Abstract
A major inhibitor of diagnostic PCR in human plasma was identified and the mechanism of inhibition was characterized. Human blood was divided by centrifugation into buffy coat, plasma, platelets, and erythrocytes. All these blood fractions were found to be highly inhibitory to a standardized PCR mixture containing the thermostable DNA polymerase AmpliTaq Gold. PCR inhibitors in human plasma were purified by chromatographic procedures and were characterized by a process of elimination, so that the PCR-inhibitory effects of plasma fractions were tested after each purification step. The major inhibitor in human plasma, as determined by size-exclusion chromatography, anion-exchange chromatography, and chromatofocusing, was found to be immunoglobulin G (IgG) on the basis of N-terminal amino acid sequencing and electrophoretic analysis of the purified polypeptide. When different concentrations of purified plasma IgG (PIgG) were added to PCR mixtures containing 11 different thermostable DNA polymerases and 1 ng of Listeria monocytogenes DNA as template DNA, the only polymerase that resisted inhibition was rTth. The inhibitory effect was reduced when PIgG was heated at 95°C before it was added to PCR or after the addition of excess nontarget DNA to the PCR mixture. However, heating of PIgG together with target DNA at 95°C was found to block the amplification. Inhibition by PIgG may be due to an interaction with single-stranded DNA, which makes the target DNA unavailable for 10 of the DNA polymerases tested. The results show the danger of using boiling as a method of sample pretreatment or using a hot start prior to PCR. The effect of plasma PCR inhibition could be removed by mixing plasma with DNA-agarose beads prior to amplification, while plasma PCR inhibitors were found to bind to the DNA-agarose beads.
Blood samples are extensively used for PCR-based diagnosis of microbial infections and genetic diseases (4). However, the usefulness of diagnostic PCR is limited, in part, by the presence of several factors in blood that reduce the amplification efficiency, such as heme (3), anticoagulants like EDTA (26) and heparin (13), and high concentrations of leukocyte DNA and other unknown inhibitors (10). Among the mechanisms by which the inhibitors may act are the following (31): (i) interference with cell lysis, (ii) degradation or capture of the nucleic acids, or (iii) inactivation of the thermostable DNA polymerase. Various sample preparation methods have been developed to remove or to reduce the effects of PCR inhibitors in blood without understanding the mechanisms of inhibition (1, 12, 16, 32). Characterization of PCR inhibitors is an important step in the development of efficient sample preparation methods, which can overcome the effects of PCR inhibitors. It has been noted that the plasma fraction of whole blood is inhibitory to PCR (16), which indicates the presence of inhibitors other than heme in blood. Plasma contains a large variety of possible inhibitors ranging from ions and low-molecular-mass solutes to proteins (19). Forty-nine of 84 plasma polypeptides have been characterized by two-dimensional electrophoresis (6), and most of the undissociated plasma proteins have molecular masses from approximately 40 kDa up to several million daltons (5).
The aim of the study described here was to identify and to characterize the major inhibitor of PCR in human plasma by fractionating plasma by size-exclusion chromatography, anion-exchange chromatography, and chromatofocusing. The amplification inhibition with various fractions was determined in a standardized PCR assay containing the thermostable DNA polymerase AmpliTaq Gold. The effect of the major PCR inhibitor in human plasma on 11 commercial thermostable DNA polymerases was also investigated.
MATERIALS AND METHODS
Template DNA.
DNA of Listeria monocytogenes 167 vet, which was obtained from Swedish Meats R&D, Kävlinge, Sweden, was used as the target DNA in this study. Extraction of DNA was performed in accordance with a standard technique described by Sambrook et al. (27). The technique was modified by the addition of 30 U of mutanolysin (Sigma Chemical Co., St. Louis, Mo.) per ml to the lysis solution. The concentration of DNA was determined spectrophotometrically (27).
PCR assay and incubation conditions.
The volume of the PCR mixture was 25 μl. All the PCR mixtures contained 0.5 μM (each) primers rU8 and LM2 (18, 25), and 0.2 mM (each) deoxyribonucleoside triphosphates. Reaction buffers for the DNA polymerases were as specified by the manufacturers (Table 1). The reaction mixtures were subjected to 30 cycles consisting of heat denaturation at 94°C for 40 s, primer annealing at 53°C for 40 s, and DNA extension at 72°C for 40 s. Finally, the samples were maintained at 72°C for 7 min for the final extension of DNA. These incubation conditions were the same for all amplification reactions except those containing AmpliTaq Gold, since this polymerase requires a hot start (95°C for 10 min). Incubation was carried out in a model 2400 thermal cycler (Perkin-Elmer Cetus, Norwalk, Conn.).
TABLE 1.
