Abstract
Both sepsis and acute respiratory distress syndrome (ARDS) rely on imprecise clinical definitions leading to heterogeneity, which has contributed to negative trials. Because circulating protein/DNA complexes have been implicated in sepsis and ARDS, we aimed to develop a proteomic signature of DNA-bound proteins to discriminate between children with sepsis with and without ARDS. We performed a prospective case-control study in 12 children with sepsis with ARDS matched to 12 children with sepsis without ARDS on age, severity of illness score, and source of infection. We performed co-immunoprecipitation and downstream proteomics in plasma collected ≤ 24 h of intensive care unit admission. Expression profiles were generated, and a random forest classifier was used on differentially expressed proteins to develop a signature which discriminated ARDS. The classifier was tested in six independent blinded samples. Neutrophil and nucleosome proteins were over-represented in ARDS, including two S100A proteins, superoxide dismutase (SOD), and three histones. Random forest produced a 10-protein signature that accurately discriminated between children with sepsis with and without ARDS. This classifier perfectly assigned six independent blinded samples as having ARDS or not. We validated higher expression of the most informative discriminating protein, galectin-3-binding protein, in children with ARDS. Our methodology has applicability to isolation of DNA-bound proteins from plasma. Our results support the premise of a molecular definition of ARDS, and give preliminary insight into why some children with sepsis, but not others, develop ARDS.
Keywords: ARDS, damage-associated molecular patterns, DAMPs, histones, PARDS
INTRODUCTION
How diverse inciting insults can result in acute respiratory distress syndrome (ARDS) (1) remains unknown. Sepsis, defined as organ dysfunction caused by a pathologic host response to an infection (2), is the most common etiology of ARDS in both adults (3) and children (4). However, both sepsis and ARDS are clinical definitions, which lead to misclassification, imprecision, and heterogeneity, all of which have contributed to negative trials. More precise definitions for ARDS may reduce heterogeneity, assist with trial selection, and allow insight into mechanisms elucidating why some patients with sepsis develop ARDS, while others do not.
Plasma biomarkers have been proposed to more precisely define ARDS (5, 6). Studies in adults have implicated nucleosomes, the histone/DNA complexes resulting from nuclear chromatin degradation released after cellular damage, as pathogenic in sepsis (7–9), aspiration (10), and trauma-related ARDS (11). Recently, we demonstrated nucleosomes were elevated in pediatric ARDS compared with controls without ARDS (12, 13). Based on these results, and given that other nonhistone protein/DNA complexes have been implicated in cancer (14), we reasoned that a systematic analysis of plasma DNA-bound proteins in children with sepsis could potentially differentiate those with and without ARDS. We additionally posited that identification of DNA-bound proteins in ARDS may provide insight into the pathogenesis of lung injury in sepsis, with a specific hypothesis that neutrophil-derived proteins from programmed neutrophil death would be implicated.
METHODS
Study Design and Patient Selection
We aimed to compare DNA-bound proteins from subjects with and without ARDS after co-immunoprecipitation (co-IP) with anti-ds DNA antibody as bait, with the resulting elution subject to liquid chromatography tandem mass spectrometry (LC-MS/MS). We performed a case-control study nested within a cohort study of ARDS at the Children’s Hospital of Philadelphia (CHOP), approved by the Institutional Review Board (CHOP IRB 13-010578), with written or verbal informed consent obtained before enrollment (13, 15). Consecutive patients were screened for (Berlin) ARDS (1) between July 2014 and June 2019: 1) acute (≤7 days of risk factor) respiratory failure requiring invasive ventilation, 2) arterial access, 3) 1 mo < age < 18 yr, 4) two consecutive / ≤ 300 separated by ≥1 h on positive end-expiratory pressure ≥ 5 cmH2O, and 5) bilateral infiltrates. Exclusion criteria were as follows: 1) respiratory failure from cardiac failure, 2) exacerbation of chronic lung disease, 3) ventilator dependence, 4) cyanotic heart disease, 5) ventilation for >7 days before / ≤ 300, or 6) ARDS established outside of CHOP.
For comparison, we enrolled 25 intubated children with ARDS risk factors screened for the ARDS cohort, but not meeting oxygenation (/ > 300) and radiographic criteria. From this group, we identified 12 subjects meeting Sepsis-3 criteria (2) and matched these 12 children with sepsis without ARDS to 12 subjects with ARDS based on age categories (≤2 yr, 2–6 yr, 6–12 yr, 12–15 yr, 15 to <18 yr), Pediatric Risk of Mortality (PRISM) III score (±2), and pulmonary versus nonpulmonary source of sepsis. We ensured that the 12 subjects with ARDS met Sepsis-3 criteria by confirming organ dysfunction and lactate levels.
Blood Collection and Co-Immunoprecipitation
For the parent ARDS cohort study, we collected blood ≤24 h of ARDS onset. For this study, to harmonize with subjects with non-ARDS sepsis, we ensured that blood for all subjects (with and without ARDS) was collected ≤24 h of pediatric intensive care unit (PICU) admission in citrated tubes (Becton, Dickinson; Franklin Lakes, NJ), centrifuged within 30 min (2,000 g, 20 min, 20°C) to generate platelet-poor plasma, and stored at −80°C until analysis. For co-IP, 100 µL of plasma was precleared with Protein A/G Plus Agarose (Pierce, Thermo Fisher Scientific, Waltham, MA) and underwent co-IP (Pierce) using anti-ds DNA IgG2A (Abcam, Cambridge, MA). Anti-ds DNA antibody was covalently bound to an amine-reactive resin, making the elution enriched for target DNA-associated proteins, with minimal contamination by antibody. This protocol generated ∼1 µg of protein per sample, quantified using Nanodrop.
In-Solution Digestion
Samples were precipitated using acetone/trichloroacetic acid (TCA) (8 vol/1 vol). The pellet was solubilized and digested with the iST kit (PreOmics GmbH, Martinsried, Germany), reduced, and alkylated by addition of sodium deoxycholate (SDC) buffer containing tris(2-carboxyethyl)phosphine (TCEP) and 2-chloroacetamide, heated to 95°C for 10 min. Proteins were hydrolyzed for 1.5 h at 37°C by adding LysC and trypsin. Resulting peptides were desalted, dried by vacuum centrifugation, and reconstituted in 0.1% trifluoroacetic acid (TFA) containing iRT peptides (Biognosys, Schlieren, Switzerland).
