Abstract
Currently, there are a number of approved antiviral agents for use in the treatment of viral infections. However, many instances exist in which the use of a second antiviral agent would be beneficial because it would allow the option of either an alternative or a combination therapeutic approach. Accordingly, virus-encoded proteases have emerged as new targets for antiviral intervention. Molecular studies have indicated that viral proteases play a critical role in the life cycle of many viruses by effecting the cleavage of high-molecular-weight viral polyprotein precursors to yield functional products or by catalyzing the processing of the structural proteins necessary for assembly and morphogenesis of virus particles. This review summarizes some of the important general features of virus-encoded proteases and highlights new advances and/or specific challenges that are associated with the research and development of viral protease inhibitors. Specifically, the viral proteases encoded by the herpesvirus, retrovirus, hepatitis C virus, and human rhinovirus families are discussed.
Although drugs capable of inhibiting virus replication were described in the scientific literature as early as the 1950s (9, 59), only recently has the development of new antiviral agents with activity against virus-specific functions made rapid progress. To date, 20 different antiviral chemotherapeutic agents have been approved for use in the treatment of individuals infected with a variety of different viruses. Although the majority of these agents are used primarily for the treatment of herpesvirus and human immunodeficiency virus (HIV) infections, respiratory syncytial virus and influenza A virus infections can also be treated (148a). Since the discovery that viruses contain nucleic acid genomes, which undergo replication as part of the virus life cycle, the early antiviral drug design efforts paralleled those in the research and development of antiproliferative agents for the treatment of cancer. Accordingly, the majority of the approved antiviral agents are nucleoside analogs which act by inhibiting viral DNA synthesis (herpesvirus) or viral reverse transcription (HIV).
Despite these advances, the use of most of these antiviral chemotherapeutic agents was characterized by limited clinical efficacy, adverse side effects, and suboptimal pharmacokinetics. Of equal concern was the emergence of drug-resistant viral strains in individuals who required chronic therapy for effective clinical management of their infection, since the development of drug-resistant variants can severely affect and limit subsequent treatment options. Due to these concerns, it was clear that the development of new antiviral agents with activity against new virus-specific targets was warranted. Recent technological advances have facilitated greater understanding of the molecular biology and biochemistry of the viral enzymes which are involved in the viral life cycle. In particular, viral enzymes that are essential for the production of infectious virus represent potential therapeutic targets. Research and development of inhibitors directed to these antiviral targets has been aided by other advances such as high-throughput screening of compound libraries and rationally based drug approaches based on X-ray crystallography.
During the last decade, preclinical research efforts have centered on virus-encoded proteases as potential targets for antiviral intervention (40, 78, 86). These studies have indicated that viral proteases are an absolute requirement in the life cycle of many viruses, either by effecting the cleavage of high-molecular-weight precursor viral proteins to yield functional products or by catalyzing the processing of the structural proteins necessary for assembly and morphogenesis of viral particles. Furthermore, the clinical efficacy of antiviral agents designed to target proteases has been demonstrated in HIV-infected individuals whose therapeutic regimens contain one of four recently approved HIV specific protease inhibitors.
This review will summarize some of the important general features of virus-encoded proteases, specifically highlighting new advances and the specific challenges associated with the design, discovery, and subsequent development of viral protease inhibitors. Although viral proteases play critical roles in the life cycle of many different virus families, this review focuses on the proteases encoded by the herpesvirus, retrovirus, hepatitis C virus (HCV), and human rhinovirus (HRV) families. Detailed information describing the structure and function of viral proteases has been extensively reviewed by other authors (40, 78, 86) and is not covered here.
HUMAN IMMUNODEFICIENCY VIRUS PROTEASE
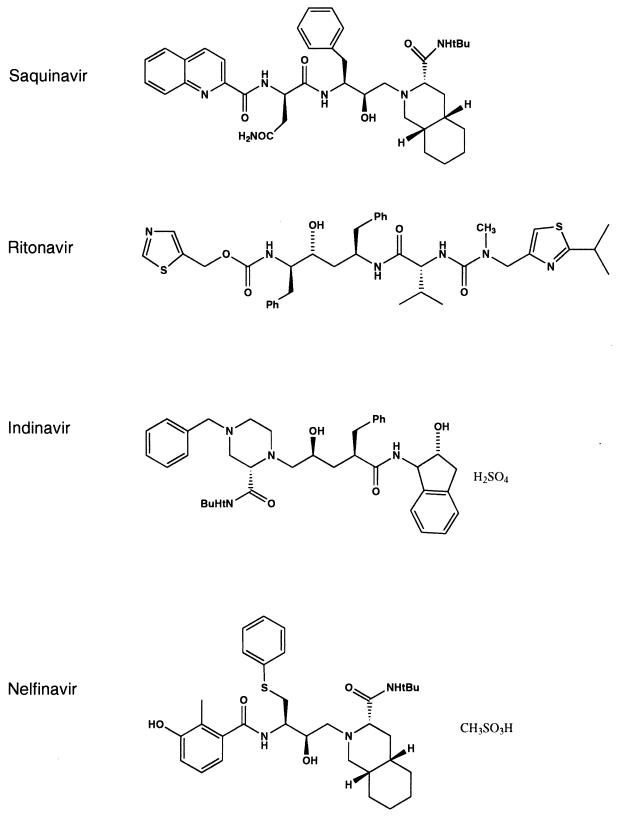
Since the identification of HIV as the causative agent of AIDS, significant efforts were directed toward the research and development of a variety of antiviral chemotherapeutic agents. The first agents that were entered into clinical trials and received approval were nucleoside analogs which target reverse transcriptase (RT), an enzyme that functions early in the HIV life cycle. In general, reverse transcriptase inhibitors (RTIs) appear to have only moderate clinical efficacy when administered in a monotherapy regimen (57, 74, 145). Their use is further limited by adverse side effects. A second class of RTIs is the nonnucleoside RTIs. Although the members of this group of structurally unrelated compounds were shown to have potent activity in vivo, the rapid emergence of highly resistant drug variants limited their clinical efficacy (131, 135). An alternative viral target for intervention is the HIV protease. The use of HIV protease as a potential target was validated in experiments which showed that mutations in the protease resulted in the production of defective, noninfectious virus (77, 84, 124). The demonstration of inhibitors with potent antiviral activity in cell culture-based assays (13, 115, 117, 122, 154) provided further evidence for HIV protease as a promising target for antiviral drug therapy. Data from subsequent clinical trials have demonstrated the safety and efficacy of HIV protease inhibitors and have, to date, led to the approval of four protease inhibitors: saquinavir (Invirase, Ro 31-8959), indinavir (Crixivan, MK-639), ritonavir (Norvir, ABT-538), and, most recently, nelfinavir (VIRACEPT, AG1343) (Fig. 1). Several other compounds, e.g., amprenavir (141W94/VX-478), DMP450, PNU-140690, ABT-378, and PD178390, are currently undergoing preclinical or clinical evaluation (15, 34–36, 68, 88, 112, 127).
FIG. 1.
HIV protease inhibitors. The chemical structures of the clinically approved HIV protease inhibitors saquinavir, ritonavir, indinavir, and nelfinavir are as shown.
Structure and Function
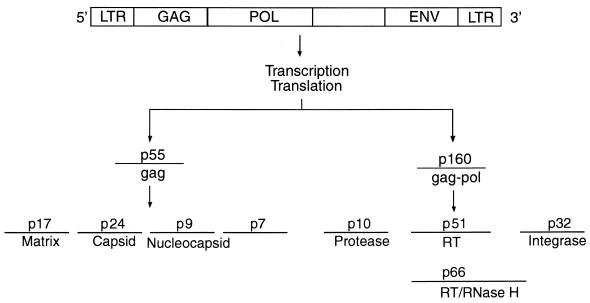
As with all retroviruses, the HIV genome is composed of three major genes, referred to as gag, pol, and env (Fig. 2). Transcription and translation of the gag and pol regions of the virus genome result in the production of two large precursor polyproteins, p55 (gag) and a ribosomal frameshift product p160 (gag-pol) (5, 124, 157). Following a poorly understood autocatalytic event, the protease is released and subsequently cleaves p55 to yield gag structural proteins p17 (matrix), p24 (capsid), p9 (nucleocapsid), and p7 (a protein of unknown function). The pol gene product is likewise cleaved to yield three enzyme products, protease, reverse transcriptase/RNase H, and integrase. In all, HIV protease-mediated processing of the p55 and p160 polyproteins occurs at nine different cleavage sites. The unique nature of some of the scissile bonds, e.g., Phe-Pro, Phe-Leu, and Phe-Thr, can be used in the design of inhibitors that demonstrate a high degree of selectivity toward HIV protease.
FIG. 2.
Genetic organization of the HIV genome. gag (p55) and gag-pol (p160) precursor proteins are cleaved by HIV protease to yield structural proteins (p17, p24, p9, and p7), protease, RT/RNase H, and integrase.
The HIV-1 protease is a member of the aspartic acid protease family and thus is structurally related to host aspartic acid proteases, which include renin, cathepsin D, gastrin, and pepsin (5, 124, 157). Structurally, HIV-1 protease is a homodimer, composed of two noncovalently associated, structurally identical polyprotein chains, each consisting of 99 amino acids. The active site of the enzyme is formed at the dimer interface and contains two catalytic aspartic acid residues, each of which is contributed by the subunits to form two Asp-Thr-Gly motifs (residues 25 to 27). The enzyme active site contains well-defined subsites in which inhibitor or substrate side chains participate in tight binding interactions. Two identical flexible flap regions project over the active site and also participate in the binding of inhibitors and substrates.
Protease Inhibitors In Vitro
Many HIV-1 protease inhibitors described to date have resulted from the screening of compound libraries or from rational drug design efforts based on solved three-dimensional (3-D) X-ray crystal structures of HIV-1 protease (5, 13, 89, 102, 163, 164). The earliest HIV protease inhibitors originated from inhibitors of renin. These were peptidic in nature, where the scissile P1-P1′ amide bond was replaced with nonhydrolyzable transition state isosteres. The utility of these compounds was limited, however, by poor oral bioavailability and metabolic instability. Improvements in the design of inhibitors led to molecules with lower molecular weight that were peptidic. To date, many HIV protease inhibitors have been synthesized; the design and structure of these compounds have been reviewed (5, 102, 163, 164).
As a representative HIV protease inhibitor, nelfinavir was discovered by using a rational drug design approach aided by 3-D X-ray crystal structure determination (76). In cell-based assays, nelfinavir demonstrated potent activity against the replication of several laboratory and clinical HIV-1 or HIV-2 isolates including pyridinone- and zidovudine (ZDV)-resistant strains (122). In drug combination experiments, nelfinavir produced additive to synergistic interactions when combined with other antiretroviral agents (118). As with other protease inhibitors, nelfinavir also inhibited virus replication in HIV-1 chronically infected cells. In contrast, RTIs which target an early preintegration stage in the viral life cycle, are not effective against chronically infected cells (117). The latter observation suggests that added benefit may be obtained when compounds which inhibit different viral targets are used in combination in drug therapeutic regimens.
Compounds that inhibit HIV-1 proteolytic processing result in the formation of defective particles which contain precursor gag (p55) and little or no processed p24. Interestingly, purification of defective HIV-1 particles from cells that have been incubated with the protease inhibitors indinavir or ritonavir and then incubated in drug-free medium has been shown to result in the resumption of proteolytic processing and restoration of viral infectivity (113, 154). However, in similar experiments with nelfinavir or saquinavir, proteolytic processing and virus infectivity were only partially restored 36 h after drug removal (113, 123). This suggests that HIV protease-mediated proteolytic processing may be slow to recover from inhibition by specific protease inhibitors, which is potentially a beneficial feature if drug levels in plasma temporarily fall below the therapeutic range.
In general, protease inhibitors are highly protein bound. A consequence of protein binding is to reduce the amount of inhibitor that is available to interact with the viral target. Protein binding can be studied in in vitro experiments by including serum proteins (albumin or plasma α1-acid glycoprotein) in antiviral activity assays and measuring subsequent reductions in antiviral activity. Although no precise correlation can be made between the degree of protein binding and subsequent clinical efficacy, the observed reduction in the in vitro antiviral activity of the protease inhibitor SC-52151 in the presence of α1-acid glycoprotein has been postulated to explain the lack of clinical efficacy demonstrated in vivo (92).
Protease Inhibitors In Vivo
Clinical trials in which protease inhibitors, e.g., indinavir, saquinavir, ritonavir, and nelfinavir, were evaluated as monotherapy demonstrated the potency of this new class of drugs, as evidenced by significant reductions in HIV-1 RNA levels in plasma and increases in CD4 cell counts during the first week of treatment (18, 29, 35, 100, 104, 110, 111). However, since the ability of these agents to adequately suppress virus replication is often limited, thereby facilitating the selection of drug-resistant HIV variants, treatment regimens have been broadened to include combination therapy. Protease inhibitors in combination with RTIs were shown to result in more significant antiviral responses, which were sustained for a longer duration (19, 27, 28, 35, 58, 104, 110, 111, 134, 155, 156). An example of the potency of a combination therapeutic approach was seen in clinical studies assessing the combination of nelfinavir with lamuvidine (3TC) and ZDV. In patients receiving nelfinavir-3TC-ZDV therapy, a reduction in HIV-1 RNA levels in plasma to <400 and <50 copies/ml was observed in 75 and 61% of patients, respectively, out to 21 months (27, 134).
