Abstract
Background
Our understanding of the cocirculation of infrequently targeted respiratory pathogens and their contribution to symptoms during the coronavirus disease 2019 (COVID-19) pandemic is currently limited. This research aims at (1) understanding the epidemiology of respiratory pathogens since the start of the pandemic, (2) assessing the contribution of non-severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)/influenza/respiratory syncytial virus (RSV) respiratory pathogens to symptoms, and (3) evaluating coinfection rates in SARS-CoV-2-positive patients, both vaccinated and unvaccinated.
Methods
Retrospective analysis of respiratory pathogens identified by the Johns Hopkins Diagnostic Laboratory between December 2019 and October 2021 was performed. In addition, we assessed the contribution of respiratory pathogens other than SARS-CoV-2 to symptomatic disease by retesting 2 cohorts of specimens that were (1) collected from symptomatic patients and (2) received limited respiratory pathogen testing. The first cohort was patients who tested negative by the standard-of-care SARS-CoV-2/influenza/RSV testing. The second was a cohort of SARS-CoV-2-positive, symptomatic, fully COVID-19 immunized and unimmunized patients.
Results
Between December 2019 and October 2021, a total of 11 806, 62 829, and 579 666 specimens were tested for an extended respiratory panel, influenza/RSV with or without SARS-CoV-2 panel, or SARS-CoV-2, respectively. Positivity rates of different targets differed between different months and were impacted by the COVID-19 pandemic. The SARS-CoV-2-negative cohort had 8.5% positivity for other respiratory pathogens that included primarily enterovirus/rhinovirus (5.8%). In the SARS-CoV-2-positive cohort, no other respiratory pathogens were detected.
Conclusions
The COVID-19 pandemic impacted the circulation of certain respiratory pathogens. Other respiratory viral pathogens were associated with symptomatic infections; however, coinfections with SARS-CoV-2 were highly uncommon.
Keywords: coinfections, COVID-19, SARS-CoV-2
The COVID-19 pandemic impacted the circulation of seasonal respiratory viruses, however, viral pathogens other than SARS-CoV-2 were associated with symptomatic infections. Coinfections with SARS-CoV-2 remains uncommon even in fully immunized individuals.
The concurrent circulation of respiratory pathogens coupled with the algorithms of laboratory diagnosis challenges a clear understanding of the association of infrequently screened pathogens with symptomatic infections and severity of disease. The disease caused by respiratory pathogens can be challenging to differentiate clinically; however, a specific laboratory diagnosis facilitates proper patient management and assists with implementing infection control measures [1, 2]. When severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and influenza viruses cocirculate, the Centers for Disease Control and Prevention (CDC) guides providers to order testing for both viruses, mainly when the patient requires hospital admission [3]. The multiplex panels that target larger numbers of respiratory pathogens are recommended for limited patient populations that include immunocompromised and hospitalized patients [4]. However, with the widespread circulation of SARS-CoV-2, testing has been primarily targeted for this virus, and it has included testing of symptomatic patients and screening of asymptomatic individuals to limit viral transmission [5].
Multiple assays that allow the simultaneous detection of influenza and SARS-CoV-2 were authorized by the US Food and Drug Administration as early as the beginning of the 2020–2021 influenza season, and some of them added the detection of respiratory syncytial virus (RSV) [6]. At Johns Hopkins Hospital, algorithms are in place to test symptomatic patients for influenza A/B, RSV, and SARS-CoV-2 [7], and an extended respiratory panel is primarily run for patients who are immunocompromised or hospitalized per the Infectious Diseases Society of America (IDSA) guidelines [8]. In addition, large-scale SARS-CoV-2 standalone testing is performed for symptomatic and asymptomatic patients. This algorithm of testing may have some limitations that include the following: (1) symptomatic patients with a positive SARS-CoV-2 result are not likely to receive testing for other respiratory pathogens, which limits our understanding of SARS-CoV-2 coinfections and their contribution to symptoms and disease severity; and (2) most symptomatic patients who have a negative result by the influenza A/B/RSV/SARS-CoV-2 panel do not receive additional testing for other respiratory pathogens, which limits the data on the epidemiology of these pathogens to a restricted patient population that receives the extended panel testing. In addition, (1) the impact of other variables including the unique nature of the respiratory season of 2020–2021 that remarkably did not witness influenza circulation and (2) the discrepancies in measures of social distancing, masking, and infection control in different times and locations throughout the year warrant a better surveillance of the circulating respiratory pathogens to better understand their contribution to symptoms and disease.