Mass Spectrometry Data Acquisition, Spectral Library Generation, and Protein Quantification
Samples were analyzed on a QExactive HF mass spectrometer (Thermo Fisher Scientific) coupled with an Ultimate 3000 nano UPLC system and EasySpray source, using data-independent acquisition (DIA) and data-dependent acquisition (DDA). Tryptic digests were spiked with iRT standards (Biognosys) and separated by reverse-phase high-purification liquid chromatography (RP-HPLC) on a nanocapillary column, 75 μm id × 50 cm 2 μm PepMap RSLC C18 column at 50°C. Mobile phase A consisted of 0.1% formic acid and mobile phase B of 0.1% formic acid/acetonitrile. Peptides were eluted into the mass spectrometer at 210 nL/min with each RP-HPLC run comprising a 125-min gradient from 1% to 5% B in 15 min, 5%–45% B in 110 min for DDA, and 140 min for DIA. For DDA, the mass spectrometer was set to repetitively scan m/z from 300 to 1,400 (R = 240,000) followed by data-dependent tandem mass spectrometry (MS/MS) scans on the 20 most abundant ions, minimum automatic gain control (AGC) 1⋅e4, dynamic exclusion with a repeat count of 1, repeat duration of 30 s (R = 15,000). Fourier transform mass spectrometry (FTMS) full scan AGC target value was 3⋅e6, while MSn AGC was 1⋅e5, respectively. MSn injection time was 160 ms; microscans were set at 1. Rejection of unassigned and 1+,6–8 charge states was set. The raw files for DIA were collected at one full MS scan at 120,000 resolution, a scan range of 300–1,650 m/z, an AGC target of 3⋅e6, and a maximum inject time of 60 ms. This was followed by 22 (DIA) isolation windows with varying sizes at 30,000 resolution, an AGC target of 3⋅e6, injection times set to auto, loop count, and msx count of 1. Default charge state was 4, first mass fixed at 200 m/z, and normalized collision energy (NCE) per window stepped at 25.5, 27, and 30.
MS/MS raw files for the DDA were searched against a reference human protein sequence database including reviewed isoforms from the Uniprot database using MaxQuant v.1.6.1.0 (16). Trypsin was specified as enzyme with two possible missed cleavages. Carbamidomethyl of cysteine was specified as fixed modification and protein N-terminal acetylation and oxidation of methionine were considered variable modifications. The MS/MS tolerance FTMS was set to 20 ppm. A false discovery rate of 1% was set for peptide and protein identification. Remaining parameters were set to default. MaxQuant output was used to generate the project-specific spectral library for DIA. This library was supplemented with additional library generated using direct DIA approach in Spectronaut 13 (Biognosys) (17).
Data Analysis
Perseus (v.1.6.6.0) and Spectronaut 13 (Biognosys) were used for proteomic processing and analysis (18). Data were log2 transformed and normalized by subtracting the median. We filtered to have eight values in at least one (with or without ARDS) grouping. Subsequently, data were processed in R v.3.6.2. Data were imputed using bpca implemented in pcaMethods package v.1.78.0 (Bioconductor, 3.10) (19) and batch effects were removed using the ComBAT method in R sva v.3.32.1 (20). To compare data between the groups, Student’s t test was used to identify differentially expressed proteins, and volcano plots were generated using EnhanceVolcano v.1.2 (Bioconductor, release 3.10) to visualize the affected proteins while comparing different groups. Lists of differentially abundant proteins were then sorted based on the P value < 0.05 and |log2FC| > 1, yielding a prioritized list, from which we selected the top 50 for Gene Ontology (GO) Biological Functional enrichment and the top 20 for predictive analyses. A single ARDS sample was excluded as an extreme outlier at this stage. By examining Pearson correlations for the top 20 proteins, we removed one of each pair of correlated proteins with R > 0.90 (based on their normalized intensities), reducing the significant set of proteins to 17.
Next, using these 17 proteins, we wished to test them together to see how well they could predict the clinical state (sample class) of ARDS versus non-ARDS. We used a random forests-based classification algorithm from the R package “random forests” v.4.6-14. Random forests are a machine learning algorithm that uses an ensemble of randomized decision trees based on a subset of chosen input features (here, proteins) and predicts sample class (ARDS or non-ARDS) from the majority vote (21). Another advantage of random forests is that the features (proteins) that are most important in making a classification decision can be identified. Here, we were interested in which sets of these 17 proteins were most useful in predicting whether a sample should be classified as ARDS or not. Using the selected features, the random forests-based model was run to identify those proteins that correctly predicted the sample class (ARDS or non-ARDS). Our benchmark was to try different random combination of these top proteins until the prediction (out of bag) error was less than 0.1. We tested accuracy of the predictive model using an unbiased approach in an independent cohort (n = 6) double-blinded by a clinical coordinator. Finally, we assessed differences of the most informative protein, galectin-3-binding protein (Gal-3BP), between children with sepsis with and without ARDS using enzyme-linked immunosorbent assay (ELISA) (R & D Systems, Minneapolis, MN).
RESULTS
Description of the Cases and Control Subjects
Twelve children with sepsis and ARDS were matched to 12 children with sepsis without ARDS on age, PRISM III severity of illness score, and pulmonary versus nonpulmonary source of sepsis (Table 1). Subjects with and without ARDS were well-matched on severity of illness and sepsis etiology. There was no significant difference in mortality between children with sepsis with and without ARDS. Of the five ARDS nonsurvivors, two died of refractory hypoxemia and three of multisystem organ failure; the sole septic non-ARDS nonsurvivor died of multisystem organ failure.
Table 1.
