Abstract
Epidemiologic research requires identification of Chlamydia trachomatis serovars and detection of mixed infection. Antibody-based serotyping is unworkable when specimens are urine or vaginal swabs. We developed a reverse dot blot (RDB) to screen for multiple serotypes in these specimens. RDB yielded the predicted results on all artificially mixed samples and on seven of eight clinically mixed samples.
There are ∼17 serovars of Chlamydia trachomatis which cause either ocular or sexually transmitted infection (25–27). The immunodominant major outer membrane protein (MOMP), encoded by the omp1 gene (1, 5–7, 15, 25, 26, 29, 30), is the most variable genetic marker known for chlamydiae, making it a useful epidemiologic tool.
Serotyping of chlamydia is unnecessary for clinical diagnosis. However, unequivocal strain identification for clinical samples is required in epidemiologic research. While most specimens from chlamydia-infected individuals contain only one serovar, 2 to 15% of infections contain two or more (2–4, 8, 16, 28).
Antibody-based procedures such as fluorescent antibody (FA) staining, enzyme immunoassay, or radioimmunoassay are commonly used for both serotyping and detection of multiple serotypes but require cell culture of chlamydiae. DNA sequencing does not require cell culture, provides serovar identification, and has detected multiple serovars (3, 4, 8). However, sequencing of a mixed sample yields ambiguous results at best because the serovars in the mixture cannot be resolved unless the omp1 PCR products are cloned and multiple clones are then sequenced.
We are studying chlamydia transmission between sexual contacts and reinfection patterns in a large group of adolescent women where unambiguous identification of serotypes and detection of mixed infection are essential. We use PCR tests to detect chlamydiae from self-administered vaginal swabs and/or urine, and we use sequencing to determine the serovar. Since sequencing does not reliably detect mixed infections, we developed a reverse dot blot (RDB) procedure as a screen. We first PCR amplify the omp1 gene and then hybridize labeled amplicons to serotype-specific omp1 oligonucleotides. RDB can detect multiple serovars in a specimen and does not require chlamydia culture or cloning.
Chlamydial elementary bodies (EB) were prepared (17) from strains A/571-B/OT, B/TW-5/OT, Ba/Ap-2/OT, C/TW- 3/OT, D/UW-3/Cx, E/UW-5/Cx, F/UW-6/Ur, G/UW-57/Cx, H/UW-4/Cx, I/UW-12/Ur, J/UW-36/Cx, and K/UW-53/Cx. Eight archived clinical specimens were used: 52, 61A, 154, 229, 814, 831, 900, and 910 (see Table 2).
TABLE 2.
FA results with ratio of IFU, DNA sequencing results, and RDB results for mixed-serovar specimens
| Specimen | Date of isolation (mo/yr) | Serovars (ratio) determined by FA staininga | DNA sequencing result | RDB result |
|---|---|---|---|---|
| 52 | 4/92 | E | E | E |
| 61A | 5/92 | E + F (1:6) | Mixture | E + F |
| 154 | 8/93 | E + J (1:5) | Mixture | E + J |
| 229 | 2/94 | D +F (ratio not available) | Mixture | D + F |
| 814 | 1/92 | D + F (1:1) | F | D + F |
| 831 | 2/92 | D + E (2:1) | D | D + E |
| 900 | 6/92 | E + I (2:1) | E | E + Ia |
| 910 | 7/92 | E + J (1:10) | Ja | E + Ja + Ia |
Specimen 52 was originally typed in 1996 as a mixture of E and D, although inclusion counts were not obtained to determine the ratio of the two serovars in the mixture. The isolate went through three passages in culture between 1996 and 2001. One passage was used to provide materials for the RDB, and one was used to provide materials for the retyping in 2001. When specimen 52 was retyped for this study, only serovar E was found.
DNA was extracted and omp1 amplified by PCR from clinical and laboratory samples as previously described (23, 24). For laboratory-created mixtures, DNAs (1 to 100 ng) from two different serovars were mixed together in the PCR at ratios ranging from 1:1 to 1:100. PCR products were purified using the QIAquick PCR purification kit (QIAgen), quantified, and labeled with digoxigenin (DIG) using the DIG Chem-link kit (Roche Molecular Biochemicals, Indianapolis, Ind.) according to instructions. The PCR probes were either used immediately or stored at −20°C.
