Abstract
Background
Although survival rates for infants born extremely preterm (gestation < 28 weeks) have improved significantly in recent decades, neurodevelopmental impairment remains a major concern. Children born extremely preterm remain at high risk for cognitive impairment from early childhood to adulthood. However, there is limited evidence on genetic factors associated with cognitive impairment in this population.
Methods
First, we used a latent profile analysis (LPA) approach to characterize neurocognitive function at age 10 for children born extremely preterm. Children were classified into two groups: (1) no or low cognitive impairment, and (2) moderate-to-severe cognitive impairment. Second, we performed TOPMed-based genotype imputation on samples with genotype array data (n = 528). Third, we then conducted a genome-wide association study (GWAS) for LPA-inferred cognitive impairment. Finally, computational analysis was conducted to explore potential mechanisms underlying the variant x LPA association.
Results
We identified two loci reaching genome-wide significance (p value < 5e-8): TEA domain transcription factor 4 (TEAD4 at rs11829294, p value = 2.40e-8) and syntaxin 18 (STX18 at rs79453226, p value = 1.91e-8). Integrative analysis with brain expression quantitative trait loci (eQTL), chromatin conformation, and epigenomic annotations suggests tetraspanin 9 (TSPAN9) and protein arginine methyltransferase 8 (PRMT8) as potential functional genes underlying the GWAS signal at the TEAD4 locus.
Conclusions
We conducted a novel computational analysis by utilizing an LPA-inferred phenotype with genetics data for the first time. This study suggests that rs11829294 and its LD buddies have potential regulatory roles on genes that could impact neurocognitive impairment for extreme preterm born children.
Supplementary Information
The online version contains supplementary material available at 10.1186/s11689-022-09429-x.
Keywords: Cognitive impairment, Neurodevelopment, Preterm children, Genome-wide association study (GWAS), Latent profile analysis (LPA), Genetic mechanisms
Background
Extreme prematurity (birth < 28 weeks of gestation) remains one of the leading causes of neonatal morbidity and mortality in the USA [1]. Although survival rates for infants born extremely preterm have improved dramatically in recent decades, children born extremely preterm remain at higher risk for cognitive impairment, with lower average general intelligence and executive function deficit [2–6] and 9-fold higher risk of severe cognitive impairment compared to children born full-term [7–13]. Adverse neurodevelopmental outcomes, such as cognitive impairment, affect ~ 1 million preterm infants born each year [14] and may persist through adulthood [15–18]. Although cognitive impairment is not always severe, even mild deficits can have substantial impact, resulting in a spectrum of outcomes from difficulties in school to inability to lead an independent adult life [19]. Specific problems can include deficits in executive function, language, learning and memory, attention, perceptual-motor function, and social cognition [3, 6, 20, 21], which negatively affect well-being [19]. Cognitive impairment has life-long effects on quality of life, with significant familial and social capital costs. Although precise data are limited [22], lifetime costs collectively for children born in 2000 with intellectual disability alone are estimated at $51.2 billion [23].
Despite substantial research efforts to understand neurodevelopment outcomes, we know remarkably little about genetic factors and molecular mechanisms influencing cognitive function in preterm children. Previous genetic studies have discovered hundreds of genetic variants that can predispose children to neurodevelopmental disorders including autism spectrum disorder [24], attention deficit disorder with hyperactivity [25], intellectual disability [26, 27], specific language impairment [28], specific learning disorders [29], and childhood onset schizophrenia [30]. Some genetic studies have evaluated genetic risk factors for neurodevelopmental outcomes for preterm children or children with low birth weight [31–36]. MAOA was found to be associated with mental development throughout early childhood among preterm children [31]. A variant rs4074134 of BDNF, and a rare insertion/deletion in the intron region of SLC6A4 were significant predictors of cognitive performance at school age in a study of genetic risk factors for poor cognitive development in children with low birth weight [36]. With preterm infants from a randomized controlled trial (RCT) examining antenatal exposure to corticosteroids, Clark et al. found variants of IL1B, IL4R, and IL6 associated with lower scores on the Bayley’s Scales of Infant Development and developmental delay at age 2 [34], and Costantine et al. [33] found that variants of VIP and GRIN3A were associated with cerebral palsy. A COMT variant was associated with reduced corpus callosum size in adults with history of preterm birth [32].
However, previous studies do not explain the pathways through which these variants or genes might influence the risk of poor cognitive outcomes, and few genome-wide association studies (GWAS) examined the genomic regions associated with cognitive function among children born extremely preterm. Therefore, identifying genetic factors that are associated with children’s cognitive function and understanding related mechanisms are necessary to develop earlier screening assessments and effective precision interventions and understand why some preterm children of the same gestational age do worse than others. To advance along these directions, we utilized samples from the extremely low gestational age newborns (ELGAN) cohort [37], the largest US-based study of children born extremely preterm, to identify genetic factors associated with cognitive impairment at age 10 years. Finally, integrative analysis with brain expression quantitative trait loci (eQTL) and chromatin interactome data was performed to identify potential causal variants and functional genes underlying the GWAS associations.
Methods
Study participants
ELGAN is a multicenter cohort study originally designed to identify exposures increasing risk of structural and functional neurologic disorders in children born extremely preterm [37]. A total of 1506 infants born before the 28th week of gestation and 1249 mothers were enrolled during the years 2002–2004. Study participants were enrolled at 14 hospitals in the United States to achieve a large enough sample size and generalizability. The enrollment and consent procedures were approved by the individual institutional review boards. At the age of 10 years, 889 of the surviving children returned for follow up (ELGAN2, 92% of the 966 who were recruited for this phase of the ELGAN Study) and were assessed for cognition capacity, learning abilities, and impairments in executive function [7]. Of these children, 528 had genotype data available for analysis and thus constitute the sample size of this paper. Table 1 summarizes demographic information for the ELGAN2 cohort and the ELGAN2 subset with genetic data (n = 528) we used in our analysis.
Table 1.
Participant characteristics of the ELGAN2 subset and ELGAN2 cohort
| Variable name | ELGAN2 subset (N = 528) | ELGAN2 (N = 889) |
|---|---|---|
| n (% or SD) | n (% or SD) | |
| Infant sex | ||
| Male | 274 (51.9%) | 455 (51.2%) |
| Female | 254 (48.1%) | 434 (48.8%) |
| Cognitive impairment | ||
| No/Low | 390 (73.9%) | 660 (74.2%) |
| Moderate/Severe | 138 (26.1%) | 214 (24.1%) |
| Not reported | 0 | 15 (1.7%) |
| Gestational age | 26.1 (1.27) | 26.1 (1.28) |
| Maternal education | ||
| ≤ 12 years | 205 (38.8%) | 355 (39.9%) |
| 13–15 years | 119 (22.5%) | 202 (22.7%) |
| 16+ years | 204 (38.6%) | 306 (34.4%) |
| Not reported | 0 | 26 (2.9%) |
| Maternal smoking | ||
| Yes | 128 (24.2%) | 215 (24.2%) |
| No | 400 (75.8%) | 655 (73.7%) |
| Not reported | 0 | 19 (2.1%) |
| Race | ||
| White | 342 (64.8%) | 554 (62.3%) |
| Black | 133 (25.2%) | 227 (25.5%) |
| Other | 53 (10.0%) | 98 (11.0%) |
| Not reported | 0 | 10 (1.1%) |
| Public insurance | ||
| Yes | 167 (31.6%) | 307 (34.5%) |
| No | 361 (68.4%) | 568 (63.9%) |
| Not reported | 0 | 14 (1.6%) |
| Multiple births | ||
| Yes | 189 (35.8%) | 313 (35.2%) |
| No | 339 (64.2%) | 576 (64.8%) |
Cognitive function at age 10 years
Cognitive function at age 10 years was assessed with latent profile analysis (LPA) [38], which empirically identifies subgroups of children who share similar profiles on a set of measures. The LPA included 9 cognitive measures including verbal and nonverbal intelligence quotient (IQ) and several measures of executive function (EF). IQ was assessed with the School-Age Differential Ability Scales–II (DAS-II) Verbal and Nonverbal Reasoning scales. EF was assessed with two subtests from the DAS-II and five subtests from the Developmental NEuroPSYchological Assessment-II (NEPSY-II). Working memory was evaluated with the DAS-II Recall of Digits Backwards and Recall Sequential Order test. The NEPSY-II Auditory Attention and Auditory Response Set, Animal Sorting Inhibition, and Inhibition Switching subtests were utilized to examine auditory attention and set switching, concept generation and mental flexibility, and simple inhibition and inhibition shifting, respectively [7]. It has been shown that characterizing cognitive function using measures of executive function in addition to IQ better discriminates the academic performance and educational needs of children born extremely preterm [38]. LPA classifies subjects who share a similar pattern of scores on the measured variables, while maximizing the difference in scoring patterns across distinct profiles [39]. It assigns subjects into a finite number of profiles by identifying the most likely model that describes the heterogeneity of data, which is known as finite mixture models.