In this study, we provide detailed data on the circulation of respiratory pathogens in the time frame between December 2019 and October 2021 from Johns Hopkins Clinical Virology Laboratory, which serves a large geographical area in the National Capital Region. In addition, we provide data on retesting a cohort of symptomatic patients who were negative for influenza A/B/RSV/SARS-CoV-2 and a cohort of symptomatic patients who tested positive for SARS-CoV-2 (including a fully coronavirus disease 2019 [COVID-19] immunized group [breakthrough infections]) for respiratory pathogens that included enterovirus/rhinovirus, parechovirus, adenovirus, parainfluenza viruses (human parainfluenza virus [HPIV]1, HPIV2, HPIV3), endemic coronaviruses (human coronavirus [HCoV]-OC43, HCoV-HKU1, HCoV-229E, and HCoV-NL63), in addition to Mycoplasma pneumoniae and Chlamydia pneumoniae to assess the potential contribution of these pathogens to symptoms.
MATERIALS AND METHODS
Patient Consent Statement
The research was performed with a waiver of consent with a protocol approved by the Johns Hopkins Institutional Review Board committee (IRB00221396). Remnant nasopharyngeal (NP) clinical swab specimens from patients after the standard-of-care testing was used. Clinical testing is performed for inpatients and outpatients across the Johns Hopkins Medical System, which includes 5 acute care hospitals and more than 40 ambulatory care offices. Severe acute respiratory syndrome coronavirus 2 testing is also performed through screening and testing services at several long-term care facilities in the State of Maryland and mobile outreach clinics.
Sample Selection
Cohort 1
The NP samples (N = 294) were randomly selected based on the criteria of being from patients with respiratory symptoms who had tested negative for influenza A/B, RSV, and SARS-CoV-2 and did not have any additional respiratory testing done (age range, 0 to >60 years). Of note, symptomatic illness was defined based on the ordering clinician and is a questionnaire included in the clinical charts for all patients receiving SARS-CoV-2 testing, which started system wide at the end of 2020. Specimens were collected in Universal Transport Media (UTM) between October 2020 and October 2021 and were kept frozen at −80°C until nucleic acid extraction.
Cohort 2
The NP samples (N = 179) were selected from symptomatic patients, 71 of which were “Vaccine breakthrough infections” based on the CDC definition: positive SARS-CoV-2 test results more than 14 days after the second shot for Pfizer/BioNTech BNT162b2 and Moderna mRNA-1273 or 14 days after the J&J/Janssen shot, and 108 were NP samples from symptomatic SARS-CoV-2 positive unvaccinated patients. Specimens were identified through the Johns Hopkins SARS-CoV-2 genomic surveillance [9, 10] and had collection dates between February and September 2021 (age range, 0 to >60 years).
Assays Used for Clinical Diagnosis
Respiratory viral testing at Johns Hopkins between December 2019 and October 2021 was performed using the Cepheid panels Xpert Xpress Flu/RSV and Xpert Xpress SARS-CoV-2/Flu/RSV [2] and the Roche (previously GenMark) Respiratory Pathogen Panel (RP) and RP2. The RP panel detects the following targets: adenovirus, HCoV-229E, HCoV-HKU1, HCoV-NL63, HCoV-OC43, human metapneumovirus (hMPV), enterovirus/rhinovirus, influenza A/A H1/A H1-2009/A H3, influenza B, parainfluenza (PIV)1–4, RSV A, RSV B, C pneumoniae, and M pneumoniae. The RP2 panel includes all targets from the RP plus SARS-CoV-2.
Before December 16, 2019, the Luminex NxTAG Respiratory Pathogen Panel (RPP) was used for extended respiratory testing. The RPP includes the following targets: adenovirus, HCoV-229E, HCoV-HKU1, HCoV-NL63, HCoV-OC43, hMPV, enterovirus/rhinovirus, influenza A/A H1/A H1-2009/A H3, influenza B, PIV1–4, RSV A, RSV B, bocavirus, C pneumoniae, and M pneumoniae [8].
Molecular assays used for SARS-CoV-2 diagnosis include the following: RealStar SARS-CoV-2 RT-PCR (Altona Diagnostics), Xpert Xpress SARS-CoV-2/Flu/RSV (Cepheid), NeuMoDx SARS-CoV-2 (QIAGEN), Cobas SARS-CoV-2 (Roche), ePlex RP2 (Roche), Aptima SARS-CoV-2 (Hologic), and Accula SARS-CoV-2 assays (Thermo Fisher Scientific) [2, 8, 11, 12].
Additional Testing for Cohorts 1 and 2
Controls
Adenovirus (Exact Diagnostics, Fort Worth, TX; starting concentration 1 E6 log10 IU/mL) was diluted into UTM to a concentration of 1 E3 IU/mL and used as an extraction control. Each of the 4 endemic coronaviruses (using previous positive patients’ samples) were used as amplification controls in the multiplex coronavirus laboratory-developed test ([LDT] see below for assay details). Simplexa extraction and amplification control (SEAC) deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) were added as an internal control to all samples to serve as an amplification control and asses polymerase chain reaction (PCR) inhibition in the DiaSorin Molecular LDT.
