Significance
Understanding autophagy regulation is instrumental in developing therapeutic interventions for autophagy-associated disease. Here, we identified SNAI2 as a regulator of autophagy from a genome-wide screen in HeLa cells. Upon energy stress, SNAI2 is transcriptionally activated by FOXO3 and interacts with FOXO3 to form a feed-forward regulatory loop to reinforce the expression of autophagy genes. Of note, SNAI2-increased FOXO3-DNA binding abrogates CRM1-dependent FOXO3 nuclear export, illuminating a pivotal role of DNA in the nuclear retention of nucleocytoplasmic shuttling proteins. Moreover, a dFoxO-Snail feed-forward loop regulates both autophagy and cell size in Drosophila, suggesting this evolutionarily conserved regulatory loop is engaged in more physiological activities.
Keywords: SNAI2, FOXO3, snail, dFoxO, Drosophila
Abstract
Autophagy is a highly conserved programmed degradation process that regulates a variety of physiological and pathological activities in health, aging, and disease. To identify additional factors that modulate autophagy, we utilized serum-free starvation or Torin1 to induce autophagy in HeLa cells for unbiased mRNA-sequencing analysis and identified SNAI2, a crucial player in epithelial-to-mesenchymal transition and cancer progression, as a regulator of autophagy. Mechanistically, SNAI2 promotes autophagy by physically interacting with FOXO3 and enhancing FOXO3 binding affinity to its response elements in autophagy-related genes. Intriguingly, binding to the DNA targets appears necessary and sufficient for FOXO3 to antagonize its CRM1-dependent nuclear export, illustrating a critical role of DNA in regulating protein nuclear localization. Moreover, stress-elevated SNAI2 expression is mediated by FOXO3, which activates SNAI2 transcription by directly binding to its promoter. Herein, FOXO3 and SNAI2 form a coherent feed-forward regulatory loop to reinforce autophagy genes induction in response to energy stress. Strikingly, a dFoxO-Snail feed-forward circuit also regulates autophagy in Drosophila, suggesting this mechanism is evolutionarily conserved from fly to human.
Autophagy is an evolutionary conserved digestive pathway that captures, degrades, and recycles dysfunctional organelles, intracellular microbes, and pathogenic proteins (1–3). Autophagy occurs in response to stress conditions, such as nutrient deprivation, DNA damage, infection, or hypoxia, to maintain cellular homeostasis (4). Autophagy operates at low baseline levels, and its disruption may result in accumulation of inclusion bodies composed by misfolded protein aggregates and degenerating organelles (5), leading to various diseases (3, 5). Over the past decades, extensive studies have significantly expanded our understanding of the molecular mechanisms underlying autophagy and the roles of autophagy in physiology and pathophysiology (6–8), yet the mechanism that regulates autophagy has not been fully understood.
Since first discovered in Drosophila ∼30 y ago (9), Snail transcription factors (TFs) have been extensively studied as transcriptional repressors to modulate epithelial-to-mesenchymal transition in cancer progression (10–12). However, additional study has also shown that Snail not only represses but also activates gene expression via a specific motif (13). In support of this, an activator role of Snail has been reasonably shown by genetic studies and biochemical assays (14–17). In addition, a convincing body of literature has demonstrated that Snail TFs also play well-characterized roles in development, cell survival, immune regulation, stem cell biology, and metabolism (13, 18–26). Yet, the role of Snail TFs in autophagy remains unknown.
The FoxO TFs represent an evolutionarily conserved family of TFs shuttling between the cytoplasm and nucleus (27, 28). Dysfunction of the shuttling system in response to external stimuli has been implicated in various biological processes, such as metabolism, longevity, and cancer (29–31). Additional studies have also characterized the role of dFoxO in controlling endoplasmic reticulum stress and tumor overgrowth (32, 33). FoxO TFs are principally regulated by two distinct mechanisms: posttranslational modifications and protein–protein interactions (29). Central to the posttranslational modifications is well-studied phosphorylation (34). Mechanistically, phosphorylated FoxOs by a plethora of kinases are reversibly sequestered in the cytoplasm via interaction with 14-3-3 or altered association with the nuclear export complex (29). In addition, the transcriptional activity of FoxO TFs is affected by physical interaction with specific binding partners (29, 35). For example, FANCD2 forms a complex with FOXO3 in response to oxidative stress and increases expression of FOXO3-controlled antioxidant genes for cell survival (36). Yet, little is known about the mechanism that regulates nuclear retention of FoxO TFs in physiological condition.
To identify additional modulators of autophagy, we took advantage of serum deprivation- or Torin1-induced autophagy in HeLa cells for messenger RNA-sequencing (mRNA-seq) analysis and identified that SNAI2, a well-known TF in epithelial-to-mesenchymal transition, plays a critical role in autophagy. Herein, SNAI2 promotes autophagy by concertedly cooperating with FOXO3 to activate expression of PIK3CA and ULK1. Mechanistically, SNAI2 physically interacts with FOXO3 and enhances FOXO3 binding affinity to its response elements in transcriptional target genes. Importantly, binding to the DNA targets appears necessary and sufficient for FOXO3 to antagonize its CRM1-mediated nuclear export, which results in FOXO3 nuclear accumulation. In addition, SNAI2 acts as a transcriptional target of FOXO3 in response to energy stress, thus forming a coherent feed-forward loop with FOXO3 in autophagy induction. Moreover, Snail, the Drosophila ortholog of SNAI2, also regulates autophagy in a feed-forward regulatory circuit with dFoxO, suggesting the function and mechanism of Snail TFs in autophagy are evolutionarily conserved from fly to human.
Results
SNAI2 Functions as a Regulator of Autophagy.
To unearth key factors that regulate autophagy, we utilized extracellularly serum-free (SF) starvation or Torin1, a well-described mammalian target of rapamycin (mTOR)C1/2 inhibitor (37–40), to treat HeLa cells for autophagy induction and subsequently performed RNA-seq and Venn plotting analysis (Fig. 1 A and B and Datasets S1–S3). We identified that mRNA levels of 120 genes were significantly up-regulated compared with the controls (Fig. 1B), among which SNAI2 mRNA was markedly elevated. Given that SNIA2 was of paramount importance to a broad spectrum of physiological and pathophysiological activities, yet the role of Snail TFs in autophagy has not been reported, SNAI2 was selected for further investigation. Reverse transcription-quantitative real-time PCR (RT-qPCR)assay confirmed that Torin1 or SF treatment resulted in increased SNAI2 mRNA (Fig. 1C and SI Appendix, Fig. S1A). Moreover, SNAI2 protein level was up-regulated by Torin1 (Fig. 1D) or another mTOR inhibitor rapamycin (Fig. 1E), implying a potential role of SNAI2 in autophagy. In agreement with this assumption, knockdown of SNAI2 (SI Appendix, Fig. S1B) significantly attenuated Torin1- or rapamycin-induced autophagy in the presence (Fig. 1F and SI Appendix, Fig. S1C) or absence of the lysosome inhibitor Bafilomycin A1 (Baf-A1) or chloroquine (CQ) (Fig. 1G and SI Appendix, Fig. S1D), but exhibited no effect on basal autophagy (SI Appendix, Fig. S1E). To determine whether SNAI2 could promote autophagy, we expressed Flag-tagged SNAI2 in 293T or HeLa cells. We found that enforced SNAI2 expression is not sufficient to trigger autophagy by itself (SI Appendix, Fig. S1 G and H), but was able to enhance autophagy in the presence of Baf-A1 or CQ (SI Appendix, Fig. S1 F and I). In addition, SNAI2 expression promoted autophagy with 2.5-h treatment of rapamycin, while this treatment alone is not sufficient for autophagy induction (Fig. 1H). Taken together, these data suggest that SNAI2 functions is a positive regulator of autophagy, and that SNAI2 may require a cofactor to initiate autophagy.
Fig. 1.