To determine the optimal number of profiles, LPA was fit to the data, and Bayesian information criteria (BIC) [40], sample-size-adjusted Bayesian information criteria (SSABIC) [41], and Lo–Mendell–Rubin-adjusted (LMR) likelihood ratio test [42] were used to assess model fit. Children were categorized by their most likely latent profile for further analysis. In this sample, a four-profile model provided the best fit for the data [38]. For our analysis, we used a binary classification that grouped participants into two previously validated distinct profile groups (LPAx) [38, 43]: no or low cognitive impairment and moderate-to-severe cognitive impairment.
Genotype data
Genomic DNA was isolated from umbilical cords and genotyping was performed using Illumina 1 Million Quad (Illumina Inc, San Diego, California). This work was done as part of the candidate gene analysis of severe intraventricular hemorrhage (IVH) in preterm born infants [44], where infants with birth weights 500–1250 g and severe grades IVH and neonates with normal cranial ultrasounds were enrolled prospectively at 24 universities. A subset of ELGAN participants were provided as additional samples along with samples from a few other studies in the IVH study [44].
We performed variant level and sample level quality control (QC) on genotype data. For variant level QC, we excluded variants with call rate < 90% or minor allele frequency (MAF) < 1%. For sample level QC, we excluded samples with missing rate > 10%. These resulted in 700,845 SNPs and 528 samples using plink v.1.90 [45, 46].
Genotype imputation
Starting with the quality controlled (QCed) genotype data, we used the Michigan imputation server [47] for phasing and imputation using TOPMed freeze 5 [48] as the reference panel. Specifically, Eagle [49] was used for phasing and Minimac4 [50, 51] was used for imputation. We performed strand matching by dropping ambiguous (i.e., A/T or C/G) SNPs and by flipping non-ambiguous SNPs that were initially in − strand when compared to alleles in the + strand observed in the TOPMed freeze 5 reference panel. Genotype data was lifted over to genome build hg38. In total, we obtained ~ 34 million well imputed variants, with 5.5 million variants having MAF > 5%, 10.5 million variants having MAF > 1%, and 17.4 million variants having MAF > 0.5%. Well imputed variants were defined by having Rsq > 0.8, where Rsq is the estimated imputation quality metric [50, 51]. To evaluate the imputation accuracy, we randomly selected 5% of the genotyped variants on chromosome 1 and performed genotype imputation with the rest 95% genotyped variants, again with the TOPMed freeze 5 reference panel. The 5% masked genotyped variants were saved for imputation quality evaluation. Specifically, we calculated the squared Pearson correlation between imputed genotypes and true observed genotypes. For the 1956 variants on chromosome 1 tested, the mean squared Pearson correlation was 0.97, which suggests that most variants were well imputed even with a relatively small sample size (Fig. S1, Additional file 1), consistent with what has been reported in the literature [52–54].
Genome-wide association analysis
We used EPACTS 3.3.0 [55] for single variant association testing. To account for relatedness among samples, we used the EMMAX (Efficient Mixed Model Association eXpedited) test [56], which is an efficient implementation of mixed model association accounting for sample structure including population structure and hidden relatedness. Biallelic SNPs with MAF > 2% (did not account for relatedness) and Rsq > 0.8 were included in the analysis. In total, 8,535,130 variants were included in the association analysis. For the 528 samples who had genotype and covariates data available, we inferred kinship matrix using EPACTS and top 10 principal components (PCs) from the genotype data using PLINK. We performed the association test on the outcome LPAx, a binary outcome that classifies children into no or low cognitive impairment and moderate-severe cognitive impairment groups. The covariates for the single variant association analysis included gestational age, maternal education, maternal race, sex of the infant, and top 10 PCs. We performed sensitivity analysis for variants showing suggestive signals by further adjusting interaction terms between covariates.
Results
Association analysis results
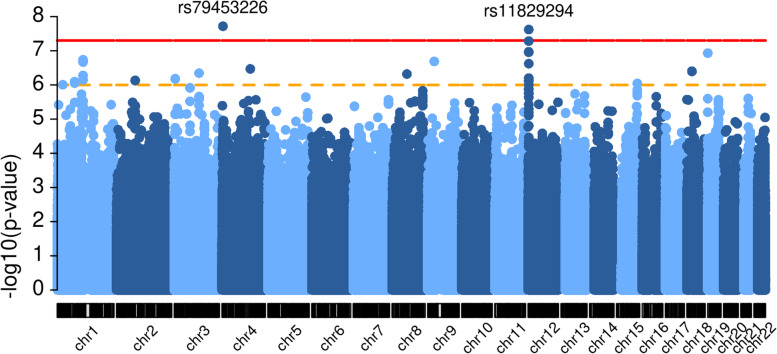
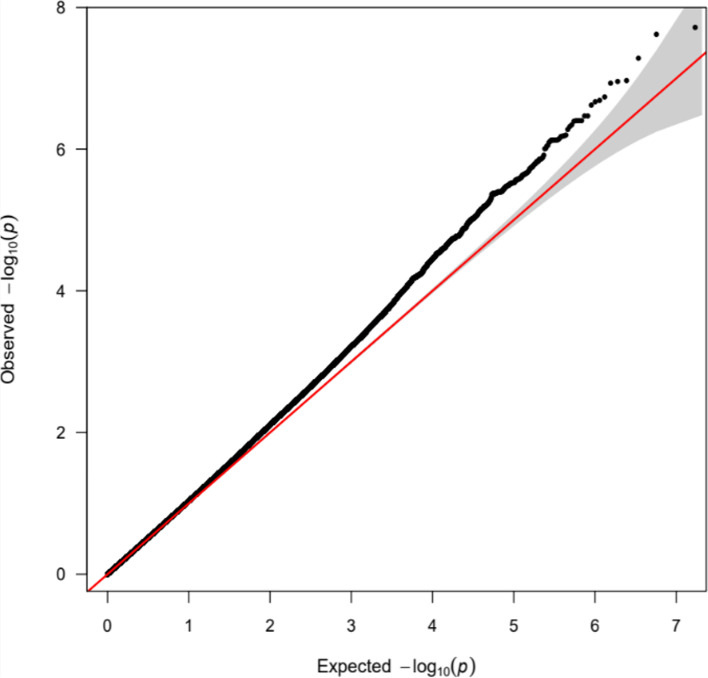
We conducted a GWAS on LPAx of 528 samples from the ELGAN2 cohort. We identified two genome-wide significant loci from the 8,535,130 variants tested: STX18 and TEAD4, which are located on chromosome 4 and chromosome 12, respectively (Fig. 1). The index SNPs are rs79453226 (MAF = 0.036) and rs11829294 (MAF = 0.145) at the STX18 and TEAD4 loci, respectively. The genomic inflation factor λ, which measures the inflation in the test statistics and is calculated as the ratio of the median of the empirically observed distribution of the test statistics to the expected median (median of a chi-square distribution with one degree of freedom), is 1.038, which suggests no significant inflation of test statistics or excess false positive rate (Fig. 2). Table 2 shows genome-wide significant variants, and suggestive variants with p values less than 1e-6. For this set of suggestive variants, we performed additional association analysis by including interaction terms between non-genetic covariates (gestational age, maternal education, maternal race, and sex of the infant). We evaluated each interaction term separately, and we found that the two genome-wide significant loci remained significant and most suggestive loci had similar significance levels as in the original model (Fig. S2, Additional file 1), indicating that our top loci are robust to the further adjustment of interactions between covariates.
Fig. 1.
Manhattan plot. The Manhattan plot visualizes the association of SNPs along the genome with the LPAx trait. X-axis represents genomic location and y-axis represents -log10(p value). Each dot represents a SNP tested. SNPs above the red horizontal line, which marks the 5 × 10−8 are considered genome-wide significant. This plot was generated using the R package karyoploteR [57]. NCBI build 38
Fig. 2.
QQ plot. A quantile-quantile (Q-Q) plot is used to characterize the extent to which the observed distribution of the test statistics follows the expected null distribution. This plot was generated using the R package qqman [58]
Table 2.