Twelve oligonucleotides were designed based on published omp1 sequences (1, 9–12, 19–23, 31) (Amitof Biotech, Alston, Mass.) and made to hybridized specifically to serovars A to K (Table 1). The C-type oligonucleotide based on the published C/TW-3/OT sequence (22) failed to hybridize and was redesigned based on new sequencing of C/TW-3/OT (GenBank accession no. AF352789). A positive-control oligonucleotide was designed to hybridize with all serovars (Table 1). Poly(dT) tails were added to the 3′ ends of oligonucleotides via a terminal transferase reaction to facilitate binding to membranes. Tailed oligonucleotides were either used immediately or stored at −20°C.
TABLE 1.
Serotype-specific omp1 oligonucleotides used in the RDB
| Oligonucleotide | Location (nt, region)a | Sequence (5′ → 3′) | Temp (°C) |
|---|---|---|---|
| MOMP | 1059, CS5 | GATGAAATCTAGAAAATCTTGCGGTATTGC | 60.6 |
| A-type | 496, VS2 | CAATCTTCTGGCTTTGATACAGCGAAT | 59.9 |
| B-type | 930, VS4 | AGCCGAGACTATCTTTGATCTTACCACTC | 59.5 |
| C-type | 988, VS4 | GGAAGTGTGGTCTCTGCCGGAA | 59.9 |
| D-type | 263, VS1 | CTACAACTGATACAGGCAATAGTGCAGCT | 59.4 |
| E-type | 550, CS3 | GAACTTTACACAGATACTGCCTTCTCTTGG | 59.3 |
| F-type | 497, VS2 | CCACGAAACCTGCTGCAGATAGTATTC | 60.2 |
| G-type | 986, VS4 | GTGTAGTCGCAGCTAACTCGGAAGG | 59.4 |
| H-type | 262, VS1 | CCTACTACCAACGATGCAGCAGACTT | 58.9 |
| I-type | 494, VS2 | CACAATCTTCTAACTTTAATACAGCGAAGCTT | 59.4 |
| J-type | 494, VS2 | CACAAGCTTCTAGCTTTAATACAGCGAATC | 59.5 |
| Ja-type | 262, VS1 | CCTACTGTCAGCGATGTAGCAGGCT | 59.9 |
| K-type | 531, VS2 | TAACACTGCTTTGGATCGAGCTGTG | 59.4 |
The nucleotide (nt) location given for each oligonucleotide is specific for that particular omp1 gene, e.g., the A-type oligonucleotide hybridizes at nucleotide 496 in the ompIA gene. The position of the MOMP, probe (positive control) is based on the omp1A sequence. CS, conserved region of MOMP; VS, variable region of MOMP.
All protocols for hybridization and detection are found in the DIG System User's Guide for Filter Hybridization (Roche). Positively charged nylon membranes (Roche) were cut into 2-by 7.5-cm strips. Three picomoles of each poly(dT)-tailed oligonucleotide was spotted onto the strips at 0.5-cm intervals. The spots were air dried and then UV cross-linked to strips using the UV Stratalinker (Stratagene). Strips were prehybridized at 45°C for 30 min and then hybridized at 42°C for 90 min. After posthybridization washes, detection was done using an anti-DIG–alkaline phosphatase conjugate and nitroblue tetrazolium (NBT)–5-bromo-4-chloro-3-indolylphosphate (BCIP) color substrate. Development was carried out in the dark for 1 to 2 h. Strips were air dried and photographed.
Specificity.
