ABSTRACT
The assembly and egress of alphaherpesviruses, including herpes simplex virus 1 (HSV-1) and pseudorabies virus (PRV), within neurons is poorly understood. A key unresolved question is the structure of the viral particle that moves by anterograde transport along the axon, and two alternative mechanisms have been described. In the “married” model, capsids acquire their envelopes in the cell body and then traffic along axons as enveloped virions within a bounding organelle. In the “separate” model, nonenveloped capsids travel from the cell body into and along the axon, eventually encountering their envelopment organelles at a distal site, such as the nerve cell terminal. Here, we describe an “envelopment trap” to test these models using the dominant negative terminal endosomal sorting complex required for transport (ESCRT) component VPS4-EQ. Green fluorescent protein (GFP)-tagged VPS4-EQ was used to arrest HSV-1 or PRV capsid envelopment, inhibit downstream trafficking, and GFP-label envelopment intermediates. We found that GFP-VPS4-EQ inhibited trafficking of HSV-1 capsids into and along the neurites and axons of mouse CAD cells and rat embryonic primary cortical neurons, consistent with egress via the married pathway. In contrast, transport of HSV-1 capsids was unaffected in the neurites of human SK-N-SH neuroblastoma cells, consistent with the separate mechanism. Unexpectedly, PRV (generally thought to utilize the married pathway) also appeared to employ the separate mechanism in SK-N-SH cells. We propose that apparent differences in the methods of HSV-1 and PRV egress are more likely a reflection of the host neuron in which transport is studied rather than true biological differences between the viruses themselves.
IMPORTANCE Alphaherpesviruses, including herpes simplex virus 1 (HSV-1) and pseudorabies virus (PRV), are pathogens of the nervous system. They replicate in the nerve cell body and then travel great distances along axons to reach nerve termini and spread to adjacent epithelial cells; however, key aspects of how these viruses travel along axons remain controversial. Here, we test two alternative mechanisms for transport, the married and separate models, by blocking envelope assembly, a critical step in viral egress. When we arrest formation of the viral envelope using a mutated component of the cellular ESCRT apparatus, we find that entry of viral particles into axons is blocked in some types of neurons but not others. This approach allows us to determine whether envelope assembly occurs prior to entry of viruses into axons or afterwards and, thus, to distinguish between the alternative models for viral transport.
KEYWORDS: ESCRT, HSV-1, PRV, anterograde transport, envelopment, neuronal egress
INTRODUCTION
Alphaherpesviruses, including herpes simplex virus 1 (HSV-1) and pseudorabies virus (PRV), assemble their capsids and then package them with the viral double-stranded DNA (dsDNA) genome in the infected cell nucleus (1–4). Packaged capsids then bud into the inner nuclear membrane to generate a primary enveloped virion in the perinuclear space (5–11). Fusion of this primary envelope with the outer nuclear membrane delivers “naked” nonenveloped capsids into the cytoplasm, where they undergo a final, secondary envelopment by budding into the lumen of membrane-bounded organelles (4, 12–17). Nonenveloped capsids prior to cytoplasmic envelopment, and the cytoplasmic organelles that acquire enveloped virions within their lumen, are capable of recruiting molecular motors in order to traffic along microtubules in an anterograde direction during assembly and egress (18–24).
The envelopment and trafficking of alphaherpesviruses within neurons is a particularly complex process. A major unresolved question is the structure of the viral particle that departs the cell body and then moves by microtubule-directed anterograde transport along the axon. Two alternative mechanisms for this process have been described (21, 24–28). In the “married” model, capsids emerge from the cell nucleus, acquire their envelopes in the neuronal cell body, and then traffic along axons as enveloped virions contained within the lumen of a bounding organelle. Alternatively, in the “separate” model, nonenveloped (naked) capsids travel from the nucleus into the axon and then along the axon to eventually encounter their envelopment organelles at a distal site, such as the nerve cell terminal. Distinguishing between these two possibilities is important because they have profoundly different implications for the molecular mechanisms of alphaherpesvirus transport and assembly. In the married model, the envelopment organelle may contain one or multiple enveloped viral particles, and the molecular motors driving anterograde traffic along microtubules must be recruited to the surrounding bilayer of this organelle (23, 24, 29, 30). In the separate model, nonenveloped naked capsids travel along axons as individual particles, and molecular motors would likely be recruited directly to the proteinaceous surface of the nucleocapsid or its surrounding tegument (18, 19, 23, 24, 31).
Evidence for and against the married and separate models for HSV-1 and PRV trafficking has led to much discussion (21, 24–27, 32). Imaging and genetic studies have resulted in a general consensus that PRV favors the married route, acquiring its envelope in the neuronal cell body. However, the situation for HSV-1 is more contentious. Data for transport of nonenveloped HSV-1 capsids along axons, in accord with the separate model, has been reported in explanted human fetal dorsal root ganglia (DRG), rat DRG, rat trigeminal ganglia (TG), embryonic rat hippocampal neurons, mouse cortical neurons, and cultured human SK-N-SH neuroblastoma cells (33–39). In contrast, evidence in favor of the married model has been described during HSV-1 infection of differentiated mouse Cath.a-derived (CAD) neurons (a derivative of a catecholaminergic central nervous system cell line [28, 40, 41]) and in rat, mouse, and chick DRG (42–44).
How can these conflicting data be reconciled? One suggestion is that HSV-1 is capable of utilizing both the separate and married transport pathways (24, 28, 32, 45, 46), though which host or viral factors might influence the choice of mechanism remain completely unknown. Alternatively, it has been suggested that HSV-1 (like PRV) traffics exclusively by the married model and that the apparently nonenveloped HSV-1 capsids observed in axons are an artifact of technical limitations in immunofluorescence (IF) and electron microscopy (EM) (25). Specifically, limited antibody sensitivity and poor antibody accessibility to the crowded, multilayered interior of enveloped, tegument-bound, organelle-enclosed virions might confound attempts to detect envelope antigens associated with HSV-1 capsids, giving axonal enveloped virions the appearance of naked capsids (25). Similarly, EM studies identifying nonenveloped HSV-1 capsids within axons, trafficking in accord with the separate model (35, 36), have been reinterpreted as instead depicting axonal dense-core vesicles or capsids present within dendrites rather than undergoing egress along axons (25). These alternative interpretations have, in turn, been challenged (26).
In the face of these problems, we sought to develop a functional assay for the site of HSV-1 capsid envelopment within neurons to complement the imaging studies detailed above. Like many families of enveloped viruses (17, 47, 48), Herpesviridae exploit the cellular endosomal sorting complex required for transport (ESCRT) machinery to construct and drive scission of their envelopes (17, 22, 49–51). The ESCRT apparatus manipulates organellar membranes during a variety of normal cellular processes via the deposition of ESCRT-III, a spring-like filament that drives the deformation of planar bilayers into vesicles and tubules (48, 52–55). HSV-1 may bypass the normal cellular pathways of ESCRT complex assembly (17, 49, 53, 56) and instead polymerize ESCRT-III filaments directly on the surface of the tegument-bound capsid to support envelope deformation (57). When budding is complete, the ESCRT-III filament constricts the envelope at its neck to catalyze scission, a process accompanied by disassembly and removal of ESCRT-III via the hexameric AAA ATPase vacuolar protein sorting 4 (VPS4) (53, 54, 58). Mutation of the VPS4 ATPase active site residue glutamate233 to glutamine generates the allele VPS4-EQ, which is able to bind ATP but not hydrolyze it (59). VPS4-EQ exerts a dominant negative effect since hexamers containing a mixture of wild-type VPS4 and VPS4-EQ subunits can associate with the ESCRT-III filament but are unable to drive its disassembly, irreversibly “locking” the VPS4 hexamer onto the bud neck and preventing completion of scission (59–62). As expected for viruses that utilize ESCRT-III to build their envelopes, expression of VPS4-EQ blocks completion of HSV-1 and PRV envelopment (22, 50) and arrests HSV-1 capsids in a partially membrane-enclosed state (50, 51). Using green fluorescent protein (GFP)-tagged alleles of VPS4 and VPS4-EQ, we were able to image trapped HSV-1 and PRV envelopment intermediates (22, 51, 63). Interestingly, unlike nonenveloped capsids and organelle-associated enveloped virions, VPS4-EQ-arrested and partially enveloped capsids were unable to traffic along microtubules in an in vitro system, which may reflect an assembly checkpoint during viral egress (20, 22, 64).
We reasoned that GFP-VPS4-EQ could be used as an “envelopment trap” in order to test whether alphaherpesviruses were utilizing the married or separate mechanisms of traffic within neurons. In the married pathway, GFP-VPS4-EQ should arrest HSV-1 or PRV capsid envelopment in the neuronal cell body. The trapped envelopment intermediates should exhibit reduced transport along microtubules and, thus, an impaired ability to travel from the cell body into axons. Furthermore, even if the block in traffic is incomplete, the majority of virions participating in the married pathway will become bound to GFP-VPS4-EQ in the cell body and, as a consequence, be fluorescently tagged with GFP by the time they enter the axon. In contrast, capsids following the separate mechanism will, by definition, not engage the envelopment apparatus in the cell body and should not spring the GFP-VPS4-EQ trap before entering the axon (21, 24–27). The microtubule-mediated trafficking of such capsids would thus be unaffected, they should travel from the cell body into the axon in normal numbers, and they should not become fluorescently labeled by GFP.
RESULTS
Consequences of GFP-VPS4 or GFP-VPS4-EQ expression for HSV-1 assembly and trafficking in differentiated CAD cells.
We used transient transfection to introduce plasmids expressing GFP-VPS4-EQ and control wild-type GFP-VPS4 into CAD cells (40, 41, 65), a well-established model system for the study of alphaherpesvirus neuronal trafficking (42, 45, 66–68). The plasmids were designed to express GFP-VPS4 and GFP-VPS4-EQ only following addition of doxycycline since long-term expression of dominant negative GFP-VPS4-EQ can be toxic to cells (22). Following plasmid transfection, the CAD cells were serum-starved to drive differentiation and formation of axon-like neurites and then infected by HSV-1 and doxycycline added to induce GFP-VPS4 or GFP-VPS4-EQ expression. After 20 h of HSV-1 infection, cells were fixed and mounted for imaging. Infected CAD cells were imaged in the red and green channels to visualize HSV-1 capsid-associated mCherry-UL25 fluorescence and GFP-tagged alleles of VPS4, respectively. Representative examples of infected CAD cell neurites used for quantitation of HSV-1 entry are shown in Fig. 1A.