SNAI2 is identified as a positive regulator of autophagy. (A) Heatmap of statistically differential gene expression between dissect control, Torin1-A, Torin1-B treatment, or SF starvation HeLa cells, respectively. Columns in green indicate decreased genes, whereas columns in red demonstrate increased genes. A, 1 μM; B, 250 nM. (B) A Venn diagram shows overlapping up-regulated genes between 1-μM and 250-nM Torin1 treatment, compared with control. (C) RT-qPCR analysis of SNAI2 mRNA level. HeLa cells were treated with DMSO as negative control and 250 nM or 1 μM Torin1 for 4h. (D) Immunoblot analysis of SNAI2 protein level and LC3-II/LC3-I in HeLa cells treated by DMSO or 250nM Torin1 for 4 h. (E) Immunoblot analysis of SNAI2 protein level and LC3-II/LC3-I in 293T cells treated with or without 1 μM rapamycin for 4 h. (F) Immunoblot analysis of 250nM Torin1-induced autophagy in HeLa cells treated by nonspecific siRNA (siCtrl) or two independent siRNA targeting SNAI2 (referred to as siSNAI2-1 and siSNAI2-2). Cells were treated by 20 μM CQ for 24 h to inhibit lysosome activity. (G) Immunoblot analysis of rapamycin-induced autophagy in 293T cells treated by siCtrl or siSNAI2-1. 293T cells were treated by 1 μM Rapamycin for 4 h. (H) Immunoblot analysis of rapamycin-induced autophagy in 293T cells in the absence or presence of transiently transfected Flag-SNAI2 for 48 h. 293T cells were treated by 1 μM rapamycin for 2.5 h before harvest. DMSO acts as negative control for Torin 1 or rapamycin treatment in 293T or HeLa cells. ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05. ns, no significant difference.
SNAI2 Interacts with and Promotes FOXO3-Mediated Autophagy.
Given that SNAI2 encodes a TF, its cofactor in autophagy induction is most likely a TF as well. From large scale yeast two-hybrid (Y2H) assays for protein–protein interactions on nearly all sequence-specific Drosophila TFs (41), we noted an interaction between Snail and dFoxO, the Drosophila orthologs of SNAI2 and FOXO3, respectively. As FoxO family proteins are well-known regulators of autophagy (42–45), we presumed that SNAI2 potentially binds to FOXO3, and promotes FOXO3-mediated autophagy. To test this assumption, we performed coimmunoprecipitation (co-IP) experiments to validate the interaction between SNAI2 and FOXO3 in both 293T and HeLa cells. Intriguingly, FOXO3 specifically interacted with SNAI2, but not SNAI1 and SNAI3 (Fig. 2 A and B). To verify whether SNAI2 cooperates with FOXO3 to induce autophagy, we first confirmed that overexpression of FOXO3 or FOXO33A in 293T cells indeed induced autophagy, as reported previously (42). Intriguingly, FOXO3- or FOXO33A-activated autophagy was significantly augmented upon SNAI2 overexpression (Fig. 2 C and D), and was largely abrogated by SNAI2 knockdown, as indicated by LC3-II/LC3-I (Fig. 2 E–H) and immunofluorescence staining (Fig. 2I). Taken together, the above data indicate that SNAI2 interacts with and promotes FOXO3-mediated autophagy.
Fig. 2.
SNAI2 interacts with and promotes FOXO3-mediated autophagy. (A and B) Immunoblot analysis of FOXO3 in input (also termed as whole-cell lysate, WCL) and anti-Flag immunoprecipitates. The plasmids encoding Flag-SNAI1, Flag-SNAI2, and Flag-SNAI3 were transiently transfected into 293T or HeLa cells for 48 h. (C) Immunoblot analysis of LC3-II/LC3-I in the ectopically expressed Flag-FOXO3 with or without HA-SNAI2 coexpression in 293T cells. (D) Immunofluorescence analysis of LC3 in HeLa cells overexpressing Flag-FOXO3 or Flag-FOXO3A coupled with or without HA-SNAI2 overexpression. (Scale bar, 10 μm.) (E and F) Immunoblot analysis of LC3-II/LC3-I in 293T cells that ectopically expressed Flag-FOXO3 or Flag-FOXO3A with or without SNAI2 knockdown. (G and H) Empty control vector, Flag-FOXO3 or Flag-FOXO33A were transfected into siSNAI2 untreated or treated HeLa cells and immunoblot analysis of LC3-II/LC3-I was performed. (I) Immunofluorescence analysis of LC3 in HeLa cells overexpressing Flag-FOXO3 or Flag-FOXO3A with or without SNAI2 knockdown. (Scale bar, 10 μm.) ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05.
SNAI2/FOXO3 Synergistically Activates Autophagy-Related Genes Transcription.
To investigate the mechanism by which SNAI2 cooperates with FOXO3 to induce autophagy, we performed RT-qPCR analysis to check the expression of FOXO3 target genes involved in autophagy (42, 46–48), and found that FOXO3-activated transcriptions of ULK1 and PIK3CA were further augmented upon SNAI2 expression (Fig. 3A). Intriguingly, SNAI2 also promoted the nuclear localized FOXO33A-activated ULK1 and PIK3CA transcription (Fig. 3B). One possible explanation for this is that SNAI2 may increase FOXO3 binding to its DNA targets in PIK3CA and ULK1. To interrogate this, we carried out chromatin immunoprecipitation (ChIP) experiments and confirmed that FOXO3 could bind to the three known targets in the PIK3CA promoter (42, 49) (Fig. 3 C and D). More importantly, levels of DNA fragments immunoprecipitated by FOXO3 were apparently increased by coexpressing SNAI2 in a dose-dependent manner (Fig. 3D), indicating that SNAI2 enhances FOXO3 binding to its response elements in PIK3CA promoter. Likewise, SNAI2 boosts FOXO3 binding to its targets located in the third intron of ULK1 (Fig. 3 E and F). Taken together, these data suggest that SNAI2 generally promotes FOXO3 binding to its DNA targets in autophagy-related genes.
Fig. 3.
SNAI2/FOXO3 synergistically activate transcription of autophagy genes. (A) RT-qPCR analysis of FOXO3 target genes related to autophagy. Flag-FOXO3 was transfected into 293T cells with or without coexpression of Flag-SNAI2 for 48 h. (B) ULK1 and PIK3CA mRNA level were judged by RT-qPCR analysis. Flag-FOXO33A was transfected into 293T cells with or without Flag-SNAI2. (C) Schematic view of PIK3CA promoter with three FOXO3 binding sites indicated as red bar. Four fragments were shown as amplicons. (D) Relative DNA enrichment in ChIP experiments. (E) Structure of the ULK1 locus. The third intron of ULK1 contains two FOXO3 binding sites indicated as red bar. A and B indicate amplicons used for ChIP. (F) ChIP-qPCR analysis of amplicon-A/B in Flag-IP reactions. (G) Relative DNA enrichment in ChIP experiments was determined by qPCR. (H and I) Analysis of FOXO-luc activity by SNAI2 with or without FOXO33A coexpression in HeLa and 293T cells. ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05. ns, no difference.
Next, we examined the mechanism by which SNAI2 promotes FOXO3-DNA binding, and surmised that SNAI2 may directly bind to its own recognition sites next to FOXO3 binding motifs or engage as a FOXO3-interacting cofactor to indirectly bind to FOXO3 response elements. To discriminate between these two possibilities, we transfected exogenous FOXO-RES, which doesn’t include SNAI2 binding sites, into 293T cells, and then performed ChIP experiments. We found that SNAI2 alone failed to bind FOXO-RES, whereas increased FOXO-RES was enriched by SNAI2 upon FOXO3 coexpression (Fig. 3G). Supporting this, SNAI2 synergistically promoted FOXO33A-activated 4X FOXO-luc activity in both HeLa and 293T cells (Fig. 3 H and I). Cumulatively, these data suggest that SNAI2 acts as a FOXO3-interacting partner, increases FOXO3 binding affinity to its responsive elements and reinforces transcription of autophagy-related genes, such as PIK3CA and ULK1.
Snail–dFoxO Complex Promotes Autophagy in Drosophila.
