Abstract
The availability of millions of SARS-CoV-2 (Severe Acute Respiratory Syndrome-Coronavirus-2) sequences in public databases such as GISAID (Global Initiative on Sharing All Influenza Data) and EMBL-EBI (European Molecular Biology Laboratory-European Bioinformatics Institute) (the United Kingdom) allows a detailed study of the evolution, genomic diversity, and dynamics of a virus such as never before. Here, we identify novel variants and subtypes of SARS-CoV-2 by clustering sequences in adapting methods originally designed for haplotyping intrahost viral populations. We asses our results using clustering entropy—the first time it has been used in this context. Our clustering approach reaches lower entropies compared with other methods, and we are able to boost this even further through gap filling and Monte Carlo-based entropy minimization. Moreover, our method clearly identifies the well-known Alpha variant in the U.K. and GISAID data sets, and is also able to detect the much less represented (<1% of the sequences) Beta (South Africa), Epsilon (California), and Gamma and Zeta (Brazil) variants in the GISAID data set. Finally, we show that each variant identified has high selective fitness, based on the growth rate of its cluster over time. This demonstrates that our clustering approach is a viable alternative for detecting even rare subtypes in very large data sets.
Keywords: clustering, entropy, fitness, genomic surveillance, viral subtypes, viral variants
1. Introduction
The novel coronavirus SARS-CoV-2 (Severe Acute Respiratory Syndrome-Coronavirus-2), which is responsible for the COVID-19 disease, was first detected in Wuhan, China, at the end of 2019 (Wu et al., 2020; Zhou et al., 2020). COVID-19 was declared a global pandemic in March 2020 by the World Health Organization (WHO). According to recent data from the WHO (WHO, 2021), there have been almost 4 million deaths due to COVID-19, and there have been hundreds of millions of confirmed cases so far, while over 3 billion vaccine doses have been administered. As the virus continues to spread throughout countries and regions across the globe, it continues to mutate, as seen in the genomic variations among the millions of sequences, which are available in public databases such as GISAID (Global Initiative on Sharing All Influenza Data) (Elbe and Buckland-Merrett, 2017). This mutational variability can be used to understand the evolution, genomic diversity, and dynamics of SARS-CoV-2, and to generate hypotheses on how the virus has evolved and spread since it first originated.
An important part of this dynamics are the subsets of sequences (or subtypes) that vary more than others in terms of genomic content, which continue to emerge. In some cases, these subtypes appear in an atypically large number or are associated with an extremely high growth rate, indicating a possible fitness advantage (transmissibility, evasion from therapies or vaccines, etc.) of this genomic variation. The best example of this is the Alpha (or B.1.1.7; Rambaut et al., 2020) variant, which differs from the typical sequence by about 30 mutations, and comprises hundreds of thousands of the currently available sequences. The Alpha variant was first detected in the United Kingdom at the end of summer 2020, where it grew to more than a third of the infected population in the United Kingdom by mid-December 2020, as seen in the EMBL-EBI (European Molecular Biology Laboratory-European Bioinformatics Institute) (the United Kingdom) database EMBL-EBI (2020). One of the first variants of concern (VoCs), the Alpha variant has undergone much investigation, some studies (Volz et al., 2021) showing it to be between 40% and 80% more transmissible. This variant is now found in countries all over the world, some for which it is the dominant variant (e.g., the United States). Despite this, the origins of the Alpha variant are still contested, popular hypotheses including immunocompromised patients, the loss of records, or even minks as an intermediary host. There are now roughly a dozen variants identified around the globe (Table 1)—an interesting question is whether they have or could potentially have the same degree of divergence as the Alpha variant.
Table 1.
Some Known Variants of SARS-CoV-2 (Severe Acute Respiratory Syndrome-Coronavirus-2)
| Variant | Region | Lineage | S/Gen. | Source |
|---|---|---|---|---|
| Gamma | Brazil | P.1(B.1.1.28.1) | 10/21 | Naveca et al. (2021b) |
| Zeta | Brazil | P.2(B.1.1.28.2) | 1/5 | |
| Epsilon | California | B.1.427/B.1.429 | 3/5 | Zhang et al. (2021) |
| Iota | New York | B.1.526 | 6/16 | West et al. (2021) |
| Beta | South Africa | B.1.351 | 9/21 | Galloway et al. (2021) |
| Alpha | The United Kingdom | B.1.1.7 | 8/17 | |
| Kappa | India | B.1.167.1 | 8/17 | Yadav et al. (2021) |
| Delta | India | B.1.167.2 | 8/17 |
Known variants of SARS-CoV-2. The five columns, starting from the left, are: Variant (Greek name); Region where it was first identified; PANGOLIN Lineage identifier; Number of mutations on the S gene/entire genome; and Source.
The typical approach that is used to recover such knowledge from viral sequences is to construct a phylogenetic tree (Hadfield et al., 2018; du Plessis et al., 2021) of evolution. However, with the high computational complexity of building a tree, more than a million sequences pose a scalability challenge for such methods (Hadfield et al., 2018; du Plessis et al., 2021; Vrbik et al., 2015). An orthogonal approach to trees is to build transmission networks of infection—the structure of the network revealing general trends. In Skums et al. (2020), the authors show that such a network is scale-free, that is, few genomic variants are responsible for the majority of possible transmissions. A third alternative to studying the mutational variability of SARS-CoV-2 that we use here is to cluster sets of sequences. While individual sequences are often unique, the sheer number of sequences available is expected to unveil meaningful groups and trends. Moreover, since most clustering techniques are much faster than, for example, tree building (Hadfield et al., 2018; du Plessis et al., 2021), such an approach can easily scale to the full size of the current data sets to leverage this information. The idea is that clusters of similar sequences should correspond to variants and subtypes, such as the Alpha variant mentioned above.
In this work, we cluster sequences by adapting methods that were originally designed for finding viral haplotypes from intrahost viral populations. The idea is that we use, for example, CliqueSNV (Clique Single Nucleotide Variation) (Knyazev et al., 2020), to find haplotypes in the massively interhost viral population, using them as cluster centers in categorical clustering algorithms such as k-modes (Huang, 1997) to find subtypes. A measure we use to assess the clustering approaches, in the absence of a ground truth, is clustering entropy. This notion was introduced in Li et al. (2004), where the authors show that minimizing clustering entropy is equivalent to maximizing the likelihood that the set of sequences is generated from a set of subtypes, closely modeling this setting of viral sequence evolution. Moreover, Li et al. (2004) show that clustering entropy is a convex function, allowing us to apply general optimization techniques such as the Monte Carlo method to minimizing entropy directly, as the objective. Finally, we use the subtypes found from our clustering techniques to patch gaps in the sequences, as an alternative to filling in the missing data with the reference genome, for example. This applies in particular to sequences collected before March, when SARS-CoV-2 sequencing and alignment were still being refined.