Reaction buffers for the DNA polymerases
| DNA polymerase | Reaction buffer components |
|---|---|
| AmpliTaq Gold | 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 2.5 mM MgCl2, and 0.75 U of AmpliTaq Gold DNA polymerase (Perkin-Elmer Cetus) |
| DyNAzyme II | 10 mM Tris-HCl (pH 8.8; 25°C), 1.5 mM MgCl2, 50 mM KCl, 0.1% Triton X-100, and 1 U of DyNAzyme II DNA polymerase (FINNZYMES OY, Riihitontuntie, Espoo, Finland) |
| DyNAzyme EXL | 50 mM Tris-HCl (pH 9.0; 25°C), 2.5 mM MgCl2, 15 mM (NH4)2SO4, 0.1% Triton X-100, and 1 U DyNAzyme EXT DNA polymerase (FINNZYMES OY) |
| Expand high fidelity | 1× Expand H F buffer, 1.5 mM MgCl2, and 1.33 U of Expand H F PCR system (Roche Molecular Biochemicals, Basel, Switzerland) |
| HotTub | 50 mM Tris-HCl (pH 9.0), 20 mM (NH4)2SO4, 2.5 mM MgCl2, and 0.75 U of HotTub DNA polymerase (Amersham Pharmacia Biotech) |
| Pwo | 10 mM Tris-HCl (pH 8.85; 20°C), 25 mM KCl, 5 mM (NH4)2SO4, 1.5 mM MgCl2, and 1.25 U of Pwo DNA polymerase (Roche Molecular Biochemicals) |
| rTth | 10 mM Tris-HCl (pH 8.3), 5% (vol/vol) glycerol, 0.1 M KCl, 0.05% (w/vol) Tween 20, 0.75 mM EGTA ([ethylene glycol-bis (β-aminoethyl ether)]-N, N, N′, N′-tetraacetic acid) 2.5 mM MgCl2, and 1.25 U of rTth DNA polymerase (Perkin-Elmer Cetus) |
| Taq | 10 mM Tris-HCl, 1.5 mM MgCl2, 50 mM KCl (pH 8.3; 20°C), and 0.75 U of Taq DNA polymerase (Roche Molecular Biochemicals) |
| Tfl | 20 mM Tris-acetate (pH 9.0), 10 mM (NH4)2SO4, 75 mM potassium acetate, 0.05% Tween 20, 2.5 mM MgSO4, and 0.5 U of Tfl DNA polymerase (Promega Corporation, Madison, Wis.) |
| Tli | 10 mM Tris-HCl (pH 9.0; 25°C), 0.1% Triton X-100, 50 mM KCl, 2.5 mM MgCl2, and 0.3 U of Tli DNA polymerase (Promega Corporation) |
| Ultma | 10 mM Tris-HCl (pH 8.8; room temperature), 10 mM KCl, 0.002% (vol/vol) Tween 20, 2.5 mM MgCl2, and 0.75 U of Ultma DNA polymerase (Perkin-Elmer Cetus) |
Preparation of blood sample.
The blood sample used was drawn from a healthy person in a quadruple blood bag (CPD; Baxter S.A., Maurpas, France). The bag was centrifuged in a cold centrifuge (Hettich, Tuttlingen, Germany) at 2,810 × g for 9 min. Plasma and platelets were extracted in one bag, and buffy coat and a portion of erythrocytes were extracted in another bag by using the Optipress plasma extractor (Baxter). Adsol was added to the erythrocytes. The plasma bag was recentrifuged at 1,200 × g for 7 min, plasma was extracted into an empty bag, and the concentrated platelets were suspended in 60 ml of plasma. Each blood fraction was poured into sterile, 1.5-ml Eppendorf tubes, flash frozen in liquid nitrogen, and stored at −80°C. The frozen samples were thawed at room temperature before use.
Purification of PCR inhibitors in human plasma by FPLC.
The ability of different plasma fractions to inhibit PCR was evaluated by the addition of 5 μl of the different fractions to PCR mixtures containing 1 ng of L. monocytogenes DNA. The PCR inhibitors were purified by a chromatographic procedure with a fast protein liquid chromatography (FPLC) system (Amersham Pharmacia Biotech, Uppsala, Sweden) containing two model P-500 high-precision pumps, a model LCC-501 plus liquid chromatography controller, three motor valves (one MV-7 and two MV-8), and a model REC 102 recorder. The elution was monitored with a UV-M II control unit (at 280 nm), and fractions were collected with a model FRAC-200 fraction collector. All the buffers and solutions were filtered through 0.2-μm-pore-size AcroCap membrane filters (Gelman Sciences, Ann Arbor, Mich.) and were degassed before use. A Hiload 16/60 Superdex 200 gel filtration prepacked column (Amersham Pharmacia Biotech) was equilibrated with a buffer consisting of 20 mM Tris-HCl and 100 mM NaCl (pH 7.2). The column was calibrated with blue dextran, ferritin, aldolase, ovalbumin, and RNase A (Amersham Pharmacia Biotech). The plasma was thawed at room temperature and was filtered through a 0.2-μm-pore-size Minisart membrane filter (Sartorious, Goettingen, Germany). A sample consisting of 2 ml of plasma was injected into the column. The plasma components were eluted with a buffer consisting of 20 mM Tris-HCl and 100 mM NaCl (pH 7.2) at a flow rate of 1.0 ml/min. The fractions were collected, dialyzed overnight against 20 mM Tris-HCl (pH 8.6) by using dialysis tubing with a cutoff of 12 to 14 kDa (Spectra/Por, Houston, Tex.), and tested for their ability to inhibit the amplification capacity of AmpliTaq Gold. The inhibitory fractions were filtered through a 0.2-μm-pore-size Minisart membrane filter and were injected into a Mono Q HR 5/5 anion-exchange column (Amersham Pharmacia Biotech) and eluted with 20 mM Tris-HCl (pH 8.6) and a sodium chloride