Significant and suggestive association results for LPAx
| rsID | Chra | Positiona | REF | ALT | P value | MAF | Locus | Effect size (s.e.) |
|---|---|---|---|---|---|---|---|---|
| rs79453226 | chr4 | 4483114 | G | C | 1.91e-08 | 0.036 | STX18 (intron) | 0.421 (0.074) |
| rs11829294 | chr12 | 3014153 | C | T | 2.40e-08 | 0.145 | TEAD4 (intron) | − 0.231 (0.041) |
| rs10774094 | chr12 | 3014630 | C | A | 5.21e-08 | 0.160 | TEAD4 (intron) | − 0.214 (0.039) |
| rs12322215 | chr12 | 3001421 | G | T | 1.08e-07 | 0.142 | TEAD4 (intron) | − 0.215 (0.040) |
| rs10128796 | chr12 | 3003552 | G | A | 1.11e-07 | 0.142 | TEAD4 (intron) | − 0.214 (0.040) |
| rs73916918 | chr19 | 376264 | C | T | 1.17e-07 | 0.020 | THEG (5’ UTR) | 0.522 (0.097) |
| rs59359613 | chr1 | 113154555 | C | T | 1.83e-07 | 0.023 | intergenic | 0.465 (0.088) |
| rs16913588 | chr9 | 28733517 | T | C | 2.05e-07 | 0.036 | intergenic | 0.396 (0.075) |
| rs58545250 | chr1 | 113172866 | T | C | 2.14e-07 | 0.024 | RP11-389O22.4 (downstream) | 0.438 (0.083) |
| rs61114884 | chr12 | 3004684 | T | A | 2.39e-07 | 0.135 | TEAD4 (intron) | − 0.210 (0.040) |
| rs28411755 | chr4 | 124309484 | T | C | 3.40e-07 | 0.025 | intergenic | 0.431 (0.083) |
| rs7657348 | chr4 | 124310584 | A | G | 3.40e-07 | 0.025 | intergenic | 0.431 (0.083) |
| rs76946462 | chr18 | 22326760 | A | G | 3.96e-07 | 0.057 | intergenic | 0.278 (0.054) |
| rs77039990 | chr18 | 22327483 | G | A | 3.97e-07 | 0.057 | intergenic | 0.278 (0.054) |
| rs76500624 | chr18 | 22326662 | G | A | 3.97e-07 | 0.057 | intergenic | 0.278 (0.054) |
| rs75050632 | chr18 | 22327123 | G | A | 4.03e-07 | 0.057 | intergenic | 0.277 (0.054) |
| rs115606157 | chr3 | 108839197 | T | G | 4.53e-07 | 0.030 | TRAT1 (intron) | 0.385 (0.07) |
| rs73690518 | chr8 | 65242418 | C | T | 4.80e-07 | 0.048 | intergenic | 0.355 (0.070) |
| rs17031018 | chr1 | 113100296 | A | G | 5.27e-07 | 0.035 | LRIG2 (intron) | 0.377 (0.074) |
| rs61917974 | chr12 | 3011978 | T | C | 6.33e-07 | 0.109 | TEAD4 (intron) | − 0.223 (0.044) |
| rs12296242 | chr12 | 3006641 | G | C | 6.49e-07 | 0.133 | TEAD4 (intron) | − 0.202 (0.040) |
| rs143601180 | chr3 | 4370781 | A | G | 6.56e-07 | 0.032 | SUMF1 (intron) | 0.383 (0.076) |
| rs79946490 | chr3 | 4385952 | C | T | 6.64e-07 | 0.032 | SUMF1 (intron) | 0.383 (0.076) |
| rs17031120 | chr1 | 113144809 | T | C | 7.12e-07 | 0.021 | intergenic | 0.462 (0.092) |
| rs2163633 | chr2 | 81884390 | C | A | 7.36e-07 | 0.046 | intergenic | − 0.350 (0.070) |
| rs6716465 | chr2 | 81871292 | G | C | 7.43e-07 | 0.045 | intergenic | − 0.350 (0.070) |
| rs11062457 | chr12 | 3010236 | C | T | 7.44e-07 | 0.145 | TEAD4 (intron) | − 0.201 (0.040) |
| rs72921448 | chr2 | 81824332 | T | C | 7.44e-07 | 0.045 | intergenic | − 0.350 (0.070) |
| rs2286647 | chr12 | 3010912 | C | T | 7.46e-07 | 0.145 | TEAD4 (intron) | − 0.201 (0.040) |
| rs116629423 | chr2 | 81858789 | A | G | 7.47e-07 | 0.045 | intergenic | − 0.350 (0.070) |
| rs143923810 | chr12 | 2988024 | C | T | 7.73e-07 | 0.139 | TEAD4 (intron) | − 0.194 (0.039) |
| rs10493588 | chr1 | 76227682 | C | T | 8.04e-07 | 0.057 | ST6GALNAC3 (intron) | 0.279 (0.056) |
| rs17098434 | chr1 | 76232427 | G | A | 8.85e-07 | 0.057 | ST6GALNAC3 (intron) | 0.277 (0.056) |
| rs8025099 | chr15 | 91488748 | C | A | 9.06e-07 | 0.486 | CRAT37 (intron) | − 0.129 (0.026) |
| rs12318430 | chr12 | 3006040 | C | A | 9.75e-07 | 0.132 | TEAD4 (intron) | − 0.201 (0.040) |
| rs9424366 | chr1 | 24475103 | G | C | 9.86e-07 | 0.036 | NIPAL3 (downstream) | 0.349 (0.070) |
Ordered by significance
aNCBI build 38
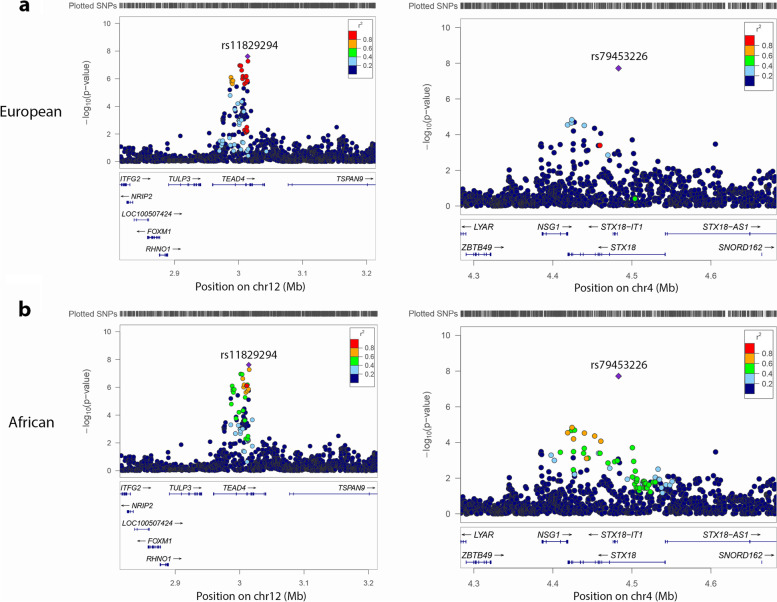
Figure 3 shows LocusZoom [59] plots for the two genome-wide significant loci, with linkage disequilibrium (LD) from TOP-LD [60], calculated using TOPMed European and African participants. We can see that in the European population, the lead variant rs11829294 in the TEAD4 region has a number of LD tags (e.g., 21 variants with r2 ≥ 0.8) and some of them had highly significant p values; by contrast, the lead variant rs79453226 in the STX18 region has fewer LD tags (2 variants with r2 ≥ 0.8) that showed suggestive association (Fig. 3a). In the African population, the lead variant rs11829294 in the TEAD4 region has only 2 LD tags, which did not have significant or suggestive association, and the lead variant rs79453226 in the STX18 region has no LD tag with r2 ≥ 0.8 (Fig. 3b).
Fig. 3.