FIG 1.
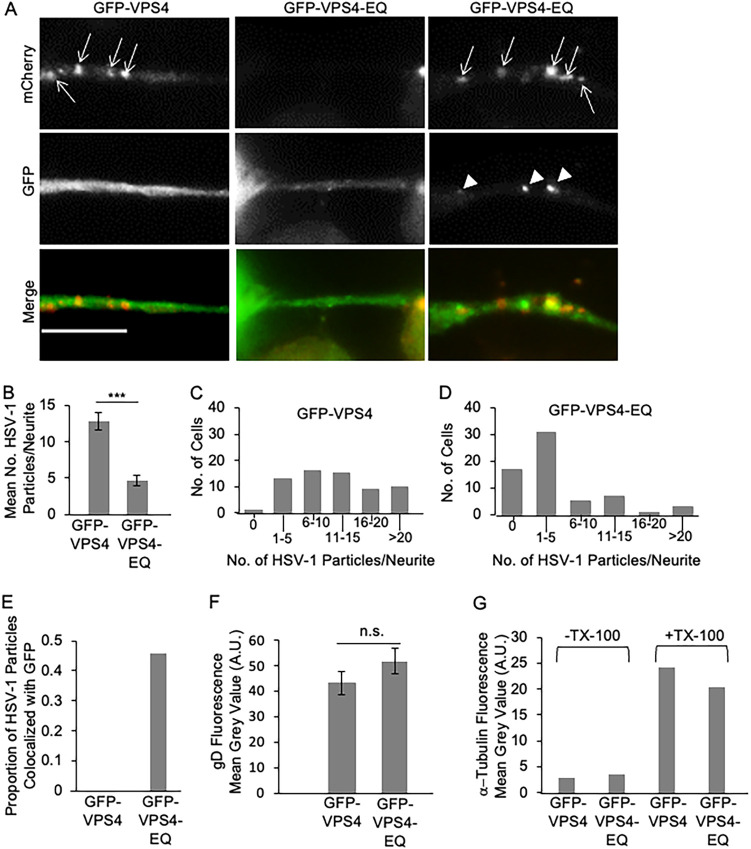
Transport of HSV-1 particles in the neurites of CAD cells transfected to express GFP-VPS4 or GFP-VPS4-EQ. Undifferentiated CAD cells were transfected with plasmids constructed to express GFP-VPS4 or GFP-VPS4-EQ under the control of a tetracycline-inducible promoter. They were then differentiated, infected with HSV-1, and GFP-VPS4 or GFP-VPS4-EQ expression induced by addition of doxycycline. After 20 h, cells were fixed, incubated with primary and fluorescent secondary antibodies as appropriate, and imaged by fluorescence microscopy. (A) Gallery of neurites from GFP-VPS4-expressing (left column) or GFP-VPS4-EQ-expressing (middle and right columns) CAD cells. Neurites were imaged in the red channel to detect puncta corresponding to mCherry-tagged HSV-1 capsids (top row, white arrows) or the green channel to detect GFP-tagged VPS4 alleles (middle row, white arrowheads indicate puncta enriched in GFP fluorescence). Bottom row shows merged images. Scale bar, 10 μm. (B) Number of mCherry-fluorescent HSV-1 capsid-associated puncta per neurite. Plotted values are the mean and standard deviation from the mean for neurites from 63 CAD cells expressing GFP-VPS4 (total of 822 puncta) or 63 CAD cells expressing GFP-VPS4-EQ (total of 297 puncta). (C, D) Frequency (y axes) at which neurites of infected CAD cells expressing GFP-VPS4 or GFP-VPS4-EQ (respectively) contain numbers of HSV-1 puncta in the ranges indicated (x axes). (E) Proportion of mCherry-fluorescent puncta, similar to those scored in panel B, exhibiting colocalization with GFP-VPS4 (total of 293 puncta) or GFP-VPS4-EQ fluorescence (total of 401 puncta) at levels enriched above that of background (see the text for more details). (F) Cell surface anti-gD immunofluorescence intensity plotted in arbitrary units (A.U.) for fixed, nonpermeabilized HSV-1-infected CAD cells expressing GFP-VPS4 or GFP-VPS4-EQ. (G) Similar experiment to panel F but plotting the intensity of anti-α-tubulin immunofluorescence displayed by fixed, Triton X-100 permeabilized (+TX-100) or nonpermeabilized (−TX-100) HSV-1-infected CAD cells expressing GFP-VPS4 or GFP-VPS4-EQ. For panels in this and all figures, no significant difference (n.s.) corresponds to P > 0.05; ***, P < 0.001 (Student's t test).
HSV-1 capsid-associated red puncta were readily visualized in the long axon-like neurites of differentiated CAD cells expressing wild-type control GFP-VPS4 (Fig. 1A, left column). In contrast, the neurites of GFP-VPS4-EQ-expressing cells contained reduced numbers of red puncta (Fig. 1A, right column) or none at all (Fig. 1A, middle column). We counted the numbers of HSV-1 capsid-associated particles seen in the neurites of cells expressing GFP-VPS4 or GFP-VPS4-EQ. GFP-VPS4-EQ expression reduced the mean number of HSV-1 particles entering axons by 64% compared to the wild-type GFP-VPS4 control (Fig. 1B). This overall decrease in the mean number of neurite-associated puncta was a result of a dramatic shift in the frequency distribution of virions within neurites. In control (GFP-VPS4-expressing) CAD cells, more than three-quarters of all neurites (50 of 63 counted) contained 6 or more viral particles, with the remainder exhibiting between 1 and 5 (Fig. 1C). This distribution was reversed upon GFP-VPS4-EQ expression; three-quarters of all neurites (48 of 63 counted) contained 5 or fewer viral particles, and more than one-quarter of the neurites (17 of 63) held no virions at all (Fig. 1C).
Some HSV-1 capsid-associated particles were nevertheless seen to enter axons in the presence of GFP-VPS4-EQ (Fig. 1B and D). To investigate the origin of these virions, we tested whether they were bound to GFP-VPS4-EQ, which would indicate that they arose from capsids that had attempted to undergo envelopment in the cell body. A complication of this analysis is that GFP-VPS4 and GFP-VPS4-EQ exist not only as peripheral membrane proteins but also diffusely distributed in the cytoplasm (53, 54, 61, 62), filling the cell body and neurite (GFP-VPS4 and GFP-VPS4-EQ fluorescence in neurites is shown in Fig. 1A). We, therefore, only considered capsid-associated HSV-1 puncta to be bound to GFP-VPS4 or GFP-VPS4-EQ if the intensity of the colocalizing GFP fluorescence exceeded the background intensity of diffuse GFP fluorescence in the cytoplasm (see Materials and Methods). These data are summarized in Fig. 1E. GFP-VPS4 was never found to be enriched on axonal HSV-1 particles above background fluorescence levels in the cytoplasm, which was expected since the wild-type allele of VPS4 departs from assembling virions upon the completion of envelopment (17, 53, 54, 61, 62). In contrast, 46% of axonal HSV-1 particles colocalized with GFP-VPS4-EQ at sites where GFP-VPS4-EQ fluorescence was enriched above that in the cytoplasm (Fig. 1E).
Taking the data in Fig. 1A to E together, GFP-VPS4-EQ expression reduced the number of HSV-1 capsid-associated particles entering CAD neurites to about 36% of that seen in the GFP-VPS4 control. Of that subpopulation able to enter the neurite, 46% colocalized with enriched levels of GFP-VPS4-EQ. This means that GFP-VPS4-EQ expression results in ∼80% of the HSV-1 particles that would normally enter neurites now no longer doing so, or entering the neurite in association with GFP-VPS4-EQ. This is consistent with ∼80% of HSV-1 capsids attempting envelopment in the CAD cell body, in accord with the predictions of the married model. However, this number is likely to be an underestimate since the GFP-VPS4-EQ-mediated dominant negative envelopment block is not 100% efficient (22) and because any neurite-localized puncta bound to GFP-VPS4-EQ at a concentration similar to that in the cytoplasm would be ignored in our study. As described below (see Fig. 3), essentially all HSV-1 particles able to enter the neurites of CAD cells expressing GFP-VPS4-EQ also exhibit profoundly suppressed motion, suggesting that the vast majority of them (not just the 46% identified by imaging in Fig. 1E) are indeed associated with GFP-VPS4-EQ.
FIG 3.
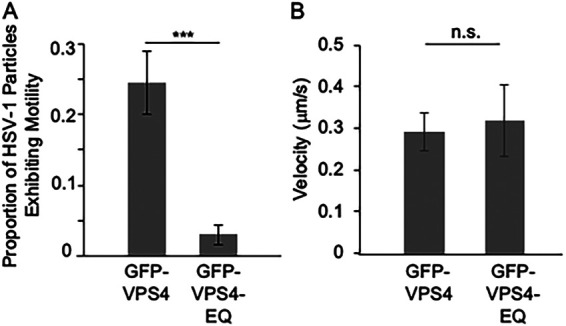
Motility of HSV-1 viral particles in the neurites of living CAD cells in the presence of GFP-VPS4 or GFP-VPS4-EQ. Differentiated, HSV-1-, and baculovirus-infected CAD cells similar to those in Fig. 2 were prepared and HSV-1-associated puncta imaged in living neurites. (A) Proportion of total HSV-1 puncta per neurite that moved a distance at least three times their own diameter during the 60-s period of image analysis. Plotted value is mean and standard deviation from the mean for neurites from 33 GFP-VPS4-expressing or 33 GFP-VPS4-EQ-expressing CAD cells containing a total of 470 and 273 puncta, respectively. (B) Mean velocity and standard deviation from the mean of a subset of the motile HSV-1 particles scored in panel A. The velocity of 10 motile puncta was measured in the neurites of GFP-VPS4-expressing CAD cells and eight in the neurites of GFP-VPS4-EQ-expressing CAD cells.