Demographics of the matched cohort
| Variable | No ARDS (n = 12) | ARDS (n = 12) | P Value |
|---|---|---|---|
| Age, yr | 10.9 [4.6, 17.3] | 11.3 [4.4, 14.7] | 0.544 |
| Female, % | 4 (33) | 7 (58) | 0.414 |
| Pediatric Risk of Mortality III | 11 [9, 15] | 11.5 [9, 15.5] | 0.547 |
| Vasopressor score | 14 [6, 19] | 13 [4, 38] | 0.424 |
| Lactate, mmol/L | 4.5 [2.7, 5.2] | 2.4 [2.2, 3.7] | 0.140 |
| Sepsis etiology, % | 0.883 | ||
| Pulmonary | 2 (17) | 2 (17) | |
| Blood | 4 (33) | 6 (50) | |
| Abdomen | 4 (33) | 3 (25) | |
| Genitourinary/renal | 1 (8) | 0 | |
| Skin/soft tissue | 0 | 1 (8) | |
| Central nervous system | 1 (8) | 0 | |
| Immunocompromised, % | 5 (42) | 6 (50) | 1 |
| Oxygenation | |||
| / | 366 [349, 467] | 223 [153, 277] | 0.012 |
| Oxygenation index | 3.1 [1.6, 3.8] | 8.1 [5.3, 12] | <0.001 |
| Berlin ARDS severity, % | |||
| Mild | 6 (50) | ||
| Moderate | 4 (33) | ||
| Severe | 2 (17) | ||
| Nonpulmonary organ failures, % | |||
| Neurologic | 0 | 0 | 1 |
| Cardiovascular | 12 (100) | 12 (100) | 1 |
| Liver | 3 (25) | 6 (50) | 0.400 |
| Kidney | 5 (42) | 7 (58) | 0.684 |
| Hematologic | 5 (42) | 6 (50) | 1 |
| PICU mortality, % | 1 (8) | 5 (42) | 0.059 |
ARDS, acute respiratory distress syndrome; PICU, pediatric intensive care unit.
Co-Immunoprecipitation and Differential Protein Expression
We compared DNA-bound proteins from subjects with and without ARDS after co-IP and LC-MS/MS. Differentially expressed DNA-bound proteins (Fig. 1) segregated using unsupervised hierarchical clustering. Differentially abundant proteins were sorted based on P value < 0.05 and |log2FC| > 1, yielding a prioritized list, from which we selected the top 50 for enrichment analysis. Neutrophil and nucleosome proteins were over-represented in ARDS samples, including two S100A proteins, superoxide dismutase (SOD), and three histones. Gene Ontology (GO) enrichment analysis (Supplemental Fig. S1; all Supplemental Material is available at https://doi.org/10.6084/m9.figshare.17159387.v1) demonstrated signals for leukocyte migration, proteasome, and neutrophil activation. GO Cellular Component analysis suggested most proteins were extracellular and cytosolic, rather than nuclear (Supplemental Fig. S2), which was less consistent with nucleosome origins.
Figure 1.
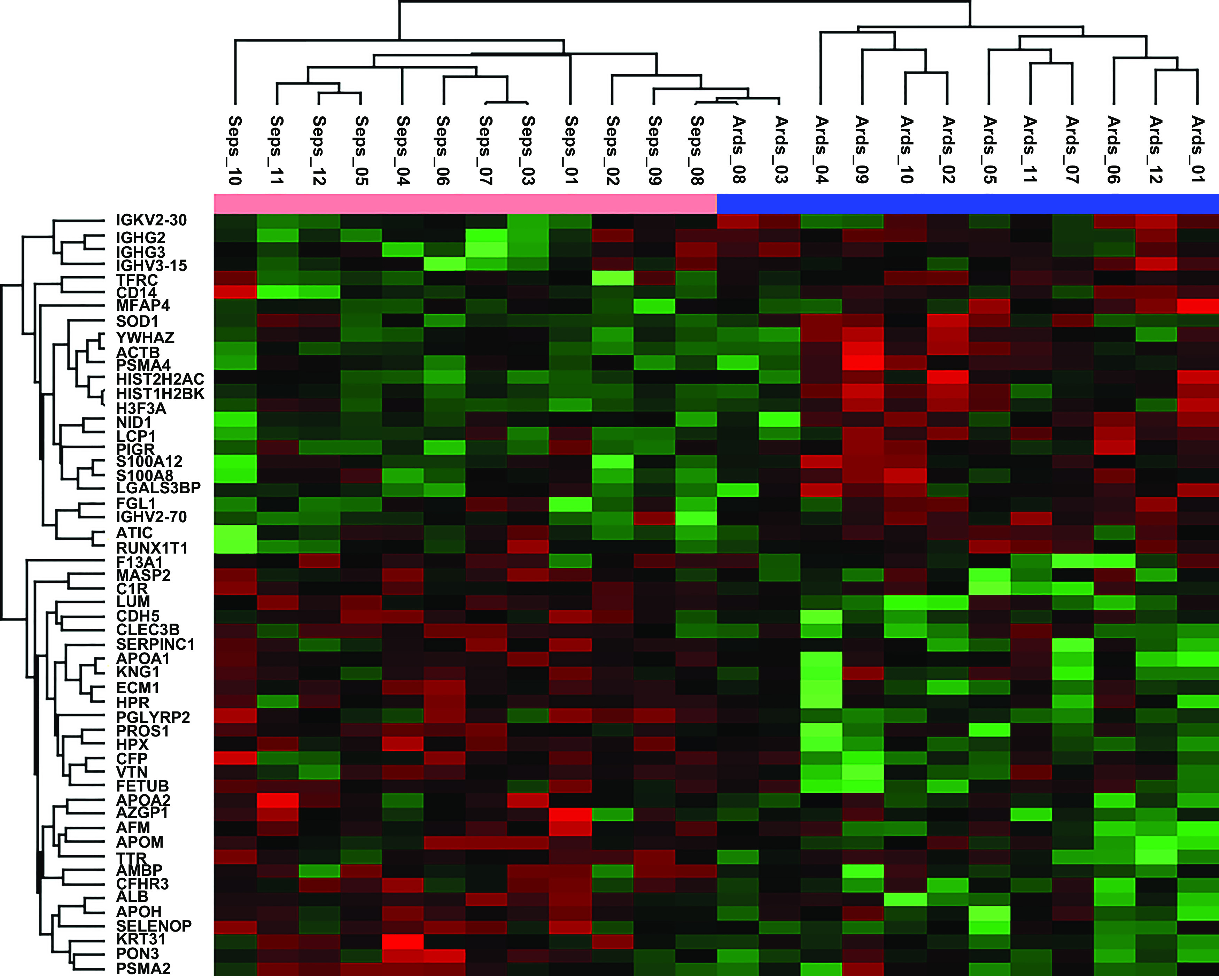
Unsupervised hierarchical clustering of the top 50 differentially expressed DNA-bound proteins between children with sepsis with (blue along top axis) and without (pink along top axis) acute respiratory distress syndrome (ARDS). Red (upregulated) and green (downregulated) shadings represent increasing and decreasing fold change.