Locus zoom plots for the two genome-wide significant loci. Colors represent linkage disequilibrium r2 values calculated from TOPMed individuals with the lead SNP in each plot. a Locus zoom plots with linkage disequilibrium r2 values calculated from TOPMed European ancestry individuals. b Locus zoom plots with linkage disequilibrium r2 values calculated from TOPMed African ancestry individuals. NCBI build 38
Epigenetic functional annotations
To further investigate the two loci identified for potential mechanisms, we examined several functional annotation metrics, including the CADD phred score [61] and the fathmm MKL score [62]. CADD phred score measures the deleteriousness of variants and is computed as -10*log10(rank/total). A CADD phred score of ≥ 10 indicates that the variant is predicted to be among the 10% most deleterious variants in the human genome, a score of ≥ 20 indicates among the 1% most deleterious. The fathmm MKL score predicts the functional consequences of variants where values above 0.5 are generally considered deleterious, and values below 0.5 neutral or benign. We also looked at the Genehancer feature [63] and the genes predicted by Genehancer. Table 3 shows functional annotations for variants that passed the suggestive p value threshold (p value < 1e-6). We observed that variants rs9424366, rs79946490, rs58545250, and rs17031018 were among the top 10% most deleterious in the human genome, and variant rs16913588 was predicted to be deleterious (with a fathmm MKL score of 0.97). Several variants were assigned by Genehancer as falling into enhancer regions with target genesTSPAN9, ITPR1, and CLIC4. These results provide evidence that some of the variants might have deleterious effects that are relevant to neurocognitive development in preterm children and suggest additional genes that might be functionally related.
Table 3.
Epigenetic functional annotations for selected genome-wide significant and suggestive associations
| rsID | P value | CADD phred | FathmmMKL | Genehancer feature | Genehancer connected gene | Locus |
|---|---|---|---|---|---|---|
| rs11829294 | 2.40e-08 | 3.728 | 0.21 | enhancer | TSPAN9 | TEAD4 (intron) |
| rs10774094 | 5.21e-08 | 0.805 | 0.10 | enhancer | TSPAN9 | TEAD4 (intron) |
| rs16913588 | 2.05e-07 | 7.525 | 0.97 | – | – | intergenic |
| rs58545250 | 2.14e-07 | 9.661 | 0.49 | – | – | RP11-389O22.4 (downstream) |
| rs61114884 | 2.39e-07 | 3.602 | 0.15 | enhancer | TSPAN9 | TEAD4 (intron) |
| rs17031018 | 5.27e-07 | 9.16 | 0.30 | – | – | LRIG2 (intron) |
| rs79946490 | 6.64e-07 | 10.19 | 0.23 | enhancer | ITPR1 | SUMF1 (intron) |
| rs11062457 | 7.44e-07 | 0.362 | 0.13 | enhancer | TSPAN9 | TEAD4 (intron) |
| rs2286647 | 7.46e-07 | 0.16 | 0.07 | enhancer | TSPAN9 | TEAD4 (intron) |
| rs143923810 | 7.73e-07 | 1.518 | 0.04 | enhancer | TSPAN9 | TEAD4 (intron) |
| rs9424366 | 9.86e-07 | 13.82 | 0.13 | enhancer | CLIC4 | NIPAL3 (downstream) |
Chromatin interactions
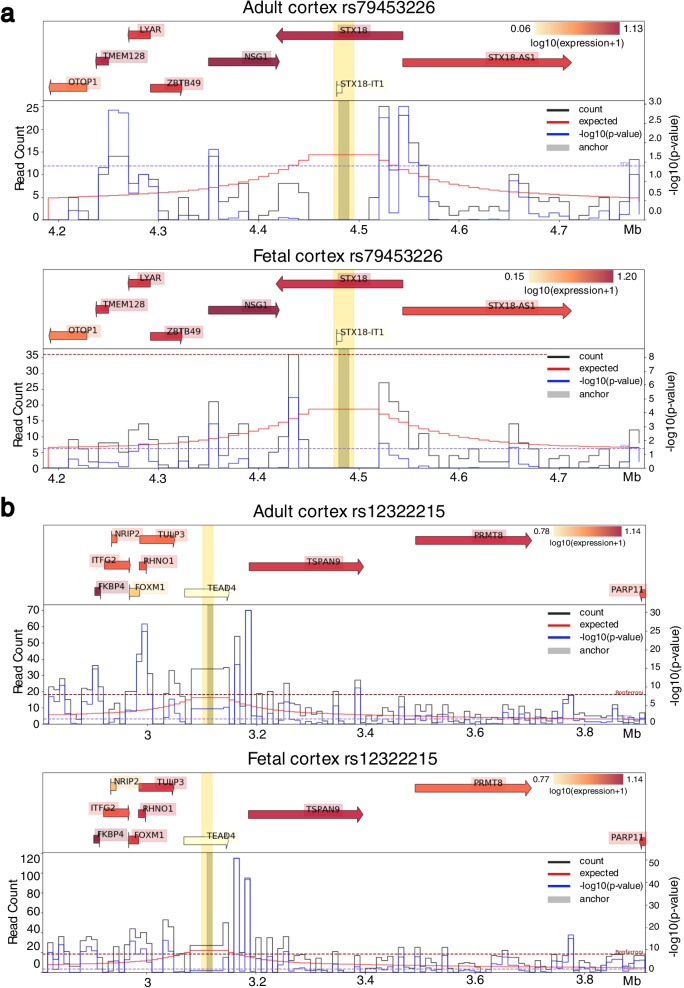
We examined chromatin conformation data for additional functional implications based on physical contacts from Hi-C and alike technologies. Figure 4 shows virtual 4C plots generated by HUGIn2 [64] for the top two loci in adult cortex and fetal cortex Hi-C data [65]. HUGIn2 is a web-based viewer of genome-wide chromatin conformation data to explore chromatin spatial organization across multiple human cell lines and primary tissues. HUGIn2 can additionally incorporate data from multiple sources including genetic variants, chromatin organization features (e.g., topologically associating domains (TADs) [66], frequently interacting regions (FIREs) [67]), gene expression, and epigenetic annotations. For our purpose, we examined ± 500 kb regions around each locus. Significant chromatin interactions between the putative regulatory regions (harboring some GWAS variant(s)) and promoters of genes suggest the likely causal or effector genes regulated by the GWAS variant(s). The results were consistent between adult cortex and fetal cortex. The variant rs79453226 at the STX18 locus was linked to the promoter regions of several genes, including STX18 and NSG1 (Fig. 4a), and the variant rs12322215 at the TEAD4 locus was linked to FKBP4, FOXM1, RHNO1, TULP3, TSPAN9, and PRMT8 (Fig. 4b).
Fig. 4.
Virtual 4C plots. Centered at a rs79453226 b rs12322215 in adult cortex and fetal cortex. The bin containing the anchor position is indicated as a thick grey vertical bar. Different genes or regions can be highlighted in yellow. On the top is gene expression data with gene locations. Each gene is indicated by an arrow pointing the direction of transcription. The start site is indicated by the tail of the arrow. Each gene is labeled by its common name and highlighted in red indicating the expression level: the deeper the red color the higher the expression. On the bottom is the chromatin interaction Hi-C data that is plotted as a virtual 4C plot with the given anchor position. The black line shows the observed counts, the red line shows the expected counts, and the blue line shows the -log10(p value). The range of the -log10(p value) is plotted on the y-axis on the right while the range of the count data is shown on the left. The x-axis is the genomic location in Mb. NCBI build 37
Overlapping with brain eQTL
Next, we investigated whether we could find any brain eQTL signals among the top variants. We examined all variants with LD r2 ≥ 0.6 with variants that passed the suggestive p value threshold (p value < 1e-6) using LD calculated from TOPMed European ancestry samples. Table 4 shows variants overlapped with commonMind eQTL [68] with FDR < 5%. Multiple brain eQTLs for PRMT8 on chromosome 12 in LD with the index SNP rs11829294 were identified.
Table 4.
Variants overlapped with commonMind eQTL
| rsID | Gene | Chra | Positiona | FDR | Index SNP | LD r2 with the index SNP |
|---|---|---|---|---|---|---|
| rs143923810 | PRMT8 | chr12 | 2988024 | 0.010 | rs11829294 | 0.724 |
| rs7302783 | PRMT8 | chr12 | 2989245 | 0.010 | rs11829294 | 0.724 |
| rs7302789 | PRMT8 | chr12 | 2989254 | 0.010 | rs11829294 | 0.720 |
| rs10082968 | PRMT8 | chr12 | 2990125 | 0.025 | rs11829294 | 0.720 |
| rs12322215 | PRMT8 | chr12 | 3001421 | 0.048 | rs11829294 | 0.883 |
| rs10128796 | PRMT8 | chr12 | 3003552 | 0.045 | rs11829294 | 0.883 |
aNCBI build 38
Overlapping with selective sweeps
We also examined whether the identified top variants overlapped with selective sweeps detected by S/HIC [69]. Table 5 shows multiple variants in LD (r2 ≥ 0.6) with the top variants overlapped with selective sweeps: all except one on chromosome 12 at the TEAD4 locus and one in an intergenic locus on chromosome 2. We found that some variants at the TEAD4 locus were located in soft sweep regions, where selection on standing variation produced qualitatively different skews in LD and allele frequencies.
