Abstract
DNA methylation is one of the most important epigenetic mechanisms to regulate gene expression, which is highly dynamic during development and specifically maintained in somatic cells. Aberrant DNA methylation patterns are strongly associated with human diseases including cancer. How are the cell-specific DNA methylation patterns established or disturbed is a pivotal question in developmental biology and cancer epigenetics. Currently, compelling evidence has emerged that long non-coding RNA (lncRNA) mediates DNA methylation in both physiological and pathological conditions. In this review, we provide an overview of the current understanding of lncRNA-mediated DNA methylation, with emphasis on the roles of this mechanism in cancer, which to the best of our knowledge, has not been systematically summarized. In addition, we also discuss the potential clinical applications of this mechanism in RNA-targeting drug development.
Keywords: lncRNA, DNA methylation, Non-coding RNA, DNMT, TET, Cancer
Background
DNA methylation is the methyl modification on the fifth carbon of cytosines (5-methylcytosine, 5mC) typically found in the context of symmetrical CpG dinucleotides in mammals [1, 2]. It is estimated that 70–80% of CpG sites in the mammalian genome are methylated [3], excluding specific regions called CpG islands (CGIs). CGIs are CpG-rich sequences of about 1 kilo-base (kb) in length that mostly exist in gene promoters [4]. Approximately 60% of human gene promoters contain CGIs [5].
DNA methylation is established by DNA methyltransferases (DNMTs). In the simplified but widely accepted ‘division of labor’ model, it is proposed that DNMT3A and DNMT3B are essential for de novo DNA methylation, while DNMT1 is for methylation maintenance during DNA replication [6]. Ten-eleven translocation (TET) family of enzymes (TET1, TET2, and TET3) oppose the actions of the DNMT family by oxidation of 5mC, followed by replication-dependent dilution or thymine DNA glycosylase (TDG)-dependent base excision repair, leading to active DNA demethylation [7–9].
Genome-scale analysis revealed distinct DNA methylation patterns across different cell types, developmental stages, and in response to different stimuli [3, 10, 11]. Aberrant DNA methylation pattern is associated with diseases, including cancer [12–15]. In cancer cells, whereas the general DNA methylation levels are reduced, the CGIs are hypermethylated in a cancer-specific manner [16, 17]. These observations raised a fundamental question: how does the cell type-specific DNA methylation pattern established across the genome? It is well-demonstrated that histone modification and chromosome remodeling [18], as well as transcriptional factors, play key roles in the regulation of DNA methylation genome-wide and in site-specific manner [19–22]. Studies in recent years have accumulated compelling evidence to suggest that long non-coding RNA (lncRNA) is another important regulator of DNA methylation, especially in cancer.
While less than 2% of the human genome encodes proteins, nearly three-quarters can be actively transcribed into non-coding RNAs [23], amongst the ones typically with length more than 200 nucleotides are cataloged as lncRNAs. According to a current statistical analysis, there are more than 173,112 annotated lncRNAs transcribed from 96,411 genomic loci [24]. It is demonstrated that lncRNAs play versatile roles in development and diseases including cancer [25–27]. In the nucleus, lncRNAs regulate chromatin remodeling and transcription; In the cytoplasm, lncRNAs regulate translation and mRNA turnover (reviewed in ref. [27]). There is accumulating evidence up to date showing that lncRNAs mediate DNA methylation via multiple manners, thereby regulating target gene expression in diverse physiological and pathological processes. In this review, we summarize our current understanding of lncRNA-mediated DNA methylation, with emphasis on the functions of this mechanism in cancer. The future direction and potential clinical application are also discussed.
LncRNAs recruit DNA methyltransferases
More than a decade ago, it was discovered that lncRNAs transcribed from the promoter of rRNA genes (rDNA) regulate DNA methylation and transcription of rDNA [28]. Later, it was demonstrated that this kind of lncRNA interacts with rDNA promoter and forms a DNA: RNA triplex, which is recognized by DNMT3B to epigenetically regulate rDNA expression [29, 30]. Although it is still unclear if this is a common model nowadays, a variety of lncRNAs have been reported to recruit DNMTs and regulate target gene expression, playing key roles in mesoderm commitment [31], muscle regeneration [32, 33], neural differentiation [34], adipogenesis [35], mental disorder [36], cardiovascular diseases [37–40], osteoarthritis [41], as well as types of cancer (Table 1).
Table 1.
LncRNAs mediate DNA methylation in cancer
| lncRNA | Role | Factor | Target | Function | Cancer | Ref |
|---|---|---|---|---|---|---|
| TINCR | Recruit | DNMT1 | miR-503-5p | Regulate EGFR expression | BC | [42] |
| MROS-1 | Recruit | DNMT3A | PRUNE2 | Nodal metastases | OC | [43] |
| HOTAIR | Recruit | DNMT1 | PTEN | Cell proliferation, invasion and migration | CML | [44] |
| LINC00887 | Recruit | DNMT1 | CA9 | Suppress oncogenic CA9 | TSCC | [45] |
| LINC00472 | Recruit | DNMTs | MCM6 | Inhibited tumor growth and metastasis | TNBC | [46] |
| LINC01270 | Recruit | DNMTs | GSTP1 | Promote tumorigenesis and drug resistance | EC | [47] |
| DLX6-AS1 | Recruit | DNMT1 | LARGE | Promotes Lymph Node Metastasis | PCa | [48] |
| HOTAIR | Recruit | DNMTs | MTHFR | chemoresistance | EC | [49] |
| ADAMTS9-AS2 | Recruit | DNMT1/3 | CDH3 | Inhibits proliferation, invasion, and migration | EC | [50] |
| IRAIN | Recruit | DNMT1/3 | VEGFA | Suppresses tumor growth | RC | [51] |
| PVT1 | Recruit | DNMT1 | miR-18b-5p | Promotes proliferation | GBC | [52] |
| DLX6-AS1 | Recruit | DNMTs | CADM1 | Maintenance of cancer stem cells | HCC | [53] |
| BZRAP1-AS1 | Recruit | DNMT3b | THBS1 | Promotes angiogenesis | HCC | [54] |
| KCNQ1OT1 | Recruit | DNMT1 | Kcnq1 | Promotes chemoresistance | OSA | [55] |
| PYCARD-AS1 | Recruit | DNMT1, G9a | PYCARD | Regulates apoptosis | BC | [56] |
| MIR210HG | Recruit | DNMT1 | CACNA2D2 | Promotes proliferation and invasion | NSCLC | [57] |
| HAGLR | Recruit | DNMT1 | E2F1 | Suppresses tumor growth | LUAD | [58] |
| DACOR1 | Recruit | DNMT1 | Genome-wide | CRC | [59, 60] | |
| LINC00628 | Recruit | DNMTs | LAMA3 | Promotes tumorigenesis and drug resistance | LUAD | [61] |
| PVT1 | Recruit | DNMT1 | BNIP3 | Promotes cell proliferation | GC | [62] |
| HOTAIR | Recruit | DNMT3B | HOXA5 | Promotes cell proliferation | AML | [63] |
| MALAT1 | Mitochondrial DNA | Control metabolic Reprogramming | HCC | [64] | ||
| HOTAIR | Upregulate | DNMT3b | PTEN | Doxorubicin resistance | AML | [65] |
| RP11-159K7.2 | Upregulate | DNMT3A | Promotes cell growth and invasion | LSCC | [66] | |
| GAS5 | Down-regulate | DNMTs | miR-424 | Suppresses multiple malignant phenotypes | Glioma | [67] |
| lnc-OIP5-AS1 | Upregulate | DNMT1 | pre-miR-218–1 | Promote cell motility and proliferation | KS | [68] |
| Linc-GALH | Ubiquitinate | DNMT1 | Gankyrin | Promotes metastasis | HCC | [69] |
| LUCAT1 | Inhibits ubiquitination | DNMT1 | tumor-suppressor genes | Promotes tumor formation and metastasis | ESCC | [70] |
| HOTAIR | Upregulate (via EZH2) | DNMTs | miR-122 | Activate Cyclin G1 and promote tumorigenicity | HCC | [71] |
| HOTAIR | Upregulate | DNMT1/3B | HOXA1 | Multidrug resistance | SCLC | [72] |
| H19 | Upregulate | TET3 | MED12 | Promotes cell proliferation | UL | [73] |
| DBCCR1-003 | Sequestrate | DNMT1 | DBCCR1 | Inhibits cell growth | BCa | [74] |
| TTTY15 | Sequestrate | DNMT3A | TBX4 | Suppresses metastasis | NSCLC | [75] |
| HOTAIRM1 | Sequestrate | G9a/EZH2/ DNMTs | HOXA1 | Promotes tumor growth and invasion | GBM | [76] |