The results of these clinical trials have led to the development of a list of guidelines that serve to recommend both the appropriate time at which to initiate antiviral therapy and the preferred antiviral drug combinations that are used for highly active antiretroviral therapy (18). Although the current recommended three-drug regimen includes the initial use of a protease inhibitor (e.g., indinavir, nelfinavir, and ritonavir) in combination with RTIs, the long-term benefit and efficacy of additional antiviral drug combinations are still undergoing evaluation. Although the results of clinical trials that evaluated the efficacy of indinavir, saquinavir, nelfinavir, and ritonavir are extensively detailed in other reviews (18, 19, 35, 104, 110, 111, 156), the key findings are briefly described here.
Saquinavir was the first protease inhibitor to receive approval for use in combination with nucleoside analogs. In a clinical efficacy trial, the two-drug regimen of saquinavir with zalcitabine (ddC) resulted in a reduced risk of disease progression in patients compared with patients on saquinavir or ddC monotherapy regimens (19, 33, 104, 110, 111, 156). The original hard-tablet formulation of saquinavir exhibits only modest antiviral activity due primarily to its low oral bioavailability (4%) and extensive first-pass hepatic metabolism by the cytochrome P-450 3A4 (CYP 3A4) system. An enhanced oral formulation of saquinavir was shown to have increased bioavailability (12%) and is currently undergoing clinical evaluation. Saquinavir is generally well tolerated, with the majority of patients reporting mild gastrointestinal disturbances and nausea as the most common adverse effects.
Clinical evaluation of ritonavir demonstrates that high drug levels in plasma are achieved, resulting in significant reductions in HIV-1 RNA levels in plasma and increases in CD4 cell counts (19, 35, 100, 104, 110, 111, 156). In a clinical efficacy trial in patients with advanced HIV disease, ritonavir significantly reduced the risk of disease progression or death when combined with RTI-containing antiretroviral regimens compared to the use of RTI-containing regimens alone. Ritonavir is a potent inhibitor of CYP 3A4. The concurrent use of ritonavir with agents which are metabolized by CYP 3A, e.g., various analgesics, antiarrhythmic agents, antibiotics, anticoagulants, anticonvulsants, antiemetics, and antifungal agents, may result in increased levels of these drugs in plasma. Due to the potential for adverse effects, the use of ritonavir with several of these medications is contraindicated. More recently, it was shown that ritonavir significantly enhanced the level of saquinavir in plasma (80, 107). A ritonavir-saquinavir combination regimen is currently undergoing clinical evaluation (17). The most common side effects associated with ritonavir therapy include nausea, vomiting, circumoral paresthesia, and an elevation of triglyceride and cholesterol levels.
Indinavir therapy was also shown to significantly lower HIV-1 RNA levels in plasma and to increase CD4 cell counts when administered as monotherapy and in combination drug regimens (19, 35, 58, 104, 110, 111, 156). Indinavir is rapidly absorbed after oral administration and must be taken in the fasted state since its bioavailability is reduced by food intake. The major side effects associated with indinavir therapy include increased hyperbilirubinemia and nephrolithiasis.
The fourth protease inhibitor approved for treatment of HIV-1 infection, nelfinavir has demonstrated safety and efficacy as monotherapy as well as in combination with stavudine (d4T) alone or ZDV plus 3TC (19, 27, 29, 35, 51, 104, 110, 111, 134, 142, 156). More recently, twice-daily dosing of nelfinavir with two RTIs has been shown to significantly reduce HIV-1 RNA levels (142). The major side effect associated with nelfinavir therapy is mild to moderate diarrhea. In addition, an increased incidence in hyperglycemia, diabetes, or fat redistribution has been reported for a small percentage of patients receiving therapy that includes either saquinavir, ritanovir, indinavir, or nelfinavir (39, 81, 96, 132). The mechanism behind the association of these side effects and the use of protease inhibitor therapy is at present unknown and warrants further investigation.
Drug resistance.
Overall, clinical trial results have confirmed the utility of HIV protease inhibitors as integral components of potent antiviral therapies. It is now understood that the goal of antiretroviral therapy should be to maximally suppress viral replication. Loss of suppression of viral replication will most probably lead to the emergence of drug-resistant viral variants which have the potential to exhibit reduced susceptibility to all classes of drugs including the currently available protease inhibitors (31, 34, 71, 72, 109, 138, 141, 155). In many cases, these protease-resistant viral variants were shown to exhibit cross-resistance to other protease inhibitors (30, 31, 34). Thus, an understanding of HIV genotypic and phenotypic changes which occur during the course of antiviral therapy is essential when considering treatment with sequential protease inhibitor-containing regimens.
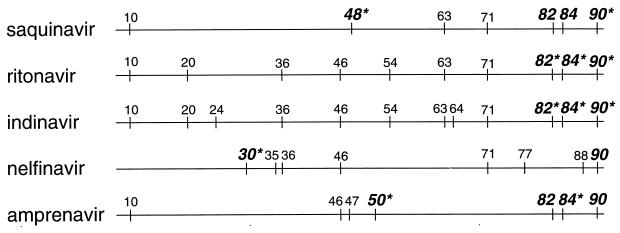
Viral resistance to protease inhibitors has been examined in in vitro serial-passage studies of HIV-1 in the presence of increasing concentrations of a given protease inhibitor (67, 73, 99, 117, 122, 123, 144, 149). The reconstruction of observed changes within the protease gene in HIV proviral clones and subsequent in vitro susceptibility testing were used to identify amino acid substitutions associated with drug resistance. These studies showed that the majority of phenotypic resistance can be attributed to a few specific amino acid substitutions which occur within conserved regions of protease. The vast majority of these substitutions occur at residues that interact directly (e.g., residues 30, 48, 50, 82, and 84) with substrate or inhibitor binding, whereas a few occur at residues that interact indirectly (e.g., residue 90) (Fig. 3). In addition, the concurrent or sequential emergence of amino acid substitutions which do not themselves confer resistance (e.g., residues 10, 20, 35, 36, 46, and 71) frequently arises during in vitro selection. These changes may play a compensatory role by indirectly affecting viral viability.
FIG. 3.
Amino acid substitutions within protease identified in HIV-1 isolates from patients treated with saquinavir, ritonavir, indinavir, nelfinavir, and amprenavir. Amino acid residues that interact with substrate or inhibitors during binding are identified in bold; substitutions at those residues that appear at a high incidence are indicated by asterisks.
The genotypic changes observed in the in vitro serial-passage studies have been somewhat predictive of the amino acid changes which confer resistance to protease inhibitors in vivo (72, 99, 109, 138, 141). Genotypic analysis of HIV isolates obtained from patients receiving indinavir treatment revealed that the protease gene may contain up to 11 different amino acid substitutions and that the majority of isolates contain substitutions at residues 82 and 46 (30, 31). Similarly, HIV isolates obtained from patients receiving ritonavir therapy contain frequent amino acid substitutions at residues 10, 54, 71, and 82 (109, 141). In patients treated with saquinavir, G48V and L90M are the predominant substitutions observed; substitutions at residues 82 and 84 were also detected in vivo but at reduced incidence (33, 71, 72, 90, 138). The genotypic changes observed in HIV isolates obtained from patients receiving nelfinavir therapy have identified a novel aspartic-acid-to-asparagine substitution at residue 30 (D30N) (98, 118, 122). In isolates from these patients, amino acid substitutions at residues 48, 82, and 84 have not been observed, although L90M has appeared in a minority of the isolates.
Since the trend in antiviral therapy is quickly extending to encompass the use of sequential and combination protease inhibitor therapeutic regimens, an understanding of the sensitivity of in vivo protease inhibitor-resistant variants to other protease inhibitors is essential. In vitro susceptibility testing on HIV-1 isolates obtained from patients treated with protease inhibitors demonstrated broad phenotypic cross-resistance among isolates (30, 31, 34, 62, 109, 141). An examination of 19 indinavir-resistant clinical isolates demonstrated that all the isolates were cross-resistant to ritonavir and between 60 and 80% were cross-resistant to both saquinavir and amprenavir (30). Similarly, in two separate studies on ritonavir-resistant clinical isolates, most of the isolates exhibited cross-resistance to indinavir and reduced susceptibility to saquinavir (109, 141). In contrast, only minimal or no cross-resistance was observed for isolates obtained from patients treated with saquinavir- or nelfinavir-containing drug regimens, respectively (34, 63, 120, 121). This latter observation may allow patients who experience clinical failure with some protease inhibitor-containing regimens to derive benefit from therapeutic regimens containing other protease inhibitors. Promising data in support of such salvage regimens were seen in a preliminary study with patients who failed on nelfinavir-containing regimens and whose HIV-1 isolates contained predominantly a substitution at residue 30. These patients demonstrated a significant reduction in HIV-1 RNA levels when switched to a ritonavir- and saquinavir-containing regimen (61, 148). While these data are encouraging, the predictive powers of the viral genotype and/or phenotype on clinical response await further substantiation from larger clinical trials.
An explanation of the ability of specific amino acid substitutions to confer resistance has been recently aided by the analysis and molecular modeling of protease and inhibitor cocrystal structures (5, 24, 44, 76). In the case of nelfinavir resistance, crystallographic analyses reveal that the D30N substitution most probably confers resistance by weakening the hydrogen bond between protease and one of the side groups of nelfinavir (5, 76). The destabilization resulting from the D30N substitution appears to disrupt an interaction that is important only for nelfinavir binding to the protease. This specificity of nelfinavir may explain the lack of cross-resistance of D30N-containing viruses to other clinically approved protease inhibitors. In a similar manner, both indinavir and ritonavir have been optimized to form strong hydrophobic interactions with protease at residue 82 and are adversely affected by substitutions at this residue. Accumulation of two or more amino acid substitutions at residues located at or near the active site (e.g., 82 and 90, 84 and 90, 48 and 90, and 82 and 84 and 90) was also reported in isolates obtained from patients treated with these inhibitors (30, 31, 116, 120). In these cases, binding to many inhibitors is significantly perturbed, resulting in broad cross-resistance among the class of protease inhibitors tested. Isolates containing these multiple changes were demonstrated to be broadly cross-resistant to all protease inhibitors.
Future Considerations
HIV protease inhibitors have emerged as potent antiretroviral chemotherapeutic agents that, in combination with RTIs, have resulted in prolonged suppression of viral replication. Although long-term efficacy data (>2 years) are not yet available, it is generally acknowledged that such factors as the high replication rate of HIV and the long half-lives of latently infected cell populations will continue to pose a challenge for the eventual successful eradication of HIV (47, 66, 125, 160, 165). An additional challenge is the emergence of drug-resistant variants in patients receiving prolonged antiviral therapy, since in many instances, resistance to one protease inhibitor confers cross-resistance to other protease inhibitors. An understanding of the viral resistance profiles which arise in patients receiving protease inhibitor-containing regimens will aid in the design of the most appropriate therapeutic strategies for patients infected with HIV.
PICORNAVIRUS PROTEASE
Picornaviruses comprise one of the largest families of medically important human pathogens. Enteroviruses, a group within the picornaviruses, are associated with a variety of different clinical syndromes including upper respiratory tract illness; aseptic meningitis; encephalitis; hand, foot, and mouth disease; and myocarditis (32, 106, 133). Another medically important group of picornaviruses is the HRV group; which is composed of more than 100 different serotypes. Together, HRV infections represent the single major cause of the common cold and as such are associated with a significant acute morbidity. Although HRV-induced upper respiratory tract illness is often mild and self-limiting, HRV infection may lead to serious medical complications. Rhinovirus colds can ultimately result in sinusitis, otitis media, and lower respiratory tract illnesses including exacerbations of asthma, cystic fibrosis, and bronchitis (in individuals with underlying respiratory disorders) (6). Although no effective antiviral therapies are currently available for either the prevention or the treatment of diseases caused by HRV infection, considerable progress has recently been made in the discovery and development of new antirhinovirus drugs. To date, numerous compounds with potent in vitro activity against HRV have been described (20, 37, 105). The majority of these compounds bind to the virus capsid and inhibit either viral attachment or subsequent uncoating.