To validate our approaches, we use data from both the GISAID and EMBL-EBI (the United Kingdom) databases mentioned above (Table 2). In a general comparison of methods and parameter settings on data from GISAID, we show that our CliqueSNV-based approach can achieve the lowest clustering entropy. What is interesting is that our gap filling approach allowed each method to lower its entropy even more. We then tested out a Monte Carlo-based entropy minimization technique to show that it gives our method an even further edge on lowering the entropy. We compared various methods on their ability to find subtypes in the U.K. data set, verifying this with the “ground truth” clusters that arise from metadata tagging each sequence with its lineage (e.g., B.1.1.7, the Alpha variant).
Table 2.
Data Sets Used in the Experiments
| Data set | Database | Start | End | No. sequences |
|---|---|---|---|---|
| GISAID 1 | GISAID | December 24, 2019 | November 7, 2020 | 209,334 |
| GISAID 1A | GISAID | December 24, 2019 | March 5, 2020 | 3688 |
| United Kingdom | EMBL-EBI | January 24, 2020 | December 29, 2020 | 88,008 |
| GISAID 2 | GISAID | December 24, 2019 | April 4, 2021 | 1,000,982 |
The four data sets that are used in the experiments of Section 5. The five columns, starting from the left, are: Name we use here; Database, it is from GISAID (Elbe and Buckland-Merrett, 2017) or EMBL-EBI (EMBL-EBI, 2020); Earliest collection date of any sequence; Latest collection date; and Number of sequences. Note that GISAID 1A is a subset of GISAID 1, which is, in turn, a subset of GISAID 2.
Our CliqueSNV-based approach identified the Alpha variant with a significantly higher precision and specificity compared with other methods, based on these metadata. This reinforces the notion that clustering entropy is an appropriate measure of the quality of clustering in this context. We then used our CliqueSNV-based clustering approach to identify subtypes in the GISAID data set, again verifying this from metadata. While our method clearly identified the Alpha variant, it was also able to detect the lesser represented Beta (South Africa), Epsilon (California), and Gamma and Zeta (Brazil) variants with specificities around 50%. What is particularly interesting about this is that these lesser represented variants comprise less than 1% of the GISAID data set (which contains more than one million sequences), yet our method was able to detect them with a much higher specificity. Finally, we validate our approaches to finding subtypes using the fitness coefficient, a third measure of clustering quality orthogonal to both entropy and specificity mentioned above. The fitness coefficient, introduced in Skums et al. (2012), is an assessment of the selective fitness of a subtype, based on the number of sequences in the corresponding cluster, and the rate at which this grows over time. Our results show that fitness tends to corroborate with these other two measures, further strengthening our results. This demonstrates that we can use clustering for the identification and surveillance of new variants, which have the potential to grow quickly, or become a threat to public health.
This article is structured as follows. In Section 2, we detail the various approaches to clustering, gap filling, and entropy minimization that we use in this work. In Section 3, we specify the measures that we used to assess the clustering approaches—clustering entropy and the fitness coefficient. In Section 4, we give details on the data sets that we use in this study, as well as some of the known variants that we can expect to find in their metadata. In Section 5, we report the results of the experiments we performed to assess the various clustering approaches, gap filling, entropy minimization, and the identification of subtypes. Section 6 then concludes the article with a discussion of the contributions of our approach, in light of these results.
2. Methods Used in Clustering
We outline in this section all of the methods that were used in clustering nucleotide sequences of the SARS-CoV-2 virus.
2.1. CliqueSNV
Since we are clustering viral sequences to identify subtypes, we propose to use currently existing tools that were developed to identify subtypes in intrahost viral populations from NGS (Next Generation Sequencing) data reviewed in Knyazev et al. (2021), such as Savage (Baaijens et al., 2017), PredictHaplo (Prabhakaran et al., 2014), and aBayesQR (Ahn and Vikalo, 2018). However, our setting is slightly different, where the data consist of large collections of interhost consensus sequences gathered from different regions and countries around the world (Elbe and Buckland-Merrett, 2017; EMBL-EBI, 2020). We expect, however, that such tools are appropriate at this scale: now the “host” is an entire region or country, and we reconstruct the subtypes, or variants, and their dynamics within these regions or countries. The SARS-CoV-2 sequences in GISAID are consensus sequences of approximate length 30K. Such sequences by quality and length have similar properties as PacBio reads. We choose CliqueSNV since it performed very well on PacBio reads (Knyazev et al., 2020).
2.2. k-modes
Since nucleotide sequences can be viewed as objects on categorical attributes—the attributes are the genomic sites, and the categories are A, C, G, T (and −, a gap)—we use k-modes (Huang, 1997, 1998) for clustering. The k-modes approach is almost identical to k-means (Anderberg, 1973; MacQueen et al., 1967), but it is based on the notion of mode (rather than the Euclidean mean), making it appropriate for clustering categorical data. Indeed, the Euclidean mean of three nucleotides has little meaning in this context, and may not even be well-defined, for example, in cases where the “distance” from A to G is different than from G to A. Similar observations were made in the context of cancer mutation profiles (Ciccolella et al., 2020), in the form of absence/presence information. Treating these as categories, in using k-modes (rather than as 0's and 1's, in using k-means) resulted in a clustering approach (Ciccolella et al., 2021a) that, when used as a preprocessing step, allowed cancer phylogeny building methods to attain a higher accuracy (Ciccolella et al., 2021b), and in some cases with much lower runtimes (Jahn et al., 2016). We briefly describe the k-modes approach in the context of clustering nucleotide sequences as follows.
The mode q of a cluster C of sequences on categorical attributes is another “sequence” on , which minimizes
| (1) |
where d is some categorical dissimilarity measure (e.g., Hamming distance) between the sequences we are considering. Note that q is not necessarily an element of C. For a set S of sequences on attributes , we are given some initial set of k cluster “centers” (each on ). The k-modes approach (similarly to k-means) then operates according to the iteration:
compute the dissimilarity between each sequence and each center ;
assign each sequence to the closest center based on the first step, resulting in a clustering with k clusters; and
compute the mode of each cluster from the second step, resulting in a new set of k centers;
until convergence, that is, the clustering does not change after an iteration.
In this article, we cluster sequences of SARS-CoV-2 with k-modes using three different ways to compute the initial set Q of cluster centers, and using two different dissimilarity measures d. The three ways to compute the initial set of cluster centers are as follows:
-
1.
choose k random sequences from the data set;
-
2.
choose k centers that are maximally pairwise distant from each other; and
-
3.
use the centers (the subtypes) found by CliqueSNV.
The two different dissimilarity measures that we use are (1) the Hamming distance, and (2) the TN-93 distance (Tamura and Nei, 1993).
2.3. MeShClust
For comparison purposes, we also apply methods designed for clustering metagenomics and multiviral sequencing data. We clustered the sequences using MeShClust (James et al., 2018), an unsupervised machine-learning method that aims to provide highly accurate clustering without the need for user-specified similarity parameters (these are learned).
However, this approach is intended for use with data sets containing genomes of multiple different viruses. In particular, it was validated on a data set containing 96 sequences of average length of 3K–12K, coming from nine different viruses. On the contrary, SARS-CoV-2 data sets usually contain several hundred thousand sequences of a single virus, with genome length averaging around 30K.