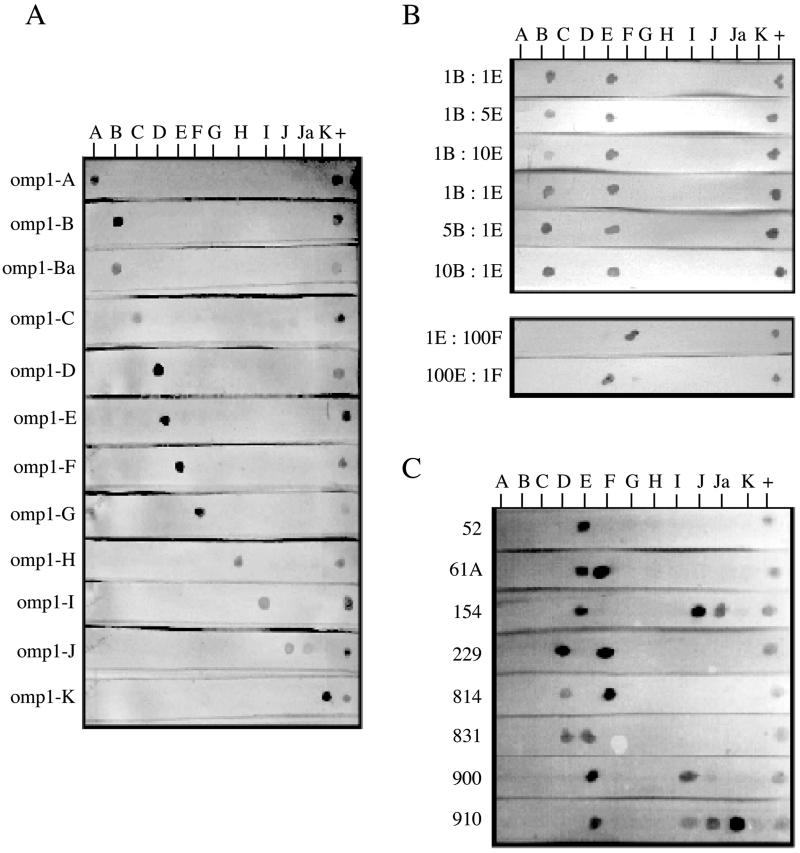
Oligonucleotides hybridized specifically with their corresponding omp1 PCR products (Fig. 1A). Cross-hybridization was found between J and Ja oligonucleotides and the J probe (FA staining does not distinguish J from Ja), although the spot created by the exact match is darker (Fig 1C; compare 154, which contains J, and 910, which contains Ja). Cross-hybridization was seen between the J oligonucleotide and the C probe. Cross-hybridization can also be found in FA staining between C- and J-specific monoclonal antibodies (12, 18). However, even in areas where trachoma is endemic, serovar C is rarely found in urogenital infections (3, 4). Therefore, the importance of this cross-reaction is negligible.
FIG. 1.
RDB results on laboratory and clinical samples. Serotype-specific oligonucleotides are bound in the order A to K, from left to right, on nylon strips. PCR products labeled with DIG are hybridized to the strips. Positive hybridization is visualized as a dark-purple spot on the membrane. (A) MOMP PCR products generated from purified EB are hybridized to strips containing all 12 serotype-specific oligonucleotides and the positive-control oligonucleotide. (B) Laboratory mixed samples were generated by mixing known quantities of DNAs purified from EB of serotypes B, E, and F at different ratios. Positive hybridization results are visualized as dark-purple spots on the membrane. The strips contain all 12 serotype-specific oligonucleotides and the positive-control oligonucleotide as in panel A. (C) RDB hybridization results from clinical specimens containing multiple serovars of chlamydia. PCR products were generated from clinical specimens after they had been expanded in cell culture and prior to their expansion in cell culture (results identical to those shown here).
Mixed-serovar laboratory samples.
DNAs from serovars B and E in one set and from serovars F and I in another were mixed at different ratios. Spots for both serovars were visible (Fig. 1B). DNAs from serovars E and F were mixed in different ratios. For all ratios tested, we were able to visualize both E and F. In the 1:100-ng ratio mixture, the 1-ng spot is light but still visible (Fig. 1B).
Mixed-serovar clinical specimens.
Eight archived samples known to contain more than one serovar of chlamydia by FA staining were evaluated. The number of inclusion-forming units (IFU) for each specimen had been quantified (Table 2), except for isolates 52 and 229. FA and RDB results were identical except for isolates 52 and 910 (Fig. 1C). FA staining detected serovars D and E in specimen 52, while RDB detected only serovar E. In specimen 910, FA staining detected serovars E and J or Ja, while RDB detected E, Ja, and a faint spot corresponding to serovar Ia. Specimens 52 and 910 were retyped by FA staining. Upon retyping, specimen 52 contained only serovar E chlamydiae. The isolate had been passaged in culture three times since the initial typing in 1996.
It is possible that during expansion of specimen 52 in culture, E cells outgrew D cells. For specimen 910, it is possible that the additional Ia spot detected by RDB was a contaminant in the expanded culture or PCR. Nested PCR (23) was done on an aliquot of the original clinical specimen (stored in transport medium at −70°C). Results of RDB on the original specimen were identical to results for the expanded isolates (data not shown).