To investigate whether the SNAI–FOXO complex promotes autophagy in vivo and whether this mechanism is evolutionarily conserved, we checked Drosophila Snail (Sna) and dFoxO, the ortholog of SNAI2 and FOXO3, respectively. Consistent with mammalian data, overexpression of Sna driven by ptc-Gal4 in Drosophila wing imaginal discs induced robust autophagy as measured by both Lyso Tracker red incorporation and Atg8a-pmCherry puncta accumulation (Fig. 4 B and I), two well-described markers for autolysosome and autophagosome, respectively (50, 51). Intriguingly, ectopic Sna-induced autophagy was dramatically suppressed by heterozygous dFoxO mutation (dFoxO△94) or RNAi-mediated dFoxO knockdown (Fig. 4 C, D, J, and K), indicating that Sna modulates dFoxO-dependent autophagy in vivo. Additionally, we found that FOXO33A- and dFoxO-induced autophagy was strikingly antagonized by two independent sna interference RNA (RNAi) (Fig. 4 E–G and L–N and SI Appendix, Fig. S2 A–C), suggesting that FOXO3/dFoxO activity depends on Sna.
Fig. 4.
Sna conservatively promotes autophagy in Drosophila. (A–N) Fluorescence micrographs of Drosophila wing imaginal discs are shown. Lysotracker staining or Atg8a-pmCherry puncta accumulation was performed as autophagy markers. (Scale bar, 20 μm.) (O) A schematic drawing summarizing the binding activities of all Sna and dFoxO fragments. In the right panel, symbols “+” and “−” indicate strong binding or weak binding/no binding, respectively. (P) Immunoblot analysis of Flag-dFoxO and Myc-Sna in input and anti-Flag or anti-Myc immunoprecipitates. Plasmids encoding Flag-dFoxO and Myc-Sna were transiently cotransfected into S2 cells for 48 h before harvest. (Q) Immunoblot analysis of interaction between full-length dFoxO with SnaC but not SnaN. (R) Flag-dFoxOA and Flag-dFoxOB but not Flag-dFoxOC interacts with Myc-SnaC. (S) Flag-dFoxOA+B but not Flag-dFoxOD interacts with Myc-SnaC. (T) Immunoblot analysis of Myc-SnaC in input and anti-Flag immunoprecipitates. Myc-SnaC interacts with Flag-dFoxOE.
Given that Snail and dFoxO are mutually required to trigger autophagy in Drosophila, we checked the physical interaction between Sna and dFoxO. We found that dFoxO reciprocally interacted with Sna as determined by co-IP assay (Fig. 4 O and P). Following extensive mapping experiments, we found that the C-terminal half of Sna (amino acids 246 to 390, referred to as SnaC) is responsible for the interaction with dFoxO (Fig. 4Q). On the other hand, the dFoxO N-terminal fragment (dFoxOA, amino acids 1 to 175) and middle fragment (dFoxOB, amino acids 176 to 445) but not C-terminal one (dFoxOC, amino acids 446 to 622) interacted with SnaC (Fig. 4R), implying the region near A/B boundary is crucial for interaction with Sna. Consistently, the segment of amino acids 155 to 225 (dFoxOE) was necessary and sufficient for dFoxO’s interaction with SnaC (Fig. 4 S and T). The DNA binding motifs in Sna and dFoxO are the zinc finger and forkhead box, respectively. Intriguingly, all five Sna zinc fingers are located in SnaC, and dFoxOE includes part of dFoxO forkhead box, we wondered whether DNA is required for the Sna–dFoxO (SnaC–dFoxOE) interaction. To address this, we digested DNA with DNase I in cell lysates prior to the co-IP experiments. We found that Sna–dFoxO interaction was not affected upon DNase I treatment (SI Appendix, Fig. S3 A and B), suggesting DNA binding is not a prerequisite for the interaction between Sna and dFoxO. In support of this, we performed GST-pull down assays and found direct interactions between dFoxO/FOXO3 and Sna/SNAI2 (SI Appendix, Fig. S3 C and D).
Snail/SNAI2 Conservatively Promotes dFoxO/FOXO3 Nuclear Accumulation.
In support of the direct physical interaction between Snail and dFoxO, we found that Snail, mainly localized in the nucleus, promoted dFoxO nuclear accumulation in S2 cells (Fig. 5A), and in the Drosophila third-instar eye imaginal discs, posterior to the morphogenetic furrow when driven by GMR-Gal4 (Fig. 5B). In addition, we also checked the localization of truncated Snail and dFoxO. As judged by immunostaining analysis, we found that nuclear localized SnaC (NLS-SnaC) was able to restrict dFoxOA+B but not dFoxOD in the nucleus both in vitro and in vivo (SI Appendix, Figs. S4 and S5 A–C). More intriguingly, dFoxOE was sufficient to be sequestered in nucleus by NLS-SnaC (SI Appendix, Fig. S5 D–F), further corroborating the role of Snail–dFoxO interaction in vivo.
Fig. 5.
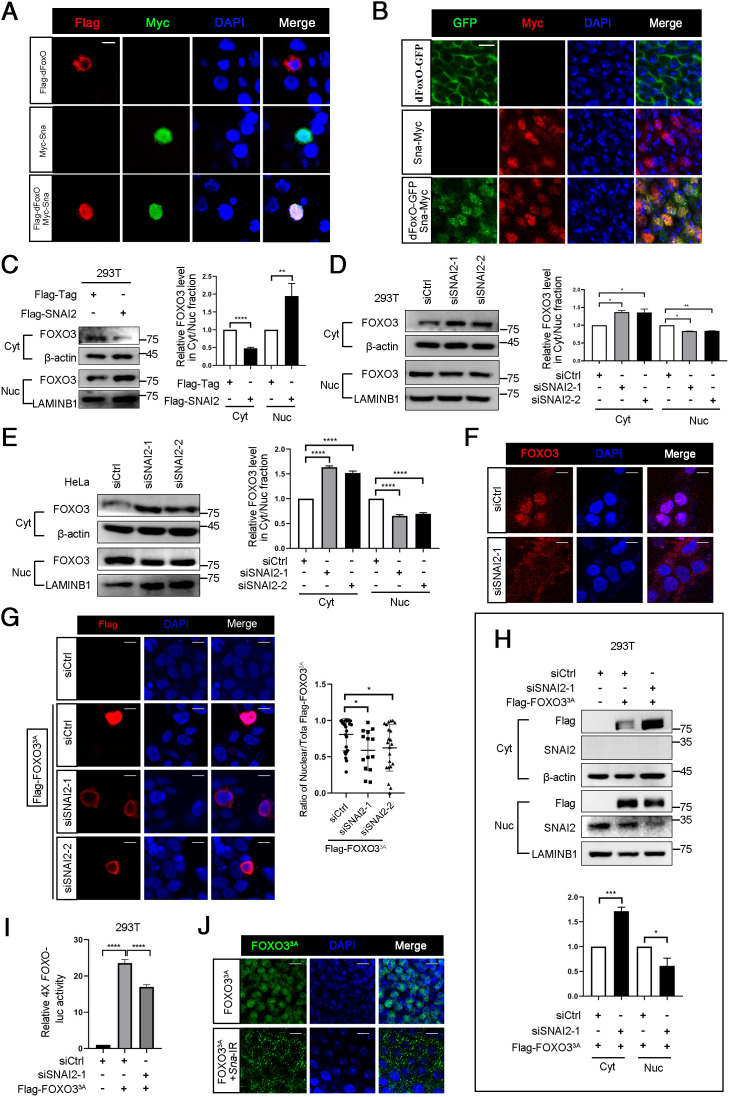
Snail/SNAI2 conservatively promotes nuclear accumulation of dFoxO/FOXO3 from Drosophila to human. (A) Immunofluorescence analysis of Flag-dFoxO cellular localization in S2 cells with or without Myc-Sna overexpression. (Scale bar, 5 μm.) (B) Fluorescence micrographs of Drosophila eye imaginal discs showing subcellular localization of dFoxO-GFP and Sna-Myc. Areas posterior to the morphogenetic furrow are shown. (Scale bar, 5 μm.) (C) Nucleocytoplasmic separation analysis of cellular localization of endogenous FOXO3 in 293T cells transfected by Flag-SNAI2 for 48 h. Cyt, cytoplasm; Nuc, nucleus. (D) Nucleocytoplasmic separation analysis of cellular localization of endogenous FOXO3 in SNAI2 knockdown 293T cells. (E) Cellular localization of endogenous FOXO3 was determined by Nucleocytoplasmic separation analysis in SNAI2 knockdown HeLa cells. (F) Immunofluorescence analysis of FOXO3 cellular localization in SNAI2 knockdown HeLa cells. (Scale bar, 10 μm.) (G) SNAI2 was silenced by siRNA for 24 h and then Flag-FOXO33A was transiently transfected into HeLa cells for another 48 h. Immunofluorescence analysis of Flag-FOXO33A cellular localization change was performed. (Scale bar, 10 μm.) (H) Cellular localization of Flag-FOXO33A in 293T cells was measured by nucleocytoplasmic separation analysis. Flag-FOXO33A was transfected into 293T cells with SNAI2 knockdown by siRNA for 48 h. (I) Relative 4X FOXO-luc activity as measured by Double Luciferase Assay in Flag-FOXO33A overexpressing 293T cells with or without SNAI2 depletion. (J) Immunofluorescence analysis of FOXO33A cellular localization in the presence or absence of Sna in Drosophila eye imaginal discs are shown. (Scale bar, 5 μm.) Areas posterior to the morphogenetic furrow are shown. ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05.
