SUMMARY
WhiB7 represents a distinct subclass of transcription factors in the WhiB-Like (Wbl) family, a unique group of iron-sulfur (4Fe-4S] cluster-containing proteins exclusive to the phylum of Actinobacteria. In Mycobacterium tuberculosis (Mtb), WhiB7 interacts with domain 4 of the primary sigma factor (σA4) in the RNA polymerase holoenzyme and activates genes involved in multiple drug resistance and redox homeostasis. Here, we report crystal structures of the WhiB7:σA4 complex alone and bound to its target promoter DNA at 1.55-Å and 2.6-Å resolution, respectively. These structures show how WhiB7 regulates gene expression by interacting with both σA4 and the AT-rich sequence upstream of the –35 promoter DNA via its C-terminal DNA-binding motif, the AT-hook. By combining comparative structural analysis of the two high-resolution σA4-bound Wbl structures with molecular and biochemical approaches, we identify the structural basis of the functional divergence between the two distinct subclasses of Wbl proteins in Mtb.
In brief
WhiB7 is an Fe-S cluster-bound transcription factor in mycobacteria that is activated by multiple antibiotics. Wan et al. provide an atomic view of how WhiB7 activates transcription by binding to σA and AT-rich promoter DNA, and they identify the structural motifs that enable WhiB7 to function differently from its paralog WhiB1.
Graphical Abstract

INTRODUCTION
The WhiB-like (Wbl) family proteins are a unique group of iron-sulfur ([4Fe-4S]) cluster-bound proteins first discovered in Streptomyces and widely distributed in the phylum Actinobacteria, which includes the notorious bacterial pathogen Mycobacterium tuberculosis (Mtb) (Davis and Chater, 1992; Soliveri et al., 2000). Wbl proteins play versatile roles in bacterial growth, development, and stress responses (see recent reviews in Bush, 2018; Chandra and Chater, 2014; and Saini et al., 2012). All members in the Wbl family share a [4Fe-4S] cluster binding motif comprising four invariant cysteines and a G[I/V/L]W[G/A]G motif that is often referred to as the b turn (Figure 1A; Figure S1). The putative DNA-binding motif in the C terminus is less defined, featuring a cluster of positively charged residues (Arg/Lys) in most Wbl proteins.
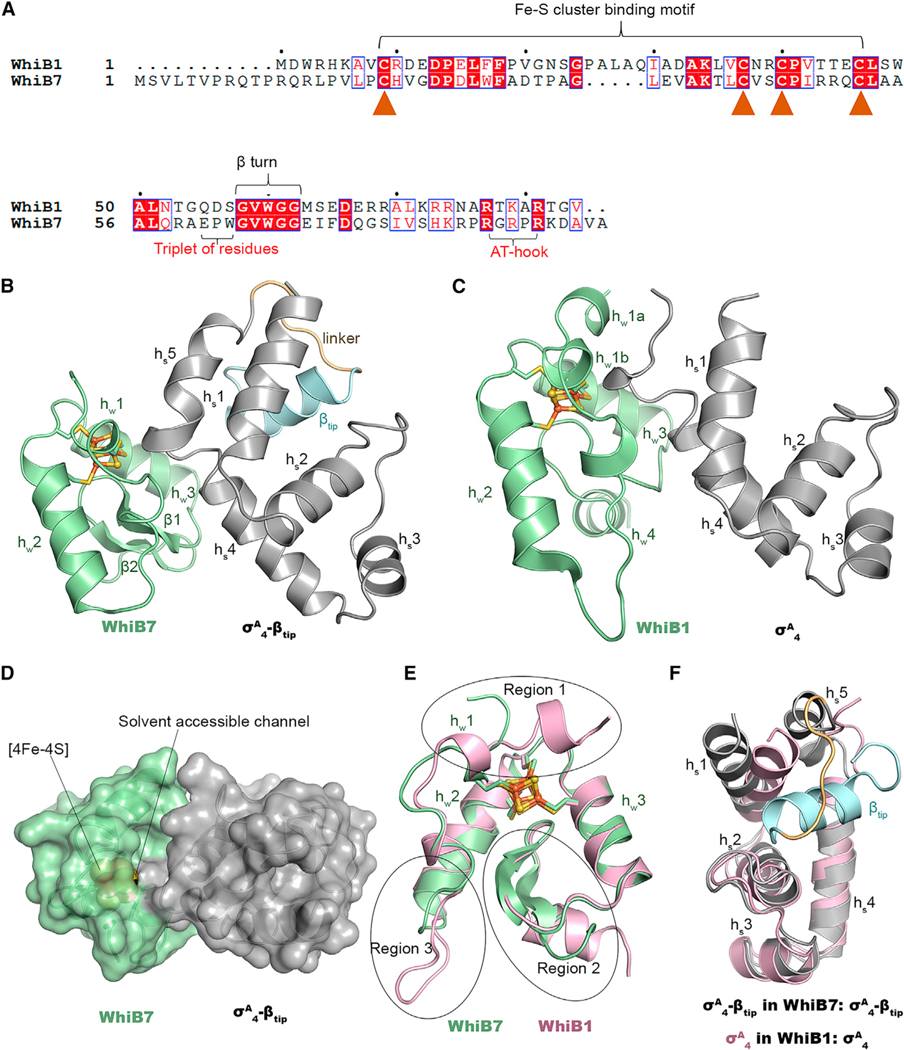
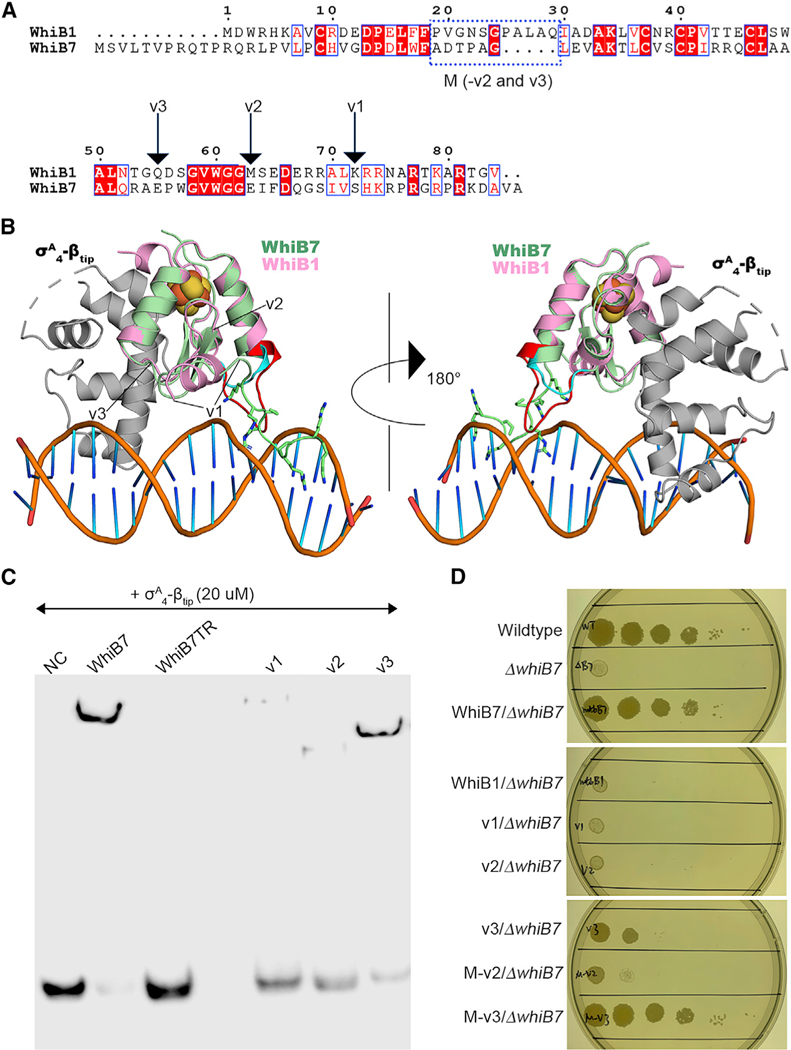
Figure 1. Comparison of the overall structures between the WhiB7TR:σA4-βtip complex and the WhiB1:σA4 complex.
(A) Sequence alignment of Mtb WhiB1 and WhiB7. The conserved cysteines in the Fe-S cluster binding motif are indicated by solid brown triangles. The WhiB7 subclass signature motifs (the triplet-peptide sequence and the AT-hook) are indicated with red fonts.
(B and C) Cartoon representations of (B) WhiB7TR:σA4-βtip and (C) WhiB1:σA4 (PDB: 6ONO) for side-by-side comparison.
(D) Surface representation of the WhiB7TR:σA4-βtip structure, with a water-accessible channel from the complex surface toward the [4Fe-4S] cluster indicated by lines.
(E) Overlay of WhiB1 (pink) and WhiB7 (pale green) in the two Wbl complexes. Three regions (regions 1–3) with significant differences between the two structures are indicated by circles.
(F) 3D structural comparison of σA4 in WhiB7TR:σA4-βtip and WhiB1:σA4.
In all structures, the [4Fe-4S] cluster is shown in ball-and-stick representation, with Fe atoms represented in orange and S atoms represented in yellow.
In Mtb, seven Wbl proteins share ~30%–50% sequence identity, display different physicochemical properties in vitro, and play crucial roles in the virulence and antibiotic resistance of the pathogen (Alam et al., 2009; Burian et al., 2013; Casonato et al., 2012; Chawla et al., 2012, 2018; Chen et al., 2016; Larsson et al., 2012; Morris et al., 2005; Raghunand and Bishai, 2006; Smith et al., 2010, 2012; Steyn et al., 2002; Wu et al., 2017). Five Wbl members of Mtb (WhiB1–4 and WhiB7) represent different subclasses of Wbl proteins widely distributed in Actinomycetes (Bush, 2018; Chandra and Chater, 2014; Saini et al., 2012). In contrast, WhiB6 is specific to mycobacteria, and WhiB5 is only found in mycobacterial pathogens. Among the five non-mycobacteria-specific Mtb Wbl proteins, WhiB1 is an essential, nitric oxide-sensitive transcription factor that is required for mycobacterial growth (Saini et al., 2012; Smith et al., 2010). It is closer to WhiB2–4 by sequence analysis and distinct from the WhiB7 subclass (Figure S1). The WhiB7 subclass features two subclass-specific signature motifs that are not found in the other four Wbl subclasses: the triplet of residues (EPW) adjacent to the N-terminal β turn and a canonical AT-hook (RGRP) in the C-terminal DNA-binding motif (Figure S1) (Burian et al., 2012, 2013; Morris et al., 2005). The AT-hook is a DNA-binding motif that preferably binds to the minor groove of AT-rich B-form DNA (Aravind and Landsman, 1998; Fonfría-Subirós et al., 2012; Huth et al., 1997). It was first identified in the high-mobility-group protein HMGA and is found prevalent in eukaryotic nuclear proteins but rare in bacteria (Aravind and Landsman, 1998). Previous studies on HMGA proteins in eukaryotes suggest that the AT-hook increases DNA-binding affinity, induces DNA conformational changes, and thus orchestrates with the partner transcription factors for transcriptional regulation and modulation of DNA architecture (Fonfría-Subirós et al., 2012; Honda et al., 2016; Morrison et al., 2017; Rodríguez et al., 2015). Little is known about the mode of action of the AT-hook-containing transcription factors in bacteria.
Mtb WhiB7 is a transcription activator that is upregulated by antibiotics with diverse structures and mechanisms of action and confers resistance to multiple antibiotics, including two second-line anti-tuberculosis drugs (streptomycin and kanamycin) (Burian et al., 2012; Morris et al., 2005; Ramón-García et al., 2013). WhiB7 has also been linked to redox homeostasis, as deletion of whiB7 resulted in a decrease in cellular levels of mycothiol, the major low-molecular-weight thiol in Actinobacteria, and a shift toward a higher ratio of the oxidized mycothiol (Burian et al., 2012). Previous studies indicate that the antibiotic activation of whiB7 involves two coordinated processes: repression of early termination occurring at the upstream of the whiB7 gene and upregulation of the promoter activity (Burian and Thompson, 2018; Burian et al., 2013; Reeves et al., 2013). However, the underlying mechanism of the whiB7 activation by antibiotics is not fully understood. Upon activation, the [4Fe-4S] cluster-bound (holo-)WhiB7 binds to domain 4 of the σ70-family primary sigma factor σA (σA4) in the RNA polymerase (RNAP) holoenzyme and upregulates genes (including itself) in the whiB7 regulon in an AT-hook-dependent manner (Burian et al., 2012, 2013; Morris et al., 2005). It is unclear how the AT-hook is used in WhiB7 to activate gene expression. Domain 4 of σA is a highly conserved motif among the σ70-family primary sigma factors in bacteria, which recognizes the –35 element and interacts with many transcription factors for regulating gene expression (Feng et al., 2016; Gomez et al., 1998; Lee et al., 2012). Several other Wbl proteins also regulate gene expression via interaction with σA, indicating a shared mode of action by Wbl proteins (Feng et al., 2016; Kudhair et al., 2017; Stewart et al., 2020; Steyn et al., 2002; Wan et al., 2020).