gradient (0 to 0.5 M) for 30 min at a flow rate of 1 ml/min. Peak fractions were collected and dialyzed overnight against 20 mM Tris-HCl (pH 8.6) and were tested for their ability to inhibit the amplification capacity of AmpliTaq Gold. Chromatofocusing was performed with a Mono P HR 5/20 column (Amersham Pharmacia Biotech). The starting buffer was 25 mM ethanolamine (pH 9.4; Merck, Darmstadt, Germany); the eluent consisted of 5% (vol/vol) Polybuffer 96 (Amersham Pharmacia Biotech) and 50 mM NaCl (pH 5.5). The inhibitory fractions collected from the Mono Q column were dialyzed overnight against 25 mM ethanolamine (pH 9.4) and were injected into the Mono P column with a 50-ml Superloop (Amersham Pharmacia Biotech). Chromatofocusing was performed at a flow rate of 0.7 ml/min with 40 ml of the Polybuffer-NaCl eluent. The flowthrough was collected and dialyzed overnight against 20 mM Tris-HCl (pH 8.6). This fraction was subsequently concentrated by using the Mono Q column (Amersham Pharmacia Biotech). The inhibitor was eluted with a sodium chloride gradient (0 to 1 M in 20 mM Tris-HCl [pH 8.6]) for 15 min at a flow rate of 1 ml/min. The protein solution was dialyzed overnight against 20 mM Tris-HCl (pH 7.2) and was tested for its ability to inhibit the amplification capacity of AmpliTaq Gold.
SDS-PAGE and N-terminal amino acid sequencing of the major plasma PCR inhibitor.
All the chemicals used for sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) were obtained from Bio-Rad (Hercules, Calif.). The low-molecular-mass standard markers were obtained from Amersham Pharmacia Biotech. The plasma PCR inhibitor was analyzed by discontinuous SDS-PAGE with a 12% polyacrylamide gel, as described by Laemmli (17), with a Mini-gel apparatus (Bio-Rad). The protein bands in the gel were visualized with Coomassie brilliant blue R-250 or were subjected to electroblotting onto polyvinylidene difluoride membranes for N-terminal sequencing, as described by Ausubel et al. (7). The membrane was stained with Coomassie brilliant blue, and a 50-kDa band was excised and sent for N-terminal sequencing. Edman degradation was performed by the Department of Plant Biology at the Swedish University of Agricultural Sciences, Uppsala, Sweden.
Monoclonal antibodies.
The monoclonal immunoglobulins immunoglobulin G1 (IgG1) (MIgG1) and IgG3 (MIgG3), which are specific for cytomegalovirus glycoprotein B, were obtained from the Department of Immunotechnology, Lund University, Lund, Sweden. They were produced in NS0 myeloma cells transfected with plasmids encoding these proteins (11). Briefly, the variable-region genes encoding this specific immunoglobulin (22, 23) were cloned into the pTIF-g1 vector and the pTIF-g3 vector (11) (which encode the MIgG1 and MIgG3 immunoglobulin heavy chains, respectively) and the pAG4622 vector (which encodes the light chain) (9). Following transformation and selection of stable cell-producing lines, antibodies were produced in miniPERM fermentors (Hereaus Instruments GmbH, Hanau, Germany). The antibodies were purified by ammonium sulfate precipitation and chromatography on HiTrap Protein G columns (Amersham Pharmacia Biotech).
Effect of DNA-agarose on the inhibitory effect of human plasma.
A sample consisting of 2 ml of human plasma was thawed at room temperature and was added to 8 ml of DNA-agarose beads (Amersham Pharmacia Biotech) in buffer A (20 mM Tris-HCl [pH 8.3], 5% [vol/vol] glycerol, 50 mM KCl, 1 mM EDTA), and the mixture was shaken slowly for 5 min. The mixture was filtered through a 50-ml Duran glass filter funnel (porosity, 3; Schott Glas, Mainz, Germany) with the aid of a vacuum pump. The DNA-agarose beads were washed with 10 ml of buffer A. The molecules that bound to the DNA-agarose beads were eluted with 10 ml of buffer B (20 mM Tris-HCl [pH 8.3], 5% glycerol, 1 M KCl). Both filtrates were dialyzed overnight against buffer A. After dialysis, the inhibitory effects of both fractions were studied in two independent experiments by adding different concentrations of the fractions to PCR mixtures containing AmpliTaq Gold and 1 ng of L. monocytogenes DNA.
RESULTS
PCR inhibition by different blood fractions.
To evaluate the inhibitory effect of buffy coat, plasma, platelets, and erythrocytes on the amplification capacity of AmpliTaq Gold, different sample concentrations (20, 2, 0.2, and 0.02% [vol/vol]) were added to PCR mixtures containing 1 ng of L. monocytogenes DNA (Table 2). All the blood fractions were found to be highly inhibitory to AmpliTaq Gold, and the concentrations of the fractions had to be reduced to 0.002% (vol/vol) in the PCR mixtures to detect the amplicons in both replicates.
TABLE 2.