In HeLa cells, the expression of VPS4-EQ has been reported to result in a 2- to 3-fold increase in the numbers of enveloped HSV-1 capsids accumulating in the perinuclear space, interpreted as an inhibition of the extent (or possibly the kinetics) of egress from the nucleus (69). However, the same study showed that VPS4-EQ did not diminish the numbers of HSV-1 capsids reaching the cytoplasm (69), consistent with earlier quantitative electron and fluorescence microscopy data showing that VPS4-EQ had no effect on HSV-1 or PRV capsid egress from the nucleus in HEK293 cells (22, 50). Nevertheless, to test whether VPS4-EQ might be interfering with capsid egress from the CAD cell nucleus, we examined the cell bodies of HSV-1-infected CADs and quantitated the intensity of extranuclear mCherry-UL25 fluorescence. Although the intensity of cytoplasmic fluorescence varied from cell to cell, we found no significant difference (P = 0.74) between CAD cells expressing GFP-VPS4 and those expressing GFP-VPS4-EQ (Table 1).
TABLE 1.
Nonnuclear mCherry-UL25 fluorescence in HSV-1-infected CAD cell bodies
| Allele of VPS4 expressed | No. of cell bodies counteda | Mean extranuclear fluorescence (AU)b | SD |
|---|---|---|---|
| GFP-VPS4 | 32 | 27.2 | 9.4 |
| GFP-VPS4-EQ | 33 | 28.2 | 10.0 |
In each case, cell bodies were selected from 21 microscopic fields prepared from two independent experiments.
AU, arbitrary units of fluorescence.
Trafficking of the HSV-1 envelope glycoprotein gD to the cell surface is unaffected by GFP-VPS4-EQ expression.
Because the ESCRT apparatus is involved in many aspects of cellular organelle manipulation and intracellular protein trafficking (52–55), interference with ESCRT function by VPS4-EQ expression can have deleterious effects. To test whether intracellular protein trafficking is occurring normally under conditions in which HSV-1 envelopment and transport was affected, we examined delivery of the HSV-1-encoded envelope glycoprotein gD to the CAD cell surface. HSV-1-infected CAD cells were prepared and fixed for imaging in an identical manner to those in Fig. 1A to E and then cell-surface levels of gD detected using a mouse anti-gD antibody and a fluorescent anti-mouse secondary antibody. Levels of CAD cell-surface gD-derived fluorescence (including that on cell bodies and neurites) were then determined. As shown in Fig. 1F, the quantity of gD delivered to the cell surface was identical for CAD cells expressing GFP-VPS4 or GFP-VPS4-EQ, indicating no gross abnormalities in intracellular protein synthesis, processing, and vesicular trafficking. This study measured only gD that had been delivered to the cell surface, since under identical conditions, the extremely abundant intracellular antigen α-tubulin was essentially undetectable unless the CAD cells were permeabilized with detergent (Fig. 1G).
Delivery of GFP-VPS4 and GFP-VPS4-EQ to CAD cells using recombinant baculoviruses.
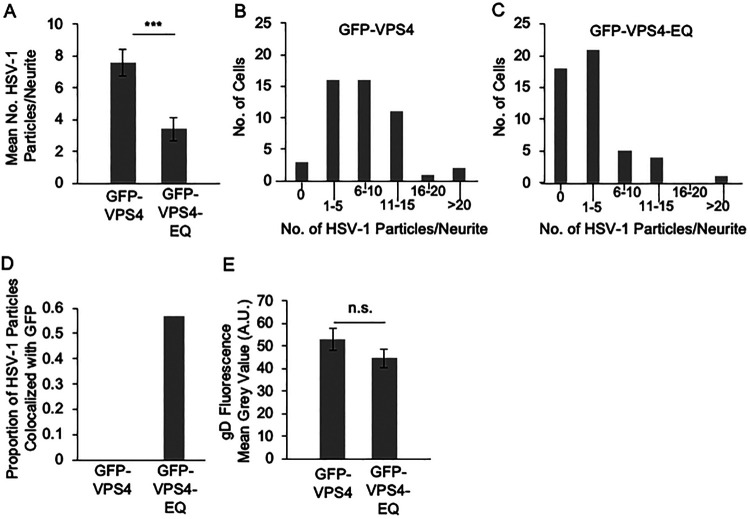
To extend our studies to a broader range of neurons, including those not readily susceptible to efficient plasmid transfection, we transferred the GFP-VPS4 and GFP-VPS4-EQ open reading frames to baculovirus expression vectors using the ViraPower BacMam expression system (Invitrogen, Life Technologies). These recombinant baculoviruses express GFP-VPS4 and GFP-VPS4-EQ from the cytomegalovirus (CMV) immediate early promoter, and the baculovirus particles are pseudotyped by the vesicular stomatitis virus G protein (VSV-G), supporting efficient entry into a range of neurons in addition to differentiated CAD cells (67). CAD cells were serum-starved to drive differentiation and formation of neurites, and then infected by HSV-1 and subsequently baculoviruses to express GFP-VPS4 or GFP-VPS4-EQ. After 20 h of HSV-1 infection, cells were fixed and mounted for imaging exactly as before. We once again counted the mean number of HSV-1 particles entering CAD cell neurites (Fig. 2A) and the distribution of the frequency with which HSV-1 particles were found in neurites in the presence of GFP-VPS4 or GFP-VPS4-EQ (Fig. 2B and C). We also scored the colocalization of HSV-1 particles in neurites with GFP-VPS4 or GFP-VPS4-EQ (Fig. 2D). Results were essentially identical for those seen in our plasmid transfection studies and suggested that at least ∼80% of HSV-1 particles are trafficking by the married pathway in CAD cells under these conditions. We also confirmed that gD trafficking to the host cell surface was unaffected by GFP-VPS4-EQ expression under these conditions (Fig. 2E).
FIG 2.
Transport of HSV-1 particles in the axons of CAD cells infected by baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. CAD cells were differentiated and then infected with HSV-1 and subsequently with baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. After 20 h, cells were fixed, incubated with primary and fluorescent secondary antibodies as appropriate, and imaged by fluorescence microscopy. (A) Number of mCherry-fluorescent HSV-1 capsid-associated puncta per neurite. Plotted values are the mean and standard deviation from the mean for neurites from 49 CAD cells expressing GFP-VPS4 (total of 374 puncta) or 49 CAD cells expressing GFP-VPS4-EQ (total of 169 puncta). (B, C) Frequency (y axes) at which neurites of infected CAD cells expressing GFP-VPS4 or GFP-VPS4-EQ (respectively) contain numbers of HSV-1 puncta in the ranges indicated (x axes). (D) Proportion of mCherry-fluorescent puncta, similar to those scored in panel A, exhibiting colocalization with GFP-VPS4 (total of 77 puncta) or GFP-VPS4-EQ fluorescence (total of 97 puncta) at levels enriched above that of background. (E) Cell surface anti-gD immunofluorescence intensity for fixed, nonpermeabilized HSV-1-infected CAD cells expressing GFP-VPS4 or GFP-VPS4-EQ.
In Fig. 1 and 2, we examined the consequences of VPS4-EQ-GFP expression for the numbers of HSV-1 particles found in neurites at 20 h postinfection. This endpoint analysis would allow slow-moving virions to gradually accumulate in the neurite over time and could, therefore, underestimate the consequences of VPS4-EQ-GFP expression for inhibition of rates of HSV-1 transport into and along the process. To address this issue, we infected differentiated CAD cells with HSV-1 and baculoviruses as above and then imaged neurite-associated HSV-1 viral particles in the living CAD cells as they trafficked in real time. We counted the total numbers of HSV-1 particles in neurites, scored how many were undergoing motion during the 60-s period of imaging, and determined their velocity. As shown in Fig. 3A, approximately 25% of the HSV-1 particles in the neurites of GFP-VPS4-expressing cells underwent motion as defined by travelling at least three times their diameter during the period of imaging. This is similar to the efficiency of microtubule-directed motion in our in vitro system (20, 22, 64, 68). The mean velocity of transport was approximately 0.3 μm/s (Fig. 3B), which is within 2-fold of that reported for HSV-1 anterograde transport in explanted human fetal dorsal root ganglia and primary mouse forebrain cortical neurons (0.56 to 0.83 μm/s and 0.44 to 0.8 μm/s, respectively) (36, 39). In contrast, of 273 viral particles imaged in the neurites of 33 VPS4-EQ-GFP-expressing CAD cells, only 10 of them, or ∼3% of the total, exhibited motility (Fig. 3A). Those few that were able to move did so with a velocity similar to puncta in control GFP-VPS4-expressing neurons (Fig. 3B), suggesting that they represent rare virions that were able to complete ESCRT-mediated envelopment despite the dominant negative effects of GFP-VPS4-EQ. We conclude that GFP-VPS4-EQ expression blocks, or reduces, the ability of essentially all HSV-1 particles (∼97%) to traffic along neurite microtubules. Those few capable of motility presumably accumulate in the neurites over time, giving rise to the reduced neurite entry phenotype seen in Fig. 1 and 2.
Consequences of GFP-VPS4 or GFP-VPS4-EQ expression for HSV-1 assembly and trafficking in embryonic rat primary cortical neurons.
Cryo-electron tomography studies of HSV-1-infected embryonic rat hippocampal axons reported the presence of predominantly naked capsids in axons and putative secondary envelopment sites at axon terminals, consistent with the separate model of transport (35). However, this conclusion has been challenged (25). We, therefore, decided to compare the assembly and trafficking of HSV-1 and PRV during GFP-VPS4 and GFP-VPS4-EQ expression in primary rat cortical neurons (PRCN) isolated from day 17 rat embryos.