To examine whether Sna-induced dFoxO nuclear accumulation is evolutionarily conserved, we checked the subcellular localization of SNAI2 and FOXO3 in mammalian cells. In agreement with the fly data, SNAI2 overexpression increased FOXO3 protein level in nucleus but decreased that in cytoplasm as measured by nuclei-cytoplasm fractionation (Fig. 5C), while FOXO3 subcellular localization was not altered upon SNAI1 or SNAI3 expression (SI Appendix, Fig. S6A). Moreover, knockdown of SNAI2 significantly increased FOXO3 in cytoplasm but decreased FOXO3 in nucleus in 293T and HeLa cells (Fig. 5 D and E), which were further confirmed by immunofluorescence staining (Fig. 5F). In support of this, SNAI2 ablation potently attenuated the activity of a FOXO-luc reporter harboring four FOXO binding sites in the promoter region (SI Appendix, Fig. S6 B and C). To rule out the possibility that SNAI2 regulates FOXO3 localization by altering 14-3-3 expression, we performed immunoblotting assay and found that altered SNAI2 expression did not affected total or individual 14-3-3 protein level (SI Appendix, Fig. S6 D–F).
To further verify the effect of SNAI2 on FOXO3 subcellular localization, we transfected into HeLa cells an active FOXO3 (Flag-FOXO33A), in which all three phosphorylation residues were mutated to alanine residues to mimic a nonphosphorylated state, and thus could not be arrested in cytoplasm by 14-3-3. Intriguingly, the nuclear localized Flag-FOXO33A was shuttled to cytoplasm upon SNAI2 knockdown as measured by immunofluorescence (Fig. 5G) and nuclei-cytoplasm fractionation (Fig. 5H). Consistently, Flag-FOXO33A–induced FOXO-luc activity was significantly attenuated by SNAI2 depletion (Fig. 5I). Furthermore, expression of FOXO33A by GMR-Gal4 in Drosophila eye imaginal discs also exhibited nuclear localization, but was shuttled to cytoplasm upon Snail (Sna) removal (Fig. 5J). Cumulatively, these data suggest that Snail/SNAI2 conservatively promote dFoxO/FOXO3 nuclear accumulation, which are consistent with previous observations that nuclear proteins could induce nuclear accumulation of nucleocytoplasmic shuttling proteins via direct interaction (52–54).
SNAI2 Inhibits FOXO3 Nuclear Export by Enhancing FOXO3-DNA Binding.
We next sought to dissect the mechanism by which SNAI2 regulates FOXO3 nuclear accumulation. It’s well-described that nuclear sequestration of a protein usually results from increased nuclear import or decreased nuclear export (52). As a constantly nuclear localized protein (The Human Protein Atlas, https://www.proteinatlas.org/), SNAI2 is more likely to reduce the nuclear export, rather than enhance the nuclear import, of FOXO3, which shuttles frequently between cytoplasm and nucleus. Consistent with previous findings that nuclear exportation of FOXO3 depends on CRM1 (29), we confirmed in a co-IP assay that CRM1 indeed could interact with both ectopically expressed and endogenous FOXO3 (Fig. 6 A–C). Intriguingly, ectopic SNAI2 abrogated the interaction between FOXO3 and CRM1 in both 293T and HeLa cells (Fig. 6 A–C), suggesting that SNAI2 promotes FOXO3 nuclear accumulation by impeding CRM1-dependent FOXO3 nuclear export. As a nucleocytoplasmic shuttling protein, nuclear FOXO3 could bind to either DNA targets for transcription or CRM1 for nuclear export. In this scenery, FOXO3 is competed by DNA and CRM1, while SNAI2 may impede FOXO3–CRM1 interaction by enhancing FOXO3-DNA binding, as in the case of PIK3CA and ULK1 (Fig. 3 D, F, and G).
Fig. 6.
SNAI2 impedes FOXO3-CRM1 interaction via enhancing FOXO3-DNA binding. (A–C) Co-IP analysis of CRM1-FOXO3 interaction with or without coexpression of SNAI2 in 293T or HeLa cells. (D) Analysis of DNase I treatment on SNAI2-reduced interaction between CRM1 and FOXO3 in 293T cells. (E and F) Effects of exogenous FOXO-luc vector or synthetic DNA fragment containing 4X FOXO3 responsive element sequence (FOXO-RES) on CRM1–FOXO3 interaction in 293T or HeLa cells. Ctrl-luc vector or Ctrl-RES fragment without FOXO target sequence serves as the negative control for FOXO-luc or FOXO-RES, respectively. Ctr-RES: 5′-GGGGGGCTATAAAAGGGGGTGGGGGCGTTCGTCCTCACTCT-3′; FOXO-RES: 5′-CTCGATGATCAAGTAAACAACTATGTAAACAAGATCAAGTAAACAACTATGTAAACAAGCGCG-3′.
To test this hypothesis, we used DNase I to digest DNA prior to the co-IP experiments, and found that FOXO3–CRM1 interaction was no longer impeded by ectopic SNAI2 upon DNase I treatment (Fig. 6D), suggesting that DNA binding may be a prerequisite for SNAI2 to block FOXO3–CRM1 interaction. Intriguingly, FOXO3–CRM1 interaction was significantly increased upon DNase I treatment (Fig. 6D), consistent with the assumption that DNA competes against CRM1 for FOXO3 binding. To examine whether increased FOXO3 targeting to DNA could sufficiently reduce FOXO3–CRM1 interaction, we transfected exogenous luciferase plasmids into 293T cells. Compared with the control plasmid (Ctrl-luc), FOXO-luc plasmid harboring 4X FOXO3 binding motifs (4X FOXO-luc) effectively blocked FOXO3–CRM1 association (Fig. 6E). Consistently, FOXO-luc vector largely abrogated CRM1 interaction with exogenously expressed FOXO33A (SI Appendix, Fig. S7S). To rule out the impact of luciferase protein in above experiments, we transfected synthetic DNA fragment containing 4X FREs (FOXO-RES) into HeLa cells and 293T cells, and confirmed that FOXO-RES was able to block the interaction between FOXO3 and CRM1, compared with the control fragment (Fig. 6F and SI Appendix, Fig. S7B). Collectively, these data suggest that DNA binding is essential for FOXO3 to antagonize its CRM1-mediated nuclear export, and that SNAI2 promotes FOXO3 nuclear retention by enhancing FOXO3 binding to its DNA targets.
FOXO3 Mediates Extracellular Stress Induced Up-Regulation of SNAI2.
We then sought to illustrate the upstream mechanism by which extracellular stress increase SNAI2 expression (Fig. 1 C–E). It has been shown that coherent feed-forward loop is one of the best-studied transcription regulation networks that control the expression of genes (55). In addition, as previously reported, dephosphorylated FOXO3 would translocate into the nucleus in response to energy limitation (28, 56–59). Hence, it is plausible that FOXO3 as a TF may directly activate SNAI2 expression. To test this possibility, we first checked whether Torin1-induced SNAI2 expression depends on FOXO3. As judged by RT-qPCR and Western blot analysis, FOXO3 knockdown significantly antagonized Torin1-induced SNAI2 up-regulation at both the mRNA and protein level (Fig. 7 A and B). Similarly, FOXO3 depletion also suppressed rapamycin-induced SNAI2 expression (SI Appendix, Fig. S8A). Moreover, overexpression of Flag-FOXO33A markedly elevated SNAI2 mRNA and protein level as measured by RT-qPCR and Western blot assay, respectively (Fig. 7 C and D). These data suggest that FOXO3 is necessary and sufficient for drug-induced SNAI2 expression.