Effects of different concentrations of buffy coat, plasma, platelets, and erythrocytes on PCRa
| Concn (%) | PCR resultsb
|
|||
|---|---|---|---|---|
| Buffy coat | Platelet concentrate | Plasma | Erythrocytes | |
| 20 | −, − | −, − | −, − | −, − |
| 2 | −, − | −, − | −, − | −, − |
| 0.2 | −, − | −, − | −, − | −, − |
| 0.02 | −, − | ±, − | ±, − | −, − |
| 0.002 | +, + | +, + | +, + | ±, + |
Percentage (vol/vol) of different blood fractions in the PCR mixtures containing 1 ng of L. monocytogenes DNA and AmpliTaq Gold.
Results of two independent PCRs. Symbols: +, band of high intensity; ±, band of low intensity; −, no band.
Purification of plasma PCR inhibitors.
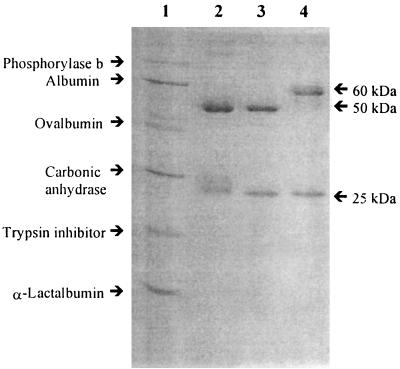
The strategy selected to identify plasma PCR inhibitors was to test the inhibitory effect of the different plasma fractions after each purification step and to continue purification of the fractions inhibitory to AmpliTaq Gold. Size-exclusion chromatography was used to divide the plasma into six main fractions, fractions P1 to P6 (Table 3). Only the fractions containing high-molecular-mass components, fractions P1 to P3, were found to be inhibitory to AmpliTaq Gold. When different dilutions of these three fractions were added to the PCR mixture of AmpliTaq Gold to compare their inhibitory effects, the second fraction (fraction P2) was found to be the most inhibitory and had to be diluted 200 times to overcome its inhibitory effects. When fraction P2 was separated on the Mono Q column, the first fraction was found to be highly inhibitory and had to be diluted 100 times to remove the inhibitory activity. This fraction was injected on a Mono P column, and most of the protein was collected in the flowthrough. The flowthrough which was concentrated by the Mono Q column was found to be highly inhibitory and had to be diluted 100 times to remove the inhibitory activity. SDS-PAGE analysis of the plasma PCR inhibitor showed two protein bands with approximate molecular masses of 25 and 50 kDa (Fig. 1). The N-terminal sequence of the larger band (-EVQLVESGGGLVQPGGSLRL-) showed 100% identity with the N-terminal sequence of the heavy chain of immunoglobulins. The electrophoresis pattern of the purified plasma IgG (PIgG) was compared with those of MIgG1 (150 kDa) and MIgG3 (170 kDa). PIgG had an electrophoretic pattern similar to that of MIgG1 (25 and 50 kDa). Analysis of PIgG, MIgG1, and MIgG3 by SDS-PAGE without boiling and the addition of β-mercaptoethanol to check the purity of the native protein showed that PIgG had only one band with migration similar to that of MIgG1 band (data not shown). To evaluate the inhibitory effects of PIgG, MIgG1, and MIgG2, they were added to PCR mixtures containing AmpliTaq Gold and 1 ng of L. monocytogenes DNA (Fig. 2). The concentrations of PIgG, MIgG1, and MIgG3 had to be reduced to approximately 0.07 to 0.08 μg per reaction tube to relieve the inhibitory effects against AmpliTaq Gold.
TABLE 3.
Inhibitory effects of different size-exclusion chromatography fractions of plasma on PCRa
| Peak no. | Molecular massb (kDa) | PCR resultsc at dilution of:
|
||||
|---|---|---|---|---|---|---|
| Undiluted | 1:10 | 1:50 | 1:100 | 1:200 | ||
| P1 | ∼800 | − | − | + | + | + |
| P2 | ∼200 | − | − | − | − | + |
| P3 | ∼80 | − | − | + | + | + |
| P4 | <13 | + | + | + | + | + |
| P5 | <13 | + | + | + | + | + |
| P6 | <13 | + | + | + | + | + |
Ability of AmpliTaq Gold to amplify 1 ng of L. monocytogenes DNA in the presence of 5 μl of different dilutions of plasma fractions.
Peaks P4, P5, and P6 were outside the range of the size marker.
Symbols: +, band of high intensity; −, no band.
FIG. 1.
SDS-PAGE of PIgG in comparison with SDS-PAGE of MIgG1 and MIgG3. Electrophoretic separation was carried out by SDS-PAGE with 12% polyacrylamide gels. Proteins were detected with Coomassie brilliant blue. Lane 1, low-molecular-mass protein standard (Amersham Pharmacia Biotech); lane 2, PIgG; lane 3, MIgG1; lane 4, MIgG3.
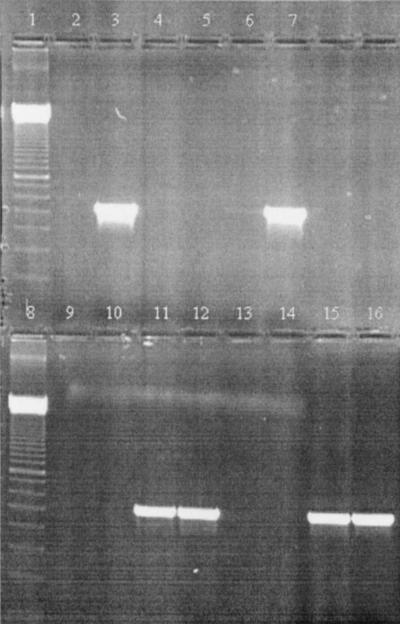
FIG. 2.