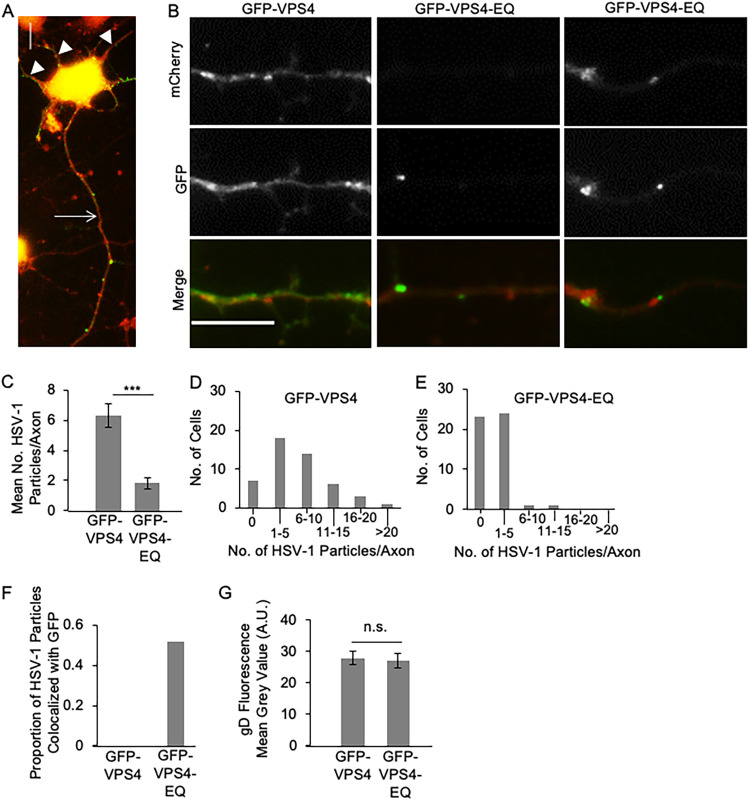
The majority of HSV-1-infected PRCN exhibited one long distinctive axonal process and several shorter dendrites (Fig. 4A). We were careful to count numbers of HSV-1 capsid-associated puncta exclusively in the unique long, axon-like process. Similar to our results in CAD neurons (Fig. 1 and 2), GFP-VPS4-EQ expression reduced the mean number of HSV-1 puncta entering the axon like processes of PRCN to about 28% of that in GFP-VPS4 control cells (Fig. 4A to C). This reflected a change in the frequency with which viral particles were distributed in the PRCN axons; approximately half of all GFP-VPS4-expressing control axons (23 of 48 counted) contained between 10 and 20 HSV-1-associated puncta, and fewer than 15% (7 of 48 counted) contained none at all (Fig. 4D). In contrast, half of all GFP-VPS4-EQ-expressing PRCN contained 5 or fewer viral particles in their axons, and the rest contained none (Fig. 4E). Of those virions able to reach the axon in the presence of the dominant negative allele of VPS4, approximately half colocalized with areas of GFP fluorescence enriched above that in the cytoplasm, suggesting the presence of bound GFP-VPS4-EQ (Fig. 4F). Delivery of the reporter gD to the PRCN cell surface was unaffected by GFP-VPS4-EQ, showing that there is no general defect in PRCN intercellular protein trafficking (Fig. 4G).
FIG 4.
Transport of HSV-1 particles in the axons of primary rat cortical neurons infected by baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. PRCN were isolated and infected with HSV-1 and baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. After 20 h, cells were fixed, incubated with primary and fluorescent secondary antibodies as appropriate, and puncta in the single long axon imaged by fluorescence microscopy. (A) Representative image of an HSV-1-infected PRCN showing the single, long axon-like process (white arrow) within which were scored viral puncta, and the cell body with short dendrite-like processes (white arrowheads). The image is a merge of the red channel (mCherry-tagged HSV-1 capsids) and green channel (GFP-VPS4). (B) Gallery of axons from GFP-VPS4-expressing (left column) or GFP-VPS4-EQ-expressing (middle and right columns) PRCN. Axons are imaged in the red channel to detect mCherry-tagged HSV-1 capsids (top row) or the green channel to detect GFP-tagged VPS4 alleles (middle row). Bottom row shows merged images. Scale bar, 10 μm. (C) Number of mCherry-fluorescent HSV-1 capsid-associated puncta per axon. Plotted values are the mean and standard deviation from the mean for 49 PRCN expressing GFP-VPS4 (total of 316 puncta) or 49 PRCN expressing GFP-VPS4-EQ (total of 89 puncta). (D, E) Frequency (y axes) at which axons of infected PRCN, expressing GFP-VPS4 or GFP-VPS4-EQ, respectively, contain numbers of HSV-1 puncta in the ranges indicated (x axes). (F) Proportion of mCherry-fluorescent puncta, similar to those scored in panel C, exhibiting colocalization with GFP-VPS4 (total of 47 puncta) or GFP-VPS4-EQ fluorescence (total of 33 puncta) at levels enriched above that of background. (G) Cell surface anti-gD immunofluorescence intensity for fixed, nonpermeabilized HSV-1-infected PRCN expressing GFP-VPS4 or GFP-VPS4-EQ.
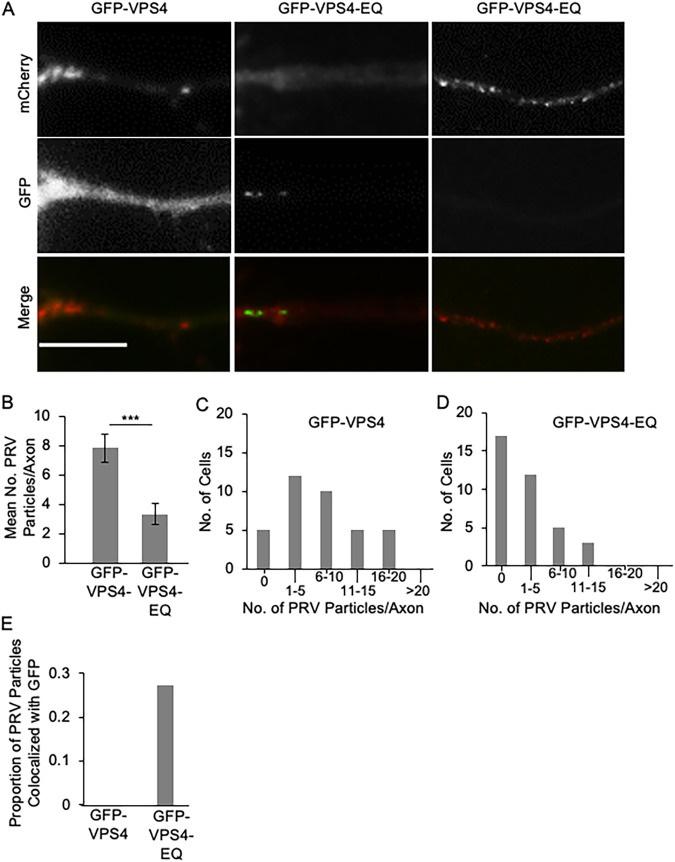
The data in Fig. 4 are consistent with at least ∼87% of HSV-1 particles utilizing the married pathway for egress. We next compared these data with the trafficking properties of PRV in PRCN, since PRV is considered to follow only the married model for transport. GFP-VPS4-EQ expression reduced the average number of PRV particles entering PRCN axons to 42% of that in control cells (Fig. 5A and B). In control, GFP-VPS4-expressing PRCN, over half of all axons (20 of 37 counted) contained between 6 and 20 PRV-associated puncta, and fewer than 15% of them (5 of 37 counted) contained no virions at all (Fig. 5C). In contrast, more than three-quarters of GFP-VPS4-EQ-expressing PRCN cells (29 of 37 counted) contained 5 or fewer viral particles in their axons, and almost half of them (17 of 37 counted) contained none (Fig. 5D). Of those reduced numbers of PRV puncta able to reach PRCN axons in the presence of GFP-VPS4-EQ, slightly over a quarter colocalized with regions of GFP fluorescence enriched above that in the cytoplasm, suggesting the presence of bound GFP-VPS4-EQ (Fig. 5E). Together, these data suggest that GFP-VPS4-EQ expression results in ∼69% of the PRV virions that would normally enter PRCN axons becoming trapped in the neuronal cell body, or entering the axon in association with GFP-VPS4-EQ, in accord with the predictions of the married model. We conclude that HSV-1 capsids utilize the married pathway for egress in PRCN neurons to at least the same extent as PRV (∼87% and ∼69% for HSV-1 and PRV, respectively).
FIG 5.
Transport of PRV particles in the axons of primary rat cortical neurons infected by baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. Similar study to that in Fig. 4 but examining the numbers of viral puncta in PRCN axons during a PRV infection. (A) Gallery of axons similar to that depicted in Fig. 4B, except that the red channel depicts mCherry-tagged PRV capsids. Scale bar, 10 μm. (B to D) Mean number of PRV puncta per axon and frequency distribution in the presence of GFP-VPS4 or GFP-VPS4-EQ. Axons were scored from PRV-infected PRCN-expressing GFP-VPS4 or GFP-VPS4-EQ (37 cells in each case, containing 290 and 120 axonal PRV puncta, respectively). (E) Proportion of mCherry-fluorescent puncta similar to those scored in panel B exhibiting colocalization with GFP-VPS4 (total of 74 puncta) or GFP-VPS4-EQ fluorescence (total of 77 puncta) at levels enriched above that of background.
Consequences of GFP-VPS4 or GFP-VPS4-EQ expression for HSV-1 and PRV assembly and trafficking in human SK-N-SH neuroblastoma cells.
Human neuroblastoma SK-N-SH cells are frequently described as an example of a neural cell line in which 80 to 95% of egressing HSV-1 particles traffic from the cell body along the neurite by the separate mechanism (32–34, 70). We therefore infected differentiated SK-N-SH cells with HSV-1 and examined viral trafficking in the presence of baculovirus-expressed GFP-VPS4 and GFP-VPS4-EQ (as before, we also demonstrated that delivery of the reporter envelope glycoprotein gD to the cell surface was unaffected by expression of GFP-VPS4 or GFP-VPS4-EQ [data not shown]). The results (Fig. 6) were strikingly different from those obtained for CAD cells and PRCN. In SK-N-SH cells, the expression of GFP-VPS4-EQ had no effect on the mean number of HSV-1 capsid-associated puncta entering neurites compared to that of the GFP-VPS4 control (Fig. 6A and B). Furthermore, the frequency distribution of virions within neurites was similar in GFP-VPS4 and GFP-VPS4-EQ-expressing cells (Fig. 6C and D). Some viral puncta were found to colocalize with enriched areas of GFP-VPS4-EQ (Fig. 6E); however, these were fewer than 20% of the HSV-1 particles in the neurites, a frequency two and a half times lower than that seen in CAD cells and PRCN (compare Fig. 1E, 2D, 4F, and 6E). Together, these data indicate that in SK-N-SH cells, HSV-1 transport into neurites is not detectably affected by the envelopment trap and that at least 80% of the viral particles (those not inhibited by or tagged with GFP-VPS4-EQ) appear to travel by the separate model. This quantity is strikingly similar to that reported in earlier studies of HSV-1 trafficking in this cell line (32).