Although many of these compounds have been evaluated in clinical trials, in only a few instances have reductions in disease severity, illness frequency, and/or viremia been achieved (6, 37). Efforts to develop antirhinovirus drugs have been unsuccessful due to lack of efficacy, problems in achieving adequate drug levels in nasal epithelium, and/or problems with toxicity. However, recent clinical trials evaluating the intranasal administration of a soluble form of the HRV receptor intercellular cell adhesion molecule (ICAM-1) have demonstrated reductions in disease severity and in levels of infectious virus (152). However the costs associated with the development of a recombinant protein may preclude the development of this chemotherapeutic agent. Another potential antirhinovirus drug is Pleconaril (VP-63843), an inhibitor of the viral uncoating process, which has demonstrated efficacy in reducing the symptoms associated with upper respiratory tract disease caused by coxsackievirus A21 (49, 140). Recently, clinical trials designed to evaluate the efficacy of Pleconaril against human HRV have been initiated. In addition to agents that inhibit an early stage in the viral life cycle, recent attention has focused on the virus-encoded 2A and 3C proteases, which are required for HRV replication (79, 91). Antiviral agents designed to inhibit 3C protease have shown considerable potential and therefore are the focus of this review.
Structure and Function
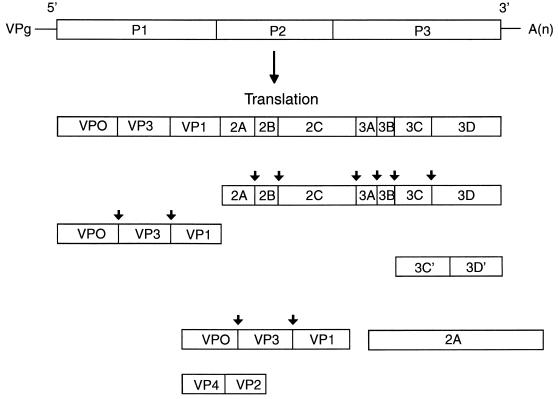
The picornaviruses genome is a positive-sense single strand of RNA of about 7,500 nucleotides, which encodes a single large polyprotein precursor (32, 40, 78, 86, 106, 133). The polyprotein precursor first undergoes an autocatalytic cleavage by the 2A protease to release P1, the precursor to the viral capsid proteins (Fig. 4). The 2A protease also mediates an alternative cleavage to generate the 3C′ and 3D′ products. All other cleavages of the polyprotein, with the exception of the late autocatalytic cleavage of VP0 during final viral assembly, are catalyzed by the virus-encoded 3C protease. These reactions occur most often between Gln-Gly pairs and are mediated by either 3C protease itself or 3C fused to the viral RNA-dependent RNA polymerase 3D (3CD). Early studies comparing sequence homologies and predicted secondary structures suggested that 3C was a member of the trypsin family of serine proteases (40, 53, 86). However, studies with cysteine protease inhibitors, together with results of oligonucleotide-mediated site-directed mutagenesis experiments, demonstrated an absolute requirement for the highly conserved cysteine residue for viral replication (40, 78, 79, 91). Resolution of the 3-D structure by X-ray crystallography subsequently revealed that 3C protease has the polypeptide backbone of a serine protease but that the active-site serine nucleophile is instead a cysteine sulfhydryl (3, 103).
FIG. 4.
Genetic organization of the picornavirus genome. The viral RNA contains a small basic protein, VPg, at the 5′ end and a polyadenylic acid [A(n)] sequence at the 3′ end. The viral RNA is divided into three coding regions, designated P1, P2, and P3. Translation of viral RNA yields a large polyprotein, which is subsequently cleaved by 2A protease to release P1. With the exception of the autocatalytic cleavage of VP0 and the 2A-mediated alternative cleavages of 3C′ and 3D′, all the cleavages are mediated by 3C protease. 3C cleavage sites are denoted by arrows.
Protease Inhibitors In Vitro
Several different anti-picornavirus compounds which target 3C protease have been designed based on the known sequence of the preferred peptide substrate, Leu-Phe-Gln (41, 42, 60, 75, 159). These compounds were further modified, using the solved 3-D structure of 3C protease (103), to produce molecules that have potent enzymatic and antiviral activity. In general, these compounds are composed of various glutamine isosteres in the P1 subsite and contain a chemical moiety capable of reacting with the active-site cysteine residue. Recently, a novel series of 2,3-dioxindoles (isatins), which contain a reactive α-keto-amide group, were found to exhibit potent activity against the enzyme, but no antiviral activity was detected in cell-based assays (158). A second class of compounds, which demonstrated potent enzyme inhibitory activity but only modest antiviral activity in cell-based assays, included the reversible tetra- or tripeptidyl aldehydes (60, 75, 159). More recently, a third class of irreversible tripeptidic inhibitors, which contain various Michael acceptor moieties, was described (41, 42). Representative compounds from this class not only have potent activity against the enzyme but also have demonstrated potent antiviral activity.
Although the metabolic instability associated with peptidic aldehydes may preclude their development as antiviral drugs, the results of studies with such a compound, AG6084 (Fig. 5), have identified some possibly unique attributes of 3C protease inhibitors (48, 119, 159, 168). A significant degree of homology within the 3C coding region, including strict conservation of the active-site residues, has been demonstrated across the 100 or more different HRV serotypes and even among several related picornaviruses (53). Accordingly, AG6084 effectively inhibited the replication of both diverse HRV serotypes and the related picornaviruses coxsackieviruses A21 and B3, enterovirus 70, and echovirus 11 (48). Furthermore, in a single-cycle, time-of-addition assay, AG6084 could be added late in the HRV life cycle and still reduce the levels of infectious virus. In contrast, compounds targeting viral attachment or uncoating were ineffective when added only 1 h after infection. Although the pathogenesis of the common cold is not completely understood, it is apparent that the cascade of host inflammatory mediators released as a result of the viral infection play an important role. In this regard, it is encouraging that AG6084 decreases interleukin-6 and interleukin-8 production in viral infected BEAS-2B cells, a transformed human bronchial cell line (168). Direct evidence of the inhibition of 3C proteolytic activity in infected cells treated with AG6084 was provided by sodium dodecyl sulfate-polyacrylamide gel electrophoresis analysis of radiolabeled proteins, which showed a dose-dependent accumulation of viral precursor polyproteins and a reduction in the amounts of processed protein products.
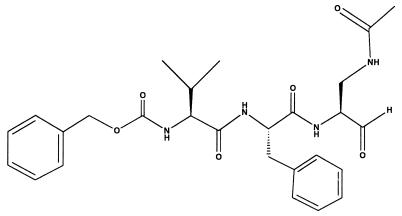
FIG. 5.
HRV protease inhibitor. The chemical structure of the rhinovirus protease inhibitor AG6084 is as shown.
Future Considerations
Although the discovery and development of antiviral agents capable of providing clinical benefit for the treatment of picornavirus infections have been the focus of considerable research efforts, no antiviral chemotherapeutic agents for this indication are yet available. Ideally, antirhinoviral agents should be effective against a majority of virus serotypes when administered either prophylactically or therapeutically. The finding of clinical efficacy following the administration of sICAM-1 and the initiation of rhinovirus clinical trials with Pleconaril, a capsid-binding molecule, should encourage new enthusiasm. Recent descriptions of 3C protease inhibitors as having a broad spectrum of antiviral activity against diverse rhinovirus serotypes and antiviral efficacy when added throughout the viral life cycle highlight the advantages of this target and make these compounds promising clinical candidates.
HERPESVIRUS PROTEASE
Human cytomegalovirus (CMV), herpes simplex virus type 1 (HSV-1), and herpes simplex virus type 2 (HSV-2) are important human pathogens which are responsible for a wide range of clinical manifestations in infected individuals. Over the last decade, the incidence and severity of HSV and CMV infections caused by the reactivation of latent virus have increased due to the rise in the number of immunocompromised patients following chemotherapy, transplantation, or infection with HIV. Currently, all the antiviral agents that are approved for the treatment of HSV and CMV infection act by interfering with viral DNA synthesis (21). For the treatment of HSV infection, the most commonly used chemotherapeutic agents are acyclovir, valaciclovir, and famciclovir, whereas for the treatment of CMV infection, ganciclovir, foscarnet, and cidofovir are used. With the exception of acyclovir, these chemotherapeutic agents have only moderate clinical efficacy and are associated with serious side effects, which often necessitate the coadministration of additional drugs to counter toxicity. In addition, as with all antiviral agents, the emergence of drug-resistant virus strains may be a critical problem during the course of prolonged therapy, since drug resistance can often confer cross-resistance to other approved antiviral agents, thus limiting treatment options. Clearly, an antiviral agent which inhibits HSV and/or CMV replication by a different mechanism of action would be an important addition to the available therapeutic regimens. A potential virus-specific target currently under investigation is the viral protease involved in capsid assembly. Extensive research efforts have led to a better understanding of the biochemistry of capsid proteases and have recently culminated in the resolution of the 3-D X-ray crystal structure of the CMV capsid protease, assemblin.
Structure and Function
CMV strains, like other members of the herpesvirus family, are large (>200-kb) DNA-containing viruses which replicate in the nucleus of an infected host cell (108). An integral step of the virus replicative process is the intranuclear packaging of newly synthesized DNA into capsids, which go on to acquire viral tegument and envelope proteins before leaving the infected cell. The CMV protease, assemblin (162), plays a central role in the complex processes of capsid assembly and maturation through its proteolytic processing of the CMV assembly protein precursor (139). Assemblin-mediated proteolytic processing is proposed to be essential for CMV viability, since it has been shown that an HSV assemblin mutant yields noninfectious virus (50).
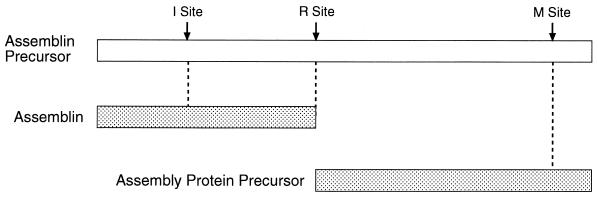
Assemblin is synthesized as a precursor polyprotein together with the assembly protein precursor (Fig. 6) (52). Assemblin-mediated cleavage at the release site frees assemblin from the assembly protein precursor, while cleavage at the maturation results in the mature form of the assembly protein. Assemblin-mediated cleavage at a third site, the internal site, releases the two chains of assemblin. This latter cleavage, however, is not essential for assemblin activity (70, 114). The assemblin precursor and the assembly protein precursors interact with the major capsid protein and facilitate its transport into the nucleus, where multimerization leads to the generation of intermediate capsids. Within the intermediate capsids, assemblin-mediated cleavage at the maturation site induces conformational changes that result in a capsid form which is competent for the packaging of virion DNA.
FIG. 6.
Diagram of the CMV assemblin protein precursor. Assemblin-mediated cleavage at the release (R) site of the assemblin protein precursor releases assemblin from the assembly protein precursor. Assemblin-mediated cleavage at the internal (I) site results in the two-chain form of assemblin. The mature form of the assembly protein is generated by assemblin-mediated cleavage at the maturation (M) site, located within the assembly protein precursor. Assemblin cleavage sites are denoted by arrows.
Recently, the resolution of the 3-D structure of assemblin by X-ray crystallography was reported by several laboratories (23, 128, 143, 151). Analysis of the assemblin crystal structure revealed distinct differences from that of cellular serine proteases. Most striking are the unique backbone fold of assemblin and its active-site residues of Ser-His-His, which represent a novel variation of the classic Ser-His-Asp triad of trypsin-like proteases. The functional form of the enzyme is proposed to be a homodimer. The active sites, where one site is contributed by each monomer, are located along the surface of the enzyme and are well separated from each other. While the assemblin active site is somewhat shallow, an understanding of both its topology and the specific recognition requirements for substrate cleavage will facilitate the rational design of substrate-based inhibitors.
Protease Inhibitors In Vitro
The kinetic activity and substrate specificity of assemblin have been extensively studied in vitro (10, 16, 69, 97, 126). The well-defined substrate specificity of assemblin has made possible the design of peptidomimetic inhibitors as evidenced by the in vitro inhibitory activity of a peptidyl inhibitor based on the maturation site cleavage junction (69). The knowledge gained from substrate specificity studies, together with an understanding of the active-site topology of assemblin from crystal structure analyses, should further facilitate the rational design of inhibitors. Discoveries such as the recent cocrystalization of assemblin with a tetrapeptide aldehyde inhibitor (23) will also aid rational-design efforts by providing a better understanding of enzyme-inhibitor binding.
A strategy that has been used in both rational-design and screening efforts is the design or selection of inhibitors that form a covalent enzyme-inhibitor complex via the catalytic serine residue of assemblin. Recent reports that describe the in vitro inhibitory activity and the antiviral activity of a subclass of benzoxazinone compounds (2) and several inhibitors in the class of monocyclic β-lactams (87) validate this approach. However, definitive proof that the antiviral effect observed with these compounds in a cell-based assay results directly from the inhibition of assemblin activity remains to be shown. Yet another approach to assemblin inhibition is to target the conserved cysteine residues. As demonstrated with CL13933 (12) and flavin (11), these compounds inhibit in vitro assemblin activity through the formation of specific intramolecular disulfide bonds between cysteine residues located near the catalytic serine residue. The disruption of assemblin dimerization is another potential approach in which the use of rational drug design methods could target the deep binding pocket of the dimer interface (23, 143).