The lack of atomic-resolution structural information of Wbl proteins hinders the mechanistic understanding of their diverse physiochemical properties and versatile biological functions. To date, Mtb WhiB1 is the only Wbl protein that has been structurally characterized in atomic detail, first by nuclear magnetic resonance of holo-WhiB1 (Kudhair et al., 2017). We subsequently reported a 1.85-Å crystal structure of WhiB1 in complex with σA (WhiB1:σA ), which reveals an unexpectedly tight interaction at the molecular interface and supports a new molecular mechanism of transcriptional regulation by WhiB1 in Actinobacteria (Wan et al., 2020). Here, we report crystal structures of WhiB7 in complex with σA, without and with its own promoter DNA (PwhiB7), at 1.55 Å and 2.6 Å, respectively. In both structures, σA is fused with the RNAP β-subunit flap tip helix (βtip) linked by an artificial sequence to stabilize its conformation in the RNAP holoenzyme (Blanco et al., 2011; Campbell et al., 2002). Thus, these structures represent the WhiB7-dependent transcription activation subcomplexes. Together with biochemical and site-directed mutagenetic studies, we provide an atomic view of how WhiB7 activates target genes by interacting with σA and stabilizing its binding to the –35 element via the AT-hook. Structural comparison with the σA4-bound WhiB1 allows us to identify the key structural motifs that explain how these two Wbl proteins are coordinated in Mtb to balance active cell growth and antibiotic stress response by anchoring to the same binding site on σA4.
RESULTS
Structure of the σA4-bound Mtb WhiB7 compared with the WhiB1:σA4 complex
We first attempted to crystalize a truncated WhiB7 (WhiB7TR; aa 1–79 without the AT-hook region) in complex with σA4 following the strategy used for structure determination of WhiB1:σA4 but without success (Wan et al., 2020). To stabilize the complex of WhiB7 and σA4 for crystallization, we designed a chimeric protein denoted σA4-βtip by fusing σA4 (containing aa 446–528) with βtip of the RNAP β-subunit (containing aa 815–829) linked by the artificial sequence GSSGSG, mimicking the contacts between the two subunits in the RNAP holoenzyme (Blanco et al., 2011; Campbell et al., 2002). Phases were determined experimentally using the single-wavelength anomalous diffraction (SAD) data from the crystal of the SeMet-substituted WhiB7TR:σA4-βtip complex (STAR Methods). The final model of the WhiB7TR:σA4-βtip complex was refined at 1.55 Å (STAR Methods; Table 1).
Table 1.
Data collection and refinement statistics for WhiB7TR:σA4-βtip and WhiB7:σA4-βtip:DNA
| Data for: | ||
|---|---|---|
|
|
||
| Statistic | WhiB7TR:σA4-βtip | WhiB7:σA4-βtip:DNA |
|
| ||
| Data collectiona | ||
|
| ||
| Space group | C2221 | P3221 |
| Cell dimensions | ||
| a, b, c (Å ) | 54.6, 61.0, 210.2 | 69.3, 69.3, 167.4 |
| α, β, γ (°) | 90.0, 90.0, 90.0 | 90.0, 106.8, 90.0 |
| Wavelength (Å) | 0.9795 | 0.9790 |
| Resolution (Å) | 50–1.55 (1.61–1.55) | 50–2.60 (2.64–2.60) |
| R merge | 0.115 (1.37) | 0.101 (2.09) |
| I/σI | 40.0 (2.1) | 33.1 (0.81) |
| Completeness (%) | 99.3 (95.7) | 98.1 (83.3) |
| Multiplicity | 52.1 (46.1) | 26.4 (15.1) |
| Anomalous multiplicity | 26.5 (24.7) | — |
| No. of unique reflections | 51,219 (4,871) | 14,301 (1,216) |
| CC1/2 (%) | 100.0 (80.5) | 99.4 (88.5) |
|
| ||
| Refinement | ||
|
| ||
| Resolution (Å) | 50–1.55 (1.60–1.55) | 50–2.60 (2.70–2.60) |
| No. of molecules per asymmetric unit | 2 | 1 |
| Rwork/Rfree | 0.187/0.207 | 0.224/0.254 |
|
| ||
| Included residue no. | ||
|
| ||
| WhiB7 | 16–79; 16–79b | 16–88 |
| σA4 | 452–526; 452–528b | 454–525 |
| βtip | 815–829; 815–827b | 815–827 |
| No. of atoms | 2,861 | 1,987 |
| Protein | 2,508 | 1,979 |
| DNA | — | 732 |
| Ligand | 16 | 8 |
| Water | 337 | |
| B factors (Å2) | 31.53 | 122.67 |
| Protein | 30.27 | 115.8 |
| DNA | — | 131.9 |
| Ligand | 18.99 | 100.43 |
| Water | 35.52 | |
|
| ||
| RMSDs | ||
|
| ||
| Bond lengths (Å) | 0.01 | 0.01 |
| Bond angles (°) | 1.98 | 0.95 |
|
| ||
| Ramachandran statistics (%) | ||
|
| ||
| Favored regions | 93.32 | 94.08 |
| Allowed regions | 2.68 | 5.92 |
| Outliers | 0 | 0 |
The highest resolution shell statistics are shown in parentheses.
The two sets of the residues are for each of the two complexes in the asymmetric unit.
The overall architecture of the WhiB7TR:σA4-βtip complex is comparable to that of the WhiB1:σA4 complex, both of which are dominated by alpha helices and with a fully occupied [4Fe-4S] cluster (Figures 1B and 1C). More importantly, the molecular interface of both the complexes is centered in the [4Fe-4S] cluster binding pocket and involved in the C terminus of helix hs4 in σA4.
Structural comparison indicates that the core helices surrounding the [4Fe-4S] cluster are conserved between WhiB1 and WhiB7 in the two σA4-bound complexes, despite their low sequence identity; however, there are significant structural differences in three regions (Figures 1B–1E). WhiB7 and WhiB1 superimpose with a Cα root-mean-square deviation (RMSDCα) of 0.68 Å for the 37 aligned residues located in helices hw1b and hw2–3 of WhiB1 (Figure 1E). In contrast, the structural elements in the N- and C-terminal regions of WhiB7 (regions 1 and 2, respectively; Figure 1E) are distinct from the corresponding regions in WhiB1. The N-terminal residues of WhiB1 form two short helical structures (hw1a and hw1b) hovering over the Fe-S cluster binding pocket (Figures 1C and 1E). By comparison, the first 15 residues in the N-terminal WhiB7 are not visible in the electron density map, and the backbone of P16 points away from the Fe-S cluster, leading to a more solvent-exposed cluster in WhiB7:σA4-βtip (Figures 1D and 1E). In the C terminus, WhiB1 features a long loop containing the b turn, which is followed by helix hw4 (Figures 1C and 1E). However, the residues in WhiB7 corresponding to the loop region of WhiB1 are replaced by a more ordered β hairpin followed by a random coil that is expected to connect to the AT-hook (Figures 1B and 1E). The β hairpin in WhiB7 is stabilized by the hydrophobic interactions between the side chains of two pairs of conserved residues in WhiB7 beside the backbone hydrogen bonds: V65••••I70 (3.85 Å apart) on one side; and W66•••F71 (3.65 Å apart) on the other side (Figure S2). Consistent with the structural differences in this region, sequence alignment shows that the residues around the β hairpin, including the WhiB7 signature triplet residues (EPW) and the b turn motif, are highly conserved within the WhiB7 subclass but are generally less conserved in the WhiB1 subclass, except for the b turn (Figure S1). As described in the following text, these two regions underline the structural basis for the observed differences in how these two Wbl proteins interact with σA4, react to O2, and interact with target DNA. In addition, the loop connecting helices hw1 and hw2 in WhiB7 (region 3 in Figure 1E) is shorter than the corresponding region in WhiB1. It is correlated with a gap in the sequence alignment, where WhiB7 lacks the corresponding residues of P25–Q29 in WhiB1 located between the C terminus of the loop and the N-terminal helix hw2 (Figures 1A and 1E). The significance of this structural difference to their functions is investigated later in the context of the structural comparison with WhiB7:σA4-βtip in the DNA-bound form.
As expected, βtip forms extensive interactions with four (hs1 and hs3–5) out of the five helices in σA4 of the WhiB7TR:σA4-βtip structure and stabilizes σA4 in a physiologically relevant conformation closely resembling that in the RNAP holoenzyme-DNA complex (PDB: 6EEC) (Blanco et al., 2011; Boyaci et al., 2019). The importance of βtip in maintaining the helical arrangement and stabilizing hs5 in σA4 of the RNAP holoenzyme has been demonstrated in previous studies (Blanco et al., 2011; Geszvain et al., 2004; Kuznedelov et al., 2002) and is evident in the structural overlay of σA4 in WhiB7TR: σA4-βtip with WhiB1: σA4 (Figure 1F). Quantitatively, 3D structural alignment of σA4 in the two Wbl structures against the RNAP holoenzyme-DNA complex (PDB: 6EEC) results in a RMSDCα of 0.65 Å for WhiB7TR:σA4-βtip versus 1.84 Å for WhiB1:σA4.
WhiB7TR:σA4-βtip shares a similar molecular interface as that of WhiB1:σA4 within the [4Fe-4S] cluster binding pocket but with several notable differences. The average buried surface area between WhiB7 and σA4 in the WhiB7TR:σA4-βtip complex (576 Å ± 3 Å) is noticeably smaller than that of the WhiB1:σA4 complex (645 Å ± 27 Å). As observed in the WhiB1:σA4 complex, H516 of σA4 in the WhiB7TR:σA4-βtip structure orchestrates hydrophobic interactions with the surrounding aromatic residues (W28, F29, and W66, corresponding to F17, F18, and W60 in WhiB1) and forms a hydrogen-bond network with the Fe-S cluster and the backbone O of V65 in WhiB7 (Figures 2A and 2B). The pull-down assays confirm the importance of H516 in these interactions (Figure S3). A key aromatic residue (W3) in the N-terminal region of WhiB1 is missing in WhiB7 (Figures 2A and 2B). W3 is invariant in the WhiB1 subclass and plays a crucial role in Fe-S cluster stability and complex formation (Wan et al., 2020). With the absence of a W3 counterpart in WhiB7, we observe that the contributions from other aromatic residues weigh in on the cluster stability and the interaction between WhiB7 and σA4. All the three σA4-contacting aromatic residues (W28, F29, and W66) in WhiB7 are required for forming the complex with σA4 in vitro and conferring antibiotic resistance in Mycobacterium smegmatis (Msm) with deletion of the whiB7 gene (ΔwhiB7) (STAR Methods; Figures 2E and 2F; Figure S3). In contrast, the effect of WhiB1 W60 on either σA4 binding or the function of WhiB1 in Msm is negligible (Wan et al., 2020). W28 and W66 are also important for the stability of the [4Fe-4S] cluster in WhiB7, as we observe the cluster degradation and protein aggregation of the Ala-substituted mutant proteins when expressed alone (Figures S3E and S3F). Structural information of the free WhiB7 is needed to elaborate their roles in the stability of the Fe-S cluster and protein folding. W63 at the molecular interface with σA4 outside of the Fe-S cluster binding pocket also shows a contribution to complex formation in the pull-down assay and the function of WhiB7 by the antibiotic sensitivity test, although its effect is not as pronounced as that of the core aromatic residues within the cluster binding pocket (Figure 2; Figure S3). W63 is in the signature triplet-residue ‘‘EPW’’ motif of the WhiB7 subclass and not conserved in WhiB1. Nonetheless, our previous study showed that W49 of WhiB1, also located at the molecular interface of the complex outside of the Fe-S cluster binding pocket, plays a role in stabilizing the Fe-S cluster in WhiB1 when expressed alone, but not when co-expressed with σA4 (Kudhair et al., 2017; Wan et al., 2020). It would be interesting to test whether these two Trp residues are functionally related, although they are not aligned in the 3D structural comparison (Figure 2B).