2.4. Monte Carlo-based entropy minimization
We use clustering entropy (Li et al., 2004) to assess the various clustering methods that we propose in this work (see Section 3.1). For this reason, we also use a technique aimed directly at minimizing clustering entropy as the objective. We first define clustering entropy in the following.
Formally, we have a set S of aligned nucleotide sequences on the set X of genomic sites. Since they are aligned, sequences can be viewed as rows of a matrix and, when restricted to a site , can be viewed as columns of this matrix. Let be the four nucleotides, not counting the gap (–) character. Using the notation of Li et al. (2004), the entropy of a subset C (a cluster) of sequences from S at site is then
| (2) |
Note that —the probability that a sequence has nucleotide a at site x—essentially amounts to the relative frequency of nucleotide in C at site x. The entropy of subset C of sequences on a subset X of sites is then
| (3) |
that is, we simply sum up the entropies at the individual sites. Since the set of sites will always correspond to the SNV (Single Nucleotide Variation) sites of our sequences, we use simply for the entropy of a subset (a cluster) of sequences from hereon. The expected entropy (Li et al., 2004) of a clustering of sequences is then
| (4) |
where , the number of elements in cluster Ci, and n is the total number of sequences. For completeness, the total entropy of a clustering is simply the sum
| (5) |
of the individual entropies of each cluster (not weighted by ni).
In Li et al. (2004), the authors prove that the entropy [Eq. (4)] is a convex function, allowing any optimization procedure to reach a global minimum. It is because of this property that we can use techniques aimed directly at minimizing clustering entropy as the objective. The Monte Carlo method is a broad class of computational algorithms that rely on repeated random sampling to optimize some criteria. In this context, we are randomly sampling clusterings of sequences to minimize [Eq. (4)]. The basic idea is that we start with some clustering—note that the clustering corresponding to placing all sequences in the same cluster has maximum entropy, by definition. The Monte Carlo process then operates according to the iteration as follows:
from the current clustering, randomly pick a sequence from some cluster and place it into another cluster, resulting in a new clustering;
compute the entropy [Eq. (4)] of the new clustering; and
accept this new clustering, if the entropy has decreased, otherwise keep the current clustering;
until convergence, that is, the clustering does not change after some number of iterations.
In Li et al. (2004), the authors prove the concept of applying the Monte Carlo method to entropy minimization by implementing a very basic procedure similar to the above, and then demonstrate it on a small data set. Since our data sets are on a much larger scale (millions of sequences on 30K genomic sites), the basic iteration that randomly samples a single sequence in each iteration would need many iterations for a very small improvement. For this reason we apply the following preprocessing step, to improve the convergence. Rather than using all (30K) columns, we first sort the columns according to their (unclustered) entropy value. We then select the n columns, or tags, with highest entropy. Next, we then run the above Monte Carlo process on the reduced data set with the n tags. This results in a clustering (of the rows), to which we then apply the original set of all columns.
2.5. Filling gaps
Finally, the set of SARS-CoV-2 sequences that we deal with contain missing nucleotides, due to gaps or deletions. This is particularly true with GISAID sequences collected from December 2019 to the end of March 2020, when sequencing, alignment, and so on were less refined. This is further complicated by the presence of deletions, which could be confused with gaps.
Here, we attempt to use the clustering obtained by some clustering method to fill the gaps. That is, rather than uniformly filling all sequences with, for example, the reference genome, we fill each sequence with the center of its cluster. The idea is that if a clustering performs well, then the sequences of a cluster should correspond to a subtype. In this case, the center—a consensus sequence of this subtype—should be much closer to any sequence of its cluster than the reference genome, resulting in a more accurate filling of the gaps.
3. Measures for Assessing Clustering Quality
In this section, we present two measures for assessing clustering quality, to compare the various clustering methods that we outlined in the previous section. The first measure is clustering entropy, an internal evaluation criterion that reflects the underlying processes that generate a set of viral sequences. The second is a measure of the selective fitness of clusters, based on how their rates of change in size vary over time.
3.1. Clustering entropy
Since we are comparing various clustering methods without knowing a ground truth, we need to consider an internal evaluation criterion. Many of the commonly used criteria require some notion of distance, or dissimilarity measure, between the objects being clustered. For example, criteria such as the Calinski–Harabasz Index (Caliński and Harabasz, 1974) or the Gap Statistic (Tibshirani et al., 2001) rely on the Euclidean distance, while the Davies–Bouldin Index (Davies and Bouldin, 1979) or the Silhouette Coefficient (Rousseeuw, 1987) require this distance (or dissimilarity) to be a metric. For the same reason that we use k-modes for clustering—sequences are objects on categorical attributes, which take values A, C, G, T (and −, a gap)—criteria based on the Euclidean distance are not appropriate. Moreover, because the various dissimilarity measures that we use within the k-modes framework for clustering are not Euclidean (Hamming distance), or even a metric [TN-93 distance, see Table 1 of Tamura and Nei (1993)], even criteria such as the Davies–Bouldin Index or Silhouette Coefficient would not apply.
The clustering entropy [Li et al. (2004); Eqs. (4) and (5)] is an internal evaluation criterion that was shown to generalize any distance-based criterion, and does not even require any notion of distance or dissimilarity. Hence, for the reasons mentioned above, the clustering entropy criterion is appropriate in our case. Moreover, clustering entropy naturally reflects the fact that the population of viral sequences comes from a number of subtypes. Clustering entropy can be formally derived using a likelihood principle based on Bernoulli mixture models. In these mixture models, the observed data are thought of as coming from a number of different latent classes. In Li et al. (2004), the authors prove that minimizing clustering entropy is equivalent to maximizing the likelihood that a set of objects are generated from a set of (k) classes. This reflects the underlying processes that generate a set of viral sequences: that they evolved from a set of (k) subtypes.
This relates closely to the widely used notion of sequence logo (Schneider and Stephens, 1990): a graphical representation of a set of aligned sequences, which conveys at each position both the relative frequency of each base (or residue) and the amount of information (the entropy) in bits. A clustering of viral sequences of low entropy then relates to a reliable set of sequence logos (in terms of information), and can hence shed light on the possible biological function of the viral subtype that each such logo (or related motif) represents.
3.2. Fitness
We use a mathematical model proposed in Skums et al. (2012) for the calculation of a numerical measure of the fitness of a quasispecies. This model is used here to calculate the fitness of a cluster, based on how the rate of change in size (number of sequences it contains) varies over time. For a set of clusters, denotes the size of cluster Ci at a particular time t. The fitness coefficient is calculated using hi, which is the cumulative sum of the Xi. It follows that is the total infected population size at time t. Each is normalized over , which is denoted by , that is,
| (6) |
Using cubic splines, and are interpolated over the time period and the derivatives and are calculated. The fitness function gi for each cluster Ci is then defined as follows:
| (7) |
The fitness coefficient ri, which is the average fitness over the time period T (composed of the times t) for cluster Ci, is then
| (8) |
To reduce sampling error, we use the Poisson distribution to draw random samples. For each cluster at time t, a sufficiently large number of random samples are drawn from the Poisson distribution on as the expectation of the interval. Then is replaced by the mean value of these random samples. This is repeated a sufficiently large number of times (e.g., 100) to calculate a set of Poisson-distributed sizes. The fitness coefficient calculation is then applied on each repetition separately and a confidence interval of this fitness coefficient is obtained.