Future Considerations
While a variety of compounds which target DNA synthesis are currently available for the treatment of HSV and CMV infections, dose-limiting toxicities and drug resistance often complicate their therapeutic usage. Therefore, the recent advances in the research into inhibitors which target the virus capsid protease represent a potential therapeutic alternative.
HEPATITIS C VIRUS PROTEASE
HCV is one of the major causes of parenterally transmitted non-A, non-B hepatitis worldwide. HCV infection is characterized by the high rate (>70%) at which acute infection progresses to chronic infection (4). Chronic HCV infection may lead to progressive liver injury, cirrhosis, and, in some cases, hepatocellular carcinoma. Currently, there are no specific antiviral agents available for the treatment of HCV infection. Although alpha interferon therapy is often used in the treatment of HCV-induced moderate or severe liver disease, only a minority of patients exhibit a sustained response (137). Additionally, a vaccine to prevent HCV infection is not yet available, and it remains uncertain whether vaccine development will be complicated by the existence of multiple HCV genotypes as well as viral variation within infected individuals (101, 161). The presence of viral heterogeneity may increase the likelihood that drug-resistant virus will emerge in infected individuals unless antiviral therapy effectively suppresses virus replication. Most recently, several of the HCV-encoded enzymes, specifically the NS3 protease and NS5B RNA polymerase, have been the focus of intensive research, in vitro screening, and/or rational drug design efforts.
Structure and Function
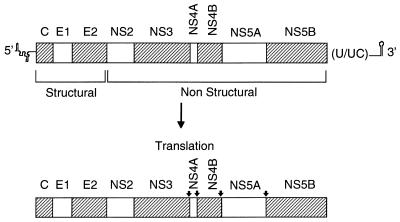
HCV has been classified in the flavivirus family in a genus separate from that of the flaviviruses and the pestiviruses (130). Although the study of HCV replication is limited by the lack of an efficient cell-based replication system, an understanding of replicative events has been inferred from analogies made to the flaviviruses, pestiviruses, and other positive-strand RNA viruses. Figure 7 shows a schematic representation of the HCV genome. As in the pestiviruses, translation of the large open reading frame occurs by a cap-independent mechanism and results in the production of a polyprotein of 3,010 to 3,030 amino acids. Proteolytic processing of the structural proteins (the nucleocapsid protein or core [C]) (136) and two envelope glycoproteins, E1 and E2 (129), is accomplished by the action of host cell signal peptidases. Cleavage of the nonstructural proteins (NS4A, NS4B, NS5A, and NS5B) is mediated by the action of the NS2/3 protease (55, 65) or the NS3 protease (8, 43, 55, 150). NS4A is a cofactor for NS3 (8, 45, 93), and NS5B is an RNA-dependent RNA polymerase (14). Functions for the NS4B and NS5A proteins have yet to be defined.
FIG. 7.
Genetic organization of the HCV genome. The 5′ and 3′ noncoding regions that flank the genomic coding region are shown. Translation of the coding region results in a polyprotein that contains both the structural gene products (core, E1, and E2) and the nonstructural gene products (NS2 to NS5B). NS3 protease cleavage sites are denoted by arrows.
The NS2/3 protease has been shown to mediate cleavage at the 2/3 junction site (Fig. 7) (54, 64). In contrast, the NS3 protease is required for multiple cleavages within the nonstructural segment of the polyprotein, specifically the 3/4A, 4A/4B, 4B/5A, and 5A/5B junction sites (7, 43, 55, 150). Although NS3 protease is presumed to be essential for HCV viability, definitive proof of its necessity has been hampered by the lack of an infectious molecular clone that can be used in cell-based experiments. However, two independent HCV infectious molecular clones have recently been developed and have been shown to replicate in chimpanzees (85, 167). The requirement for NS3 in the HCV life cycle may be validated in these clones by using oligonucleotide-mediated site-directed mutagenesis to inactivate the NS3 catalytic serine residue and then determining whether infectious virus is produced in chimpanzees. Until these experiments are performed, the necessity for NS3 is inferred from cell-based experiments using the related yellow fever virus (YFV) and bovine viral diarrhea virus (BVDV). Mutagenesis of the YFV (22) and BVDV (166) NS3 protease homologs has shown that NS3 protease activity is essential for YFV and BVDV replication.
Protease Inhibitors In Vitro
The determination of the 3-D X-ray crystal structure of the NS3 protease (82, 95) was recently reported. Analysis of the NS3 protease crystal structure revealed a monomeric enzyme with two domains and a structural zinc site. The observed folding of the two domains and the relative order and spacing of the catalytic triad (His, Asp, Ser) place NS3 in the trypsin family of serine proteases. Compared to cellular serine proteases, the substrate binding groove of the NS3 protease is shallow. This extended shallow substrate binding groove represents a challenge to the rational design of inhibitors, since the number of potential inhibitor-enzyme contacts is diminished. However, in vitro studies probing the substrate specificity of NS3 demonstrate stereochemical requirements that differ from those of cellular serine proteases (83, 146, 169). These differences, together with the crystallographic information, should facilitate the design of inhibitors which have a high degree of selectivity to NS3.
Many of the in vitro assays used to characterize NS3 protease activity have been adapted to screen compound libraries or to evaluate rationally designed inhibitors. To date, several thiazolidine derivatives (147) and a phenanthrenequinone compound (26) have demonstrated in vitro NS3 inhibitory activity. Additionally, two in vitro affinity selection techniques have been used to isolate RNA aptamers (153) and a derivative of human pancreatic trypsin inhibitor (38), both of which have been shown to inhibit NS3 protease activity.
Cell-based assays.
Since the evaluation of inhibitors obtained from rational drug design efforts or in vitro screens against HCV itself in a standard cell-based antiviral assay is not a practical option, several alternative approaches are being pursued. One approach involves the evaluation of inhibitors in reporter gene systems. In these cell-based systems, the transcription (65) or the detection (94) of a given reporter gene product has been engineered to be dependent upon NS3 protease activity. A second approach, which is perhaps more reflective of the context and localization of NS3 in HCV-infected cells, is the construction of virus chimeras. These virus chimeras have been designed to require one or more NS3-mediated cleavage events during the course of the viral life cycle before infectious virus can be produced. Recently, two NS3-containing Sindbis virus chimeras (25, 46) and an NS3-containing poliovirus chimera (56) have been described. An additional approach to the evaluation of potential NS3 inhibitors is the use of the HCV-related pestivirus BVDV as a surrogate for virus infection. However, whether inhibitors which demonstrate activity against the related BVDV NS3 protease will also be effective against the HCV NS3 protease is unknown.
Future Considerations
The treatment of HCV infection is an unmet clinical need, since there are currently no approved antiviral agents available for therapeutic use. The NS3 protease is the most extensively studied target to date for the development of an HCV antiviral agent. Other potential antiviral targets include the HCV NS3 helicase domain and NS5B RNA polymerase; accordingly, research in these areas is ongoing. The development of an HCV antiviral agent presents some unique challenges that must be addressed. One of these challenges is the lack of an efficient cell-based assay system, but several alternative approaches are being used to overcome this obstacle. In addition, as a greater understanding of the clinical pathology and progression of HCV infection becomes available, the design and appropriate use of the therapeutic antiviral agents currently under development will follow.
ACKNOWLEDGMENTS
We thank Rose Najarian for help in preparation of the manuscript, Muizz Hasham for help in preparation of the graphics, and Steve Worland for critical reading of the manuscript.
REFERENCES
- 1. Reference deleted.
- 2.Abood N A, Schretzman L A, Flynn D L, Houseman K A, Wittwer A J, Dilworth V M, Hippenmeyer P J, Holwerda B C. Inhibition of human cytomegalovirus protease by benzoxazinones and evidence of antiviral activity in cell culture. Bioorg Med Chem Lett. 1997;7:2105–2108. [Google Scholar]
- 3.Allaire M, Chernaia M M, Malcom B A, James M N. Picornaviral 3C cycteine proteinases have a fold similar to chymotrypsin-like serine proteinases. Nature. 1994;369:72–76. doi: 10.1038/369072a0. [DOI] [PubMed] [Google Scholar]
- 4.Alter M J. Epidemiology of hepatitis C in the west. Semin Liver Dis. 1995;15:5–14. doi: 10.1055/s-2007-1007259. [DOI] [PubMed] [Google Scholar]
- 5.Appelt K. Inhibitors of HIV-1 protease. In: Veerapandian P, editor. Structure-based drug design. New York, N.Y: Marcel Dekker, Inc.; 1997. pp. 1–39. [Google Scholar]
- 6.Arruda E, Hayden F G. Clinical studies of antiviral agents for picornaviral infections. In: Jeffries D J, De Clerq E, editors. Antiviral chemotherapy. Chichester, United Kingdom: John Wiley & Sons, Ltd.; 1995. pp. 321–355. [Google Scholar]
- 7.Bartenschlager R, Ahlborn-Laake L, Mous J, Jacobsen H. Nonstructural protein 3 of the hepatitis C virus encodes a serine-type proteinase required for cleavage at the NS3/4 and NS4/5 junctions. J Virol. 1993;67:3835–3844. doi: 10.1128/jvi.67.7.3835-3844.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bartenschlager R, Ahlborn-Laake L, Mous J, Jacobsen H. Kinetic and structural analyses of hepatitis C virus polyprotein processing. J Virol. 1994;68:5045–5055. doi: 10.1128/jvi.68.8.5045-5055.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bauer D J. The antiviral and synergic actions of isatin ithiosemicarbazone and certain phenoxypyrimidines in vaccinia infection in mice. Br J Exp Pathol. 1955;36:105–114. [PMC free article] [PubMed] [Google Scholar]
- 10.Baum E Z, Bebernitz G A, Hulmes J D, Muzithras V P, Jones T R, Gluzman Y. Expression and analysis of the human cytomegalovirus UL80-encoded protease: identification of autoproteolytic sites. J Virol. 1993;67:497–506. doi: 10.1128/jvi.67.1.497-506.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Baum E Z, Ding W D, Siegel M M, Hulmes J D, Bebernitz G A, Sridharan L, Tabei K, Krishnamurthy G, Carofiglio T, Groves J T, Bloom J D, DiGrandi M, Bradley M, Ellestad G, Seddon A P, Gluzman Y. Flavins inhibit human cytomegalovirus UL80 protease via disulfide bond formation. Biochemistry. 1996;35:5847–5855. doi: 10.1021/bi9529972. [DOI] [PubMed] [Google Scholar]
- 12.Baum E Z, Siegel M M, Bebernitz G A, Hulmes J D, Sridharan L, Sun L, Tabei K, Johnston S H, Wildey M J, Nygaard J, Jones T R, Gluzman Y. Inhibition of human cytomegalovirus UL80 protease by specific intramolecular disulfide bond formation. Biochemistry. 1996;35:5838–5846. doi: 10.1021/bi952996+. [DOI] [PubMed] [Google Scholar]
- 13.Bechtold C M, Patick A K, Alam M, Greytok J, Tino J A, Chen P, Gordon E, Ahmad S, Barrish J C, Zahler R, Lin P F, Colonno R. Antiviral properties of aminodiol inhibitors against human immunodeficiency virus and protease. Antimicrob Agents Chemother. 1995;39:374–379. doi: 10.1128/aac.39.2.374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Behrens S-E, Tomei L, DeFrancesco R. Identification and properties of the RNA-dependent RNA polymerase of hepatitis C virus. EMBO J. 1996;15:12–22. [PMC free article] [PubMed] [Google Scholar]