Nucleic Acid Extraction
Automated nucleic acid extraction was performed using the Chemagic 360 instrument (PerkinElmer, Austin, TX) with software version v6.3.0.3. Specimen extraction used a sample volume of 300 µL and elution volume of 60 µL. Extraction was performed using the manufacturer’s protocol.
Nucleic Acid Amplification and Detection
DiaSorin Molecular reagents were used to run 4 separate multiplexed assays: HPIV (included HPIV1–3), enterovirus (included enterovirus and parechovirus), adenovirus (included adenovirus 3’ and 5’ hexon), and pneumoniae (included M pneumoniae and C pneumoniae). The total amplification reaction volume was 10 µL (consisting of 5 or 6 µL reagent mix and 4 or 5 µL extracted sample). The reagent mix for RNA targets included 4 µL of 2.5× Universal Master Mix, 0.2 µL SEAC RNA primer pair, 0.5 µL reverse transcriptase, 0.2 µL each of target primer pair and 0.7 or 1 µL of water (HPIV mix or enterovirus mix, respectively). The reagent mix for DNA targets included 4 µL of 2.5× Universal Master Mix, 0.2 µL SEAC DNA primer pair, 0.2 µL each of target pair, and 0.4 µL water. Reverse-transcription PCR (RT-PCR) for RNA targets was performed using the 96-well Universal Disc on the LIAISON MDX (DiaSorin) using the following cycling conditions: 2 holds at 47°C for 600 seconds and 97°C for 200 seconds, followed by 40 cycles of 97°C for 5 seconds and 58°C for 30 seconds. The PCR for DNA targets used the following cycling conditions: hold at 97°C for 120 seconds, followed by 40 cycles of 97°C for 10 seconds and 60°C for 30 seconds. All assays used a cycle threshold cutoff of 40.
An LDT TaqMan real-time RT-PCR (rRT-PCR) was used to run the multiplex assay for the endemic coronaviruses (HCoV-OC43, HKU1, 229E, and NL63) [13]. The total reaction volume was 20 µL, including 5 µL TaqPath 1-Step RT-qPCR Master Mix (Applied Biosystems), 8.4 µL water, 0.18 µL each of the forward and reverse primers for HCoV-HKU1, NL63, and OC43 (100 µM) and HCoV-229E (50 µM), 0.04 µL of each probe (100 µM for HCoV-HKU1, HCoV-NL63, and HCoV-OC43; 50 µM for HCoV-229E), and 5 µL of the template. The rRT-PCR was performed using the Prism 7500 Sequence Detection System (Applied Biosystems) using the following cycling conditions: 1 cycle at 25°C for 2 minutes, 1 cycle at 50°C for 15 minutes, 1 cycle at 95°C for 2 minutes then 40 cycles at 95°C for 3 seconds and 60°C for 30 seconds.
RESULTS
Respiratory Testing and Positivity Trends
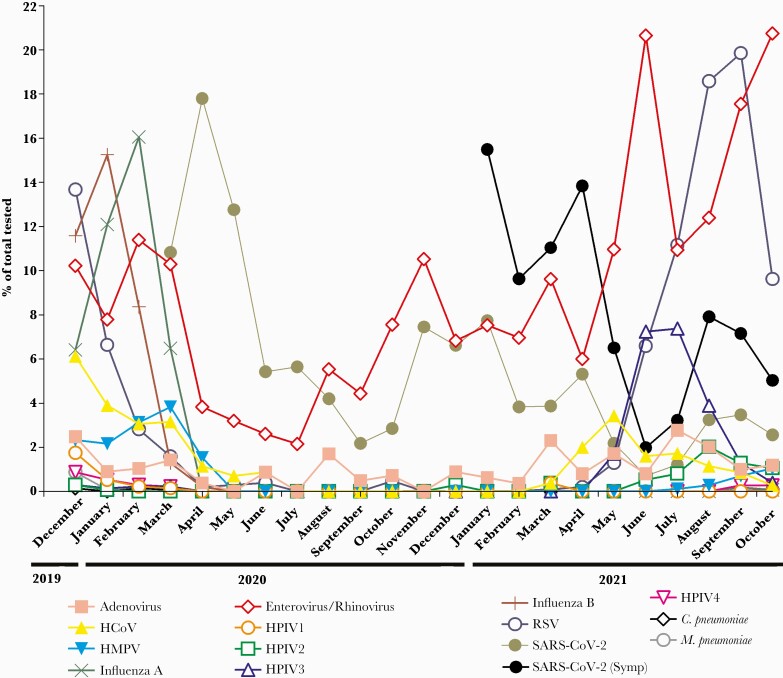
In the time frame between December 2019 and the end of October 2021, a total of 74 635 respiratory samples were tested with both the Cepheid (62 829) and ePlex (11 806) panels. Of these, there were a total of 10 055 single-target positive samples detected (excluding SARS-CoV-2) and 245 samples positive for more than 1 target (Supplementary Table S1 and Table 1), with an overall positivity of 13.8%, excluding SARS-CoV-2. In the same time frame, 579 666 SARS-CoV-2 tests were performed with a total of 29 019 positives diagnosed by all platforms (average 5% positivity). Before testing for SARS-CoV-2 started (March 2020) and before the COVID-19 pandemic (between December 2019 and February 2020), the highest positivity rates were for influenza A (peak of 16.1 % in February 2020), influenza B (peak of 15.3% in January 2020), and RSV (peak of 13.7 % in December 2019) (Figure 1). Severe acute respiratory syndrome coronavirus 2 testing started on March 11, 2020 at Johns Hopkins laboratory, the same date the World Health Organization (WHO) declared COVID-19 as a pandemic [12, 14]. After the pandemic, the months between March 2020 and April 2021 showed a predominance in SARS-CoV-2 positivity (peak of 17.8% in April 2020) (Figure 1 and Supplementary Table S1). It is notable that enterovirus/rhinovirus was still detectable in this time frame with a positivity that peaked at 10.5% in November 2020. All other targets at that time frame showed no to very minimal circulation. When symptomatic SARS-CoV-2 positivity was separated from asymptomatic screening (based on the test order [Figure 1, starting from January 2021]), it was notable that SARS-CoV-2 positivity was higher than enterovirus/rhinovirus (Figure 1). On May 2021, other respiratory viruses’ positivity started to outcompete SARS-CoV-2 (Figure 1). A predominance of enterovirus/rhinovirus (peak of 20.7% in October 2021) was notable followed by RSV (19.9% in September 2021) and HPIV3 (7.3% in July 2021) (Figure 1).
Table 1.
Coinfections With Respiratory Viral Pathogens Between December 2019 and October 2021 of Samples Diagnosed at Johns Hopkins Microbiology Laboratorya
| 2019 | 2020 | 2021 | Total | ||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| December | January | February | March | April | May | June | July | August | September | October | November | December | January | February | March | April | May | June | July | August | September | October | |||
| Total | 685 | 1335 | 1343 | 1332 | 261 | 281 | 230 | 233 | 235 | 203 | 278 | 266 | 337 | 319 | 273 | 260 | 250 | 292 | 373 | 868 | 694 | 701 | 757 | 11 806 | |
| Positives for 2 or more targets | 31 | 31 | 27 | 25 | 1 | 0 | 1 | 4 | 3 | 0 | 1 | 0 | 2 | 3 | 0 | 4 | 0 | 8 | 15 | 30 | 26 | 18 | 15 | 245 | |
| %coinfections | 4.53 | 2.32 | 2.01 | 1.88 | 0.38 | 0.00 | 0.43 | 1.72 | 1.28 | 0.00 | 0.36 | 0.00 | 0.59 | 0.94 | 0.00 | 1.54 | 0.00 | 2.74 | 4.02 | 3.46 | 3.75 | 2.57 | 1.98 | 2.08 | % to total |
| Influenza A | 4 | 4 | 6 | 6 | 1 | 1 | 1 | 23 | 9.4 | ||||||||||||||||
| Influenza B | 4 | 4 | 5 | 2 | 2 | 1 | 18 | 7.3 | |||||||||||||||||
| RSV | 14 | 8 | 4 | 2 | 1 | 1 | 1 | 6 | 5 | 13 | 10 | 14 | 79 | 32.2 | |||||||||||
| Adenovirus | 4 | 8 | 9 | 7 | 1 | 1 | 1 | 3 | 1 | 1 | 2 | 1 | 3 | 6 | 4 | 1 | 1 | 54 | 22.0 | ||||||
| HCoV | 10 | 5 | 8 | 8 | 1 | 1 | 4 | 1 | 3 | 3 | 1 | 45 | 18.4 | ||||||||||||
| HMPV | 5 | 8 | 4 | 6 | 1 | 2 | 5 | 31 | 12.7 | ||||||||||||||||
| Enterovirus/rhinovirus | 20 | 14 | 19 | 17 | 1 | 3 | 1 | 1 | 3 | 3 | 6 | 12 | 20 | 21 | 18 | 13 | 172 | 70.2 | |||||||
| HPIV1 | 1 | 1 | 2 | 0.8 | |||||||||||||||||||||
| HPIV2 | 1 | 1 | 1 | 1 | 7 | 5 | 3 | 3 | 22 | 9.0 | |||||||||||||||
| HPIV3 | 3 | 7 | 18 | 5 | 2 | 35 | 14.3 | ||||||||||||||||||
| HPIV4 | 5 | 4 | 1 | 10 | 4.1 | ||||||||||||||||||||
| Mycoplasma pneumoniae | 1 | 1 | 1 | 1 | 4 | 1.6 | |||||||||||||||||||
| SARS-CoV-2 | 1 | 1 | 2 | 0.8 | |||||||||||||||||||||
Abbreviations: HCoV, human coronaviruses; HMPV, human metapneumovirus; HPIV, human parainfluenza virus; RSV, respiratory syncytial virus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2.
