ABSTRACT
Technical advances in metagenomics and metatranscriptomics have dramatically accelerated virus discovery in recent years. “Chuviruses” were first described in 2015 as obscure negative-sense RNA viruses of diverse arthropods. Although “chuviruses” first appeared to be members of the negarnaviricot order Mononegavirales in phylogenetic analyses using RNA-directed RNA polymerase sequences, further characterization revealed unusual gene orders in genomes that are nonsegmented, segmented, and/or possibly circular. Consequently, a separate order, Jingchuvirales, was established to include a monospecific family, Chuviridae. Recently, it has become apparent that jingchuvirals are broadly distributed and are therefore likely of ecological and economic importance. Here, we describe recent and ongoing efforts to create the necessary taxonomic framework to accommodate the expected flood of novel viruses belonging to the order.
KEYWORDS: Chuviridae, chuvirus, Jingchuvirales, mivirus, virus classification, virus nomenclature, virus taxonomy
INTRODUCTION
Viruses are polyphyletic mobile genetic elements that have been classified into six realms and several lower unassigned taxa (1–5). The realm Riboviria includes all taxa for viruses encoding RNA-directed RNA polymerases (RdRps) (kingdom Orthornavirae) or reverse transcriptases (RTs) (kingdom Pararnavirae). These two hallmark enzymes are used for phylogenetic megataxonomic assignments (2, 6–8). Specifically, RdRp-encoding negative-sense RNA viruses all belong to the orthornaviran phylum Negarnaviricota, which includes two subphyla, Haploviricotina (e.g., Aspiviridae and Mononegavirales) and Polyploviricotina (e.g., Bunyavirales and Orthomyxoviridae) (2, 6–8).
In 2015, a metagenomic study aiming at the discovery of novel RNA viruses in arthropods detected a novel clade of haploviricotines in chelicerates, crustaceans, and insects sampled in China (9). These viruses, designated “chuviruses” after the historic Chǔ (楚) region in China (roughly the area where the sampled arthropods were collected), clustered with mononegavirals in RdRp phylogenies (9). Mononegavirals typically have nonsegmented RNA genomes; only two genera, rhabdovirid Dichorhavirus and Varicosavirus, harbor viruses with bisegmented genomes. In addition, mononegaviral genomes have five canonical genes that encode structural proteins that are mostly organized in the same order (albeit often with interspersed additional genes), 3′-N-P-M-(G)-L-5′, encoding nucleoprotein, polymerase cofactor, matrix protein, glycoprotein, and large protein with its RdRp domain, respectively (10). On the other hand, “chuvirus” genomes typically encode only two to three genes arranged in a different order, (G)-(x)-N-(x)-L or L-(G)-N-(x) (with x being one or several open reading frames encoding unknown proteins), and are circular nonsegmented (type I) (e.g., Tǎchéng tick virus 4), circular bisegmented (type II) (e.g., Wēnzhōu crab virus 2), or linear nonsegmented or segmented (type III) (e.g., Wēnlǐng crustacean virus 15 and Lampyris noctiluca chuvirus-like virus 1, respectively) RNAs (9) (Fig. 1). This unusual arrangement (and the first evidence that single-stranded RNA viruses may have circular genomes) justified the establishment of a sister order to Mononegavirales in one class, Monjiviricetes. This new order, named Jingchuvirales (Jīngchǔ [荆楚] is a synonym for Chǔ in Chinese), initially included one family, Chuviridae, with a single genus, Mivirus (named after Mǐ [芈], the name of a king of Chǔ), for 30 species (7).
FIG 1.
Genome organization of representatives of each of the established five jingchuviral families. Genomes are drawn to scale (3′→5′). L, large protein gene; HP, hypothetical protein gene; N, nucleoprotein gene; G, glycoprotein gene.
During recent years, it has become clear that jingchuvirals are not exotic but highly diverse and likely globally distributed in arthropods of all major subphyla (Chelicerata, Crustacea, Hexapoda, and Myriapoda) as well as being associated with eucestodes, fish, mammals, reptiles, and sea anemones (11–48). In addition, jingchuvirals were detected in platyhelminths (30), cnidaria (40), and nematodes (27), suggesting that these viruses are broadly distributed and that numerous new, related viruses will be discovered once intense sampling of animals of diverse phyla commences. A recent preprint indicates that jingchuvirals also may infect humans and even be possibly associated with febrile disease (49). Finally, endogenous jingchuviral-like elements (sequences of jingchuviral origin integrated into host genomes) were discovered in numerous nonhuman genomes (e.g., crustaceans; coleopteran, dipteran, isopteran, hemipteran, hymenopteran, and lepidopteran insects; and myriapods) (9, 50, 51), indicating that these viruses have interacted with their hosts for eons.
METHODS AND RESULTS
The International Committee on Taxonomy of Viruses (ICTV) Animal dsRNA and ssRNA Viruses Subcommittee recently established the ICTV Jingchuvirales Study Group (https://talk.ictvonline.org) to create a more structured framework for jingchuviral classification.
As a first step, the genome sequences of the originally classified 30 miviruses were collected from the National Center for Biotechnology Information (NCBI) GenBank database. Existing taxonomic classifiers (as defined by the NCBI) were then used for the identification of related sequences, their metadata, as well as the publication source(s). Using this approach, over 100 L gene sequences were retrieved. Sequences were initially aligned using ClustalW (52), refined using Multiple Sequence Comparison by Log Expectation (MUSCLE) (53), and manually curated in Geneious R9 (54). ProtTest (55) was used to estimate the best amino acid substitution model for the data set (LG [Le Gascuel matrix] + I [invariant sites] + Γ [gamma distribution] + F [using observed amino acid frequencies]). The tree was estimated using PhyML 3.0 (56), a subtree pruning and regrafting (SPR) topology searching algorithm, and a Bayes branch support algorithm.