4. Data Sets
In this section, we outline the data sets that we used in the experiments of the next section. We first give a brief overview of well-known subtypes, or variants, from the literature, and then describe the three data sets we use, which are known to contain different proportions of these variants.
4.1. Known variants
Since its emergence in November 2019 (Deasy et al., 2020), SARS-CoV-2 has evolved into different variants. Divergences in mutation at the genomic level have been observed in different regions of the world as new infectious variants are emerging. The following is a description of some of the well-known variants to date. A more complete list can be found in Table 1.
4.1.1. Alpha variant (the United Kingdom)
The Alpha variant, also known as the B.1.1.7 variant of SARS-CoV-2, was first identified in Kent, the United Kingdom, in late summer to early autumn 2020. It has the highest transmissibility of any lineage, with a 50%–100% reproduction rate (Volz et al., 2021a). The first case was reported on December 14, 2020, and this variant is now detected in over 30 countries, with more than 15 thousand people affected worldwide (Galloway et al., 2021). Of the many genomic mutations that characterize this variant, it has a 69/70 deletion and a mutation at position 501, which affects the conformation of the receptor binding domain (RBD) of the spike protein of SARS-CoV-2. It has 17 mutations that include 14 amino acids and 3 in-frame deletions at open-reading frame (ORF) 1 a/b, ORF 8, spike (S), and N gene regions. These mutations have biological implications and have resulted in diagnostic failures (Ramírez et al., 2021).
4.1.2. Beta variant (South Africa)
The first case of the Beta variant, also known as B.1.351, was identified in Nelson Mandela Bay, South Africa, in October 2020. This lineage was predominant by the end of November 2020 in the Eastern and Western Cape Provinces of South Africa. By January 2021, there were 415 known cases of infection with this variant, found in 13 different countries. This variant has eight mutations in the S gene region, including three mutations SK417N, E484K, and N501Y that affect the RBD of the spike protein. These three mutations can be the reason for increased transmissibility, and can also lead to alterations in conformation that could pose a challenge for the effectiveness of vaccines (Galloway et al., 2021; Tang et al., 2021; Zucman et al., 2021).
4.1.3. Gamma and Zeta variants (Brazil)
The Gamma variant, also known as P.1(B.1.1.28.1), was initially identified in February 2020, in Japanese travelers coming from Amazonas State, Brazil. It was first reported in a 29-year-old female resident of Amazonas State. The P.1 lineage has mutations K417T, E484K, and N501Y in the S gene region, which affect the RBD of the spike protein. The Zeta variant, also known as P.2(B.1.1.28.2), was first identified in Rio de Janeiro, Brazil. It shares the mutation E484K with the Gamma variant (Naveca et al., 2021a).
4.1.4. Epsilon variants (California, USA)
In July 2020, the first case of the Epsilon variants, also known as the CAL.20C or B.1.427/B.1.429 variants of SARS-CoV-2, was identified in Los Angeles County, California, USA. The Cedars-Sinai Medical Center reported that the second B.1.429 Epsilon variant contains five mutations at ORF 1 a (I4205V), ORF 1 b (D1183Y), and S gene mutations S13I, W152C, and L452R. Mutation L452R is correlated with higher infectivity (Zhang et al., 2021). The Epsilon variants are spreading in the United States and in 29 other countries (McCallum et al., 2021).
4.1.5. Iota variant (New York, USA)
The Iota variant, also known as B.1.526, was first found in November 2020 in New York, United States. At that time, the number of sequences of the Iota variant comprised less than 1% of all sequences in the GISAID database (Elbe and Buckland-Merrett, 2017). Scientists from Caltech noticed a surge in the growth of this number by roughly one-third by February 2021. This variant has mutations L5F, T95I, D253G, E484K or S477N, D614G, and A701V in the S gene region—mutations E484K and S477N affecting the RBD of the spike protein. Note that the E484K mutation causes attenuation in in vitro neutralization, and is found in other VoCs (Thompson et al., 2021; West et al., 2021), such as the Beta, Gamma, and Zeta variants, described above.
4.2. Data sets used
In our experiments, we use four different data sets, three of which are various snapshots of the GISAID (Elbe and Buckland-Merrett, 2017) database at different time points, and the fourth is a data set obtained from the EMBL-EBI (EMBL-EBI, 2020) database in the United Kingdom. These data sets are summarized in Table 2, and then each one is explained in more detail in its corresponding subsection below.
4.2.1. GISAID 1
The first data set consists of sequences submitted to the GISAID (Elbe and Buckland-Merrett, 2017) database up until November 2020. This data set contains sequences from all over the world. Since this data set covers the period of time from December 2019 to March 2020, some of these sequences have a sizeable number of gaps.
4.2.2. GISAID 1A
The second data set consists of sequences submitted to GISAID up until the beginning of March 2020. This smaller data set, a subset of GISAID 1, was designed to test out the Monte Carlo optimization procedure described in Section 2.4.
4.2.3. The United Kingdom
The third data set consists of sequences submitted to the EMBL-EBI (EMBL-EBI, 2020) database from the end of January 2020 to the end of December 2020. Since this database is in the United Kingdom, and given the collection period, this data set contains a sizeable portion of the Alpha variant.
4.2.4. GISAID 2
The third data set consists of all sequences submitted to GISAID up until April 2021. Since many of the known variants mentioned above have been well-documented by April 2021, this data set contains a sizable portion of sequences annotated as being from the Alpha, Beta, Gamma, Epsilon, and Zeta variants. Such labels correspond to “ground truth clusters” for which we can compute the precision, specificity, F1 score, and so on of a clustering obtained with a given method.
5. Experimental Results
In this section, we report the results of our approaches of clustering and gap filling using the four data sets mentioned in Section 4.2 above. For all data sets, we align the sequences and trim the first and last 50 bp of the aligned sequences. We use default parameters for running CliqueSNV to find initial cluster centers, in all cases setting the minimum cluster frequency to be at least 1% of the population. We refer to the approach of using CliqueSNV to find the initial centers, followed by clustering with k-modes as our CliqueSNV-based approach (setting 3 of Section 2.2).
These experiments, and their results, are grouped as follows: a general comparison of clustering and gap filling approaches, using the GISAID 1 data set (Section 5.1); a test of our Monte Carlo-based entropy minimization procedure introduced in Section 2.4, using the GISAID 1A data set (Section 5.2); and a demonstration of the use of various clustering methods for finding subtypes, using the U.K. and GISAID 2 data sets (Section 5.3).
5.1. Comparison of clustering approaches
Using the GISAID 1 data set, our CliqueSNV-based approach identified at most 66 subtypes (smallest k that achieves minimum cluster frequency ), which vary in proportion between December 2019 and November 2020. We report the relative distributions over time of these different subtypes in Figures 1 and 2, in a similar way to that of Figure 3 of du Plessis et al. (2021).