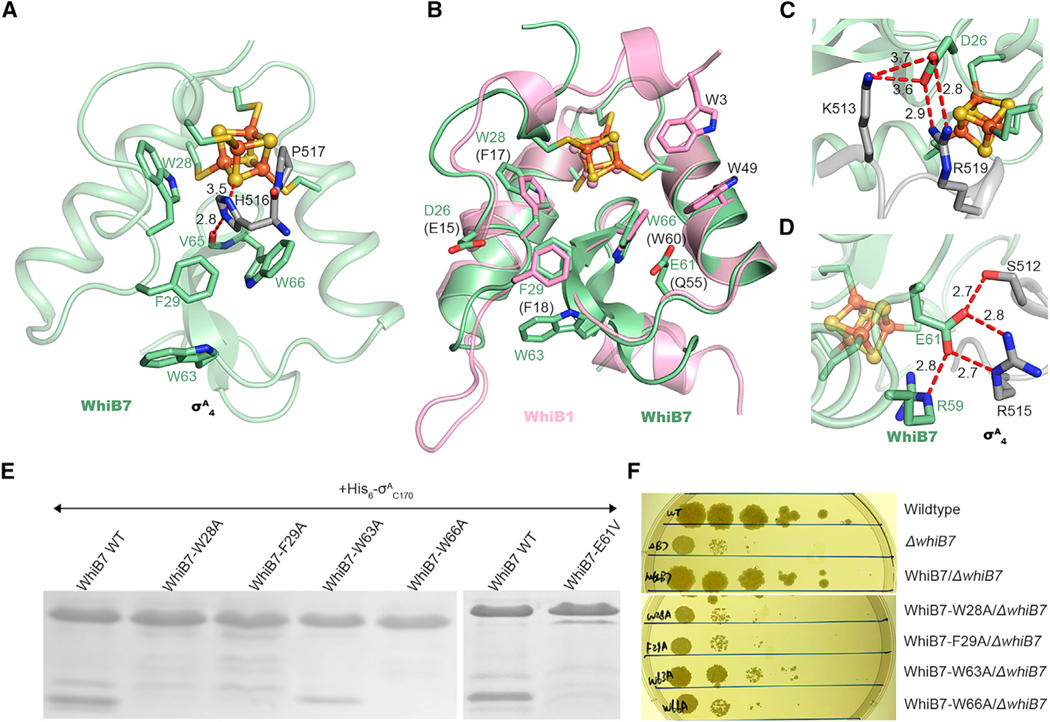
Figure 2. Molecular interface of WhiB7 and σA4 in comparison to WhiB1.
(A) Close-up view of the molecular interface between WhiB7 (pale green) and σA4 (gray) in the [4Fe-4S] cluster binding pocket, with H516 and P517 of σA4 protruded into the cluster binding pocket and surrounded by the highly conserved aromatic residues, W28, F29, and W66, within 4 Å. W63, the only aromatic residue at the molecular interface but outside of the cluster binding site, is also shown for comparison. The H516-centered hydrogen bonding network is highlighted by red dashed lines.
(B) Structural overlay of the key residues in WhiB7 (pale green) and WhiB1 (pink), respectively, at the interaction interface with σA4.
(C and D) Highlights of the two clusters of polar contacts between WhiB7 (pale green) and σA4 (gray) centered on D26 (C) and E61 (D), respectively.
(E) SDS-PAGE analyses of the samples from the co-expression and affinity purification of tagless WhiB7 (wild-type [WT] and mutants as indicated) and His6-σAC170. Full gel images and the UV-visible spectra of these samples are shown in Figure S3.
(F) Spectinomycin sensitivity test by spotted dilutions (10—1–10—5) of Msm WT and the whiB7 deletion mutant (ΔwhiB7) alone or complemented by Mtb whiB7 (WT or mutant as indicated).
Our structural and functional analyses suggest that the reduced hydrophobic contacts between WhiB7 and σA4 due to the absence of W3 are at least partially compensated by the enhanced hydrophilic interactions at the molecular interface of WhiB7 and σA4 (Figures 2C–2F; Figure S3). On both sides of helix hs4 in σA4 are two clusters of strong hydrophilic networks centered on D26 and E61 of WhiB7, respectively (Figures 2C and 2D). In particular, E61 is part of the WhiB7 subclass signature triplet-residue ‘‘EPW’’ motif and has been previously shown to be functionally essential (Burian et al., 2013). Consistently, we found that an E61V mutation in WhiB7 disrupts the complex in the pull-down assay (Figure 2E; Figure S3). In contrast, mutation of the corresponding polar residues in WhiB1 (E15 and Q55, corresponding to D26 and E61, respectively, in WhiB7) did not affect the stability of the complex, as judged by the pull-down assays (Wan et al., 2020).
O2 sensitivity of the [4Fe-4S] cluster in the σA4-bound WhiB7 and WhiB1
The local protein environment of the Fe-S cluster binding pocket in WhiB7TR:σA4-βtip is dominated by hydrophobic residues resembling that of WhiB1: σA4, except for the N-terminal region, which lacks a counterpart of WhiB1 W3 and thus leads to a more solvent-accessible pore toward the cluster of WhiB7TR: σA4-βtip (region 1 in Figure 1E; see also Figures 3A and 3B). A solvent-accessible path is found from the surface toward the [4Fe-4S] cluster in WhiB7: σA4-βtip with the minimum bottleneck of 1.4 Å, which is the assumed radius cutoff for solvent molecules. By comparison, the largest tunnel for WhiB1: σA4 has a minimum bottleneck of 0.8 Å (Wan et al., 2020).
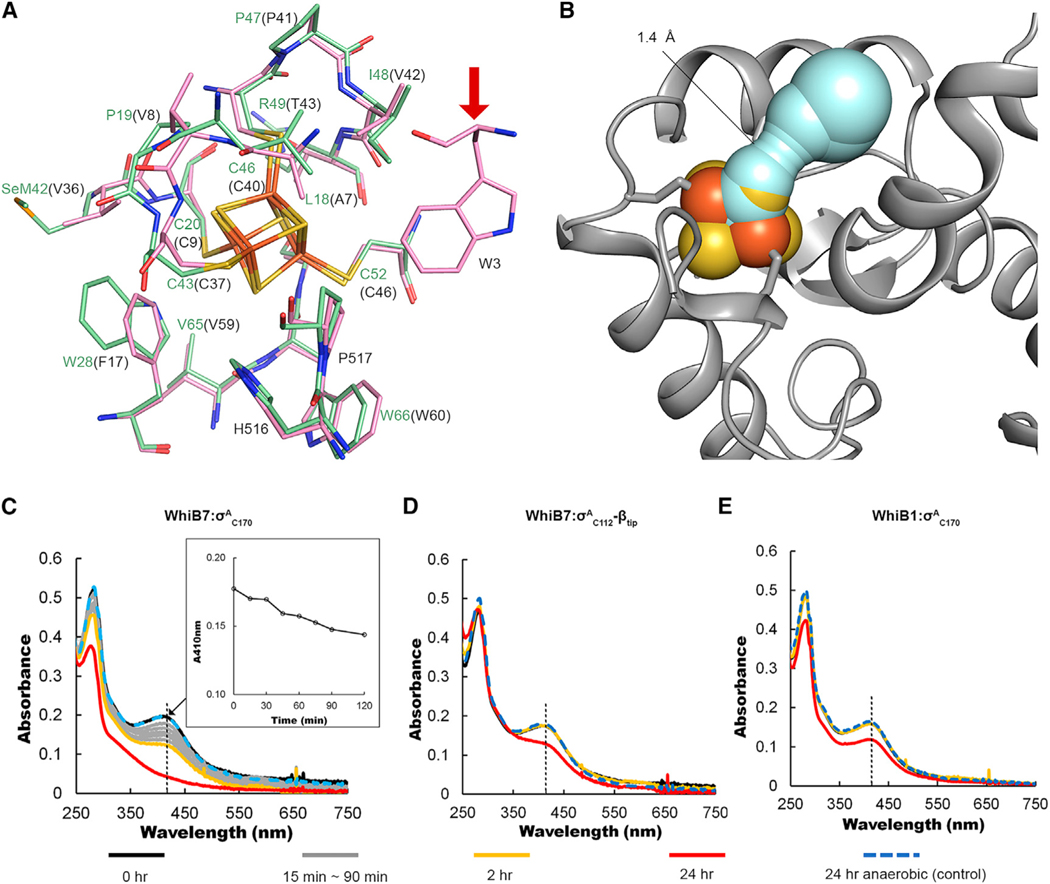
Figure 3. Comparison of the local environment and stability of the [4Fe-4S] cluster in the σA4-bound Wbl complexes.
(A) Comparison of the residues within 4 Å of the [4Fe-4S] cluster in the σA4-bound WhiB7 (pale green) and WhiB1 (gray). The residues surrounding the clusters are well conserved between the two complexes, except for W3 in WhiB1 (indicated by a red arrow).
(B) The solvent-accessible path from the complex surface toward the [4Fe-4S] cluster in WhiB7TR: σA4-βtip calculated by CAVER using a shell radius of 1.4 Å as the radius cutoff for solvent molecules.
(C–E) Monitoring the [4Fe-4S] cluster degradation in (C) the σA4-bound WhiB7, (D) the σA4-βtip-bound WhiB7, and (E) the σA4-bound WhiB1 in the air O2-saturated Tris buffer by UV-visible spectroscopy. A decrease in the intensity of the absorption peak around 410 nm, as highlighted by the black dashed lines, is indicative of the cluster loss. As a control, the UV-visible spectra were collected on the same protein samples kept under anaerobic conditions for 24 h (24-h anaerobic) to rule out the possible difference in thermal stability of the complexes.
In the O2 sensitivity test, we observe 50% cluster loss in the σA4-bound WhiB7 by 2 h in an aerobic environment, while the [4Fe-4S] cluster in the σA4-bound WhiB1 is unreactive to O2 under the same experimental conditions (Figures 3C and 3E). We note that the O2-mediated [4Fe-4S] cluster degradation in WhiB7: σA4 is considerably slower than measured in dedicated O2 sensors such as the fumarate and nitrate reduction regulatory protein FNR, which has a much more solvent-exposed [4Fe-4S] cluster (Kiley and Beinert, 1998; Partridge et al., 2009). However, the O2-sensitive [4Fe-4S] cluster in WhiB7 is unfavorable for an antibiotic-responsive transcription factor, considering that many antibiotics induce oxidative stress (Kohanski et al., 2010). Because βtip stabilizes the σA4 conformation in σA4-bound WhiB7, we thus tested whether βtip also has an impact on the stability of the cluster in the complex. As shown in Figure 3D, the [4Fe-4S] cluster in WhiB7: σA4-βtip is stable in an aerobic environment for 2 h, underlying the importance of maintaining the σA4 conformation in the RNAP holoenzyme on the cluster stability in WhiB7: σA4-βtip. This finding is comparable to that of the previous study on the [4Fe-4S]-containing nitric oxide sensor NsrR from Streptomyces coelicolor (ScNsrR), which is reportedly unreactive to O2, with no loss of cluster observed up to 43 min and only a marginal (~8%) cluster loss observed by 2 h (Crack et al., 2015). ScNsrR has a similar solvent-accessible path to the [4Fe-4S] cluster but with a non-Cys ligand (D8) bound to the Fe-S cluster, which may affect the cluster stability (Volbeda et al., 2017). Additionally, it remains to be tested whether the disordered N-terminal peptide in WhiB7 plays a role in shielding the cluster from O2.
Structural basis of the WhiB7-mediated transcription activation
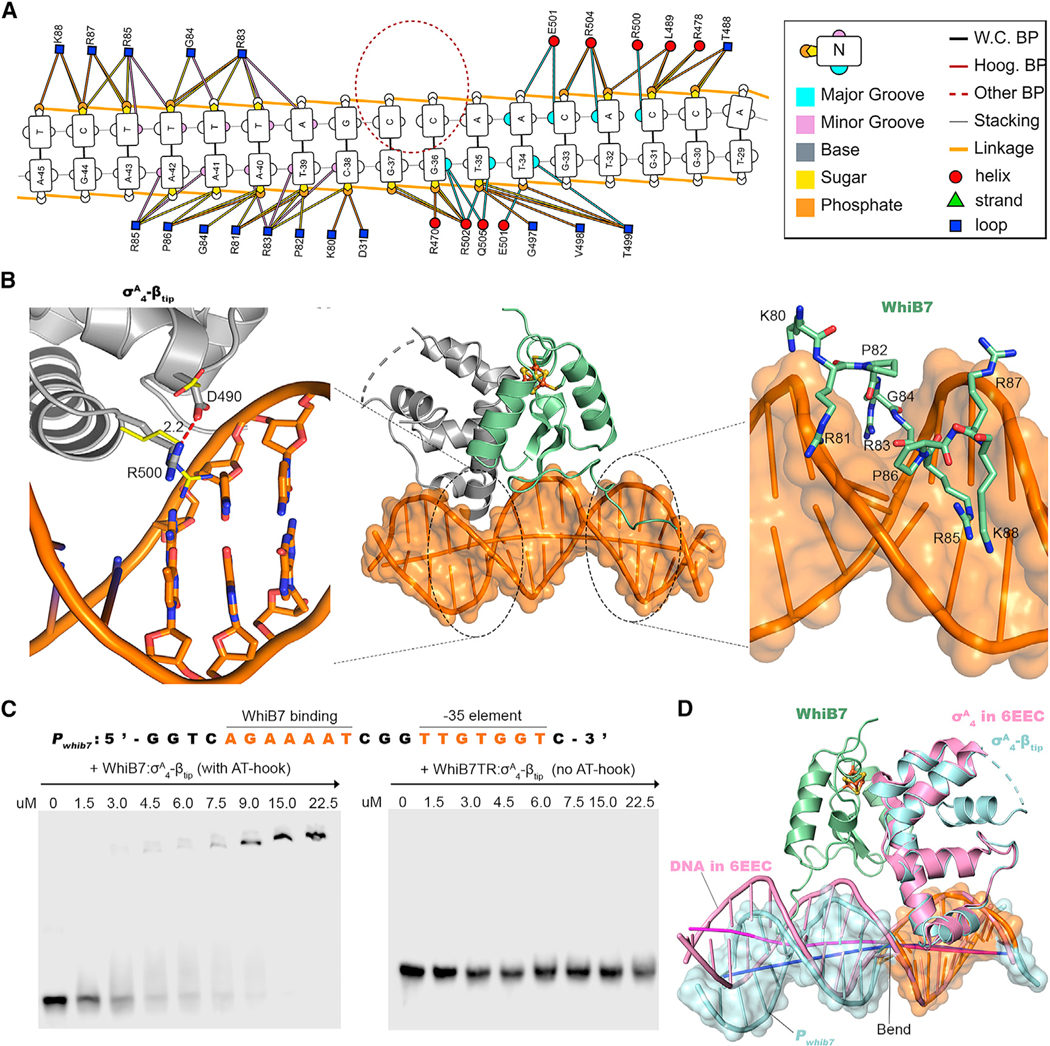
To gain structural insights into how WhiB7 activates gene expression by interacting with σA4 and its target gene promoters, we co-crystallized WhiB7: σA4-βtip and an 18-bp Pwhib7 including the –35 element and the upstream WhiB7 binding site with an overhang of C/G at the 5ʹ end (STAR Methods; Figure 4). The WhiB7TR: σA4-βtip structure was used as the search model for phase determination by molecular replacement. The crystal structure of the WhiB7: σA4-βtip:PwhiB7 complex was refined at 2.6 Å, with one copy of the complex per asymmetric unit (Table 1). The WhiB7: σA4-βtip:PwhiB7 structure can be superimposed over WhiB7TR: σA4-βtipwith a RMSDCα of 0.68 Å, indicating no significant conformational changes upon DNA binding. The electron density is well resolved and allows unambiguous assignment of the nucleotides in the DNA helix and the backbones of the AT-hook residues in WhiB7, while the electron density around the side chains of the Lys and Arg residues in the AT-hook is less pronounced (Figures 4A and 4B). In the crystal structure of WhiB7: σA4-βtip:PwhiB7, the symmetry-related DNA double helixes are packed head-to-tail to form a pseudo-continuous double helix (Figure 4C), which may have facilitated crystal packing.
