Abstract
Simple Summary
Lynch-like syndrome (LLS) is defined as colorectal cancer cases with microsatellite instability (MSI) and loss of expression of MLH1, MSH2, MSH6, or PMS2 by immunohistochemistry (IHC) in the absence of a germline mutation in these genes that cannot be explained by BRAF mutation or MLH1 hypermethylation. The application of the universal strategy for the diagnosis of Lynch syndrome (LS) in all CRCs is leading to an increase in the incidence of cases of LLS. It has been described that risk of cancer in relatives of LLS patients is in between of that found in Lynch syndrome families and sporadic cases. That makes LLS patients and their families a challenging group for which the origin of CRC is unknown, being a mixture between unidentified hereditary CRC and sporadic cases. The potential causes of LLS are discussed in this review, as well as methods for identification of truly hereditary cases.
Abstract
Lynch syndrome is an autosomal dominant disorder caused by germline mutations in DNA mismatch repair (MMR) system genes, such as MLH1, MSH2, MSH6, or PMS2. It is the most common hereditary colorectal cancer syndrome. Screening is regularly performed by using microsatellite instability (MSI) or immunohistochemistry for the MMR proteins in tumor samples. However, in a proportion of cases, MSI is found or MMR immunohistochemistry is impaired in the absence of a germline mutation in MMR genes, BRAF mutation, or MLH1 hypermethylation. These cases are defined as Lynch-like syndrome. Patients with Lynch-like syndrome represent a mixture of truly hereditary and sporadic cases, with a risk of colorectal cancer in first-degree relatives that is between the risk of Lynch syndrome in families and relatives of sporadic colon cancer cases. Although multiple approaches have been suggested to distinguish between hereditary and sporadic cases, a homogeneous testing protocol and consensus on the adequate classification of these patients is still lacking. For this reason, management of Lynch-like syndrome and prevention of cancer in these families is clinically challenging. This review explains the concept of Lynch-like syndrome, potential mechanisms for its development, and methods for adequately distinguishing between sporadic and hereditary cases of this entity.
Keywords: lynch syndrome, lynch-like syndrome, hereditary cancer, colorectal cancer, DNA mismatch repair genes
1. Introduction
Lynch-like syndrome (LLS) is defined as colorectal cancer cases with microsatellite instability (MSI) and loss of expression of MLH1, MSH2, MSH6, or PMS2 by immunohistochemistry (IHC) in the absence of a germline mutation in these genes that cannot be explained by BRAF mutation or MLH1 hypermethylation [1]. Managing these cases is challenging because the subsequent carcinogenic process is yet to be unveiled. LLS is probably caused by somatic mutations in the mismatch repair (MMR) genes, and, therefore, it is sporadic [2,3]. However, patients with LLS and their relatives have an increased risk of colorectal cancer (CRC), suggesting a possibility of inherited risk. Thus, the most probable scenario is that LLS represents a mixture of sporadic MSI cases, unidentified Lynch syndrome (LS) cases, and possibly other hereditary cases of yet-to-be-determined origin [1,4]. Differentiating between both sporadic and hereditary origin has been a challenge, mainly due to the difficulty in conducting mutational somatic studies of CRC samples. In this review, the characteristics of LLS cases, the potential causes, and recommendations on managing these cases are discussed.
2. Carcinogenic Pathways in Colorectal Cancer
CRC is the third most commonly diagnosed cancer after breast and lung cancer, accounting for 10% of all cancers diagnosed worldwide, or approximately 2 million cases every year [5]. Approximately 70–80% of CRC are considered to be sporadic [6].
Based on molecular characteristics, CRC can be subclassified into tumors that exhibit chromosomal instability (CIN) and tumors with MSI. CIN includes structural abnormalities or changes in the number of chromosomes [7]. Missegregation of chromosomes can lead to both the activation of oncogenes, such as KRAS, and tumor-suppressor effects, such as inactivation of APC [8,9]. Approximately 70–80% of colorectal tumors have chromosomal abnormalities associated with poor prognosis [10]. CIN causes tumor progression by increasing genomic alterations, making the tumor more aggressive and drug-resistant [11]. Most tumors with CIN are microsatellite stable (MSS) [12].
Between 20% and 30% of CRCs arise through the serrated pathway of carcinogenesis [13]. The serrated pathway includes tumors with MSS and MSI. Serrated polyps are the precursor lesion in this pathway. A majority of serrated polyps (80–90%) are benign lesions, with only a minority developing dysplasia. However, after dysplasia, carcinogenesis is accelerated and CRC develops [14]. Most CpG island methylator phenotype (CIMP) tumors are MSS or have low MSI, but if hypermethylation affects MLH1, CIMP tumors can also have high MSI. BRAF mutation is the original somatic event in this pathway, with CIMP as an early feature [13].
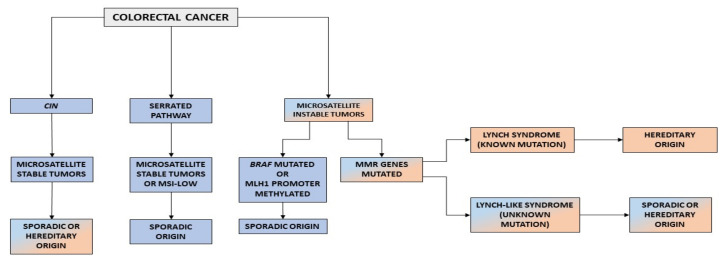
Finally, tumors with MSI represent 15% of all CRCs and are MMR-deficient (MMR-D) [15,16,17]. The role of MMR system is to correct mismatches and small insertion/deletions that occur during DNA replication. Microsatellites are repetitive sequences that are prone to the accumulation of errors [18]. When there is a deficiency in the MMR machinery, these errors cannot be corrected, resulting in microsatellites that are different in size and MSI [19]. On the one hand, the most frequent cause of MMR-D is the hypermethylation of MLH1, which occurs in 80% of tumor with MSI tumors, as a consequence of CIMP [20]. On the other hand, in a percentage of the remaining cases, MMR-D is caused by germline mutations in the MMR genes MLH1, MSH2, MSH6, and PMS2 [21]. Currently, three CRC phenotypes are related to the presence of MSI: MSI sporadic tumors, LS, and LLS (Figure 1).
Figure 1.
Carcinogenic pathways in CRC. CIN, chromosomal instability.
3. Lynch Syndrome
LS is the most common hereditary cancer syndrome and accounts for approximately 3% of all CRCs [22]. LS is an autosomal dominant disorder caused by germline mutations in MLH1, MSH2, MSH6, and PMS2, as well as deletions in EPCAM. Germline EPCAM deletions result in methylation of the surrounding genomic region, affecting the MSH2 promoter located 18 Kb downstream. As a consequence, MSH2 gene expression is silenced [23]. Constitutional epigenetic silencing of MLH1 [24,25,26,27,28,29,30,31,32,33,34,35,36] and hypermethylation of MSH2 as a consequence of EPCAM deletion [29] have been rarely reported in some families.
LS patients develop multiple tumors, most frequently colorectal and endometrial [37], but also upper gastrointestinal, ovarian, biliary, urinary, brain, non-melanoma skin, and prostate tumors [38]. LS patients are diagnosed at an early age; with a mean age of diagnosis of around 45 years, they develop cancer a mean of 23 years earlier than the general population [39]. Lynch tumors develop faster than sporadic CRC [40]. LS patients have an increased risk of synchronous and metachronous neoplasias. Approximately 7% of LS patients have multiple cancers when diagnosed [40,41]. LS tumors are poorly differentiated, and some present with mucinous features, a medullary growth pattern or showing infiltrating lymphocytes [42,43]. Moreover, their location is predominately in the proximal colon [40,44].
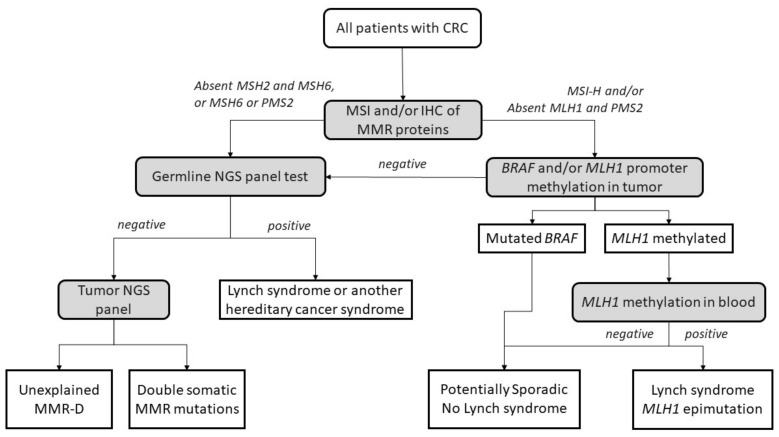
The diagnostic algorithm for LS starts by testing tumors for MSI and/or loss of immunochemical expression of MMR proteins. The Jerusalem guidelines, so-called ‘universal screening’, recommend screening all CRCs and endometrial patients <70 years old for MSI or MMR-D [45]. If MLH1 is lost in IHC, the tumor should then be tested for methylation of the promoter of MLH1 and/or the BRAF V600E mutation to rule out sporadic CIMP tumors. If testing negative, patients are submitted to germline testing, which includes the sequencing and the analysis of deletions and duplications in the appropriate MMR genes. Germline testing results confirm an LS diagnosis [46,47], whereas, if MLH1 is methylated in a tumor, a complementary MLH1 methylation study in blood should be performed to identify constitutional epimutation of MLH1 [48] (Figure 2).