| 91H | Repel | DNMTs | H19/IGF2 locus | Promotes tumorigenesis | BC | [77] |
| HOTAIR | Recruit (via EZH2) | HOXA1 | Multidrug resistance | SCLC | [78] | |
| SNHG3 | Recruit (via EZH2) | MED18 | Promotes cell migration and invasion | GC | [79] | |
| HOXB13-AS1 | Recruit (via EZH2) | DNMT3B | HOXB13 | Promotes cell proliferation | Glioma | [80] |
| Lnc-LALC | Recruit (via EZH2) | DNMTs | LZTS1 | Liver metastasis | CRC | [81] |
| HOTAIR | Recruit (via EZH2) | DNMT1 | miR-454-3p | Promotes tumor growth | CS | [82] |
| GIHCG | Recruit (via EZH2) | DNMT1 | miR-200b/a/429 | Promotes tumor growth and metastasis | HCC | [83] |
| LINC00630 | Restrict (via EZH2) | DNMT3B | BEX1 | Suppresses cell apoptosis and promotes radio-resistance | CRC | [84] |
| Lnc34a | Recruit (via PHB2) | DNMT3A | miR-34a | Promotes cell proliferation | CRC | [85] |
| H19 | Inhibit (via inhibiting SAHH) | DNMT3b | Beclin1 | Induces autophagy activation and tamoxifen resistance | BC | [86] |
| LINC00662 | Regulate | MAT1A/ SAHH | Activates SAM-dependent oncogenes | HCC | [87] | |
| SNHG6 | Regulate (via miRNAs) | MAT1A, MAT2A | Genome-wide | HCC | [88] | |
| H19 | Inhibit (via inhibiting SAHH) | DNMTs | LINE-1 | Benzo [a]pyrene (BaP) carcinogenesis | Lung cancer | [89] |
| MAGI2-AS3 | Recruit | TET2 | LRIG1 | Inhibits the self-renewal of leukaemic stem cells | AML | [90] |
| SSTR5-AS1 | Recruit | TET1 | E-cadherin | Inhibits tumor progression and metastasis | LSCC | [91] |
| SATB2-AS1 | Recruit (via GADD45A) | TETs | SATB2 | Inhibits cell metastasis and regulates immune response | CRC | [92] |
Abbreviations: BC Breast cancer, OC Oral cancer, CML Chronic myeloid leukemia, TSCC Tongue squamous cell carcinoma, TNBC Triple-negative breast cancer, EC Esophageal cancer, PCa Prostate cancer, RC Renal carcinoma, GBC Gallbladder cancer, HCC Hepatocellular carcinoma, OSA Osteosarcoma, NSCLC Non-small cell lung cancer, LUAD Lung adenocarcinoma, CRC Colorectal cancer, GC Gastric cancer, AML Acute myeloid leukemia, LSCC Laryngeal squamous cell carcinoma, KS Kaposi’s sarcoma, ESCC Esophageal squamous cell carcinoma, SCLC small-cell lung cancer, UL Uterine leiomyomas, GBM Glioblastoma multiforme, CS Chondrosarcoma
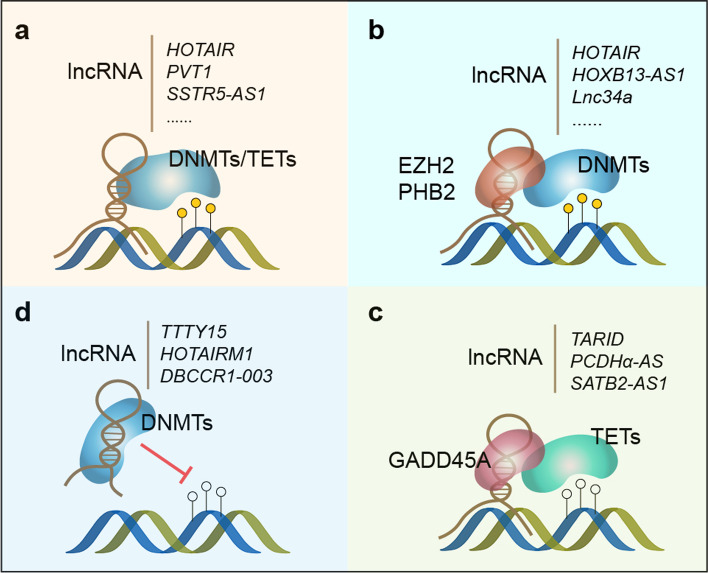
Using an optimized RIP-seq method, Merry et al. identified 148 lncRNAs interacting with DNMT1 in colon cancer cells [59], and the following investigation showed that one of these lncRNAs, DACOR1, could recruit DNMT1 and reprogram genome-wide DNA methylation [60]. Currently, a growing number of studies suggest that lncRNA might recruit DNMTs directly to specific targets (Fig. 1a), including both protein-coding genes [43, 44, 46, 47, 49–51, 55, 57, 58, 62, 63] and non-coding genes such as miRNA [42, 52, 93]. For instance, in esophageal cancer (EC), lncRNA ADAMTS9-AS2 was reported to recruit DNMT1/3 to CDH3 promoter, inhibiting the cancer cell proliferation, invasion, and migration [50]. Two other lncRNAs, HOTAIR and LINC01270 might recruit DNMTs to the promoters of MTHFR and GSTP1 respectively, leading to chemoresistance in EC [47, 49]. In lung adenocarcinoma (LUAD), lncRNA HAGLR was identified as a tumor suppressor by recruiting DNMT1 to the promoter of E2F1 to inhibit tumor growth [58]. A recent study revealed a more complex scenario, in which the authors identified two novel variants of lncRNA LINC00887, and showed that the short form variant suppressed Carbonic Anhydrase IX (CA9) by recruiting DNMT1 to its promoter, while the long-form variant activated CA9's transcription via interacting with HIF1α [45]. The two variants were supposed to differentially respond to hypoxia and oppositely control the progression of tongue squamous carcinoma [45].
Fig. 1.
LncRNA interacts with DNMTs/TETs. a. LncRNAs directly recruit DNMTs/ TETs. b. LncRNAs indirectly recruit DNMTs via EZH2/PHB2. c. LncRNAs indirectly recruit TETs via GADD45A. d. LncRNAs sequestrate DNMTs
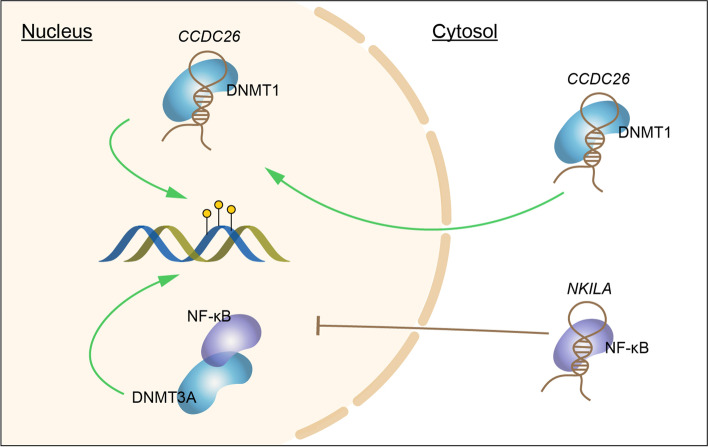
Meanwhile, several groups also proposed that lncRNAs could recruit DNMT indirectly through the mediation of other factors (Fig. 1b). It was previously proposed that the polycomb group (PcG) protein EZH2 (Enhancer of Zeste homolog 2) interacts with DNMT and associates with DNMT activity [94]. Studies in recent years demonstrated in diverse cancers that lncRNAs might regulate DNA methylation of target genes via association with EZH2, promoting tumor growth [80, 82], metastasis [79, 81, 83] and radio-resistance [84]. Alternatively, EZH2 might regulate DNA methylation by the formation of H3K27me3 histone modification [78], while the molecular mechanism involved in H3K27me3-induced DNA methylation is unclear. Apart from histone modifier EZH2, two transcriptional regulators, NF-κB and PHB2 were also reported to interact with DNMT3A [85, 95]. LncRNA NKILA was identified as a suppressor of NF-κB by sequestering NF-κB in cytoplasm [96]. Upon proinflammatory stimuli, NF-κB is released from the sequestration and translocated into the nucleus (Fig. 2). DNMT3A is then recruited to the promoter of KLF4 by NF-κB, repressing KLF4 transcription by DNA methylation [95]. Another study by Wang et al. reported a lncRNA called Lnc34a, which could interact with Prohibitin 2 (PHB2) and then recruit DNMT3A to miR-34a promoter, silencing miR-34a expression and promoting colorectal cancer growth [85]. PHB2 is a multi-functional protein that can shuttle between nucleus and mitochondria [97]. Interestingly, the nuclear-encoded lncRNA MALAT1 was recently discovered to be transported into mitochondria and to regulate the methylation status of mitochondrial DNA in hepatocellular carcinoma [64], yet the detailed mechanism is unclear.
Fig. 2.
LncRNAs regulate DNMT activity via nucleocytoplasmic shuttling. LncRNA CCDC26 interacts with DNMT1 and promotes its localization from the cytosol to the nucleus. LncRNA NKILA sequesters NF-κB in the cytoplasm, which hinders NF-κB recruitment of DNMT3A in the nucleus
While most of the reported function of lncRNA recruitment of DNMT is to target DNMT to specific genomic sites or regions, recent work from Jones et al. proposed a different model, in which the lncRNA CCDC26 specifically interacts with DNMT1 and promote its localization from the cytosol to nucleus (Fig. 2), while removal of CCDC26 leads to genome-wide hypomethylation, increasing double-stranded DNA breaks and inducing cell death [98]. More investigation is needed to confirm if the interaction is direct and to reveal the detailed mechanisms.
LncRNAs recruit TET enzymes
TET (Ten-eleven Translocation)-mediated 5mC oxidation is responsible for the active erasure of DNA methylation [99]. Studies from recent years have revealed that a subset of lncRNAs has the potential to interact with TETs and regulate DNA methylation (Table 1).
In some cases, lncRNA directly interacts with TETs and recruits them to specific targets (Fig. 1a). It was demonstrated that lncRNA Oplr16 binds to the Oct4 promoter, orchestrating the promoter-enhancer loops and then interacts with TET2 by the 3' region of Oplr16 [100]. Similarly, Du et al. identified two motifs in lncRNA Platr10 that interact with Oct4 promoter and TET1 respectively, thus inducing TET1- mediated DNA demethylation at specific site [101]. A research by Zhou et al. suggested that lncRNA TETILA regulates TET2 subcellular localization and enzymatic activity by binding to the DSBH (double-stranded β-helix) domain of TET2 [102]. In acute myeloid leukemia, lncRNA MAGI2-AS3 recruits TET2 to LRIG1 promoter, inducing up-regulation of LRIG1 and inhibition of leukemic stem cell self-renewal [90]. Interestingly, using RNA reverse transcription-associated trap sequencing (RAT-seq) approach to profile genome-wide interaction targets for lncRNAs in mice, a recent study reported that lncRNA Peblr20 recruits TET2 to the enhancer of Pou5F1 and activates the enhancer-transcribed RNAs [103]. Whether a similar mechanism exists in humans especially in cancer development remains uninvestigated.