FIG 6.
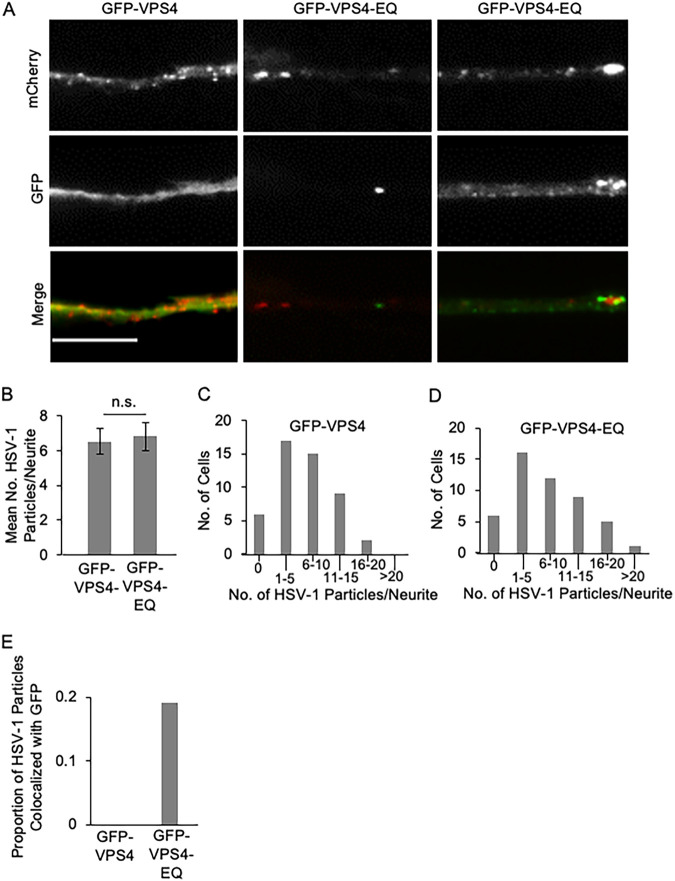
Transport of HSV-1 particles in the neurites of SK-N-SH cells infected by baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. Differentiated SK-N-SH cells were infected with HSV-1 and subsequently with baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. After 20 h, cells were fixed and puncta in the neurite imaged by fluorescence microscopy. (A) Gallery of neurites from GFP-VPS4-expressing (left column) or GFP-VPS4-EQ-expressing (middle and right columns) SK-N-SH cells. Neurites are imaged in the red channel to detect mCherry-tagged HSV-1 capsids (top row) or the green channel to detect GFP-tagged VPS4 alleles (middle row). Bottom row shows merged images. Scale bar, 10 μm. (B) Number of mCherry-fluorescent HSV-1 capsid-associated puncta per neurite. Plotted values are the mean and standard deviation from the mean for 49 SK-N-SH cells expressing GFP-VPS4 (total of 335 puncta) or 49 SK-N-SH cells expressing GFP-VPS4-EQ (total of 347 puncta). (C, D) Frequency (y axes) at which infected SK-N-SH cells expressing GFP-VPS4 or GFP-VPS4-EQ, respectively, contain numbers of HSV-1 puncta in their neurites in the ranges indicated (x axes). (E) Proportion of mCherry-fluorescent puncta, similar to those scored in panel B, exhibiting colocalization with GFP-VPS4 (total of 207 puncta) or GFP-VPS4-EQ fluorescence (total of 214 puncta) at levels enriched above that of background.
As detailed in the introduction, PRV is thought to only employ the married mechanism for egress (see the introduction for details). If this is the case, PRV transport into the neurites of SK-N-SH cells should be susceptible to the envelopment trap. We, therefore, repeated the above studies in PRV-infected SK-N-SH cells. As we observed for HSV-1, the expression of GFP-VPS4-EQ in SK-N-SH cells had no significant effect upon the mean number of PRV capsid-associated puncta within neurites (Fig. 7A and B) and did not alter the frequency distribution of virions within neurites (Fig. 7C and D) compared to the GFP-VPS4 control. Approximately 27% of PRV puncta colocalized with enriched areas of GFP-VPS4-EQ (Fig. 7E), which was slightly higher than that seen for HSV-1 (20%) (Fig. 6E). Unexpectedly, therefore, PRV appears to follow a similar mechanism of egress to HSV-1 in SK-N-SH cells, one in which most viral capsids are not arrested by the envelopment trap in the cell body and which is most consistent with the separate model of transport.
FIG 7.
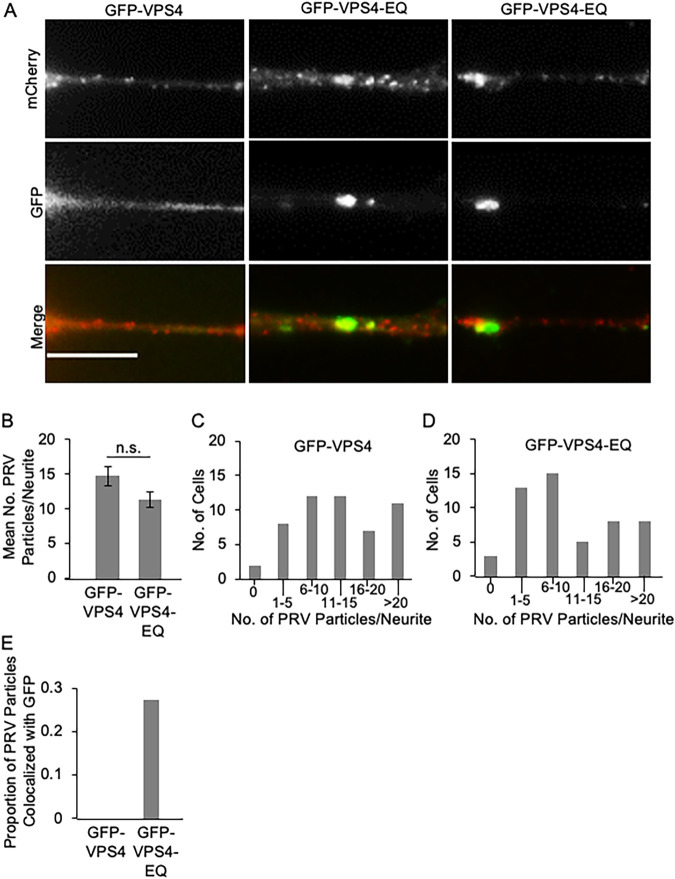
Transport of PRV particles in the neurites of SK-N-SH cells infected by baculoviruses expressing GFP-VPS4 or GFP-VPS4-EQ. Similar study to that in Fig. 6 but examining the numbers of viral puncta in SK-N-SH neurites during a PRV infection. (A) Gallery of neurites similar to that depicted in Fig. 6A, except that the red channel depicts mCherry-tagged PRV capsids. Scale bar, 10 μm. (B to D) Mean number of PRV puncta per neurite, and frequency distribution, in the presence of GFP-VPS4 or GFP-VPS4-EQ. PRV-infected, GFP-VPS4-expressing or GFP-VPS4-EQ-expressing, SK-N-SH cells were scored (52 cells in each case, containing 709 and 572 neurite-associated PRV puncta, respectively). (E) Proportion of mCherry-fluorescent puncta similar to those scored in panel B exhibiting colocalization with GFP-VPS4 (total of 135 puncta) or GFP-VPS4-EQ fluorescence (total of 209 puncta) at levels enriched above that of background.
DISCUSSION
Several complementary approaches have been used to determine the structure of alphaherpesvirus particles undergoing anterograde traffic in neuronal axons. These methods each have limitations; antibodies to capsid or tegument proteins may be subject to restricted access to enveloped capsids and tegument within carrier vesicles, while viral genetic studies are complicated by the possibility of redundancy with other viral genes or secondary effects resulting from a given gene product having multiple functions (25, 26, 42–44). Similarly, ultrastructural analyses describing naked HSV capsids in axons (35, 36) have been critiqued as possibly misidentifying neuronal dense core vesicles as capsids or as representing nonenveloped capsids within dendrites rather than axons (25), though these suggestions have been challenged (26).
In this study, we have taken an alternative, functional approach to this question, using the dominant negative VPS4-EQ molecule to arrest envelopment in alphaherpesvirus-infected cultured neural cell lines and explanted primary neurons. By manipulating a cellular protein rather than a viral one, we are able to perform all of our experiments using viruses that are wild type (other than the presence of a fluorescently tagged capsid subunit), and we can directly compare the behavior of HSV-1 and PRV in identical cells that are differentiated, cultured, and imaged under the same conditions. Moreover, the biological role and biochemical properties of VPS4 and its substrate ESCRT-III are understood in considerable detail, extremely highly conserved between cells (53, 54, 61, 71, 72), and expression of VPS4-EQ results in an envelope-trapped terminal capsid phenotype that is well defined (17, 22, 50, 51). The ability to block alphaherpesvirus envelopment and simultaneously GFP tag the envelopment intermediate using GFP-VPS4-EQ, therefore, makes it possible to perform a direct test of the married and separate pathways.
Differentiated mouse CAD neurons (40, 41, 65) are a well-established system for study of alphaherpesvirus egress and one in which HSV-1 has been proposed to traffic by the married model (42, 45, 66–68). Our data are consistent with this conclusion. When GFP-VPS4-EQ was expressed from plasmid or baculovirus vectors, at least 80% of trafficking HSV-1 capsids were either blocked from entering the axon-like neurite, exhibited greatly reduced frequency of motility within the neurite, or had become labeled with GFP, indicating a prior attempt to undergo envelopment (Fig. 1-3). Having established the efficacy of the envelopment trap in this system, we then extended our studies to two host cell backgrounds where HSV-1 has been reported to traffic by the separate model as detailed below.