ARDS Protein Signature
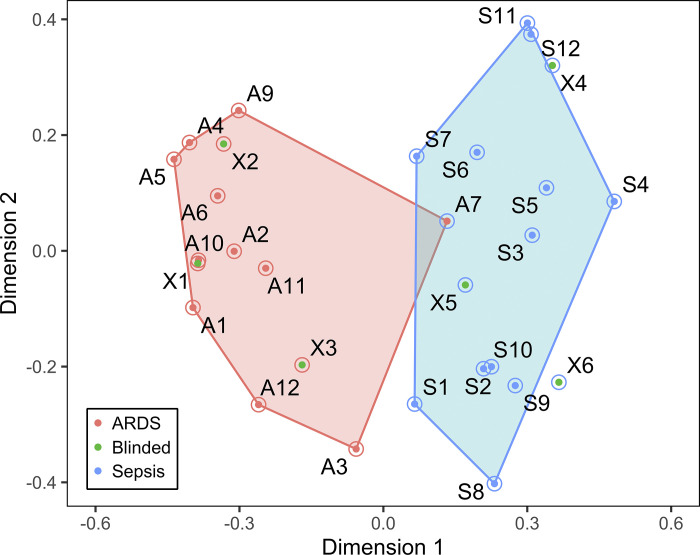
Next, we sought to develop a minimal protein signature to predict ARDS. Proteins were sorted according to relative predictor importance in their ability to discriminate the presence of ARDS (Supplemental Fig. S3). Filtering based on P value < 0.05 and |log2-fold change| > 1 reduced the list to 20 potential predictors. Removal of highly correlated (r > 0.90) proteins resulted in 17 potential predictors. A single ARDS sample with a genetic syndrome, moderate ARDS, and nonpulmonary sepsis was excluded as an extreme outlier to improve performance. Using these 17 proteins, we used a random forest classifier to test which subset most efficiently discriminated subjects with and without ARDS. Our final list for the predictive model consisted of 10 proteins (Fig. 2 and Supplemental Fig. S4). The model demonstrated a conservative (unadjusted for matching variables) area under the receiver operating characteristic curve of 0.96 (95% confidence interval, 0.88–1), a sensitivity of 0.92, and a specificity of 1. This model was tested against six new samples (3 children with and 3 without ARDS), which were coded and re-coded for double-blinding before co-IP, LC-MS/MS, and analysis. The classifier demonstrated perfect assignment of these six samples (Fig. 2).
Figure 2.
Proximity plot of first two dimensions of random forest classifier-derived top 10 protein predictor, and distribution of acute respiratory distress syndrome (ARDS; pink) and non-ARDS (blue) subjects from derivation set. A single ARDS subject is misclassified. The independent samples (n = 6, labeled X) are also shown.
Galectin-3-Binding Protein Discriminates ARDS in Children with Sepsis
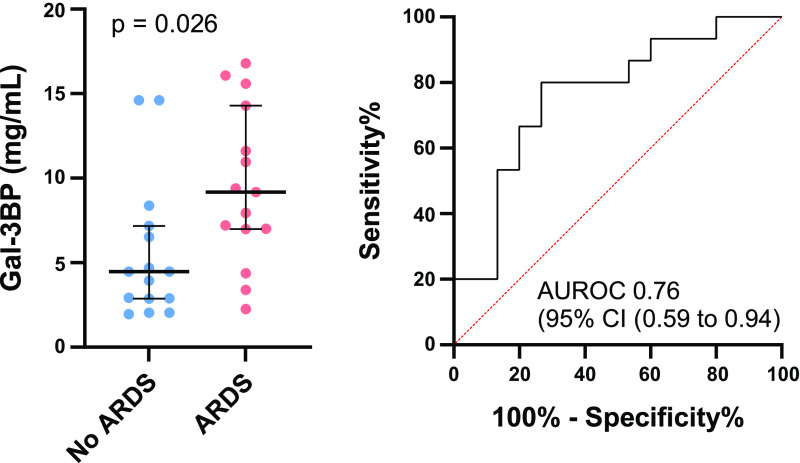
Galectin-3-binding protein (Gal-3BP) was the most informative protein in the random forest analysis for discriminating the presence or absence of ARDS (Supplemental Fig. S3). We tested the ability of this protein alone to discriminate ARDS in children with sepsis (Fig. 3). Gal-3BP was higher in children with sepsis with ARDS (P = 0.026), relative to without, and discriminated the presence of ARDS with an area under the receiving operator characteristic curve of 0.76 (95% confidence interval, 0.59–0.94).
Figure 3.
Differences in galectin-3-binding protein (Gal-3BP) between children with sepsis with and without acute respiratory distress syndrome (ARDS). Gal-3BP modestly discriminates the presence of ARDS, with an area under the receiver operating characteristic (AUROC) curve of 0.76 (95% CI, 0.59–0.94). CI, confidence interval.
DISCUSSION
Using a combination of proteomics and informatics, we developed an ARDS signature for children with sepsis composed of circulating DNA-bound proteins, with good performance in a small independent test cohort. A simple ELISA of Gal-3BP, the most informative protein in the random forest classifier, demonstrated modest ability to discriminate ARDS. As neutrophil-derived proteins were enriched in ARDS, further investigation of these differentially expressed proteins may provide insight into the role of neutrophil death in the pathogenesis of ARDS.
The over-representation of neutrophil proteins and histones in the 17 predictor proteins support neutrophil extracellular traps (NETs) as a potential source for elevated DNA-bound proteins in ARDS, suggesting a role for NET degradation in the pathogenesis of ARDS in sepsis. Histones have been implicated as causal for both sepsis (7–9, 22) and ARDS (10, 11, 23). In mouse models, circulating histones were more toxic to lungs than other organs (11). Our results also support our earlier work showing elevated nucleosomes in children with ARDS relative to intubated children without ARDS (13).