There is also evidence supporting an indirect model (Fig. 1c), in which lncRNAs recruit TET via GADD45A. It was first reported by Arab et al. that an antisense lncRNA from TCF21 gene locus termed TARID might recruit GADD45A (growth arrest and DNA-damage-inducible, alpha), and GADD45A then recruits TET to the promoter of its partner gene and induce its activation by DNA demethylation [104]. In the following work, the authors further showed that TARID forms an R-loop at the TCF21 promoter to recruit GADD45A [105]. It was speculated that lncRNA PCDHα-AS might function in a similar mechanism to recruit TET3 via GADD45A, driving stochastic promoter choice to establish a neuronal surface identity code for circuit assembly [106]. In colorectal cancer (CRC), lncRNA SATB2-AS1 directly recruits WDR5 and GADD45A, promoting SATB2 transcription by histone modification, as well as DNA demethylation [92], which inhibits cell metastasis and regulates the immune response in CRC. Recently, a database was created, with a comprehensive list of R-loops and their respective regulatory proteins [107], which might serve as a useful resource to identify novel lncRNAs with the potential to recruit GADD45A via formation of R-loops.
LncRNAs repel/ sequestrate DNA methyltransferases
While most of the current reports suggest the DNMT-recruiting role of lncRNAs, some lncRNAs are also shown to repel or sequestrate DNMT to negatively regulate DNA methylation (Fig. 1d and Table 1).
It was first reported by Di Ruscio et al. that a lncRNA arising from the CEBPA gene locus binds to DNMT1 and prevents CEBPA promoter methylation [108]. The lncRNA DBCCR1-003 was reported to function similarly to suppress DBCCR1 promoter methylation by sequestrating DNMT1 and eventually to inhibit cell growth in bladder cancer [74]. In non-small cell lung cancer, lncRNA TTTY15 interacts with DNMT3A and inhibits the binding of DNMT3A to TBX4 promoter, while the lower expression level of TTTY15 is associated with tumor metastasis [75]. In glioblastoma, lncRNA HOTAIRM1 was suggested to interact with several epigenetic factors including DNMT1/3A/3B to sequester them away from HOXA1 promoter [76]. In breast cancer, it was discovered that lncRNA 91H, which is transcribed from the antisense orientation of H19, promotes oncogenesis by masking methylation site on the H19 promoter, inducing the oncogenic H19 overexpression [77].
LncRNAs control SAM/ SAH level to regulate DNMT activity
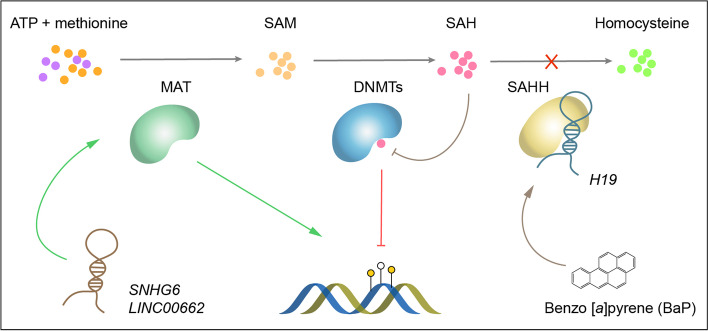
DNMT catalyzes transmethylation reactions using S-adenosylmethionine (SAM) as the methyl group donor, yielding S-adenosylhomocysteine (SAH) as a by-product, which is also a strong feedback inhibitor of DNMT [6]. In mammals, SAM is biosynthesized by methionine adenosyltransferase (MAT) from ATP and methionine [109], while SAH is reversibly cleaved into adenosine and homocysteine by S-adenosylhomocysteine hydrolase (SAHH, also known as AdoHcy hydrolase, AHCY), which is essential to prevent accumulation of SAH [109], thereby relieving its inhibition to DNMT (Fig. 3).
Fig. 3.
LncRNAs control SAM/SAH level to regulate DNMT activity. S-adenosylhomocysteine (SAM) is biosynthesized by MAT (methionine adenosyltransferase) and converted to SAH (S-adenosylhomocysteine) by DNMTs. SAH is also a strong feedback inhibitor of DNMTs and it can be cleaved by S-adenosylhomocysteine hydrolase (SAHH). LncRNAs control SAM/SAH level by interacting with MAT or SAHH, and carcinogens such as benzo [a]pyrene (BaP) might enhance the interaction
It was proposed that lncRNA H19 binds to and inhibits SAHH, leading to genome-wide methylation changes at numerous gene loci [110]. Afterward, this mechanism was verified in embryonic hematopoietic stem cell development [111], odontogenic differentiation [112], metabolic abnormality [113] and neurodegenerative diseases [114]. In breast cancer, it was demonstrated that H19 inhibits SAHH, resulting in the accumulation of SAH, which restricts DNMT3B from methylating Beclin1 promoter and inducing the upregulation of Beclin1 and subsequently initiates autophagy, contributing to tamoxifen resistance [86]. Interestingly, the interaction of H19 and SAHH might be enhanced by Benzo [a]pyrene (BaP), which is a potent carcinogen, especially in lung cancer [89].
Other than the SAH level regulated by SAHH, the SAM level regulated by MAT is another factor affecting DNMT activity (Fig. 3). MAT has several homologs and isoenzymes, among which, MAT1A is mainly expressed in adulthood, serving as a marker for the normal differentiated liver. While MAT2A is a marker for rapid liver growth and dedifferentiation, which is transcriptionally induced in hepatocellular carcinoma (HCC) [109]. It was reported that the oncogenic lncRNA SNHG6 upregulates MAT2A expression as a competitive endogenous RNA (ceRNA) to sponge miR-1297, while down-regulates MAT1A translation by suppressing nucleocytoplasmic shuttling of MAT1A mRNA, thereby causing genome-wide hypomethylation and promoting HCC [88]. Recently, the same group of investigators identified a novel lncRNA named LINC00662 that was shown to decay MAT1A mRNA by RNA–RNA interactions and degrades SAHH protein by ubiquitination [87]. These studies revealed a pathway regulating the level of SAM/SAH to further control DNMT activity, with broad functions in cancer and other diseases.
LncRNAs regulate the expression of DNMTs/ TETs
There is compelling evidence showing that lncRNAs control the expression of DNMTs and TETs at diverse levels to regulate DNA methylation (Table 1 and Fig. 4). It was reported that lncRNAs promote or suppress DNMT expression, playing key roles in osteogenesis [115], macrophage polarization [116], as well as cell invasion in Kaposi's sarcoma [68] and chemoresistance in small cell lung cancer [72] and acute myeloid leukemia [65]. Several molecular mechanisms of lncRNA’s regulatory effect on DNMTs or TETs have been elucidated (Fig. 4).
Fig. 4.
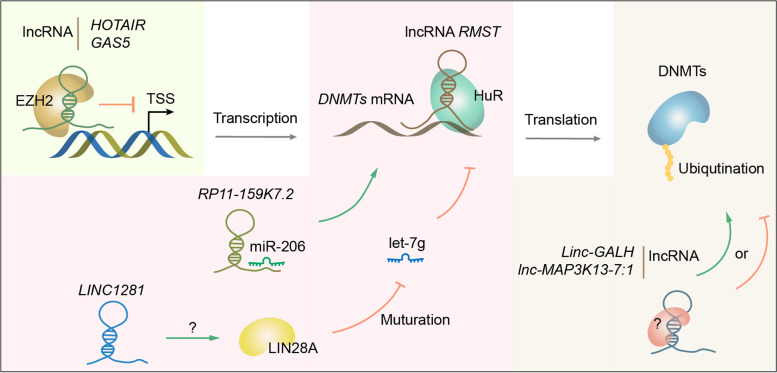
LncRNAs regulate the expression of DNMTs/ TETs at diverse levels. Firstly, lncRNAs might regulate the transcription of DNMTs via interaction with EZH2 to form repressive chromatin; Secondly, lncRNAs can stabilize DNMTs mRNA by recruiting HuR or as a miRNA sponge; Thirdly, lncRNAs regulate DNMT on the protein level by promoting or inhibiting its ubiquitination
The first mechanism is to regulate the transcription, as demonstrated in malignant glioma, where lncRNA GAS5 directly interacts with EZH2 and stimulates the formation of polycomb repressive complex 2 (PRC2), thereby transcriptionally suppressing DNMT [67]. There is also a report suggesting that EZH2 is recruited by lncRNA HOTAIR to upregulate DNMT, while the mechanism is unclear [71].
The second mechanism is to regulate the stability of DNMT mRNA, where lncRNA functions as a mediator to upregulating DNMT by interaction with the stabilizing factor HuR [117], or as a ceRNA to sponge specific miRNA, thereby upregulating DNMT [66]. The latter mechanism was also discovered in TET regulation, where estradiol and progesterone upregulate lncRNA H19 to suppress miRNA Let-7 and stabilize TET3 mRNA, activating key fibroid-promoting genes in uterine leiomyomas [73]. LncRNA might also exert this effect via a more indirect manner, as demonstrated for LINC1281, which stabilizes the expression of Let-7 miRNA, thus down-regulating its targets DNMT3A/B [118].