On the basis of live cell imaging and cryo-electron tomography studies, Ibiricu and colleagues reported the presence of predominantly naked capsids in the axons of HSV-1-infected embryonic rat hippocampal neurons (35). However, we found that the envelopment trap reduced the number of HSV-1 puncta entering PRCN axons by 72%, and half of those that reached the axon were bound to GFP-VPS4-EQ, suggesting that they had previously encountered the envelopment apparatus (Fig. 4). In fact, HSV-1 entry into axons was inhibited by the envelopment trap to at least the same extent as seen for PRV (Fig. 4 and 5). These results are consistent with the married model for traffic in these neurons and are difficult to reconcile with earlier morphological data (35) unless those studies were visualizing capsid transport within dendrites as has been suggested (25).
The best characterized cell culture system for HSV-1 transport by the separate mechanism is the human neuroblastoma cell line SK-N-SH (32–34, 70). Consistent with the predictions of the separate model and in striking contrast to our results with HSV-1 in CAD and PRCN cells, the envelopment trap had no measurable effect upon HSV-1 entry into SK-N-SH neurites (Fig. 6). We next tested the effect of the envelopment trap upon PRV trafficking in SK-N-SH cells, expecting to find that GFP-VPS4-EQ expression would reduce the frequency of entry of PRV particles into neurites. To our surprise, the results for PRV and HSV-1 were identical (Fig. 6 and 7), suggesting that both of these alphaherpesviruses follow the separate pathway in this background. Remarkably, to our knowledge, this study is the first to directly compare HSV-1 and PRV trafficking side by side in SK-N-SH cells, using the same culture, differentiation, and imaging conditions.
How can we reconcile this finding with the general consensus that PRV utilizes the married mechanism (21, 24–27, 32) for egress? One possibility is that, as has been proposed for HSV-1 (24, 28, 32, 45, 46), PRV can employ both the separate and married transport pathways. However, there is no precedent for PRV traffic by the separate route in any other neuronal cell type. Given this, an attractive alternative explanation for our data is that cultured, differentiated SK-N-SH cells may fail to tightly restrict the delivery of cargo from the cell body into the neurite. It is certainly clear that nonenveloped HSV-1 and PRV capsids can recruit kinesin motors for anterograde traffic within the cytoplasm prior to envelopment (19, 23, 24, 31), and it is reasonable to suppose that such capsids would be capable of entering axons or neurites if the cargo-filtering zone in the proximal region of those processes is poorly established (73–75). This possibility has precedent in PRV egress from rat superior cervical ganglia (SCG). The transport of PRV virions from the cell body to the axons of mature rat SCG is critically dependent upon US9p (24, 76), a virally encoded membrane protein that may recruit the kinesin-3 motor KIF1A to the organelle surrounding the enveloped viral particle (24, 30, 68). The requirement for US9p is, however, considerably less stringent in immature SCG, showing that alternative transport pathways can deliver viral particles into the axon when neuronal differentiation is incomplete (77).
In summary, our use of an ESCRT-based trap to perturb capsid envelopment has revealed substantial differences in the trafficking properties of alphaherpesviruses among several different kinds of neuron and neural cell lines. Unexpectedly, we did not see differences between HSV-1 and PRV transport in these studies, and we hypothesize that data in support of the married or separate pathways for egress may reflect more upon the properties of the infected neuron than differences between the two alphaherpesviruses. In the past, the married or separate pathways have been assigned to egressing HSV-1 and PRV virions during infection of neurons isolated from a variety of sources and cultured and imaged by many different investigators (33, 34, 36, 37, 39, 42, 44). Future study of this question would benefit from a side by side, simultaneous comparison of HSV-1 and PRV egress from such neurons, combining ultrastructural and light microscopy analyses with functional perturbation of envelope assembly using approaches such as the one detailed here.
MATERIALS AND METHODS
CAD cell culture, transfection, differentiation, and infection.
CAD cells were maintained in Dulbecco’s modified Eagle medium (DMEM)/F12 containing 15 mM HEPES (Gibco Laboratories) and supplemented with 10% fetal calf serum (FCS) (Peak Serum, Inc.) and 1% penicillin-streptomycin (PS) (Gibco Laboratories). Where appropriate, undifferentiated CAD cells were transfected with the tetracycline-inducible expression vectors pTetOne-GFP-VPS4 and pTetOne-GFP-VPS4-EQ (see below) using Lipofectamine 3000 (Invitrogen) according to the manufacturers protocol. At 24 h posttransfection, they were processed for differentiation and infection as described below,
For differentiation, CAD cells (whether transfected or untransfected) were seeded at a density of 1.5 × 104 to 3 × 104 cells/cm2 onto glass coverslips precoated with 10 μg/mL poly-l-lysine (Sigma-Aldrich). After attachment, cells were washed twice with phosphate-buffered saline (PBS) (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH 7.4) and then incubated in differentiation medium comprising DMEM/F12 supplemented with 10 μg/mL transferrin (Sigma-Aldrich), 50 ng/mL sodium selenite (Sigma-Aldrich), and 1% PS. Differentiation medium was replaced after 3 days and cells infected with HSV-1 at a multiplicity of infection (MOI) of 20 after 5 days. Where appropriate, transfected plasmid gene expression was induced by addition of 200 ng/mL doxycycline 1 h after initiation of HSV-1 infection. Alternatively, baculoviruses were used to infect CAD cells at the same time as HSV-1 infection. After 20 h of HSV-1 infection, cells were fixed and processed for imaging or immunocytochemistry as described below.
PRCN isolation, culture, and infection.
Primary cortical neurons (PRCN) were prepared from embryonic day 18 rats as previously described (78). All experiments complied with all relevant ethical regulations for animal testing and research and were approved by the Albert Einstein College of Medicine Institutional Animal Care and Use Committee (IACUC, IACUC protocol number 0001209). Rats were housed and handled at the Albert Einstein College of Medicine. Euthanasia was done with a lethal dose of CO2 gas delivered by compressed CO2 canisters using the recommended gas flow rate prior to decapitation. The Panel on Euthanasia of the American Veterinary Medical Association recommends these procedures.
PRCN were seeded onto glass coverslips precoated with 100 μg/mL poly-l-lysine and were maintained in Neurobasal medium containing GlutaMAX, B-27 supplement (Gibco Laboratories), and 1% PS. PRCN were infected with HSV-1 or PRV at an MOI of 20 after 5 days of culture in vitro. Where appropriate, baculoviruses were used to infect the PRCN cells at the same time as infection with HSV-1 or PRV.
SK-N-SH cell culture, differentiation, and infection.
SK-N-SH cells were maintained in culture and transferred to glass coverslips precoated with poly-l-lysine, exactly as described above for CAD cells. For differentiation and infection, coverslips of SK-N-SH cells were washed twice with PBS and then changed to differentiation medium (5:1 mixture of DMEM/F12 and Medium 199 [Gibco Laboratories]) supplemented with 5% FCS, 1% PS, and 10 μM retinoic acid (Sigma-Aldrich). Differentiation medium was replaced on days 3 and 5. At day 7, cells were infected with HSV-1 or PRV at an MOI of 20. Where appropriate, baculoviruses were used to infect SK-N-SH cells at the same time as infection by HSV-1 or PRV.
HSV-1 and PRV strains and preparation of stocks.
HSV-1 strain GS6807 is derived from a self-excising strain F infectious clone, which encodes a UL25-mCherry capsid fusion (79) and has been previously described (56). Stocks were grown and titered on Vero cells grown in DMEM (Gibco Laboratories) supplemented with 10% newborn calf serum (NCS) (Peak Serum, Inc.) and 1% PS. PRV strain GS4282 was derived from the pBecker3 infectious clone (80), encodes a UL25-mCherry capsid fusion, and was previously described (68, 81). Working stocks were prepared on porcine kidney PK15 cells and titers determined on Vero cells as above. HSV-1 strain GS6807 and PRV strain GS4284 are both kind gifts from Gregory A. Smith.
For infection of CAD, PRCN, or SK-N-SH cells, virus stocks were prepared from the cell culture medium of HSV-1-infected Vero or PRV-infected PK15 cells as follows. After removal of infected cells by centrifugation at 2,000 × g, culture medium was centrifuged at 50,000 × g for 1 h and pelleted virus allowed to resuspended overnight in PBS at 4°C. The pellet was further resuspended by pipetting and sonication, and then incubated with 250 U Benzonase Nuclease (Sigma-Aldrich) for 1 h at 4°C before freezing viral aliquots at −80°C. Stocks were titrated on Vero cells.
Construction of tetracycline-inducible GFP-VPS4 and GFP-VPS4-EQ-expression plasmids.
Plasmids encoding GFP fusions to the amino termini of the A isoforms of wild-type VPS4 and VPS4-EQ, termed VPS4Awt and VPS4AEQ, respectively, have been previously described (82) and were a kind gift from Margaret Kielian. The GFP-VPS4 and GFP-VPS4-EQ open reading frames were amplified from these plasmids by conventional PCR techniques using the forward primer, 5′-CCCTCGTAAAGAATTCTAGAGCCACCATGGTGAGCAAG GGCGAGGAGCTG-3′, and the reverse primer, 5′-GCAGAGATCTGGATCCCTATTAACTCTCTTGCCCAAAGTCCTC-3′. These primers were designed so that the underlined termini were homologous to the termini of plasmid pTetOne (TaKaRa Bio USA, Inc.) following its digestion with the restriction enzymes EcoRI and BamHI. The digested pTetOne vector and the amplified GFP-VPS4 and GFP-VPS4-EQ DNA fragments were recombined using the In-Fusion HD cloning system (TaKaRa Bio USA, Inc.) generating plasmids pTetOne-GFP-VPS4 and pTetOne-GFP-VPS4-EQ.
Construction and propagation of recombinant baculoviruses expressing GFP-VPS4 and GFP-VPS4-EQ.