FIG. 1.
Subtype distribution (GISAID data set, 15-day window, relative count).
FIG. 2.
Subtype distribution (GISAID data set, cumulative, relative count).
FIG. 3.
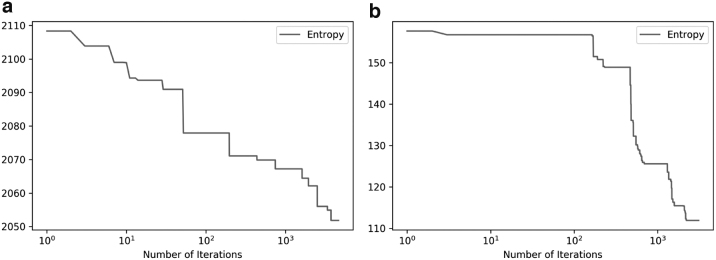
The entropy descent of our Monte Carlo method applied to the initial clustering obtained by CliqueSNV-based clustering of the GISAID 1A data set after having preprocessed to (a) tags and (b) tags. Note that in the latter table, the entropy is in terms of just the 1000 tags—the optimal clustering in terms of these 1000 tags then applied to the original set of all columns, for the final entropy 945.9 seen in Table 6. Note that a threshold of (see Section 2.4) was used in both cases.
Table 3 gives an assessment of all clusterings (and gap fillings) of the GISAID 1 data set that were computed for the various settings mentioned in Section 2.2, in terms of both the expected entropy [Eq. (4)] and total entropy [Eq. (5)]. While any form of clustering achieves a better expected (and total) entropy than not clustering at all, our CliqueSNV-based approach tends to outperform all other forms of clustering using either Hamming or TN-93 distance. This demonstrates that CliqueSNV finds meaningful centers in these interhost viral data. Based on these results, from hereon we use only the Hamming distance setting of our CliqueSNV-based clustering (setting 3a of Section 2.2, second-last line of Table 3), and the random centers' initialization and Hamming distance setting (setting 1a of Section 2.2, second line of Table 3) of k-modes, unless otherwise indicated. Finally, by filling gaps in sequences based on the center of its cluster, we achieve an even lower expected (and total) entropy. This highlights the value of a cluster-based approach for filling gaps. For example, the entropy of the data set without clustering remained high even after filling gaps, which would, by definition, be based on the center for the entire data set, which is effectively the reference genome.
Table 3.
Entropy of All Clusterings of GISAID 1 Data Set Computed
| |
Without gap filling |
With gap filling |
||
|---|---|---|---|---|
| k-modes setting (initialization + distance) | Expected entropy | Total entropy | Expected entropy | Total entropy |
| Without clustering | 9536.89 | 9536.89 | 8417.89 | 8417.89 |
| Random centers + hamming | 123.00 | 3170.60 | 109.21 | 2474.30 |
| Random centers + TN-93 | 127.32 | 4401.18 | 111.05 | 3470.03 |
| Pairwise distant + hamming | 422.65 | 4651.23 | 294.98 | 3629.47 |
| Pairwise distant + TN-93 | 273.34 | 3500.14 | 256.44 | 3007.07 |
| CliqueSNV + hamming | 110.58 | 2585.29 | 90.42 | 2308.95 |
| CliqueSNV + TN-93 | 121.87 | 2379.46 | 100.85 | 2117.40 |
The expected entropy [Eq. (4)] and total entropy [Eq. (5)] of the sequences of the GISAID 1 data set without clustering (i.e., considered a single cluster containing all sequences), and when clustering using each of the six combinations of settings mentioned in Section 2.2, both without filling gaps and with gap filling.
Table 4 reports the runtimes of the various stages of our CliqueSNV-based clustering approach, and Table 5 compares the overall runtimes of CliqueSNV-based clustering and the k-modes approach. We note, given the latter table, that the CliqueSNV-based approach had a slightly lower runtime than the k-modes approach, despite it performing best overall.
Table 4.
Runtime of Each Stage of the CliqueSNV-Based Approach
| Stage | Time (seconds) |
|---|---|
| CliqueSNV (finding initial centers) | 2405.08 |
| Clustering (with k-modes) | 2324.34 |
| Gap filling | 2740.32 |
| Entropy computation | 1254.22 |
| Total | 8723.96 |
Runtime of each of the different stages of the CliqueSNV-based approach on the GISAID 1 data set. All stages were executed on a PC with an Intel(R) Xeon(R) CPU X5550 2.67GHz x2 with 8 cores per CPU, DIMM DDR3 1333 MHz RAM 4Gb x12, and running the CentOS 6.4 operating system.
Table 5.
Runtimes of CliqueSNV-Based and k-Modes Clustering
| Clustering method | Time (seconds) |
|---|---|
| CliqueSNV-based | 4729.42 |
| k-modes | 4922.44 |
Runtimes of CliqueSNV-based and k-modes (random centers + Hamming) for the GISAID 1 data set. Both methods were executed on a PC with an Intel(R) Xeon(R) CPU X5550 2.67GHz x2 with 8 cores per CPU, DIMM DDR3 1333 MHz RAM 4Gb x12, and running the CentOS 6.4 operating system.
5.2. Entropy minimization
The main goal of entropy minimization is to make further gains on the performance of existing clustering techniques. Hence, we apply our Monte Carlo-based procedure described in Section 2.4 to the clustering obtained by our CliqueSNV-based method (the most performant method), which identified 28 subtypes in the GISAID 1A data set. As a baseline for comparison, we also produce a random clustering of the data into 28 clusters. Table 6 reports results of our Monte Carlo-based entropy minimization procedure on these two initial clusterings when preprocessing to various different numbers n of tags. Initial clustering with our CliqueSNV-based method followed by our Monte Carlo procedure with 1000 tags achieved the largest decrease in entropy, from 2093.6 to 945.9, as well as the best overall final clustered entropy. The results for CliqueSNV-based clustering indicate that a local (probably global) minimum sits somewhere between 1500 and 500, in terms of the optimal number n of tags to select to achieve the best results. While the results for random clustering were considerably worse, there seems to be a trend toward better entropy with reduced numbers of tags. Finally, Figure 3a and b depicts the entropy descent of our Monte Carlo method applied to the initial CliqueSNV-based clustering for 28,000 and 1000 tags, respectively. The latter shows more (relative) improvement in the entropy, indicating that selecting a subset of tags can allow the Monte Carlo iteration to approach closer to the optimum entropy with fewer iterations.
Table 6.
Entropy Minimization with the GISAID 1A Data Set
| Initial | Tags | Iterations | Initial entropy | Final entropy |
|---|---|---|---|---|
| CliqueSNV based | 28,000 | 4625 | 2051.8 | |
| 1500 | 2650 | 1166.3 | ||
| 1000 | 3100 | 2093.6 | 945.9 | |
| 750 | 3300 | 971.8 | ||
| 500 | 8100 | 1117.3 | ||
| Random | 28,000 | 1100 | 4458.7 | |
| 1000 | 1975 | 4479.1 | 4324.5 | |
| 750 | 6175 | 4115.2 | ||
| 500 | 4600 | 4098.7 |
Monte Carlo-based entropy minimization applied to the GISAID 1A data set for various initial clusterings and numbers of tags. The five columns, starting from the left, are: Initial clustering (to clusters); Number n of tags selected in the preprocessing; Cumulative number of iterations of the Monte Carlo procedure; Initial clustering entropy; Final clustering entropy after the number of iterations reported in column 3. Note that a threshold of (see Section 2.4) was used in all cases.