Inhibitory effects of different concentrations of PIgG, MIgG1, and MIgG3 on a PCR mixture containing AmpliTaq Gold and 1 ng of L. monocytogenes DNA. Lanes 1 and 8, 100-bp molecular mass marker (Amersham Pharmacia Biotech); lane 2, negative control; lane 3, positive control; lane 4, 0.82 μg of PIgG; lane 5, 0.41 μg of PIgG; lane 6, 0.082 μg of PIgG; lane 7, 0.041 μg of PIgG; lane 9, 0.68 μg of MIgG1; lane 10, 0.34 μg of MIgG1; lane 11, 0.068 μg of MIgG1; lane 12, 0.034 μg of MIgG1; lane 13, 0.77 μg of MIgG3; lane 14, 0.39 μg of MIgG3; lane 15, 0.077 μg of MIgG3; lane 16, 0.039 μg of MIgG3.
Effects of different concentrations of PIgG on the amplification capacities of 11 DNA polymerases.
The capacities of 11 DNA polymerases to amplify 1 ng of L. monocytogenes DNA in pure water and in the presence of different concentrations of PIgG were studied (Table 4). It was found that only rTth DNA polymerase could amplify the specific PCR product in the presence of undiluted PIgG. The bands produced by Tfl DNA polymerase were weak, and no product was detected in the PCR tube containing 0.082 μg of PIgG per reaction tube. Reduction of the amount of PIgG to 0.82 μg per reaction tube allowed amplification by DyNAzyme II DNA polymerase. The enzymes AmpliTaq Gold, DyNAzyme EXL, Expand high fidelity, Taq, and Tli amplified the L. monocytogenes DNA, in both replicates, only when the concentration of PIgG was reduced to 0.041 μg per reaction tube. All PIgG concentrations tested were completely inhibitory to the enzymes HotTub, Pwo, and Ultma.
TABLE 4.
Inhibitory effects of different concentrations of PIgG on the amplification capacities of 11 thermostable DNA polymerasesa
| DNA polymerase | PCR result(s)b with the following IgG concn (μg/reaction tube):
|
|||||
|---|---|---|---|---|---|---|
| 0 | 4.1 | 0.82 | 0.41 | 0.082 | 0.041 | |
| AmpliTaq Gold | +, + | −, − | −, − | −, − | −, ± | ±, + |
| DyNAzyme II | +, + | −, −, − | ±, −, ± | ±, −, ± | +, −, ± | +, +, ± |
| DyNAzyme EXL | +, + | −, − | −, − | −, − | −, − | ±, + |
| Expand high fidelity | + | − | − | − | − | + |
| HotTub | +, + | −, − | −, − | −, − | −, − | −, − |
| Pwo | +, + | −, − | −, − | −, − | −, − | −, − |
| rTth | +, + | +, ±, ± | +, ±, ± | +, ±, ± | +, +, + | +, +, + |
| Taq | +, + | −, − | −, − | −, − | −, − | +, + |
| Tfl | +, + | ±, −, ± | ±, −, ± | ±, ±, − | −, −, − | ±, +, − |
| Tli | +, + | −, − | −, − | −, − | ±, − | +, + |
| Ultma | +, ± | −, − | −, − | −, − | −, − | −, − |
Concentration (wt/vol) of IgG in the PCR mixtures containing 1 ng of L. monocytogenes DNA.
Results of one to three independent PCRs. Symbols: +, band of high intensity; ±, band of low intensity; −, no band.
Effect of PIgG on amplification capacities of AmpliTaq Gold and rTth.
In the absence of PIgG the detection limits for AmpliTaq Gold and rTth were 10 pg of L. monocytogenes DNA for both enzymes (Table 5). The detection limit for AmpliTaq Gold was reduced to 0.1 μg by adding 0.41 μg of PIgG to the reaction mixtures, while the detection limit for rTth DNA polymerase was reduced by 2 log units. Heating of PIgG at 95°C for 10 min before it was added to the reaction mixture of AmpliTaq Gold increased the amplification sensitivity by 2 log units. However, the amplification sensitivities of AmpliTaq Gold and rTth were dramatically reduced to 1 and 0.1 μg, respectively, when PIgG was heated with L. monocytogenes DNA before they were added to the reaction mixtures. Furthermore, the addition of 1.25 ng of bacteriophage λ DNA to the PCR mixtures containing 0.41 μg of PIgG per reaction tube relieved the inhibition, so that the detection limit was the same as that in the absence of PIgG.
TABLE 5.