- 15.Borin M T, Wang Y, Schneck D W, Li H, Brewer J E, Daenzer C L. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Multiple-dose, safety, tolerance and pharmacokinetics of the protease inhibitor PNU-140690 in healthy volunteers, abstr. 648; p. 201. [Google Scholar]
- 16.Burck P J, Berg D H, Luk T P, Sassmannshausen L M, Wakulchik M, Smith D P, Hsiung H M, Becker G W, Gibson W, Villareal E C. Human cytomegalovirus maturational proteinase: expression in Escherichia coli, purification, and enzymatic characterization by using peptide substrate mimics of natural cleavage sites. J Virol. 1994;68:2937–2946. doi: 10.1128/jvi.68.5.2937-2946.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cameron D W, Japour A, Mellors J W, Farthing C, Cohen C, Markowitz M, Poretz D, Follansbee S, Ho D, McMahon D, Berg J, Nieves J, Xu Y, Rode R, Salgo M, Leonard J, Sun E. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Antiretroviral safety and durability of ritonavir (RIT)-saquinavir (SQV) in protease inhibitor-naive patients in year two of follow-up, abstr. 388; p. 152. [Google Scholar]
- 18.Carpenter C, Fischl M A, Hammer S M, Hirsch M S, Jacobsen D M, Katzenstein D A, Montaner J S G, Richman D D, Saag M S, Schooley R T, Thompson M A, Vella S, Yeni P G, Volberding P A. Antiretroviral therapy for HIV infection in 1997. JAMA. 1997;277:1962–1969. [PubMed] [Google Scholar]
- 19.Carr A, Cooper D A. HIV protease inhibitors. AIDS. 1996;10:S151–S157. doi: 10.1097/00002030-199601001-00021. [DOI] [PubMed] [Google Scholar]
- 20.Carrasco L. Picornavirus inhibitors. Pharmacol Ther. 1994;64:119–128. doi: 10.1016/0163-7258(94)90040-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cassady K A, Whitley R J. New therapeutic approaches to the alphaherpesvirus infections. J Antimicrob Chemother. 1997;39:119–128. doi: 10.1093/jac/39.2.119. [DOI] [PubMed] [Google Scholar]
- 22.Chambers T J, Weir R C, Grakoui A, Court D W, Bazan J F, Fletterick R J, Rice C M. Evidence that the N-terminal domain of nonstructural protein NS3 from yellow fever virus is a serine protease responsible for site-specific cleavages in the viral polyprotein. Proc Natl Acad Sci USA. 1990;87:8898–8902. doi: 10.1073/pnas.87.22.8898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chen P, Tsuge H, Almassay R J, Gribskov C L, Katoh S, Vanderpool D L, Margosiak S A, Pinko C, Matthews D A, Kan C-C. Structure of the human cytomegalovirus protease catalytic domain reveals a novel serine protease fold and catalytic triad. Cell. 1996;86:835–843. doi: 10.1016/s0092-8674(00)80157-9. [DOI] [PubMed] [Google Scholar]
- 24.Chen Z, Schock H B, Hall D, Chen E, Kuo L C. Three-dimensional structure of a mutant HIV-1 protease displaying cross-resistance to all protease inhibitors in clinical trials. J Biol Chem. 1995;270:21433–21436. doi: 10.1074/jbc.270.37.21433. [DOI] [PubMed] [Google Scholar]
- 25.Cho Y-G, Moon H-S, Sung Y-C. Construction of hepatitis C-SIN virus recombinants with replicative dependency on hepatitis C virus serine protease activity. J Virol Methods. 1997;65:201–207. doi: 10.1016/s0166-0934(97)02183-6. [DOI] [PubMed] [Google Scholar]
- 26.Chu M, Mierzwa R, Truumees I, King A, Patel M, Berrie R, Hart A, Butkiewicz N, Mahapatra B D, Chan T-M, Puar M S. Structure of Sch 68631: a new hepatitis C proteinase inhibitor from Stretomyces sp. Tetrahedron Lett. 1996;37:7229–7232. [Google Scholar]
- 27.Clendeninn N, Quart B, Anderson R, Knowles M, Chang Y. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Analysis of long-term virologic data from the VIRACEPT® 511 protocol using 3 HIV-RNA assays, abstr. 372; p. 148. [Google Scholar]
- 28.Collier A C, Coombs R W, Schoenfeld D A, Bassett R L, Timpone J, Baruch A, Jones M, Facey K, Whitacre C, McAuliffe V J, Friedman H M, Merigan T C, Reichman R C, Hooper C, Corey L. Treatment of human immunodeficiency virus infection with saquinavir, zidovudine and zalcitabine. N Engl J Med. 1996;334:1011–1017. doi: 10.1056/NEJM199604183341602. [DOI] [PubMed] [Google Scholar]
- 29.Conant M, Markowitz M, Hurley A, Ho D, Peterkin J, Chapman S. Abstracts of the XI International Conference on AIDS. 1996. A randomized phase II dose range-finding study of the HIV protease inhibitor VIRACEPT® as monotherapy in HIV positive patients, abstr. 2129; p. 286. [Google Scholar]
- 30.Condra J H, Holder D J, Schleif W A, Blahy O M, Danovich R M, Gabryelski L J, Graham D J, Laird D, Quintero J C, Rhodes A, Robbins H L, Roth E, Shivaprakash M, Yang T, Chodakewitz J A, Deutsch P J, Leavitt R Y, Massari F E, Mellors J W, Squires K E, Steigbigel R T, Teppler H, Emini E A. Genetic correlates of in vivo viral resistance to indinavir, a human immunodeficiency virus type 1 protease inhibitor. J Virol. 1996;70:8270–8276. doi: 10.1128/jvi.70.12.8270-8276.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Condra J H, Schleif W A, Blahy O M, Gabryelski L J, Graham D J, Quintero J C, Rhodes A, Robbins H L, Roth E, Shivaprakash M. In vivo emergence of HIV-1 variants resistant to multiple protease inhibitors. Nature. 1995;374:569–571. doi: 10.1038/374569a0. [DOI] [PubMed] [Google Scholar]
- 32.Couch R B. Rhinoviruses. In: Fields B N, Knipe D M, editors. Virology. New York, N.Y: Raven Press; 1990. pp. 607–629. [Google Scholar]
- 33.Craig C, O’Sullivan E, Cammack N. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Increased exposure to the HIV protease inhibitor saquinavir (SQV) does not alter the nature of key resistance mutations, abstr. 398; p. 154. [Google Scholar]
- 34.Craig J C, Duncan I B, Gilbert S, Jacobsen H, Jupp R, Moffatt A, Race E, Roberts N A, Mills J S, Mous J, Sheldon J, Tomlinson P W, Whittaker L N. Abstracts of the 5th International Workshop: HIV Drug Resistance. 1996. Treatment with saquinavir (Invirase™) should leave the majority of patients the option to use other HIV proteinase inhibitors, abstr. 32; p. 32. [Google Scholar]
- 35.Deeks S G, Smith M, Holodniv M, Kahn J O. HIV-1 protease inhibitors. JAMA. 1997;277:145–153. [PubMed] [Google Scholar]
- 36.De Pasquale M P, Murphy R, Gulick R, Smeaton L, Sommadossi J-P, Degruttola V, Caliendo A, Kuritzkes D, Sutton L, Savara A, D’Aquila R. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Mutations selected in HIV plasma RNA during 141W94 therapy, abstr. 546; p. 182. [Google Scholar]
- 37.Diana G D, Pevear D C. Antipicornavirus drugs: current status. London, United Kingdom: International Medical Press Ltd; 1997. pp. 401–408. [Google Scholar]
- 38.Dimasi N, Martin F, Volpari C, Brunetti M, Biasiol G, Altamura S, Cortese R, de Francesco R, Steinkühler C, Sollazzo M. Characterization of engineered hepatitis C virus NS3 protease inhibitors affinity selected from human pancreatic secretory trypsin inhibitor and minibody repertoires. J Virol. 1997;71:7461–7469. doi: 10.1128/jvi.71.10.7461-7469.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Dong B J, Gruta C, Legg J, Balano K, Goldschmit R H. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Diabetes and use of protease inhibitors, abstr. 416; p. 157. [Google Scholar]
- 40.Dougherty W G, Semler B L. Expression of virus-encoded proteinases: functional and structural similarities with cellular enzymes. Microbiol Rev. 1993;57:781–822. doi: 10.1128/mr.57.4.781-822.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Dragovich P S, Webber S E, Babine R E, Fuhrman S A, Patick A K, Matthews D A, Lee C A, Reich S H, Prins T J, Marakovits J T, Littlefield E S, Zhou R, Tikhe J, Ford C E, Wallace M, Meador J W, Ferre R A, Brown E L, Binford S L, Harr J E V, DeLisle D M, Worland S T. Structure-based design, synthesis, and biological evaluation of irreversible human rhinovirus 3C protease inhibitors. 1. Michael acceptor structure-activity studies. J Med Chem. 1998;48:2806–2818. doi: 10.1021/jm980068d. [DOI] [PubMed] [Google Scholar]
- 42.Dragovich P S, Webber S E, Babine R E, Fuhrman S A, Patick A K, Matthews D A, Reich S H, Marakovits J T, Prins T J, Zhou R, Tikhe J, Littlefield E S, Bleckman T M, Wallace M, Little T, Ford C E, Meador J W, Ferre R A, Brown E L, Binford S L, DeLisle D M, Worland S T. Structure-based design, synthesis, and biological evaluation of irreversible human rhinovirus 3C protease inhibitors. 2. Peptide structure-activity studies. J Med Chem. 1998;41:2819–2834. doi: 10.1021/jm9800696. [DOI] [PubMed] [Google Scholar]
- 43.Eckart M R, Selby M, Masiarz F, Lee C, Berger K, Crawford C, Kuo C, Kuo G, Houghton M, Choo Q-L. The hepatitis C virus encodes a serine proteinase involved in processing of the putative nonstructural proteins from the viral polyprotein precursor. Biochem Biophys Res Commun. 1993;192:399–406. doi: 10.1006/bbrc.1993.1429. [DOI] [PubMed] [Google Scholar]
- 44.Erickson J W, Burt S K. Structural mechanisms of HIV drug resistance. Annu Rev Pharmacol Toxicol. 1996;36:545–571. doi: 10.1146/annurev.pa.36.040196.002553. [DOI] [PubMed] [Google Scholar]
- 45.Failla C, Tomei L, DeFrancesco R. Both NS3 and NS4A are required for proteolytic processing of hepatitis C virus nonstructural proteins. J Virol. 1994;68:3753–3760. doi: 10.1128/jvi.68.6.3753-3760.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Filocamo G, Pacini L, Migliaccio G. Chimeric Sindbis viruses dependent on the NS3 protease of hepatitis C virus. J Virol. 1997;71:1417–1427. doi: 10.1128/jvi.71.2.1417-1427.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Finzi D, Hermankova M, Pierson T, Carruth L M, Buck C, Chaisson R E, Quinn T C, Chadwick K, Margolick J, Brookmeyer R, Gallant J, Markowitz M, Ho D D, Richman D D, Siliciano R F. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science. 1997;278:1295–1300. doi: 10.1126/science.278.5341.1295. [DOI] [PubMed] [Google Scholar]
- 48.Ford C E, Binford S, Fuhrman S, Webber S E, Okano K, Patick A K. Abstracts of the 10th International Conference on Antiviral Research. 1997. Rhinovirus 3C protease inhibitors are efficacious against several related picornaviruses, abstr. 173; p. 34:A90. [Google Scholar]
- 49.Fromtling R A, Castañer J. VP-63843 Pleconaril WIN-63843. Drugs Future. 1997;22:40–44. [Google Scholar]
- 50.Gao M, Matusick-Kumar L, Hurlburrt W, DiTusa S F, Newcomb W W, Brown J C, McCann III P J, Deckman I, Colonno R J. The protease of herpes simplex virus type 1 is essential for functional capsid formation and viral growth. J Virol. 1994;68:3702–3712. doi: 10.1128/jvi.68.6.3702-3712.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Gathe J, Jr, Burkhardt B, Hawley P, Conant M, Peterkin J, Chapman S. Abstracts of the XI International Conference on AIDS. 1996. A randomized phase II study of VIRACEPT®, a novel HIV protease inhibitor, used in combination with stavudine vs. stavudine alone, abstr. 413. [Google Scholar]
- 52.Gibson W. Structure and assembly of the virion. Intervirology. 1996;39:389–400. doi: 10.1159/000150509. [DOI] [PubMed] [Google Scholar]
- 53.Gorbalenya A E, Blinov V M, Donchenko A P. Poliovirus-encoded proteinase 3C: a possible evolutionary link between cellular serine and cysteine proteinase families. FEBS Lett. 1986;194:253–259. doi: 10.1016/0014-5793(86)80095-3. [DOI] [PubMed] [Google Scholar]