The third mechanism is to regulate DNMT at the protein level. Current studies mainly focus on protein degradation by ubiquitination (Fig. 4). It was reported by several groups that lncRNAs serve as a protein-binding scaffold and induce ubiquitin-mediated DNMT protein degradation, epigenetically regulating target gene expression in obesity-mediated beta cell dysfunction [119], polycystic ovary syndrome [120] and hepatocellular carcinoma (HCC) [69]. The detailed mechanism involving the role of lncRNA in DNMT ubiquitination is largely unknown and warrant more deep investigation. In esophageal squamous cell carcinoma, a distinct model was proposed, in which, the lncRNA LUCAT1 binds DNMT1 to protect it from ubiquitination, while LUCAT1 knock-down promotes ubiquitination of DNMT1 through UHRF1 (Ubiquitin-Like PHD and RING Finger Domain-Containing Protein 1) [70]. However, it is well established that UHRF1 deposits dual mono- ubiquitination on the H3 histone tail and PCNA-associated factor 15 (PAF15) for direct DNMT1 recruitment and DNA methylation maintenance [121–123], while its roles in the mediation of DNMT1 ubiquitination need further validation and investigation.
Conclusions and discussions
Studies in recent years have revealed the multi-faceted role of lncRNA in regulating DNA methylation. Firstly, lncRNAs can recruit or repel DNA modifiers (DNMTs/ TETs) to specific gene targets (Fig. 1; Fig. 2); Secondly, lncRNAs can regulate DNMT activity by controlling the level of DNMT cofactor SAM/ SAH (Fig. 3); Lastly, lncRNAs can regulate the expression of DNMTs/ TETs per se at multiple levels (Fig. 4). All these mechanisms have been investigated in development and disease, with emphasized roles in cancer.
While most of the studies focused on the DNA methylation of the gene promoters, there is also a recent report highlighting the gene-body methylation mediated by a lncRNA by recruiting DNMT3A, which facilitates transcription of CTSG in dermatomyositic myoideum [124]. Whether this mechanism exists in cancer needs further investigation.
Although this review mainly discussed the lncRNA function in mediating DNA methylation, another two issues should be noted. The first is that lncRNAs are in turn regulated targets of DNA methylation [125–128]; The second is that lncRNAs also mediate other epigenetic alterations such as histone modification and chromosome remodeling [129–136]. These issues provide an additional layer of gene expression regulation to form complex crosstalk between lncRNA, transcriptional factors, and various epigenetic modifications. More elaborate investigations are warranted to reveal the common mechanisms.
Perspectives
The emerging roles of lncRNAs in cancer through the mediation of DNA methylation suggest novel applications in drug development. While there are currently no drugs targeting lncRNA based exactly on this mechanism, relevant studies shed light on this field (Fig. 5).
Fig. 5.
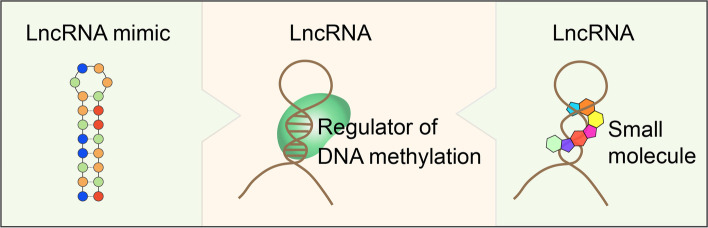
Potential therapeutic strategies for lncRNA targeting based on lncRNA- mediated DNA methylation. The middle panel shows the interaction between lncRNA and DNA methylation regulators, which could be specifically interrupted by small molecular compounds (right panel). Alternatively, the regulators could also be modulated by lncRNA mimics (left panel)
One direction is to design lncRNA mimics to regulate the activity of their target proteins, which was recently applied in treating a rare disease of phenylketonuria, where a lncRNA HULC was identified to interact with phenylalanine hydroxylase (PAH) and to modulate the enzymatic activities of PAH. In their work, the authors constructed a lncRNA mimic that rescues PAH enzymatic activity in HULC-deficient cells and mouse models, which showed the therapeutic potential for phenylketonuria [137].
Another direction is to design small molecules directly targeting lncRNA-protein interactions [138–141]. Based on the structural insight of the interaction between lncRNA HOTAIR and EZH2, Ren et al. conducted a high-throughput virtual screening and identified a compound that selectively interrupts the lncRNA-protein interaction and inhibits cancer cell invasion and migration [142].
Owing to the fast progress of RNA structural biology and screening technologies, as well as the in-depth mechanistic studies and drug delivery technologies, it is reasonable to expect that RNA-targeting will emerge as a growing therapeutic strategy for human disorders, especially cancer.
Acknowledgements
Not applicable.
Abbreviations
- lncRNA
Long non-coding RNA
- CGIs
CpG islands
- DNMTs
DNA methyltransferases
- TET
Ten-eleven translocation
- TDG
Thymine DNA glycosylase
- PcG
Polycomb group
- PRC2
Polycomb repressive complex 2
- EZH2
Enhancer of Zeste homolog 2
- GADD45A
Growth arrest and DNA-damage-inducible alpha
- SAM
S-adenosylmethionine
- SAH
S-adenosylhomocysteine
- MAT
Methionine adenosyltransferase
- SAHH
S-adenosylhomocysteine hydrolase
- ceRNA
Competitive endogenous RNA
- BC
Breast cancer
- OC
Oral cancer
- CML
Chronic myeloid leukemia
- TSCC
Tongue squamous cell carcinoma
- TNBC
Triple-negative breast cancer
- EC
Esophageal cancer
- PCa
Prostate cancer
- RC
Renal carcinoma
- GBC
Gallbladder cancer
- HCC
Hepatocellular carcinoma
- OSA
Osteosarcoma
- NSCLC
Non-small cell lung cancer
- LUAD
Lung adenocarcinoma
- CRC
Colorectal cancer
- GC
Gastric cancer
- AML
Acute myeloid leukemia
- LSCC
Laryngeal squamous cell carcinoma
- KS
Kaposi’s sarcoma
- ESCC
Esophageal squamous cell carcinoma
- SCLC
Small-cell lung cancer
- UL
Uterine leiomyomas
- GBM
Glioblastoma multiforme
- CS
Chondrosarcoma
Authors’ contributions
W.H. and H.L. and Q.Y. retrieved literature; W.H. and H.L. wrote the manuscript and prepared the figures; W.X. and D.W. critically revised the manuscript. All authors have read and approved the final manuscript.
Funding
This work was financially supported by the National Natural Science Foundation of China (32000426 and 31971335) to Huang W and Wang DO; the National Key R&D Program of China (2021YFC2009302) to Li H; Special Funds for Transformation and Upgrading of Industrial Informatization of Industry and Information Technology Department of Jiangsu in 2020 to Xiao W; Xingliao Talents Program (XLYC1802007) and Department of Education of Liaoning Province (1911520092) to Wang DO.
Availability of data and materials
Not applicable.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no conflict of interest.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Wanxu Huang and Hua Li contributed equally to this work.
Contributor Information
Wanxu Huang, Email: huangwx@syphu.edu.cn.
Wei Xiao, Email: xw_kanion@163.com.