AmpliTaq Gold and rTth DNA polymerase detection sensitivity in water and in the presence of PIgG and effects of heating of PIgG and L. monocytogenes DNA together and adding bacteriophage λ DNAa
| Amt of L. monocytogenes DNA (g)/reaction tube | PCR resultb
|
|||||||
|---|---|---|---|---|---|---|---|---|
| Water
|
IgGc
|
Heated IgG, AmpliTaqd | Heated IgG and DNAe
|
IgG and λ DNA,f AmpliTaq | ||||
| AmpliTaq | rTth | AmpliTaq | rTth | AmpliTaq | rTth | |||
| 10−6 | + | + | + | + | + | + | + | + |
| 10−7 | + | + | + | + | + | − | + | + |
| 10−8 | + | + | − | + | + | − | − | + |
| 10−9 | + | + | − | ± | ± | − | − | + |
| 10−10 | + | + | − | − | − | − | − | + |
| 10−11 | + | + | − | − | − | − | − | + |
| 10−12 | − | − | − | − | − | − | − | − |
Ability of AmpliTaq Gold and rTth DNA polymerases to amplify different concentrations of L. monocytogenes DNA in water and in the presence of 0.41 μg PIgG and the effects of heating PIgG together with different concentrations of L. monocytogenes DNA and the addition of 1.25 ng of bacteriophage λ DNA on the detection level.
PCR results: +, band of high intensity; ±, band of low intensity; −, no band.
PCR detection in the presence of 0.41 μg of PIgG.
PCR detection in the presence of 0.41 μg of PIgG heated at 95°C for 10 min.
PCR detection in the presence of 0.41 μg of PIgG heated together with L. monocytogenes DNA at 95°C for 10 min.
PCR detection in the presence of 0.41 μg of PIgG and 1.25 ng of bacteriophage λ DNA.
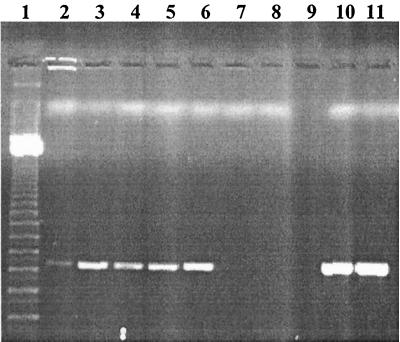
To investigate the mechanism of PCR inhibition by PIgG, human plasma was mixed with single-stranded DNA (ssDNA)–agarose beads prior to amplification (Fig. 3). The plasma filtrate containing the molecules that did not bind to DNA-agarose was found not to be inhibitory to the amplification capacity of AmpliTaq Gold in the presence of 1 ng of L. monocytogenes DNA. In contrast, the plasma components eluted with buffer with high salt concentrations were found to be inhibitory to AmpliTaq Gold, and dilution 50 times was needed to remove the inhibitory activity.
FIG. 3.
Removal of the inhibition of human plasma by DNA-agarose beads. Lane 1, 100-bp molecular mass marker (Amersham Pharmacia Biotech); lanes 2 to 6, different dilutions of human plasma molecules that did not bind to DNA-agarose beads (undiluted and diluted 1:5, 1:10, 1:50, and 1:100, respectively); lanes 7 to 11, different dilutions of human plasma molecules that did bind to DNA-agarose (undiluted and diluted 1:5, 1:10, 1:50, and 1:100, respectively).
DISCUSSION
Blood is highly inhibitory to PCR (1, 24). It has previously been found that the DNA polymerases from Thermus aquaticus, AmpliTaq Gold and Taq DNA polymerases, were totally inhibited in the presence of 0.004% (vol/vol) blood in the PCR mixture (2). This can partially be attributed to the presence of heme and anticoagulants which chelate Mg2+. In the present study, blood fractions without hemoglobin were also found to be highly inhibitory to the PCR. N-terminal sequencing and SDS-PAGE identified PIgG as the major PCR inhibitor in plasma. The inhibitory effect of the cellular fractions may be partially due to PIgG. All thermostable DNA polymerases tested except rTth DNA polymerase were inhibited by PIgG. This explains the ability of rTth DNA polymerases to amplify DNA in the presence of 20% (vol/vol) blood without reduced amplification sensitivity (2). The resistance of rTth DNA polymerase to PCR inhibitors has been noted in other samples, such as aqueous and vitreous fluids of the eye (30), feces (2), and phenol (14). The resistance of rTth to inhibitors has been suggested to be due to the unique structural properties of the Thermus thermophilus DNA polymerase (Tth) and its ability to maintain both DNA- and RNA-dependent DNA polymerase activities (14).
The ability of MIgG1 and MIgG3 to inhibit AmpliTaq Gold suggests that this inhibition is a general effect of immunoglobulins and is not related to specific clones of immunoglobulins. PCR inhibition by PIgG was assumed to be due to its ability to interact with target DNA (ssDNA). This assumption was based on the observations that (i) PCR inhibition was increased when target DNA was heated with PIgG, (ii) the addition of bacteriophage λ DNA reduced PIgG inhibition, (iii) the mixing of plasma with ssDNA-agarose beads reduced the inhibitory effect of plasma on AmpliTaq Gold, and (iv) the analysis of PIgG DNA interaction by mobility shift assay of DNA-binding proteins showed that a PIgG-DNA complex was formed only when PIgG was heated with ssDNA probes at 95°C for 10 min (data not shown).