The two S100 proteins enriched in ARDS, S100A8 and S100A12, have also been implicated in lung disease (24). S100A8 (25) and S100A12 (26) both promote neutrophil and monocyte chemotaxis, consistent with the heightened innate immune response to the lungs characterizing early ARDS. Both S100A proteins, like histones, function as damage-associated molecular patterns (DAMPs), as they are themselves released from neutrophils and monocytes in response to infection. S100A12 can be induced by interleukin-6 (27), which is also elevated in ARDS (6, 28). Overall, the protein signature enriched in ARDS strongly implicates neutrophil turnover as a key pathogenic mechanism contributing to ARDS pathogenesis, potentially via generation of DAMPs causing additional inflammation and tissue damage.
However, it is unclear whether the proteins comprising the signature are released from cells already bound to DNA, or whether they are freely circulating and encounter cell-free DNA during sepsis (29). Cellular component analysis suggested that most proteins higher in ARDS were extracellular or cytosolic, not nuclear, which does not support a predominantly nucleosomal or nuclear signature. Multiple types of regulated cell death have been implicated in ARDS other than NETosis, including necroptosis (30), pyroptosis (31, 32), ferroptosis (33), and classic apoptosis (34, 35). Unregulated cell death from necrosis has also been implicated in ARDS (36), and is highly inflammatory. Thus, given the multiple types of cell death from diverse cells in both sepsis and ARDS, firm conclusions cannot be made regarding the origins of the DNA-bound proteins enriched in the patients with ARDS. However, the ability of the protein signature to discriminate ARDS has utility even without precise delineation of the source of the proteins.
Gal-3BP was the most informative protein for discriminating ARDS in the random forest. Gal-3BP is a secreted glycoprotein of the scavenger receptor cysteine-rich domain family that binds to galectins 1, 3, and 7, originally described in the secretome of several cancer cells (37). High Gal-3BP levels have been found in a myriad of viral and bacterial infections (38, 39). Mice deficient in Gal-3BP are more susceptible to LPS, with higher levels of proinflammatory cytokines (40, 41). Overall, Gal-3BP appears to be induced by microbial infections and has a role in regulating innate immunity. A recent study found that circulating Gal-3BP levels were correlated with greater Covid-19 severity (42), consistent with our results of Gal-3BP as a novel marker for discriminating children with sepsis with ARDS from those without ARDS. Interestingly, the polymeric immunoglobulin receptor (PIGR), a transmembrane protein implicated in epithelial cell immune complex transcytosis, was also elevated in subjects with ARDS. PIGR has also been associated with Covid-19 severity, with higher levels seen in actively viremic patients, correlated with S100A8 levels (43). Detection of elevated Gal-3BP in our study, and of PIGR in other studies, also raises the possibility that proteins we identified as elevated in ARDS do not necessarily need to undergo a co-IP step for detection.
To our knowledge, this is the first use of anti-ds DNA co-IP for enrichment of DNA-bound proteins, which may have utility for characterization of protein/DNA complexes in other inflammatory conditions. Clinically, sepsis and ARDS demonstrate overlap in presentation and biomarker profiles (44–46). However, a protein signature for ARDS may assist with more precise diagnosis and allow insight into the pathogenesis of ARDS. Given differences in co-morbidities, epidemiology, and outcomes between pediatric ARDS and adults (4), it is unknown how well this signature would perform in adults with sepsis, and validation in additional pediatric and adult septic population with and without ARDS is necessary.
The heterogeneity and imprecision of the definition of ARDS has contributed to the paucity of therapies. Although preliminary, our data suggest the potential for a more precise molecular definition for ARDS. Understanding the proteomic signature of ARDS would permit greater understanding of the underlying pathophysiology and potentially permit treatments targeted at shared underlying pathways. It is notable that we found a distinct protein signature despite half of subjects having mild ARDS according to Berlin criteria. Hypoxemia only weakly correlated with mortality risk in both adult (1) and pediatric (4) ARDS, particularly at onset, and it is possible that our proteomic signature was capturing underlying physiology more precisely than a single blood-gas measurement. Larger sample sizes will be necessary to assess the generalizability and utility of these proteins for identifying ARDS and for assessing whether they correlate with severity categories.
Our study has limitations. Both derivation and test cohorts were small and from a single center, raising generalizability concerns. However, demographics, severity of illness, and outcomes were similar to other cohorts (4, 47, 48). The co-IP and proteomic processing are cumbersome and impractical for a realistic enrichment strategy; however, identification of a subset of proteins, such as Gal-3BP, raises the potential for development of a multiplex of the most discriminative proteins with rapid turnaround. Additional work is necessary to determine whether the DNA-bound nature of these proteins is a necessary component of this signature, or whether a simple multiplex can discriminate ARDS without an anti-ds DNA co-IP. Both subjects with and without ARDS met criteria for sepsis, meaning that our signature may not extrapolate to noninfectious ARDS. Given that >70% of pediatric ARDS is caused by either pneumonia or nonpulmonary sepsis (4, 49), our signature should prove useful for most cases of ARDS. Overall, our results provide the beginnings of a molecular definition of ARDS, and give preliminary insight into why some children with sepsis, but not others, develop ARDS.
SUPPLEMENTAL DATA
Supplemental Figs. S1–S4: https://doi.org/10.6084/m9.figshare.17159387.v1.
GRANTS
This work was supported by National Institutes of Health Grants K23 HL136688 (to N.Y.) and R01 HL148054 (to N.Y.).
DISCLOSURES
N. Yehya’s institution receives funding from Pfizer outside of the scope of this manuscript. None of the other authors has any conflicts of interest, financial or otherwise, to disclose.