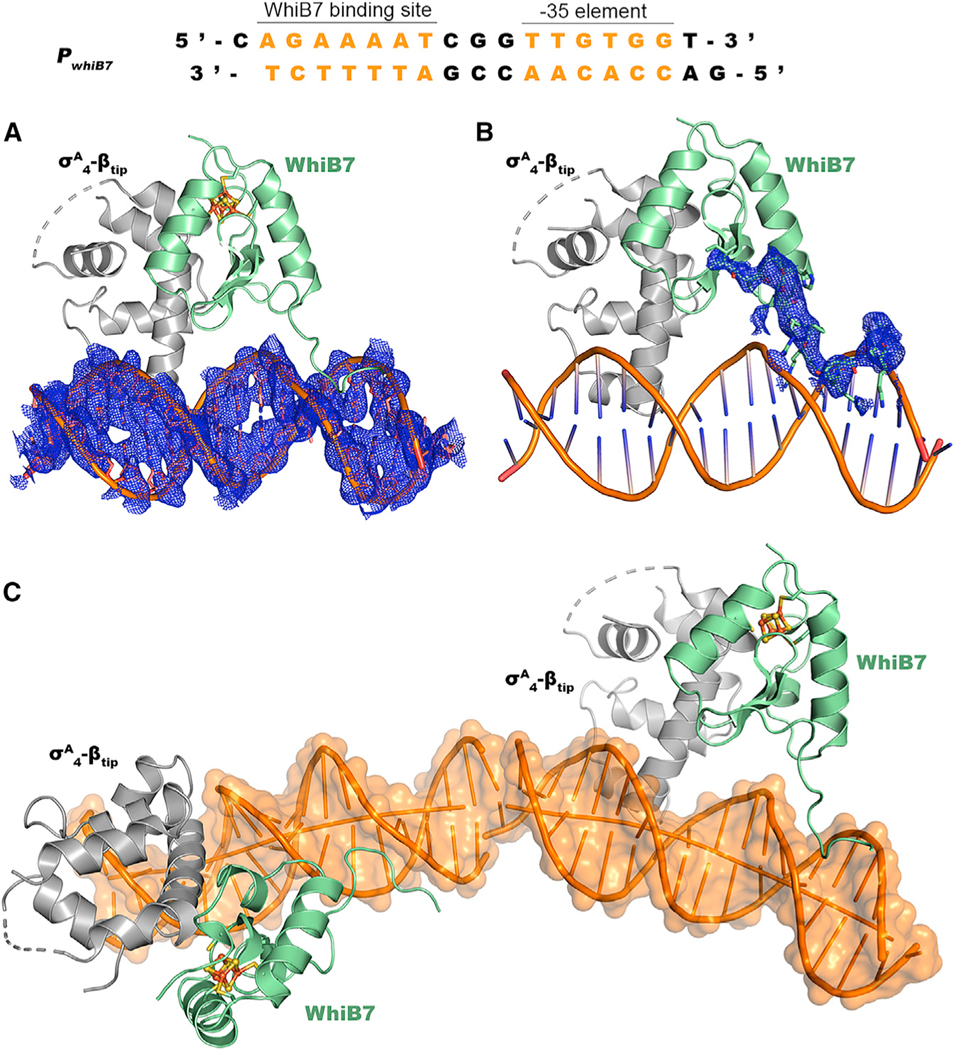
Figure 4. Structural characterization of the WhiB7:σA4-βtip:PwhiB7 complex.
(A and B) Simulated-annealing 2Fo-Fc composite omit maps around the PwhiB7 DNA helix and the AT-hook in WhiB7, respectively, contoured at 1σ.
(C) Head-to-tail packing of the symmetry-related WhiB7:σA4-βtip:PwhiB7 complexes in two neighboring asymmetric units. The joint of the two DNA helixes is indicated by a blue circle.
Examination of the protein-DNA contacts in the crystal structure of WhiB7: σA4-βtip:PwhiB7 reveals two DNA anchoring sites in the WhiB7: σA4-βtip complex that bind to PwhiB7 (Figures 5A and 5B). The DNA binding motif of σA4 inserts into the major groove of the 35 hexanucleotide site, while the AT-hook of WhiB7 slides into the minor groove immediately upstream of the σA4 binding site. The AT-hook of WhiB7 interacts with the minor groove DNA and closely resembles the reported structures of the AT-hook motif in complex with the 12-bp AT-rich DNA (Fonfría-Subirós et al., 2012; Huth et al., 1997). The central RGR motif of the AT- hook lies in the minor groove of the DNA and forms extensive polar and nonpolar contacts (Figures 5A and 5B).
Figure 5. Interactions between the WhiB7:σA4-βtip complex and PwhiB7.
(A) Overview of the DNA contacts by the AT-hook of WhiB7 and the DNA binding motif σA4. Nucleotide numbering is based on the position of –35 element. The nucleotides of the DNA helix and the residues of the protein complex at the protein:DNA interface are indicated in color, using the scheme shown in the color key on the right and are described in detail in the legend of Figure S5. All the base pairs in PwhiB7 are Watson-Crick base pairs (W.C. BP) without any Hoogsteen base pairs (Hoog. BP). A break of base-base stacking is indicated by a red circle, correlated with the starting point of DNA bending shown in (D).
(B) Close-up view of interactions between the WhiB7:σA4-βtip complex and PwhiB7, with a focus on the DNA-binding motif of σA4 on the left and the AT-hook of WhiB7on the right. Residues R500 and D490 of σA4 are indicated with sticks, while the same pair of residues in the RNAP holoenzyme-DNA complex (PDB: 6EEC) are indicated with thin lines, with the carbon atoms represented in yellow.
(C) The AT-hook-dependent interaction between the WhiB7: σA4-βtip complex and PwhiB7 (sequence as indicated) as analyzed by the EMSAs.
(D) AT-hook of WhiB7-induced DNA bending indicated by overlaying of σA4 in the WhiB7: σA4-βtip:PwhiB7 structure with the the RNAP holoenzyme-DNA complex (PDB: 6EEC).
Structural comparison suggests that the AT-hook of WhiB7 binding to PwhiB7 may compensate for the reduced σA4:DNA contacts in WhiB7: σA4-βtip:PwhiB7. The surface contact area between σA4 and PwhiB7 in WhiB7:σA4-βtip PwhiB7 is estimated at 536.4 Å2 by DNAproDB (Sagendorf et al., 2020). This buried surface area is, on average, about 18% smaller than that observed in other reported structures in which σA4 (and its homologs) interacts with 35 element alone without requiring a transcription activator (e.g., 685.8 Å2 in 6EEC, 649.2 Å2 in 6EDT, and 628.0 Å2 in 1KU7) (Figure S4A; Boyaci et al., 2019; Campbell et al., 2002). One of the most noticeable differences is at the highly conserved residue R500 of σA4 with its counterpart R584 in E. coli and R409 in Thermus aquaticus (Taq) (Figure 5B; Figure S4). As shown in Figure S4A, R500 in Mtb σA4 (and its homologs) forms multiple polar interactions via its side chain with the nucleotides in the major groove of the –35 hexanucleotide region and is 6.5 Å away from D490. However, R500 in the WhiB7: σA4-βtip:PwhiB7 structure extends away from the DNA helix and forms a strong hydrogen bond with OD2 of D490 (2.2 Å) (Figure 5B; Figure S4C). The altered conformation of R500 appears to be correlated with WhiB7:σA4-βtip binding to PwhiB7, based on the structural comparison between WhiB7: σA4-βtip:PwhiB7 and WhiB7TR: σA4-βtip (Figure S4D). While the importance of R500 in transcription initiation has been well established in the previous studies (Campbell et al., 2002; Gregory et al., 2005; Siegele et al., 1989), it remains to be tested whether D490 is involved in fine-tuning transcriptional activity in coordination with a σA4-dependent transcription activator. The importance of the WhiB7 AT-hook for phoretic mobility shift assay (EMSA). As shown in Figure 5C, the interaction between the WhiB7: σA4-βtip complex and PwhiB7 strongly depends on the presence of the AT-hook in WhiB7. Deletion of the AT-hook in WhiB7 completely abolishes the DNA binding under the experimental conditions.
The WhiB7 binding site in PwhiB7 contains an A-tract, which refers to a short run of consecutive adenine-thymine base pairs (for An, n ≥ 4 for asymmetric A-tracts; or, for AnTn, n ≥ 2 for symmetric A-tracts) (Leroy et al., 1988; Marini et al., 1982; Wu and Crothers, 1984). Extensive studies in the past have shown that the structures of A-tract DNA are distinct from those of canonical B-form DNA, including the narrow minor groove, high propeller twist, and DNA bending toward the AT-rich minor groove (see the review in Haran and Mohanty, 2009). Our results from the DNA structure analyses of WhiB7:σA4-βtip:PwhiB7 suggest that the AT-hook binding to PwhiB7 may alter the DNA structure. The central axis of PwhiB7 calculated by CURVES+ slightly bends at the junction between the major and minor grooves away from the WhiB7: σA4-βtip complex, which is correlated with a break of base-base stacking between the –36 and –37 nucleotides and likely resulted from the cooperative DNA binding by WhiB7 and σA4 (Figures 5A and 5D) (Lavery et al., 2009). The DNA bending in WhiB7: σA4-βtip:PwhiB7 is evident when superimposing the structure over the RNAP holoenzyme-DNA complex (PDB: 6EEC) (Figure 5D; Boyaci et al., 2019). The direction of PwhiB7 bending is opposite to that of the expected intrinsic bending in A-tract DNA. Although we cannot completely rule out a sequence-specific effect on the DNA conformation, this observation is in line with previous reports that the AT-hook may straighten or reverse DNA bending based on the circular permutation analysis (Chen et al., 2010; Falvo et al., 1995). We also note that the central minor-groove width of PwhiB7 around the WhiB7 binding site (~7 Å) is similar to that of classical B-form DNA and the AT-hook-bound DNA, while it is significantly larger than expected for A-tract DNA (Figure S5) (Aravind and Landsman, 1998; Fonfría-Subirós et al., 2012). The narrow minor groove is one of the most fundamental properties for the AT-rich B-form DNA, of which the central minor groove width is typically smaller than 4 Å (Hizver et al., 2001; Stella et al., 2010; Yoon et al., 1988).