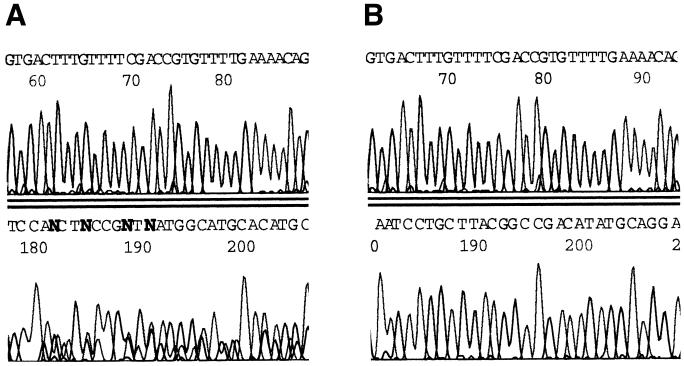
The eight clinical mixed samples were sequenced using MOMP-87, which extends through variable and constant regions of omp1 (23, 24) (Fig. 2 and Table 2). Either one discernible sequence or an ambiguous mixture was obtained. Thus, sequencing did not dependably determine (i) that a mixture was present in the sample or (ii) the identity of the serotypes in the mixture.
FIG. 2.
Automated DNA nucleotide sequencing of the omp1 genes of clinical specimens containing multiple serovars. The omp1 gene was amplified from all eight clinical isolates with mixed serovars and sequenced with the MOMP-87 primer (23), which begins at approximately base 87 in the omp1 gene and extends though conserved regions (CS) 1, 2, and 3 and variable regions (VS) 1 and 2 of the omp1 gene. Sequencing electropherogram results are from specimen 154, which contains serovars E and J in a 1:5 ratio (A), and from specimen 900, which contains serovars E and Ia in a 2:1 ratio (B) in both examples are, on the top row, bases ∼197 to 229 in CS1 and, on the bottom row, bases ∼318 to 345, near the end of VS1 of the omp1 gene. Evidence of a mixture can be seen in specimen 154 but not in specimen 900.
The omp1 gene of C. trachomatis has been a useful marker in epidemiologic studies (3, 4, 28) and for the study of transmission patterns (13, 14). We are examining the transmission of discrete strains within two study populations and use the omp1 gene as a molecular marker. Because these are large studies, the clinical specimens are self-collected vaginal swabs and urine, with primary chlamydia detection by diagnostic PCR. Culture is not performed, making screening for mixed infections by FA staining impossible. DNA sequencing does not reliably detect mixed samples (Fig. 2 and Table 2), and even if mixtures are detected, serovar determination requires cloning of amplicons followed by sequencing of multiple clones. As an example, serovars E and Ja are present at a ratio of 1:10 in specimen 910. Using the binomial equation, if we sequence 20 clones, there is only a 70% chance of finding one serovar E clone.
The eight clinical specimens tested were the only ones in our archives dating back to 1986 that contained more than one serovar detected by FA staining and for which both the original specimen and the expanded isolate could be found. Upon retyping for this study, only serovar E could be found in specimen 52. Thus, RDB results agreed with FA results in seven of eight mixed-serovar specimens. While RDB results on these samples were satisfactory, further testing of this procedure is recommended.
For large epidemiologic studies, when chlamydia culture is not an option, RDB is a simple and sensitive technique for screening and identification of multiple serovars of chlamydia in a clinical specimen. Although we use omp1 sequencing for primary strain identification, RDB could also be used to identify the serovar of chlamydia present in a clinical sample if serovar-specific antibodies are not available or if chlamydia culture is not done. In addition, RDB can be modified for detection of other organisms isolated from human infections, or in a multiplex format to detect several different organisms, such as C. trachomatis and Neisseria gonorrhoeae, in a single clinical specimen.
Acknowledgments
I thank Rebecca Gast, Woods Hole Oceanographic Institute, for technical advice, and Greg Toth for superb technical assistance.
This work was supported by NIH grant U19AI43924, Mid-America Adolescent Sexually Transmitted Diseases Cooperative Research Center (project 2), awarded by the National Institute of Allergy and Infectious Diseases.