Fig. 7.
FOXO3 mediates energy stress-induced SNAI2 up-regulation. (A) RT-qPCR analysis of SNAI2 mRNA level. FOXO3 knockdown or control HeLa cells were subject to 0.25-μM Torin1 treatment for 4 h. (B) Immunoblot analysis of SNAI2 protein level in FOXO3 knockdown or control HeLa cells with or without 0.25-μM Torin1 treatment for 4 h. (C) RT-qPCR analysis of SNAI2 mRNA level in HeLa and 293T cells with or without Flag-FOXO33A expression. (D) Immunoblot analysis of SNAI2 protein level. Control or Flag-FOXO33A was transfected into 293T or HeLa cells for 48 h. (E) Schematic view of SNAI2 promoter. The red bar represents presumptive FOXO3 binding sites, the green fragment shows the ChIP assay target regions. E1 and E2 (deleting core sequence) were fragments used to drive luciferase reporters in Double Luciferase Assay. (F) Relative luciferase activity driven by E1 or E2 in the absence or presence of FOXO33A. (G) Relative DNA enrichment in ChIP experiments was determined by qPCR. (H) RT-qPCR analysis of SNAI2 mRNA level in response to normal or SF medium for 4 h. (I) Immunoblot analysis of SNAI2 protein level in FOXO3 knockdown or control HeLa cells with or without SF medium treatment for 4 h. (J) RT-qPCR analysis of sna mRNA in ptc > dFoxO or FOXO33A fly wing imaginal discs. ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05.
To investigate whether FOXO3 directly promotes SNAI2 transcription, we examined the SNAI2 cis-regulatory region and identified a putative FOXO3 binding motif (ATAAAC) about −1.8 kb in the upstream region (Fig. 7E). To verify this motif, we generated two firefly luciferase reporters driven by the E1 fragment harboring the motif or E2 fragment deleting the core sequence-ATAAAC from E1. Luciferase expression driven by E1 was markedly up-regulated by Flag-FOXO33A, which was significantly abrogated by deleting the core sequence in E2, as evidenced by double luciferase analysis in 293T and HeLa cells (Fig. 7F and SI Appendix, Fig. S8B). Meanwhile, we performed a ChIP experiment and found that region A containing the potential binding site but not region B as a negative control was appreciably enriched by FOXO3 (Fig. 7G and SI Appendix, Fig. S8C). Collectively, these data suggest that FOXO3 was the key factor for drug-induced SNAI2 expression via direct transcriptional regulation.
To mimic the physiological stress that cells may encounter in vivo, we used SF medium to induce starvation in HeLa cells. Starvation obviously increased SNAI2 expression at both the mRNA and protein levels (Fig. 7H and SI Appendix, Figs. S1A), which was significantly retarded by FOXO3 knockdown (Fig. 7I). Taken together, these data suggest that, in response to starvation stress, FOXO3 activates the expression of SNAI2, which collaborates with FOXO3 in a coherent feed-forward regulatory loop to reinforce transcription of autophagy-related genes (Fig. 8).
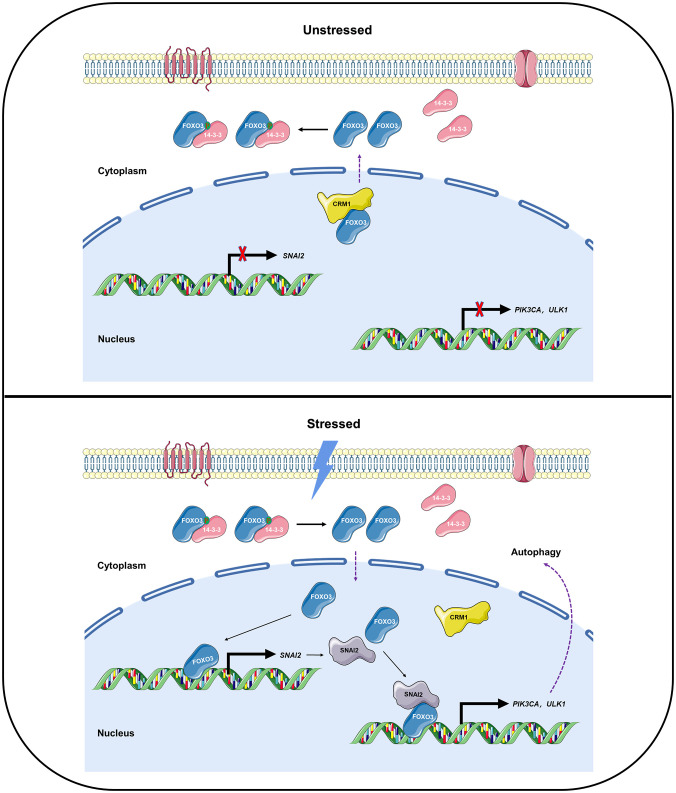
Fig. 8.
Model for a conserved FOXO3-SNAI2 feed-forward loop in autophagy. Under normal conditions (unstressed), CRM1-dependent nuclear exported-FOXO3 is phosphorylated and then sequestered in cytoplasm via interaction with 14-3-3. Upon energy stress conditions like starvation (Stressed), dephosphorylated FOXO3 translocates into nucleus and activates SNAI2 transcription. Then SNAI2 interacts with and enhances FOXO3 binding affinity to response elements in autophagy-related genes like PIK3CA and ULK1 for autophagy induction. More notably, the mechanism underlying SNAI2-decreased FOXO3 nuclear export lies in increased FOXO3-DNA binding in the nucleus. Hence, we propose the model of a coherent FOXO3-SNAI2 feed-forward loop in autophagy, which is conserved from Drosophila to human.
Moreover, consistent with the foregoing finding that FOXO3 activates SNAI2 transcription in mammalian cells, overexpression of dFoxO or FOXO33A in Drosophila wing imaginal discs driven by ptc-Gal4 also significantly increased sna transcription (Fig. 7J), suggesting the dFoxO/FOXO3-Snail/SNAI2 feed-forward regulatory loop is evolutionarily conserved from fly to human.
Snail Functions as a General Cofactor of dFoxO.
Accumulated evidence has shown that the FoxO TFs are involved in a broad spectrum of activities, such as serving as a tumor suppressor (60, 61), cell size, and autophagy regulators (42, 44, 62). It is intriguing that Sna also modulates cell size in Drosophila wing imaginal discs (12), but the mechanism is yet to be determined. To ascertain whether Sna-induced cell size reduction depends on dFoxO, we produced clones in wing margin bristles. Compared with controls (SI Appendix, Fig. S2F), Sna-expressing bristles showed reduced size (SI Appendix, Fig. S2 G and K), which was significantly ameliorated in heterozygous dFoxO mutants (SI Appendix, Fig. S2 H and K). Consistent with previous study, overexpression of dFoxO apparently decreased bristle size (SI Appendix, Fig. S2 I and K), which was significantly rescued by knockdown of Sna (SI Appendix, Fig. S2 J and K). To interrogate the target gene responsible for dFoxO/Snail-mediated cell size control, we checked the expression of 4E-BP, a known transcriptional target of dFoxO and an inhibitor of cell growth (63, 64). We found that 4E-BP expression was up-regulated by expressing Sna or dFoxO, and synergistically by expressing both, while Dp100 served as a negative control (SI Appendix, Fig. S2L). In support of this, Sna cooperated with dFoxO to promote the activity of 4E-BP promoter-luc (SI Appendix, Fig. S2E), which harbors dFoxO binding site and serves as a reporter for dFoxO transcriptional activity (65, 66). Collectively, these data suggest Snail may act as a general cofactor of dFoxO.