Database entries suggesting incomplete genomes (defined here as genomes consisting of less than L- and N-encoding fragments or segments), containing apparently misassembled genomes (e.g., missing transcriptional initiation and/or termination sequences, obvious duplication of long stretches of identical sequences, clearly fragmented open reading frames [ORFs], or missing ORFs), or lacking host or geographical metadata were manually removed from the data set. A combination of L protein amino acid identity distribution (Fig. 2) and phylogenetic topology based on maximum likelihood tree estimations (Fig. 3) was then used to create taxon demarcation criteria within the order Jingchuvirales.
FIG 2.
The distribution of L protein percent identities guides the taxonomical demarcation of jingchuvirals. New jingchuviral families, genera, and species should be established when the amino acid identities of two complete L sequences are <21%, <31%, and <90%, respectively. Demarcations were originally established on the basis of all available coding-complete “chu-like” RdRp sequences in GenBank on 3 March 2020 (57). Jingchuviral classification strategies follow previous taxonomical classification efforts that used pairwise sequencing comparison methods (59–64).
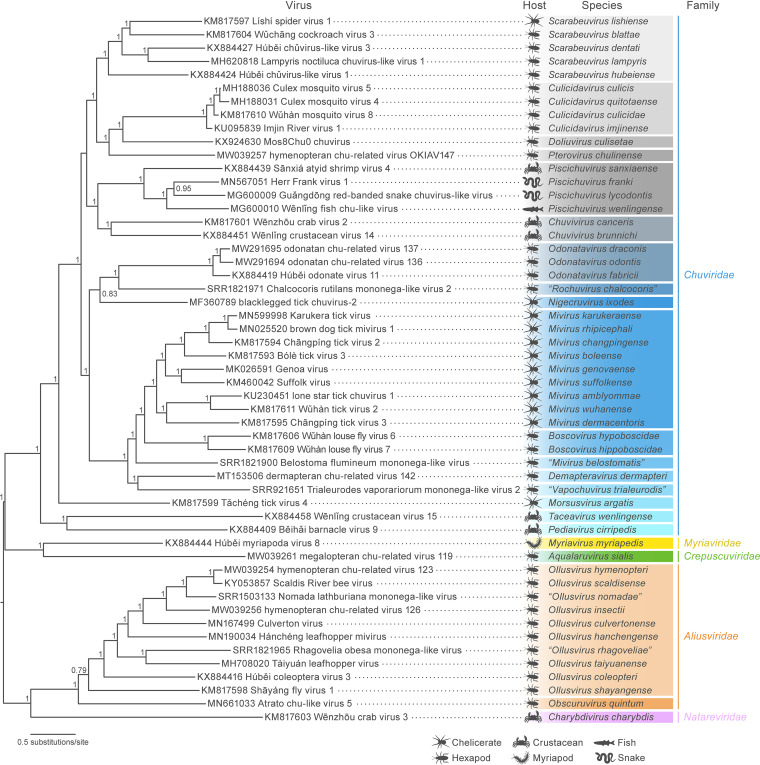
FIG 3.
Maximum likelihood phylogenetic tree estimated using 53 complete jingchuviral L protein sequences. Tree branches are scaled by substitutions per site. Support values are shown as decimal values and were calculated from 500 replicates.
This first analysis resulted in the ICTV-accepted 2021 reorganization of the order to include 5 families, 19 genera, and 48 species (Fig. 3 and Table 1) (4, 57). In early 2021, this analysis was repeated, and a taxonomic proposal (TaxoProp) was submitted to the ICTV to expand the order (Fig. 3 and Table 1) (58). L gene nucleotide and L amino acid residue identities across all jingchuvirals of this analysis are listed in Tables S1 and S2 in the supplemental material. All jingchuviral families and genera that include more than one species clustered with well-supported nodes. In contrast, all currently monospecific families/genera are associated with long branch lengths due to high sequence diversity and a lack of sampling.
TABLE 1.
ICTV-accepted taxonomy of the monjiviricete order Jingchuvirales (Riboviria: Orthornavirae: Negarnaviricota: Haploviricotina) as of March 2021a and newly proposed changes currently under ICTV considerationb
| Family and genus | Species | Virus (abbreviation) | Presumed host(s) | Reference(s) |
|---|---|---|---|---|
| Aliusviridae | ||||
| Obscuruvirus | Obscuruvirus quintum | Atrato chu-like virus 5 (AClV-5) | Culicid mosquito (Psorophora albipes (Theobald, 1907)) | Unpublished |
| Ollusvirus | Ollusvirus coleopteri | Húběi coleoptera virus 3 (HbCV-3) | Unspecified beetle (Coleoptera sp.) | 27 |
| Ollusvirus culvertonense | Culverton virus (CvV) | Hercules’ flea (Macropsylla hercules Rothschild, 1905) | 11 | |
| Ollusvirus hanchengense | Hánchéng leafhopper mivirus (HLMV) | Leafhopper (Psammotettix alienus (Dahlbom, 1850)) | 12 | |
| Ollusvirus hymenopteri | hymenopteran chu-related virus 123 (HCrV-123) | Megachilid bee (Dioxys cincta (Jurine, 1807)) | 13 | |
| Ollusvirus insectii | hymenopteran chu-related virus 126 (HCrV-126) | Crabronid wasp (Oxybelus bipunctatus Olivier, 1812) | 13 | |
| “Ollusvirus nomadae” | Nomada lathburiana mononega-like virus (NLmlV) | Apid bee (Nomada lathburiana (Kirby, 1802)) | Unpublished, 25 | |
| “Ollusvirus rhagoveliae” | Rhagovelia obesa mononega-like virus (ROmlV) | Veliid water strider (Rhagovelia obesa Uhler, 1871) | Unpublished, 25 | |
| Ollusvirus scaldisense | Scaldis River bee virus (SRBV) | European orchard bee (Osmia cornuta (Latreille, 1805)) | 14 | |
| Ollusvirus shayangense | Shāyáng fly virus 1 (SyFV-1) | Pepper fruit fly (Atherigona orientalis Schiner, 1868) | 9 | |
| Ollusvirus taiyuanense | Tàiyuán leafhopper virus (TYLeV) | Leafhopper (Psammotettix alienus (Dahlbom, 1850)) | 15 | |
| Chuviridae | ||||
| Boscovirus | Boscovirus hippoboscidae | Wǔhàn louse fly virus 7 (WhLFV-7) | Unspecific louse fly (Hippoboscidae sp.) | 9 |
| Boscovirus hypoboscidae | Wǔhàn louse fly virus 6 (WhLFV-6) | Unspecific louse fly (Hippoboscidae sp.) | 9 | |
| Chuvivirus | Chuvivirus brunnichi | Wēnlǐng crustacean virus 14 (WlCV-14) | Asian paddle crab (Charybdis japonica (A. Milne-Edwards, 1861)) | 9 |
| Chuvivirus canceris | Wēnzhōu crab virus 2 (WzCV-2) | Unspecified crustacean | 27 | |