When superimposing the WhiB1: σA4 and WhiB7: σA4-βtip:PwhiB7 structures, it becomes evident that the C-terminal structural arrangement in WhiB7 is preferred over that in WhiB1 to place the AT-hook in a favorable conformation for DNA binding (region 2 in Figure 1E; see also Figure 6). In particular, we anticipate that the b hairpin is critical to orient the C-terminal loop toward the minor groove of the DNA helix, while the long loop between the β hairpin and the AT-hook provides the flexibility needed for the AT-hook to slide into the minor groove of the DNA when in close proximity. Additionally, the long loop between helices hw1 and hw2 of WhiB1 (region 3 in Figure 1E; see also Figure 6B) may interfere with AT-hook binding to DNA despite its flexible nature, while the shorter loop of WhiB7 in this region is optimal for its binding to target promoter DNA. To verify our hypothesis, we have designed three chimeras (v1–v3) by fusing a truncated N-terminal WhiB1 with the C-terminal WhiB7 at the designed locations (Figures 6A and 6B) and examined their ability for DNA binding in vitro and for complementing the function of WhiB7 in the ΔwhiB7 strain. As expected, exclusion of either the entire or partial sequence corresponding to the WhiB7 β hairpin (v1 and v2, respectively) disrupts the DNA binding in the EMSA and does not confer antibiotic resistance in the ΔwhiB7 strain, despite the finding that both chimeras contain the AT-hook motif and can bind to σA4 in the pull-down assays (Figures 6C and 6D; Figures S6A and S6B). In contrast, chimera v3, which contains the intact b hairpin and AT-hook of WhiB7, binds to the PwhiB7 promoter in the EMSA and partially complements the function of WhiB7 in the ΔwhiB7 strain, based on the antibiotic sensitivity test (Figures 6C and 6D; Figure S6C). We cannot conclude unambiguously from the EMSAs whether chimera v3 has a slightly lower affinity to the promoter DNA compared to the wild-type WhiB7. Upon induction in the ΔwhiB7 strain, the abundance of the chimera v3 protein is the highest among the testing samples in the western blot analysis (Figure S6D). Therefore, we can rule out the possibility that chimera v3 does not fully complement WhiB7 in the ΔwhiB7 strain because of the low level of the protein. To evaluate the impact of the middle-loop region (region 3 in Figure 1E) on the function of WhiB7, we made a modification on both chimeras v2 and v3 by replacing the peptide sequence in WhiB1 with the corresponding peptide in WhiB7. Intriguingly, this modification results in a functional chimera (chimera M-v3) that is comparable to the wild-type WhiB7 based on the antibiotic sensitivity test (Figure 6D). Chimera M-v2 also shows a marginal improvement in conferring antibiotic resistance, compared to Chimera v2 (Figure 6D). Therefore, these observations confirm our hypothesis stated earlier that the structural arrangements in the middle loop and the C terminus of WhiB7 are critical for its function.
Figure 6. Identification of the key structural motifs in WhiB7 for transcriptional activation.
(A and B) Illustration of the WhiB1-WhiB7 chimera design in the aligned protein sequences and the superimposed 3D structures of WhiB1 (pink) and WhiB7 (pale green), respectively, in complex with σA4 (gray). The residues in the AT-hook motif are indicated with sticks. σA4 in the WhiB1: σA4 complex is not shown in the figure for the clarity. Chimera v1 includes the N-terminal WhiB1 (aa 1–71) and the C-terminal AT-hook of WhiB7 (aa 78–92). Chimera v2 includes the N-terminal WhiB1 (aa 1–62) and the C-terminal peptide of WhiB7 corresponding to the second strand of the b hairpin and the AT-hook of WhiB7 (aa 69–92). Chimera v3 includes the N-terminal WhiB1 (aa 1–54) and the C-terminal peptide of WhiB7 corresponding to the intact β hairpin and the AT-hook of WhiB7 (aa 61–92). Chimeras M-v2 and M-v3 include an additional modification on chimeras v2 and v3, respectively, with a replacement of the peptide in WhiB1, highlighted in red in (B), with the corresponding peptide in WhiB7 (cyan), as indicated in the blue frame in (A). This modification is made based on the structural difference identified in region 3 of Figure 1E.
(C) EMSAs of the σA4-βtip-bound WhiB1-WhiB7 chimeras. The PwhiB7 DNA sequence used in the assays is shown in Figure 5C. All the proteins used in the assay are in complex with His6- σAC112-βtip. The wild-type WhiB7 is used as the reference. No protein sample was added in the negative control (NC).
(D) Spectinomycin sensitivity test by spotted dilutions (10–1–10–5) of Msm wild-type, the whiB7 deletion mutant (ΔwhiB7) alone or complemented by MtbWhiB7 (wild-type), MtbWhiB1 (wild-type), or a chimera as indicated.
DISCUSSION
In this work, we describe the interaction of WhiB7 with σA4 and PwhiB7 in atomic detail and reveal the structural basis for how the [4Fe-4S] cluster and the AT-hook of WhiB7 are utilized to orchestrate transcription activation with σA4. A detailed structural comparison with the σA4-bound WhiB1, complemented with molecular and biochemical approaches, uncovers the key structural motifs underlying the differences in biological functions of these two distinct subclasses of Wbl proteins.
WhiB7 is a monomeric transcription activator with a single AT- hook DNA binding motif, which is expected to have low DNA affinity and specificity by itself (Aravind and Landsman, 1998). Results from the structural analyses and EMSAs in this study indicate that WhiB7 achieves higher DNA binding affinity and specificity by forming a tight heterodimeric complex with σA4 centered in the [4Fe-4S] binding site and coordinating the binding, assuming that the free PwhiB7 has the characteristics of A-tract DNA. Although our study strongly supports that the WhiB7 AT-hook enhances σA4 binding to the target promoter, further studies are needed to elucidate the role of the AT-hook in WhiB7-dependent gene activation and determine whether the DNA binding by the WhiB7 AT-hook in synergy with RNAP σA4 facilitates the open complex formation or enhances promoter escape. In addition, the interactions between the WhiB7 AT-hook and the AT-rich minor-groove DNA resemble that of eukaryotic AT-hook-containing proteins (Bagga et al., 2000; Chase et al., 1999; Falvo et al., 1995; Fonfría-Subirós et al., 2012; Huth et al., 1997), suggesting a shared molecular mechanism of DNA binding and transcriptional regulation in eukaryotes and prokaryotes. The crystal structure of the WhiB7:σA4-βtip:PwhiB7 complex represents the first atomic view of how an AT-hook-containing protein may interact with target DNA cooperatively with its partner transcription factor; thus, it will also be instrumental for the mechanistic understanding of the transcription regulation by AT-hook-containing transcription factors in eukaryotes.
Our structural and biochemical analyses demonstrate that WhiB1 and WhiB7 share the helical structural arrangement around the [4Fe-4S] cluster binding pocket and the anchoring site on σA4 featuring a pair of invariant residues H516-P517. By structural comparison and in vivo studies, we have identified key structural determinants for a functional WhiB7. We have also defined the structural basis underlying the functional difference between WhiB1 and WhiB7. We highlight the significance of the absence of an aromatic residue in WhiB7 that is functionally equivalent to W3 in WhiB1. Our biochemical analysis supports a model in which the absence of a WhiB1-W3 counterpart takes a heavy toll on the stability of the [4Fe-4S] cluster and the WhiB7:σA4 complex. This finding is also consistent with the previous report that WhiB7 activation by antibiotics was boosted by a reducing agent, while WhiB7 was not responsive to oxidative stress induced by diamide (Burian et al., 2012). We note that the O2 sensitivity of the [4Fe-4S] cluster is dramatically reduced in the WhiB7:σA4-βtip complex, in which the conformation of σA4 represents that of the RNAP holoenzyme. This finding provides a plausible mechanism for how WhiB7, with a more O2 sensitive [4Fe-4S] cluster, maintains the cluster-dependent interaction with σA4 in the RNAP holoenzyme and activates transcription of the target genes in the event of antibiotic stress. It remains to be tested whether the WhiB7-responsive antibiotics induce the production of metabolic intermediates that stabilize the Fe-S cluster in WhiB7, either by generating an overall more reductive environment in the cell or by directly binding and sealing the Fe-S cluster binding pocket in the σA4-bound WhiB7.
The information presented in this study will also lead to a better mechanistic understanding of the functional divergence of other Wbl proteins. Most Mtb Wbl proteins bind to σA4, with the exception of WhiB2 and WhiB5, which are undetermined (Feng et al., 2016; Kudhair et al., 2017; Steyn et al., 2002; Wan et al., 2020). Our structural and biochemical studies indicate that WhiB1 and WhiB7, representing two distinct subclasses of Wbl proteins, share a unique interaction interface with σA4 that features a cluster of highly conserved hydrophobic residues, including F17, F18, and W60 in WhiB1 (corresponding to W28, F29, and W66 in WhiB7) surrounding the H516-P517 pair of σA4. These residues are highly conserved in all Mtb Wbl proteins, including WhiB2 and WhiB5 (Figure S1). Supported by structural modeling of Wbl proteins using the σA4-bound WhiB1 and WhiB7 structures as templates, it is reasonable to propose that all Mtb Wbl proteins, including WhiB2 and WhiB5, may share the same anchoring point on σA4. We are in the process of verifying this hypothesis by using structural and biochemical approaches. In addition, our success in using the σA4-βtip chimera to stabilize the WhiB7:σA4 complex for the crystallographic study establishes a feasible strategy for structural characterization of Wbl proteins in complex with σA4 at atomic resolution, which has been a major technical challenge and hinders an in-depth mechanistic understanding of the functional divergence of Wbl proteins.
Limitations of the study
Although our analyses demonstrate that the structure of WhiB7: σA4-βtip-bound PwhiB7 DNA is distinct from that of A-track B-form DNA, the flanking DNA sequence around the WhiB7 binding site and/or σA4 binding may have an impact on the DNA structure. Therefore, future studies on the structure of the free PwhiB7 DNA and in the RNAP holoenzyme-bound form are needed to elaborate the impact of WhiB7 AT-hook binding on the DNA structure of target promoters. In addition, the crystal structures reported in this study do not include RNAP. Thus, we cannot determine whether WhiB7 may also have loose contacts with other components in RNAP.
STAR★METHODS
Detailed methods are provided in the online version of this paper and include the following:
RESOURCE AVAILABILITY
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead contact, LiMei Zhang (lzhang30@unl.edu).
Materials availability
All unique/stable reagents generated in this study are available from the Lead Contact, LiMei Zhang (lzhang30@unl.edu). We may require a completed Materials Transfer Agreement if appropriate.
Data and code availability
Atomic coordinates and structure factors have been deposited in the RCSB Protein Data Bank (PDB) under the accession codes 7KUG and 7KUF for the WhiB7:σA4-βtip complex and the WhiB7: σA4-βtip:PwhiB7 complex, respectively.
EXPERIMENTAL MODEL AND SUBJECT DETAILS
E. coli strains
All the E. coli strains were grown in Luria–Bertani (LB) media and at 37°C, 200 rpm, unless otherwise specified.
Mycobacterial strains
Mycobacterium smegmatis MC2 155 (Msm) and the related mutant without and with a complementary gene generated in this study were grown in either Middlebrook 7H9 broth or on 7H10 agar (BD Difco™) supplemented with 10% (v/v) ADS (2% dextrose, 5% bovine serum albumin and 0.85% NaCl), 0.2% (v/v) glycerol, 0.05% (v/v) Tween 80 (Sigma). When appropriate, the media were supplemented with antibiotics at following concentrations: ampicillin, 100 mg/ml; hygromycin, 100 mg/ml for E. coli and 50 mg/ml for Msm; kanamycin, 50 mg/ml for E. coli and 25 mg/ml for Msm; spectinomycin, 100 mg/ml for E. coli and 12 mg/ml for Msm used in antibiotic sensitivity test.
The bacterial strains and plasmids used in this study are listed in the key resources table and Table S1.