Discussion
The autophagy degradation pathway plays a fundamental role in organism homeostasis via sequestering dangerous cargo, such as large protein aggregates and entire damaged organelles into autolysosome for digestion, and its dysfunction has been implicated in various human diseases (3, 5, 67, 68). Over the past decades, extraordinary advances have been made to understand the initiation and regulation of autophagy, yet the regulatory mechanism of this important process has not been fully elucidated. In our present study, we performed SF starvation or Torin1 treatment as autophagy inducer in HeLa cells, exploited RNA-seq analysis and identified SNAI2 as a crucial regulator of autophagy. We found that SNAI2 expression was significantly up-regulated at both mRNA and protein levels during autophagy induction. Interestingly, SNAI2 was not only required but also sufficient to promote autophagy. Mechanistically, SNAI2 specifically interacts with and tethers FOXO3 in the nucleus to augment the expression of target genes, such as ULK1 and PIK3CA. Of note, SNAI2 enhances the binding affinity of FOXO3 to its DNA target sites, which blocks FOXO3–CRM1 interaction for nuclear export.
In addition, we provide evidence that FOXO3 mediates the induction of SNAI2 by starvation, Torin1, or rapamycin treatment. Further investigations show that FOXO3 directly activates the transcription of SNAI2 by binding to its promoter. Thus, we conclude that FOXO3 forms a positive feed-forward loop with SNAI2 to reinforce expression of autophagy target genes, like PIK3CA and ULK1 in response to energy stress. Moreover, the activity of SNAI2 has been retained by its Drosophila ortholog Snail, which facilitates dFoxO activity by enhancing its nuclear localization. In addition, ptc > dFoxO also activates sna transcription in fly wing imaginal discs, thus forming an evolutionarily conserved feed-forward gene-regulatory circuit for autophagy induction in Drosophila. Given that there’s yet no report about the role of Snail TFs in autophagy regulation, our findings provide evidence that, as a dFoxO or FOXO3 target gene, Snail or SNAI2 acts in a feed-forward loop to promote dFoxO/FOXO3-mediated autophagy from fly to human. In addition, we found that CRM1-mediated FOXO3 nuclear export was impeded by FOXO3-bound DNA, which supports the role of DNA in regulating the ins and outs of shuttling proteins (69, 70). Snail-enhanced FOXO-DNA binding may cause spatial segregation of FOXO from CRM1, or interfere with FOXO–CRM1 interaction, thus decreasing FOXO nuclear-cytoplasmic shuttling and increasing its nuclear accumulation.
While SNAI2 expression alone does not affect basal autophagy in 293T and HeLa cells (SI Appendix, Fig. S1 G and H), ectopic Sna in Drosophila wing imaginal discs is able to trigger autophagosome formation (Fig. 4 B and I). As Snail regulates autophagy via FoxOs, this discrepancy might be caused by different expression levels of the endogenous FoxO proteins in mammalian cells and Drosophila. Indeed, FOXO3 level is low in 293T and HeLa cells (https://www.proteinatlas.org/), whereas endogenous dFoxO expression is high in fly imaginal discs (https://flybase.org/). To test this assumption, we found from the Human Protein Atlas database that HSkCM cells show relatively high FOXO3 level (SI Appendix, Fig. S9A). We performed RT-qPCR and Western blot experiments to confirm that FOXO3 expression in HSkCM cells is indeed higher than that in HeLa cells (SI Appendix, Fig. S9 B and C). Consistent with our hypothesis, SNAI2 overexpression in HSkCM cells sufficed to induce autophagy as measured by LC3-II/LC3-I (SI Appendix, Fig. S9D), supporting SNAI2 as a positive regulator of autophagy.
While dFoxO is the only fly FOXO gene, four FOXO genes, including FOXO1, FOXO3, FOXO4, and FOXO6, were found in mammals (28). Both FOXO1 and FOXO3 have been shown to regulate autophagy. Consistently, we found FOXO1 and FOXO3, but not FOXO4 and FOXO6, were up-regulated about twofold upon Torin1 treatment or SF starvation (Datasets S1–S3). In our study, FOXO3 not only promotes SNAI2 transcription but also interacts with SNAI2 in both 293T and HeLa cells. To test whether any other FOXOs is involved in the regulation of SNAI2, we performed co-IP and RT-qPCR experiments. We found that only FOXO3 but not FOXO1/4/6 could interact with SNAI2 in 293T and HeLa cells (SI Appendix, Fig. S10A). In addition, FOXO1/4/6 were not able to activate SNAI2 transcription in 293T and HeLa cells (SI Appendix, Fig. S10 B–D), suggesting that SNAI2 is specifically regulated by FOXO3.
Extensive studies have demonstrated that FoxO TFs, mainly functioning under the control of insulin and insulin-like peptide signaling, are phosphorylated by the kinase Akt, the main downstream mediators of the signaling, and are retained in cytoplasm by 14-3-3 (27, 71, 72). It’s well known that FoxO TFs are involved in the regulation of diverse processes, including development, metabolism, stem cell maintenance, and longevity (44, 73). Central to the regulation of FoxO activities is the shuttling system between the cytoplasm and nucleus (27, 59, 74, 75), hence, identifying novel regulators of FoxO TFs subcellular localization will provide additional approach to modulate FoxO-related pathological disorders. In current study, we present evidence that Snail/SNAI2 is an evolutionarily conserved regulator of dFoxO/FOXO3 shuttling system. Further investigation is needed to explore whether SNAI2 is involved in other physiological and pathological functions of FOXO3.
Materials and Methods
Expression Constructs.
To generate Flag-dFoxO, Flag-dFoxOA, Flag-dFoxOB, Flag-dFoxOC, Flag-dFoxOD, Flag-dFoxOE, Myc-Snail, SnailC-Myc, Myc-SnailC, Myc-SnailN, and HA-Snail, we amplified the corresponding complementary DNA (cDNA) fragments using Q5 DNA polymerase (New England Biolabs, M0491L), and cloned them into the pUAST-Myc, pUAST-Flag, or pUAST-HA backbone vector. For expression in human cells, constructs including Flag-SNAI1, Flag-SNAI2, Flag-SNAI3, and HA-SNAI2 were amplified using HeLa cell cDNA as template and subcloned into the p3xFlag-Myc-CMV or pCDNA3.0-HA vector. Plasmids encoding Flag-FOXO3WT and Flag-FOXO33A, were gifts from Weiguo Zhu, Peking University, Beijing, China. Plasmids, including pCMV-mFoxo1, pFlag-FOXO4 and pFlag-FOXO6 were obtained from Bio-Research Innovation Center Suzhou, China. All of the constructs generated were confirmed by DNA sequencing.
Cell Culture and Transfection.
S2 cells were cultured in Drosophila Schneider’s Medium (Gibco, 21720) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 100 U/mL penicillin, and 100 μg/mL streptomycin (Invitrogen) at 25 °C in a humidified air atmosphere. Human 293T, HeLa, and HSkCM cells were maintained at 37 °C in DMEM (Gibco) containing 10% FBS, 100 U/mL penicillin, and 100 μg/mL streptomycin in a humidified incubator with 5% CO2. Transfection of S2 cells was performed with the Effectene Transfection Reagent (Qiagen, 301427) according to the manufacturer’s instructions. For 293T and HeLa cells, we utilized Lipofectamine 3000 (Invitrogen, L3000015) for DNA plasmids transfection and Lipofectamine RNAiMAX (Invitrogen, 13778150) for small-interfering RNA (siRNA) transfection following the instructions. For SNAI2 knockdown, a mixture of two different siRNAs was used. The sequences are as follows:
siRNA-SNAI2-1: 5′-GAAUGUCUCUCCUGCACAATT-3′(sense);
siRNA-SNAI2-1: 5′-UUGUGCAGGAGAGACAUUCTT-3′(antisense);
siRNA-SNAI2-2: 5′-GCGCCCUGAAGAUGCAUAUTT-3′ (sense);
siRNA-SNAI2-2: 5′-AUAUGCAUCUUCAGGGCGCTT-3′ (antisense);
siRNA-Control: 5′-UUCUCCGAACGUGUCACGUTT-3′(sense);
siRNA-Control: 5′-ACGUGACACGUUCGGAGAATT-3′ (antisense).
Immunofluorescence and LysoTracker Red Staining.