| Culicidavirus | Culicidavirus culicidae | Wǔhàn mosquito virus 8 (WhMV-8) | Culicid mosquito (Culex tritaeniorhynchus Giles, 1901) | 9 |
| Culicidavirus culicis | Culex mosquito virus 5 (ClMV-5) | Culicid mosquito (Culex tritaeniorhynchus Giles, 1901) | 16 | |
| Culicidavirus imjinense | Imjin River virus 1 (IjRV-1) | Culicid mosquito (Culex bitaeniorhynchus Giles, 1901) | 17 | |
| Culicidavirus quitotaense | Culex mosquito virus 4 (ClMV-4) | Culicid mosquito (Culex tritaeniorhynchus Giles, 1901) | 16 | |
| Demapteravirus | Demapteravirus dermapteri | dermapteran chu-related virus 142 (DCrV-142) | Anisolabid earwig (Gonolabis marginalis (Dohrn, 1864)) | 13 |
| Doliuvirus | Doliuvirus culisetae | Mos8Chu0 chuvirus (MoCV) | Culicid mosquito (Culiseta minnesotae Barr, 1957) | Unpublished |
| Mivirus | “Mivirus belostomatis” | Belostoma flumineum mononega-like virus (BFmlV) | Giant water bug (Belostoma flumineum Say, 1832) | Unpublished, 25 |
| Mivirus boleense | Bólè tick virus 3 (BTV-3) | Ixodid tick (Hyalomma asiaticum Schülze and Schlottke, 1929) | 9 | |
| Mivirus changpingense | Chāngpíng tick virus 2 (CpTV-2) | Ixodid tick (Dermacentor sp.) | 9 | |
| Mivirus dermacentoris | Chāngpíng tick virus 3 (CpTV-3) | Ixodid tick (Dermacentor sp.) | 9 | |
| Tǎchéng tick virus 5 (TcTV-5) | Ixodid tick (Dermacentor marginatus Sulzer, 1776) | 9 | ||
| Mivirus genovaense | Genoa virus (GeV) | Australian paralysis tick (Ixodes holocyclus Neumann, 1899) | 18 | |
| Mivirus karukeraense | Karukera tick virus (KtV) | Ixodid ticks {Asian blue tick [Rhipicephalus microplus (Canestrini, 1888)] and tropical bont tick (Amblyomma variegatum Fabricius, 1794)} | 19 | |
| Mivirus rhipicephali | brown dog tick mivirus 1 (BDTMV-1) | Brown dog tick (Rhipicephalus sanguineus (Latreille, 1806)) | 20 | |
| Mivirus amblyommae | lone star tick chuvirus 1 (LSTCV-1) | Lone star tick (Amblyomma americanum (Linnaeus, 1758)) | 21 | |
| Mivirus suffolkense | Suffolk virus (SFKV) | Deer tick (Ixodes scapularis Say, 1821) | 21 | |
| Mivirus wuhanense | Wǔhàn tick virus 2 (WhTV-2) | Asian blue tick (Rhipicephalus microplus (Canestrini, 1888)) | 9 | |
| Morsusvirus | Morsusvirus argatis | Tǎchéng tick virus 4 (TcTV-4) | Ixodid tick (Argas miniatus Koch, 1844) | 9 |
| Nigecruvirus | Nigecruvirus ixodes | blacklegged tick chuvirus 2 (BlTC-2) | Deer tick (Ixodes scapularis Say, 1821) | 21 |
| Odonatavirus | Odonatavirus draconis | odonatan chu-related virus 137 (OCrV-137) | Common flatwing (Austroargiolestes icteromelas (Selys, 1862)) | 13 |
| Odonatavirus fabricii | Húběi odonate virus 11 (HbOV-11) | Unspecified dragonfly/damselfly (Odonata sp.) | 27 | |
| Odonatavirus odontis | odonatan chu-related virus 136 (OCrV-136) | Whitewater rockmaster (Diphlebia lestoides (Selys, 1853)) | 13 | |
| Pediavirus | Pediavirus cirripedis | Běihǎi barnacle virus 9 (BhBV-9) | Unspecified barnacle (Cirripedia sp.) | 27 |
| Piscichuvirus | Piscichuvirus franki | Herr Frank virus 1 (HFrV-1) | Boa constrictor (Boa constrictor Linnaeus, 1758) | 23 |
| Piscichuvirus lycodontis | Guǎngdōng red-banded snake chuvirus-like virus (GRSCV) | Colubrid snake (Lycodon rufozonatus (Cantor, 1842)) | 27 | |
| Piscichuvirus sanxiaense | Sānxiá atyid shrimp virus 4 (SxASC-4) | Unspecified atyid shrimp | 27 | |
| Piscichuvirus wenlingense | Wēnlǐng fish chu-like virus (WFClV) | Longspine snipefish (Macroramphosus scolopax (Linnaeus, 1758)) | 27 | |
| Pterovirus | Pterovirus chulinense | hymenopteran chu-related virus OKIAV147 (HCrV-147) | Aphelinid wasp (Aphelinus abdominalis (Dalman, 1820)) | 13 |
| “Rochuvirus” | “Rochuvirus chalcocoris” | Chalcocoris rutilans mononega-like virus 2 (CRmlV-2) | Stink bug (Chalcocoris rutilans (Fabricius, 1781)) | Unpublished, 25 |
| Scarabeuvirus | Scarabeuvirus blattae | Wǔchāng cockroach virus 3 (WcLFV-3) | German cockroach (Blattella germanica Linnaeus, 1767) | 9 |
| Scarabeuvirus dentati | Húběi chǔvirus-like virus 3 (HbCLV-3) | Unspecified dragonfly/damselfly (Odonata sp.) | 27 | |
| Scarabeuvirus hubeiense | Húběi chǔvirus-like virus 1 (HbCLV-1) | Unspecified dragonfly/damselfly (Odonata sp.) | 27 | |
| Scarabeuvirus lampyris | Lampyris noctiluca chuvirus-like virus 1 (LNClV-1) | Common glow-worm (Lampyris noctiluca (Linnaeus, 1767)) | 24 | |
| Scarabeuvirus lishiense | Líshí spider virus 1 (LsSV-1) | Common house spider (Parasteatoda tepidariorum (C. L. Koch, 1841)) | 9 | |
| Taceavirus | Taceavirus wenlingense | Wēnlǐng crustacean virus 15 (WlCV-15) | Unspecified crustacean | 27 |