Detailed combinations of coinfections with 1 or more targets are shown in Supplementary Table S2.
Figure 1.
Respiratory virus positivity rates from December 2019 to October 2021. Total numbers tested and positive tests for each target are detailed in Supplementary Table S1. HMPV, human metapneumovirus; HPIV, human parainfluenza virus; RSV, respiratory syncytial virus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SARS-CoV-2 (Symp), indicates symptomatic testing positivity.
There was a notable decrease in the numbers of respiratory panel testing conducted between December 2020 and March 2021 with an overall 75% reduction compared to the same time frame in 2019–2020 (1189 and 4695 tests, respectively) (Supplementary Table S1). Between December 2019 and March 2020, an overall positivity rate of 27.3% was recorded, with the overwhelming majority being influenza A (10.1%), enterovirus/rhinovirus (9.9%), followed by influenza B (8.5%) and RSV (5.4%). Although less respiratory highly multiplexed panels were run in the same time frame in 2020–2021, the overall positivity rate was also remarkably less than the previous year (0.5%) if SARS-CoV-2 was excluded. In addition to SARS-CoV-2, which had an overall positivity of 5.7%, the main circulating viruses were enterovirus/rhinovirus (7.7%) and adenovirus (1%), and both had positivity rates comparable to the same time frame from the previous year (Supplementary Table S1).
Respiratory Viral Coinfections Trends
Coinfections with more than 1 target were infrequently encountered (2.1%) (Table 1), with only 2 specimens that were positive for SARS-CoV-2 and enterovirus/rhinovirus in January 2021 or SARS-CoV-2, enterovirus/rhinovirus, and adenovirus in August 2021 (of a total of 173 SARS-CoV-2 positives by the extended respiratory panel [RP2]). Enterovirus/rhinovirus was the most frequently detected target in cases of coinfections in all months (70.2%) (Table 1), and the monthly trends correlated with the overall positivity trends (Figure 1). Detailed combinations of coinfections with 2 or more targets are shown in Supplementary Table S2. The months between May and October 2021, which witnessed a rebound in other respiratory viruses than SARS-CoV-2 and an increase in enterovirus/ rhinovirus positivity, correlated with the highest coinfection rates since the beginning of the pandemic with a peak in the month of June (Table 1). It is notable that patients with more than 1 target detected in this time frame were mostly pediatric patients (18 years or younger [89%], 2 years or younger [63.5%]).
Identification of Nonsevere Acute Respiratory Syndrome Coronavirus 2 Respiratory Pathogens in Symptomatic Patients
To identify the correlation of other respiratory pathogens than influenza, RSV, and SARS-CoV-2 as well as coinfections with SARS-CoV-2 with symptomatic respiratory infections, we performed additional testing using 2 cohorts. The first cohort of patients had samples collected and were negative for influenza A/B, RSV, and SARS-CoV-2 testing between October 2020 and October 2021. Between October 2020 and March 2021, detailed chart reviews were performed to verify the symptomatic presentation when patients were tested. Of the 200 patients during this time frame, 98.5% were outpatients. Our data showed that 118 of the patients with available detailed information in the clinical records presented with symptoms of viral illness that included fever, cough, congestion, sore throat, headache, or exacerbation of asthma. Fifteen patients were admitted (7.5%) secondary to respiratory symptoms. Assessment of the symptomatic status was added as a questionnaire to the clinical charts in the beginning of 2021, and hence negative samples from patients tested in September and October 2021 were all from patients with respiratory symptoms. Additional testing of the total of 294 samples for pathogens associated with respiratory disease showed that 25 samples were positive for 1 of the 8 tested analytes. Seventeen samples were positive for enterovirus/rhinovirus, 6 were adenovirus positive, 1 was positive for HCoV-OC43, and 1 was positive for HPIV2 (Table 2).
Table 2.