REFERENCES
- 1.Baehr W, Zhang Y-X, Joseph T, Su H, Nano F E, Everett K D E, Caldwell H D. Mapping antigenic domains expressed by Chlamydia trachomatis major outer membrane protein genes. Proc Natl Acad Sci USA. 1988;85:4000–4004. doi: 10.1073/pnas.85.11.4000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Batteiger B E, Lennington W, Newhall W J, Katz B P, Morrison H T, Jones R B. Correlation of infecting serovar and local inflammation in genital chlamydial infections. J Infect Dis. 1989;160:332–336. doi: 10.1093/infdis/160.2.332. [DOI] [PubMed] [Google Scholar]
- 3.Brunham R C, Kimani J, Bwayo J, Maitha G, Maclean I, Yang C, Shen C, Roman S, Nagelkerke N J D, Cheang M, Plummer F A. The epidemiology of Chlamydia trachomatis within a sexually transmitted diseases core group. J Infect Dis. 1996;173:950–956. doi: 10.1093/infdis/173.4.950. [DOI] [PubMed] [Google Scholar]
- 4.Brunham R C, Yang C, Maclean I, Kimani J, Maitha G, Plummer F A. Chlamydia trachomatis from individuals in a sexually transmitted disease core group exhibit frequent sequence variation in the major outer membrane protein (omp1) gene. J Clin Investig. 1994;94:458–463. doi: 10.1172/JCI117347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Caldwell H D, Kromhout J, Schachter J. Purification and partial characterization of the major outer membrane protein of Chlamydia trachomatis. Infect Immun. 1981;31:1161–1176. doi: 10.1128/iai.31.3.1161-1176.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Caldwell H D, Judd R C. Structural analysis of chlamydial major outer membrane proteins. Infect Immun. 1982;38:960–968. doi: 10.1128/iai.38.3.960-968.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Caldwell H D, Schachter J. Antigenic analysis of the major outer membrane protein of Chlamydia spp. Infect Immun. 1982;35:1024–1031. doi: 10.1128/iai.35.3.1024-1031.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dean D, Oudens E, Bolan G, Padian N, Schachter J. Major outer membrane protein variants of Chlamydia trachomatis are associated with severe upper genital tract infections and histopathology in San Francisco. J Infect Dis. 1995;172:1013–1022. doi: 10.1093/infdis/172.4.1013. [DOI] [PubMed] [Google Scholar]
- 9.Farencena A, Comanducci M, Donati M, Ratti G, Cevenini R. Characterization of a new isolate of Chlamydia trachomatis which lacks the common plasmid and has properties of biovar trachoma. Infect Immun. 1997;65:2965–2969. doi: 10.1128/iai.65.7.2965-2969.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hamilton P T, Malinowski D P. Nucleotide sequence of the major outer membrane protein gene from Chlamydia trachomatis serovar H. Nucleic Acids Res. 1989;17:8366. doi: 10.1093/nar/17.20.8366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hayes L J, Clark I N. Nucleotide sequence of the major outer membrane protein gene of Chlamydia trachomatis strain A/SA1/OT. Nucleic Acids Res. 1990;18:6136. doi: 10.1093/nar/18.20.6136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hayes L J, Pickett M A, Conlan J W, Ferris S, Everson J S, Ward M E, Clark I N. The major outer membrane proteins of Chlamydia trachomatis serovars A and B: intra-serovar amino acid changes do not alter specificities of serovar- and C subspecies-reactive antibody-binding domains. J Gen Microbiol. 1990;136:1559–1566. doi: 10.1099/00221287-136-8-1559. [DOI] [PubMed] [Google Scholar]
- 13.Hayes L J, Bailey R L, Mabey D C W, Clarke I N, Picket M A, Watt P J, Ward M E. Genotyping of Chlamydia trachomatis from a trachoma-endemic village in The Gambia by a nested polymerase chain reaction: identification of strain variants. J Infect Dis. 1992;166:1173–1177. doi: 10.1093/infdis/166.5.1173. [DOI] [PubMed] [Google Scholar]
- 14.Hayes L J, Pecharatana S, Bailey R L, Hampton T J, Picket M A, Mabey D C W, Watt P J, Ward M E. Extent and kinetics of genetic change in the omp1 gene of Chlamydia trachomatis in two villages with endemic trachoma. J Infect Dis. 1995;172:268–272. doi: 10.1093/infdis/172.1.268. [DOI] [PubMed] [Google Scholar]