| “Vapochuvirus” | “Vapochuvirus trialeurodis” | Trialeurodes vaporariorum mononega-like virus 2 (TVmlV-2) | Greenhouse whitefly (Trialeurodes vaporariorum Westwood, 1856) | 26 |
| Crepuscuviridae | ||||
| Aqualaruvirus | Aqualaruvirus sialis | megalopteran chu-related virus 119 (MCrV-119) | Unspecified corydalid fly | 13 |
| Myriaviridae | ||||
| Myriavirus | Myriavirus myriapedis | Húběi myriapoda virus 8 (HbMV-8) | Unspecified millipede/centipede (Myriapoda sp.) | 27 |
| Natareviridae | ||||
| Charybdivirus | Charybdivirus charybdis | Wēnzhōu crab virus 3 (WzCV-3) | Asian paddle crab (Charybdis japonica (A. Milne-Edwards, 1861)) | 9 |
DISCUSSION
The sheer number of (often fragmented, incomplete, and/or possibly misassembled) jingchuviral-like sequences in public databases indicates that the order Jingchuvirales will have to be expanded rather dramatically in the near future. The increased understanding of jingchuviral distribution and diversity is almost diametrically opposed to the molecular understanding of these viruses or their impacts on ecosystems. Thus far, with one possible exception reported in a preprint (49), not a single jingchuviral has been isolated in culture, and functional genomic studies are lacking. As a result, there is no certainty regarding the circularity and/or completeness of a given jingchuviral genome in sequence data sets, as pertinent genomic segments could have been overlooked, and genome termini are often not clearly defined. Likewise, the functions of individual jingchuviral proteins have not yet been systematically examined. Host specificity also remains unclear: whereas it is quite certain that jingchuvirals replicate in arthropods (9), and there is experimental evidence that they are vertically transmitted in cestodes (30), it is less certain whether jingchuvirals truly infect other animals (fish, mammals, reptiles, and sea anemones) or whether they are contaminants stemming from other organisms (such as arthropods) associated with these animals. It also remains to be determined whether jingchuvirals are commensals, mutualists, or parasites of their hosts and how they shape host population dynamics. The presented taxonomy framework will aid in the rapid classification of newly discovered jingchuvirals and presents a minimal set of criteria to make official classification possible. Once jingchuvirals become better characterized, it may be possible to expand classification criteria to include entire genome comparisons or at least to define taxa based on gene nucleotide identities of concatenated open reading frames and/or the identities of their concatenated expression products. In addition, the isolation of jingchuvirals in culture and/or the characterization of individual jingchuviral proteins using plasmid-based expression systems may aid in the inclusion of additional biological criteria in taxon demarcation. Nomenclature guidelines, in particular jingchuviral names, still have to be developed to decrease potential confusion (e.g., currently, several virus names contain the infix “chu-like,” albeit these viruses do not cluster with chuvirids, and many jingchuviral names are needlessly complex). Harmonizing virus names and officially expanding the classification of this understudied virus group will make jingchuvirals potentially more visible to the general virology community and thereby create interest in studying their biology.
DATA CITATION
The taxonomic proposal (TaxoProp) on the reorganization of order Jingchuvirales (2020.026M.R.Jingchuvirales), which was written by the authors of this article (57) and subsequently ratified by the International Committee on Taxonomy of Viruses (ICTV) in March 2021 (1), can be found at the ICTV website (https://talk.ictvonline.org). Draft TaxoProp 2021.015M.Ud.v1.Jingchuvirales_2ngen_10nsp (58), which was also written by the authors of this article, proposes to further expand this taxonomy and is currently under consideration by the ICTV.
ACKNOWLEDGMENTS
We are grateful to Peter J. Walker (University of Queensland, Australia) for pointing out assembly errors in various jingchuviral GenBank and SRA entries and critically reviewing both taxonomic proposals underlying this work. We thank Anya Crane and Jiro Wada (Integrated Research Facility at Fort Detrick, National Institute of Allergy and Infectious Diseases [NIAID], National Institutes of Health, Fort Detrick, Frederick, MD, USA) for critically editing the manuscript and for preparing figures, respectively. N.D.P., N.M.D., S.J., S.P., M.S., and J.H.K. are members of the 2020–2023 International Committee on Taxonomy of Viruses Jingchuvirales Study Group.