Cohorts From Symptomatic Patients Tested for Respiratory Viral Targets That Included Adenovirus, HCoV-229E, HCoV-HKU1, HCoV-NL63, HCoV-OC43, HMPV, Enterovirus/Rhinovirus, and HPIV1–4 in Addition to Chlamydia pneumoniae and Mycoplasma pneumoniae
| Cohort 1 (Samples From Symptomatic Patients Tested Clinically and Were Negative for SARS-CoV-2/Influenza/RSV) | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| October 2020 | November 2020 | December 2020 | January 2021 | February 2021 | March 2021 | April 2021 | May 2021 | July 2021 | August 2021 | September 2021 | October 2021 | Total | |
| Total tested | 16 | 37 | 37 | 36 | 37 | 37 | 48 | 46 | 294 | ||||
| Total positives | 3 | 2 | 2 | 6 | 2 | 3 | 1 | 6 | 25 | ||||
| %Positivity | 18.8 | 5.4 | 5.4 | 16.7 | 5.4 | 8.1 | 2.1 | 13.0 | 8.5 | ||||
| Targets detected | Enterovirus/rhinovirus (3) | Enterovirus/ rhinovirus (1) | Enterovirus/ rhinovirus (1) | Enterovirus/ rhinovirus (4) | Enterovirus/rhinovirus (2) | Enterovirus/rhinovirus (2) | Enterovirus/ rhinovirus (1) | Enterovirus/ rhinovirus (3) | 17 | ||||
| Adenovirus (1) | Adenovirus (1) | Adenovirus (1) | Adenovirus (3) | 6 | |||||||||
| HPIV2 (1) | 1 | ||||||||||||
| HCoV-OC43 (1) | 1 | ||||||||||||
| Cohort 2 (Samples From Symptomatic Patients Tested Clinically and Were Positive for SARS-CoV-2) | |||||||||||||
| Total tested | 1 | 12 | 33 | 3 | 35 | 4 | 91 | 179 | |||||
| Fully vaccinated | 71 | ||||||||||||
| Unvaccinated | 108 | ||||||||||||
Abbreviations: HCoV, human coronaviruses; HMPV, human metapneumovirus; HPIV, human parainfluenza virus; RSV, respiratory syncytial virus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2.
The second cohort of patients was SARS-CoV-2-positive symptomatic patients (N = 179), and 39.7% of them were symptomatic, fully COVID-19-vaccinated individuals. Thirty-five samples (41.7%) had a collection date in July 2021, 33 (39.3%) had a collection date in April 2021, and 91 (50.8%) were collected in September 2021 (Table 2). Testing these samples for other respiratory pathogens revealed no coinfections with any of the tested respiratory pathogens, and SARS-CoV-2 was the sole pathogen detected.
DISCUSSION
Severe acute respiratory syndrome coronavirus 2 caused a devastating pandemic that was declared by the WHO on March 11, 2020. Throughout the initial one and half years of the pandemic, different measures for limiting the spread of the virus were implemented, including but not limited to facial coverings and masking, closures and adaptations of schools and businesses, restrictions of gatherings, periodic stay-at-home orders, and restrictions of international travel [15]. Other measures included hand hygiene, case isolations, and contact tracing in an attempt to limit further transmission. Fortunately, associated with these infection control measures was a marked impact on the transmission of other respiratory viral infections, especially influenza and RSV [16–19]. At the Johns Hopkins Clinical Microbiology Laboratory, a marked reduction in influenza and RSV positivity was noted in late 2020–early 2021 compared to the same time frame from the previous year, 2019–2020. However, enterovirus/rhinovirus and adenovirus positivity rates were similar. Our data indicate that even though the circulation of some of the respiratory viruses was impacted by the COVID-19 pandemic, enterovirus/rhinovirus and adenovirus were likely circulating in rates similar to the prepandemic. The discrepancy of the impact of the pandemic on the circulation of respiratory viruses indicates that intrinsic viral characteristics might have influenced their circulation. This could include the viral structure (nonenveloped viruses: enterovirus/rhinovirus and adenovirus; enveloped viruses: influenza, HPIV, HMPV, HCoV, and RSV) that could impact the viral survival and sensitivity to environmental conditions and disinfectants.
Large-scale SARS-CoV-2 testing was essential for diagnosis, screening, and contact tracing. In the State of Maryland, at the time of writing this manuscript (November 13, 2021), a total of 14 548 741 tests were performed with 570 806 confirmed cases (https://coronavirus.maryland.gov/). Severe acute respiratory syndrome coronavirus 2 testing in the clinical diagnostic laboratories has become a front-line order for patients with symptomatic respiratory disease [20]. Integrating SARS-CoV-2 testing impacted the diagnostic algorithms of other respiratory viruses, especially with the limited circulation of influenza and RSV. At Johns Hopkins Clinical Microbiology Laboratory, SARS-CoV-2 testing volumes exceeded testing for other respiratory pathogens by approximately 8-fold since the beginning of the pandemic. However, our data showed that other respiratory viruses started to increase in frequency after May 2021, including enterovirus/rhinovirus, RSV, and HPIV3. Our data also showed that enterovirus/rhinovirus maintained a constant circulation that increased dramatically starting in June 2021. Our data also show that enterovirus/rhinovirus was the detectable pathogens from 5.8% of samples from symptomatic patients who were only tested for SARS-CoV-2, influenza, and RSV. This highlights the importance of considering the association of these viruses with the increase in symptomatic influenza-like illness in the recent months when their positivity was the predominant.