The GFP-VPS4 and GFP-VPS4-EQ open reading frames were amplified from plasmids pTetOne-GFP-VPS4 and pTetOne-GFP-VPS4-EQ by conventional PCR techniques using the forward primer 5′-CACCATGGTGAGCAAGGGCGAG-3′ and the reverse primer 5′-CTATTAACTCTCTTGCCCAAA-3′. The primers were designed so that the 5′ end of the PCR product (corresponding to the region that is underlined in the sequence of the forward primer) will be invaded by the single-stranded overhanging end of the commercially available linearized pENTR/d-TOPO Gateway entry vector (Invitrogen). The PCR products were then recombined with the pENTR/d-TOPO vector using pENTR directional TOPO cloning methodology as specified by the manufacturer (Invitrogen). The resulting pENTR/d-TOPO constructs were used to transfer the GFP-VPS4 and GFP-VPS4-EQ open reading frames into plasmid vector BacMam pCMV-DEST (ViraPower BacMam expression system; Invitrogen) by attR/attL LR recombination using methodology specified by the manufacturer (Invitrogen). Finally, the pCMV-DEST-based constructs were transformed into DH10Bac cells (Thermo Fisher) for transposition into a bacmid containing the baculovirus genome. Following isolation of desired recombinants, bacmid DNA was prepared and transfected into Sf9 insect cells (Invitrogen) using Cellfectin II reagent (Invitrogen).
Baculoviruses were propagated and amplified on cultures of Sf9 cells in suspension (1 × 106 cells/mL in Sf-900 III SFM medium supplemented with 100 U/mL PS [HyClone]) infected at an MOI from 0.05 to 0.1 and then incubated at 27°C for 3 days. Baculoviruses were concentrated by centrifugation for 1 h at 50,000 × g. Viral pellets were resuspended overnight in Sf-900 III SFM medium supplemented with 3% FCS, and then further resuspended by pipetting and sonication. Stocks were titrated on Sf9 cells with a 1% agarose overlay and plaques counted after 10 days.
Fluorescence and immunofluorescence imaging of infected, fixed cells.
Coverslips of HSV-1 or PRV-infected CAD, SK-N-SH, or PRCN cells were washed gently with PBS and fixed for 15 min at room temperature with 4% paraformaldehyde in PBS. Coverslips were washed again with PBS and either mounted with ProLong diamond antifade agent (Invitrogen) or processed for immunocytochemistry as follows. Cells were left unpermeabilized or permeabilized with 0.1% Triton X-100 in PBS for 15 min at room temperature, washed in PBS, and blocked in 20% NCS in PBS for 1 h. They were then incubated for 1 h with mouse monoclonal anti-gD (Abcam; Cat no. ab6507) or anti-α-tubulin (Sigma-Aldrich; Cat No. T9026) antibodies or with no primary antibody as a control. The coverslips were then washed and incubated for 1 h with Alexa Fluor Plus 647 donkey anti-mouse IgG (Invitrogen), washed with PBS, and mounted as above.
All imaging studies were performed in the Analytical Imaging Facility (AIF) of the Albert Einstein College of Medicine. Fluorescent images of fixed cells were collected on a Zeiss Axio Observer Z1 microscope using Axiovision software (for CAD cells transiently transfected with plasmids) or an Inverted Olympus IX81 microscope using IP Lab 4.0.8 software (all other studies), in both cases using a 60× 1.4 NA oil lens. Anti-gD and anti-α-tubulin immunofluorescence imaging conditions were chosen such that control samples, prepared in the absence of a primary antibody, appeared black. The intensity of anti-gD and anti-α-tubulin immunofluorescence was then measured using the Fiji software “mean gray value” analysis tool. For quantitation of mCherry-UL25 capsid fluorescence in the cytoplasm of the CAD cell body, infected CAD cells were fixed as described above, except that they were mounted using ProLong Diamond Antifade agent containing 4′,6-diamidino-2-phenylindole (DAPI). Cells expressing GFP-VPS4 or GFP-VPS4-EQ were identified by imaging in the green channel and DAPI fluorescence in the blue channel used to define the location of the nucleus. The intensity of cell body mCherry-UL25 fluorescence lying outside the perimeter of the nucleus was then measured using the Fiji software “mean gray value” analysis tool.
Numbers of fluorescent viral puncta in neurites were counted using the open-source software Fiji (http://fiji.sc) (83, 84). HSV-1 and PRV mCherry capsid colocalization with GFP-VPS4 or GFP-VPS4-EQ was scored using Fiji software as follows. Neurites or axons were imaged in the green and red channels and the “analyze particles” tool used to identify green or red puncta and mask lower intensity levels of background fluorescence in the cytoplasm. The masked red and green images were then combined to identify mCherry-tagged viral puncta that colocalized with above-background levels of GFP-VPS4 or GFP-VPS4-EQ fluorescence.
Live cell fluorescence imaging.
Undifferentiated CAD cells were seeded at a density of 0.5 × 105 to 1 × 105 cells/well in 2-well chambered coverslips (Ibidi) then differentiated and infected with HSV-1 and baculoviruses exactly as described above. After 20 h of HSV-1 infection, the cells were washed gently with prewarmed PBS and changed to DMEM/F12 medium lacking phenol red (Gibco Laboratories) and supplemented with 10% FCS. Living cells were imaged using an Inverted Nikon ECLIPSE Ti-E microscope and 60× air lens with heated stage using Nikon Elements software. Time lapse images were captured at 1-s or 2-s intervals for 60 to 120 s and compiled using Fiji software.
ACKNOWLEDGMENTS
This work was supported by National Institutes of Health grants R01 AI125244 (to D.W.W.) and R01 NS118820 (to B.A.J.). J.B. acknowledges support from the Institutional AIDS training grant, “Training in HIV/AIDS Pathogenesis; Basic and Translational Research” T32 AI007501.
Microscopic image collection and analysis was performed in the Analytical Imaging Facility of the Albert Einstein College of Medicine with help from Andrea Briceno and support from NCI cancer center grant P30CA013330.
Contributor Information
Duncan W. Wilson, Email: duncan.wilson@einsteinmed.edu.
Lori Frappier, University of Toronto.
REFERENCES
- 1.Heming JD, Conway JF, Homa FL. 2017. Herpesvirus capsid assembly and DNA packaging. Adv Anat Embryol Cell Biol 223:119–142. 10.1007/978-3-319-53168-7_6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Dai X, Zhou ZH. 2018. Structure of the herpes simplex virus 1 capsid with associated tegument protein complexes. Science 360:eaao7298. 10.1126/science.aao7298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.El Bilali N, Khadivjam B, Bonneil E, Thibault P, Lippe R. 2021. Proteomics of herpes simplex virus type 1 nuclear capsids. J Virol 95:e01842-19. 10.1128/JVI.01842-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ahmad I, Wilson DW. 2020. HSV-1 cytoplasmic envelopment and egress. Int J Mol Sci 21:5969. 10.3390/ijms21175969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Draganova EB, Thorsen MK, Heldwein EE. 2021. Nuclear egress. Curr Issues Mol Biol 41:125–170. 10.21775/cimb.041.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Draganova EB, Zhang J, Zhou ZH, Heldwein EE. 2020. Structural basis for capsid recruitment and coat formation during HSV-1 nuclear egress. Elife 9:e56627. 10.7554/eLife.56627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bigalke JM, Heldwein EE. 2017. Have NEC coat, will travel: structural basis of membrane budding during nuclear egress in herpesviruses. Adv Virus Res 97:107–141. 10.1016/bs.aivir.2016.07.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bigalke JM, Heldwein EE. 2016. Nuclear exodus: herpesviruses lead the way. Annu Rev Virol 3:387–409. 10.1146/annurev-virology-110615-042215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bigalke JM, Heldwein EE. 2015. Structural basis of membrane budding by the nuclear egress complex of herpesviruses. EMBO J 34:2921–2936. 10.15252/embj.201592359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Mettenleiter TC. 2016. Vesicular nucleo-cytoplasmic transport-herpesviruses as pioneers in cell biology. Viruses 8:266. 10.3390/v8100266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Banfield BW. 2019. Beyond the NEC: modulation of herpes simplex virus nuclear egress by viral and cellular components. Curr Clin Micro Rep 6:1–9. 10.1007/s40588-019-0112-7. [DOI] [Google Scholar]
- 12.Crump C. 2018. Virus assembly and egress of HSV. Adv Exp Med Biol 1045:23–44. 10.1007/978-981-10-7230-7_2. [DOI] [PubMed] [Google Scholar]
- 13.Owen DJ, Crump CM, Graham SC. 2015. Tegument assembly and secondary envelopment of alphaherpesviruses. Viruses 7:5084–5114. 10.3390/v7092861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hollinshead M, Johns HL, Sayers CL, Gonzalez-Lopez C, Smith GL, Elliott G. 2012. Endocytic tubules regulated by Rab GTPases 5 and 11 are used for envelopment of herpes simplex virus. EMBO J 31:4204–4220. 10.1038/emboj.2012.262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Henaff D, Radtke K, Lippe R. 2012. Herpesviruses exploit several host compartments for envelopment. Traffic 13:1443–1449. 10.1111/j.1600-0854.2012.01399.x. [DOI] [PubMed] [Google Scholar]
- 16.Harley CA, Dasgupta A, Wilson DW. 2001. Characterization of herpes simplex virus-containing organelles by subcellular fractionation: role for organelle acidification in assembly of infectious particles. J Virol 75:1236–1251. 10.1128/JVI.75.3.1236-1251.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Barnes J, Wilson DW. 2019. Seeking closure: how do herpesviruses recruit the cellular ESCRT apparatus? J Virol 93:e00392-19. 10.1128/JVI.00392-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wolfstein A, Nagel CH, Radtke K, Dohner K, Allan VJ, Sodeik B. 2006. The inner tegument promotes herpes simplex virus capsid motility along microtubules in vitro. Traffic 7:227–237. 10.1111/j.1600-0854.2005.00379.x. [DOI] [PubMed] [Google Scholar]