5.3. Finding subtypes
One of the important goals of clustering in this context is to identify subtypes, for example, VoCs. Here we demonstrate the ability of our clustering approaches to finding subtypes in the U.K. data set, and then in the much larger GISAID 2 data set.
5.3.1. The U.K. data set
Using the U.K. data set, our CliqueSNV-based approach identified 15 subtypes. Since the data here are over a shorter time span (are smaller) and more uniform, a k of 15 was sufficient for the minimum cluster frequency to be at least 1% of the population. On the contrary, MeShClust (James et al., 2018) was only able to find 3 clusters in this data set. Table 7 reports the F1 score of the methods we compared. Our CliqueSNV-based approach outperformed other methods by a large margin in producing a clustering with all sequences of the Alpha variant residing in a single cluster, while 1.30% of the sequences in this cluster do not belong to the Alpha variant. In the clustering produced by the k-modes approach, the sequences of the Alpha variant were spread over five clusters, while one cluster contained 97.45% of the Alpha variant sequences. However, 86.54% of the sequences of this cluster did not belong to the Alpha lineage. MeShClust, on the contrary, produced a clustering with all Alpha variant sequences residing in a single cluster, where 90.68% of the sequences in this cluster do not belong to the Alpha variant.
Table 7.
F1 Score of Various Clustering Approaches on the U.K. Data Set
| Method | F1 score | F1 largest cluster |
|---|---|---|
| CliqueSNV based | 0.99 | 0.99 |
| k-modes | 0.003 | 0.24 |
| MeShClust | 0.11 | 0.11 |
The F1 score of the clustering produced by a method (column 1) with respect to the sequences of the Alpha variant (column 2), and of the cluster containing the largest number of sequences with the Alpha variant (column 3).
We report the relative distributions of these different subtypes in Figure 4 over the period of time between the beginning of October 2020, when the first case of the Alpha variant was reported in the United Kingdom, and the middle of December, when this variant comprised more than one-third of all sequences. We report a weekly moving average because a weekly oscillation in the SARS-CoV-2 data has been noted in Bukhari et al. (2020). One will notice, in Figure 4, the sharp increase of the relative proportion of a certain subtype (in the bottom right corner) to more than a third of the population. We confirm from metadata that this corresponds to the Alpha variant that was first identified in studies such as Volz et al. (2021b).
FIG. 4.
Subtype distribution (the U.K. data set, weekly window, relative count) produced our CliqueSNV-based clustering method. The subtype in the bottom right corner contributes to sequences that correspond to the Alpha variant.
When restricting the clusters returned by our CliqueSNV-based approach to the final 1-week interval of Figure 4, leading up to mid-December 2020, all Alpha variant sequences appear in a single cluster (among a total of 15). In the clusters returned by the k-modes approach, on the contrary, sequences of the Alpha variant are spread over 13 clusters, with counts ranging from 1 to 6327 Alpha variant sequences per cluster. MeShClust again produced only 3 clusters, with a single cluster containing all Alpha variant sequences, when restricted to this final interval, while 90.86% of the sequences in this cluster did not belong to the Alpha variant. The expected entropy of our CliqueSNV-based approach and the k-modes approach was 75.73 and 94.16, respectively, while the total entropy was 986.48 and 2074.12, respectively. This illustrates the ability of our clustering to identify subtypes that are known in the literature. Interestingly enough, the study of Volz et al. (2021b) is based on an approach of building a phylogenetic tree. This demonstrates our approach, which is based on clustering sequences, as a viable alternative.
Our CliqueSNV-based clustering method was able to detect one subtype that tends to dominate the population in this U.K. data set, in attaining good entropy and F1 scores. However, we wanted to further validate if this is consistent with other independent measures of quality, such as the cluster-based fitness coefficient that we detail in Section 3.2. To compute this, we chose our time points t to be intervals of 1 week over the period from the beginning of October to the middle of December, exactly as in Figure 4. The size of each cluster Ci (of clusters) for every week t was obtained, and each fitness coefficient ri was computed according to Eq. (8). To reduce sampling error, we drew 2000 random samples from the Poisson distribution on according to Section 3.2. We repeated this 100 times, and we report in Table 8 the 95% confidence interval of the top five clusters, sorted by interval lower bound. We note that similar results are obtained with either Hamming or TN-93 distance, with TN-93 distance corresponding to slightly higher fitness coefficients. We confirm that in either case, the mostly highly ranked cluster in terms of fitness (with cluster ID 6) corresponds to the cluster containing all of the sequences pertaining to the Alpha variant from above. This highlights the ability of our clustering-based approach for detecting, based purely on sequence content, novel subtypes that have the potential of becoming dominant in the population.
Table 8.
Fitness Coefficients of the Clusters of the U.K. Data Set
| Distance | Rank | Cluster ID | Interval lower B | Interval upper B |
|---|---|---|---|---|
| Hamming | 1 | 6 | 1.343 | 1.504 |
| 2 | 3 | 0.354 | 0.369 | |
| 3 | 14 | 0.284 | 0.324 | |
| 4 | 8 | 0.08691 | 0.0881 | |
| 5 | 2 | 0.08690 | 0.0878 | |
| TN-93 | 1 | 6 | 1.390 | 1.510 |
| 2 | 2 | 0.789 | 0.795 | |
| 3 | 3 | 0.351 | 0.364 | |
| 4 | 14 | 0.353 | 0.390 | |
| 5 | 8 | 0.086 | 0.0869 |
The 95% confidence interval of the top five fitness coefficients, according to the interval lower bound, of the 15 clusters of the U.K. data set obtained using our CliqueSNV-based clustering method with Hamming distance and TN-93 distance, respectively.
5.3.2. The GISAID 2 data set
Since our CliqueSNV-based clustering approach was able to clearly pinpoint the Alpha variant within the U.K. data set, we tested it also on the GISAID 2 data set, which contains many of the variants listed in Table 1. CliqueSNV-based clustering identified 36 subtypes in this data set. We first computed fitness coefficients ri [Eq. (8)] for these 36 clusters using 1-week time intervals t. Table 9 reports the 95% confidence interval due to subsampling (see Section 3.2) of the top and bottom five clusters, sorted by interval lower bound. One will notice immediately that the fitness coefficient is much more evenly distributed across the clusters of this data set, compared with the U.K. data set (Table 8).
Table 9.