AUTHOR CONTRIBUTIONS
N.Y., S.S.M., S.H.S., and G.S.W. conceived and designed research; N.Y., H.F., G.G.L., L.A.S., J.M.T., and S.H.S. performed experiments; N.Y., H.F., D.M.T., G.G.L., L.A.S., J.M.T., and S.H.S. analyzed data; N.Y., H.F., D.M.T., G.G.L., and L.A.S. interpreted results of experiments; N.Y., H.F., and D.M.T. prepared figures; N.Y. drafted manuscript; N.Y., H.F., D.M.T., G.G.L., L.A.S., J.M.T., S.S.M., S.H.S., and G.S.W. edited and revised manuscript; N.Y., H.F., D.M.T., G.G.L., L.A.S., J.M.T., S.S.M., S.H.S., and G.S.W. approved final version of manuscript.
REFERENCES
- 1.Force ADT, Ranieri VM, Rubenfeld GD, Thompson BT, Ferguson ND, Caldwell E, Fan E, Camporota L, Slutsky AS; ARDS Definition Task Force. Acute respiratory distress syndrome: the Berlin Definition. JAMA 307: 2526–2533, 2012. doi: 10.1001/jama.2012.5669. [DOI] [PubMed] [Google Scholar]
- 2.Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, Bellomo R, Bernard GR, Chiche JD, Coopersmith CM, Hotchkiss RS, Levy MM, Marshall JC, Martin GS, Opal SM, Rubenfeld GD, van der Poll T, Vincent JL, Angus DC. The Third International Consensus Definitions for sepsis and septic shock (sepsis-3). JAMA 315: 801–810, 2016. doi: 10.1001/jama.2016.0287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bellani G, Laffey JG, Pham T, Fan E, Brochard L, Esteban A, Gattinoni L, van Haren F, Larsson A, McAuley DF, Ranieri M, Rubenfeld G, Thompson BT, Wrigge H, Slutsky AS, Pesenti A, Investigators LS, Group ET; ESICM Trials Group. Epidemiology, patterns of care, and mortality for patients with acute respiratory distress syndrome in intensive care units in 50 countries. JAMA 315: 788–800, 2016. doi: 10.1001/jama.2016.0291. [DOI] [PubMed] [Google Scholar]
- 4.Khemani RG, Smith L, Lopez-Fernandez YM, Kwok J, Morzov R, Klein MJ, Yehya N, Willson D, Kneyber MCJ, Lillie J, Fernandez A, Newth CJL, Jouvet P, Thomas NJ, Pediatric Acute Respiratory Distress Syndrome I, Epidemiology I, Pediatric Acute Lung I, Sepsis Investigators N; Pediatric Acute Lung Injury and Sepsis Investigators (PALISI) Network. Paediatric acute respiratory distress syndrome incidence and epidemiology (PARDIE): an international, observational study. Lancet Respir Med 7: 115–128, 2019. doi: 10.1016/S2213-2600(18)30344-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Calfee CS, Delucchi K, Parsons PE, Thompson BT, Ware LB, Matthay MA, Network NA. Subphenotypes in acute respiratory distress syndrome: latent class analysis of data from two randomised controlled trials. Lancet Respir Med 2: 611–620, 2014. doi: 10.1016/S2213-2600(14)70097-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Calfee CS, Janz DR, Bernard GR, May AK, Kangelaris KN, Matthay MA, Ware LB. Distinct molecular phenotypes of direct vs indirect ARDS in single-center and multicenter studies. Chest 147: 1539–1548, 2015. doi: 10.1378/chest.14-2454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Xu J, Zhang X, Pelayo R, Monestier M, Ammollo CT, Semeraro F, Taylor FB, Esmon NL, Lupu F, Esmon CT. Extracellular histones are major mediators of death in sepsis. Nat Med 15: 1318–1321, 2009. doi: 10.1038/nm.2053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Alhamdi Y, Abrams ST, Cheng Z, Jing S, Su D, Liu Z, Lane S, Welters I, Wang G, Toh CH. Circulating histones are major mediators of cardiac injury in patients with sepsis. Crit Care Med 43: 2094–2103, 2015. doi: 10.1097/CCM.0000000000001162. [DOI] [PubMed] [Google Scholar]
- 9.Raffray L, Douchet I, Augusto JF, Youssef J, Contin-Bordes C, Richez C, Duffau P, Truchetet ME, Moreau JF, Cazanave C, Leroux L, Mourrissoux G, Camou F, Clouzeau B, Jeannin P, Delneste Y, Gabinski C, Guisset O, Lazaro E, Blanco P. Septic shock sera containing circulating histones induce dendritic cell-regulated necrosis in fatal septic shock patients. Crit Care Med 43: e107–e116, 2015. doi: 10.1097/CCM.0000000000000879. [DOI] [PubMed] [Google Scholar]
- 10.Zhang Y, Wen Z, Guan L, Jiang P, Gu T, Zhao J, Lv X, Wen T. Extracellular histones play an inflammatory role in acid aspiration-induced acute respiratory distress syndrome. Anesthesiology 122: 127–139, 2015. doi: 10.1097/ALN.0000000000000429. [DOI] [PubMed] [Google Scholar]
- 11.Abrams ST, Zhang N, Manson J, Liu T, Dart C, Baluwa F, Wang SS, Brohi K, Kipar A, Yu W, Wang G, Toh CH. Circulating histones are mediators of trauma-associated lung injury. Am J Respir Crit Care Med 187: 160–169, 2013. doi: 10.1164/rccm.201206-1037OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yehya N, Fazelinia H, Lawrence GG, Spruce LA, Mai MV, Worthen GS, Christie JD. Plasma nucleosomes are associated with mortality in pediatric acute respiratory distress syndrome. Crit Care Med 49: 1149–1158, 2021. doi: 10.1097/CCM.0000000000004923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yehya N, Thomas NJ, Margulies SS. Circulating nucleosomes are associated with mortality in pediatric acute respiratory distress syndrome. Am J Physiol Lung Cell Mol Physiol 310: L1177–L1184, 2016. doi: 10.1152/ajplung.00067.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tamkovich SN, Tutanov OS, Serdukov DS, Belenikin MS, Shlikht AG, Kirushina NA, Voytsitskiy VE, Tsentalovich YP, Tkachuk VA, Laktionov PP. Protein content of circulating nucleoprotein complexes. Adv Exp Med Biol 924: 133–136, 2016. doi: 10.1007/978-3-319-42044-8_26. [DOI] [PubMed] [Google Scholar]