With the widespread increase in breakthrough symptomatic SARS-CoV-2 infections associated with the predominance of the Delta variant of concern since June 2021 [21], it is essential to assess the contribution of other respiratory circulating viruses to symptoms in this population of patients. Our data indicate that symptomatic SARS-CoV-2-positive patients, whether vaccinated or unvaccinated, are not likely to be coinfected with other respiratory pathogens, consistent with previous reports that showed a low frequency of SARS-CoV-2 coinfections [22–25]. It is notable that this cohort of patients had sample collection times that spanned from March to September 2021 and coincided with increases in the circulation of enterovirus/rhinovirus, RSV, and HPIV3 (Figure 1).
The use of syndromic panels has been limited to specific patient populations; hence, we thought that sampling negative SARS-CoV-2 specimens for an extended panel of respiratory targets could give us an idea about the epidemiology of these pathogens in an unbiased setting. Of the 294 samples tested, only 25 were positive (8.5%); 17 enterovirus/rhinovirus, 6 adenovirus, 1 HCoV-OC43, and 1 HPIV2. It is notable that the positivity rate from this cohort was comparable to the positivity rate using the extended diagnostic panel, and the trends of the detected pathogens were consistent with what was detectable by the ePlex panels (Supplementary Table S1), highlighting the relevance of using the diagnostic panel results for surveillance and epidemiology even if their use is limited to a specific population of patients.
The limitations of the study include its retrospective nature and the lack of controlling the timing of sample collections in relation to the course of the symptoms or disease. In addition, for outpatients who tested positive for respiratory viruses, only limited clinical data sets are available through the patients’ electronic medical records, which limits assessing the association among symptoms, outcomes, and targets positivity. In addition, the detection of more than one target might reflect prolonged shedding of one of the detected targets due to a past infection, rather than a true coinfection.
CONCLUSIONS
In summary, our data show that the COVID-19 pandemic impacted the circulation of certain respiratory viruses. The large-scale testing of other respiratory pathogens might not be warranted for SARS-CoV-2-positive patients due to the infrequent coinfections with SARS-CoV-2. However, extended respiratory panels are beneficial for understanding the trends in the circulation of other respiratory pathogens that are not frequently tested for in symptomatic patients.
Supplementary Data
Supplementary materials are available at Open Forum Infectious Diseases online. Consisting of data provided by the authors to benefit the reader, the posted materials are not copyedited and are the sole responsibility of the authors, so questions or comments should be addressed to the corresponding author.
Supplementary Table S1. Respiratory Pathogen Testing and Total Positives Per Target at Johns Hopkins Between December 2019 and October 2021.
Supplementary Table S2. Coinfections with respiratory viral pathogens between December 2019 and October 2021 of samples diagnosed at Johns Hopkins Microbiology laboratory. HMPV, human metapneumovirus; HPIV, human parainfluenza virus; RSV, respiratory syncytial virus.
Acknowledgments
This study was only possible with the unique efforts of the Johns Hopkins Clinical Microbiology Laboratory faculty and staff.
Disclaimer. The views expressed in this manuscript are those of the authors and do not necessarily represent the views of the National Institute of Biomedical Imaging and Bioengineering; the National Heart, Lung, and Blood Institute; the National Institutes of Health; or the US Department of Health and Human Services.
Financial support. The study was funded by DiaSorin Molecular and Johns Hopkins University. H. H. M. is supported by the HIV Prevention Trials Network sponsored by the National Institute of Allergy and Infectious Diseases, the Johns Hopkins Center of Excellence in Influenza Research and Surveillance (HHSN272201400007C), National Institute on Drug Abuse, National Institute of Mental Health, and Office of AIDS Research, of the National Institutes of Health (NIH), US Department of Health and Human Services (UM1 AI068613), the NIH RADx-Tech program (3U54HL143541-02S2), National Institute of Health RADx-UP initiative (Grant R01 DA045556-04S1), Centers for Disease Control and Prevention (contract 75D30121C11061), the Johns Hopkins University President’s Fund Research Response, the Johns Hopkins Department of Pathology, and the Maryland Department of Health.
Potential conflicts of interest. All authors: No reported conflicts of interest. All authors have submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest.
References
- 1. Solomon DA, Sherman AC, Kanjilal S.. Influenza in the COVID-19 era. JAMA 2020; 324:1342–3. [DOI] [PubMed] [Google Scholar]
- 2. Mostafa HH, Carroll KC, Hicken R, et al. . Multi-center evaluation of the cepheid Xpert (R) Xpress SARS-CoV-2/Flu/RSV test. J Clin Microbiol 2020. doi: 10.1128/JCM.02955-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Centers for Disease Control and Prevention. Testing guidance for clinicians when SARS-CoV-2 and influenza viruses are co-circulating. Available at: https://www.cdc.gov/flu/professionals/diagnosis/testing-guidance-for-clinicians.htm. Accessed 7 February 2022.