Figure 2.
Universal screening strategy for Lynch-syndrome patients. CRC, colorectal cancer; MSI, microsatellite instability; IHC, immunohistochemistry; MMR, mismatch repair; MMR-D, mismatch repair deficiency; NGS, next-generation sequencing. Adapted from Valle et al. [49].
4. Lynch-like Syndrome
Up to 50% of cases of suspected LS in patients with CRC that test positive for MMR-D by IHC or MSI do not have any germline mutation in an MMR gene, BRAF alteration or MLH1 hypermethylation [1]. These cases are defined as Lynch-like syndrome (LLS). LLS possibly describes a heterogeneous group of conditions that possibly includes a mix between sporadic and hereditary cases. Although multiple approaches have been suggested to distinguish between hereditary and sporadic cases, accurate testing and a consensus on the adequate classification of these patients are still lacking.
4.1. Demographics
The risk of developing CRC and other LS-related cancers in LLS cohorts and their first-degree relatives is lower than in LS patients. However, LLS patients are more likely to have CRC than sporadic cases [1,44,50,51]. Moreover, the age at diagnosis follows the same pattern, and it is higher than in LS but lower than in sporadic cases [1,52]. LLS tumors share the pathological characteristics of MSI CRC, as they are mainly located on the proximal colon, frequently have a large size, and usually present a higher concentration of infiltrated lymphocytes [1,4,44]. On the other hand, no differences in sex have been found. Some series show a predominance of female patients, but without significant differences [1,53].
4.2. Family History
LLS patients can be clinically differentiated into two groups. One group includes patients with family history that suggests a hereditary origin. These families probably have a hereditary factor that predisposes them to a high risk of CRC. However, in these cases, the genetic alteration is unknown. Furthermore, there are other LLS families who do not have a history of cancer. In that case, it is possible to find a double somatic mutation in MMR genes that explains their MSI. These cases could be considered sporadic [4,52,54]. As noted previously, LLS cases are a heterogeneous group that includes sporadic and inherited cases, and it is necessary to define molecular tools to efficiently differentiate between both groups. Moreover, there are no differences in the clinical or pathological characteristics that differentiate between hereditary and sporadic cases [54]. The validation and implementation of molecular analysis of MMR genes in tumors as a part of routine diagnosis is still a challenge in many laboratories.
4.3. Pathology and IHC
All CRCs undergo IHC for MMR proteins MLH1, MSH2, MSH6, and PMS2 and/or MSI testing [45]. Tumors are considered MMR-D when they exhibit a loss of expression of the MMR proteins by IHC. Universal IHC or MSI testing increases the detection of LLS [53]. IHC has shown that the main defective protein in LLS tumors is MLH1 [55]. Picó et al., found that 50% of LLS patients lack MLH1/PMS2 and 27.9% lack MSH2/MSH6 expression [54]. These results are similar to previous studies that obtained the same data. Perez-Carbonell et al., identified a lack of MLH1 protein in 29 of 62 patients, followed by MSH2 and MSH6 loss (19 patients), and Overbeek et al., exposed MLH1 deficiency in 5 of 16 families [51,53]. Although one publication reported that grade 1 dysplasia was predominant in LLS, in contrast to grade 3 in LS [11,14], no other histological characteristics have been found [4,56].
4.4. Cancer Risk
First-degree relatives of patients with LLS CRC have an increased risk of CRC and non-CRC LS-related cancers. However, this risk is lower than that found in LS families. This can be seen in Rodriguez-Soler et al., who estimated a standardized incidence ratio (SIR) for CRC of 2.12 in LLS cases versus 6.04 in LS patients. In addition, Picó et al., estimated an SIR for CRC (4.25 in LS vs. 2.08 in LLS) and LS-associated neoplasms (5.01 in LS vs. 2.04 in LLS) [1,4,50]. Regarding non-CRC cancers associated with LS, they estimated an SIR of 1.69 in LLS families vs. 2.81 in LS patients [1,52]. These results are supported by Win et al., who showed that the risk of a first-degree relative of an LS patient developing CRC (hazard ratio (HR) = 5.37) is higher than in LLS (HR = 2.06) and MMR-D non-LS groups (HR = 1.04) (50). LS families also have a higher risk of developing endometrial tumors. Other studies have shown an excess risk of pancreatic cancer in LLS families [4]. Due to this increased risk of developing CRC in LLS families, if it is not possible to safely identify truly sporadic cases, LLS families should be considered a high-risk group, and some authors suggest screening colonoscopies every 3 years for first-degree relatives of LLS patients [57].
5. Potential Causes of Lynch-like Syndrome
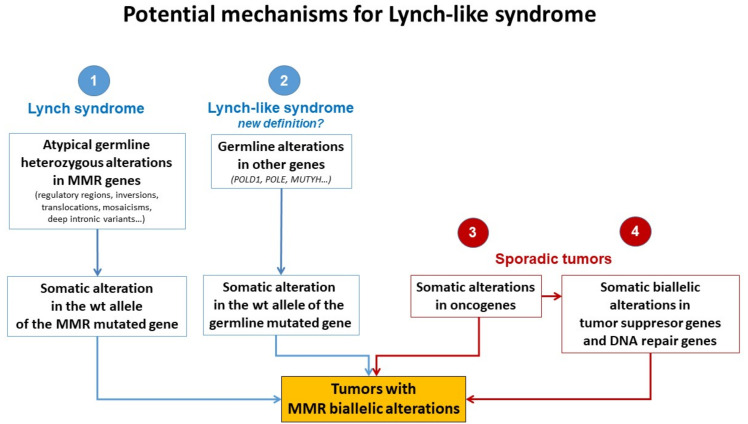
Different plausible causes to explain the origin of LLS tumors have been described. According to the hereditary origin, unknown mechanisms or germline mutations in other genes than those involved in the classical MMR system could mimic the Lynch phenotype with MMR-D. In addition, some LLS cases could be LS with unidentified germline MMR mutations. In contrast, LLS could be due to somatic defects in genes related to tumor onset and progression or due to biallelic alterations in MMR genes outside MLH1 promoter methylation [2,3,58], thus having a sporadic origin. A frequent explanation for LLS cases that should always be ruled out is false-positive IHC/MSI results, which represent approximately 19% of cases in some series [59], and confirmation of MSI and IHC status should be the first step before classifying a case as LLS. Figure 3 describes different potential causes of LLS. It is important to clarify that, if some LLS cases after their molecular analysis can be classified in another category, they will no longer be considered LLS. They will be included in the surveillance program of the new group.
Figure 3.
Potential mechanisms for Lynch-like syndrome. MMR, mismatch repair; wt, wild type. Adapted from Pico et al. [54].
5.1. Germline Mutations in Other Genes Affecting the MMR System
The fact that LLS patients are younger at diagnosis than sporadic cases and some of them have a family history of LS-related neoplasias suggests that germline mutations in other genes could also be involved in cancer development in some of these cases (Figure 3). It is important to distinguish whether MMR-D is driving tumor formation or is a secondary event. Germline mutations in MUTYH and POLE have been reported in some patients with MMR-D [31,60,61,62]. Mutations in MUTYH have been previously associated with MUTYH-attenuated polyposis [31,62]. In addition, mutations in MUTYH have been described in MLH1-methylated tumors [31,62]. Approximately 1–3% of LLS cases carry biallelic mutations in MUTYH [31,62]. In addition, mutations in the exonuclease domain of POLE and POLD1 cause a hypermutator phenotype that confers a high predisposition to developing attenuated colorectal polyposis at an early age. POLE and POLD1 mutations may be associated with MMR-D in some cases due to MMR mutations secondary to the hypermutator phenotype [60,61,63,64,65].
On the other hand, Xavier et al., found potentially pathogenic variants in a group of genes involved in the regulation of cellular activity (EXO1, POLD1, RFC1, and RPA1) [55]. EXO1 is related to the union of MLH1 and MSH2, and a mutation in these genes may trigger MMR-D [66]. In addition, RPA1 and POLD1 are associated with harmful effects in tumors with mutations in these genes [67,68]. RFC1 has been described in the development of different malignancies. Huang et al., noted the presence of a variation of this gene in a plasmatic cell tumor [69]. Moreover, somatic mutations in RFC1 were reported in 10.2% of uterine carcinomas and 5.5% of CRCs [70]. This gene also plays an important role in genomic integrity because it is a member of the BRCA1-associated genomic surveillance complex [71]. Golubicki et al., found unknown variants in four genes (POLE, ERCC6, RAD54L, and PALB2) in a group of LLS patients [72]. ERCC6 and PALB2 have been associated with CRC [61,73,74], and the PALB2 variant was previously reported in a suspected case of LS [75].
Next-generation sequencing (NGS) studies have allowed the identification of pathogenic variants that could be candidates for familial CRC with unknown genetic basis. Recently published studies have identified pathogenic variants in genes that maintain DNA integrity resulting in a variety of clinical phenotypes. Germline variants in NTHL1 cause adenomatous polyposis and CRC [76]; MCM9 variants are associated with hereditary mixed polyposis, CRC, and primary ovarian failure [65,77]; and variants in FAN1 cause hereditary CRC by impairing DNA repair [78]. Following this line of inquiry, variants in BUB1 and BUB3 [79], SETD2 [80], WRN [81], BARD1 [81], MCPH1 [81], and REV3L [81] have been found in the germline analysis of LLS cases, linking the mutation of WRN, BARD1, MCPH1, and REV3L for the first time with CRC.