- 15.Yehya N, Thomas NJ, Meyer NJ, Christie JD, Berg RA, Margulies SS. Circulating markers of endothelial and alveolar epithelial dysfunction are associated with mortality in pediatric acute respiratory distress syndrome. Intensive Care Med 42: 1137–1145, 2016. doi: 10.1007/s00134-016-4352-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tyanova S, Temu T, Cox J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc 11: 2301–2319, 2016. doi: 10.1038/nprot.2016.136. [DOI] [PubMed] [Google Scholar]
- 17.Bruderer R, Bernhardt OM, Gandhi T, Miladinovic SM, Cheng LY, Messner S, Ehrenberger T, Zanotelli V, Butscheid Y, Escher C, Vitek O, Rinner O, Reiter L. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol Cell Proteomics 14: 1400–1410, 2015. doi: 10.1074/mcp.M114.044305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tyanova S, Temu T, Sinitcyn P, Carlson A, Hein MY, Geiger T, Mann M, Cox J. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods 13: 731–740, 2016. doi: 10.1038/nmeth.3901. [DOI] [PubMed] [Google Scholar]
- 19.Oba S, Sato MA, Takemasa I, Monden M, Matsubara K, Ishii S. A Bayesian missing value estimation method for gene expression profile data. Bioinformatics 19: 2088–2096, 2003. doi: 10.1093/bioinformatics/btg287. [DOI] [PubMed] [Google Scholar]
- 20.Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8: 118–127, 2007. doi: 10.1093/biostatistics/kxj037. [DOI] [PubMed] [Google Scholar]
- 21.Breiman L. Random forests. Mach Learn 45: 5–32, 2001. doi: 10.1023/A:1010933404324. [DOI] [Google Scholar]
- 22.Ekaney ML, Otto GP, Sossdorf M, Sponholz C, Boehringer M, Loesche W, Rittirsch D, Wilharm A, Kurzai O, Bauer M, Claus RA. Impact of plasma histones in human sepsis and their contribution to cellular injury and inflammation. Crit Care 18: 543, 2014. doi: 10.1186/s13054-014-0543-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wygrecka M, Kosanovic D, Wujak L, Reppe K, Henneke I, Frey H, Didiasova M, Kwapiszewska G, Marsh LM, Baal N, Hackstein H, Zakrzewicz D, Muller-Redetzky HC, de Maat S, Maas C, Nolte MW, Panousis C, Schermuly RT, Seeger W, Witzenrath M, Schaefer L, Markart P. Antihistone properties of C1 esterase inhibitor protect against lung injury. Am J Respir Crit Care Med 196: 186–199, 2017. doi: 10.1164/rccm.201604-0712OC. [DOI] [PubMed] [Google Scholar]
- 24.Lorenz E, Muhlebach MS, Tessier PA, Alexis NE, Duncan Hite R, Seeds MC, Peden DB, Meredith W. Different expression ratio of S100A8/A9 and S100A12 in acute and chronic lung diseases. Respir Med 102: 567–573, 2008. doi: 10.1016/j.rmed.2007.11.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ryckman C, Vandal K, Rouleau P, Talbot M, Tessier PA. Proinflammatory activities of S100: proteins S100A8, S100A9, and S100A8/A9 induce neutrophil chemotaxis and adhesion. J Immunol 170: 3233–3242, 2003. doi: 10.4049/jimmunol.170.6.3233. [DOI] [PubMed] [Google Scholar]
- 26.Rouleau P, Vandal K, Ryckman C, Poubelle PE, Boivin A, Talbot M, Tessier PA. The calcium-binding protein S100A12 induces neutrophil adhesion, migration, and release from bone marrow in mouse at concentrations similar to those found in human inflammatory arthritis. Clin Immunol 107: 46–54, 2003. doi: 10.1016/s1521-6616(02)00043-8. [DOI] [PubMed] [Google Scholar]
- 27.Hasegawa T, Kosaki A, Kimura T, Matsubara H, Mori Y, Okigaki M, Masaki H, Toyoda N, Inoue-Shibata M, Kimura Y, Nishikawa M, Iwasaka T. The regulation of EN-RAGE (S100A12) gene expression in human THP-1 macrophages. Atherosclerosis 171: 211–218, 2003. doi: 10.1016/j.atherosclerosis.2003.08.021. [DOI] [PubMed] [Google Scholar]
- 28.Sinha P, Delucchi KL, McAuley DF, O'Kane CM, Matthay MA, Calfee CS. Development and validation of parsimonious algorithms to classify acute respiratory distress syndrome phenotypes: a secondary analysis of randomised controlled trials. Lancet Respir Med 8: 247–257, 2020. doi: 10.1016/S2213-2600(19)30369-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hotz MJ, Qing D, Shashaty MGS, Zhang P, Faust H, Sondheimer N, Rivella S, Worthen GS, Mangalmurti NS. Red blood cells homeostatically bind mitochondrial DNA through TLR9 to maintain quiescence and to prevent lung injury. Am J Respir Crit Care Med 197: 470–480, 2018. doi: 10.1164/rccm.201706-1161OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Qing DY, Conegliano D, Shashaty MG, Seo J, Reilly JP, Worthen GS, Huh D, Meyer NJ, Mangalmurti NS. Red blood cells induce necroptosis of lung endothelial cells and increase susceptibility to lung inflammation. Am J Respir Crit Care Med 190: 1243–1254, 2014. doi: 10.1164/rccm.201406-1095OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Cohen TS, Prince AS. Activation of inflammasome signaling mediates pathology of acute P. aeruginosa pneumonia. J Clin Invest 123: 1630–1637, 2013. doi: 10.1172/JCI66142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cheng KT, Xiong S, Ye Z, Hong Z, Di A, Tsang KM, Gao X, An S, Mittal M, Vogel SM, Miao EA, Rehman J, Malik AB. Caspase-11-mediated endothelial pyroptosis underlies endotoxemia-induced lung injury. J Clin Invest 127: 4124–4135, 2017. doi: 10.1172/JCI94495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Liu P, Feng Y, Li H, Chen X, Wang G, Xu S, Li Y, Zhao L. Ferrostatin-1 alleviates lipopolysaccharide-induced acute lung injury via inhibiting ferroptosis. Cell Mol Biol Lett 25: 10, 2020. doi: 10.1186/s11658-020-00205-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Matute-Bello G, Liles WC, Steinberg KP, Kiener PA, Mongovin S, Chi EY, Jonas M, Martin TR. Soluble Fas ligand induces epithelial cell apoptosis in humans with acute lung injury (ARDS). J Immunol 163: 2217–2225, 1999. [PubMed] [Google Scholar]