Immunoglobulins are sensitive to heat denaturation, and in this study it was found that heating of PIgG decreased the inhibition of AmpliTaq Gold. It was also found that heating of PIgG with the DNA template increased the inhibition of the DNA polymerases. This may be due to the formation of a complex between PIgG and the DNA template which blocked the amplification. A possible explanation of why PCR is not inhibited by immunoglobulins when used in one of the “hot-start” strategies for inactivation of the DNA polymerase (15, 29) may be the formation of a complex of immunoglobulins with the DNA polymerase and their lack of freedom to interact with the target DNA. Heating will denature the immunoglobulins, which will subsequently not be able to interact with the target DNA and to inhibit the PCR. Also, these results might explain the nature of a potent PCR inhibitor of Taq DNA polymerase copurified with human genomic DNA, which was not inactivated by boiling of the DNA (10).
In a study by Mantero et al. (21), boiling was used to exclude heat-labile proteins by assuming that the high denaturation temperature in the thermal cycler could prevent interference of these proteins. A hot start (heating of the reaction mixture at 95°C for 10 min) has also been used to avoid undesirable PCR products due to nonspecific annealing and primer elongation. Several reports indicate the interference of the reverse transcription step with PCR amplification in reverse transcription-PCR (8, 20, 28). This interference was removed by increasing the template concentration or including the T4 gene 32 protein during the reverse transcription-PCR step when the reverse transcriptase (RT) was present (8). This inhibitory effect may be similar to the inhibitory effects of immunoglobulins, and heating of RT (thermolabile) with the DNA template at 100°C for 5 min may have formed a complex between RT and the DNA template which blocked amplification by Taq DNA polymerase.
In conclusion, characterization of PCR inhibitors is a necessity for the development of more efficient sample preparation methods, which will allow the PCR detection of pathogens in complex biological samples such as blood. This is the first study in which PCR inhibition by immunoglobulins has been reported. The inhibition was found to take place through an interaction with target ssDNA. The results of this study also showed that heating of reaction mixtures containing immunoglobulins or any protein that can interact with DNA, such as thermolabile RT, is strongly unfavorable.
ACKNOWLEDGMENTS
We are grateful to Mats Ohlin for kindly providing us with the monoclonal immunoglobulins (MIgG1 and MIgG3). We thank Nidal Irsheed for help in drawing blood and separation of the blood into the four fractions. We also thank Jan-Åke Jönsson for valuable discussions.
This work was supported by the Swedish Council for Forestry and Agricultural Research, the Swedish National Board for Industrial and Technical Development, and Fysiografiska Sällskapet in Lund.
REFERENCES
- 1.Abu Al-Soud W, Lantz P-G, Bäckman A, Olcén P, Rådström P. A sample preparation method which facilitates detection of bacteria in blood cultures by the polymerase chain reaction. J Microbiol Methods. 1998;32:217–224. [Google Scholar]
- 2.Abu Al-Soud W, Rådström P. Capacity of nine thermostable DNA polymerases to mediate DNA amplification in the presence of PCR-inhibiting samples. Appl Environ Microbiol. 1998;64:3748–3753. doi: 10.1128/aem.64.10.3748-3753.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Akane A, Matsubara K, Nakamura H, Takahashi S, Kimura K. Identification of the heme compound copurified with deoxyribonucleic acid (DNA) from bloodstrains, a major inhibitor of polymerase chain reaction (PCR) amplification. J Forensic Sci. 1994;39:362–372. [PubMed] [Google Scholar]
- 4.Altwegg M, Verhoef J. Amplification methods in diagnostic microbiology. J Microbiol Methods. 1995;23:3–138. [Google Scholar]
- 5.Anderson L, Anderson N G. High resolution two-dimensional electrophoresis of human plasma proteins. Proc Natl Acad Sci USA. 1977;74:5421–5425. doi: 10.1073/pnas.74.12.5421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Anderson N L, Anderson N G. A two-dimensional gel database of human plasma proteins. Electrophoresis. 1991;12:883–906. doi: 10.1002/elps.1150121108. [DOI] [PubMed] [Google Scholar]
- 7.Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Current protocols in molecular biology. Vol. 2. New York, N.Y: John Wiley & Sons, Inc.; 1989. pp. 10.19.1–10.19.12. [Google Scholar]
- 8.Chandler D P, Wagnon C A, Bolton H., Jr Reverse transcriptase (RT) inhibition of PCR at low concentrations of template and its implications for quantitative RT-PCR. Appl Environ Microbiol. 1998;64:669–677. doi: 10.1128/aem.64.2.669-677.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Coloma M J, Hastings A, Wims L A, Morrison S L. Novel vectors for the expression of antibody molecules using variable regions generated by polymerase chain reaction. J Immunol Methods. 1992;152:89–104. doi: 10.1016/0022-1759(92)90092-8. [DOI] [PubMed] [Google Scholar]