References
- 1.Greenberg MVC, Bourc'his D. The diverse roles of DNA methylation in mammalian development and disease. Nat Rev Mol Cell Biol. 2019;20:590–607. doi: 10.1038/s41580-019-0159-6. [DOI] [PubMed] [Google Scholar]
- 2.Suzuki MM, Bird A. DNA methylation landscapes: provocative insights from epigenomics. Nat Rev Genet. 2008;9:465–476. doi: 10.1038/nrg2341. [DOI] [PubMed] [Google Scholar]
- 3.Ziller MJ, Gu H, Muller F, Donaghey J, Tsai LT, Kohlbacher O, et al. Charting a dynamic DNA methylation landscape of the human genome. Nature. 2013;500:477–481. doi: 10.1038/nature12433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Deaton AM, Bird A. CpG islands and the regulation of transcription. Genes Dev. 2011;25:1010–1022. doi: 10.1101/gad.2037511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Li E, Zhang Y. DNA methylation in mammals. Cold Spring Harb Perspect Biol. 2014;6:a019133. doi: 10.1101/cshperspect.a019133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Lyko F. The DNA methyltransferase family: a versatile toolkit for epigenetic regulation. Nat Rev Genet. 2018;19:81–92. doi: 10.1038/nrg.2017.80. [DOI] [PubMed] [Google Scholar]
- 7.Wu X, Zhang Y. TET-mediated active DNA demethylation: mechanism, function and beyond. Nat Rev Genet. 2017;18:517–534. doi: 10.1038/nrg.2017.33. [DOI] [PubMed] [Google Scholar]
- 8.Rasmussen KD, Helin K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 2016;30:733–750. doi: 10.1101/gad.276568.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wu SC, Zhang Y. Active DNA demethylation: many roads lead to Rome. Nat Rev Mol Cell Biol. 2010;11:607–620. doi: 10.1038/nrm2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature. 2009;462:315–322. doi: 10.1038/nature08514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Meissner A, Mikkelsen TS, Gu H, Wernig M, Hanna J, Sivachenko A, et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature. 2008;454:766–770. doi: 10.1038/nature07107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhao SG, Chen WS, Li H, Foye A, Zhang M, Sjostrom M, et al. The DNA methylation landscape of advanced prostate cancer. Nat Genet. 2020;52:778–789. doi: 10.1038/s41588-020-0648-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Phillips RE, Soshnev AA, Allis CD. Epigenomic Reprogramming as a Driver of Malignant Glioma. Cancer Cell. 2020;38:647–660. doi: 10.1016/j.ccell.2020.08.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sina AA, Carrascosa LG, Liang Z, Grewal YS, Wardiana A, Shiddiky MJA, et al. Epigenetically reprogrammed methylation landscape drives the DNA self-assembly and serves as a universal cancer biomarker. Nat Commun. 2018;9:4915. doi: 10.1038/s41467-018-07214-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Reddington JP, Sproul D, Meehan RR. DNA methylation reprogramming in cancer: does it act by re-configuring the binding landscape of Polycomb repressive complexes? BioEssays. 2014;36:134–140. doi: 10.1002/bies.201300130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Baylin SB, Jones PA. Epigenetic Determinants of Cancer. Cold Spring Harb Perspect Biol. 2016;8(9):a019505. doi: 10.1101/cshperspect.a019505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Nishiyama A, Nakanishi M. Navigating the DNA methylation landscape of cancer. Trends Genet. 2021;37:1012–1027. doi: 10.1016/j.tig.2021.05.002. [DOI] [PubMed] [Google Scholar]
- 18.Du J, Johnson LM, Jacobsen SE, Patel DJ. DNA methylation pathways and their crosstalk with histone methylation. Nat Rev Mol Cell Biol. 2015;16:519–532. doi: 10.1038/nrm4043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhu H, Wang G, Qian J. Transcription factors as readers and effectors of DNA methylation. Nat Rev Genet. 2016;17:551–565. doi: 10.1038/nrg.2016.83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Heberle E, Bardet AF. Sensitivity of transcription factors to DNA methylation. Essays Biochem. 2019;63:727–741. doi: 10.1042/EBC20190033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Schubeler D. Function and information content of DNA methylation. Nature. 2015;517:321–326. doi: 10.1038/nature14192. [DOI] [PubMed] [Google Scholar]
- 22.Blattler A, Farnham PJ. Cross-talk between site-specific transcription factors and DNA methylation states. J Biol Chem. 2013;288:34287–34294. doi: 10.1074/jbc.R113.512517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, et al. Landscape of transcription in human cells. Nature. 2012;489:101–108. doi: 10.1038/nature11233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zhao L, Wang J, Li Y, Song T, Wu Y, Fang S, et al. NONCODEV6: an updated database dedicated to long non-coding RNA annotation in both animals and plants. Nucleic Acids Res. 2021;49:D165–D171. doi: 10.1093/nar/gkaa1046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Huarte M. The emerging role of lncRNAs in cancer. Nat Med. 2015;21:1253–1261. doi: 10.1038/nm.3981. [DOI] [PubMed] [Google Scholar]
- 26.Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152:1298–1307. doi: 10.1016/j.cell.2013.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Yao RW, Wang Y, Chen LL. Cellular functions of long noncoding RNAs. Nat Cell Biol. 2019;21:542–551. doi: 10.1038/s41556-019-0311-8. [DOI] [PubMed] [Google Scholar]
- 28.Mayer C, Schmitz KM, Li J, Grummt I, Santoro R. Intergenic transcripts regulate the epigenetic state of rRNA genes. Mol Cell. 2006;22:351–361. doi: 10.1016/j.molcel.2006.03.028. [DOI] [PubMed] [Google Scholar]
- 29.Bierhoff H, Schmitz K, Maass F, Ye J, Grummt I. Noncoding transcripts in sense and antisense orientation regulate the epigenetic state of ribosomal RNA genes. Cold Spring Harb Symp Quant Biol. 2010;75:357–364. doi: 10.1101/sqb.2010.75.060. [DOI] [PubMed] [Google Scholar]
- 30.Schmitz KM, Mayer C, Postepska A, Grummt I. Interaction of noncoding RNA with the rDNA promoter mediates recruitment of DNMT3b and silencing of rRNA genes. Genes Dev. 2010;24:2264–2269. doi: 10.1101/gad.590910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Frank S, Ahuja G, Bartsch D, Russ N, Yao W, Kuo JC, et al. yylncT Defines a Class of Divergently Transcribed lncRNAs and Safeguards the T-mediated Mesodermal Commitment of Human PSCs. Cell Stem Cell. 2019;24:318–27 e8. doi: 10.1016/j.stem.2018.11.005. [DOI] [PubMed] [Google Scholar]
- 32.Ponnusamy M, Liu F, Zhang YH, Li RB, Zhai M, Liu F, et al. Long Noncoding RNA CPR (Cardiomyocyte Proliferation Regulator) Regulates Cardiomyocyte Proliferation and Cardiac Repair. Circulation. 2019;139:2668–2684. doi: 10.1161/CIRCULATIONAHA.118.035832. [DOI] [PubMed] [Google Scholar]
- 33.Wang L, Zhao Y, Bao X, Zhu X, Kwok YK, Sun K, et al. LncRNA Dum interacts with Dnmts to regulate Dppa2 expression during myogenic differentiation and muscle regeneration. Cell Res. 2015;25:335–350. doi: 10.1038/cr.2015.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chalei V, Sansom SN, Kong L, Lee S, Montiel JF, Vance KW, et al. The long non-coding RNA Dali is an epigenetic regulator of neural differentiation. Elife. 2014;3:e04530. doi: 10.7554/eLife.04530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yi F, Zhang P, Wang Y, Xu Y, Zhang Z, Ma W, et al. Long non-coding RNA slincRAD functions in methylation regulation during the early stage of mouse adipogenesis. RNA Biol. 2019;16:1401–1413. doi: 10.1080/15476286.2019.1631643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ni C, Jiang W, Wang Z, Wang Z, Zhang J, Zheng X, et al. LncRNA-AC006129.1 reactivates a SOCS3-mediated anti-inflammatory response through DNA methylation-mediated CIC downregulation in schizophrenia. Mol Psychiatry. 2020;26(8):4511–4528. doi: 10.1038/s41380-020-0662-3. [DOI] [PubMed] [Google Scholar]
- 37.Deng Y, Chen D, Gao F, Lv H, Zhang G, Sun X, et al. Silencing of Long Non-coding RNA GAS5 Suppresses Neuron Cell Apoptosis and Nerve Injury in Ischemic Stroke Through Inhibiting DNMT3B-Dependent MAP4K4 Methylation. Transl Stroke Res. 2020;11:950–966. doi: 10.1007/s12975-019-00770-3. [DOI] [PubMed] [Google Scholar]
- 38.Xie Z, Wang Q, Hu S. Coordination of PRKCA/PRKCA-AS1 interplay facilitates DNA methyltransferase 1 recruitment on DNA methylation to affect protein kinase C alpha transcription in mitral valve of rheumatic heart disease. Bioengineered. 2021;12:5904–5915. doi: 10.1080/21655979.2021.1971482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Li H, Han S, Sun Q, Yao Y, Li S, Yuan C, et al. Long non-coding RNA CDKN2B-AS1 reduces inflammatory response and promotes cholesterol efflux in atherosclerosis by inhibiting ADAM10 expression. Aging (Albany NY) 2019;11:1695–1715. doi: 10.18632/aging.101863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wang Y, Yang X, Jiang A, Wang W, Li J, Wen J. Methylation-dependent transcriptional repression of RUNX3 by KCNQ1OT1 regulates mouse cardiac microvascular endothelial cell viability and inflammatory response following myocardial infarction. FASEB J. 2019;33:13145–13160. doi: 10.1096/fj.201900310R. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 41.Chen H, Yang S, Shao R. Long non-coding XIST raises methylation of TIMP-3 promoter to regulate collagen degradation in osteoarthritic chondrocytes after tibial plateau fracture. Arthritis Res Ther. 2019;21:271. doi: 10.1186/s13075-019-2033-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wang Q, Liu J, You Z, Yin Y, Liu L, Kang Y, et al. LncRNA TINCR favors tumorigenesis via STAT3-TINCR-EGFR-feedback loop by recruiting DNMT1 and acting as a competing endogenous RNA in human breast cancer. Cell Death Dis. 2021;12:83. doi: 10.1038/s41419-020-03188-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Su SC, Yeh CM, Lin CW, Hsieh YH, Chuang CY, Tang CH, et al. A novel melatonin-regulated lncRNA suppresses TPA-induced oral cancer cell motility through replenishing PRUNE2 expression. J Pineal Res. 2021;71:e12760. doi: 10.1111/jpi.12760. [DOI] [PubMed] [Google Scholar]