5.2. Hereditary Cases: Unknown Mutations in MMR Genes
In some cases, LLS patients are actually LS patients whose pathogenic variants have not been identified (Figure 3). Current techniques cannot easily identify complex and cryptic mutations. Intronic regions, structural changes such as inversions, and/or copy number variation (CNV) are rarely analyzed genetic changes but may play an important role in unveiling mutations in these patients. For example, the mutation 478 bp upstream of exon 2 in MSH2 creates a canonical splice donor site. The pseudo-exon that is created contains a stop codon that results in a truncated protein [65,82].
Structural changes have been found in some families, such as the inversion of MSH2 exons 1–7 in 10 families in North America [83,84,85] and the inversion of MSH2 exons 2–6 in two families in Australia [86]. Another example of structural genetic changes is the MLH1-LRRFIP2 fusion after a paracentric inversion of chromosome 3 [87] or deletion in that same locus [65,88]. Moreover, Hellen et al., show a retrotranspositional insertion in PMS2 mediated by LINE-1 between exon 7 and 8 [89].
Regulatory regions of MMR genes should also be taken into account. In some cases, variants in the promoter region of MMR have been associated with reduced promoter activity or transcriptional silencing of the allele [80,90]. The accumulation of mutations in the 3′UTR of genes affects mRNA stability and, therefore, protein expression. Germline 3′ UTR mutations in MLH1 have been associated with loss of expression [91]. On the other hand, abnormal regulation of protein expression by miRNA could cause a loss of MMR gene expression. High levels of miRNA-21 downregulate MSH2 and MSH6 and have been found in CRC with loss of MSH2 expression [92]. The same has been seen with MLH1 and miRNA-155 [92]. These examples show the importance of more extensive sequencing methods to detect complex mutations in families of patients with MMR-D and without germline mutations by routine procedures.
Somatic mosaicism could also account for some LLS cases. For instance, Sourrouille et al., described a case of somatic mosaicism in MSH2 after de novo mutation of this gene [58]. Another study described somatic mosaicism in a woman with synchronous gynecological tumors at 44 years old. The MLH1 mutation was only present in 20% of the allele fraction in normal tissue, but her sister and father, who were also affected with LS-related tumors, carried the same mutation [93]. A recent study reported a case of de novo somatic mosaicism in which the MLH1 mutation was detected in the tumor and at a lower level in peripheral blood but not in any other family member [94]. Mosaicism can be detected using highly sensitive NGS with high coverage, and more genetic-driven cases could be correctly identified.
Another important factor to consider is the presence of variants of uncertain significance (VUS) in approximately 30% of cases [95]. Some of them could be pathogenic but cannot be classified due to the absence of clinical, molecular, or functional evidence. Families carrying VUS are managed based on their family history of cancer until further variant classification is available [65].
5.3. Somatic Alteration in Other Cancer Genes or Epigenetic Structures
Some patients with sporadic cancers do not exhibit any alteration in MMR genes but lack these proteins. In these cases, other molecular mechanisms could be leading to MMR-D and a MSI phenotype. These tumors could be due to somatic alterations in cancer genes or epigenetic events outside of the MMR system [96].
Li et al., found a relationship between the epigenetic histone marker H3K36me3 and the MMR system. H3K36me3 recruits the mismatch recognition protein hMutSα (MSH2-MSH6) onto chromatin interacting with MSH6. This protein and the histone methyltransferase SETD2, which also acts in the trimethylation of H3K36, are required for activation of the MMR system [97]. Li et al., demonstrated that depletion of SETD2 and/or H3K36me3 in cells resulted in an MMR-D mutator phenotype, providing a molecular explanation of tumors that are positive for MSI with the MMR-D phenotype [98]. Another important molecule is proliferation cell nuclear antigen (PCNA), which plays an important role during DNA replication. Ortega et al., showed that the phosphorylation of PCNA by epidermal growth factor receptor (EGFR) alters its interaction with MMR proteins, revealing another possible mechanism of cancer development via suppression of MMR function [99].
AT-rich interaction domain 1A (ARID1A) is mutated in a large proportion of tumors [100]. These proteins interact with MSH2, recruiting it to chromatin during DNA replication. Shen et al., demonstrated that impairment of ARID1A contributes to MMR-D [101]. In addition, a somatic exonuclease domain mutation in POLE would be involved in phenocopying defective MMR DNA in 25% of unexplained endometrial cancers with MSI [102].
Local inflammation also promotes genetic and epigenetic alterations in CRC [103] and has been determined to be an important factor in damage to the MMR system [104]. An increase in the concentration of proinflammatory cytokine IL-6 has been demonstrated to alter MMR function. IL6 can activate STAT3, which drives MSH3 out of the nucleus and prevents it from performing its nuclear function [105]. In the same way, high levels of reactive oxygen species (ROS) can induce DNA damage, resulting in MMR-D. Chang et al., showed that non-cytotoxic H2O2 can damage MMR complexes, triggering a reduction in these proteins [106].
Somatic methylation could also explain the MMR-D present in some LLS cases (Figure 3). Many tumor suppressor genes are methylated in sporadic cancers, including RB [107,108], VHL [109], and BRCA1 [110], as well as MLH1 promoter hypermethylation in sporadic CRC caused by the CIMP phenotype [111]. Recently, Buckley et al., reported an association between the methylation of SHPRH and MSI burden [112]. In addition, epimutations in MLH1 and MSH2 have been reported in some families [24,25,26,27,28,29,30,32,33,34,35,36], but other MMR genes can also be targets of somatic methylation [65].
5.4. Somatic Biallelic Alteration in MMR
Recent studies have shown that somatic mutations in MMR genes are responsible for a proportion of LLS cases [2,3,58,113,114]. In one study, 17 CRC cases with MSI were screened for somatic mutations, resulting in one out of seven MLH1-D tumors with two somatic mutations in MLH1 and three out of eight MSH2-D tumors with two somatic mutations in MSH2 [58]. Moreover, the combination of somatic mutation and loss of heterozygosity (LOH) as a second hit was studied in 25 tumors, finding two somatic hits in 13 tumors (8/18 in MLH1 and 5/7 in MSH2) [2]. Porka et al., identified two somatic events in MMR genes in 11 out of 14 tumors and a somatic mutation and LOH in 10 out of 11 (4/10 in MLH1, 5/10 MSH2, and 1/10 PMS2), whereas only one tumor was characterized as having two somatic mutations (MSH6) [56]. Lefol et al., also looked for somatic variants, LOH, and single events in 85% [97/113] of LLS tumors (85%), but double somatic hits were found in only 63% (72/113) [114]. Another study looked for CNV in addition to mutation and LOH in 40 LLS cases; 16/24 carried double somatic hits in MLH1 and 5/12 in MSH2 [3]. Vargas-Parra et al., obtained the same results; they found a double somatic hit in MSH2 and MSH6 in five tumors from four LLS patients (80). All of these results agree with those obtained by Xicola et al., who showed a somatic mutation in one MMR allele and LOH in the other allele in 4/9 LLS cases (3/4 in MLH1 and 1/4 in MSH2) [81]. It seems that the most common double somatic hit is a somatic mutation combined with LOH, followed by two somatic mutations. Based on these studies, the number of cases explained by somatic inactivation could be approximately 50% (32–82%) of LLS tumors [2,3,58]. Nevertheless, the percentage of tumors with a double somatic hit could be underestimated because some tumors also exhibit VUS or uninformative LOH, and the causative effect on the MSI phenotype is uncertain. That point is reinforced with the study of Elze et al., where they show a proportion of somatic deficient MMR tumors with somatic exon deletion that is not detectable by sequencing [115].
When comparing tumors with double somatic alterations to LS tumors, no significant histopathological difference was found [116]. Tumor sequencing is the adequate way to evaluate double somatic mutations. Therefore, tumor sequencing should be considered to clarify sporadic versus hereditary causes of unexplained MMR-D [65,117].
Different studies investigated promoter methylation of MMR genes in LLS patients. Methylation of MSH2 was only found in one [117,118] out of 53 LLS patients with loss of expression of MSH2 by IHC studied [80,117,118]. The MSH6 promoter was unmethylated in 108 patients with LLS and MMR-D [80,119,120], and the same happened with the PMS2 promoter in 100 cases with loss of expression of PMS2 or MLH1 who were negative for PMS2 promoter methylation [121]. In summary, based on these studies, somatic promoter hypermethylation of MMR genes does not seem to be the underlying cause of MMR-D in these unexplained tumors.
Therefore, there is a subgroup of LLS that can be explained by double somatic inactivation, and these cases should probably be excluded from the LLS classification due to the probable sporadic origin. However, this approach still has some open questions, because there is no standardized universally accepted technique or protocol for differentiating these cases. Moreover, the biallelic somatic inactivation of MMR genes can also be due to any of the previously described mechanisms, some of them generated by germline genetic alterations. Classifying patients as sporadic or potentially hereditary cases should also be the subject of clinical validation by adequately comparing pedigrees, with long-term follow-up of these families in order to find differences in the incidence of CRC and other LS-related disorders. When some groups advocate for generalization of a somatic study of LLS cases, it is necessary to reach a consensus on how to perform such a study and which cases could be confidently considered as sporadic with no indication for follow-up of patients and relatives. This algorithm has not been clinically validated. Table 1 show a summary of potential causes of LLS.
Table 1.