Table 5.
Variants overlapped with selective sweeps
| Chra | Positiona | Start | End | Selective sweeps |
|---|---|---|---|---|
| chr12 | 3095812 | 2800000 | 3100000 | CEU: soft, GWD: soft, LWK: soft, PEL: soft, YRI: soft |
| chr12 | 3097190 | 2800000 | 3100000 | CEU: soft, GWD: soft, LWK: soft, PEL: soft, YRI: soft |
| chr12 | 3098411 | 2800000 | 3100000 | CEU: soft, GWD: soft, LWK: soft, PEL: soft, YRI: soft |
| chr12 | 3098420 | 2800000 | 3100000 | CEU: soft, GWD: soft, LWK: soft, PEL: soft, YRI: soft |
| chr12 | 3099291 | 2800000 | 3100000 | CEU: soft, GWD: soft, LWK: soft, PEL: soft, YRI: soft |
| chr2 | 82111514 | 82100000 | 82200000 | GWD: soft, YRI: soft |
Population: CEU (UT, USA), GWD (Western Divisions, the Gambia), LWK (Webuye, Kenya), PEL (Lima, Peru), YRI (Ibadan, Nigeria). Start and end are start and end positions of selective sweep regions
aNCBI build 37
Discussion
Cognitive impairment is highly prevalent among children born extremely preterm. Yet, limited evidence is available on the genetic factors that may contribute to this kind of impairment. In this study, we aimed to identify genetic factors that are associated with children’s cognitive function and understand related genetic mechanisms by utilizing samples from the ELGAN cohort. Leveraging an LPA-derived phenotype and genetics data, we identified two genome-wide significant loci in our genome-wide association analysis for LPAx (a data-derived cognitive impairment outcome): TEAD4 (rs11829294, p value = 2.40e-8) and STX18 (rs79453226, p value = 1.91e-8).
We utilized chromatin conformation data from multiple human cell lines and primary tissues to see whether there are significant chromatin interactions between the two genome-wide significant loci and their neighboring regions. In adult cortex and fetal cortex, we found that variant rs12322215 (p value = 1.08e-07) in high LD with rs11829294 (r2 = 0.883) is linked to promoter regions of a few genes including TSPAN9 and PRMT8 (Fig. 4). Furthermore, the association at the TEAD4 locus rs11829294 and a few other variants that showed suggestive significance at the same locus were assigned by Genehancer as falling into the enhancer region of TSPAN9 (Table 3). We also observed TSPAN9 is highly expressed in both adult cortex and fetal cortex but not in hippocampus, and we did not observe similar chromatin interactions in hippocampus (Fig. 4). TSPAN9 is located at chr12:3,077,355-3,286,564 (GRCh38/hg38) and is one of tetraspanins, a superfamily of glycoproteins that function as “organizers” of cell membranes by recruiting other receptors and signaling proteins into tetraspanin-enriched microdomains and induce normal platelet activation [70]. Those proteins mediate signal transduction events that play a role in the regulation of cell development, activation, growth, and motility. TSPAN9 is highly expressed in normal brain tissues, including cerebellum and cerebellar hemisphere [71]. These pieces of evidence suggest the potential regulatory role of rs11829294 and its LD buddies on the TSPAN9 gene that could impact cognitive development among children born extremely preterm.
We also performed integrative analysis with brain eQTL to identify potential functional genes underlying the genome-wide significant association. A few brain eQTL for PRMT8 were found to be in high LD with rs11829294 (Table 4). PRMT8 is a member of the protein arginine N-methyltransferase (PRMT) family, which mediates protein arginine methylation, a common post-translational modification that has been implicated in signal transduction, RNA processing, transcriptional regulation, and DNA repair [72]. PRMT8 was found to be associated with the plasma membrane and has a tissue-specific expression in brain. Specifically, it is highly expressed in nucleus accumbens (basal ganglia), putamen (basal ganglia), cortex, caudate (basal ganglia), frontal cortex (Brodmann area 9), and anterior cingulate cortex (Brodmann area 24) [71]. It was also identified as a tissue-restricted enzyme responsible for proper asymmetric dimethylarginine (ADMA) level in postmitotic neurons where PRMT8-dependent arginine methylation is required for neuroprotection against age-related increased of cellular stress [73]. Moreover, PRMT8 in human embryonic stem cells (hESCs) plays an important role not only in maintaining pluripotency but also in controlling mesodermal differentiation [74]. Along with the evidence that variant rs12322215 is linked to the promoter region of PRMT8, we conclude that PRMT8 is another biologically plausible gene regulated by eQTL at the TEAD4 locus that could have potential effects on cognitive impairment among preterm children.
Additionally, we found that multiple variants in LD with rs11829294 overlapped with selective sweeps detected by S/HIC (Table 5). There is evidence that soft sweeps are widespread and account for the vast majority of recent human adaptation, and positive selection may often proceed via “soft sweeps” acting on mutations already present within a population. Furthermore, linked positive selection affects patterns of variation across much of the genome, and may increase the frequencies of deleterious mutations [69]. Therefore, these variants in soft sweep regions provide additional evidence for the associations found at the TEAD4 locus being biologically plausible and potentially causal.
For rs79453226, we found that it is linked to promoter regions of STX18 and NSG1 (Fig. 4). We did not find as much evidence for the STX18 locus supporting the significant association as for the TEAD4 locus. By examining LD r2 values calculated from TOPMed, we observed that rs11829294 and rs79453226 have different LD structures at their loci (Fig. 3). Specifically, rs11829294 has a number of LD buddies with r2 ≥ 0.8 showing suggestive association with LPAx in the European population. In contrast, rs79453226 has fewer LD buddies and is not in high LD with any of the suggestive variants. Therefore, rs11829294 is more likely to tag effects from causal variants than rs79453226.
With association results and other information considered, we did not have direct evidence indicating TEAD4 and STX18 as causal genes. However, there is evidence that TEAD4 and STX18 are related to placental development and brain respectively.TEAD4 is a member of the TEAD transcription factor family, which is best known for transcriptional output of the Hippo signaling pathway and has been implicated in processes such as development, cell growth and proliferation, tissue homeostasis, and regeneration [75]. TEADs have been found to be evolutionarily conserved, and have been shown to play important roles in various biological processes and human disease [76, 77]. Mouse knockout studies showed that TEAD4 is specifically required for embryo implantation and trophectoderm lineage determination [78, 79], which play important roles in placental development. TEAD4 null mice are embryonic lethal due to failure in embryo implantation; however, disruption of TEAD4 after embryo implantation results in normal development [78, 79]. TEADs seem to have important biological functions, but studies thoroughly characterizing TEAD function and regulation are lacking. In the future, we can utilize genome-wide DNA methylation, mRNA, and miRNA data from the placenta to study this gene more closely. The gene STX18 encodes a member of the syntaxin family of soluble N-ethylmaleimide-sensitive factor attachment protein receptors (SNAREs) which is part of a membrane tethering complex that includes other SNAREs and several peripheral membrane proteins, and is involved in vesicular transport between the endoplasmic reticulum (ER) and the Golgi complex [80]. It has also been shown that STX18 is important for the organization of two ER subdomains, smooth/rough ER membranes and ER exit sites by mediating the fusion of retrograde membrane carriers with the ER membrane [81]. Knockdown of STX18 caused a global change in ER membrane architecture, leading to the segregation of the smooth and rough ER. Moreover, the organization of ER exit sites was markedly changed concomitantly with dispersion of the ER-Golgi intermediate compartment and the Golgi complex. Variants in STX18 were previously found to be associated with brain volume measurement and neuroimaging measurement [82, 83].
One limitation of our analysis is that our results may not be generalizable to children who are not extreme premature. Another issue is the small sample size, although we were able to impute most variants well (Fig. S1, Additional file), it limits the statistical power of the association analysis. The few genome-wide significant single variant associations we found, and the non-statistically significant heritability estimate also suggest the need for better powered analyses (Additional file 1). It is also possible that variants included in our analyses are in low or moderate LD with true causal variants which are rare and cannot be well-imputed in the ELGAN2 cohort. While ELGAN2 is the largest cohort with genotype and long-term cognitive assessment for extremely preterm children currently available in the USA, in the future we hope to study a larger population with longitudinal data of cognitive function, to investigate whether there are genetic variants that interact with perinatal and neonatal immune factors to increase risk for development of trajectories of impaired cognitive function.