- 44.Song H, Chen L, Liu W, Xu X, Zhou Y, Zhu J, et al. Depleting long noncoding RNA HOTAIR attenuates chronic myelocytic leukemia progression by binding to DNA methyltransferase 1 and inhibiting PTEN gene promoter methylation. Cell Death Dis. 2021;12:440. doi: 10.1038/s41419-021-03637-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Shen T, Xia W, Min S, Yang Z, Cheng L, Wang W, et al. A pair of long intergenic non-coding RNA LINC00887 variants act antagonistically to control Carbonic Anhydrase IX transcription upon hypoxia in tongue squamous carcinoma progression. BMC Biol. 2021;19:192. doi: 10.1186/s12915-021-01112-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Shao G, Fan X, Zhang P, Liu X, Huang L, Ji S. Methylation-dependent MCM6 repression induced by LINC00472 inhibits triple-negative breast cancer metastasis by disturbing the MEK/ERK signaling pathway. Aging (Albany NY) 2021;13:4962–4975. doi: 10.18632/aging.103568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Li N, Zhao Z, Miao F, Cai S, Liu P, Yu Y, et al. Silencing of long non-coding RNA LINC01270 inhibits esophageal cancer progression and enhances chemosensitivity to 5-fluorouracil by mediating GSTP1methylation. Cancer Gene Ther. 2021;28:471–485. doi: 10.1038/s41417-020-00232-1. [DOI] [PubMed] [Google Scholar]
- 48.Zhao Z, Liang S, Sun F. LncRNA DLX6-AS1 Promotes Malignant Phenotype and Lymph Node Metastasis in Prostate Cancer by Inducing LARGE Methylation. Front Oncol. 2020;10:1172. doi: 10.3389/fonc.2020.01172. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 49.Zhang S, Zheng F, Zhang L, Huang Z, Huang X, Pan Z, et al. LncRNA HOTAIR-mediated MTHFR methylation inhibits 5-fluorouracil sensitivity in esophageal cancer cells. J Exp Clin Cancer Res. 2020;39:131. doi: 10.1186/s13046-020-01610-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Liu D, Wu K, Yang Y, Zhu D, Zhang C, Zhao S. Long noncoding RNA ADAMTS9-AS2 suppresses the progression of esophageal cancer by mediating CDH3 promoter methylation. Mol Carcinog. 2020;59:32–44. doi: 10.1002/mc.23126. [DOI] [PubMed] [Google Scholar]
- 51.Li Y, Luo Q, Li Z, Wang Y, Zhu C, Li T, et al. Long Non-coding RNA IRAIN Inhibits VEGFA Expression via Enhancing Its DNA Methylation Leading to Tumor Suppression in Renal Carcinoma. Front Oncol. 2020;10:1082. doi: 10.3389/fonc.2020.01082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Jin L, Cai Q, Wang S, Wang S, Wang J, Quan Z. Long noncoding RNA PVT1 promoted gallbladder cancer proliferation by epigenetically suppressing miR-18b-5p via DNA methylation. Cell Death Dis. 2020;11:871. doi: 10.1038/s41419-020-03080-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wu DM, Zheng ZH, Zhang YB, Fan SH, Zhang ZF, Wang YJ, et al. Down-regulated lncRNA DLX6-AS1 inhibits tumorigenesis through STAT3 signaling pathway by suppressing CADM1 promoter methylation in liver cancer stem cells. J Exp Clin Cancer Res. 2019;38:237. doi: 10.1186/s13046-019-1239-3. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 54.Wang W, Chen G, Wang B, Yuan Z, Liu G, Niu B, et al. Long non-coding RNA BZRAP1-AS1 silencing suppresses tumor angiogenesis in hepatocellular carcinoma by mediating THBS1 methylation. J Transl Med. 2019;17:421. doi: 10.1186/s12967-019-02145-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Qi X, Yu XJ, Wang XM, Song TN, Zhang J, Guo XZ, et al. Knockdown of KCNQ1OT1 Suppresses Cell Invasion and Sensitizes Osteosarcoma Cells to CDDP by Upregulating DNMT1-Mediated Kcnq1 Expression. Mol Ther Nucleic Acids. 2019;17:804–818. doi: 10.1016/j.omtn.2019.06.010. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 56.Miao H, Wang L, Zhan H, Dai J, Chang Y, Wu F, et al. A long noncoding RNA distributed in both nucleus and cytoplasm operates in the PYCARD-regulated apoptosis by coordinating the epigenetic and translational regulation. PLoS Genet. 2019;15:e1008144. doi: 10.1371/journal.pgen.1008144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kang X, Kong F, Huang K, Li L, Li Z, Wang X, et al. LncRNA MIR210HG promotes proliferation and invasion of non-small cell lung cancer by upregulating methylation of CACNA2D2 promoter via binding to DNMT1. Onco Targets Ther. 2019;12:3779–3790. doi: 10.2147/OTT.S189468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Guo X, Chen Z, Zhao L, Cheng D, Song W, Zhang X. Long non-coding RNA-HAGLR suppressed tumor growth of lung adenocarcinoma through epigenetically silencing E2F1. Exp Cell Res. 2019;382:111461. doi: 10.1016/j.yexcr.2019.06.006. [DOI] [PubMed] [Google Scholar]
- 59.Merry CR, Forrest ME, Sabers JN, Beard L, Gao XH, Hatzoglou M, et al. DNMT1-associated long non-coding RNAs regulate global gene expression and DNA methylation in colon cancer. Hum Mol Genet. 2015;24:6240–6253. doi: 10.1093/hmg/ddv343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Somasundaram S, Forrest ME, Moinova H, Cohen A, Varadan V, LaFramboise T, et al. The DNMT1-associated lincRNA DACOR1 reprograms genome-wide DNA methylation in colon cancer. Clin Epigenetics. 2018;10:127. doi: 10.1186/s13148-018-0555-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Xu SF, Zheng Y, Zhang L, Wang P, Niu CM, Wu T, et al. Long Non-coding RNA LINC00628 Interacts Epigenetically with the LAMA3 Promoter and Contributes to Lung Adenocarcinoma. Mol Ther Nucleic Acids. 2019;18:166–182. doi: 10.1016/j.omtn.2019.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Xin L, Lu H, Liu C, Zeng F, Yuan YW, Wu Y, et al. Methionine deficiency promoted mitophagy via lncRNA PVT1-mediated promoter demethylation of BNIP3 in gastric cancer. Int J Biochem Cell Biol. 2021;141:106100. doi: 10.1016/j.biocel.2021.106100. [DOI] [PubMed] [Google Scholar]
- 63.Wang SL, Huang Y, Su R, Yu YY. Silencing long non-coding RNA HOTAIR exerts anti-oncogenic effect on human acute myeloid leukemia via demethylation of HOXA5 by inhibiting Dnmt3b. Cancer Cell Int. 2019;19:114. doi: 10.1186/s12935-019-0808-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Zhao Y, Zhou L, Li H, Sun T, Wen X, Li X, et al. Nuclear-Encoded lncRNA MALAT1 Epigenetically Controls Metabolic Reprogramming in HCC Cells through the Mitophagy Pathway. Mol Ther Nucleic Acids. 2021;23:264–276. doi: 10.1016/j.omtn.2020.09.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zhou W, Xu S, Chen X, Wang C. HOTAIR suppresses PTEN via DNMT3b and confers drug resistance in acute myeloid leukemia. Hematology. 2021;26:170–178. doi: 10.1080/16078454.2021.1880733. [DOI] [PubMed] [Google Scholar]
- 66.Wang X, Yu B, Jin Q, Zhang J, Yan B, Yang L, et al. Regulation of laryngeal squamous cell cancer progression by the lncRNA RP11–159K7.2/miR-206/DNMT3A axis. J Cell Mol Med. 2020;24:6781–95. doi: 10.1111/jcmm.15331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Jin C, Zhao J, Zhang ZP, Wu M, Li J, Xiao GL, et al. Long non-coding RNA GAS5, by up-regulating PRC2 and targeting the promoter methylation of miR-424, suppresses multiple malignant phenotypes of glioma. J Neurooncol. 2020;148:529–543. doi: 10.1007/s11060-020-03544-2. [DOI] [PubMed] [Google Scholar]
- 68.Li W, Wang Q, Feng Q, Wang F, Yan Q, Gao SJ, et al. Oncogenic KSHV-encoded interferon regulatory factor upregulates HMGB2 and CMPK1 expression to promote cell invasion by disrupting a complex lncRNA-OIP5-AS1/miR-218–5p network. PLoS Pathog. 2019;15:e1007578. doi: 10.1371/journal.ppat.1007578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Xu X, Lou Y, Tang J, Teng Y, Zhang Z, Yin Y, et al. The long non-coding RNA Linc-GALH promotes hepatocellular carcinoma metastasis via epigenetically regulating Gankyrin. Cell Death Dis. 2019;10:86. doi: 10.1038/s41419-019-1348-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Yoon JH, You BH, Park CH, Kim YJ, Nam JW, Lee SK. The long noncoding RNA LUCAT1 promotes tumorigenesis by controlling ubiquitination and stability of DNA methyltransferase 1 in esophageal squamous cell carcinoma. Cancer Lett. 2018;417:47–57. doi: 10.1016/j.canlet.2017.12.016. [DOI] [PubMed] [Google Scholar]
- 71.Cheng D, Deng J, Zhang B, He X, Meng Z, Li G, et al. LncRNA HOTAIR epigenetically suppresses miR-122 expression in hepatocellular carcinoma via DNA methylation. EBioMedicine. 2018;36:159–170. doi: 10.1016/j.ebiom.2018.08.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Fang S, Gao H, Tong Y, Yang J, Tang R, Niu Y, et al. Long noncoding RNA-HOTAIR affects chemoresistance by regulating HOXA1 methylation in small cell lung cancer cells. Lab Invest. 2016;96:60–68. doi: 10.1038/labinvest.2015.123. [DOI] [PubMed] [Google Scholar]