Conceptualization, N.D.P., N.M.D., S.J., S.P., M.S., and J.H.K.; methodology, N.D.P., N.M.D., S.J., S.P., and M.S.; formal analysis, N.D.P.; data curation, N.D.P.; writing—original draft preparation, N.D.P. and J.H.K.; writing—review and editing, all authors; supervision, N.D.P. and J.H.K.; project administration, N.D.P. All authors reviewed and approved the manuscript. N.D.P. and J.H.K. wrote the initial manuscript draft. The remaining authors performed analyses, reviewed and edited the paper, and were involved in original taxonomic proposal writing. The order of these authors in the byline is alphabetical by last name.
This work was supported in part through a Laulima Government Solutions, LLC, prime contract with the U.S. NIAID under contract no. HHSN272201800013C. J.H.K. performed this work as an employee of Tunnell Government Services (TGS), a subcontractor of Laulima Government Solutions, LLC, under contract no. HHSN272201800013C. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results. The content of this publication does not necessarily reflect the views or policies of the U.S. Department of Health and Human Services (HHS) or of the institutions and companies affiliated with the authors. Mention of trade names, commercial products, or organizations does not imply endorsement by the U.S. Government.
This work did not include any work with humans or animals.
We declare no conflict of interest.
Footnotes
Supplemental material is available online only.
Contributor Information
Nicholas Di Paola, Email: nicholas.dipaola.civ@mail.mil.
Nicole R. Buan, University of Nebraska—Lincoln
REFERENCES
- 1.Walker PJ, Siddell SG, Lefkowitz EJ, Mushegian AR, Adriaenssens E, Alfenas-Zerbini P, Davison AJ, Dempsey DM, Dutilh BE, García ML, Harrach B, Harrison RL, Hendrickson RC, Junglen S, Knowles NJ, Krupovic M, Kuhn JH, Lambert AJ, Łobocka M, Nibert M, Oksanen HM, Orton RJ, Robertson D, Rubino L, Sabanadzovic S, Simmonds P, Smith DB, Suzuki N, Dooerslaer KV, Vandamme A, Varsani A, Zerbini FM. 2021. Changes to virus taxonomy and to the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2021). Arch Virol 166:2633–2648. 10.1007/s00705-021-05156-1. [DOI] [PubMed] [Google Scholar]
- 2.Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini FM, Kuhn JH. 2020. Global organization and proposed megataxonomy of the virus world. Microbiol Mol Biol Rev 84:e00061-19. 10.1128/MMBR.00061-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Krupovic M, Kuhn JH, Wang F, Baquero DP, Dolja VV, Egelman EH, Prangishvili D, Koonin EV. 2021. Adnaviria: a new realm for archaeal filamentous viruses with linear A-form double-stranded DNA genomes. J Virol 95:e00673-21. 10.1128/JVI.00673-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Walker PJ, Siddell SG, Lefkowitz EJ, Mushegian AR, Adriaenssens EM, Dempsey DM, Dutilh BE, Harrach B, Harrison RL, Hendrickson RC, Junglen S, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Nibert M, Orton RJ, Rubino L, Sabanadzovic S, Simmonds P, Varsani A, Zerbini FM, Davison AJ. 2020. Changes to virus taxonomy and the statutes ratified by the International Committee on Taxonomy of Viruses (2020). Arch Virol 165:2737–2748. 10.1007/s00705-020-04752-x. [DOI] [PubMed] [Google Scholar]
- 5.Koonin EV, Dolja VV, Krupovic M, Kuhn JH. 2021. Viruses defined by the position of the virosphere within the replicator space. Microbiol Mol Biol Rev 85:e00193-20. 10.1128/MMBR.00193-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wolf YI, Kazlauskas D, Iranzo J, Lucía-Sanz A, Kuhn JH, Krupovic M, Dolja VV, Koonin EV. 2018. Origins and evolution of the global RNA virome. mBio 9:e02329-18. 10.1128/mBio.02329-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Siddell SG, Walker PJ, Lefkowitz EJ, Mushegian AR, Adams MJ, Dutilh BE, Gorbalenya AE, Harrach B, Harrison RL, Junglen S, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Nibert M, Rubino L, Sabanadzovic S, Sanfaçon H, Simmonds P, Varsani A, Zerbini FM, Davison AJ. 2019. Additional changes to taxonomy ratified in a special vote by the International Committee on Taxonomy of Viruses (October 2018). Arch Virol 164:943–946. 10.1007/s00705-018-04136-2. [DOI] [PubMed] [Google Scholar]
- 8.Kuhn JH, Wolf YI, Krupovic M, Zhang Y-Z, Maes P, Dolja VV, Koonin EV. 2019. Classify viruses—the gain is worth the pain. Nature 566:318–320. 10.1038/d41586-019-00599-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li C-X, Shi M, Tian J-H, Lin X-D, Kang Y-J, Chen L-J, Qin X-C, Xu J, Holmes EC, Zhang Y-Z. 2015. Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. Elife 4:e05378. 10.7554/eLife.05378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Easton A, Pringle CR. 2011. Order Mononegavirales, p 653–657. In King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (ed), Virus taxonomy. Classification and nomenclature of viruses. Ninth report of the International Committee on Taxonomy of Viruses. Elsevier Academic Press, London, United Kingdom. [Google Scholar]
- 11.Harvey E, Rose K, Eden J-S, Lawrence A, Doggett SL, Holmes EC. 2019. Identification of diverse arthropod associated viruses in native Australian fleas. Virology 535:189–199. 10.1016/j.virol.2019.07.010. [DOI] [PubMed] [Google Scholar]
- 12.Han X, Wang H, Wu N, Liu W, Cao M, Wang X. 2020. Leafhopper Psammotettix alienus hosts chuviruses with different genomic structures. Virus Res 285:197992. 10.1016/j.virusres.2020.197992. [DOI] [PubMed] [Google Scholar]