Fitness Coefficients of the Clusters of the GISAID 2 Data Set
| Rank | Cluster ID | Interval lower B | Interval upper B |
|---|---|---|---|
| 1 | 1 | 0.0601 | 0.0602 |
| 2 | 17 | 0.0486 | 0.0489 |
| 3 | 21 | 0.0463 | 0.0463 |
| 4 | 20 | 0.0456 | 0.0457 |
| 5 | 35 | 0.0440 | 0.0440 |
| 32 | 4 | 0.0143 | 0.0143 |
| 33 | 29 | 0.0138 | 0.0138 |
| 34 | 28 | 0.0120 | 0.0120 |
| 35 | 32 | 0.0118 | 0.0118 |
| 36 | 34 | 0.0110 | 0.0110 |
The 95% confidence interval of the top and bottom five fitness coefficients, according to the interval lower bound, of the 36 clusters of the GISAID 2 data set obtained using our CliqueSNV-based clustering method. The mean () standard deviation () of the interval lower and upper bounds is and , respectively.
Table 10 reports some the variants found by our CliqueSNV-based approach in terms of specificity, F1 score, and fitness rank (Table 9). Notice that specificity/F1 score generally decrease with rank and cluster size, as would be expected. Exceptions to this trend are the Gamma/Zeta variant in F1 score versus rank (having a high F1 score for its rank) and the Epsilon variant (having a large cluster size for its F1 score and rank). Finally, since the GISAID 2 data set contains more than 1 million sequences, the Gamma/Zeta, Beta, and Epsilon variants comprise less than 1% of the sequences, yet our CliqueSNV based was still able to identify them with specificities around 50% and F1 scores . This demonstrates the ability of our clustering approach to detect rare subtypes in very large sets of sequences.
Table 10.
Variants Found in the GISAID 2 Data Set Using CliqueSNV-Based Clustering
| Variant | ID | Specificity, % | F1 | Rank | Size |
|---|---|---|---|---|---|
| Alpha (the United Kingdom) | 1 | 93.16 | 0.96 | 1 | 265,255 |
| Gamma and Zeta (Brazil) | 25 | 51.21 | 0.68 | 7 | 1892 |
| Beta (South Africa) | 21 | 45.85 | 0.62 | 3 | 2754 |
| Epsilon (California) | 13 | 41.08 | 0.58 | 13 | 9251 |
Specificity, F1 score, and fitness rank (Table 9) of the cluster containing the largest number of sequences of the corresponding variant.
6. Conclusions
In this work, we successfully adapted a method CliqueSNV (Knyazev et al., 2020), originally designed for discovering viral haplotypes in an intrahost population, to finding subtypes of SARS-CoV-2 in the (massively interhost) global population. We use clustering entropy (Li et al., 2004) to assess the quality of a clustering—a notion that naturally reflects the underlying processes from which a set of viral subtypes arise. We introduce two additional techniques that boost the entropy even further, namely, gap filling and Monte Carlo entropy minimization. The former is useful for sequences collected before March 2020 when collection and sequencing were not yet refined, while the latter is possible because clustering entropy is convex (Li et al., 2004), allowing optimization techniques aimed directly at minimizing entropy as the objective. We show that our CliqueSNV-based clustering method outperforms other techniques in terms of low entropy, and the further improvements in entropy that can be obtained with gap filling and Monte Carlo minimization.
We then turned to data sets obtained from the GISAID and EMBL-EBI (the United Kingdom) databases to identify viral subtypes. Our method was able to most clearly identify the Alpha variant in the U.K. data set, with a single cluster containing all sequences with a specificity . These results tended to be in agreement with the entropies obtained, as well as with the measure of selective fitness introduced in Section 3.2. In the GISAID data set, which contains over one million sequences, our CliqueSNV-based method was able to clearly identify the Alpha variant, and also the lesser represented Beta (South Africa), Epsilon (California), and Gamma and Zeta (Brazil) variants. What is interesting about this is that these lesser represented variants comprise a few thousand sequences each ( of the sequences), and yet our method was able to cluster them with specificities around 50%, corroborating again with the fitness coefficient. This demonstrates the approach of clustering as a viable and scalable alternative for detecting even the rarest subtypes at an early stage of development.
An immediate future work is a more full exploration of how our Monte Carlo entropy minimization approach can be made faster and more scalable to large data sets. Ideas include parallelization of our current approach, the design of data structures that can be more efficiently updated, or heuristics beyond our use of tags. The use of optimization techniques other than the Monte Carlo method is a possibility as well. Since CliqueSNV (Knyazev et al., 2020) is a relatively new technique, possible advancements in its ability to better detect viral haplotypes within an intrahost population would likely carry over to improvements to finding subtypes in the interhost population setting of this work. Finally, while we provide a viable alternative to building phylogenetic trees [e.g., du Plessis et al. (2021) for detecting subtypes, it would be interesting to explore how these could be combined [as in e.g., Ciccolella et al. (2021a)].
Author Disclosure Statement
The authors declare they have no conflicting financial interests.
Funding Information
A.M., S.K., B.S., R.H., and A.Z. were partially supported by the National Science Foundation (NSF; Grant No. 1564899) and the National Institutes of Health (NIH; Grant No. 1R01EB025022-01). F.M. and P.S. were partially supported by the NIH (Grant No. 1R01EB025022-01) and NSF (Grant No. 2047828). A.M., B.S., and S.K. were partially supported by the GSU Molecular Basis of Disease Fellowship. M.P. was supported by a Georgia State University/Computer Science startup grant.
References
- Ahn, S., and Vikalo, H.. 2018. abayesqr: A bayesian method for reconstruction of viral populations characterized by low diversity. J. Comput. Biol. 25, 637–648. [DOI] [PubMed] [Google Scholar]
- Anderberg, M.R. 1973. Cluster Analysis for Applications. Academic Press, New York, NY. [Google Scholar]
- Baaijens, J., Aabidine, A., Rivals, E., et al. 2017. De novo assembly of viral quasispecies using overlap graphs. Gen. Res. 27, 835–848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukhari, Q., Jameel, Y., Massaro, J.M., et al. 2020. Periodic oscillations in daily reported infections and deaths for coronavirus disease 2019. JAMA Netw. Open. 3, e2017521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caliński, T., and Harabasz, J.. 1974. A dendrite method for cluster analysis. Commun. Stat. 3, 1–27. [Google Scholar]
- Ciccolella, S., Patterson, M., Bonizzoni, P., et al. 2021. a. Effective clustering for single cell sequencing cancer data. IEEE J. Biomed. Health Inform. 2021, 34003758, 1–11. [DOI] [PubMed] [Google Scholar]
- Ciccolella, S., Ricketts, C., Soto Gomez, M., et al. 2021. b. Inferring cancer progression from single-cell sequencing while allowing mutation losses. Bioinformatics 37, 326–333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciccolella, S., Soto Gomez, M., Patterson, M., et al. 2020. gpps: An ilp-based approach for inferring cancer progression with mutation losses from single cell data. BMC Bioinform. 21, 1–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies, D.L., and Bouldin, D.W.. 1979. A cluster separation measure. IEEE Trans. Pattern Analysis Mach. Intell. PAMI-1, 224–227. [PubMed] [Google Scholar]
- Deasy, J., Rocheteau, E., Kohler, K., et al. 2020. Forecasting ultra-early intensive care strain from COVID-19 in England. medRxiv.