- 73.Cao T, Jiang Y, Wang Z, Zhang N, Al-Hendy A, Mamillapalli R, et al. H19 lncRNA identified as a master regulator of genes that drive uterine leiomyomas. Oncogene. 2019;38:5356–5366. doi: 10.1038/s41388-019-0808-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Qi D, Li J, Que B, Su J, Li M, Zhang C, et al. Long non-coding RNA DBCCR1-003 regulate the expression of DBCCR1 via DNMT1 in bladder cancer. Cancer Cell Int. 2016;16:81. doi: 10.1186/s12935-016-0356-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Lai IL, Chang YS, Chan WL, Lee YT, Yen JC, Yang CA, et al. Male-Specific Long Noncoding RNA TTTY15 Inhibits Non-Small Cell Lung Cancer Proliferation and Metastasis via TBX4. Int J Mol Sci. 2019;20(14):3473. doi: 10.3390/ijms20143473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Li Q, Dong C, Cui J, Wang Y, Hong X. Over-expressed lncRNA HOTAIRM1 promotes tumor growth and invasion through up-regulating HOXA1 and sequestering G9a/EZH2/Dnmts away from the HOXA1 gene in glioblastoma multiforme. J Exp Clin Cancer Res. 2018;37:265. doi: 10.1186/s13046-018-0941-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Vennin C, Spruyt N, Robin YM, Chassat T, Le Bourhis X, Adriaenssens E. The long non-coding RNA 91H increases aggressive phenotype of breast cancer cells and up-regulates H19/IGF2 expression through epigenetic modifications. Cancer Lett. 2017;385:198–206. doi: 10.1016/j.canlet.2016.10.023. [DOI] [PubMed] [Google Scholar]
- 78.Fang S, Shen Y, Chen B, Wu Y, Jia L, Li Y, et al. H3K27me3 induces multidrug resistance in small cell lung cancer by affecting HOXA1 DNA methylation via regulation of the lncRNA HOTAIR. Ann Transl Med. 2018;6:440. doi: 10.21037/atm.2018.10.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Xuan Y, Wang Y. Long non-coding RNA SNHG3 promotes progression of gastric cancer by regulating neighboring MED18 gene methylation. Cell Death Dis. 2019;10:694. doi: 10.1038/s41419-019-1940-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Xiong Y, Kuang W, Lu S, Guo H, Wu M, Ye M, et al. Long noncoding RNA HOXB13-AS1 regulates HOXB13 gene methylation by interacting with EZH2 in glioma. Cancer Med. 2018;7:4718–4728. doi: 10.1002/cam4.1718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Zhang C, Wang L, Jin C, Zhou J, Peng C, Wang Y, et al. Long non-coding RNA Lnc-LALC facilitates colorectal cancer liver metastasis via epigenetically silencing LZTS1. Cell Death Dis. 2021;12:224. doi: 10.1038/s41419-021-03461-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Bao X, Ren T, Huang Y, Sun K, Wang S, Liu K, et al. Knockdown of long non-coding RNA HOTAIR increases miR-454–3p by targeting Stat3 and Atg12 to inhibit chondrosarcoma growth. Cell Death Dis. 2017;8:e2605. doi: 10.1038/cddis.2017.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Sui CJ, Zhou YM, Shen WF, Dai BH, Lu JJ, Zhang MF, et al. Long noncoding RNA GIHCG promotes hepatocellular carcinoma progression through epigenetically regulating miR-200b/a/429. J Mol Med (Berl) 2016;94:1281–1296. doi: 10.1007/s00109-016-1442-z. [DOI] [PubMed] [Google Scholar]
- 84.Liu F, Huang W, Hong J, Cai C, Zhang W, Zhang J, et al. Long noncoding RNA LINC00630 promotes radio-resistance by regulating BEX1 gene methylation in colorectal cancer cells. IUBMB Life. 2020;72:1404–1414. doi: 10.1002/iub.2263. [DOI] [PubMed] [Google Scholar]
- 85.Wang L, Bu P, Ai Y, Srinivasan T, Chen HJ, Xiang K, et al. A long non-coding RNA targets microRNA miR-34a to regulate colon cancer stem cell asymmetric division. Elife. 2016;5:e14620. doi: 10.7554/eLife.14620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Wang J, Xie S, Yang J, Xiong H, Jia Y, Zhou Y, et al. The long noncoding RNA H19 promotes tamoxifen resistance in breast cancer via autophagy. J Hematol Oncol. 2019;12:81. doi: 10.1186/s13045-019-0747-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Guo T, Gong C, Wu P, Battaglia-Hsu SF, Feng J, Liu P, et al. LINC00662 promotes hepatocellular carcinoma progression via altering genomic methylation profiles. Cell Death Differ. 2020;27:2191–2205. doi: 10.1038/s41418-020-0494-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Guo T, Wang H, Liu P, Xiao Y, Wu P, Wang Y, et al. SNHG6 Acts as a Genome-Wide Hypomethylation Trigger via Coupling of miR-1297-Mediated S-Adenosylmethionine-Dependent Positive Feedback Loops. Cancer Res. 2018;78:3849–3864. doi: 10.1158/0008-5472.CAN-17-3833. [DOI] [PubMed] [Google Scholar]
- 89.Fu Y, Wang W, Li X, Liu Y, Niu Y, Zhang B, et al. LncRNA H19 interacts with S-adenosylhomocysteine hydrolase to regulate LINE-1 Methylation in human lung-derived cells exposed to Benzo[a]pyrene. Chemosphere. 2018;207:84–90. doi: 10.1016/j.chemosphere.2018.05.048. [DOI] [PubMed] [Google Scholar]
- 90.Chen L, Fan X, Zhu J, Chen X, Liu Y, Zhou H. LncRNA MAGI2-AS3 inhibits the self-renewal of leukaemic stem cells by promoting TET2-dependent DNA demethylation of the LRIG1 promoter in acute myeloid leukaemia. RNA Biol. 2020;17:784–793. doi: 10.1080/15476286.2020.1726637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Wang B, Zhao L, Chi W, Cao H, Cui W, Meng W. Aberrant methylation-mediated downregulation of lncRNA SSTR5-AS1 promotes progression and metastasis of laryngeal squamous cell carcinoma. Epigenetics Chromatin. 2019;12:35. doi: 10.1186/s13072-019-0283-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Xu M, Xu X, Pan B, Chen X, Lin K, Zeng K, et al. LncRNA SATB2-AS1 inhibits tumor metastasis and affects the tumor immune cell microenvironment in colorectal cancer by regulating SATB2. Mol Cancer. 2019;18:135. doi: 10.1186/s12943-019-1063-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Hu S, Yao Y, Hu X, Zhu Y. LncRNA DCST1-AS1 downregulates miR-29b through methylation in glioblastoma (GBM) to promote cancer cell proliferation. Clin Transl Oncol. 2020;22:2230–2235. doi: 10.1007/s12094-020-02363-1. [DOI] [PubMed] [Google Scholar]
- 94.Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature. 2006;439:871–874. doi: 10.1038/nature04431. [DOI] [PubMed] [Google Scholar]
- 95.Zhu X, Du J, Yu J, Guo R, Feng Y, Qiao L, et al. LncRNA NKILA regulates endothelium inflammation by controlling a NF-kappaB/KLF4 positive feedback loop. J Mol Cell Cardiol. 2019;126:60–69. doi: 10.1016/j.yjmcc.2018.11.001. [DOI] [PubMed] [Google Scholar]
- 96.Liu B, Sun L, Liu Q, Gong C, Yao Y, Lv X, et al. A cytoplasmic NF-kappaB interacting long noncoding RNA blocks IkappaB phosphorylation and suppresses breast cancer metastasis. Cancer Cell. 2015;27:370–381. doi: 10.1016/j.ccell.2015.02.004. [DOI] [PubMed] [Google Scholar]
- 97.Artal-Sanz M, Tavernarakis N. Prohibitin and mitochondrial biology. Trends Endocrinol Metab. 2009;20:394–401. doi: 10.1016/j.tem.2009.04.004. [DOI] [PubMed] [Google Scholar]
- 98.Jones R, Wijesinghe S, Wilson C, Halsall J, Liloglou T, Kanhere A. A long intergenic non-coding RNA regulates nuclear localization of DNA methyl transferase-1. iScience. 2021;24:102273. doi: 10.1016/j.isci.2021.102273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Wu H, Zhang Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell. 2014;156:45–68. doi: 10.1016/j.cell.2013.12.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Jia L, Wang Y, Wang C, Du Z, Zhang S, Wen X, et al. Oplr16 serves as a novel chromatin factor to control stem cell fate by modulating pluripotency-specific chromosomal looping and TET2-mediated DNA demethylation. Nucleic Acids Res. 2020;48:3935–3948. doi: 10.1093/nar/gkaa097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Du Z, Wen X, Wang Y, Jia L, Zhang S, Liu Y, et al. Chromatin lncRNA Platr10 controls stem cell pluripotency by coordinating an intrachromosomal regulatory network. Genome Biol. 2021;22:233. doi: 10.1186/s13059-021-02444-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Zhou L, Ren M, Zeng T, Wang W, Wang X, Hu M, et al. TET2-interacting long noncoding RNA promotes active DNA demethylation of the MMP-9 promoter in diabetic wound healing. Cell Death Dis. 2019;10:813. doi: 10.1038/s41419-019-2047-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Wang C, Jia L, Wang Y, Du Z, Zhou L, Wen X, et al. Genome-wide interaction target profiling reveals a novel Peblr20-eRNA activation pathway to control stem cell pluripotency. Theranostics. 2020;10:353–370. doi: 10.7150/thno.39093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Arab K, Park YJ, Lindroth AM, Schafer A, Oakes C, Weichenhan D, et al. Long noncoding RNA TARID directs demethylation and activation of the tumor suppressor TCF21 via GADD45A. Mol Cell. 2014;55:604–614. doi: 10.1016/j.molcel.2014.06.031. [DOI] [PubMed] [Google Scholar]