- 10.de Franchis R, Cross N C P, Foulkes N S, Cox T M. A potent inhibitor of Taq polymerase copurifies with human genomic DNA. Nucleic Acids Res. 1988;16:10355. doi: 10.1093/nar/16.21.10355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Furebring C, Ohlin M, Pettersson S, Borrebaeck C A. Evaluation of novel control elements by construction of eukaryotic expression vectors. Gene. 1997;188:191–198. doi: 10.1016/s0378-1119(96)00791-3. [DOI] [PubMed] [Google Scholar]
- 12.Hassan-King M, Baldeh I, Secka O, Falade A, Greenwood B. Detection of Streptococcus pneumoniae DNA in blood cultures by PCR. J Clin Microbiol. 1994;32:1721–1724. doi: 10.1128/jcm.32.7.1721-1724.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Izraeli S, Pfleiderer C, Lion T. Detection of gene expression by PCR amplification of RNA derived from frozen heparinized whole blood. Nucleic Acids Res. 1991;19:6051. doi: 10.1093/nar/19.21.6051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Katcher H L, Schwartz I. A distinctive property of Tth DNA polymerase: enzymatic amplification in the presence of phenol. BioTechniques. 1994;16:84–92. [PubMed] [Google Scholar]
- 15.Kellogg D E, Rybalkin I, Chen S, Mukhamamedova N, Vlasik T, Siebert P D, Chenchik A. TaqStart Antibody™: “hot start” PCR facilitated by neutralizing monoclonal antibody directed against Taq DNA polymerase. BioTechniques. 1994;16:1134–1137. [PubMed] [Google Scholar]
- 16.Klein A, Barsuk R, Dagan S, Nusbaum O, Shouval D, Galun E. Comparison of methods for extraction of nucleic acid from hemolytic serum for PCR amplification of hepatitis B virus DNA sequences. J Clin Microbiol. 1997;35:1897–1899. doi: 10.1128/jcm.35.7.1897-1899.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Laemmli U K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- 18.Lantz P-G, Tjerneld F, Borch E, Hahn-Hägerdal B, Rådström P. Enhanced sensitivity in PCR detection of Listeria monocytogenes in soft cheese through use of an aqueous two-phase system as a sample preparation method. Appl Environ Microbiol. 1994;60:3416–3418. doi: 10.1128/aem.60.9.3416-3418.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lehninger A L, Nelson D L, Cox M M. Principles of biochemistry. 2nd ed. New York, N.Y: Worth Publishers; 1993. p. 745. [Google Scholar]
- 20.Mallet F, Oriol G, Mary C, Verrier B, Mandrand B. Continuous RT-PCR using AMV-RT and Taq DNA polymerase: characterization. BioTechniques. 1995;18:678–687. [PubMed] [Google Scholar]
- 21.Mantero G, Zonaro A, Albertini A, Bertolo P, Primi D. DNA enzyme immunoassay: general method for detecting products of polymerase chain reaction. Clin Chem. 1991;37:422–429. [PubMed] [Google Scholar]
- 22.Ohlin M, Owman H, Mach M, Borrebaeck C A. Light chain shuffling of a high affinity antibody results in a drift in epitope recognition. Mol Immunol. 1996;33:47–56. doi: 10.1016/0161-5890(95)00123-9. [DOI] [PubMed] [Google Scholar]
- 23.Ohlin M, Sundqvist V A, Mach M, Wahren B, Borrebaeck C A. Fine specificity of the human immune response to the major neutralization epitopes expressed on cytomegalovirus gp58/116 (gB), as determined with human monoclonal antibodies. J Virol. 1993;67:703–710. doi: 10.1128/jvi.67.2.703-710.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Panaccio M, Lew A. PCR based diagnosis in the presence of 8% (v/v) blood. Nucleic Acids Res. 1991;19:1151. doi: 10.1093/nar/19.5.1151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rådström P, Bäckman A, Qian N, Kragsbjerg P, Påhlson C, Olcén P. Detection of bacterial DNA in cerebrospinal fluid by an assay for simultaneous detection of Neisseria meningitidis, Haemophilus influenzae, and streptococci using a seminested PCR strategy. J Clin Microbiol. 1994;32:2738–2744. doi: 10.1128/jcm.32.11.2738-2744.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rossen L, Nøskov P, Holmstrøm K, Rasmussen O F. Inhibition of PCR by components of food samples, microbial diagnostic assays and DNA-extraction solution. Int J Food Microbiol. 1992;17:37–45. doi: 10.1016/0168-1605(92)90017-w. [DOI] [PubMed] [Google Scholar]
- 27.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. pp. E3–E5. [Google Scholar]
- 28.Sellner L N, Coelen R J, Mackenzie J S. Reverse transcriptase inhibits Taq polymerase activity. Nucleic Acids Res. 1992;20:1487–1490. doi: 10.1093/nar/20.7.1487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Westfall B, Sitaraman K, Solus J, Hughes J, Rashtchian A. Improved PCR specificity and yield with platinum™ TAQ DNA polymerase. BioTechniques. 1997;19:46–48. [Google Scholar]
- 30.Wiedbrauk D L, Werner J C, Drevon A M. Inhibition of PCR by aqueous and vitreous fluids. J Clin Microbiol. 1995;33:2643–2646. doi: 10.1128/jcm.33.10.2643-2646.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wilson I G. Inhibition and facilitation of nucleic acid amplification. Appl Environ Microbiol. 1997;63:3741–3751. doi: 10.1128/aem.63.10.3741-3751.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhang Y, Isaacman D, Wadowsky R, Rydquist-White J, Post J, Ehrlich G. Detection of Streptococcus pneumoniae in whole blood by PCR. J Clin Microbiol. 1995;33:596–601. doi: 10.1128/jcm.33.3.596-601.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]