- 54.Grakoui A, McCourt D W, Wychowski C, Feinstone S M, Rice C M. A second hepatitis C virus-encoded proteinase. Proc Natl Acad Sci USA. 1993;90:10583–10587. doi: 10.1073/pnas.90.22.10583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Grakoui A, McCourt D W, Wychowski C, Feinstone S M, Rice C M. Characterization of the hepatitis C virus-encoded serine proteinase: determination of proteinase-dependent polyprotein cleavage site. J Virol. 1993;67:2832–2843. doi: 10.1128/jvi.67.5.2832-2843.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Hahm B, Back S H, Lee T G, Wimmer E, Jang S K. Generation of a novel poliovirus with a requirement of hepatitis C virus protease NS3 activity. Virology. 1996;226:318–326. doi: 10.1006/viro.1996.0659. [DOI] [PubMed] [Google Scholar]
- 57.Hammer S M, Katzenstein D A, Hughes M D, Gendacker H, Schooley R T, Haubrich R H, Henry W K, Lederman M M, Phair J P, Nui M, Hirsch M S, Merigan T C. A trial comparing nucleoside monotherapy with combination therapy in HIV-infected adults with CD4 cell counts from 200 to 500 per cubic millimeter. AIDS Clinical Trials Group Study 175 Study Team. N Engl J Med. 1996;355:1081–1090. doi: 10.1056/NEJM199610103351501. [DOI] [PubMed] [Google Scholar]
- 58.Hammer S M, Squires K E, Hughes M D, Grimes J M, Demeter L M, Currier J S, Eron J J, Feinberg J E, Balfour H H, Deyton L R, Chodakewitz J A, Fischl M A. A controlled trial of two nucleoside analogues plus indinavir in persons with human immunodeficiency virus infection and CD4 cell counts of 200 per cubic millimeter or less. N Engl J Med. 1997;337:725–733. doi: 10.1056/NEJM199709113371101. [DOI] [PubMed] [Google Scholar]
- 59.Hamre D, Bernstein J, Donovick R. Activity of p-aminobenzaldehyde, 3-thiosemicarbazone in the chick embryo and in the mouse. Proc Soc Exp Biol Med. 1950;73:275–278. [Google Scholar]
- 60.Heinz B A, Tang J, Labus J M, Chadwell F W, Kaldor S W, Hammond M. Simple in vitro translation assay to analyze inhibitors of rhinovirus proteases. Antimicrob Agents Chemother. 1996;40:267–270. doi: 10.1128/aac.40.1.267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Henry K, Kane E, Melroe H, Simpson J, Patick A, Winslow D. Program and Abstracts of the 37th Interscience Conference on Antimicrobial Agents and Chemotherapy. Washington, D.C: American Society for Microbiology; 1997. Experience with a ritonavir/saquinavir based regimen for the treatment of HIV-infection in subjects developing increased viral loads while receiving nelfinavir, abstr. I-204; p. 282. [Google Scholar]
- 62.Hertogs K, Mellors J W, Schel P, Cauwenberghe A V, Larder B, Kemp S, Miller V, Staszewski S, Conant M, Pauwels R. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Patterns of cross-resistance among protease inhibitors in 483 clinical HIV-1 isolates, abstr. 395; p. 153. [Google Scholar]
- 63.Hertogs K, Patick A, Schel P, Van Cauwenberge A, Markowitz M, Kuritzkes D, Anderson B, Pauwels R. Abstracts of the Sixth European Conference on Clinical Aspects and Treatment of HIV-Infection. 1997. Phenotypic resistance testing (PR-RT-Antivirogram™) of clinical HIV-1 isolates confirms the unique and different resistance pathway of nelfinavir, abstr. 906-Latebreaker. [Google Scholar]
- 64.Hijikata M, Mizushima H, Akagi T, Mori S, Kakiuchi N, Kato N, Tanaka T, Kimura K, Shimotohno K. Two distinct proteinase activities required for the processing of a putative nonstructural precursor protein of the hepatitis C virus. J Virol. 1993;67:4665–4675. doi: 10.1128/jvi.67.8.4665-4675.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Hirowatari Y, Hijikata M, Shimotohno K. A novel method for analysis of viral proteinase activity encoded by hepatitis C virus in cultured cells. Anal Biochem. 1995;225:113–120. doi: 10.1006/abio.1995.1116. [DOI] [PubMed] [Google Scholar]
- 66.Ho D D, Neuman A U, Perelson A S, Chen W, Leonard J M, Markowitz M. Rapid turnover of plasma virions and CD4 lymphocytes in HIV-1 infection. Nature. 1995;373:123–126. doi: 10.1038/373123a0. [DOI] [PubMed] [Google Scholar]
- 67.Ho D D, Toyoshima T, Mo H, Kempf D J, Norbeck D, Chen C-M, Wideburg N E, Burt S K, Erickson J W, Singh M K. Characterization of human immunodeficiency virus type 1 variants with increased resistance to a C2-symmetric protease inhibitor. J Virol. 1994;68:2016–2020. doi: 10.1128/jvi.68.3.2016-2020.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Hodge C N, Aldrich P E, Bacheler L T, Chang C H, Eyermann C J, Garber S, Grubb M, Jackson D A, Jadhav P K, Korant B, Lam P Y, Maurin M B, Meek J L, Otto M J, Rayner M M, Reid C, Sharpe T R, Shum L, Winslow D L, Erickson-Viitanen S. Improved cyclic urea inhibitors of the HIV-1 protease: synthesis, potency, resistance profile, human pharmacokinetics and X-ray crystal structure of DMP 450. Chem Biol. 1996;3:301–314. doi: 10.1016/s1074-5521(96)90110-6. [DOI] [PubMed] [Google Scholar]
- 69.Holskin B P, Bukhtiyarova M, Dunn B M, Baur P, de Chastonay J, Pennington M W. A continuous fluorescence-based assay of human cytomegalovirus protease using a peptide substrate. Anal Biochem. 1995;227:148–155. doi: 10.1006/abio.1995.1264. [DOI] [PubMed] [Google Scholar]
- 70.Holwerda B C, Wittwer A J, Duffin K L, Smith C, Toth M V, Carr L S, Wiegand R C, Bryant M L. Activity of two-chain recombinant human cytomegalovirus protease. J Biol Chem. 1994;269:25911–25915. [PubMed] [Google Scholar]
- 71.Ives K J, Jacobsen H, Galpin S A, Garaev M M, Dorrell L, Mous J, Bragman K, Weber J N. Emergence of resistant variants of HIV in vivo during monotherapy with the proteinase inhibitor saquinavir. J Antimicrob Chemother. 1997;39:771–779. doi: 10.1093/jac/39.6.771. [DOI] [PubMed] [Google Scholar]
- 72.Jacobsen H, Hänggi M, Ott M, Duncan I B, Owen S, Andreoni M, Vella S, Mous J. In vivo resistance to a human immunodeficiency virus type 1 proteinase inhibitor: mutations, kinetics and frequencies. J Infect Dis. 1996;173:1379–1387. doi: 10.1093/infdis/173.6.1379. [DOI] [PubMed] [Google Scholar]
- 73.Jacobsen H, Yasargil K, Winslow D L, Craig J C, Krohn A, Duncan I B, Mous J. Characterization of human immunodeficiency virus type 1 mutants with decreased sensitivity to proteinase inhibitor Ro 31-8959. Virology. 1995;206:527–534. doi: 10.1016/s0042-6822(95)80069-7. [DOI] [PubMed] [Google Scholar]
- 74.Kahn J O, Lagakos S W, Richman D D, Cross A, Pettinelli C, Liou S H, Brown M, Volberding P A, Crumpacker C S, Beall G, et al. A controlled trial comparing continued zidovudine with didanosine in human immunodeficiency virus infection. N Engl J Med. 1992;327:581–587. doi: 10.1056/NEJM199208273270901. [DOI] [PubMed] [Google Scholar]
- 75.Kaldor S W, Hammon M, Dressman B A, Labus J M, Chadwell F W, Kline A D, Heinz B A. Glutamine-derived aldehydes for the inhibition of human rhinovirus 3C protease. Bioorg Med Chem Lett. 1995;5:2021–2026. [Google Scholar]
- 76.Kaldor S W, Kalish V J, Davies II J F, Shetty B V, Fritz J E, Appelt K, Burgess J A, Campanale K M, Chirgadze N Y, Clawson D K, Dressman B A, Hatch S D, Khalil D A, Kosa M B, Lubbehusen P P, Myesing M A, Patick A K, Reich S H, Su K S, Tatlock J H. VIRACEPT® (Nelfinavir Mesylate, AG1343): a potent, orally bioavailable inhibitor of HIV-1 protease. J Med Chem. 1997;40:3979–3985. doi: 10.1021/jm9704098. [DOI] [PubMed] [Google Scholar]
- 77.Kaplan A H, Zack J A, Knigge M, Paul D A, Kempf D J, Norbeck D W, Swanstrom R. Partial inhibition of the human immunodeficiency virus type 1 protease results in aberrant virus assembly and the formation of noninfectious particles. J Virol. 1993;67:4050–4055. doi: 10.1128/jvi.67.7.4050-4055.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Kay J, Dunn B M. Viral proteinases: weakness in strength. Biochim Biophys Acta. 1990;1048:1–18. doi: 10.1016/0167-4781(90)90015-t. [DOI] [PubMed] [Google Scholar]
- 79.Kean K M, Howell M T, Grünert S, Girard M, Jackson R J. Substitution mutations at the putative catalytic triad of the poliovirus 3C protease have differential effects on cleavage at different sites. Virology. 1993;194:360–364. doi: 10.1006/viro.1993.1268. [DOI] [PubMed] [Google Scholar]
- 80.Kempf D J, Marsh K C, Kumar G, Rodrigues A D, Denissen J F, McDonald E, Kukulka M J, Hsu A, Granneman G R, Baroldi P A, Sun E, Pizzuti D, Plattner J J, Norbeck D W, Leonard J M. Pharmacokinetic enhancement of inhibitors of the human immunodeficiency virus protease by coadministration with ritonavir. Antimicrob Agents Chemother. 1997;41:654–660. doi: 10.1128/aac.41.3.654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Keruly J C, Chaisson R E, Moore R D. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Diabetes and hyperglycemia in patients receiving protease inhibitors, abstr. 415; p. 157. [Google Scholar]
- 82.Kim J L, Morgenstern K A, Lin C, Fox T, Dwyer M D, Landro J A, Chambers S P, Markland W, Lepre C A, O’Malley E T, Harbeson S L, Rice C M, Murcko M A, Caron P R, Thomson J A. Crystal structure of the hepatitis C virus NS3 protease domain complexed with a synthetic NS4A cofactor peptide. Cell. 1996;87:343–355. doi: 10.1016/s0092-8674(00)81351-3. [DOI] [PubMed] [Google Scholar]
- 83.Koch J O, Bartenschlager R. Determinants of substrate specificity in the NS3 serine proteinase of the hepatitis C virus. Virology. 1997;237:78–88. doi: 10.1006/viro.1997.8760. [DOI] [PubMed] [Google Scholar]
- 84.Kohl N E, Emini E A, Schleif W A, Davis L J, Heimbach J C, Dixon R A, Scolnick E M, Sigal I S. Active human immunodeficiency virus protease is required for viral infectivity. Proc Natl Acad Sci USA. 1988;85:4686–4690. doi: 10.1073/pnas.85.13.4686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Kolykhalov A A, Agapov E V, Blight K J, Mihalik K, Feinstone S M, Rice C M. Transmission of hepatitis C by intrahepatic inoculation with transcribed RNA. Science. 1997;277:570–574. doi: 10.1126/science.277.5325.570. [DOI] [PubMed] [Google Scholar]
- 86.Kräusslich H-G, Wimmer E. Viral proteinases. Annu Rev Biochem. 1988;57:701–754. doi: 10.1146/annurev.bi.57.070188.003413. [DOI] [PubMed] [Google Scholar]
- 87.Lagacé L, Dô F, Malenfant E, Déziel R, Cordingley M G. Abstracts of the 11th International Conference on Antiviral Research. 1998. Inhibition of proteinase dependent processing of the scaffold proteins of HCMV capsid by monocyclic β-lactams, abstr. 158, poster. [Google Scholar]
- 88.Lal R, Hsu A, Granneman G R, El-Shourbagy T, Johnson M, Lam W, Manning L, Japour A, Sun E. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Multiple dose safety, tolerability and pharmacokinetics of ABT-378 in combination with ritonavir, abstr. 647; p. 201. [Google Scholar]
- 89.Lam P Y, Jadhav P K, Eyermann C J, Hodge C N, Ru Y, Bacheler L T, Meek J L, Otto M J, Tayner M M, Wong Y N, et al. Rational design of potent, bioavailable, nonpeptide cyclic ureas as HIV protease inhibitors. Science. 1994;263:380–384. doi: 10.1126/science.8278812. [DOI] [PubMed] [Google Scholar]
- 90.Lawrence J, Schapiro J, Winters M, Montoya J, Zolopa A, Pesano R, Winslow D, Merigan T C. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Salvage therapy with indinavir plus nevirapine in patients previously treated with two other protease inhibitors and multiple reverse transcriptase inhibitors, abstr. 422; p. 158. [Google Scholar]
- 91.Lawson M A, Semler B L. Picornavirus protein processing—enzymes, substrates, and genetic regulation. Curr Top Microbiol Immunol. 1990;161:49–80. [PubMed] [Google Scholar]
- 92.Lazdins J K, Mestan J, Goutte G, Walker M R, Bold G, Capraro H G, Klimkait T. In vitro effect of alpha-acid glycoprotein on the anti-human immunodeficiency virus (HIV) activity of the protease inhibitor CGP 61755: a comparative study with other relevant HIV protease inhibitors. J Infect Dis. 1997;175:1063–1070. doi: 10.1086/520352. [DOI] [PubMed] [Google Scholar]