- 105.Arab K, Karaulanov E, Musheev M, Trnka P, Schafer A, Grummt I, et al. GADD45A binds R-loops and recruits TET1 to CpG island promoters. Nat Genet. 2019;51:217–223. doi: 10.1038/s41588-018-0306-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Canzio D, Nwakeze CL, Horta A, Rajkumar SM, Coffey EL, Duffy EE, et al. Antisense lncRNA Transcription Mediates DNA Demethylation to Drive Stochastic Protocadherin alpha Promoter Choice. Cell. 2019;177:639–53 e15. doi: 10.1016/j.cell.2019.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Lin R, Zhong X, Zhou Y, Geng H, Hu Q, Huang Z, et al. R-loopBase: a knowledgebase for genome-wide R-loop formation and regulation. Nucleic Acids Res. 2021;50(D1):D303–D315. doi: 10.1093/nar/gkab1103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Di Ruscio A, Ebralidze AK, Benoukraf T, Amabile G, Goff LA, Terragni J, et al. DNMT1-interacting RNAs block gene-specific DNA methylation. Nature. 2013;503:371–376. doi: 10.1038/nature12598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Lu SC, Mato JM. S-adenosylmethionine in liver health, injury, and cancer. Physiol Rev. 2012;92:1515–1542. doi: 10.1152/physrev.00047.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Zhou J, Yang L, Zhong T, Mueller M, Men Y, Zhang N, et al. H19 lncRNA alters DNA methylation genome wide by regulating S-adenosylhomocysteine hydrolase. Nat Commun. 2015;6:10221. doi: 10.1038/ncomms10221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Zhou J, Xu J, Zhang L, Liu S, Ma Y, Wen X, et al. Combined Single-Cell Profiling of lncRNAs and Functional Screening Reveals that H19 Is Pivotal for Embryonic Hematopoietic Stem Cell Development. Cell Stem Cell. 2019;24:285–98 e5. doi: 10.1016/j.stem.2018.11.023. [DOI] [PubMed] [Google Scholar]
- 112.Zeng L, Sun S, Han D, Liu Y, Liu H, Feng H, et al. Long non-coding RNA H19/SAHH axis epigenetically regulates odontogenic differentiation of human dental pulp stem cells. Cell Signal. 2018;52:65–73. doi: 10.1016/j.cellsig.2018.08.015. [DOI] [PubMed] [Google Scholar]
- 113.Deng J, Mueller M, Geng T, Shen Y, Liu Y, Hou P, et al. H19 lncRNA alters methylation and expression of Hnf4alpha in the liver of metformin-exposed fetuses. Cell Death Dis. 2017;8:e3175. doi: 10.1038/cddis.2017.392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Spinelli M, Boucard C, Ornaghi S, Schoeberlein A, Irene K, Coman D, et al. Preimplantation factor modulates oligodendrocytes by H19-induced demethylation of NCOR2. JCI Insight. 2021;6(20):e132335. doi: 10.1172/jci.insight.132335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Chen Z, Zheng J, Hong H, Chen D, Deng L, Zhang X, et al. lncRNA HOTAIRM1 promotes osteogenesis of hDFSCs by epigenetically regulating HOXA2 via DNMT1 in vitro. J Cell Physiol. 2020;235:8507–8519. doi: 10.1002/jcp.29695. [DOI] [PubMed] [Google Scholar]
- 116.Li X, Zhang Y, Pei W, Zhang M, Yang H, Zhong M, et al. LncRNA Dnmt3aos regulates Dnmt3a expression leading to aberrant DNA methylation in macrophage polarization. FASEB J. 2020;34:5077–5091. doi: 10.1096/fj.201902379R. [DOI] [PubMed] [Google Scholar]
- 117.Peng WX, Koirala P, Zhang W, Ni C, Wang Z, Yang L, et al. lncRNA RMST Enhances DNMT3 Expression through Interaction with HuR. Mol Ther. 2020;28:9–18. doi: 10.1016/j.ymthe.2019.09.024. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 118.Li MA, Amaral PP, Cheung P, Bergmann JH, Kinoshita M, Kalkan T, et al. A lncRNA fine tunes the dynamics of a cell state transition involving Lin28, let-7 and de novo DNA methylation. Elife. 2017;6:e23468. doi: 10.7554/eLife.23468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Zhang FF, Liu YH, Wang DW, Liu TS, Yang Y, Guo JM, et al. Obesity-induced reduced expression of the lncRNA ROIT impairs insulin transcription by downregulation of Nkx6.1 methylation. Diabetologia. 2020;63:811–24. doi: 10.1007/s00125-020-05090-y. [DOI] [PubMed] [Google Scholar]
- 120.Geng X, Zhao J, Huang J, Li S, Chu W, Wang WS, et al. lnc-MAP3K13-7:1 Inhibits Ovarian GC Proliferation in PCOS via DNMT1 Downregulation-Mediated CDKN1A Promoter Hypomethylation. Mol Ther. 2021;29:1279–1293. doi: 10.1016/j.ymthe.2020.11.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Nishiyama A, Mulholland CB, Bultmann S, Kori S, Endo A, Saeki Y, et al. Two distinct modes of DNMT1 recruitment ensure stable maintenance DNA methylation. Nat Commun. 2020;11:1222. doi: 10.1038/s41467-020-15006-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Petryk N, Bultmann S, Bartke T, Defossez PA. Staying true to yourself: mechanisms of DNA methylation maintenance in mammals. Nucleic Acids Res. 2021;49:3020–3032. doi: 10.1093/nar/gkaa1154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Qin W, Wolf P, Liu N, Link S, Smets M, La Mastra F, et al. DNA methylation requires a DNMT1 ubiquitin interacting motif (UIM) and histone ubiquitination. Cell Res. 2015;25:911–929. doi: 10.1038/cr.2015.72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Liang Y, Peng Y. Gene body methylation facilitates the transcription of CTSG via antisense lncRNA AL136018.1 in dermatomyositic myoideum. Cell Biol Int. 2021;45:456–62. doi: 10.1002/cbin.11508. [DOI] [PubMed] [Google Scholar]
- 125.Hadji F, Boulanger MC, Guay SP, Gaudreault N, Amellah S, Mkannez G, et al. Altered DNA Methylation of Long Noncoding RNA H19 in Calcific Aortic Valve Disease Promotes Mineralization by Silencing NOTCH1. Circulation. 2016;134:1848–1862. doi: 10.1161/CIRCULATIONAHA.116.023116. [DOI] [PubMed] [Google Scholar]
- 126.Yang Z, Xu F, Wang H, Teschendorff AE, Xie F, He Y. Pan-cancer characterization of long non-coding RNA and DNA methylation mediated transcriptional dysregulation. EBioMedicine. 2021;68:103399. doi: 10.1016/j.ebiom.2021.103399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Lu C, Wei Y, Wang X, Zhang Z, Yin J, Li W, et al. DNA-methylation-mediated activating of lncRNA SNHG12 promotes temozolomide resistance in glioblastoma. Mol Cancer. 2020;19:28. doi: 10.1186/s12943-020-1137-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Morenos L, Chatterton Z, Ng JL, Halemba MS, Parkinson-Bates M, Mechinaud F, et al. Hypermethylation and down-regulation of DLEU2 in paediatric acute myeloid leukaemia independent of embedded tumour suppressor miR-15a/16-1. Mol Cancer. 2014;13:123. doi: 10.1186/1476-4598-13-123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329:689–693. doi: 10.1126/science.1192002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Fang H, Bonora G, Lewandowski JP, Thakur J, Filippova GN, Henikoff S, et al. Trans- and cis-acting effects of Firre on epigenetic features of the inactive X chromosome. Nat Commun. 2020;11:6053. doi: 10.1038/s41467-020-19879-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Liu YW, Xia R, Lu K, Xie M, Yang F, Sun M, et al. LincRNAFEZF1-AS1 represses p21 expression to promote gastric cancer proliferation through LSD1-Mediated H3K4me2 demethylation. Mol Cancer. 2017;16:39. doi: 10.1186/s12943-017-0588-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Huang MD, Chen WM, Qi FZ, Sun M, Xu TP, Ma P, et al. Long non-coding RNA TUG1 is up-regulated in hepatocellular carcinoma and promotes cell growth and apoptosis by epigenetically silencing of KLF2. Mol Cancer. 2015;14:165. doi: 10.1186/s12943-015-0431-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Hanly DJ, Esteller M, Berdasco M. Interplay between long non-coding RNAs and epigenetic machinery: emerging targets in cancer? Philos Trans R Soc Lond B Biol Sci. 2018;373(1748):20170074. doi: 10.1098/rstb.2017.0074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011;472:120–124. doi: 10.1038/nature09819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Gu P, Chen X, Xie R, Han J, Xie W, Wang B, et al. lncRNA HOXD-AS1 Regulates Proliferation and Chemo-Resistance of Castration-Resistant Prostate Cancer via Recruiting WDR5. Mol Ther. 2017;25:1959–1973. doi: 10.1016/j.ymthe.2017.04.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Xia M, Liu J, Liu S, Chen K, Lin H, Jiang M, et al. Ash1l and lnc-Smad3 coordinate Smad3 locus accessibility to modulate iTreg polarization and T cell autoimmunity. Nat Commun. 2017;8:15818. doi: 10.1038/ncomms15818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Li Y, Tan Z, Zhang Y, Zhang Z, Hu Q, Liang K, et al. A noncoding RNA modulator potentiates phenylalanine metabolism in mice. Science. 2021;373:662–673. doi: 10.1126/science.aba4991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Warner KD, Hajdin CE, Weeks KM. Principles for targeting RNA with drug-like small molecules. Nat Rev Drug Discov. 2018;17:547–558. doi: 10.1038/nrd.2018.93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Sztuba-Solinska J, Chavez-Calvillo G, Cline SE. Unveiling the druggable RNA targets and small molecule therapeutics. Bioorg Med Chem. 2019;27:2149–2165. doi: 10.1016/j.bmc.2019.03.057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Hargrove AE. Small molecule-RNA targeting: starting with the fundamentals. Chem Commun (Camb) 2020;56:14744–14756. doi: 10.1039/d0cc06796b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Disney MD, Angelbello AJ. Rational Design of Small Molecules Targeting Oncogenic Noncoding RNAs from Sequence. Acc Chem Res. 2016;49:2698–2704. doi: 10.1021/acs.accounts.6b00326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Ren Y, Wang YF, Zhang J, Wang QX, Han L, Mei M, et al. Targeted design and identification of AC1NOD4Q to block activity of HOTAIR by abrogating the scaffold interaction with EZH2. Clin Epigenetics. 2019;11:29. doi: 10.1186/s13148-019-0624-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.