- 4. Uyeki TM, Bernstein HH, Bradley JS, et al. . Clinical practice guidelines by the Infectious Diseases Society of America: 2018 update on diagnosis, treatment, chemoprophylaxis, and institutional outbreak management of seasonal influenzaa. Clin Infect Dis 2018; 68:e1–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Centers for Disease Control and Prevention. Overview of testing for SARS-CoV-2 (COVID-19). Available at: https://www.cdc.gov/coronavirus/2019-ncov/hcp/testing-overview.html. Accessed 7 February 2022.
- 6. Centers for Disease Control and Prevention. Multiplex assays authorized for simultaneous detection of influenza viruses and SARS-CoV-2 by FDA. Available at: https://www.cdc.gov/flu/professionals/diagnosis/table-flu-covid19-detection.html. Accessed 7 February 2022.
- 7. Mostafa HH, Carroll KC, Hicken R, et al. . Multicenter evaluation of the Cepheid Xpert Xpress SARS-CoV-2/Flu/RSV test. J Clin Microbiol 2021; 59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Jarrett J, Uhteg K, Forman MS, et al. . Clinical performance of the GenMark Dx ePlex respiratory pathogen panels for upper and lower respiratory tract infections. J Clin Virol 2021; 135:104737. [DOI] [PubMed] [Google Scholar]
- 9. Morris CP, Luo CH, Amadi A, et al. . An update on SARS-CoV-2 diversity in the United States national capital region: evolution of novel and variants of concern. Clin Infect Dis 2021. doi: 10.1093/cid/ciab636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Thielen PM, Wohl S, Mehoke T, et al. . Genomic diversity of SARS-CoV-2 during early introduction into the Baltimore-Washington metropolitan area. JCI Insight 2021; 6:e144350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Mostafa HH, Hardick J, Morehead E, Miller JA, Gaydos CA, Manabe YC.. Comparison of the analytical sensitivity of seven commonly used commercial SARS-CoV-2 automated molecular assays. J Clin Virol 2020; 130:104578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Uhteg K, Jarrett J, Richards M, et al. . Comparing the analytical performance of three SARS-CoV-2 molecular diagnostic assays. J Clin Virol 2020; 127:104384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Gaunt ER, Hardie A, Claas EC, Simmonds P, Templeton KE.. Epidemiology and clinical presentations of the four human coronaviruses 229E, HKU1, NL63, and OC43 detected over 3 years using a novel multiplex real-time PCR method. J Clin Microbiol 2010; 48:2940–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Hu B, Guo H, Zhou P, Shi Z-L.. Characteristics of SARS-CoV-2 and COVID-19. Nat Rev Microbiol 2021; 19:141–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. World Health Organization. WHO health emergency dashboard. Available at: https://covid19.who.int/. Accessed 7 February 2022.
- 16. Huang QS, Wood T, Jelley L, et al. . Impact of the COVID-19 nonpharmaceutical interventions on influenza and other respiratory viral infections in New Zealand. Nat Commun 2021; 12:1001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Olsen SJ, Azziz-Baumgartner E, Budd AP, et al. . Decreased influenza activity during the COVID-19 pandemic-United States, Australia, Chile, and South Africa, 2020. Am J Transplant 2020; 20:3681–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Sherman AC, Babiker A, Sieben AJ, et al. . The effect of severe acute respiratory syndrome Coronavirus 2 (SARS-CoV-2) mitigation strategies on seasonal respiratory viruses: a tale of 2 large metropolitan centers in the United States. Clin Infect Dis 2021; 72:e154–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Uyeki TM, Wentworth DE, Jernigan DB.. Influenza activity in the US during the 2020-2021 season. JAMA 2021; 325:2247–8. [DOI] [PubMed] [Google Scholar]
- 20. Kevadiya BD, Machhi J, Herskovitz J, et al. . Diagnostics for SARS-CoV-2 infections. Nat Mater 2021; 20:593–605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Luo CH, Morris CP, Sachithanandham J, et al. . Infection with the SARS-CoV-2 delta variant is associated with higher recovery of infectious virus compared to the alpha variant in both unvaccinated and vaccinated individuals. Clin Infect Dis 2021. doi: 10.1093/cid/ciab986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Marshall NC, Kariyawasam RM, Zelyas N, Kanji JN, Diggle MA.. Broad respiratory testing to identify SARS-CoV-2 viral co-circulation and inform diagnostic stewardship in the COVID-19 pandemic. Virol J 2021; 18:93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Zhang DD, Acree ME, Ridgway JP, et al. . Characterizing coinfection in children with COVID-19: a dual center retrospective analysis. Infect Control Hosp Epidemiol 2020; 1: 3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Nowak MD, Sordillo EM, Gitman MR, Paniz Mondolfi AE.. Coinfection in SARS-CoV-2 infected patients: where are influenza virus and rhinovirus/enterovirus? J Med Virol 2020; 92:1699–700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Mostafa HH, Fissel JA, Fanelli B, et al. . Metagenomic next-generation sequencing of nasopharyngeal specimens collected from confirmed and suspect COVID-19 patients. mBio 2020; 11:e01969–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.