- 93.Lin C, Rice C M. The hepatitis C virus NS3 serine proteinase and NS4A cofactor: establishment of a cell-free trans-processing assay. Proc Natl Acad Sci USA. 1995;92:7622–7626. doi: 10.1073/pnas.92.17.7622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Lin C, Wu J-W, Hsiao K, Thompson J A, Su M S-S. Abstracts of the 4th International Meeting on Hepatitis C Virus and Related Viruses. 1997. A novel, highthroughput assay system for the hepatitis C virus NS3 serine proteinase in cos cells, abstr. 101; p. 101. [Google Scholar]
- 95.Love R A, Parge H E, Wickersham J A, Hostomsky Z, Habuka N, Moomaw E W, Adachi T, Hostomska Z. The crystal structure of hepatitis C virus NS3 proteinase reveals a trypsin-like fold and a structural zinc binding site. Cell. 1996;87:331–342. doi: 10.1016/s0092-8674(00)81350-1. [DOI] [PubMed] [Google Scholar]
- 96.Mann M, Piazza-Hepp T, Koller E, Gibert C. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Abnormal fat distribution in AIDS patients following protease inhibitor therapy, abstr. 412; p. 157. [Google Scholar]
- 97.Margosiak S A, Vanderpool D L, Sisson W, Pinko C, Kan C-C. Dimerization of the human cytomegalovirus protease: kinetic and biochemical characterization of the catalytic homodimer. Biochemistry. 1996;35:5300–5307. doi: 10.1021/bi952842u. [DOI] [PubMed] [Google Scholar]
- 98.Markowitz, M., M. Conant, A. Hurley, R. Schluger, M. Duran, J. Peterkin, S. Chapman, A. Patick, A. Henricks, G. J. Yuen, W. Hoskins, N. Clendennin, and D. D. Ho. A preliminary evaluation of nelfinavir mesylate, an inhibitor of HIV-1 protease, to treat HIV infection. J. Infect. Dis., in press. [DOI] [PubMed]
- 99.Markowitz M, Mo H, Kempf D J, Norbeck D W, Narayana Bhat T, Erickson J W, Ho D D. Selection and analysis of human immunodeficiency virus type 1 variants with increased resistance to ABT-538, a novel protease inhibitor. J Virol. 1995;69:701–706. doi: 10.1128/jvi.69.2.701-706.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Markowitz M, Saag M, Powderly W G, Hurley A M, Hsu A, Valdes J M, Henry D, Sattler F, La Marca A, Leonard J M, Ho D D. A preliminary study of ritonavir, an inhibitor of HIV-1 protease, to treat HIV-1 infection. N Engl J Med. 1995;333:1534–1539. doi: 10.1056/NEJM199512073332204. [DOI] [PubMed] [Google Scholar]
- 101.Martell M, Esteban J I, Quer J, Genesca J, Weiner A, Esteban R, Guardia J, Gomez J. Hepatitis C virus (HCV) circulates as a population of different but closely related genomes: quasispecies nature of HCV genome distribution. J Virol. 1992;66:3225–3229. doi: 10.1128/jvi.66.5.3225-3229.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Martin J A. Recent advances in the design of HIV proteinase inhibitors. Antiviral Res. 1992;17:265–278. doi: 10.1016/0166-3542(92)90022-w. [DOI] [PubMed] [Google Scholar]
- 103.Matthews D A, Smith W W, Ferre R A, Condon B, Budahazi G, Sisson W, Villafranca J E, Janson C A, McElroy H E, Gribskov C L, Worland S. Structure of human rhinovirus 3C protease reveals a trypsin-like polypeptide fold, RNA-binding site, and means for cleaving precursor polyprotein. Cell. 1994;77:761–771. doi: 10.1016/0092-8674(94)90059-0. [DOI] [PubMed] [Google Scholar]
- 104.McDonald C K, Kuritzkes D R. Human immunodeficiency virus type 1 protease inhibitors. Arch Intern Med. 1997;157:951–959. [PubMed] [Google Scholar]
- 105.McKinlay M A, Pevear D C, Rossmann M G. Treatment of the picornavirus common cold by inhibitors of viral uncoating and attachment. Annu Rev Microbiol. 1992;46:635–654. doi: 10.1146/annurev.mi.46.100192.003223. [DOI] [PubMed] [Google Scholar]
- 106.Melnick J L. Enteroviruses: polioviruses, coxsackieviruses, echoviruses, and newer enteroviruses. In: Fields B N, Knipe D M, editors. Virology. New York, N.Y: Raven Press; 1990. pp. 549–605. [Google Scholar]
- 107.Merry C, Barry M G, Mulcahy F, Ryan M, Heavey J, Tjia J F, Gibbons S E, Breckenridge A M, Back D J. Saquinavir pharmacokinetics alone and in combination with ritonavir in HIV-infected patients. AIDS. 1997;11:F29–F33. doi: 10.1097/00002030-199704000-00001. [DOI] [PubMed] [Google Scholar]
- 108.Mocarski E S., Jr . Cytomegaloviruses and their replication. In: Fields B N, Knipe D M, editors. Virology. 3rd ed. Philadelphia, Pa: Lippincott-Raven; 1996. pp. 2447–2492. [Google Scholar]
- 109.Molla A, Korneyeva M, Gao Q, Vasavanonda S, Schipper P J, Mo H-M, Markowitz M, Chernyavskiy T, Niu P, Lyons N, Hsu A, Granneman G R, Ho D D, Boucher C A B, Leonard J M, Norbeck D W, Kempf D J. Ordered accumulation of mutations in HIV protease confers resistance to ritonavir. Nat Med. 1996;2:760–766. doi: 10.1038/nm0796-760. [DOI] [PubMed] [Google Scholar]
- 110.Morris-Jones S, Moyle G, Easterbrook P J. Antiretroviral therapies in HIV-1 infection. Expert Opin Investig Drugs. 1997;6:1049–1061. doi: 10.1517/13543784.6.8.1049. [DOI] [PubMed] [Google Scholar]
- 111.Mueller B U. Antiviral chemotherapy. Curr Opin Pediatr. 1997;9:178–183. doi: 10.1097/00008480-199704000-00012. [DOI] [PubMed] [Google Scholar]
- 112.Murphy R, Degruttola V, Gulick R, D’Aguila R, Eron J, Sommadossi J P, Smeaton L, Currier J, Tung R, Kuritzkes D. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. 141W94 with or without zidovudine/3TC in patients with no prior protease inhibitor or 3TC therapy-ACTG 347, abstr. 512; p. 175. [Google Scholar]
- 113.Nascimbeni M, Paulous S, Clavel F. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Single-cycle parameters of the inhibition of HIV infectivity by protease inhibitors, abstr. 406; p. 155. [Google Scholar]
- 114.O’Boyle D R I, Wager-Smith K, Stevens J T, Weinheimer S P. The effect of internal autocleavage on kinetic properties of the human cytomegalovirus protease catalytic domain. J Biol Chem. 1995;270:4753–4758. doi: 10.1074/jbc.270.9.4753. [DOI] [PubMed] [Google Scholar]
- 115.Otto M J, Reid C D, Garber S, Lam P Y-S, Scarnati H, Bacheler L T, Rayner M M, Winslow D L. In vitro anti-human immunodeficiency virus (HIV) activity of XM323, a novel HIV protease inhibitor. Antimicrob Agents Chemother. 1993;37:2606–2611. doi: 10.1128/aac.37.12.2606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Palmer S, Shafer R, Merigan T C. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Highly drug-resistant HIV-1 isolates are cross resistant to many of the current anti-HIV compounds in clinical development, abstr. 405; p. 155. [Google Scholar]
- 117.Partaledis J A, Yamaguchi K, Tisdale M, Blair E E, Falcione C, Maschera B, Myers R E, Pazhanisamy S, Futer O, Cullinan A B, Stuver C M, Byrn R A, Livingston D J. In vitro selection and characterization of human immunodeficiency virus type 1 (HIV-1) isolates with reduced sensitivity of hydroxyethylamino sulfonamide inhibitors of HIV-1 aspartyl protease. J Virol. 1995;69:5228–5235. doi: 10.1128/jvi.69.9.5228-5235.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Patick A K, Boritzki T J, Bloom L A. Activities of the human immunodeficiency virus type 1 (HIV-1) protease inhibitor nelfinavir mesylate in combination with reverse transcriptase and protease inhibitors against acute HIV-1 infection in vitro. Antimicrob Agents Chemother. 1997;41:2159–2164. doi: 10.1128/aac.41.10.2159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Patick A K, Ford C, Binford S, Fuhrman S, Brothers M, Meador J, Webber S E, Okano K, Zalman L, Worland S. Abstracts of the 10th International Conference on Antiviral Research. 1997. Evaluation of the antiviral activity and cytotoxicity of peptidic inhibitors of human rhinovirus 3C protease, a novel target for antiviral intervention, abstr. 116. [Google Scholar]
- 120.Patick A K, Kuritzkes D, Johnson V A, Shugarts D, Bakhtiari M, Potts K E, Farnsworth A, Anderson R, Koel J L, Hazelwood J D, Nail C D, Duran M, Markowitz M, Ho D, Richman D. Abstracts of the International Workshop on HIV Drug Resistance, Treatment Strategies and Eradication. 1997. Genotypic and phenotypic analyses of HIV-1 variants isolated from patients treated with nelfinavir and other HIV-1 protease inhibitors, abstr. 18; p. 12. [Google Scholar]
- 121.Patick A K, Duran M, Cao Y, Shugarts D, Keller M R, Mazabel E, Knowles M, Chapman S, Kuritzkes D R, Ho D D, Markowitz M. Genotypic and phenotypic characterization of human immunodeficiency virus type 1 variants isolated from patients treated with the protease inhibitor nelfinavir. Antimicrob Agents Chemother. 1998;42:2637–2644. doi: 10.1128/aac.42.10.2637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Patick A K, Mo H, Markowitz M, Appelt K, Wu B, Musick L, Kalish V, Kaldor S, Reich S, Ho D, Webber S. Antiviral and resistance studies of AG1343, an orally bioavailable inhibitor of human immunodeficiency virus protease. Antimicrob Agents Chemother. 1996;40:292–297. doi: 10.1128/aac.40.2.292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Patick A K, Rose R, Greytok J, Bechtold C M, Hermsmeier M A, Chen P T, Barrish J C, Zahler R, Colonno R J, Lin P-F. Characterization of a human immunodeficiency virus type 1 variant with reduced sensitivity to an aminodiol protease inhibitor. J Virol. 1995;69:2148–2152. doi: 10.1128/jvi.69.4.2148-2152.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Peng C, Ho B K, Chang T W, Chang N T. Role of human immunodeficiency virus type 1-specific protease in core protein maturation and viral infectivity. J Virol. 1989;63:2550–2556. doi: 10.1128/jvi.63.6.2550-2556.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Perelson A S, Neuman A U, Markowitz M, Leonard J M, Ho D D. HIV-1 dynamics in vivo: virion clearance rate, infected cell life-span and viral generation time. Science. 1996;271:1582–1586. doi: 10.1126/science.271.5255.1582. [DOI] [PubMed] [Google Scholar]
- 126.Pinko C, Margosiak S A, Vaderpool D, Gutowski J C, Condon B, Kan C-C. Single-chain recombinant human cytomegalovirus protease. J Biol Chem. 1995;270:23634–23640. doi: 10.1074/jbc.270.40.23634. [DOI] [PubMed] [Google Scholar]
- 127.Poppe S M, Slade D E, Chong K-T, Hinshaw R R, Pagano P J, Markowitz M, Ho D D, Mo H, Gorman III R R, Dueweke T J, Thaisrivongs S, Tarpley W G. Antiviral activity of the dihydropyrone PNU-140690, a new nonpeptidic human immunodeficiency virus protease inhibitor. Antimicrob Agents Chemother. 1997;41:1058–1063. doi: 10.1128/aac.41.5.1058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Qiu X, Culp J S, DiLella A G, Hellmig B, Hoog S S, Janson C A, Smith W W, Abdel-Meguid S S. Unique fold and active site in cytomegalovirus protease. Nature. 1996;383:275–278. doi: 10.1038/383275a0. [DOI] [PubMed] [Google Scholar]
- 129.Ralston R, Thudium K, Berger K, Kuo C, Gervase B, Hall J, Selby M, Kuo G, Houghton M, Choo Q-L. Characterization of hepatitis C virus envelope glycoprotein complexes expressed by recombinant vaccinia viruses. J Virol. 1993;67:6753–6761. doi: 10.1128/jvi.67.11.6753-6761.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Rice C M. Flaviviridae: the viruses and their replication. In: Fields B N, Knipe D M, editors. Virology. 3rd ed. Philadelphia, Pa: Lippincott-Raven; 1996. pp. 931–959. [Google Scholar]
- 131.Richman D D, Havlir D, Corbeil J, Looney D, Ignacia C, Spector S A, Sullivan J, Cheeseman S, Barringer K, Pauletti D, et al. Nevirapine resistance mutation of human immunodeficiency virus type 1 selected during therapy. J Virol. 1994;68:1660–1666. doi: 10.1128/jvi.68.3.1660-1666.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Roth V R, Angel J B, Kravick S. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Development of a cervical fat pad following treatment with HIV-1 protease inhibitors, abstr. 411; p. 156. [Google Scholar]