KEY RESOURCES TABLE
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
|
| ||
| Antibodies | ||
|
| ||
| Monoclonal ANTI-FLAG M2 antibody, mouse | Sigma-Aldrich | Cat. No. F3165; RRID:AB_259529 |
| Purified anti-E.coli RNA Sigma-Aldrich 70 2G10, mouse | Biolegend | Cat. No. 663208; RRID: AB_2814499 |
| Peroxidase-AffiniPure Goat Anti-Mouse antibody | Jackson Immune Research | Cat. No.:115–035-068; RRID: AB_2338505 |
|
| ||
| Bacterial and virus strains | ||
|
| ||
| E.coli, BL21-Gold (DE3) | Agilent | 230132 |
| E.coli, XL1-Blue | Agilent | 200249 |
| Mycobacterium smegmatis MC2 155 | ATCC | 700084 |
| Mycobacterium smegmatis MC2 155 ΔwhiB7 | This study | N/A |
|
| ||
| Chemicals, peptides, and recombinant proteins | ||
|
| ||
| Ampicillin | Research Products International | A40040 |
| Kanamycin Monosulfate | GoldBio | K-120–25 |
| Spectinomycin Dihydrochloride | GoldBio | S-140–50 |
| Hygromycin B | GoldBio | H-270–1 |
| L-Selenomethionine | TCI America | 3211–76-5 |
| HisTrap™ High Performance | GE Healthcare | 17–5248-02 |
| HiLoad 16/600 Superdex 200 | GE Healthcare | 28–9893-35 |
|
| ||
| Critical commercial assays | ||
|
| ||
| Chemiluminescent Nucleic Acid Detection Module | Thermo scientific | 89880 |
| LightShift Chemiluminescent EMSA Kit | Thermo scientific | 20148 |
|
| ||
| Deposited data | ||
|
| ||
| Crystal structure of WhiB7TR: σA4-βtip complex | This paper | PDB: 7KUG |
| Crystal structure of WhiB7TR: σA4-βtip:PwhiB7 complex | This paper | PDB: 7KUF |
|
| ||
| Oligonucleotides | ||
|
| ||
| Pwhib7 for EMSA forward: GGTCAGAAAATCGGTTGTGGTC | This paper | N/A |
| PwhiB7 for EMSA reverse: CCAGTCTTTTAGCCAACACCAG | This paper | N/A |
| PwhiB7 for crystallization forward: CAGAAAATCGGTTGTGGT | This paper | N/A |
| PwhiB7 for crystallization reverse: TCTTTTAGCCAACACCAG | This paper | N/A |
|
| ||
| Recombinant DNA | ||
|
| ||
| pET-21b(+) | Novagen | 68741 |
| pET-28a | Novagen | 69864 |
| pKW08-Lx-Int | Addgene | 25015 |
| pCDF-1b | Novagen | 71330 |
| pETDuet-1 | Novagen | 71146 |
| pJV53-GFP | (Williams et al., 2010) | N/A |
| pUC-Hyg | (Williams et al., 2010) | N/A |
| Plasmids for expression of the C-terminal σA proteins in E. coli (see Table S1) | This paper | N/A |
| Continued REAGENT or RESOURCE |
SOURCE | IDENTIFIER |
| Plasmids for expression of the σA4-βtip proteins in E. coli (see Table S1) | This paper | N/A |
| Plasmids for expression of WhiBI in E. coli (see Table S1) | (Wan et al., 2020) | N/A |
| Plasmids for expression of the WhiB7 proteins in E. coli (see Table S1) | This paper | N/A |
| Plasmids for expression of the WhiBI-WhiB7 Chimera proteins in E. coli (see Table S1) | This paper | N/A |
| Plasmids for expression of the Wbl proteins in Msm (see Table S1) | This paper | N/A |
|
| ||
| Software and algorithms | ||
|
| ||
| XDS | (Kabsch, 2010) | https://xds.mr.mpg.de/ |
| HKL3000 | (Otwinowski and Minor, 1997) | https://www.hkl-xray.com/ |
| Phenix 1.15.2–3472 | (Adams et al., 2010) | https://phenix-online.org/ |
| COOT | (Emsley et al., 2010) | https://www2.mrc-lmb.cam.ac.uk/personal/pemsley/coot/ |
| CURVES+ | (Lavery et al., 2009) | https://bisi.ibcp.fr/tools/curves_plus/ |
| w3DNA | (Li et al., 2019) | http://web.x3dna.org/ |
| PDBePISA | (Krissinel and Henrick, 2007) | https://www.ebi.ac.uk/pdbe/pisa/ |
| DNAproDB | (Sagendorf et al., 2017) | https://dnaprodb.usc.edu/ |
| PyMol v2.3 | N/A | https://pymol.org/2/ |
| Caver 3.0 | (Chovancova et al., 2012) | https://www.caver.cz/ |
| Clustal Omega | (Sievers and Higgins, 2018) | https://www.ebi.ac.uk/Tools/msa/clustalo/ |
| ESpript 3.0 | (Robert and Gouet, 2014) | https://espript.ibcp.fr |
| WebLogo 3 | (Crooks et al., 2004) | https://weblogo.berkeley.edu/ |
METHOD DETAILS
Cloning for protein expression in E. coli
The genes encoding WhiB7 (Rv3197A, 1–92 aa) and the C-terminal domain of σA4 (Rv2703) containing M359–D528 of the protein (de- noted σAC170) were amplified by PCR from Mtb H37Rv genomic DNA (a gift from Dr. Midori Kato-Maeda’s group at the University of California, San Francisco), and subsequently cloned into pET21b(+) and pCDF-1b to express tagless WhiB7 and His6- σAC170, respectively. The resulting plasmids, pET21-MtbWhiB7 for WhiB7 and pCDF-1b-6HisMtb σAC170 for His6- σAC170, respectively, were subsequently modified for expression of the proteins with either truncation or point mutation used for crystallization and biochemical as says as described in the related sections. To confirm the expression of the WhiB7 mutants, the whiB7 gene (wild-type or mutant) was amplified from pET21-MtbWhiB7 (wild-type or mutant) by PCR into pETDuet-His6-SUMO vector (modified from pETDuet). The gene encoding the last 82 residues in σA4 with a N-terminal Strep-tag, denoted nStrep- σAC82, was amplified from pCDF-1b-6HisMtb σAC170 and cloned into pET28a(+) to express nStrep- σAC82. For expression and purification of the tagless MtbWhiB1-WhiB7 chimeras in E. coli, the DNA fragment encoding the N-terminal WhiB1 and the C-terminal WhiB7 at the designed position cloned into the NdeI-HindIII site of pET21b(+).
The pET21b-MtbWhiB7 plasmid was modified by site-directed mutagenesis to generate the pET21b-MtbWhiB7TR plasmid, which encodes a truncated WhiB7 (containing aa 1–79) without the AT-hook for the crystallographic work and the electrophoretic mobility shift assays (EMSA). To improve the stability of the protein complexes for the EMSA assays and the O2 sensitivity test, the pET28b-6HisMtb σAC112-βtip plasmid was constructed to express a chimera protein, denoted σAC112-βtip, by fusing σAC112 (containing σA4, aa 416–528) with the RNAP β-subunit flap tip helix (βtip, containing aa 815–829) linked by the artificial linker (GSSGSGC). The pET28b- 6HisMtb σAC822-βtip plasmid was constructed to express a similar chimera protein with a shorter C-terminal domain of σA (containing aa 446–528 of σA), denoted σAC82-βtip, for the crystallographic work. The pET28b-6HisMtb σAC82-βtip plasmid encoding His6- σAC82-βtip was then co-transformed with either pET21b-MtbWhiB7TR or the pET21b-MtbWhiB7 plasmid to express the WhiB7TR: σAC82-βtip and WhiB7: σAC82-βtip complexes, respectively. Two point-mutations, namely L27M and L42M of WhiB7, were introduced in the pET21-MtbWhiB7TR plasmid for experimental phasing in the X-ray crystallographic study.
The full list of plasmids used in this study can be found in the key resources table and Table S1. All the constructed plasmids were confirmed by DNA sequencing.
Protein expression and purification
The proteins used for structural and biochemical studies were overexpressed in E. coli BL21-Gold (DE3) as previously described (Wan et al., 2020). The selenomethionine (SeMet)-substituted protein was produced in E. coli BL21-Gold (DE3) using the metabolic inhibition method as previously described (Doublie, 1997). Cells were collected by centrifugation and kept at –80°C for storage.
Affinity purification was performed using a HisTrap HP column or loose NiNTA-agarose resin (GE Healthcare Life Sciences) at room temperature under strict anaerobic conditions as previously described with minor modification (Wan et al., 2020). For the purification of the WhiB7: σAC82-βtip complex, a 10 column-volume wash with 50 mM Tris-HCl pH 8.0, 1M NaCl, 1mM DTT was added in the purification procedures to remove the DNA contamination before elution. Imidazole in the eluted fractions was removed by passing through either a desalting column or a Superdex 200 column (GE Healthcare Life Sciences). The samples after each step of purification were analyzed by SDS-PAGE and by UV-Visible (UV-Vis) spectroscopy. Unless otherwise specified, the final purified proteins in 50 mM Tris-HCl, pH 8.0, 100 mM NaCl, 1mM DTT were stored in liquid nitrogen until use. Protein concentrations were estimated by the Pierce Bradford Assay Kit (Thermo Fisher Scientific) and by the absorption at 280 nm.
UV-Visible spectroscopy
The UV-Visible spectra of the purified protein were recorded using an HP 8452a diode array UV-Visible spectrophotometer (Agilent Technologies Inc.). The absorption at 410 nm, characteristic of proteins containing [4Fe-4S]2+ clusters (Crack et al., 2011; Johnson, 1998), was used to estimate the occupancy of the Fe-S cluster in the protein samples containing a Wbl protein.
Pull-down assays
The interactions between WhiB7 (wild-type and mutants) and His6-σAC170 (wild-type and mutants) were tested by the co-expression and affinity purification assay to verify the effect of single mutations in either WhiB7 or σAC170 on the interaction of the two proteins. The two plasmids containing the genes encoding His6-σAC170 and tagless WhiB7 (either wild-type or mutants), respectively, were co-transformed in E. coli BL21-Gold (DE3) strain. Protein co-expression and affinity purification by Ni-NTA Sepharose resin was carried out as described above. The purified proteins were analyzed by UV-Visible absorption spectroscopy and SDS-PAGE. To confirm that these WhiB7 mutants that do not bind to σA4 were expressed in E. coli, the two plasmids containing the genes encoding His6-SUMO-WhiB7 (wild-type and mutants) and nStrep-σAC82, respectively, were co-transformed in E. coli BL21-Gold (DE3) strain. Protein co-expression and affinity purification were carried out as described above.
O2 sensitivity test
UV-Vis spectroscopy was used to monitor the [4Fe-4S] cluster loss in the WhiB1:σAC170,WhiB7:σAC170 and WhiB7:σAC112-βtip complexes under aerobic conditions. DTT in the purified protein sample was removed by passing through a desalting column. The resulting protein sample at ~400 uM was then diluted in 1:10 ratio with air saturated Tris buffer (50 mM Tris-HCl, pH 8.0, 100 mM NaCl) at room temperature (~20°C) under aerobic conditions and divided into 250-ul aliquots. The UV-Visible spectra were collected from an aliquot every 15 min for the first 90 minutes, and then by 2 hr and 24 hr under aerobic conditions. The aliquoted samples were centrifuged at 14,000 rpm, 4°C for 5 min to remove the aggregated protein due to the cluster loss before the measurement. As a control, the same protein sample was diluted with the O2-free buffer in the anaerobic chamber, and the UV-Visible spectrum was collected by 24 hr kept under anaerobic conditions.
Electrophoretic mobility shift assay
22-bp synthetic double-stranded oligos (5ʹ-GGTCAGAAAATCGGTTGTGGTC-3ʹ) covering an AT-rich region and –35 element from the whib7 promoter were used as probes. The EMSA experiments were performed using LightShift Chemiluminescent EMSA Kit (Thermo Fisher Scientific) as previously described (Wan et al., 2020). The biotin-labeled probe (0.2 nM) was incubated with either WhiB7: σAC112-βtip (with AT-hook), WhiB7TR: σAC112-βtip (without AT-hook) or the σAC112-βtip-bound MtbWhiB1-WhiB7 chimera proteins as indicated, in the presence of 100 mM NaCl, 40 mM Tris (pH 8.0), 10 mM MgCl2, 1 mM EDTA, 0.25 mg/ml BSA, 1 mM DTT in a total volume of 20 μl. 50 ng/μl Poly (dI•dC) was included in all reactions as a non-specific competitor. After incubation on ice for 20 min, the reaction mixture was subjected to electrophoretic separation in 10% native polyacrylamide gels at 100 V in 0.5 3 TBE buffer. DNA and DNA-protein complexes in the gel were transferred to Hybond-N+ nylon membrane (GE Healthcare Life Sciences) and UV cross-linked to the membrane. The biotin-labeled DNA was detected using the LightShift Chemiluminescent EMSA Kit according to the manufacturer’s instructions.
Creation of a whiB7 deletion mutant in Msm
The whiB7 (MSMEG_1953) deletion mutant (ΔwhiB7) in Msm was generated using homologous recombination-based in-frame unmarked deletion as previously described (Mao et al., 2016; Wan et al., 2020). The resulting ΔwhiB7 strain was verified by PCR and further confirmed by DNA sequencing.
Complementation test in the ΔwhiB7 strain
For the heterologous complementation test in the ΔwhiB7 strain, the DNA fragment encoding the 3FLAG-tagged MtbWhiB7 (wild-type or mutant) or MtbWhiB1-WhiB7 chimeras was cloned into the integration plasmid pKW08-Lx-Int for expression of 3×FLAG- MtbWhiB7 from the inducible tetRO promoter (Williams et al., 2010). The resulting plasmid was transformed and integrated in the genome of the ΔwhiB7 strain for the expression of WhiB7 (wild-type, mutant or chimera) under the tetRO promoter. The expression of 3×FLAG-MtbWhiB7 (wild-type, mutant or chimera) was induced by anhydrotetracycline (aTc). The plasmid encoding 3FLAG- tagged MtbWhiB1 was constructed in the same way and was included in the study as a negative control.