Potential causes of LLS. LLS, Lynch-like syndrome; MMR, mismatch repair.
| Mutations in other Genes Affecting MMR System (Germline) |
Unknown Mutations in MMR Genes (Germline) |
Somatic Mutations in Cancer Genes (Somatic) |
Biallelic Alteration in MMR (Somatic) |
|---|---|---|---|
| MUTYH | Mutation EXON 2 MSH2 | H3K36me3 | Double somatic hit |
| POLE/POLD1 | Inversion EXON 1-7 MSH2 | SETD2 | Somatic mutations in MMR genes |
| EXO1/RFC1/RPA1 | Inversion EXON 2-6 MSH2 | PCNA | Methylation in MMR genes |
| ERCC6/RAD54L/PALB2 | MLH1-LRRFIP2 fusion | ARID1A | |
| PIK3CA | MLH1 3′ UTR mutation | POLE | |
| FAN1/MCM9 | Deep intronic variant in MSH2 | IL-6 and oxidative stress | |
| NTHL1 | miRNA 21 AND miRNA 155 | Methylation in other genes | |
| BUB1/BUB3/WRN/MCPH1/REV3L | Mosaicism | ||
| VUS |
6. Future Research
Although approximately 50–60% of LLS cases can be explained by double somatic inactivation of MMR genes [2,3,114], this does not exclude the possibility that patients carry germline variants of susceptibility that increase the risk of developing cancer. Although somatic mutational testing of tumor samples could be implemented as usual practice in select centers, this practice is still used very little in regular clinical practice. Therefore, it is important to establish firstly accurate protocols and guidelines for the study of somatic mutations in tumors and then determine whether any germline variants could trigger carcinogenesis. These discoveries could be incorporated into clinical multigene panels to improve the diagnostic algorithm for these cases.
Implementing the use of NGS technologies in the diagnostic routine will allow for the characterization of the molecular profile of patients with LLS [122]. However, a collateral effect could be finding a number of VUS with uncertain pathogenicity. This will be important to increase efforts for adequate classification of the increased number of VUS, allowing personalized diagnosis, treatment, and surveillance for LLS families, even when new alterations previously unrelated to CCR are detected. All of these advances will cost-effectively decrease mortality. Exome-based studies looking for new genes involved in LLS are ongoing in several research groups [72,123]. Approaches based on the application of simultaneous somatic and germline studies in these patients can also help differentiate between hereditary and sporadic cases. An analysis of up-front somatic testing and its applicability in clinical practice and ability to differentiate germline and somatic mutations is another interesting line for future research [124].
7. Conclusions
Although some recent studies suggest that LLS cases can be explained by somatic mutations [2,3,114], the fact that relatives of LLS patients have an increased risk of CRC implies that there may be an inherited risk factor [4]. Therefore, LLS is probably heterogeneous and composed of both hereditary and sporadic cases. Differentiating these cases is a current challenge in the management of high-risk conditions. The first step in the study of potential causes of LLS must be the confirmation of the diagnosis, including MSI and lack of expression of MMR proteins by IHC, without BRAF mutation or MLH1 hypermethylation. There are then four potential explanations for the observed MSI and MMR-D phenotype in LLS: germline alterations in other genes; atypical germline alterations in MMR genes; somatic alterations in other genes that trigger a cascade, affecting MMR expression; and somatic biallelic inactivation of MMR genes. With recent advances in and availability of NGS, it is possible to improve somatic and germline testing and possibly distinguish which force is driving carcinogenesis in these cases. Clinical validation of these findings is also needed to improve the algorithms for adequate and safe management and genetic counselling of patients.
Author Contributions
A.M.-R. and M.G.-C. contributed equally to this work. Conceptualization, A.M.-R., M.G.-C. and R.J.; investigation, A.M.-R., M.G.-C., A.G.-H., O.M., J.L.S., A.C. and R.J.; writing—original draft preparation, A.M.-R., M.G.-C. and R.J.; writing—review and editing, A.M.-R., M.G.-C., A.G.-H., O.M., J.L.S., A.C. and R.J; supervision, J.L.S., A.C. and R.J.; funding acquisition, R.J. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Instituto de Salud Carlos III (PI17/01756 and PI20/01527). Asociación Española de Gastroenterología Grants (Tamarite 2015 “Caracterización molecular del síndrome de Lynch-Like”). Fundación Instituto de Investigación Sanitaria y Biomédica de Alicante ISABIAL (UGP-21-172—UGP-20-207 and UGP-21-104). Alejandro Martínez-Roca and Oscar Murcia received PFIS grant from the Instituto de Salud Carlos III (FI18/00301-CM18/00058). Anabel García-Heredia received a Sara Borrell grant from the Instituto de Salud Carlos III (CD19/00133). Asociación para la Investigación en Gastroenterología de la Provincia de Alicante (AIGPA), a private association that promotes research in gastrointestinal diseases in Alicante, also supported the logistical aspects of the study, but declares no conflict of interest.
Conflicts of Interest
The authors declare no conflict of interest.
Footnotes
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Rodríguez–Soler M., Pérez–Carbonell L., Guarinos C., Zapater P., Castillejo A., Barberá V.M., Juárez M., Bessa X., Xicola R.M., Clofent J., et al. Risk of cancer in cases of suspected lynch syndrome without germline mutation. Gastroenterology. 2013;144:926–932. doi: 10.1053/j.gastro.2013.01.044. [DOI] [PubMed] [Google Scholar]
- 2.Mensenkamp A.R., Vogelaar I.P., van Zelst–Stams W.A., Goossens M., Ouchene H., Hendriks–Cornelissen S.J., Kwint M.P., Hoogerbrugge N., Nagtegaal I.D., Ligtenberg M.J. Somatic mutations in MLH1 and MSH2 are a frequent cause of mismatch-repair deficiency in lynch syndrome-like tumors. Gastroenterology. 2014;146:643–646. doi: 10.1053/j.gastro.2013.12.002. [DOI] [PubMed] [Google Scholar]
- 3.Geurts-Giele W.R.R., Leenen C.H.M., Dubbink H.J., Meijssen I.C., Post E., Sleddens H.F.B.M., Kuipers E.J., Goverde A., Ouweland A.M.W.V.D., Van Lier M.G.F., et al. Somatic aberrations of mismatch repair genes as a cause of microsatellite-unstable cancers. J. Pathol. 2014;234:548–559. doi: 10.1002/path.4419. [DOI] [PubMed] [Google Scholar]
- 4.Picó M.D., Sánchez-Heras A.B., Castillejo A., Giner-Calabuig M., Alustiza M., Sánchez A., Moreira L., Pellise M., Castells A., Llort G., et al. Risk of Cancer in Family Members of Patients with Lynch-Like Syndrome. Cancers. 2020;12:2225. doi: 10.3390/cancers12082225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.World Health Organisation Globocan. [(accessed on 23 January 2022)];Int. Agency Res. 2020 419:3–4. Available online: https://ascopost.com/news/december-2020/globocan-2020-database-provides-latest-global-data-on-cancer-burden-cancer-deaths. [Google Scholar]
- 6.Valle L. Genetic predisposition to colorectal cancer: Where we stand and future perspectives. World J. Gastroenterol. 2014;20:9828–9849. doi: 10.3748/wjg.v20.i29.9828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sieber O.M., Heinimann K., Tomlinson I.P.M. Genomic Instability—The Engine of Tumorigenesis? Nat. Rev. Cancer. 2003;3:701–708. doi: 10.1038/nrc1170. [DOI] [PubMed] [Google Scholar]
- 8.Grady W.M., Carethers J.M. Genomic and Epigenetic Instability in Colorectal Cancer Pathogenesis. Gastroenterology. 2008;135:1079–1099. doi: 10.1053/j.gastro.2008.07.076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yuen K.W., Desai A. The wages of CIN. J. Cell Biol. 2008;180:661–663. doi: 10.1083/jcb.200801030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Popat S., Hubner R., Houlston R.S. Systematic Review of Microsatellite Instability and Colorectal Cancer Prognosis. J. Clin. Oncol. 2005;23:609–618. doi: 10.1200/JCO.2005.01.086. [DOI] [PubMed] [Google Scholar]
- 11.Thompson S.L., Bakhoum S.F., Compton D.A. Mechanisms of Chromosomal Instability. Curr. Biol. 2010;20:R285–R295. doi: 10.1016/j.cub.2010.01.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yamamoto H., Gil J., Schwartz S., Perucho M. Frameshift mutations in Fas, Apaf-1, and Bcl-10 in gastro-intestinal cancer of the microsatellite mutator phenotype. Cell Death Differ. 2000;7:238–239. doi: 10.1038/sj.cdd.4400651. [DOI] [PubMed] [Google Scholar]