Conclusions
In this work, we present an innovative computational approach that combines LPA with multi-faceted genomic analysis to investigate potential genetic risk factors underlying cognitive impairment among children born extremely preterm. Our association analysis identified two genome-wide significant loci: TEAD4 at rs11829294 and STX18 at rs79453226. Further genomic analysis suggests that rs11829294 and its LD buddies have potential regulatory roles on likely functionally relevant genes TSPAN9 and PMRT8. This study provides new mechanistic insight into neurocognitive function among children born extremely preterm by performing an imputation-based GWAS with subsequent prioritization of causal variants and effector genes.
Supplementary Information
Additional file 1. SNP-heritability estimation with GCTA, Fig. S1, and Fig. S2.
Acknowledgements
The authors gratefully acknowledge the contributions of ELGAN participants and their families, as well as those of our colleagues. We thank Dr. Timothy C. Heeren for his expertise on the LPA analysis.
Abbreviations
- LPA
Latent profile analysis
- GWAS
Genome-wide association study
- eQTL
Expression quantitative trait loci
- RCT
Randomized controlled trial
- ELGAN
Extremely low gestational age newborns
- IQ
Intelligence quotient
- EF
Executive functioning
- DAS-II
Differential Ability Scales–II
- NEPSY-II
NEuroPSYchological Assessment-II
- BIC
Bayesian information criteria
- SSABIC
Sample-size-adjusted Bayesian information criteria
- LMR
Lo–Mendell–Rubin
- IVH
Intraventricular hemorrhage
- QC
Quality control
- MAF
Minor allele frequency
- PCs
Principal components
- GRM
Genetic relationship matrix
- REML
Restricted maximum likelihood
- Q-Q
Quantile-quantile
- LD
Linkage disequilibrium
- FIREs
Frequently interacting regions
- ADMA
Asymmetric dimethylarginine
- hESCs
Human embryonic stem cells
- SNAREs
Soluble N-ethylmaleimide-sensitive factor attachment protein receptors
- ER
Endoplasmic reticulum
Authors’ contributions
Conceptualization: YL, TMO, RCF, HPS; data analysis: WL, LH, QS, GW, YL, HPS; data curation: WL, AB, TMO, RCF, YL, HPS; funding acquisition: KCKK, RMJ, TMO, RCF, YL, HPS; methodology: WL, XT, YL, HPS; writing—original draft: WL, YL, HPS; writing—review and editing: WL, LH, QS, AB, GW, XT, KCKK, RMJ, TMO, RCF, YL, HPS. All authors have read and approved the final manuscript and acknowledged the statement.
Funding
This work was supported by the National Institutes of Health (NIH), including awards funded by the National Institute of Child Health and Human Development under grant# 5R01HD092374-05, the National Institute of Nursing Research under grant# 5R01NR019245-02, and the Office of the NIH Director grants# P50HD103573, U01DA052713, and UH3OD023348.
Availability of data and materials
The genotype data analyzed during the current study are not publicly available but are available from the corresponding author on reasonable request. Epigenetic functional annotations, chromatin interaction, brain eQTL, and selective sweeps data are publicly available.
Declarations
Ethics approval and consent to participate
Participating mothers provided informed consent following admission to the hospital, before birth, or immediately following birth. Study procedures were approved by the Institutional Review Board at each of the 14 participating ELGAN sites.
Consent for publication
Consent for publication was granted during in informed consent process.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rebecca C. Fry, Yun Li, and Hudson P. Santos are co-senior authors.
Contributor Information
Yun Li, Email: yunli@med.unc.edu.
Hudson P. Santos, Jr, hsantosj@email.unc.edu.
References
- 1.Mathews TJ, Driscoll AK. Trends in Infant Mortality in the United States, 2005-2014. NCHS Data Brief. 2017. pp. 1–8. [PubMed] [Google Scholar]
- 2.Korologou-Linden R, Anderson EL, Jones HJ, Davey Smith G, Howe LD, Stergiakouli E. Polygenic risk scores for Alzheimer’s disease, and academic achievement, cognitive and behavioural measures in children from the general population. Int J Epidemiol. 2019;48:1972–1980. doi: 10.1093/ije/dyz080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ding S, Lemyre B, Daboval T, Barrowman N, Moore GP. A meta-analysis of neurodevelopmental outcomes at 4-10 years in children born at 22-25 weeks gestation. Acta Paediatr. 2019;108:1237–1244. doi: 10.1111/apa.14693. [DOI] [PubMed] [Google Scholar]
- 4.Serenius F, Källén K, Blennow M, Ewald U, Fellman V, Holmström G, et al. Neurodevelopmental outcome in extremely preterm infants at 2.5 years after active perinatal care in Sweden. JAMA. 2013;309:1810–1820. doi: 10.1001/jama.2013.3786. [DOI] [PubMed] [Google Scholar]
- 5.Moore T, Hennessy EM, Myles J, Johnson SJ, Draper ES, Costeloe KL, et al. Neurological and developmental outcome in extremely preterm children born in England in 1995 and 2006: the EPICure studies. BMJ. 2012;345:e7961. doi: 10.1136/bmj.e7961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Pascal A, Govaert P, Oostra A, Naulaers G, Ortibus E, Van den Broeck C. Neurodevelopmental outcome in very preterm and very-low-birthweight infants born over the past decade: a meta-analytic review. Dev Med Child Neurol. 2018;60:342–355. doi: 10.1111/dmcn.13675. [DOI] [PubMed] [Google Scholar]
- 7.Joseph RM, O’Shea TM, Allred EN, Heeren T, Hirtz D, Jara H, et al. Neurocognitive and academic outcomes at age 10 years of extremely preterm newborns. Pediatrics. 2016;137(4). 10.1542/peds.2015-4343. [DOI] [PMC free article] [PubMed]
- 8.Kuban KCK, Joseph RM, O’Shea TM, Allred EN, Heeren T, Douglass L, et al. Girls and boys born before 28 weeks gestation: risks of cognitive, behavioral, and neurologic outcomes at age 10 years. J Pediatr. 2016;173:69–75.e1. doi: 10.1016/j.jpeds.2016.02.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Johnson S, Fawke J, Hennessy E, Rowell V, Thomas S, Wolke D, et al. Neurodevelopmental disability through 11 years of age in children born before 26 weeks of gestation. Pediatrics. 2009;124(2):e249-57. 10.1542/peds.2008-3743. [DOI] [PubMed]
- 10.Russ SA, Larson K, Halfon N. A national profile of childhood epilepsy and seizure disorder. Pediatrics. 2012;129(2):256–64. 10.1542/peds.2010-1371. [DOI] [PubMed]
- 11.Johnson S, Marlow N. Early and long-term outcome of infants born extremely preterm. Arch Dis Child. 2017;102:97–102. doi: 10.1136/archdischild-2015-309581. [DOI] [PubMed] [Google Scholar]
- 12.Chan E, Leong P, Malouf R, Quigley MA. Long-term cognitive and school outcomes of late-preterm and early-term births: a systematic review. Child Care Health Dev. 2016;42:297–312. doi: 10.1111/cch.12320. [DOI] [PubMed] [Google Scholar]
- 13.Van Naarden BK, Christensen D, Doernberg N, Schieve L, Rice C, Wiggins L, et al. Trends in the prevalence of autism spectrum disorder, cerebral palsy, hearing loss, intellectual disability, and vision impairment, metropolitan atlanta, 1991-2010. PLoS One. 2015;10:e0124120. doi: 10.1371/journal.pone.0124120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Blencowe H, Lee ACC, Cousens S, Bahalim A, Narwal R, Zhong N, et al. Preterm birth-associated neurodevelopmental impairment estimates at regional and global levels for 2010. Pediatr Res. 2013;74(Suppl 1):17–34. doi: 10.1038/pr.2013.204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Crowley P. Prophylactic corticosteroids for preterm birth. Cochrane Database Syst Rev. 2000;(2):CD000065. 10.1002/14651858.CD000065. [DOI] [PubMed]
- 16.Schmidt B, Anderson PJ, Doyle LW, Dewey D, Grunau RE, Asztalos EV, et al. Survival without disability to age 5 years after neonatal caffeine therapy for apnea of prematurity. JAMA. 2012;307:275–282. doi: 10.1001/jama.2011.2024. [DOI] [PubMed] [Google Scholar]
- 17.Schmidt B, Davis P, Moddemann D, Ohlsson A, Roberts RS, Saigal S, et al. Long-term effects of indomethacin prophylaxis in extremely-low-birth-weight infants. N Engl J Med. 2001;344:1966–1972. doi: 10.1056/NEJM200106283442602. [DOI] [PubMed] [Google Scholar]