- 133.Rueckert R R. Picornaviridae and their replication. In: Fields B N, Knipe D M, editors. Virology. New York, N.Y: Raven Press; 1990. pp. 507–548. [Google Scholar]
- 134.Saag M, Gersten M, Chang Y, Greenberg S L, Yu G, Clendeninn N J. Abstracts of the 35th Annual Meeting of the Infectious Diseases Society of America. 1997. Long term virological and immunological effect of the HIV protease inhibitor, VIRACEPT® (nelfinavir mesylate) in combination with zidovudine and lamivudine; p. 396. [Google Scholar]
- 135.Saag M S, Emini E A, Laskin O L, Douglas J, Lapidus W I, Schleif W A, Whitley R J, Hildebrand C, Byrnes V W, Kappes J C, et al. A short-term clinical evaluation of L-697,661, a non-nucleoside inhibitor of HIV-1 reverse transcriptase. L-697,661 Working Group. N Engl J Med. 1993;329:1065–1072. doi: 10.1056/NEJM199310073291502. [DOI] [PubMed] [Google Scholar]
- 136.Santolini E, Migliaccio G, LaMonica N. Biosynthesis and biochemical properties of the hepatitis C virus core protein. J Virol. 1994;68:3631–3641. doi: 10.1128/jvi.68.6.3631-3641.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Saracco G, Rizzetto M. The long-term efficacy of interferon alpha in chronic hepatitis C patients: a critical review. J Gastroenterol Hepatol. 1995;10:668–673. doi: 10.1111/j.1440-1746.1995.tb01369.x. [DOI] [PubMed] [Google Scholar]
- 138.Schapiro J M, Winters M A, Stewart F, Efron B, Norris J, Kozal M J, Merigan T C. The effect of high-dose saquinavir on viral load and CD4+ T-cell counts in HIV-counts in HIV-infected patients. Ann Intern Med. 1996;124:1039–1050. doi: 10.7326/0003-4819-124-12-199606150-00003. [DOI] [PubMed] [Google Scholar]
- 139.Schenk P, Woods A S, Gibson W. The 45-kilodalton protein of cytomegalovirus (Colburn) B-capsids is an amino-terminal extension form of the assembly protein. J Virol. 1991;65:1525–1529. doi: 10.1128/jvi.65.3.1525-1529.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Schiff G M, McKinlay M A, Sherwood J R. Program and Abstracts of the 36th Interscience Conference on Antimicrobial Agents and Chemotherapy. Washington, D.C: American Society for Microbiology; 1996. Oral efficacy of VP 63843 in coxsackievirus A21 infected volunteers, abstr. H43; p. 93. [Google Scholar]
- 141.Schmit J-C, Ruiz L, Clotet B, Raventos A, Tor J, Leonard J, Desmyter J, De Clercq E, Vandamme A-M. Resistance-related mutations in the HIV-1 protease gene of patients treated for one year with the protease inhibitor ritonavir (ABT-538) AIDS. 1996;10:995–999. doi: 10.1097/00002030-199610090-00010. [DOI] [PubMed] [Google Scholar]
- 142.Sension M, Elion R, Farthing C, Currier J, Lindquist C, Richardson B, Becker M. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Open label pilot studies to assess the safety and efficacy of bid dosing regimens of VIRACEPT® (nelfinavir mesylate) combined with NRTI’s in HIV-infected treatment-naive patients, abstr. 387A; p. 151. [Google Scholar]
- 143.Shieh H-S, Kurumbail R G, Stevens A M, Stegeman R A, Sturman E J, Pak J Y, Wittwer A J, Palmier M O, Wiegand R C, Holwerda B C, Stallings W C. Three-dimensional structure of human cytomegalovirus protease. Nature. 1996;383:279–282. doi: 10.1038/383279a0. [DOI] [PubMed] [Google Scholar]
- 144.Smidt M L, Potts K E, Tucker S P, Blystone L, Steibel T R, Stallings W C, McDonald J J, Pillay D, Richman D D, Bryant M L. A mutation in human immunodeficiency virus type 1 protease at position 88, located outside the active site, confers resistance to the hydroxyethylurea inhibitor SC-55389A. Antimicrob Agents Chemother. 1996;41:515–522. doi: 10.1128/aac.41.3.515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Spruance S L, Pavia A T, Mellors J W, Murphy R, Gathe J, Jr, Stool E, Jemsek J G, Dellamonica P, Cross A, Dunkle L. Clinical efficacy of monotherapy with stavudine compared with zidovudine in HIV-infected, zidovudine-experienced patients. A randomized, double-blind, controlled trial. Bristol-Myers Squibb Stavudine/019 Study Group. Ann Intern Med. 1997;126:355–363. doi: 10.7326/0003-4819-126-5-199703010-00003. [DOI] [PubMed] [Google Scholar]
- 146.Steinkuhler C, Urbani A, Tomei L, Biasiol G, Sardana M, Bianchi E, Pessi A, De Francesco R. Activity of purified hepatitis C virus protease NS3 on peptide substrates. J Virol. 1996;70:6694–6700. doi: 10.1128/jvi.70.10.6694-6700.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Sudo K, Matsumoto Y, Matsushima M, Fujiwara M, Konno K, Shimotohno K, Shigeta S, Yokota T. Novel hepatitis C virus protease inhibitors: thiazolidine derivatives. Biochem Biophys Res Commun. 1997;238:643–647. doi: 10.1006/bbrc.1997.7358. [DOI] [PubMed] [Google Scholar]
- 148.Tebas P, Kane E, Klebert M, Simpson J, Powderly W G, Henry K. Abstracts of the 5th Conference on Retroviruses and Opportunistic Infections. 1998. Virologic responses to a ritonavir/saquinavir containing regimen in patients who have previously failed nelfinavir, abstr. 510; p. 175. [Google Scholar]
- 148a.Threlkeld D S, editor. Facts and Comparisons. 1997. Facts and comparisons: loose leaf drug information service; pp. 402A–407Z. , St. Louis, Mo. [Google Scholar]
- 149.Tisdale M, Myers R E, Maschera B, Parry N R, Oliver N M, Blair E D. Cross-resistance analysis of human immunodeficiency virus type 1 variants individually selected for resistance to five different protease inhibitors. Antimicrob Agents Chemother. 1995;39:1704–1710. doi: 10.1128/aac.39.8.1704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Tomei L, Failla C, Santolini E, De Francesco R, LaMonica N. NS3 is a serine protease required for processing of hepatitis C virus polyprotein. J Virol. 1993;67:4017–4026. doi: 10.1128/jvi.67.7.4017-4026.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Tong L, Qian C, Massariol M-J, Bonneau P R, Cordingley M G, Lagacé L. A new serine-protease fold revealed by the crystal structure of human cytomegalovirus protease. Nature. 1996;383:272–275. doi: 10.1038/383272a0. [DOI] [PubMed] [Google Scholar]
- 152.Turner R B, Wecker M T, Pohl G, Witek Jr T J, Marlin S, McNally E, Hayden F G. Program and Abstracts of the 37th Interscience Conference on Antimicrobial Agents and Chemotherapy. Washington, D.C: American Society for Microbiology; 1997. Efficacy of soluble ICAM-1 (sICAM) for prevention of rhinovirus infection and illness, abstr. H-85; p. 77. [Google Scholar]
- 153.Urvil P T, Kakiuchi N, Zhou D-M, Shimotohno K, Kumar P K R, Nishikawa S. Selection of RNA aptamers that bind specifically to the NS3 protease of hepatitis C virus. Eur J Biochem. 1997;248:130–138. doi: 10.1111/j.1432-1033.1997.t01-1-00130.x. [DOI] [PubMed] [Google Scholar]
- 154.Vacca J P, Dorsey B D, Schleif W A, Levin R B, McDaniel S L, Darke P L, Zugay J, Quintero J C, Blahy O M, Roth E, Sardana V V, Schlabach A J, Graham P I, Condra J H, Gotlib L, Holloway M K, Lin J, Chen I-W, Vastag K, Ostovic D, Anderson P S, Emini E A, Huff J R. L-735,524: an orally bioavailable human immunodeficiency virus type 1 protease inhibitor. Proc Natl Acad Sci USA. 1994;91:4096–4100. doi: 10.1073/pnas.91.9.4096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Vella S, Galluzzo C, Giannini G, Pirilo M F, Duncan I, Jacobsen H, Andreoni M, Sarmati L, Ercoli L. Saquinavir/zidovudine combination in patients with advanced HIV infection and no prior antiretroviral therapy: CD4+ lymphocyte/plasma RNA changes, and emergence of HIV strains with reduced phenotypic sensitivity. Antiviral Res. 1996;29:91–93. doi: 10.1016/0166-3542(95)00926-4. [DOI] [PubMed] [Google Scholar]
- 156.Vella S, Palmisano L. Update on protease inhibitors. Antiviral Ther. 1997;2:29–37. [Google Scholar]
- 157.Von der Helm K. Retroviral proteases: structure, function and inhibition from a non-anticipated viral enzyme to the target of a most promising HIV therapy. Biol Chem. 1996;377:765–774. [PubMed] [Google Scholar]
- 158.Webber S, Tikhe J, Worland S T, Fuhrman S A, Hendrickson T F, Matthews D A, Love R A, Patick A K, Meador J W, Ferre R A, Brown E L, DeLisle D M, Ford C E, Binford S L. Design, synthesis, and evaluation of nonpeptidic inhibitors of human rhinovirus 3C protease. J Med Chem. 1996;39:5072–5082. doi: 10.1021/jm960603e. [DOI] [PubMed] [Google Scholar]
- 159.Webber S E, Okano K, Little T L, Reich S, Xin Y, Worland S T, Furhman S A, Matthews D A, Hendrickson T F, Love R A, Patick A K, Meador III J W, Ferre R A, Brown E L, Ford C E, Binford S L. Tripeptide aldehyde inhibitors of human rhinovirus 3C protease: The design, synthesis, biological evaluation and cocrystal structure solution of P1 glutamine isosteric replacements. J Med Chem. 1998;41:2786–2805. doi: 10.1021/jm980071x. [DOI] [PubMed] [Google Scholar]
- 160.Wei X, Ghosh S K, Taylor M E, Johnson V A, Emini E A, Deutsch P, Lifson J D, Bonhoeffer S, Nowak M A, Hahn B H, Saag M S, Shaw E M. Viral dynamics in human immunodeficiency virus type 1. Nature. 1995;373:117–122. doi: 10.1038/373117a0. [DOI] [PubMed] [Google Scholar]
- 161.Weiner A J, Geysen H M, Christopherson C, Hall J E, Mason T J, Saracco G, Bonino F, Crawford K, Marion C D, Crawford K A, Brunetto M, Barr P J, Miyamura T, McHutchinson J, Houghton M. Evidence for immune selection of hepatitis C virus (HCV) putative envelope glycoprotein variants: Potential role in chronic HCV infections. Proc Natl Acad Sci USA. 1992;89:3468–3472. doi: 10.1073/pnas.89.8.3468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Welch A R, Woods A S, McNally L M, Cotter R J, Gibson W. A herpesvirus maturational proteinase, assemblin: identification of its gene, putative active site domain, and cleavage site. Proc Natl Acad Sci USA. 1991;88:10792–10796. doi: 10.1073/pnas.88.23.10792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Winslow D L, Otto M J. HIV protease inhibitors. AIDS. 1995;9:S183–S192. [PubMed] [Google Scholar]
- 164.Wlodawer A, Erickson J W. Structure-based inhibitors of HIV-1 protease. Annu Rev Biochem. 1993;62:543–585. doi: 10.1146/annurev.bi.62.070193.002551. [DOI] [PubMed] [Google Scholar]
- 165.Wong J K, Hezareh M, Günthard H F, Havlir D V, Ignacio C C, Spina C A, Richman D D. Recovery of replication-competent HIV despite prolonged suppression of plasma viremia. Science. 1997;278:1291–1295. doi: 10.1126/science.278.5341.1291. [DOI] [PubMed] [Google Scholar]
- 166.Xu J, Mendez E, Caron P R, Lin C, Murcko M A, Collett M S, Rice C M. Bovine viral diarrhea virus NS3 serine proteinase: polyprotein cleavage sites, cofactor requirements, and molecular model of an enzyme essential for pestivirus replication. J Virol. 1997;71:5312–5322. doi: 10.1128/jvi.71.7.5312-5322.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Yanagi M, Purcell R H, Emerson S U, Bukh J. Transcripts from a single full-length cDNA clone of hepatitis C virus are infectious when directly transfected into the liver of a chimpanzee. Proc Natl Acad Sci USA. 1997;94:8738–8743. doi: 10.1073/pnas.94.16.8738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Zalman L S, Brothers M A, Okano K, Webber S E. Abstracts of the 10th International Conference on Antiviral Research. 1997. A novel anti-rhinoviral compound directed at 3C protease can diminish IL-6 and IL-8 production in a transformed bronchial epithelial cell line, abstr. 172; p. A89. [Google Scholar]
- 169.Zhang R, Durkin J, Windsor W T, McNemar C, Ramanathan L, Le H V. Probing the substrate specificity of hepatitis C virus NS3 serine protease by using synthetic peptides. J Virol. 1997;71:6208–6213. doi: 10.1128/jvi.71.8.6208-6213.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]