- 13.Käfer S, Paraskevopoulou S, Zirkel F, Wieseke N, Donath A, Petersen M, Jones TC, Liu S, Zhou X, Middendorf M, Junglen S, Misof B, Drosten C. 2019. Re-assessing the diversity of negative strand RNA viruses in insects. PLoS Pathog 15:e1008224. 10.1371/journal.ppat.1008224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Schoonvaere K, De Smet L, Smagghe G, Vierstraete A, Braeckman BP, de Graaf DC. 2016. Unbiased RNA shotgun metagenomics in social and solitary wild bees detects associations with eukaryote parasites and new viruses. PLoS One 11:e0168456. 10.1371/journal.pone.0168456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wang H, Liu Y, Liu W, Cao M, Wang X. 2019. Sequence analysis and genomic organization of a novel chuvirus, Tàiyuán leafhopper virus. Arch Virol 164:617–620. 10.1007/s00705-018-4075-4. [DOI] [PubMed] [Google Scholar]
- 16.Sadeghi M, Altan E, Deng X, Barker CM, Fang Y, Coffey LL, Delwart E. 2018. Virome of >12 thousand Culex mosquitoes from throughout California. Virology 523:74–88. 10.1016/j.virol.2018.07.029. [DOI] [PubMed] [Google Scholar]
- 17.Hang J, Klein TA, Kim H-C, Yang Y, Jima DD, Richardson JH, Jarman RG. 2016. Genome sequences of five arboviruses in field-captured mosquitoes in a unique rural environment of South Korea. Genome Announc 4:e01644-15. 10.1128/genomeA.01644-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Harvey E, Rose K, Eden J-S, Lo N, Abeyasuriya T, Shi M, Doggett SL, Holmes EC. 2019. Extensive diversity of RNA viruses in Australian ticks. J Virol 93:e01358-18. 10.1128/JVI.01358-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gondard M, Temmam S, Devillers E, Pinarello V, Bigot T, Chrétien D, Aprelon R, Vayssier-Taussat M, Albina E, Eloit M, Moutailler S. 2020. RNA viruses of Amblyomma variegatum and Rhipicephalus microplus and cattle susceptibility in the French Antilles. Viruses 12:144. 10.3390/v12020144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sameroff S, Tokarz R, Charles RA, Jain K, Oleynik A, Che X, Georges K, Carrington CV, Lipkin WI, Oura C. 2019. Viral diversity of tick species parasitizing cattle and dogs in Trinidad and Tobago. Sci Rep 9:10421. 10.1038/s41598-019-46914-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tokarz R, Sameroff S, Tagliafierro T, Jain K, Williams SH, Cucura DM, Rochlin I, Monzon J, Carpi G, Tufts D, Diuk-Wasser M, Brinkerhoff J, Lipkin WI. 2018. Identification of novel viruses in Amblyomma americanum, Dermacentor variabilis, and Ixodes scapularis ticks. mSphere 3:e00614-17. 10.1128/mSphere.00614-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Costa VA, Mifsud JCO, Gilligan D, Williamson JE, Holmes EC, Geoghegan JL. 2021. Metagenomic sequencing reveals a lack of virus exchange between native and invasive freshwater fish across the Murray-Darling Basin, Australia. Virus Evol 7:veab034. 10.1093/ve/veab034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Argenta FF, Hepojoki J, Smura T, Szirovicza L, Hammerschmitt ME, Driemeier D, Kipar A, Hetzel U, The BIBD Group . 2020. Identification of reptarenaviruses, hartmaniviruses, and a novel chuvirus in captive native Brazilian boa constrictors with boid inclusion body disease. J Virol 94:e00001-20. 10.1128/JVI.00001-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Viljakainen L, Borshagovski A-M, Saarenpää S, Kaitala A, Jurvansuu J. 2020. Identification and characterisation of common glow-worm RNA viruses. Virus Genes 56:236–248. 10.1007/s11262-019-01724-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chang T, Hirai J, Hunt BPV, Suttle CA. 2021. Arthropods and the evolution of RNA viruses. bioRxiv 10.1101/2021.05.30.446314. [DOI]
- 26.Wu H, Pang R, Cheng T, Xue L, Zeng H, Lei T, Chen M, Wu S, Ding Y, Zhang J, Shi M, Wu Q. 2020. Abundant and diverse RNA viruses in insects revealed by RNA-Seq analysis: ecological and evolutionary implications. mSystems 5:e00039-20. 10.1128/mSystems.00039-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Shi M, Lin X-D, Tian J-H, Chen L-J, Chen X, Li C-X, Qin X-C, Li J, Cao J-P, Eden J-S, Buchmann J, Wang W, Xu J, Holmes EC, Zhang Y-Z. 2016. Redefining the invertebrate RNA virosphere. Nature 540:539–543. 10.1038/nature20167. [DOI] [PubMed] [Google Scholar]
- 28.Sameroff S, Tokarz R, Jain K, Oleynik A, Carrington CVF, Lipkin WI, Oura CAL. 2021. Novel quaranjavirus and other viral sequences identified from ticks parasitizing hunted wildlife in Trinidad and Tobago. Ticks Tick Borne Dis 12:101730. 10.1016/j.ttbdis.2021.101730. [DOI] [PubMed] [Google Scholar]
- 29.Gómez GF, Isaza JP, Segura JA, Alzate JF, Gutiérrez LA. 2020. Metatranscriptomic virome assessment of Rhipicephalus microplus from Colombia. Ticks Tick Borne Dis 11:101426. 10.1016/j.ttbdis.2020.101426. [DOI] [PubMed] [Google Scholar]
- 30.Hahn MA, Rosario K, Lucas P, Dheilly NM. 2020. Characterization of viruses in a tapeworm: phylogenetic position, vertical transmission, and transmission to the parasitized host. ISME J 14:1755–1767. 10.1038/s41396-020-0642-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Niu J, Zhang W, Sun Q-Z, Wang J-J. 2019. Three novel RNA viruses in the spider mite Tetranychus urticae and their possible interactions with the host RNA interference response. J Invertebr Pathol 166:107228. 10.1016/j.jip.2019.107228. [DOI] [PubMed] [Google Scholar]
- 32.Maia LMS, de Lara Pinto AZ, de Carvalho MS, de Melo FL, Ribeiro BM, Slhessarenko RD. 2019. Novel viruses in mosquitoes from Brazilian Pantanal. Viruses 11:957. 10.3390/v11100957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Niu J, Li X-L, Wu Y-L, Sun Q-Z, Zhang W, Cao M, Wang J-J. 2020. RNA virome screening in diverse but ecologically related citrus pests reveals potential virus-host interactions. J Invertebr Pathol 170:107329. 10.1016/j.jip.2020.107329. [DOI] [PubMed] [Google Scholar]