Antibiotic sensitivity test
Overnight cultures of the Msm strains were subcultured in 7H9 broth containing 50 ug/mL hygromycin with the initial OD600nm of 0.05, and incubated at 37°C, 200 rpm until OD600nm of 0.4~0.6. The wild-type Msm and the ΔwhiB7 strain were grown under the same conditions without hygromycin and used as controls. All the strains were then normalized to OD600nm of 0.4. 3 ul of culture from ten-fold serial dilutions were spotted on 7H10 plates containing 12 ug/ml spectinomycin and 200 ng/ml aTc. Images were taken after 72h incubation at 37°C.
Western blot analysis
Overnight cultures of the Msm strains were subcultured in 7H9 broth containing 200 ug/mL aTc and 50 ug/ml hygromycin with the initial OD600nm of 0.05, and incubated at 37°C, 200 rpm until OD600nm of 1.0~1.5. The wild-type Msm and the ΔwhiB7 strain were grown under the same conditions without hygromycin and used as controls. All the strains were then normalized to the same OD600nm and collected by centrifugation. The cells were re-suspended with a buffer containing 50 mM Tris pH8.0, 100 mM NaCl and 5 mM DTT, and then lysed using FastPrep-24 (MP biomedicals). The 3FLAG-tagged Wbl proteins were detected by chemiluminescence using mouse anti-FLAG M2 antibody (Sigma-Aldrich) as the primary antibody and the horseradish peroxidase-coupled goat anti-mouse IgG antibody (Jackson ImmunoResearch Laboratories) as the secondary antibody in the western blot analysis. The mouse anti-RNA Sigma70 antibody (Biolegend, Clone 2G10) was used as the primary antibody for the detection of the native Msm σA, which was used as an internal reference for total protein loading control.
Crystallization
Initial crystallization screens of the WhiB7TR:σAC82-βtip complex were carried out at 18°C in a Coy anaerobic chamber using the sitting-drop vapor diffusion method, followed by optimization of the crystallization hits. The high-quality crystals were obtained by mixing 1 ml of WhiB7TR: σAC82-βtip at 80 mg/ml with an equal volume of the reservoir solution containing 0.1 M Tris pH 8.0, 10%–15% PEG8000, 2%–5% PGA.
18-bp synthetic PwhiB7 duplex DNA with a 5ʹ G/C overhang at each end (5ʹ-CAGAAAATCGGTTGTGGT-3ʹ/5ʹ-TCTTTTAGCCAA- CACCAG-3ʹ, Sigma-Aldrich) was used for co-crystallization with the WhiB7: σAC82-βtip complex to determine the WhiB7: σAC82-βtip:PwhiB7 structure based on the previously reported consensus DNA sequence of WhiB7 (Burian et al., 2012). The WhiB7: σA4-βtip complex was first mixed in a 1:1 molar ratio with 100 uM PwhiB7, and then concentrated to 40 mg/ml before crystallization. The best crystals were obtained by mixing 1 ul of the complex with the reservoir solution containing 0.2–0.3 M NH4H2PO4. All the crystals were briefly soaked in the reservoir solution with 20% glycerol for cryoprotection before flash-cooling in liquid nitrogen.
X-ray diffraction data collection
X-ray diffraction data were collected at beamlines 9–2 and 12–2 of the Stanford Synchrotron Radiation Light Source from single crystals maintained at 100 K using a 6M Pixel Array Detector. Taking use of the large size of the crystals, two sets of diffraction data were collected at two different spots on the same crystal at the Se K-edge absorption peak (12657.6 eV) to achieve high redundancy. For the WhiB7TR: σAC82-βtip crystal, 1440 diffraction images were collected over 720° in each set of data. For the WhiB7: σAC82-βtip:PwhiB7 crystal, 720 diffraction images were collected over 360° in each dataset.
Structural determination and analysis
The diffraction data of WhiB7TR: σA4-βtip and WhiB7: σA4-βtip:PwhiB7 were processed using XDS and HKL2000, respectively (Kabsch, 2010; Otwinowski and Minor, 1997). The two sets of the data were merged together for the data processing. For the WhiB7TR: σA4-βtip structure, the phase information was obtained from the SAD data using Phenix.Autosol (Adams et al., 2010). The subsequent model building and structure refinement were carried out using COOT and Phenix, respectively (Adams et al., 2010; Emsley et al., 2010). The final structure was refined at 1.55 Å, with two copies of the complex molecules per asymmetric unit, which are essentially identical except for the terminal residues and those around the linker. A fully occupied [4Fe-4S] cluster is included in both complexes. The phase for the WhiB7: σA4-βtip:PwhiB7 structure was determined by molecular replacement using the WhiB7TR: σA4-βtip structure as the search model. Phenix.autobuid was then used to build the first partial model of the PwhiB7 DNA. The assignment of the rest nucleotides was done manually, followed by multiple cycles of refinements using COOT and PHENIX, respectively (Adams et al., 2010; Emsley et al., 2010). The conserved –35 element in the σA4:DNA complex (PDB ID: 1KU7, at 2.4 Å) was used as a guide for the nucleotide assignment in the corresponding region (Campbell et al., 2002). The final structure was determined at 2.6 Å. The data collection and refinement statistics for both structures are summarized in Table 1.
Data visualization
Sequence alignments were performed using Clustal Omega and ESpript online server (https://espript.ibcp.fr), and the sequence logo was generated using WebLogo (Crooks et al., 2004; Robert and Gouet, 2014; Sievers and Higgins, 2018). The WhiB1 and WhiB7 subclass sequences used for the alignments are modified from the previous study by Chandra et al. (Chandra and Chater, 2014), and listed in the legend of Figure S1. 3D structural figures were prepared with the PyMol Molecular Graphics System v2.3 (https://pymol.org/2/). Protein-DNA interaction diagrams were generated by the online server DNAproDB (Sagendorf et al., 2017).
QUANTIFICATION AND STATISTICAL ANALYSIS
Statistical validation for the final models deposited in the RCSB Protein Data Bank (accession codes 7KUG and 7KUF) were performed using Phenix (Adams et al., 2010). Tunnel dimensions were calculated using the Caver 3.0 software with probe radii indicated in the figure (Chovancova et al., 2012). The DNA structural analyses were calculated using Curves+ and w3DNA (Lavery et al., 2009; Li et al., 2019; Zheng et al., 2009). Protein-protein interface were calculated using the online server PDBePISA (Krissinel and Henrick, 2007).
Supplementary Material
Highlights.
WhiB7 binds to the same site of the primary sigma factor σA as WhiB1
Fe-S cluster of WhiB7 is stable to O2 upon binding to s in the RNAP holoenzyme
AT-hook of WhiB7 enhances DNA binding and induces DNA conformational changes
Structural arrangements of WhiB7 are optimal for AT-hook binding to DNA
ACKNOWLEDGMENTS
We thank Dr. M.A. Wilson for thoughtful feedback on the manuscript, Dr. Elizabeth Campbell at the Rockefeller University for insightful discussions on the WhiB7-dependent transcriptional regulation, Dr. Wei Xie at the Sloan Kettering Institute for assistance with the structural refinement of the WhiB7:σA4-βtip:PwhiB7 complex, Dr. M. Kato-Maeda for the Mtb H37Rv genomic DNA, and Dr. Y. Sun for the plasmids used for site-directed mutagenesis in Msm. The authors thank the staff at beamlines 9-2 and 12-2 of SSRL for assistance during the X-ray diffraction data collection.
This work was supported in part by the National Institutes of Health (grant R35 GM138157-01 to L.-M.Z.) and by a National Science Foundation CAREER award (CLP 1846908 to L.-M.Z.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Use of the Stanford Synchrotron Radiation Lightsource (SSRL), SLAC National Accelerator Laboratory, is supported by the U.S. Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences, under contract no. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research and by the National Institute of General Medical Sciences, National Institutes of Health (including grant P41GM103393).
Footnotes
DECLARATION OF INTERESTS
The authors declare no competing interests.
INCLUSION AND DIVERSITY
One or more of the authors of this paper self-identifies as an underrepresented ethnic minority in science.
SUPPLEMENTAL INFORMATION
Supplemental information can be found online at https://doi.org/10.1016/j.molcel.2021.06.002.
REFERENCES
- Adams PD, Afonine PV, Bunkóczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse-Kunstleve RW, et al. (2010). PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr 66, 213–221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alam MS, Garg SK, and Agrawal P. (2009). Studies on structural and functional divergence among seven WhiB proteins of Mycobacterium tuberculosis H37Rv. FEBS J. 276, 76–93. [DOI] [PubMed] [Google Scholar]
- Aravind L, and Landsman D. (1998). AT-hook motifs identified in a wide variety of DNA-binding proteins. Nucleic Acids Res. 26, 4413–4421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bagga R, Michalowski S, Sabnis R, Griffith JD, and Emerson BM (2000). HMG I/Y regulates long-range enhancer-dependent transcription on DNA and chromatin by changes in DNA topology. Nucleic Acids Res. 28, 2541–2550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blanco AG, Canals A, Bernués J, Solà M, and Coll M. (2011). The structure of a transcription activation subcomplex reveals how σ70 is recruited to PhoB promoters. EMBO J. 30, 3776–3785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boyaci H, Chen J, Jansen R, Darst SA, and Campbell EA (2019). Structures of an RNA polymerase promoter melting intermediate elucidate DNA unwinding. Nature 565, 382–385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burian J, and Thompson CJ (2018). Regulatory genes coordinating antibiotic-induced changes in promoter activity and early transcriptional termination of the mycobacterial intrinsic resistance gene whiB7. Mol. Microbiol 107, 402–415. [DOI] [PubMed] [Google Scholar]
- Burian J, Ramón-García S, Sweet G, Gómez-Velasco A, Av-Gay Y, and Thompson CJ (2012). The mycobacterial transcriptional regulator whiB7 gene links redox homeostasis and intrinsic antibiotic resistance. J. Biol. Chem 287, 299–310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burian J, Yim G, Hsing M, Axerio-Cilies P, Cherkasov A, Spiegelman GB, and Thompson CJ (2013). The mycobacterial antibiotic resistance determinant WhiB7 acts as a transcriptional activator by binding the primary sigma factor SigA (RpoV). Nucleic Acids Res. 41, 10062–10076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bush MJ (2018). The actinobacterial WhiB-like (Wbl) family of transcription factors. Mol. Microbiol 110, 663–676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campbell EA, Muzzin O, Chlenov M, Sun JL, Olson CA, Weinman O, Trester-Zedlitz ML, and Darst SA (2002). Structure of the bacterial RNA polymerase promoter specificity sigma subunit. Mol. Cell 9, 527–539. [DOI] [PubMed] [Google Scholar]
- Casonato S, Cervantes Sánchez A, Haruki H, Rengifo González M, Provvedi R, Dainese E, Jaouen T, Gola S, Bini E, Vicente M, et al. (2012). WhiB5, a transcriptional regulator that contributes to Mycobacterium tuberculosis virulence and reactivation. Infect. Immun 80, 3132–3144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chandra G, and Chater KF (2014). Developmental biology of Streptomyces from the perspective of 100 actinobacterial genome sequences. FEMS Microbiol. Rev 38, 345–379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chase MB, Haga SB, Hankins WD, Williams DM, Bi Z, Strovel JW, Obriecht C, and Berg PE (1999). Binding of HMG-I(Y) elicits structural changes in a silencer of the human beta-globin gene. Am. J. Hematol 60, 27–35. [DOI] [PubMed] [Google Scholar]
- Chawla M, Parikh P, Saxena A, Munshi M, Mehta M, Mai D, Srivastava AK, Narasimhulu KV, Redding KE, Vashi N, et al. (2012). Mycobacterium tuberculosis WhiB4 regulates oxidative stress response to modulate survival and dissemination in vivo. Mol. Microbiol 85, 1148–1165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chawla M, Mishra S, Anand K, Parikh P, Mehta M, Vij M, Verma T, Singh P, Jakkala K, Verma HN, et al. (2018). Redox-dependent condensation of the mycobacterial nucleoid by WhiB4. Redox Biol. 19, 116–133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen B, Young J, and Leng F. (2010). DNA bending by the mammalian high-mobility group protein AT hook 2. Biochemistry 49, 1590–1595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Z, Hu Y, Cumming BM, Lu P, Feng L, Deng J, Steyn AJ, and Chen S. (2016). Mycobacterial WhiB6 differentially regulates ESX-1 and the Dos regulon to modulate granuloma formation and virulence in Zebrafish. Cell Rep. 16, 2512–2524. [DOI] [PubMed] [Google Scholar]