- 13.Bettington M., Walker N., Clouston A., Brown I., Leggett B., Whitehall V. The serrated pathway to colorectal carcinoma: Current concepts and challenges. Histopathology. 2013;62:367–386. doi: 10.1111/his.12055. [DOI] [PubMed] [Google Scholar]
- 14.Togo G., Shiratori Y., Okamoto M., Yamaji Y., Matsumura M., Sano T., Motojima T., Omata M. Relationship between grade of microsatellite instability and target genes of mismatch repair pathways in sporadic colorectal carcinoma. Am. J. Dig. Dis. 2001;46:1615–1622. doi: 10.1023/a:1010628831884. [DOI] [PubMed] [Google Scholar]
- 15.Boland C.R., Goel A. Microsatellite Instability in Colorectal Cancer. Gastroenterology. 2010;138:2073–2087. doi: 10.1053/j.gastro.2009.12.064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Vilar E., Gruber S.B. Microsatellite instability in colorectal cancer—The stable evidence. Nat. Rev. Clin. Oncol. 2010;7:153–162. doi: 10.1038/nrclinonc.2009.237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Imai K., Yamamoto H. Carcinogenesis and microsatellite instability: The interrelationship between genetics and epigenetics. Carcinogenesis. 2008;29:673–680. doi: 10.1093/carcin/bgm228. [DOI] [PubMed] [Google Scholar]
- 18.Wu M.-S., Lee C.W., Shun C.-T., Wang H.P., Lee W.J., Sheu J.-C., Lin J.T. Clinicopathological significance of altered loci of replication error and microsatellite instability-associated mutations in gastric cancer. Cancer Res. 1998;58:1494–1497. [PubMed] [Google Scholar]
- 19.Xicola R.M., Llor X., Pons E., Castells A., Alenda C., Piñol V., Andreu M., Castellvi-Bel S., Payá A., Jover R., et al. Performance of Different Microsatellite Marker Panels for Detection of Mismatch Repair–Deficient Colorectal Tumors. JNCI J. Natl. Cancer Inst. 2007;99:244–252. doi: 10.1093/jnci/djk033. [DOI] [PubMed] [Google Scholar]
- 20.Young J., Simms L.A., Biden K.G., Wynter C., Whitehall V., Karamatic R., George J., Goldblatt J., Walpole I., Robin S.-A., et al. Features of Colorectal Cancers with High-Level Microsatellite Instability Occurring in Familial and Sporadic Settings: Parallel Pathways of Tumorigenesis. Am. J. Pathol. 2001;159:2107–2116. doi: 10.1016/S0002-9440(10)63062-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Inamura K. Colorectal Cancers: An Update on Their Molecular Pathology. Cancers. 2018;10:26. doi: 10.3390/cancers10010026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lynch H.T., De La Chapelle A. Genetic susceptibility to non-polyposis colorectal cancer. J. Med. Genet. 1999;36:801–818. [PMC free article] [PubMed] [Google Scholar]
- 23.De La Chapelle A. Genetic predisposition to colorectal cancer. Nat. Rev. Cancer. 2004;4:769–780. doi: 10.1038/nrc1453. [DOI] [PubMed] [Google Scholar]
- 24.Gazzoli I., Loda M., Garber J., Syngal S., Kolodner R.D. A hereditary nonpolyposis colorectal carcinoma case associated with hypermethylation of the MLH1 gene in normal tissue and loss of heterozygosity of the unmethylated allele in the resulting microsatellite instability-high tumor. Cancer Res. 2002;62:3925–3928. [PubMed] [Google Scholar]
- 25.Hitchins M., Williams R., Cheong K., Halani N., Lin V.A., Packham D., Ku S., Buckle A., Hawkins N., Burn J., et al. MLH1 Germline Epimutations as a Factor in Hereditary Nonpolyposis Colorectal Cancer. Gastroenterology. 2005;129:1392–1399. doi: 10.1053/j.gastro.2005.09.003. [DOI] [PubMed] [Google Scholar]
- 26.Hitchins M.P., Wong J.J., Suthers G., Suter C.M., Martin D.I., Hawkins N.J., Ward R.L. Inheritance of a Cancer-Associated MLH1 Germ-Line Epimutation. N. Engl. J. Med. 2007;356:697–705. doi: 10.1056/NEJMoa064522. [DOI] [PubMed] [Google Scholar]
- 27.Morak M., Schackert H.K., Rahner N., Betz B., Ebert M., Walldorf C., Royer-Pokora B., Schulmann K., von Knebel-Doeberitz M., Dietmaier W., et al. Further evidence for heritability of an epimutation in one of 12 cases with MLH1 promoter methylation in blood cells clinically displaying HNPCC. Eur. J. Hum. Genet. 2008;16:804–811. doi: 10.1038/ejhg.2008.25. [DOI] [PubMed] [Google Scholar]
- 28.Hitchins M.P., Ward R. Constitutional (germline) MLH1 epimutation as an aetiological mechanism for hereditary non-polyposis colorectal cancer. J. Med. Genet. 2009;46:793–802. doi: 10.1136/jmg.2009.068122. [DOI] [PubMed] [Google Scholar]
- 29.Chan T.L., Yuen S.T., Kong C.K., Chan Y.W., Chan A.S., Ng W.F., Tsui W.Y., Lo M.W., Tam W.Y., Li V.S., et al. Heritable germline epimutation of MSH2 in a family with hereditary nonpolyposis colorectal cancer. Nat. Genet. 2006;38:1178–1183. doi: 10.1038/ng1866. [DOI] [PubMed] [Google Scholar]
- 30.Valle L., Carbonell P., Fernández V., Dotor A., Sanz M., Benítez J., Urioste M. MLH1 germline epimutations in selected patients with early-onset non-polyposis colorectal cancer. Clin. Genet. 2007;71:232–237. doi: 10.1111/j.1399-0004.2007.00751.x. [DOI] [PubMed] [Google Scholar]
- 31.Castillejo A., Vargas-Parra G., Castillejo M.I., Navarro M., Barbera V.-M., González S., Hernández-Illán E., Brunet J., Cajal T.R.Y., Balmaña J., et al. Prevalence of germline MUTYH mutations among Lynch-like syndrome patients. Eur. J. Cancer. 2014;50:2241–2250. doi: 10.1016/j.ejca.2014.05.022. [DOI] [PubMed] [Google Scholar]
- 32.Cini G., Carnevali I., Quaia M., Chiaravalli A.M., Sala P., Giacomini E., Maestro R., Tibiletti M.G., Viel A. Concomitant mutation and epimutation of the MLH1 gene in a Lynch syndrome family. Carcinogenesis. 2015;36:452–458. doi: 10.1093/carcin/bgv015. [DOI] [PubMed] [Google Scholar]
- 33.Kidambi T.D., Blanco A., Van Ziffle J., Terdiman J.P. Constitutional MLH1 methylation presenting with colonic polyposis syndrome and not Lynch syndrome. Fam. Cancer. 2016;15:275–280. doi: 10.1007/s10689-016-9868-6. [DOI] [PubMed] [Google Scholar]
- 34.Hitchins M.P. Finding the needle in a haystack: Identification of cases of Lynch syndrome with MLH1 epimutation. Fam. Cancer. 2016;15:413–422. doi: 10.1007/s10689-016-9887-3. [DOI] [PubMed] [Google Scholar]
- 35.Crépin M., Dieu M.-C., Lejeune S., Escande F., Boidin D., Porchet N., Morin G., Manouvrier S., Mathieu M., Buisine M.-P. Evidence of constitutional MLH1 epimutation associated to transgenerational inheritance of cancer susceptibility. Hum. Mutat. 2011;33:180–188. doi: 10.1002/humu.21617. [DOI] [PubMed] [Google Scholar]
- 36.Castillejo A., Hernández-Illán E., Rodriguez-Soler M., Pérez-Carbonell L., Egoavil C., Barberá V.M., Castillejo M.-I., Guarinos C., Martínez-De-Dueñas E., Juan M.-J., et al. Prevalence ofMLH1constitutional epimutations as a cause of Lynch syndrome in unselected versus selected consecutive series of patients with colorectal cancer. J. Med. Genet. 2015;52:498–502. doi: 10.1136/jmedgenet-2015-103076. [DOI] [PubMed] [Google Scholar]
- 37.Vasen A.H.F., Blanco I., Aktan-Collan K., Gopie J.P., Alonso A., Aretz S., Bernstein I., Bertario L., Burn J., Capella G., et al. Revised guidelines for the clinical management of Lynch syndrome (HNPCC): Recommendations by a group of European experts. Gut. 2013;62:812–823. doi: 10.1136/gutjnl-2012-304356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lynch H.T., Boland C.R., Rodriguez-Bigas M.A., Amos C., Lynch J.P., Lynch P.M. Who should be sent for genetic testing in hereditary colorectal cancer syndromes? J. Clin. Oncol. 2007;25:3534–3542. doi: 10.1200/JCO.2006.10.3119. [DOI] [PubMed] [Google Scholar]
- 39.Hampel H., Frankel W.L., Martin E., Arnold M., Khanduja K., Kuebler P., Nakagawa H., Sotamaa K., Prior T.W., Westman J., et al. Screening for the Lynch Syndrome (Hereditary Nonpolyposis Colorectal Cancer) N. Engl. J. Med. 2005;352:1851–1860. doi: 10.1056/NEJMoa043146. [DOI] [PubMed] [Google Scholar]
- 40.Lynch H.T., Snyder C.L., Shaw T.G., Heinen C.D., Hitchins M.P. Milestones of Lynch syndrome: 1895–2015. Nat. Cancer. 2015;15:181–194. doi: 10.1038/nrc3878. [DOI] [PubMed] [Google Scholar]
- 41.Risio M., Reato G., Di Celle P.F., Fizzotti M., Rossini F.P., Foà R. Microsatellite instability is associated with the histological features of the tumor in nonfamilial colorectal cancer. Cancer Res. 1996;56:5470–5474. [PubMed] [Google Scholar]