- 15.Hatch T P, Vance D W, Al-Hossainy E. Identification of a major envelope protein in Chlamydia spp. J Bacteriol. 1981;146:426–429. doi: 10.1128/jb.146.1.426-429.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Morre S A, Rozendaal L, van Valkengoed I G M, Boeke A J P, van Voorst Vader P C, Schirm J, de Blok S, van den Hoek J A R, van Doornum G J J, Miejer C J L M, van den Brule A J C. Urogenital Chlamydia trachomatis serovars in men and women with a symptomatic or asymptomatic infection: an association with clinical manifestations? J Clin Microbiol. 2000;38:2292–2296. doi: 10.1128/jcm.38.6.2292-2296.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Newhall W J, Batteiger B, Jones R B. Analysis of the human serologic response to proteins of Chlamydia trachomatis. Infect Immun. 1982;38:1181–1189. doi: 10.1128/iai.38.3.1181-1189.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Persson K, Osser S. Serovars of Chlamydia trachomatis causing post-abortion salpingitis. Eur J Clin Microbiol Infect Dis. 1989;8:795–798. doi: 10.1007/BF02185848. [DOI] [PubMed] [Google Scholar]
- 19.Peterson E M, Markoff B A, de la Maza L M. The major outer membrane protein nucleotide sequence of Chlamydia trachomatis, serovar E. Nucleic Acids Res. 1990;18:3414. doi: 10.1093/nar/18.11.3414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sayada C, Denamur E, Elion J. Complete sequence of the major outer membrane protein-encoding gene of Chlamydia trachomatis serovar Da∗. Gene. 1992;120:129–130. doi: 10.1016/0378-1119(92)90022-h. [DOI] [PubMed] [Google Scholar]
- 21.Sayada C, Vretou E, Orfila J, Elion J, Denamur E. Heterogeneity within the first constant segment of the major outer membrane protein gene in Chlamydia trachomatis serovar D/Da distinguishes two lineages. C R Acad Sci. 1995;318:943–949. [PubMed] [Google Scholar]
- 22.Stephens R S, Sanchez-Pescador R, Wagar E A, Inouye C, Urdea M S. Diversity of Chlamydia trachomatis major outer membrane protein genes. J Bacteriol. 1987;169:3879–3885. doi: 10.1128/jb.169.9.3879-3885.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Stothard D R, Boguslawski G, Jones R B. Phylogenetic analysis of the Chlamydia trachomatis major outer membrane protein (MOMP) and examination of potential pathogenic determinants. Infect Immun. 1998;66:3618–3625. doi: 10.1128/iai.66.8.3618-3625.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Stothard D R, Van Der Pol B, Smith N J, Jones R B. Effect of serial passage in tissue culture on sequence of omp1 from Chlamydia trachomatis clinical isolates. J Clin Microbiol. 1998;36:3686–3688. doi: 10.1128/jcm.36.12.3686-3688.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wang S P, Grayston J T, Alexander E R, Holmes K K. Simplified microimmunofluorescence test with trachoma-lymphogranuloma venereum (Chlamydia trachomatis) antigens for use as a screening test for antibody. J Clin Microbiol. 1975;1:250–255. doi: 10.1128/jcm.1.3.250-255.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wang S P, Kuo C-C, Barnes R C, Stephens R S, Grayston J T. Immunotyping of Chlamydia trachomatis with monoclonal antibodies. J Infect Dis. 1985;152:791–800. doi: 10.1093/infdis/152.4.791. [DOI] [PubMed] [Google Scholar]
- 27.Wang S P, Grayston J T. Three new serovars of Chlamydia trachomatis: Da, Ia, and L2a. J Infect Dis. 1991;163:403–405. doi: 10.1093/infdis/163.2.403. [DOI] [PubMed] [Google Scholar]
- 28.Yang C L, Maclean I, Brunham R C. DNA sequence polymorphism of the Chlamydia trachomatis omp1 gene. J Infect Dis. 1993;168:1225–1230. doi: 10.1093/infdis/168.5.1225. [DOI] [PubMed] [Google Scholar]
- 29.Yuan Y, Zhang Y-X, Watkins N G, Caldwell H D. Nucleotide and deduced amino acid sequences for the four variable domains of the major outer membrane proteins of the 15 Chlamydia trachomatis serovars. Infect Immun. 1989;57:1040–1049. doi: 10.1128/iai.57.4.1040-1049.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zhang Y-X, Stewart S, Joseph T, Taylor H R, Caldwell H D. Protective monoclonal antibodies recognize epitopes located on the major outer membrane protein of Chlamydia trachomatis. J Immunol. 1987;138:575–581. [PubMed] [Google Scholar]
- 31.Zhang Y-X, Morrison S G, Caldwell H D. The nucleotide sequence of the major outer membrane protein gene of Chlamydia trachomatis serovar F. Nucleic Acids Res. 1990;18:1061. doi: 10.1093/nar/18.4.1061. [DOI] [PMC free article] [PubMed] [Google Scholar]