Cells were washed three times with ice-cold PBS and were fixed for 15 min at room temperature with Immunol Staining Fix Solution (Beyotime, P0098) and then permeabilized with Immunostaining Permeabilization Buffer with Triton X-100 (Beyotime, P0096) for 20 min. Following permeabilization, nonspecific binding in the cells was blocked by Immunol Staining Blocking Buffer (Beyotime, P0102) for 1 h at room temperature. Then cells were incubated for 1 h with specific primary antibodies and after three washes with PBS, the cells were incubated for another 1h with secondary antibodies.
Lysosomal activity as a marker of autophagy in Drosophila wing imaginal discs was detected by the LysoTracker Red Kit (Beyotime, C1046). Imaginal discs dissected from third-instar larvae were collected in PBS and incubated with LysoTracker Red (1:3,000) for 15 min at 37 °C, washed with PBS three times prior to imaging. Primary antibodies included: rabbit anti-FoxO3A (1:200; Abcam, ab12162), mouse anti–Myc-tag (1:350, CST, 9B11), rabbit anti-FLAG antibody (1:200, Sigma). Secondary antibodies were anti-mouse CY3 (1:1,000) and anti-rabbit Alexa Flour 488 (1:500). All images were collected with a confocal microscope (Zeiss LSM 780).
Supplementary Material
Acknowledgments
We thank the Bloomington Drosophila Stock Center, the Vienna Drosophila RNAi Center, and Core Facility of Drosophila Resource and Technology, the Center for Excellence in Molecular Cell Science, the Chinese Academy of Science for fly stocks; Dr. Weiguo Zhu for the plasmids; and members of the L.X. laboratory for discussion and critical comments. This work is supported by National Natural Science Foundation of China Grant 31970536 (to L.X.), Grant 32000547 (to X.G.), and Grant 81771957 (to L.L.); Shanghai Committee of Science and Technology grants 09DZ2260100 and 19MC1910300 (to L.X.); China Postdoctoral Science Foundation Grant 2000229071 (to X.G.); and Shanghai Shenkang Three Years Action Project SHDC2020CR2054B (to Z. Lv).
Footnotes
The authors declare no competing interest.
This article is a PNAS Direct Submission.
Data Availability
All study data are included in the main text and supporting information.
References
- 1.Mizushima N., Autophagy: Process and function. Genes Dev. 21, 2861–2873 (2007). [DOI] [PubMed] [Google Scholar]
- 2.Klionsky D. J., Autophagy: From phenomenology to molecular understanding in less than a decade. Nat. Rev. Mol. Cell Biol. 8, 931–937 (2007). [DOI] [PubMed] [Google Scholar]
- 3.López-Otín C., Kroemer G., Hallmarks of health. Cell 184, 33–63 (2021). [DOI] [PubMed] [Google Scholar]
- 4.Klionsky D. J., et al. , Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Levine B., Kroemer G., Biological functions of autophagy genes: A disease perspective. Cell 176, 11–42 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dikic I., Elazar Z., Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 19, 349–364 (2018). [DOI] [PubMed] [Google Scholar]
- 7.Galluzzi L., Pietrocola F., Levine B., Kroemer G., Metabolic control of autophagy. Cell 159, 1263–1276 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mizushima N., A brief history of autophagy from cell biology to physiology and disease. Nat. Cell Biol. 20, 521–527 (2018). [DOI] [PubMed] [Google Scholar]
- 9.Alberga A., Boulay J. L., Kempe E., Dennefeld C., Haenlin M., The snail gene required for mesoderm formation in Drosophila is expressed dynamically in derivatives of all three germ layers. Development 111, 983–992 (1991). [DOI] [PubMed] [Google Scholar]
- 10.Barrallo-Gimeno A., Nieto M. A., The Snail genes as inducers of cell movement and survival: Implications in development and cancer. Development 132, 3151–3161 (2005). [DOI] [PubMed] [Google Scholar]
- 11.Campbell K., et al. , Collective cell migration and metastases induced by an epithelial-to-mesenchymal transition in Drosophila intestinal tumors. Nat. Commun. 10, 2311 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Campbell K., Lebreton G., Franch-Marro X., Casanova J., Differential roles of the Drosophila EMT-inducing transcription factors Snail and Serpent in driving primary tumour growth. PLoS Genet. 14, e1007167 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rembold M., et al. , A conserved role for Snail as a potentiator of active transcription. Genes Dev. 28, 167–181 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Reece-Hoyes J. S., et al. , The C. elegans Snail homolog CES-1 can activate gene expression in vivo and share targets with bHLH transcription factors. Nucleic Acids Res. 37, 3689–3698 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zeitlinger J., et al. , Whole-genome ChIP-chip analysis of Dorsal, Twist, and Snail suggests integration of diverse patterning processes in the Drosophila embryo. Genes Dev. 21, 385–390 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tao G., Levay A. K., Gridley T., Lincoln J., Mmp15 is a direct target of Snai1 during endothelial to mesenchymal transformation and endocardial cushion development. Dev. Biol. 359, 209–221 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Sakai D., Suzuki T., Osumi N., Wakamatsu Y., Cooperative action of Sox9, Snail2 and PKA signaling in early neural crest development. Development 133, 1323–1333 (2006). [DOI] [PubMed] [Google Scholar]
- 18.Vega S., et al. , Snail blocks the cell cycle and confers resistance to cell death. Genes Dev. 18, 1131–1143 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Inoue A., et al. , Slug, a highly conserved zinc finger transcriptional repressor, protects hematopoietic progenitor cells from radiation-induced apoptosis in vivo. Cancer Cell 2, 279–288 (2002). [DOI] [PubMed] [Google Scholar]
- 20.Kudo-Saito C., Shirako H., Takeuchi T., Kawakami Y., Cancer metastasis is accelerated through immunosuppression during Snail-induced EMT of cancer cells. Cancer Cell 15, 195–206 (2009). [DOI] [PubMed] [Google Scholar]
- 21.Lyons J. G., et al. , Snail up-regulates proinflammatory mediators and inhibits differentiation in oral keratinocytes. Cancer Res. 68, 4525–4530 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kurrey N. K., et al. , Snail and slug mediate radioresistance and chemoresistance by antagonizing p53-mediated apoptosis and acquiring a stem-like phenotype in ovarian cancer cells. Stem Cells 27, 2059–2068 (2009). [DOI] [PubMed] [Google Scholar]
- 23.Zeng J., Huynh N., Phelps B., King-Jones K., Snail synchronizes endocycling in a TOR-dependent manner to coordinate entry and escape from endoreplication pausing during the Drosophila critical weight checkpoint. PLoS Biol. 18, e3000609 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Liu Y., Bao H., Wang W., Lim H. Y., Cardiac Snail family of transcription factors directs systemic lipid metabolism in Drosophila. PLoS Genet. 15, e1008487 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cowden J., Levine M., The Snail repressor positions Notch signaling in the Drosophila embryo. Development 129, 1785–1793 (2002). [DOI] [PubMed] [Google Scholar]
- 26.Zinzen R. P., Senger K., Levine M., Papatsenko D., Computational models for neurogenic gene expression in the Drosophila embryo. Curr. Biol. 16, 1358–1365 (2006). [DOI] [PubMed] [Google Scholar]
- 27.Van Der Heide L. P., Hoekman M. F., Smidt M. P., The ins and outs of FoxO shuttling: Mechanisms of FoxO translocation and transcriptional regulation. Biochem. J. 380, 297–309 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Calnan D. R., Brunet A., The FoxO code. Oncogene 27, 2276–2288 (2008). [DOI] [PubMed] [Google Scholar]
- 29.Boccitto M., Kalb R. G., Regulation of Foxo-dependent transcription by post-translational modifications. Curr. Drug Targets 12, 1303–1310 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Tia N., et al. , Role of Forkhead Box O (FOXO) transcription factor in aging and diseases. Gene 648, 97–105 (2018). [DOI] [PubMed] [Google Scholar]
- 31.Hwangbo D. S., Gershman B., Tu M. P., Palmer M., Tatar M., Drosophila dFOXO controls lifespan and regulates insulin signalling in brain and fat body. Nature 429, 562–566 (2004). [DOI] [PubMed] [Google Scholar]