- 42.Willett C.G., Chang D.T., Czito B.G., Meyer J., Wo J. Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–337. doi: 10.1038/nature11252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Schwitalle Y., Kloor M., Eiermann S., Linnebacher M., Kienle P., Knaebel H.P., Tariverdian M., Benner A., von Knebel Doeberitz M. Immune Response Against Frameshift-Induced Neopeptides in HNPCC Patients and Healthy HNPCC Mutation Carriers. Gastroenterology. 2008;134:988–997. doi: 10.1053/j.gastro.2008.01.015. [DOI] [PubMed] [Google Scholar]
- 44.Mas-Moya J., Dudley B., Brand R.E., Thull D., Bahary N., Nikiforova M.N., Pai R.K. Clinicopathological comparison of colorectal and endometrial carcinomas in patients with Lynch-like syndrome versus patients with Lynch syndrome. Hum. Pathol. 2015;46:1616–1625. doi: 10.1016/j.humpath.2015.06.022. [DOI] [PubMed] [Google Scholar]
- 45.Boland C.R., Shike M. Report from the Jerusalem Workshop on Lynch Syndrome-Hereditary Nonpolyposis Colorectal Cancer. Gastroenterology. 2010;138:2197.e1. doi: 10.1053/j.gastro.2010.04.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Aran V., Victorino A.P., Thuler L.C., Gil Ferreira C. Colorectal Cancer: Epidemiology, Disease Mechanisms and Interventions to Reduce Onset and Mortality. Clin. Color. Cancer. 2016;15:195–203. doi: 10.1016/j.clcc.2016.02.008. [DOI] [PubMed] [Google Scholar]
- 47.Sinicrope F.A. Lynch Syndrome–Associated Colorectal Cancer. N. Engl. J. Med. 2018;379:764–773. doi: 10.1056/NEJMcp1714533. [DOI] [PubMed] [Google Scholar]
- 48.Hitchins M.P. Constitutional epimutation as a mechanism for cancer causality and heritability? Nat. Rev. Cancer. 2015;15:625–634. doi: 10.1038/nrc4001. [DOI] [PubMed] [Google Scholar]
- 49.Valle L., Vilar E., Tavtigian S.V., Stoffel E.M. Genetic predisposition to colorectal cancer: Syndromes, genes, classification of genetic variants and implications for precision medicine. J. Pathol. 2018;247:574–588. doi: 10.1002/path.5229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Win A., Buchanan D., Rosty C., MacInnis R., Dowty J., Dite G., Giles G., Southey M.C., Young J., Clendenning M., et al. Role of tumour molecular and pathology features to estimate colorectal cancer risk for first-degree relatives. Gut. 2014;64:101–110. doi: 10.1136/gutjnl-2013-306567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Overbeek L.I.H., Kets C.M., Hebeda K.M., Bodmer D., Van Der Looij E., Willems R., Goossens M., Arts N., Brunner H.G., Van Krieken J.H.J.M., et al. Patients with an unexplained microsatellite instable tumour have a low risk of familial cancer. Br. J. Cancer. 2007;96:1605–1612. doi: 10.1038/sj.bjc.6603754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Carethers J.M. Differentiating Lynch-Like From Lynch Syndrome. Gastroenterology. 2014;146:602–604. doi: 10.1053/j.gastro.2014.01.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Pérez-Carbonell L., Ruiz-Ponte C., Guarinos C., Alenda C., Payá A., Brea-Fernández A., Egoavil C., Castillejo A., Barbera V.-M., Bessa X., et al. Comparison between universal molecular screening for Lynch syndrome and revised Bethesda guidelines in a large population-based cohort of patients with colorectal cancer. Gut. 2011;61:865–872. doi: 10.1136/gutjnl-2011-300041. [DOI] [PubMed] [Google Scholar]
- 54.PPicó M.D., Castillejo A., Murcia O., Giner-Calabuig M., Alustiza M., Sánchez A., Moreira L., Pellise M., Castells A., Carrillo-Palau M., et al. Clinical and Pathological Characterization of Lynch-Like Syndrome. Clin. Gastroenterol. Hepatol. 2019;18:368–374.e1. doi: 10.1016/j.cgh.2019.06.012. [DOI] [PubMed] [Google Scholar]
- 55.Xavier A., Olsen M.F., Lavik L.A., Johansen J., Singh A.K., Sjursen W., Scott R.J., Talseth-Palmer B.A. Comprehensive mismatch repair gene panel identifies variants in patients with Lynch-like syndrome. Mol. Genet. Genom. Med. 2019;7:e850. doi: 10.1002/mgg3.850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Porkka N., Lahtinen L., Ahtiainen M., Böhm J.P., Kuopio T., Eldfors S., Mecklin J.P., Seppälä T.T., Peltomäki P. Epidemiological, clinical and molecular characterization of Lynch-like syndrome: A population-based study. Int. J. Cancer. 2019;145:87–98. doi: 10.1002/ijc.32085. [DOI] [PubMed] [Google Scholar]
- 57.Ladabaum U. What Is Lynch-like Syndrome and How Should We Manage It? Clin. Gastroenterol. Hepatol. 2020;18:294–296. doi: 10.1016/j.cgh.2019.08.009. [DOI] [PubMed] [Google Scholar]
- 58.Sourrouille I., Coulet F., Lefevre J.H., Colas C., Eyries M., Svrcek M., Bardier-Dupas A., Parc Y., Soubrier F. Somatic mosaicism and double somatic hits can lead to MSI colorectal tumors. Fam. Cancer. 2012;12:27–33. doi: 10.1007/s10689-012-9568-9. [DOI] [PubMed] [Google Scholar]
- 59.Haraldsdottir S., Hampel H., Tomsic J., Frankel W.L., Pearlman R., de la Chapelle A., Pritchard C.C. Colon and Endometrial Cancers With Mismatch Repair Deficiency Can Arise From Somatic, Rather Than Germline, Mutations. Gastroenterology. 2014;147:1308–1316.e1. doi: 10.1053/j.gastro.2014.08.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Palles C., Cazier J.B., Howarth K.M., Domingo E., Jones A.M., Broderick P., Kemp Z., Spain S.L., Guarino E., Salguero I., et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat. Genet. 2013;45:136–143. doi: 10.1038/ng.2503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Elsayed F.A., Kets C.M., Ruano D., Akker B.V.D., Mensenkamp A., Schrumpf M., Nielsen M., Wijnen J.T., Tops C.M., Ligtenberg M.J., et al. Germline variants in POLE are associated with early onset mismatch repair deficient colorectal cancer. Eur. J. Hum. Genet. 2014;23:1080–1084. doi: 10.1038/ejhg.2014.242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Morak M., Heidenreich B., Keller G., Hampel H., Laner A., De La Chapelle A., Holinski-Feder E. Biallelic MUTYH mutations can mimic Lynch syndrome. Eur. J. Hum. Genet. 2014;22:1334–1337. doi: 10.1038/ejhg.2014.15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Bellido F., Pineda M., Aiza G., Valdés-Mas R., Navarro M., Puente D.A., Pons T., González S., Iglesias S., Darder E., et al. POLE and POLD1 mutations in 529 kindred with familial colorectal cancer and/or polyposis: Review of reported cases and recommendations for genetic testing and surveillance. Genet. Med. 2016;18:325–332. doi: 10.1038/gim.2015.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Spier I., Holzapfel S., Altmüller J., Zhao B., Horpaopan S., Vogt S., Chen S., Morak M., Raeder S., Kayser K., et al. Frequency and phenotypic spectrum of germline mutations in POLE and seven other polymerase genes in 266 patients with colorectal adenomas and carcinomas. Int. J. Cancer. 2015;137:320–331. doi: 10.1002/ijc.29396. [DOI] [PubMed] [Google Scholar]
- 65.Giner-Calabuig M. Ph.D. Thesis. Universidad de Alicante; Alicante, Spain: Nov 30, 2020. Novel Germline and Somatic Processes in Mismatch Repair Deficient Tumors. [Google Scholar]
- 66.Keijzers G., Liu D., Rasmussen L.J. Exonuclease 1 and its versatile roles in DNA repair. Crit. Rev. Biochem. Mol. Biol. 2016;51:440–451. doi: 10.1080/10409238.2016.1215407. [DOI] [PubMed] [Google Scholar]
- 67.Nicolas E., Golemis E., Arora S. POLD1: Central mediator of DNA replication and repair, and implication in cancer and other pathologies. Gene. 2016;590:128–141. doi: 10.1016/j.gene.2016.06.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wang Y., Putnam C., Kane M.F., Zhang W., Edelmann L., Russell R., Carrión D.V., Chin L., Kucherlapati R., Kolodner R.D., et al. Mutation in Rpa1 results in defective DNA double-strand break repair, chromosomal instability and cancer in mice. Nat. Genet. 2005;37:750–755. doi: 10.1038/ng1587. [DOI] [PubMed] [Google Scholar]
