10.3109/00365529509096306. [DOI] [PubMed] [Google Scholar]
- 8.Gottke M, Fallone C, Barkun A, Trautmann M, Tong J, Fainsilber T, Hahn H, Korber J, Loewe A, Beech R. Genetic variability determinants of Helicobacter pylori: influence of clinical background and geographic origin of isolates. J Infect Dis. 2000;181:1674–1681. doi: 10.1086/315425. [DOI] [PubMed] [Google Scholar]
- 9.Jorgensen M, Daskalopoulus G, Warburton V, Mitchell H, Hazell S. Multiple strain colonization and metronidazole resistance in Helicobacter pylori-infected patients: identification from sequential and multiple biopsy specimens. J Infect Dis. 1996;174:631–635. doi: 10.1093/infdis/174.3.631. [DOI] [PubMed] [Google Scholar]
- 10.Kidd M, Lastovica A, Atherton J, Louw A. Heterogeneity in the Helicobacter pylori genes vacA and cagA: associated with gastroduodenal disease in South Africa? Gut. 1999;45:499–503. doi: 10.1136/gut.45.4.499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kidd M, Lastovica A, Atherton J, Louw J. Specific genotypes of Helicobacter pylori vacA and cagA, but not the presence of cagA, are associated with gastroduodenal disease in South Africa. Gastroenterology. 1999;116:G0928. [Google Scholar]
- 12.Letley D, Lastovica A, Louw J, Hawkey C, Atherton J. Allelic diversity of Helicobacter pylori vacuolating cytotoxin gene in South Africa: rarity of the vac s1a genotype and natural occurrence of an s2/m1 allele. J Clin Microbiol. 1999;37:1203–1205. doi: 10.1128/jcm.37.4.1203-1205.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Suerbaum S, Maynard Smith J, Bapumia K, Morelli G, Smith N, Kunstmann E, Dyrek L, Achtman M. Free recombination within Helicobacter pylori. Proc Natl Acad Sci USA. 1998;95:12619–12624. doi: 10.1073/pnas.95.21.12619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Taylor N, Fox J, Akopyants N, Berg D, Thompson N, Shames B, Yan L, Fontham E, Janney F, Hunter F, Correa P. Long-term colonization with single and multiple strains of Helicobacter pylori assessed by DNA fingerprinting. J Clin Microbiol. 1995;33:918–923. doi: 10.1128/jcm.33.4.918-923.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tomb J-F, White O, Kerlavage A, Clayton R, Sutton G, Fleischmann R, Ketchum K, Klenk H, Gill S, Dougherty B, Nelson K, Quackenbush J, Kirkness E, Lee N, Adams M, Venter J. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature. 1997;388:539–547. doi: 10.1038/41483. [DOI] [PubMed] [Google Scholar]
- 16.van Doorn N, Namavar F, Kusters J, van Rees E, Kuipers E, de Graaf J. Genomic DNA fingerprinting of clinical isolates of Helicobacter pylori by REP-PCR and restriction fragment end-labelling. FEMS Microbiol Lett. 1998;160:145–150. doi: 10.1111/j.1574-6968.1998.tb12904.x. [DOI] [PubMed] [Google Scholar]
- 17.Versalovic J, Koeuth T, Lupski J. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 1991;19:6823–6831. doi: 10.1093/nar/19.24.6823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Versalovic J, Kapur V, Mason E, Shah U, Koeuth T, Lupski J, Musser J. Penicillin-resistant Streptococcus pneumoniae strains recovered in Houston: identification and molecular characterization of multiple clones. J Infect Dis. 1993;167:850–856. doi: 10.1093/infdis/167.4.850. [DOI] [PubMed] [Google Scholar]
- 19.Ward J. Hierarchical grouping to optimize an objective function. J Am Stat Assoc. 1963;58:236–244. [Google Scholar]
- 20.Yamaoka Y, Kodama T, Kashima K, Graham D, Sepulveda A. Variants of the 3′ region of the cagA gene in Helicobacter pylori isolates from patients with different H. pylori-associated diseases. J Clin Microbiol. 1998;36:2258–2263. doi: 10.1128/jcm.36.8.2258-2263.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]