- 32.Nowak K., Gupta A., Stocker H., FoxO restricts growth and differentiation of cells with elevated TORC1 activity under nutrient restriction. PLoS Genet. 14, e1007347 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gupta A., Stocker H., FoxO suppresses endoplasmic reticulum stress to inhibit growth of Tsc1-deficient tissues under nutrient restriction. eLife 9, e53159 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zhao Y., Wang Y., Zhu W. G., Applications of post-translational modifications of FoxO family proteins in biological functions. J. Mol. Cell Biol. 3, 276–282 (2011). [DOI] [PubMed] [Google Scholar]
- 35.Shats I., et al. , FOXO transcription factors control E2F1 transcriptional specificity and apoptotic function. Cancer Res. 73, 6056–6067 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Li J., Du W., Maynard S., Andreassen P. R., Pang Q., Oxidative stress-specific interaction between FANCD2 and FOXO3a. Blood 115, 1545–1548 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Xu C., et al. , SIRT1 is downregulated by autophagy in senescence and ageing. Nat. Cell Biol. 22, 1170–1179 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dong X., et al. , Sorting nexin 5 mediates virus-induced autophagy and immunity. Nature 589, 456–461 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Koundouros N., et al. , Metabolic fingerprinting links oncogenic PIK3CA with enhanced arachidonic acid-derived eicosanoids. Cell 181, 1596–1611.e27 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Tiessen I., et al. , A high-throughput screen identifies the long non-coding RNA DRAIC as a regulator of autophagy. Oncogene 38, 5127–5141 (2019). [DOI] [PubMed] [Google Scholar]
- 41.Shokri L., et al. , A comprehensive Drosophila melanogaster transcription factor interactome. Cell Rep. 27, 955–970.e7 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Zhou J., et al. , FOXO3 induces FOXO1-dependent autophagy by activating the AKT1 signaling pathway. Autophagy 8, 1712–1723 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Cai J., et al. , CK1α suppresses lung tumour growth by stabilizing PTEN and inducing autophagy. Nat. Cell Biol. 20, 465–478 (2018). [DOI] [PubMed] [Google Scholar]
- 44.Zhao Y., et al. , Cytosolic FoxO1 is essential for the induction of autophagy and tumour suppressor activity. Nat. Cell Biol. 12, 665–675 (2010). [DOI] [PubMed] [Google Scholar]
- 45.Tas D., et al. , Parallel roles of transcription factors dFOXO and FER2 in the development and maintenance of dopaminergic neurons. PLoS Genet. 14, e1007271 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zhang J., et al. , Histone deacetylase inhibitors induce autophagy through FOXO1-dependent pathways. Autophagy 11, 629–642 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zhang L., Li J., Ouyang L., Liu B., Cheng Y., Unraveling the roles of Atg4 proteases from autophagy modulation to targeted cancer therapy. Cancer Lett. 373, 19–26 (2016). [DOI] [PubMed] [Google Scholar]
- 48.Cheng Z., The FoxO-autophagy axis in health and disease. Trends Endocrinol. Metab. 30, 658–671 (2019). [DOI] [PubMed] [Google Scholar]
- 49.Hui R. C., et al. , The forkhead transcription factor FOXO3a increases phosphoinositide-3 kinase/Akt activity in drug-resistant leukemic cells through induction of PIK3CA expression. Mol. Cell. Biol. 28, 5886–5898 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Mauvezin C., Ayala C., Braden C. R., Kim J., Neufeld T. P., Assays to monitor autophagy in Drosophila. Methods 68, 134–139 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Zhuang L., et al. , CHIP modulates APP-induced autophagy-dependent pathological symptoms in Drosophila. Aging Cell 19, e13070 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Fang L., et al. , SET1A-mediated mono-methylation at K342 regulates YAP activation by blocking its nuclear export and promotes tumorigenesis. Cancer Cell 34, 103–118.e9 (2018). [DOI] [PubMed] [Google Scholar]
- 53.Zhang L., et al. , The TEAD/TEF family of transcription factor Scalloped mediates Hippo signaling in organ size control. Dev. Cell 14, 377–387 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Cho Y. S., et al. , Regulation of Yki/Yap subcellular localization and Hpo signaling by a nuclear kinase PRP4K. Nat. Commun. 9, 1657 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Alon U., Network motifs: Theory and experimental approaches. Nat. Rev. Genet. 8, 450–461 (2007). [DOI] [PubMed] [Google Scholar]
- 56.Brunet A., et al. , Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell 96, 857–868 (1999). [DOI] [PubMed] [Google Scholar]
- 57.Schmeisser K., Parker J. A., Pleiotropic effects of mTOR and autophagy during development and aging. Front. Cell Dev. Biol. 7, 192 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Dwyer D. S., Horton R. Y., Aamodt E. J., Role of the evolutionarily conserved starvation response in anorexia nervosa. Mol. Psychiatry 16, 595–603 (2011). [DOI] [PubMed] [Google Scholar]
- 59.Bülow M. H., Bülow T. R., Hoch M., Pankratz M. J., Jünger M. A., Src tyrosine kinase signaling antagonizes nuclear localization of FOXO and inhibits its transcription factor activity. Sci. Rep. 4, 4048 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Greer E. L., Brunet A., FOXO transcription factors at the interface between longevity and tumor suppression. Oncogene 24, 7410–7425 (2005). [DOI] [PubMed] [Google Scholar]
- 61.Burgering B. M., Kops G. J., Cell cycle and death control: Long live Forkheads. Trends Biochem. Sci. 27, 352–360 (2002). [DOI] [PubMed] [Google Scholar]
- 62.Mukherjee S., Duttaroy A., Spargel/dPGC-1 is a new downstream effector in the insulin-TOR signaling pathway in Drosophila. Genetics 195, 433–441 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Miron M., et al. , The translational inhibitor 4E-BP is an effector of PI(3)K/Akt signalling and cell growth in Drosophila. Nat. Cell Biol. 3, 596–601 (2001). [DOI] [PubMed] [Google Scholar]
- 64.Demontis F., Perrimon N., FOXO/4E-BP signaling in Drosophila muscles regulates organism-wide proteostasis during aging. Cell 143, 813–825 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Tettweiler G., Miron M., Jenkins M., Sonenberg N., Lasko P. F., Starvation and oxidative stress resistance in Drosophila are mediated through the eIF4E-binding protein, d4E-BP. Genes Dev. 19, 1840–1843 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Jünger M. A., et al. , The Drosophila forkhead transcription factor FOXO mediates the reduction in cell number associated with reduced insulin signaling. J. Biol. 2, 20 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Mizushima N., Komatsu M., Autophagy: Renovation of cells and tissues. Cell 147, 728–741 (2011). [DOI] [PubMed] [Google Scholar]
- 68.Amaravadi R., Kimmelman A. C., White E., Recent insights into the function of autophagy in cancer. Genes Dev. 30, 1913–1930 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.McBride K. M., Reich N. C., The ins and outs of STAT1 nuclear transport. Sci. STKE 2003, RE13 (2003). [DOI] [PubMed] [Google Scholar]
- 70.McBride K. M., McDonald C., Reich N. C., Nuclear export signal located within theDNA-binding domain of the STAT1transcription factor. EMBO J. 19, 6196–6206 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Dobson M., et al. , Bimodal regulation of FoxO3 by AKT and 14-3-3. Biochim. Biophys. Acta 1813, 1453–1464 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Nielsen M. D., Luo X., Biteau B., Syverson K., Jasper H., 14-3-3 Epsilon antagonizes FoxO to control growth, apoptosis and longevity in Drosophila. Aging Cell 7, 688–699 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Bai H., Kang P., Hernandez A. M., Tatar M., Activin signaling targeted by insulin/dFOXO regulates aging and muscle proteostasis in Drosophila. PLoS Genet. 9, e1003941 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Puig O., Mattila J., Understanding Forkhead box class O function: Lessons from Drosophila melanogaster. Antioxid. Redox Signal. 14, 635–647 (2011). [DOI] [PubMed] [Google Scholar]
- 75.Latré de Laté P., et al. , Glucocorticoid-induced leucine zipper (GILZ) promotes the nuclear exclusion of FOXO3 in a Crm1-dependent manner. J. Biol. Chem. 285, 5594–5605 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All study data are included in the main text and supporting information.