- 69.Huang X., Gao Y., He J., Cai J., Ta N., Jiang H., Zhu J., Zheng J. The association between RFC1 G80A polymorphism and cancer susceptibility: Evidence from 33 studies. J. Cancer. 2016;7:144–152. doi: 10.7150/jca.13303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.The Cancer Genome Atlas Research Network. Weinstein J.N., Collisson E.A., Mills G.B., Shaw K.R.M., Ozenberger B.A., Ellrott K., Shmulevich I., Sander C., Stuart J.M. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 2013;45:1113–1120. doi: 10.1038/ng.2764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Wang Y., Cortez D., Yazdi P., Neff N., Elledge S.J., Qin J. BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures. Genes Dev. 2000;14:927–939. doi: 10.1101/gad.14.8.927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Golubicki M., Díaz-Gay M., Bonjoch L., Franch-Expósito S., Muñoz J., Cuatrecasas M., Ocaña T., Iseas S., Mendez G., Carballido M., et al. Comprehensive Genomic Characterization of Fifteen Early-Onset Lynch-Like Syndrome Colorectal Cancers. Cancers. 2021;13:1259. doi: 10.3390/cancers13061259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Yurgelun M.B., Kulke M.H., Fuchs C.S., Allen B.A., Uno H., Hornick J., Ukaegbu C.I., Brais L.K., McNamara P.G., Mayer R.J., et al. Cancer Susceptibility Gene Mutations in Individuals With Colorectal Cancer. J. Clin. Oncol. 2017;35:1086–1095. doi: 10.1200/JCO.2016.71.0012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Arora S., Yan H., Cho I., Fan H.-Y., Luo B., Gai X., Bodian D.L., Vockley J.G., Zhou Y., Handorf E.A., et al. Genetic Variants That Predispose to DNA Double-Strand Breaks in Lymphocytes From a Subset of Patients With Familial Colorectal Carcinomas. Gastroenterology. 2015;149:1872–1883.e9. doi: 10.1053/j.gastro.2015.08.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Yurgelun M.B., Allen B., Kaldate R.R., Bowles K.R., Judkins T., Kaushik P., Roa B.B., Wenstrup R.J., Hartman A.-R., Syngal S. Identification of a Variety of Mutations in Cancer Predisposition Genes in Patients With Suspected Lynch Syndrome. Gastroenterology. 2015;149:604–613.e20. doi: 10.1053/j.gastro.2015.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Weren R.D., Ligtenberg M.J., Kets C.M., De Voer R.M., Verwiel E.T., Spruijt L., van Zelst-Stams W.A., Jongmans M.C., Gilissen C., Hehir-Kwa J.Y., et al. A germline homozygous mutation in the base-excision repair gene NTHL1 causes adenomatous polyposis and colorectal cancer. Nat. Genet. 2015;47:668–671. doi: 10.1038/ng.3287. [DOI] [PubMed] [Google Scholar]
- 77.Goldberg Y., Halpern N., Hubert A., Adler S.N., Cohen S., Plesser-Duvdevani M., Pappo O., Shaag A., Meiner V. Mutated MCM9 is associated with predisposition to hereditary mixed polyposis and colorectal cancer in addition to primary ovarian failure. Cancer Genet. 2015;208:621–624. doi: 10.1016/j.cancergen.2015.10.001. [DOI] [PubMed] [Google Scholar]
- 78.Seguí N., Mina L.B., Lázaro C., Sanz-Pamplona R., Pons T., Navarro M., Bellido F., López-Doriga A., Valdés-Mas R., Pineda M., et al. Germline Mutations in FAN1 Cause Hereditary Colorectal Cancer by Impairing DNA Repair. Gastroenterology. 2015;149:563–566. doi: 10.1053/j.gastro.2015.05.056. [DOI] [PubMed] [Google Scholar]
- 79.De Voer R.M., van Kessel A.G., Weren R.D., Ligtenberg M.J., Smeets D., Fu L., Vreede L., Kamping E.J., Verwiel E.T., Hahn M.M., et al. Germline mutations in the spindle assembly checkpoint genes BUB1 and BUB3 are risk factors for colorectal cancer. Gastroenterology. 2013;145:544–547. doi: 10.1053/j.gastro.2013.06.001. [DOI] [PubMed] [Google Scholar]
- 80.Vargas-Parra G.M., González-Acosta M., Thompson B.A., Gómez C., Fernández A., Dámaso E., Pons T., Morak M., Del Valle J., Iglesias S., et al. Elucidating the molecular basis of MSH2-deficient tumors by combined germline and somatic analysis. Int. J. Cancer. 2017;141:1365–1380. doi: 10.1002/ijc.30820. [DOI] [PubMed] [Google Scholar]
- 81.Xicola R.M., Clark J.R., Carroll T., Alvikas J., Marwaha P., Regan M.R., Lopez-Giraldez F., Choi J., Emmadi R., Alagiozian-Angelova V., et al. Implication of DNA repair genes in Lynch-like syndrome. Fam. Cancer. 2019;18:331–342. doi: 10.1007/s10689-019-00128-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Clendenning M., Buchanan D., Walsh M.D., Nagler B., Rosty C., Thompson B., Spurdle A., Hopper J.L., Jenkins M., Young J. Mutation deep within an intron of MSH2 causes Lynch syndrome. Fam. Cancer. 2011;10:297–301. doi: 10.1007/s10689-011-9427-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Mork M.E., Rodriguez A., Taggart M.W., Rodriguez-Bigas M.A., Lynch P.M., Bannon S.A., You Y.N., Vilar E. Identification of MSH2 inversion of exons 1–7 in clinical evaluation of families with suspected Lynch syndrome. Fam. Cancer. 2016;16:357–361. doi: 10.1007/s10689-016-9960-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Wagner A., van der Klift H., Franken P., Wijnen J., Breukel C., Bezrookove V., Smits R., Kinarsky Y., Barrows A., Franklin B., et al. A 10-Mb paracentric inversion of chromosome arm 2p inactivatesMSH2 and is responsible for hereditary nonpolyposis colorectal cancer in a North-American kindred. Genes Chromosom. Cancer. 2002;35:49–57. doi: 10.1002/gcc.10094. [DOI] [PubMed] [Google Scholar]
- 85.Rhees J., Arnold M., Boland C.R. Inversion of exons 1–7 of the MSH2 gene is a frequent cause of unexplained Lynch syndrome in one local population. Fam. Cancer. 2013;13:219–225. doi: 10.1007/s10689-013-9688-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Liu Q., Hesson L.B., Nunez A.C., Packham D., Williams R., Ward R.L., Sloane M.A. A cryptic paracentric inversion ofMSH2exons 2–6 causes Lynch syndrome. Carcinogenesis. 2015;37:10–17. doi: 10.1093/carcin/bgv154. [DOI] [PubMed] [Google Scholar]
- 87.Morak M., Koehler U., Schackert H.K., Steinke V., Royer-Pokora B., Schulmann K., Kloor M., Höchter W., Weingart J., Keiling C., et al. Biallelic MLH1 SNP cDNA expression or constitutional promoter methylation can hide genomic rearrangements causing Lynch syndrome. J. Med. Genet. 2011;48:513–519. doi: 10.1136/jmedgenet-2011-100050. [DOI] [PubMed] [Google Scholar]
- 88.Meyer C., Brieger A., Plotz G., Weber N., Passmann S., Dingermann T., Zeuzem S., Trojan J., Marschalek R. An Interstitial Deletion at 3p21.3 Results in the Genetic Fusion of MLH1 and ITGA9 in a Lynch Syndrome Family. Clin. Cancer Res. 2009;15:762–769. doi: 10.1158/1078-0432.CCR-08-1908. [DOI] [PubMed] [Google Scholar]
- 89.Van der Klift H.M., Tops C.M., Hes F.J., Devilee P., Wijnen J.T. Insertion of an SVA element, a nonautonomous retrotransposon, inPMS2intron 7 as a novel cause of lynch syndrome. Hum. Mutat. 2012;33:1051–1055. doi: 10.1002/humu.22092. [DOI] [PubMed] [Google Scholar]
- 90.Liu Q., Thompson B., Ward R., Hesson L.B., Sloane M.A. Understanding the Pathogenicity of Noncoding Mismatch Repair Gene Promoter Variants in Lynch Syndrome. Hum. Mutat. 2016;37:417–426. doi: 10.1002/humu.22971. [DOI] [PubMed] [Google Scholar]
- 91.Wilding J.L., McGowan S., Liu Y., Bodmer W.F. Replication error deficient and proficient colorectal cancer gene expression differences caused by 3’UTR polyT sequence deletions. Proc. Natl. Acad. Sci. USA. 2010;107:21058–21063. doi: 10.1073/pnas.1015604107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Valeri N., Gasparini P., Braconi C., Paone A., Lovat F., Fabbri M., Sumani K.M., Alder H., Amadori D., Patel T., et al. MicroRNA-21 induces resistance to 5-fluorouracil by down-regulating human DNA MutS homolog 2 (hMSH2) Proc. Natl. Acad. Sci. USA. 2010;107:21098–21103. doi: 10.1073/pnas.1015541107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Pastrello C., Fornasarig M., Pin E., Berto E., Pivetta B., Viel A. Somatic mosaicism in a patient with Lynch syndrome. Am. J. Med. Genet. Part A. 2009;149A:212–215. doi: 10.1002/ajmg.a.32620. [DOI] [PubMed] [Google Scholar]