- Chovancova E, Pavelka A, Benes P, Strnad O, Brezovsky J, Kozlikova B, Gora A, Sustr V, Klvana M, Medek P, et al. (2012). CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput. Biol 8, e1002708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crack JC, Smith LJ, Stapleton MR, Peck J, Watmough NJ, Buttner MJ, Buxton RS, Green J, Oganesyan VS, Thomson AJ, and Le Brun NE (2011). Mechanistic insight into the nitrosylation of the [4Fe-4S] cluster of WhiB-like proteins. J. Am. Chem. Soc 133, 1112–1121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crack JC, Munnoch J, Dodd EL, Knowles F, Al Bassam MM, Kamali S, Holland AA, Cramer SP, Hamilton CJ, Johnson MK, et al. (2015). NsrR from Streptomyces coelicolor is a nitric oxide-sensing [4Fe-4S] cluster protein with a specialized regulatory function. J. Biol. Chem 290, 12689–12704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crooks GE, Hon G, Chandonia JM, and Brenner SE (2004). WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis NK, and Chater KF (1992). The Streptomyces coelicolor whiB gene encodes a small transcription factor-like protein dispensable for growth but essential for sporulation. Mol. Gen. Genet 232, 351–358. [DOI] [PubMed] [Google Scholar]
- Doublie S. (1997). Preparation of selenomethionyl proteins for phase determination. Methods Enzymol. 276, 523–530. [PubMed] [Google Scholar]
- Emsley P, Lohkamp B, Scott WG, and Cowtan K. (2010). Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr 66, 486–501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falvo JV, Thanos D, and Maniatis T. (1995). Reversal of intrinsic DNA bends in the IFN beta gene enhancer by transcription factors and the architectural protein HMG I(Y). Cell 83, 1101–1111. [DOI] [PubMed] [Google Scholar]
- Feng L, Chen Z, Wang Z, Hu Y, and Chen S. (2016). Genome-wide characterization of monomeric transcriptional regulators in Mycobacterium tuberculosis. Microbiology (Reading) 162, 889–897. [DOI] [PubMed] [Google Scholar]
- Fonfría-Subirós E, Acosta-Reyes F, Saperas N, Pous J, Subirana JA, and Campos JL (2012). Crystal structure of a complex of DNA with one AT-hook of HMGA1. PLoS ONE 7, e37120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geszvain K, Gruber TM, Mooney RA, Gross CA, and Landick R. (2004). A hydrophobic patch on the flap-tip helix of E.coli RNA polymerase mediates σ70 region 4 function. J. Mol. Biol 343, 569–587. [DOI] [PubMed] [Google Scholar]
- Gomez M, Doukhan L, Nair G, and Smith I. (1998). sigA is an essential gene in Mycobacterium smegmatis. Mol. Microbiol 29, 617–628. [DOI] [PubMed] [Google Scholar]
- Gregory BD, Nickels BE, Darst SA, and Hochschild A. (2005). An altered-specificity DNA-binding mutant of Escherichia coli sigma70 facilitates the analysis of sigma70 function in vivo. Mol. Microbiol 56, 1208–1219. [DOI] [PubMed] [Google Scholar]
- Haran TE, and Mohanty U. (2009). The unique structure of A-tracts and intrinsic DNA bending. Q. Rev. Biophys 42, 41–81. [DOI] [PubMed] [Google Scholar]
- Hizver J, Rozenberg H, Frolow F, Rabinovich D, and Shakked Z. (2001). DNA bending by an adenine~thymine tract and its role in gene regulation. Proc. Natl. Acad. Sci. USA 98, 8490–8495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Honda S, Bicocca VT, Gessaman JD, Rountree MR, Yokoyama A, Yu EY, Selker JML, and Selker EU (2016). Dual chromatin recognition by the histone deacetylase complex HCHC is required for proper DNA methylation in Neurospora crassa. Proc. Natl. Acad. Sci. USA 113, E6135–E6144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huth JR, Bewley CA, Nissen MS, Evans JN, Reeves R, Gronenborn AM, and Clore GM (1997). The solution structure of an HMG-I(Y)-DNA complex defines a new architectural minor groove binding motif. Nat. Struct. Biol 4, 657–665. [DOI] [PubMed] [Google Scholar]
- Johnson MK (1998). Iron-sulfur proteins: new roles for old clusters. Curr. Opin. Chem. Biol 2, 173–181. [DOI] [PubMed] [Google Scholar]
- Kabsch W. (2010). Xds. Acta Crystallogr D Biol Crystallogr 66, 125–132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiley PJ, and Beinert H. (1998). Oxygen sensing by the global regulator, FNR: the role of the iron-sulfur cluster. FEMS Microbiol. Rev 22, 341–352. [DOI] [PubMed] [Google Scholar]
- Kohanski MA, Dwyer DJ, and Collins JJ (2010). How antibiotics kill bacteria: from targets to networks. Nat. Rev. Microbiol 8, 423–435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krissinel E, and Henrick K. (2007). Inference of macromolecular assemblies from crystalline state. J. Mol. Biol 372, 774–797. [DOI] [PubMed] [Google Scholar]
- Kudhair BK, Hounslow AM, Rolfe MD, Crack JC, Hunt DM, Buxton RS, Smith LJ, Le Brun NE, Williamson MP, and Green J. (2017). Structure of a Wbl protein and implications for NO sensing by M. tuberculosis. Nat. Commun 8, 2280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuznedelov K, Minakhin L, Niedziela-Majka A, Dove SL, Rogulja D, Nickels BE, Hochschild A, Heyduk T, and Severinov K. (2002). A role for interaction of the RNA polymerase flap domain with the sigma subunit in promoter recognition. Science 295, 855–857. [DOI] [PubMed] [Google Scholar]
- Larsson C, Luna B, Ammerman NC, Maiga M, Agarwal N, and Bishai WR (2012). Gene expression of Mycobacterium tuberculosis putative transcription factors whiB1–7 in redox environments. PLoS ONE 7, e37516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lavery R, Moakher M, Maddocks JH, Petkeviciute D, and Zakrzewska K. (2009). Conformational analysis of nucleic acids revisited: Curves+. Nucleic Acids Res. 37, 5917–5929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee DJ, Minchin SD, and Busby SJ (2012). Activating transcription in bacteria. Annu. Rev. Microbiol 66, 125–152. [DOI] [PubMed] [Google Scholar]
- Leroy JL, Charretier E, Kochoyan M, and Guéron M. (1988). Evidence from base-pair kinetics for two types of adenine tract structures in solution: their relation to DNA curvature. Biochemistry 27, 8894–8898. [DOI] [PubMed] [Google Scholar]
- Li S, Olson WK, and Lu XJ (2019). Web 3DNA 2.0 for the analysis, visualization, and modeling of 3D nucleic acid structures. Nucleic Acids Res. 47 (W1), W26–W34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mao XJ, Yan MY, Zhu H, Guo XP, and Sun YC (2016). Efficient and simple generation of multiple unmarked gene deletions in Mycobacterium smegmatis. Sci. Rep 6, 22922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marini JC, Levene SD, Crothers DM, and Englund PT (1982). Bent helical structure in kinetoplast DNA. Proc. Natl. Acad. Sci. USA 79, 7664–7668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morris RP, Nguyen L, Gatfield J, Visconti K, Nguyen K, Schnappinger D, Ehrt S, Liu Y, Heifets L, Pieters J, et al. (2005). Ancestral antibiotic resistance in Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 102, 12200–12205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morrison EA, Sanchez JC, Ronan JL, Farrell DP, Varzavand K, Johnson JK, Gu BX, Crabtree GR, and Musselman CA (2017). DNA binding drives the association of BRG1/hBRM bromodomains with nucleosomes. Nat. Commun 8, 16080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otwinowski Z, and Minor W. (1997). Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326. [DOI] [PubMed] [Google Scholar]
- Partridge JD, Bodenmiller DM, Humphrys MS, and Spiro S. (2009). NsrR targets in the Escherichia coli genome: new insights into DNA sequence requirements for binding and a role for NsrR in the regulation of motility. Mol. Microbiol 73, 680–694. [DOI] [PubMed] [Google Scholar]
- Raghunand TR, and Bishai WR (2006). Mycobacterium smegmatis whmD and its homologue Mycobacterium tuberculosis whiB2 are functionally equivalent. Microbiology (Reading) 152, 2735–2747. [DOI] [PubMed] [Google Scholar]
- Ramón-García S, Ng C, Jensen PR, Dosanjh M, Burian J, Morris RP, Folcher M, Eltis LD, Grzesiek S, Nguyen L, and Thompson CJ (2013). WhiB7, an Fe-S-dependent transcription factor that activates species-specific repertoires of drug resistance determinants in actinobacteria. J. Biol. Chem 288, 34514–34528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reeves AZ, Campbell PJ, Sultana R, Malik S, Murray M, Plikaytis BB, Shinnick TM, and Posey JE (2013). Aminoglycoside cross-resistance in Mycobacterium tuberculosis due to mutations in the 5ʹ untranslated region of whiB7. Antimicrob. Agents Chemother 57, 1857–1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robert X, and Gouet P. (2014). Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodríguez J, Mosquera J, Couceiro JR, Vázquez ME, and Mascareñas JL (2015). The AT-Hook motif as a versatile minor groove anchor for promoting DNA binding of transcription factor fragments. Chem. Sci. (Camb.) 6, 4767–4771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sagendorf JM, Berman HM, and Rohs R. (2017). DNAproDB: an interactive tool for structural analysis of DNA-protein complexes. Nucleic Acids Res. 45 (W1), W89–W97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sagendorf JM, Markarian N, Berman HM, and Rohs R. (2020). DNAproDB: an expanded database and web-based tool for structural analysis of DNA-protein complexes. Nucleic Acids Res. 48 (D1), D277–D287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saini V, Farhana A, and Steyn AJ (2012). Mycobacterium tuberculosis WhiB3: a novel iron-sulfur cluster protein that regulates redox homeostasis and virulence. Antioxid. Redox Signal 16, 687–697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siegele DA, Hu JC, Walter WA, and Gross CA (1989). Altered promoter recognition by mutant forms of the sigma 70 subunit of Escherichia coli RNA polymerase. J. Mol. Biol 206, 591–603. [DOI] [PubMed] [Google Scholar]
- Sievers F, and Higgins DG (2018). Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 27, 135–145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith LJ, Stapleton MR, Fullstone GJ, Crack JC, Thomson AJ, Le Brun NE, Hunt DM, Harvey E, Adinolfi S, Buxton RS, and Green J. (2010). Mycobacterium tuberculosis WhiB1 is an essential DNA-binding protein with a nitric oxide-sensitive iron-sulfur cluster. Biochem. J 432, 417–427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith LJ, Stapleton MR, Buxton RS, and Green J. (2012). Structure-function relationships of the Mycobacterium tuberculosis transcription factor WhiB1. PLoS ONE 7, e40407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soliveri JA, Gomez J, Bishai WR, and Chater KF (2000). Multiple paralogous genes related to the Streptomyces coelicolor developmental regulatory gene whiB are present in Streptomyces and other actinomycetes. Microbiology (Reading) 146, 333–343. [DOI] [PubMed] [Google Scholar]
- Stella S, Cascio D, and Johnson RC (2010). The shape of the DNA minor groove directs binding by the DNA-bending protein Fis. Genes Dev. 24, 814–826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart MYY, Bush MJ, Crack JC, Buttner MJ, and Le Brun NE (2020). Interaction of the Streptomyces Wbl protein WhiD with the principal sigma factor σHrdB depends on the WhiD [4Fe-4S] cluster. J. Biol. Chem 295, 9752–9765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steyn AJ, Collins DM, Hondalus MK, Jacobs WR Jr., Kawakami RP, and Bloom BR (2002). Mycobacterium tuberculosis WhiB3 interacts with RpoV to affect host survival but is dispensable for in vivo growth. Proc. Natl. Acad. Sci. USA 99, 3147–3152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Volbeda A, Dodd EL, Darnault C, Crack JC, Renoux O, Hutchings MI, Le Brun NE, and Fontecilla-Camps JC (2017). Crystal structures of the NO sensor NsrR reveal how its iron-sulfur cluster modulates DNA binding. Nat. Commun 8, 15052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wan T, Li S, Beltran DG, Schacht A, Zhang L, Becker DF, and Zhang L. (2020). Structural basis of non-canonical transcriptional regulation by the σA4-bound iron-sulfur protein WhiB1 in M. tuberculosis. Nucleic Acids Res. 48, 501–516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams KJ, Joyce G, and Robertson BD (2010). Improved mycobacterial tetracycline inducible vectors. Plasmid 64, 69–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu HM, and Crothers DM (1984). The locus of sequence-directed and protein-induced DNA bending. Nature 308, 509–513. [DOI] [PubMed] [Google Scholar]
- Wu J, Ru HW, Xiang ZH, Jiang J, Wang YC, Zhang L, and Liu J. (2017). WhiB4 Regulates the PE/PPE Gene Family and is Essential for Virulence of Mycobacterium marinum. Sci. Rep 7, 3007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoon C, Privé GG, Goodsell DS, and Dickerson RE (1988). Structure of an alternating-B DNA helix and its relationship to A-tract DNA. Proc. Natl. Acad. Sci. USA 85, 6332–6336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng G, Lu XJ, and Olson WK (2009). Web 3DNA–a web server for the analysis, reconstruction, and visualization of three-dimensional nucleic-acid structures. Nucleic Acids Res. 37, W240–W246. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Atomic coordinates and structure factors have been deposited in the RCSB Protein Data Bank (PDB) under the accession codes 7KUG and 7KUF for the WhiB7:σA4-βtip complex and the WhiB7: σA4-βtip:PwhiB7 complex, respectively.