- 18.Brooks-Gunn J, McCarton CM, Casey PH, McCormick MC, Bauer CR, Bernbaum JC, et al. Early intervention in low-birth-weight premature infants. Results through age 5 years from the Infant Health and Development Program. JAMA. 1994;272:1257–1262. [PubMed] [Google Scholar]
- 19.Maxwell JR, Yellowhair TR, Oppong AY, Camacho JE, Lowe JR, Jantzie LL, et al. Cognitive development in preterm infants: multifaceted deficits reflect vulnerability of rigorous neurodevelopmental pathways. Minerva Pediatr. 2017;69:298–313. doi: 10.23736/S0026-4946.17.04905-2. [DOI] [PubMed] [Google Scholar]
- 20.Murray AL, Scratch SE, Thompson DK, Inder TE, Doyle LW, Anderson JFI, et al. Neonatal brain pathology predicts adverse attention and processing speed outcomes in very preterm and/or very low birth weight children. Neuropsychology. 2014;28:552–562. doi: 10.1037/neu0000071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Twilhaar ES, Wade RM, de Kieviet JF, van Goudoever JB, van Elburg RM, Oosterlaan J. Cognitive outcomes of children born extremely or very preterm since the 1990s and associated risk factors: a meta-analysis and meta-regression. JAMA Pediatr. 2018;172:361–367. doi: 10.1001/jamapediatrics.2017.5323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Buescher AVS, Cidav Z, Knapp M, Mandell DS. Costs of autism spectrum disorders in the United Kingdom and the United States. JAMA Pediatr. 2014;168:721–728. doi: 10.1001/jamapediatrics.2014.210. [DOI] [PubMed] [Google Scholar]
- 23.Centers for Disease Control and Prevention (CDC) Economic costs associated with mental retardation, cerebral palsy, hearing loss, and vision impairment—United States, 2003. MMWR Morb Mortal Wkly Rep. 2004;53:57–59. [PubMed] [Google Scholar]
- 24.Grove J, Ripke S, Als TD, Mattheisen M, Walters RK, Won H, et al. Identification of common genetic risk variants for autism spectrum disorder. Nat Genet. 2019;51:431–444. doi: 10.1038/s41588-019-0344-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Demontis D, Walters RK, Martin J, Mattheisen M, Als TD, Agerbo E, et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat Genet. 2019;51:63–75. doi: 10.1038/s41588-018-0269-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tatton-Brown K, Loveday C, Yost S, Clarke M, Ramsay E, Zachariou A, et al. Mutations in epigenetic regulation genes are a major cause of overgrowth with intellectual disability. Am J Hum Genet. 2017;100:725–736. doi: 10.1016/j.ajhg.2017.03.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Stessman HAF, Willemsen MH, Fenckova M, Penn O, Hoischen A, Xiong B, et al. Disruption of POGZ is associated with intellectual disability and autism spectrum disorders. Am J Hum Genet. 2016;98:541–552. doi: 10.1016/j.ajhg.2016.02.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Andres EM, Hafeez H, Yousaf A, Riazuddin S, Rice ML, Basra MAR, et al. A genome-wide analysis in consanguineous families reveals new chromosomal loci in specific language impairment (SLI) Eur J Hum Genet. 2019;27:1274–1285. doi: 10.1038/s41431-019-0398-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mehta CM, Gruen JR, Zhang H. A method for integrating neuroimaging into genetic models of learning performance. Genet Epidemiol. 2017;41:4–17. doi: 10.1002/gepi.22025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ambalavanan A, Girard SL, Ahn K, Zhou S, Dionne-Laporte A, Spiegelman D, et al. De novo variants in sporadic cases of childhood onset schizophrenia. Eur J Hum Genet. 2016;24:944–948. doi: 10.1038/ejhg.2015.218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yao N-J, Hsieh W-S, Lin C-H, Tseng C-I, Lin W-Y, Kuo P-H, et al. Interaction between prematurity and the MAOA gene on mental development in children: a longitudinal view. Front Pediatr. 2020;8:92. doi: 10.3389/fped.2020.00092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Dutt A, Shaikh M, Ganguly T, Nosarti C, Walshe M, Arranz M, et al. COMT gene polymorphism and corpus callosum morphometry in preterm born adults. Neuroimage. 2011;54:148–153. doi: 10.1016/j.neuroimage.2010.07.048. [DOI] [PubMed] [Google Scholar]
- 33.Costantine MM, Clark EAS, Lai Y, Rouse DJ, Spong CY, Mercer BM, et al. Association of polymorphisms in neuroprotection and oxidative stress genes and neurodevelopmental outcomes after preterm birth. Obstet Gynecol. 2012;120:542–550. doi: 10.1097/AOG.0b013e318265f232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Clark EAS, Mele L, Wapner RJ, Spong CY, Sorokin Y, Peaceman A, et al. Association of fetal inflammation and coagulation pathway gene polymorphisms with neurodevelopmental delay at age 2 years. Am J Obstet Gynecol. 2010;203:83.e1–83.e10. doi: 10.1016/j.ajog.2010.01.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Blair LM, Pickler RH, Anderson C. Integrative review of genetic factors influencing neurodevelopmental outcomes in preterm infants. Biol Res Nurs. 2016;18:127–137. doi: 10.1177/1099800415605379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Blair LM, Pickler RH, Gugiu PC, Ford JL, Munro CL, Anderson CM. Genetic risk factors for poor cognitive development in children with low birth weight. Biol Res Nurs. 2020;22:5–12. doi: 10.1177/1099800419869507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.O’Shea TM, Allred EN, Dammann O, Hirtz D, Kuban KCK, Paneth N, et al. The ELGAN study of the brain and related disorders in extremely low gestational age newborns. Early Hum Dev. 2009;85:719–725. doi: 10.1016/j.earlhumdev.2009.08.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Heeren T, Joseph RM, Allred EN, O’Shea TM, Leviton A, Kuban KCK. Cognitive functioning at the age of 10 years among children born extremely preterm: a latent profile approach. Pediatr Res. 2017;82:614–619. doi: 10.1038/pr.2017.82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kongsted A, Nielsen AM. Latent class analysis in health research. J Physiother. 2017;63:55–58. doi: 10.1016/j.jphys.2016.05.018. [DOI] [PubMed] [Google Scholar]
- 40.Schwarz G. Estimating the Dimension of a Model. Ann Statist. 1978;6:461–464. [Google Scholar]
- 41.Sclove SL. Application of model-selection criteria to some problems in multivariate analysis. Psychometrika. 1987;52:333–343. [Google Scholar]
- 42.Lo Y, Mendell NR, Rubin DB. Testing the number of components in a normal mixture. Biometrika. 2001;88:767–778. [Google Scholar]
- 43.Meakin CJ, Martin EM, Santos HP, Mokrova I, Kuban K, O’Shea TM, et al. Placental CpG methylation of HPA-axis genes is associated with cognitive impairment at age 10 among children born extremely preterm. Horm Behav. 2018;101:29–35. doi: 10.1016/j.yhbeh.2018.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Adén U, Lin A, Carlo W, Leviton A, Murray JC, Hallman M, et al. Candidate gene analysis: severe intraventricular hemorrhage in inborn preterm neonates. J Pediatr. 2013;163:1503–6.e1. doi: 10.1016/j.jpeds.2013.06.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chang CC, Chow CC. PLINK 1.9 [Internet]. [cited 2021 Jan 15]. Available from: http://www.cog-genomics.org/plink/1.9/
- 46.Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7. doi: 10.1186/s13742-015-0047-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Das S, Forer L, Schönherr S, Sidore C, Locke AE, Kwong A, et al. Next-generation genotype imputation service and methods. Nat Genet. 2016;48:1284–1287. doi: 10.1038/ng.3656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.TOPMed Whole Genome Sequencing Project - Freeze 5b, Phases 1 and 2 [Internet]. 2020 [cited 2021 Jan 15]. Available from: https://www.nhlbiwgs.org/topmed-whole-genome-sequencing-project-freeze-5b-phases-1-and-2