- du Plessis, L., McCrone, J.T., Zarebski, A.E., et al. 2021. Establishment and lineage dynamics of the sars-cov-2 epidemic in the UK. Science 371, 708–712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elbe, S., and Buckland-Merrett, G.. 2017. Data, disease and diplomacy: GISAID's innovative contribution to global health. Global Challenges 1, 33–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- EMBL-EBI. 2020. Wellcome Genome Campus, Hinxton, Cambridgeshire, CB10 1SD, United Kingdom.
- Galloway, S.E., Paul, P., MacCannell, D.R., et al. 2021. Emergence of SARS-CoV-2 b. 1.1. 7 lineage—United States, December 29, 2020–January 12, 2021. Morb. Mortal. Wkly. Rep. 70, 95–99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hadfield, J., Megill, C., Bell, S., et al. 2018. Nextstrain: Real-time tracking of pathogen evolution. Bioinformatics 34, 4121–4123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang, Z. 1997. A fast clustering algorithm to cluster very large categorical data sets in data mining. 3, 34–39.
- Huang, Z. 1998. Extensions to the k-means algorithm for clustering large data sets with categorical values. Data Mining Knowl. Discov. 2, 283–304. [Google Scholar]
- Jahn, K., Kuipers, J., and Beerenwinkel, N.. 2016. Tree inference for single-cell data. Gen. Biol. 17, 1–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- James, B., Luczak, B., and Girgis, H.. 2018. Meshclust: An intelligent tool for clustering DNA sequences. Nucleic Acids Res. 46, e83–e83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knyazev, S., Hughes, L., Skums, P., et al. 2021. Epidemiological data analysis of viral quasispecies in the next-generation sequencing era. Brief. Bioinform. 22, 96–108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knyazev, S., Tsyvina, V., Shankar, A., et al. 2020. Accurate assembly of minority viral haplotypes from next-generation sequencing through efficient noise reduction. Nucleic Acids Res. 49, 13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li, T., Ma, S., and Ogihara, M.. 2004. Entropy-based criterion in categorical clustering. In Proceeding of the 21st International Conference on Machine Learning (ICML), 3, 536–543. [Google Scholar]
- MacQueen, J., Le Cam, L.M., Neyman, J.. 1967. Some methods for classification and analysis of multivariate observations. In Proceedings of the Fifth Berkeley Symposium on Mathematical Statistics and Probability. 1, 281–297. [Google Scholar]
- McCallum, M., Bassi, J., Marco, A., et al. 2021. SARS-CoV-2 immune evasion by variant B.1.427/B.1.429. bioRxiv. Science 373, 648–654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naveca, F., da Costa, C., Nascimento, V., et al. 2021. a. SARS-CoV-2 reinfection by the new variant of concern (VoC) p. 1 in Amazonas, Brazil. Virological.org. Last viewed July 20, 2021.
- Naveca, F., Nascimento, V., Souza, V., et al. 2021. b. Phylogenetic relationship of sars-cov-2 sequences from amazonas with emerging brazilian variants harboring mutations e484k and n501y in the spike protein. Virological.org. Last viewed July 20, 2021.
- Prabhakaran, S., Rey, M., Zagordi, O., et al. 2014. HIV haplotype inference using a propagating dirichlet process mixture model. IEEE/ACM Trans. Comput. Biol. Bioinform. 11, 182–191. [DOI] [PubMed] [Google Scholar]
- Rambaut, A., Holmes, E.C., O'Toole, A., et al. 2020. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 5, 1403–1407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramírez, J.D., Muñoz, M., Patiño, L.H., et al. 2021. Will the emergent SARS-CoV2 B. 1.1. 7 lineage affect molecular diagnosis of COVID-19? J. Med. Virol. 93, 2566–2568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rousseeuw, P.J. 1987. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65. [Google Scholar]
- Schneider, T.D., and Stephens, R.M.. 1990. Sequence logos: A new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skums, P., Campo, D., Dimitrova, Z., et al. 2012. Numerical detection, measuring and analysis of differential interferon resistance for individual HCV intra-host variants and its influence on the therapy response. Silico Biol. 11, 263–269. [DOI] [PubMed] [Google Scholar]
- Skums, P., Kirpich, A., Icer Baykal, P., et al. 2020. Global transmission network of SARS-CoV-2: From outbreak to pandemic. medRxiv. 2020, 20041145, 1–28.
- Tamura, K., and Nei, M.. 1993. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10, 512–526. [DOI] [PubMed] [Google Scholar]
- Tang, J., Toovey, O., Harvey, K., et al. 2021. Introduction of the South African SARS-CoV-2 variant 501y.v2 into the UK. J. Infect. 82, e8–e10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson, C., Hughes, S., Ngai, S., et al. 2021. Rapid emergence and epidemiologic characteristics of the SARS-CoV-2 B.1.526 variant—New York City, New York, January 1–April 5, 2021. Morb. Mortal. Wkly. Rep. 70, 712–716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tibshirani, R., Walther, G., and Hastie, T.. 2001. Estimating the number of clusters in a data set via the gap statistic. J. R. Stat. Soc. 63, 411–423. [Google Scholar]
- Volz, E., Mishra, S., Chand, M., et al. 2021. a. Assessing transmissibility of SARS-CoV-2 lineage B. 1.1. 7 in England. Nature 593, 266–269. [DOI] [PubMed] [Google Scholar]
- Volz, E., Mishra, S., Chand, M., et al. 2021. b. Transmission of SARS-CoV-2 lineage B. 1.1. 7 in England: Insights from linking epidemiological and genetic data, 2020–12. medRxiv. Nature 593, 266–269. [DOI] [PubMed]
- Vrbik, I., Stephens, D.A., Roger, M., et al. 2015. The gap procedure: For the identification of phylogenetic clusters in HIV-1 sequence data. BMC Bioinform. 16, 1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West, A., Wertheim, J., Wang, J., et al. 2021. Detection and characterization of the SARS-CoV-2 lineage B.1.526. Nat. Commun. 12, 4886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- WHO. 2021. Update. World Health Organization (July 20, 2021). A report on the COVID-19 pandemic. [Google Scholar]
- Wu, F., Zhao, S., Yu, B., et al. 2020. A new coronavirus associated with human respiratory disease in china. Nature 579, 265–269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yadav, P., Sapkal, G., Abraham, P., et al. 2021. Neutralization of variant under investigation B.1.617 with sera of BBv152 vaccinees. Clin. Infect. Dis. ciab411. [DOI] [PubMed] [Google Scholar]
- Zhang, W., Davis, B.D., Chen, S.S., et al. 2021. Emergence of a novel SARS-CoV-2 variant in Southern California. JAMA 325, 1324–1326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou, P., Yang, X.L., Wang, X.G., et al. 2020. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zucman, N., Uhel, F., Descamps, D., et al. 2021. Severe reinfection with South African SARS-CoV-2 variant 501y.v2: A case report. Clin. Infect. Dis. ciab129. [DOI] [PMC free article] [PubMed] [Google Scholar]