- 35.Matute-Bello G, Liles WC, Radella F, Steinberg KP, Ruzinski JT, Hudson LD, Martin TR. Modulation of neutrophil apoptosis by granulocyte colony-stimulating factor and granulocyte/macrophage colony-stimulating factor during the course of acute respiratory distress syndrome. Crit Care Med 28: 1–7, 2000. doi: 10.1097/00003246-200001000-00001. [DOI] [PubMed] [Google Scholar]
- 36.Tamada N, Tojo K, Yazawa T, Goto T. Necrosis rather than apoptosis is the dominant form of alveolar epithelial cell death in lipopolysaccharide-induced experimental acute respiratory distress syndrome model. Shock 54: 128–139, 2020. doi: 10.1097/SHK.0000000000001425. [DOI] [PubMed] [Google Scholar]
- 37.Loimaranta V, Hepojoki J, Laaksoaho O, Pulliainen AT. Galectin-3-binding protein: a multitask glycoprotein with innate immunity functions in viral and bacterial infections. J Leukoc Biol 104: 777–786, 2018. doi: 10.1002/JLB.3VMR0118-036R. [DOI] [PubMed] [Google Scholar]
- 38.Iacobelli S, Natoli C, D'Egidio M, Tamburrini E, Antinori A, Ortona L. Lipoprotein 90K in human immunodeficiency virus-infected patients: a further serologic marker of progression. J Infect Dis 164: 819, 1991. doi: 10.1093/infdis/164.4.819. [DOI] [PubMed] [Google Scholar]
- 39.Liu KT, Liu YH, Chen YH, Lin CY, Huang CH, Yen MC, Kuo PL. Serum galectin-9 and galectin-3-binding protein in acute dengue virus infection. Int J Mol Sci 17: 832, 2016. doi: 10.3390/ijms17060832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Trahey M, Weissman IL. Cyclophilin C-associated protein: a normal secreted glycoprotein that down-modulates endotoxin and proinflammatory responses in vivo. Proc Natl Acad Sci USA 96: 3006–3011, 1999. doi: 10.1073/pnas.96.6.3006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hong CS, Park MR, Sun EG, Choi W, Hwang JE, Bae WK, Rhee JH, Cho SH, Chung IJ. Gal-3BP negatively regulates NF-κB signaling by inhibiting the activation of TAK1. Front Immunol 10: 1760, 2019. doi: 10.3389/fimmu.2019.01760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Messner CB, Demichev V, Wendisch D, Michalick L, White M, Freiwald A, Textoris-Taube K, Vernardis SI, Egger AS, Kreidl M, Ludwig D, Kilian C, Agostini F, Zelezniak A, Thibeault C, Pfeiffer M, Hippenstiel S, Hocke A, von Kalle C, Campbell A, Hayward C, Porteous DJ, Marioni RE, Langenberg C, Lilley KS, Kuebler WM, Mülleder M, Drosten C, Suttorp N, Witzenrath M, Kurth F, Sander LE, Ralser M. Ultra-high-throughput clinical proteomics reveals classifiers of COVID-19 infection. Cell Syst 11: 11–24.e4, 2020. doi: 10.1016/j.cels.2020.05.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Gutmann C, Takov K, Burnap SA, Singh B, Ali H, Theofilatos K, et al. SARS-CoV-2 RNAemia and proteomic trajectories inform prognostication in COVID-19 patients admitted to intensive care. Nat Commun 12: 3406, 2021. doi: 10.1038/s41467-021-23494-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ricciuto DR, dos Santos CC, Hawkes M, Toltl LJ, Conroy AL, Rajwans N, Lafferty EI, Cook DJ, Fox-Robichaud A, Kahnamoui K, Kain KC, Liaw PC, Liles WC. Angiopoietin-1 and angiopoietin-2 as clinically informative prognostic biomarkers of morbidity and mortality in severe sepsis. Crit Care Med 39: 702–710, 2011. doi: 10.1097/CCM.0b013e318206d285. [DOI] [PubMed] [Google Scholar]
- 45.Calfee CS, Gallagher D, Abbott J, Thompson BT, Matthay MA, Network NA. Plasma angiopoietin-2 in clinical acute lung injury: prognostic and pathogenetic significance. Crit Care Med 40: 1731–1737, 2012. doi: 10.1097/CCM.0b013e3182451c87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Agrawal A, Matthay MA, Kangelaris KN, Stein J, Chu JC, Imp BM, Cortez A, Abbott J, Liu KD, Calfee CS. Plasma angiopoietin-2 predicts the onset of acute lung injury in critically ill patients. Am J Respir Crit Care Med 187: 736–742, 2013. doi: 10.1164/rccm.201208-1460OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Evans IVR, Phillips GS, Alpern ER, Angus DC, Friedrich ME, Kissoon N, Lemeshow S, Levy MM, Parker MM, Terry KM, Watson RS, Weiss SL, Zimmerman J, Seymour CW. Association between the New York sepsis care mandate and in-hospital mortality for pediatric sepsis. JAMA 320: 358–367, 2018. doi: 10.1001/jama.2018.9071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Schouten LR, Veltkamp F, Bos AP, van Woensel JB, Serpa Neto A, Schultz MJ, Wosten-van Asperen RM. Incidence and mortality of acute respiratory distress syndrome in children: a systematic review and meta-analysis. Crit Care Med 44: 819–829, 2016. doi: 10.1097/CCM.0000000000001388. [DOI] [PubMed] [Google Scholar]
- 49.Yehya N, Keim G, Thomas NJ. Subtypes of pediatric acute respiratory distress syndrome have different predictors of mortality. Intensive Care Med 44: 1230–1239, 2018. doi: 10.1007/s00134-018-5286-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental Figs. S1–S4: https://doi.org/10.6084/m9.figshare.17159387.v1.