- 49.Loh P-R, Danecek P, Palamara PF, Fuchsberger C, Reshef YA, Finucane HK, et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat Genet. 2016;48:1443–1448. doi: 10.1038/ng.3679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Fuchsberger C, Abecasis GR, Hinds DA. minimac2: faster genotype imputation. Bioinformatics. 2015;31:782–784. doi: 10.1093/bioinformatics/btu704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.statgen/Minimac4 [Internet]. [cited 2021 Jan 15]. Available from: https://github.com/statgen/Minimac4
- 52.Li Y, Willer CJ, Ding J, Scheet P, Abecasis GR. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol. 2010;34:816–834. doi: 10.1002/gepi.20533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Duan Q, Liu EY, Croteau-Chonka DC, Mohlke KL, Li Y. A comprehensive SNP and indel imputability database. Bioinformatics. 2013;29:528–531. doi: 10.1093/bioinformatics/bts724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kowalski MH, Qian H, Hou Z, Rosen JD, Tapia AL, Shan Y, et al. Use of >100,000 NHLBI Trans-Omics for Precision Medicine (TOPMed) Consortium whole genome sequences improves imputation quality and detection of rare variant associations in admixed African and Hispanic/Latino populations. PLoS Genet. 2019;15:e1008500. doi: 10.1371/journal.pgen.1008500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.EPACTS - Genome Analysis Wiki [Internet]. [cited 2021 Jan 15]. Available from: https://genome.sph.umich.edu/wiki/EPACTS
- 56.Kang HM, Sul JH, Service SK. Zaitlen NA, Kong S-Y, Freimer NB, et al. Variance component model to account for sample structure in genome-wide association studies. Nat Genet. 2010;42:348–354. doi: 10.1038/ng.548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Gel B, Serra E. karyoploteR: an R/Bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics. 2017;33:3088–3090. doi: 10.1093/bioinformatics/btx346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Turner S. qqman: an R package for visualizing GWAS results using Q-Q and manhattan plots. JOSS. 2018;3:731. [Google Scholar]
- 59.Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP, et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics. 2010;26:2336–2337. doi: 10.1093/bioinformatics/btq419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.TopLD [Internet]. [cited 2021 Apr 13]. Available from: http://topld.genetics.unc.edu/topld/
- 61.Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019;47:D886–D894. doi: 10.1093/nar/gky1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Shihab HA, Rogers MF, Gough J, Mort M, Cooper DN, Day INM, et al. An integrative approach to predicting the functional effects of non-coding and coding sequence variation. Bioinformatics. 2015;31:1536–1543. doi: 10.1093/bioinformatics/btv009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Fishilevich S, Nudel R, Rappaport N, Hadar R, Plaschkes I, Iny Stein T, et al. GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database (Oxford). 2017;2017. 10.1093/database/bax028. [DOI] [PMC free article] [PubMed]
- 64.Hugin2 [Internet]. [cited 2021 Jan 16]. Available from: http://hugin2.genetics.unc.edu/Project/hugin/
- 65.Giusti-Rodriguez P, Lu L, Yang Y, Crowley CA, Liu X, Juric I, et al. Using three-dimensional regulatory chromatin interactions from adult and fetal cortex to interpret genetic results for psychiatric disorders and cognitive traits. BioRxiv. 2018. 10.1101/406330.
- 66.Fudenberg G, Imakaev M, Lu C, Goloborodko A, Abdennur N, Mirny LA. Formation of chromosomal domains by loop extrusion. Cell Rep. 2016;15:2038–2049. doi: 10.1016/j.celrep.2016.04.085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Crowley C, Yang Y, Qiu Y, Hu B, Abnousi A, Lipiński J, et al. FIREcaller: Detecting frequently interacting regions from Hi-C data. Comput Struct Biotechnol J. 2021;19:355–362. doi: 10.1016/j.csbj.2020.12.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Fromer M, Roussos P, Sieberts SK, Johnson JS, Kavanagh DH, Perumal TM, et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat Neurosci. 2016;19:1442–1453. doi: 10.1038/nn.4399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Schrider DR, Kern AD. Soft sweeps are the dominant mode of adaptation in the human genome. Mol Biol Evol. 2017;34:1863–1877. doi: 10.1093/molbev/msx154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Protty MB, Watkins NA, Colombo D, Thomas SG, Heath VL, Herbert JMJ, et al. Identification of Tspan9 as a novel platelet tetraspanin and the collagen receptor GPVI as a component of tetraspanin microdomains. Biochem J. 2009;417:391–400. doi: 10.1042/BJ20081126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.GTEx Portal [Internet]. [cited 2021 Apr 14]. Available from: https://gtexportal.org/home/
- 72.Lee J, Sayegh J, Daniel J, Clarke S, Bedford MT. PRMT8, a new membrane-bound tissue-specific member of the protein arginine methyltransferase family. J Biol Chem. 2005;280:32890–32896. doi: 10.1074/jbc.M506944200. [DOI] [PubMed] [Google Scholar]
- 73.Simandi Z, Pajer K, Karolyi K, Sieler T, Jiang L-L, Kolostyak Z, et al. Arginine methyltransferase PRMT8 provides cellular stress tolerance in aging motoneurons. J Neurosci. 2018;38:7683–7700. doi: 10.1523/JNEUROSCI.3389-17.2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Jeong H-C, Park S-J, Choi J-J, Go Y-H, Hong S-K, Kwon O-S, et al. PRMT8 controls the pluripotency and mesodermal fate of human embryonic stem cells by enhancing the PI3K/AKT/SOX2 axis. Stem Cells. 2017;35:2037–2049. doi: 10.1002/stem.2642. [DOI] [PubMed] [Google Scholar]
- 75.Lin KC, Park HW, Guan K-L. Regulation of the hippo pathway transcription factor TEAD. Trends Biochem Sci. 2017;42:862–872. doi: 10.1016/j.tibs.2017.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Jin Y, Messmer-Blust AF, Li J. The role of transcription enhancer factors in cardiovascular biology. Trends Cardiovasc Med. 2011;21:1–5. doi: 10.1016/j.tcm.2011.12.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Pobbati AV, Hong W. Emerging roles of TEAD transcription factors and its coactivators in cancers. Cancer Biol Ther. 2013;14:390–398. doi: 10.4161/cbt.23788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Yagi R, Kohn MJ, Karavanova I, Kaneko KJ, Vullhorst D, DePamphilis ML, et al. Transcription factor TEAD4 specifies the trophectoderm lineage at the beginning of mammalian development. Development. 2007;134:3827–3836. doi: 10.1242/dev.010223. [DOI] [PubMed] [Google Scholar]
- 79.Nishioka N, Yamamoto S, Kiyonari H, Sato H, Sawada A, Ota M, et al. Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech Dev. 2008;125:270–283. doi: 10.1016/j.mod.2007.11.002. [DOI] [PubMed] [Google Scholar]
- 80.Hatsuzawa K, Hirose H, Tani K, Yamamoto A, Scheller RH, Tagaya M. Syntaxin 18, a SNAP receptor that functions in the endoplasmic reticulum, intermediate compartment, and cis-Golgi vesicle trafficking. J Biol Chem. 2000;275:13713–13720. doi: 10.1074/jbc.275.18.13713. [DOI] [PubMed] [Google Scholar]
- 81.Iinuma T, Aoki T, Arasaki K, Hirose H, Yamamoto A, Samata R, et al. Role of syntaxin 18 in the organization of endoplasmic reticulum subdomains. J Cell Sci. 2009;122:1680–1690. doi: 10.1242/jcs.036103. [DOI] [PubMed] [Google Scholar]
- 82.Zhao B, Luo T, Li T, Li Y, Zhang J, Shan Y, et al. Genome-wide association analysis of 19,629 individuals identifies variants influencing regional brain volumes and refines their genetic co-architecture with cognitive and mental health traits. Nat Genet. 2019;51:1637–1644. doi: 10.1038/s41588-019-0516-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.van der Meer D, Frei O, Kaufmann T, Shadrin AA, Devor A, Smeland OB, et al. Understanding the genetic determinants of the brain with MOSTest. Nat Commun. 2020;11:3512. doi: 10.1038/s41467-020-17368-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1. SNP-heritability estimation with GCTA, Fig. S1, and Fig. S2.
Data Availability Statement
The genotype data analyzed during the current study are not publicly available but are available from the corresponding author on reasonable request. Epigenetic functional annotations, chromatin interaction, brain eQTL, and selective sweeps data are publicly available.