- 34.Modha S, Hughes J, Bianco G, Ferguson HM, Helm B, Tong L, Wilkie GS, Kohl A, Schnettler E. 2019. Metaviromics reveals unknown viral diversity in the biting midge Culicoides impunctatus. Viruses 11:865. 10.3390/v11090865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mishra N, Fagbo SF, Alagaili AN, Nitido A, Williams SH, Ng J, Lee B, Durosinlorun A, Garcia JA, Jain K, Kapoor V, Epstein JH, Briese T, Memish ZA, Olival KJ, Lipkin WI. 2019. A viral metagenomic survey identifies known and novel mammalian viruses in bats from Saudi Arabia. PLoS One 14:e0214227. 10.1371/journal.pone.0214227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.de Souza WM, Fumagalli MJ, Torres Carrasco ADO, Romeiro MF, Modha S, Seki MC, Gheller JM, Daffre S, Nunes MRT, Murcia PR, Acrani GO, Figueiredo LTM. 2018. Viral diversity of Rhipicephalus microplus parasitizing cattle in southern Brazil. Sci Rep 8:16315. 10.1038/s41598-018-34630-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Brinkmann A, Dinçer E, Polat C, Hekimoğlu O, Hacıoğlu S, Földes K, Özkul A, Öktem İMA, Nitsche A, Ergünay K. 2018. A metagenomic survey identifies Tamdy orthonairovirus as well as divergent phlebo-, rhabdo-, chu- and flavi-like viruses in Anatolia, Turkey. Ticks Tick Borne Dis 9:1173–1183. 10.1016/j.ttbdis.2018.04.017. [DOI] [PubMed] [Google Scholar]
- 38.de Lara Pinto AZ, Santos de Carvalho M, de Melo FL, Ribeiro ALM, Morais Ribeiro B, Dezengrini Slhessarenko R. 2017. Novel viruses in salivary glands of mosquitoes from sylvatic Cerrado, Midwestern Brazil. PLoS One 12:e0187429. 10.1371/journal.pone.0187429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Medd NC, Fellous S, Waldron FM, Xuéreb A, Nakai M, Cross JV, Obbard DJ. 2018. The virome of Drosophila suzukii, an invasive pest of soft fruit. Virus Evol 4:vey009. 10.1093/ve/vey009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Waldron FM, Stone GN, Obbard DJ. 2018. Metagenomic sequencing suggests a diversity of RNA interference-like responses to viruses across multicellular eukaryotes. PLoS Genet 14:e1007533. 10.1371/journal.pgen.1007533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhang Y-Y, Chen Y, Wei X, Cui J. 17 June 2021. Viromes in marine ecosystems reveal remarkable invertebrate RNA virus diversity. Sci China Life Sci 10.1007/s11427-020-1936-2. [DOI] [PubMed]
- 42.Vanmechelen B, Merino M, Vergote V, Laenen L, Thijssen M, Martí-Carreras J, Claerebout E, Maes P. 2021. Exploration of the Ixodes ricinus virosphere unveils an extensive virus diversity including novel coltiviruses and other reoviruses. Virus Evol 7:veab066. 10.1093/ve/veab066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Geoghegan JL, Di Giallonardo F, Wille M, Ortiz-Baez AS, Costa VA, Ghaly T, Mifsud JCO, Turnbull OMH, Bellwood DR, Williamson JE, Holmes EC. 2021. Virome composition in marine fish revealed by meta-transcriptomics. Virus Evol 7:veab005. 10.1093/ve/veab005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Batson J, Dudas G, Haas-Stapleton E, Kistler AL, Li LM, Logan P, Ratnasiri K, Retallack H. 2020. Single mosquito metatranscriptomics identifies vectors, emerging pathogens and reservoirs in one assay. Elife 7:e68353. 10.7554/eLife.68353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Shi C, Zhao L, Atoni E, Zeng W, Hu X, Matthijnssens J, Yuan Z, Xia H. 2020. The conservation of a core virome in Aedes mosquitoes across different developmental stages and continents. bioRxiv 10.1101/2020.04.23.058701. [DOI] [PMC free article] [PubMed]
- 46.Chong R, Shi M, Grueber CE, Holmes EC, Hogg C, Belov K, Barrs VR. 2018. Characterisation of the faecal virome of captive and wild Tasmanian devils using virus-like particles metagenomics and meta-transcriptomics. bioRxiv 10.1101/443457. [DOI]
- 47.Shi J, Shen S, Wu H, Zhang Y, Deng F. 2021. Metagenomic profiling of viruses associated with Rhipicephalus microplus ticks in Yunnan Province, China. Virol Sin 36:623–635. 10.1007/s12250-020-00319-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wang S, Zhao T, Yu X, Lin Z, Hua X, Cui L. 2020. Characterization of tick viromes collected from dogs in China. Biosaf Health 2:79–88. 10.1016/j.bsheal.2020.03.002. [DOI] [Google Scholar]
- 49.Quan L, Wang Z-D, Gao Y, Lv X, Han S, Zhang X, Shao J-W, Chen C, Li L, Hou Z-J, Sui L, Zhao Y, Wang B, Wang W, Song M. 2020. Identification of a new chuvirus associated with febrile illness in China. Res Sq Prepr 10.21203/rs.3.rs-104938/v1. [DOI]
- 50.Dezordi FZ, Dos Santos Vasconcelos CR, Rezende AM, Wallau GL. 2020. In and outs of Chuviridae endogenous viral elements: origin of a potentially new retrovirus and signature of ancient and ongoing arms race in mosquito genomes. Front Genet 11:542437. 10.3389/fgene.2020.542437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Whitfield ZJ, Dolan PT, Kunitomi M, Tassetto M, Seetin MG, Oh S, Heiner C, Paxinos E, Andino R. 2017. The diversity, structure, and function of heritable adaptive immunity sequences in the Aedes aegypti genome. Curr Biol 27:3511–3519.e7. 10.1016/j.cub.2017.09.067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Thompson JD, Gibson TJ, Higgins DG. 2002. Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinformatics Chapter 2:Unit 2.3. 10.1002/0471250953.bi0203s00. [DOI] [PubMed] [Google Scholar]
- 53.Edgar RC. 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A. 2012. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. 10.1093/bioinformatics/bts199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Abascal F, Zardoya R, Posada D. 2005. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2105. 10.1093/bioinformatics/bti263. [DOI] [PubMed] [Google Scholar]
- 56.Guindon S, Gascuel O. 2003. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704. 10.1080/10635150390235520. [DOI] [PubMed] [Google Scholar]
- 57.Di Paola N, Dheilly NM, Kuhn JH, Junglen S, Paraskevopoulou S, Postler TS, Shi M. 2021. Reorganize the order to include four new families, 18 new genera, and 22 new species (Jingchuvirales). ICTV [International Committee on Taxonomy of Viruses] TaxoProp 2020.026M.R.Jingchuvirales. https://talk.ictvonline.org/files/ictv_official_taxonomy_updates_since_the_8th_report/m/animal-dsrna-and-ssrna-viruses/11055.