- 94.Geurts-Giele W.R., Rosenberg E.H., Van Rens A., Van Leerdam M.E., Dinjens W.N., Bleeker F.E. Somatic mosaicism by a de novo MLH1 mutation as a cause of Lynch syndrome. Mol. Genet. Genom. Med. 2019;7:e00699. doi: 10.1002/mgg3.699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Thompson B.A., Spurdle A.B., Plazzer J.-P., Greenblatt M.S., Akagi K., Al-Mulla F., Bapat B., Bernstein I., Capellá G., den Dunnen J.T., et al. Application of a 5-tiered scheme for standardized classification of 2,360 unique mismatch repair gene variants in the InSiGHT locus-specific database. Nat. Genet. 2014;46:107–115. doi: 10.1038/ng.2854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Leclerc J., Vermaut C., Buisine M.-P. Diagnosis of Lynch Syndrome and Strategies to Distinguish Lynch-Related Tumors from Sporadic MSI/dMMR Tumors. Cancers. 2021;13:467. doi: 10.3390/cancers13030467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Edmunds J.W., Mahadevan L.C., Clayton A.L. Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J. 2007;27:406–420. doi: 10.1038/sj.emboj.7601967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Li F., Mao G., Tong D., Huang J., Gu L., Yang W., Li G.-M. The Histone Mark H3K36me3 Regulates Human DNA Mismatch Repair through Its Interaction with MutSα. Cell. 2013;153:590–600. doi: 10.1016/j.cell.2013.03.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Ortega J., Li J., Lee S., Tong D., Gu L., Li G.-M. Phosphorylation of PCNA by EGFR inhibits mismatch repair and promotes misincorporation during DNA synthesis. Proc. Natl. Acad. Sci. USA. 2015;112:5667–5672. doi: 10.1073/pnas.1417711112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Wu J.N., Roberts C.W.M. ARID1A mutations in cancer: Another epigenetic tumor suppressor? Cancer Discov. 2013;3:35–43. doi: 10.1158/2159-8290.CD-12-0361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Shen J., Ju Z., Zhao W., Wang L., Peng Y., Ge Z., Nagel Z.D., Zou J., Wang C., Kapoor P., et al. ARID1A deficiency promotes mutability and potentiates therapeutic antitumor immunity unleashed by immune checkpoint blockade. Nat. Med. 2018;24:556–562. doi: 10.1038/s41591-018-0012-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Billingsley C.C., Cohn D.E., Mutch D.G., Stephens J.A., Suarez A.A., Goodfellow P.J. Polymerase ε (POLE) mutations in endometrial cancer: Clinical outcomes and implications for Lynch syndrome testing. Cancer. 2015;121:386–394. doi: 10.1002/cncr.29046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Louis P., Hold G.L., Flint H.J. The gut microbiota, bacterial metabolites and colorectal cancer. Nat. Rev. Microbiol. 2014;12:661–672. doi: 10.1038/nrmicro3344. [DOI] [PubMed] [Google Scholar]
- 104.Li S.K., Martin A. Mismatch Repair and Colon Cancer: Mechanisms and Therapies Explored. Trends Mol. Med. 2016;22:274–289. doi: 10.1016/j.molmed.2016.02.003. [DOI] [PubMed] [Google Scholar]
- 105.Tseng-Rogenski S.S., Hamaya Y., Choi D.Y., Carethers J.M. Interleukin 6 Alters Localization of hMSH3, Leading to DNA Mismatch Repair Defects in Colorectal Cancer Cells. Gastroenterology. 2015;148:579–589. doi: 10.1053/j.gastro.2014.11.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Chang C.L., Marra G., Chauhan D.P., Ha H.T., Chang D.K., Ricciardiello L., Randolph A., Carethers J.M., Boland C.R. Oxidative stress inactivates the human DNA mismatch repair system. Am. J. Physiol. Cell Physiol. 2002;283:C148–C154. doi: 10.1152/ajpcell.00422.2001. [DOI] [PubMed] [Google Scholar]
- 107.Greger V., Passarge E., Messmer E., Horsthemke B. Epigenetic changes may contribute to the formation and spontaneous regression of retinoblastoma. Qual. Life Res. 1989;83:155–158. doi: 10.1007/BF00286709. [DOI] [PubMed] [Google Scholar]
- 108.Sakai T., Toguchida J., Ohtani N., Yandell D.W., Rapaport J.M., Dryja T.P. Allele-specific hypermethylation of the retinoblastoma tumor-suppressor gene. Am. J. Hum. Genet. 1991;48:880–888. [PMC free article] [PubMed] [Google Scholar]
- 109.Herman J.G., Latif F., Weng Y., Lerman M.I., Zbar B., Liu S., Samid D., Duan D.S., Gnarra J., Linehan W.M. Silencing of the VHL tumor-suppressor gene by DNA methylation in renal carcinoma. Proc. Natl. Acad. Sci. USA. 1994;91:9700–9704. doi: 10.1073/pnas.91.21.9700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Dobrovic A., Simpfendorfer D. Methylation of the BRCA1 gene in sporadic breast cancer. Cancer Res. 1997;57:3347–3350. [PubMed] [Google Scholar]
- 111.Kane M.F., Loda M., Gaida G.M., Lipman J., Mishra R., Goldman H., Jessup J.M., Kolodner R. Methylation of the hMLH1 promoter correlates with lack of expression of hMLH1 in sporadic colon tumors and mismatch repair-defective human tumor cell lines. Cancer Res. 1997;57:808–811. [PubMed] [Google Scholar]
- 112.Buckley A.R., Ideker T., Carter H., Harismendy O., Schork N.J. Exome-wide analysis of bi-allelic alterations identifies a Lynch phenotype in The Cancer Genome Atlas. Genome Med. 2018;10:69. doi: 10.1186/s13073-018-0579-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Jansen A.M., Van Wezel T., Akker B.E.V.D., García M.V., Ruano D., Tops C.M., Wagner A., Letteboer T.G., Gomez-Garcia E., Devilee P., et al. Combined mismatch repair and POLE/POLD1 defects explain unresolved suspected Lynch syndrome cancers. Eur. J. Hum. Genet. 2015;24:1089–1092. doi: 10.1038/ejhg.2015.252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Lefol C., Sohier E., Baudet C., Naïbo P., Ruano E., Grand-Masson C., Viari A., Wang Q. Acquired somatic MMR deficiency is a major cause of MSI tumor in patients suspected for “Lynch-like syndrome” including young patients. Eur. J. Hum. Genet. 2020;29:482–488. doi: 10.1038/s41431-020-00778-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Elze L., Mensenkamp A.R., Nagtegaal I.D., van Zelst-Stams W.A., de Voer R.M., Ligtenberg M.J., Dommering C.J., Hoogerbrugge N., de Jong M.M., Bleeker F.E., et al. Somatic Nonepigenetic Mismatch Repair Gene Aberrations Underly Most Mismatch Repair–Deficient Lynch-Like Tumors. Gastroenterology. 2021;160:1414–1416.e3. doi: 10.1053/j.gastro.2020.11.042. [DOI] [PubMed] [Google Scholar]
- 116.Hemminger J.A., Pearlman R., Haraldsdottir S., Knight D., Jonasson J.G., Pritchard C.C., Hampel H., Frankel W.L. Histology of colorectal adenocarcinoma with double somatic mismatch-repair mutations is indistinguishable from those caused by Lynch syndrome. Hum. Pathol. 2018;78:125–130. doi: 10.1016/j.humpath.2018.04.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Nagasaka T., Rhees J., Kloor M., Gebert J., Naomoto Y., Boland C.R., Goel A. Somatic Hypermethylation of MSH2 Is a Frequent Event in Lynch Syndrome Colorectal Cancers. Cancer Res. 2010;70:3098–3108. doi: 10.1158/0008-5472.CAN-09-3290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Rumilla K., Schowalter K.V., Lindor N.M., Thomas B.C., Mensink K.A., Gallinger S., Holter S., Newcomb P.A., Potter J., Jenkins M., et al. Frequency of Deletions of EPCAM (TACSTD1) in MSH2-Associated Lynch Syndrome Cases. J. Mol. Diagn. 2011;13:93–99. doi: 10.1016/j.jmoldx.2010.11.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Vymetalkova V.P., Slyskova J., Korenkova V., Bielik L., Langerova L., Prochazka P., Rejhova A., Schwarzova L., Pardini B., Naccarati A., et al. Molecular characteristics of mismatch repair genes in sporadic colorectal tumors in Czech patients. BMC Med. Genet. 2014;15:17. doi: 10.1186/1471-2350-15-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Moura Lima E., Ferreira Leal M., de Arruda Cardoso Smith M., Rodríguez Burbano R., Pimentel de Assumpção P., Bello M.J., Rey J.A., Ferreira de Lima F., Casartelli C. DNA mismatch repair gene methylation in gastric cancer in individuals from northern Brazil. Biocell. 2008;32:237–243. doi: 10.32604/biocell.2008.32.237. [DOI] [PubMed] [Google Scholar]
- 121.Truninger K., Menigatti M., Luz J., Russell A., Haider R., Gebbers J.O., Bannwart F., Yurtsever H., Neuweiler J., Riehle H.M., et al. Immunohistochemical analysis reveals high frequency of PMS2 defects in colorectal cancer. Gastroenterology. 2005;128:1160–1171. doi: 10.1053/j.gastro.2005.01.056. [DOI] [PubMed] [Google Scholar]
- 122.Kamps R., Brandão R.D., van den Bosch B.J., Paulussen A.D., Xanthoulea S., Blok M.J., Romano A. Next-Generation Sequencing in Oncology: Genetic Diagnosis, Risk Prediction and Cancer Classification. Int. J. Mol. Sci. 2017;18:308. doi: 10.3390/ijms18020308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Golubicki M., Bonjoch L., Acuña-Ochoa J.G., Díaz-Gay M., Muñoz J., Cuatrecasas M., Ocaña T., Iseas S., Mendez G., Cisterna D., et al. Germline biallelic Mcm8 variants are associated with early-onset Lynch-like syndrome. JCI Insight. 2020:5. doi: 10.1172/jci.insight.140698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Hampel H., Pearlman R., Beightol M., Zhao W., Jones D., Frankel W.L., Goodfellow P.J., Yilmaz A., Miller K., Bacher J., et al. Assessment of Tumor Sequencing as a Replacement for Lynch Syndrome Screening and Current Molecular Tests for Patients With Colorectal Cancer. JAMA Oncol. 2018;4:806–813. doi: 10.1001/jamaoncol.2018.0104. [DOI] [PMC free article] [PubMed] [Google Scholar]