- 58.Di Paola N, Dheilly NM, Kuhn JH, Junglen S, Paraskevopoulou S, Postler TS, Shi M. 2021. Create ten new species and two new genera in the families Aliusviridae and Chuviridae (Jingchuvirales). ICTV [International Committee on Taxonomy of Viruses] TaxoProp 2021.015M.Ud.v1.Jingchuvirales_2ngen_10nsp. https://talk.ictvonline.org/files/proposals/animal_dsrna_and_ssrna-_viruses/m/animal_rna_minus_under_consideration/12825.
- 59.Briddon RW, Martin DP, Roumagnac P, Navas-Castillo J, Fiallo-Olivé E, Moriones E, Lett J-M, Zerbini FM, Varsani A. 2018. Alphasatellitidae: a new family with two subfamilies for the classification of geminivirus- and nanovirus-associated alphasatellites. Arch Virol 163:2587–2600. 10.1007/s00705-018-3854-2. [DOI] [PubMed] [Google Scholar]
- 60.Bào Y, Amarasinghe GK, Basler CF, Bavari S, Bukreyev A, Chandran K, Dolnik O, Dye JM, Ebihara H, Formenty P, Hewson R, Kobinger GP, Leroy EM, Mühlberger E, Netesov SV, Patterson JL, Paweska JT, Smither SJ, Takada A, Towner JS, Volchkov VE, Wahl-Jensen V, Kuhn JH. 2017. Implementation of objective PASC-derived taxon demarcation criteria for official classification of filoviruses. Viruses 9:106. 10.3390/v9050106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Lauber C, Gorbalenya AE. 2012. Toward genetics-based virus taxonomy: comparative analysis of a genetics-based classification and the taxonomy of picornaviruses. J Virol 86:3905–3915. 10.1128/JVI.07174-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Radoshitzky SR, Bào Y, Buchmeier MJ, Charrel RN, Clawson AN, Clegg CS, DeRisi JL, Emonet S, Gonzalez J-P, Kuhn JH, Lukashevich IS, Peters CJ, Romanowski V, Salvato MS, Stenglein MD, de la Torre JC. 2015. Past, present, and future of arenavirus taxonomy. Arch Virol 160:1851–1874. 10.1007/s00705-015-2418-y. [DOI] [PubMed] [Google Scholar]
- 63.Kuhn JH, Dürrwald R, Bào Y, Briese T, Carbone K, Clawson AN, deRisi JL, Garten W, Jahrling PB, Kolodziejek J, Rubbenstroth D, Schwemmle M, Stenglein M, Tomonaga K, Weissenböck H, Nowotny N. 2015. Taxonomic reorganization of the family Bornaviridae. Arch Virol 160:621–632. 10.1007/s00705-014-2276-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Bao Y, Chetvernin V, Tatusova T. 2014. Improvements to pairwise sequence comparison (PASC): a genome-based Web tool for virus classification. Arch Virol 159:3293–3304. 10.1007/s00705-014-2197-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kuhn JH, Adkins S, Agwanda BR, Al Kubrusli R, Alkhovsky SV, Amarasinghe GK, Avšič-Županc T, Ayllón MA, Bahl J, Balkema-Buschmann A, Ballinger MJ, Basler CF, Bavari S, Beer M, Bejerman N, Bennett AJ, Bente DA, Bergeron É, Bird BH, Blair CD, Blasdell KR, Blystad D-R, Bojko J, Borth WB, Bradfute S, Breyta R, Briese T, Brown PA, Brown JK, Buchholz UJ, Buchmeier MJ, Bukreyev A, Burt F, Büttner C, Calisher CH, Cao M, Casas I, Chandran K, Charrel RN, Cheng Q, Chiaki Y, Chiapello M, Choi I-R, Ciuffo M, Clegg JCS, Crozier I, Dal Bó E, de la Torre JC, de Lamballerie X, de Swart RL, et al. 2021. 2021 taxonomic update of phylum Negarnaviricota (Riboviria: Orthornavirae), including the large orders Bunyavirales and Mononegavirales. Arch Virol 166:3567–3579. 10.1007/s00705-021-05266-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. Download aem.01954-21-s0001.xlsx, XLSX file, 0.03 MB (30.5KB, xlsx)
Table S2. Download aem.01954-21-s0002.xlsx, XLSX file, 0.03 MB (32.8KB, xlsx)