- 19.Radtke K, Kieneke D, Wolfstein A, Michael K, Steffen W, Scholz T, Karger A, Sodeik B. 2010. Plus- and minus-end directed microtubule motors bind simultaneously to herpes simplex virus capsids using different inner tegument structures. PLoS Pathog 6:e1000991. 10.1371/journal.ppat.1000991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Shanda SK, Wilson DW. 2008. UL36p is required for efficient transport of membrane-associated herpes simplex virus type 1 along microtubules. J Virol 82:7388–7394. 10.1128/JVI.00225-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Smith G. 2012. Herpesvirus transport to the nervous system and back again. Annu Rev Microbiol 66:153–176. 10.1146/annurev-micro-092611-150051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kharkwal H, Smith CG, Wilson DW. 2014. Blocking ESCRT-mediated envelopment inhibits microtubule-dependent trafficking of alphaherpesviruses in vitro. J Virol 88:14467–14478. 10.1128/JVI.02777-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wilson DW. 2021. Motor skills: recruitment of kinesins, myosins and dynein during assembly and egress of alphaherpesviruses. Viruses 13:1622. 10.3390/v13081622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Diwaker D, Wilson DW. 2019. Microtubule-dependent trafficking of alphaherpesviruses in the nervous system: the ins and outs. Viruses 11:1165. 10.3390/v11121165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kratchmarov R, Taylor MP, Enquist LW. 2012. Making the case: married versus separate models of alphaherpes virus anterograde transport in axons. Rev Med Virol 22:378–391. 10.1002/rmv.1724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cunningham A, Miranda-Saksena M, Diefenbach R, Johnson D. 2013. Letter in response to: making the case: married versus separate models of alphaherpes virus anterograde transport in axons. Rev Med Virol 23:414–418. 10.1002/rmv.1760. [DOI] [PubMed] [Google Scholar]
- 27.Johnson DC, Baines JD. 2011. Herpesviruses remodel host membranes for virus egress. Nat Rev Microbiol 9:382–394. 10.1038/nrmicro2559. [DOI] [PubMed] [Google Scholar]
- 28.DuRaine G, Johnson DC. 2021. Anterograde transport of alpha-herpesviruses in neuronal axons. Virology 559:65–73. 10.1016/j.virol.2021.02.011. [DOI] [PubMed] [Google Scholar]
- 29.Kratchmarov R, Kramer T, Greco TM, Taylor MP, Ch'ng TH, Cristea IM, Enquist LW. 2013. Glycoproteins gE and gI are required for efficient KIF1A-dependent anterograde axonal transport of alphaherpesvirus particles in neurons. J Virol 87:9431–9440. 10.1128/JVI.01317-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kramer T, Greco TM, Taylor MP, Ambrosini AE, Cristea IM, Enquist LW. 2012. Kinesin-3 mediates axonal sorting and directional transport of alphaherpesvirus particles in neurons. Cell Host Microbe 12:806–814. 10.1016/j.chom.2012.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sandbaumhuter M, Dohner K, Schipke J, Binz A, Pohlmann A, Sodeik B, Bauerfeind R. 2013. Cytosolic herpes simplex virus capsids not only require binding inner tegument protein pUL36 but also pUL37 for active transport prior to secondary envelopment. Cell Microbiol 15:248–269. 10.1111/cmi.12075. [DOI] [PubMed] [Google Scholar]
- 32.Wisner TW, Sugimoto K, Howard PW, Kawaguchi Y, Johnson DC. 2011. Anterograde transport of herpes simplex virus capsids in neurons by both separate and married mechanisms. J Virol 85:5919–5928. 10.1128/JVI.00116-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Snyder A, Wisner TW, Johnson DC. 2006. Herpes simplex virus capsids are transported in neuronal axons without an envelope containing the viral glycoproteins. J Virol 80:11165–11177. 10.1128/JVI.01107-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Snyder A, Bruun B, Browne HM, Johnson DC. 2007. A herpes simplex virus gD-YFP fusion glycoprotein is transported separately from viral capsids in neuronal axons. J Virol 81:8337–8340. 10.1128/JVI.00520-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ibiricu I, Huiskonen JT, Dohner K, Bradke F, Sodeik B, Grunewald K. 2011. Cryo electron tomography of herpes simplex virus during axonal transport and secondary envelopment in primary neurons. PLoS Pathog 7:e1002406. 10.1371/journal.ppat.1002406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Penfold ME, Armati P, Cunningham AL. 1994. Axonal transport of herpes simplex virions to epidermal cells: evidence for a specialized mode of virus transport and assembly. Proc Natl Acad Sci USA 91:6529–6533. 10.1073/pnas.91.14.6529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Miranda-Saksena M, Boadle RA, Aggarwal A, Tijono B, Rixon FJ, Diefenbach RJ, Cunningham AL. 2009. Herpes simplex virus utilizes the large secretory vesicle pathway for anterograde transport of tegument and envelope proteins and for viral exocytosis from growth cones of human fetal axons. J Virol 83:3187–3199. 10.1128/JVI.01579-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Saksena MM, Wakisaka H, Tijono B, Boadle RA, Rixon F, Takahashi H, Cunningham AL. 2006. Herpes simplex virus type 1 accumulation, envelopment, and exit in growth cones and varicosities in mid-distal regions of axons. J Virol 80:3592–3606. 10.1128/JVI.80.7.3592-3606.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Dong X, Zhou J, Qin HB, Xin B, Huang ZL, Li YY, Xu XM, Zhao F, Zhao CJ, Liu JJ, Luo MH, Zeng WB. 2020. Anterograde viral tracer herpes simplex virus 1 strain H129 transports primarily as capsids in cortical neuron axons. J Virol 94:e01957-19. 10.1128/JVI.01957-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Qi Y, Wang JK, McMillian M, Chikaraishi DM. 1997. Characterization of a CNS cell line, CAD, in which morphological differentiation is initiated by serum deprivation. J Neurosci 17:1217–1225. 10.1523/JNEUROSCI.17-04-01217.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Li Y, Hou LX, Aktiv A, Dahlstrom A. 2007. Studies of the central nervous system-derived CAD cell line, a suitable model for intraneuronal transport studies? J Neurosci Res 85:2601–2609. 10.1002/jnr.21216. [DOI] [PubMed] [Google Scholar]
- 42.Antinone SE, Zaichick SV, Smith GA. 2010. Resolving the assembly state of herpes simplex virus during axon transport by live-cell imaging. J Virol 84:13019–13030. 10.1128/JVI.01296-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Antinone SE, Smith GA. 2010. Retrograde axon transport of herpes simplex virus and pseudorabies virus: a live-cell comparative analysis. J Virol 84:1504–1512. 10.1128/JVI.02029-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Buch A, Muller O, Ivanova L, Dohner K, Bialy D, Bosse JB, Pohlmann A, Binz A, Hegemann M, Nagel CH, Koltzenburg M, Viejo-Borbolla A, Rosenhahn B, Bauerfeind R, Sodeik B. 2017. Inner tegument proteins of herpes simplex virus are sufficient for intracellular capsid motility in neurons but not for axonal targeting. PLoS Pathog 13:e1006813. 10.1371/journal.ppat.1006813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.DuRaine G, Wisner TW, Howard P, Williams M, Johnson DC. 2017. Herpes simplex virus gE/gI and US9 promote both envelopment and sorting of virus particles in the cytoplasm of neurons, two processes that precede anterograde transport in axons. J Virol 91:e00050-17. 10.1128/JVI.00050-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Negatsch A, Granzow H, Maresch C, Klupp BG, Fuchs W, Teifke JP, Mettenleiter TC. 2010. Ultrastructural analysis of virion formation and intraaxonal transport of herpes simplex virus type 1 in primary rat neurons. J Virol 84:13031–13035. 10.1128/JVI.01784-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Votteler J, Sundquist WI. 2013. Virus budding and the ESCRT pathway. Cell Host Microbe 14:232–241. 10.1016/j.chom.2013.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Scourfield EJ, Martin-Serrano J. 2017. Growing functions of the ESCRT machinery in cell biology and viral replication. Biochem Soc Trans 45:613–634. 10.1042/BST20160479. [DOI] [PubMed] [Google Scholar]
- 49.Pawliczek T, Crump CM. 2009. Herpes simplex virus type 1 production requires a functional ESCRT-III complex but is independent of TSG101 and ALIX expression. J Virol 83:11254–11264. 10.1128/JVI.00574-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Crump CM, Yates C, Minson T. 2007. Herpes simplex virus type 1 cytoplasmic envelopment requires functional Vps4. J Virol 81:7380–7387. 10.1128/JVI.00222-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kharkwal H, Smith CG, Wilson DW. 2016. Herpes simplex virus capsid localization to ESCRT-VPS4 complexes in the presence and absence of the large tegument protein UL36p. J Virol 90:7257–7267. 10.1128/JVI.00857-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Alonso YAM, Migliano SM, Teis D. 2016. ESCRT-III and Vps4: a dynamic multipurpose tool for membrane budding and scission. FEBS J 283:3288–3302. 10.1111/febs.13688. [DOI] [PubMed] [Google Scholar]
- 53.Christ L, Raiborg C, Wenzel EM, Campsteijn C, Stenmark H. 2017. Cellular functions and molecular mechanisms of the ESCRT membrane-scission machinery. Trends Biochem Sci 42:42–56. 10.1016/j.tibs.2016.08.016. [DOI] [PubMed] [Google Scholar]
- 54.McCullough J, Frost A, Sundquist WI. 2018. Structures, functions, and dynamics of ESCRT-III/Vps4 membrane remodeling and fission complexes. Annu Rev Cell Dev Biol 34:85–109. 10.1146/annurev-cellbio-100616-060600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Olmos Y, Carlton JG. 2016. The ESCRT machinery: new roles at new holes. Curr Opin Cell Biol 38:1–11. 10.1016/j.ceb.2015.12.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Barnes J, Wilson DW. 2020. The ESCRT-II subunit EAP20/VPS25 and the Bro1 domain proteins HD-PTP and BROX are individually dispensable for herpes simplex virus 1 replication. J Virol 94:e01641-19. 10.1128/JVI.01641-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Butt BG, Owen DJ, Jeffries CM, Ivanova L, Hill CH, Houghton JW, Ahmed MF, Antrobus R, Svergun DI, Welch JJ, Crump CM, Graham SC. 2020. Insights into herpesvirus assembly from the structure of the pUL7:pUL51 complex. Elife 9:e53789. 10.7554/eLife.53789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.McCullough J, Clippinger AK, Talledge N, Skowyra ML, Saunders MG, Naismith TV, Colf LA, Afonine P, Arthur C, Sundquist WI, Hanson PI, Frost A. 2015. Structure and membrane remodeling activity of ESCRT-III helical polymers. Science 350:1548–1551. 10.1126/science.aad8305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Sun S, Li L, Yang F, Wang X, Fan F, Yang M, Chen C, Li X, Wang HW, Sui SF. 2017. Cryo-EM structures of the ATP-bound Vps4(E233Q) hexamer and its complex with Vta1 at near-atomic resolution. Nat Commun 8:16064. 10.1038/ncomms16064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Han H, Monroe N, Votteler J, Shakya B, Sundquist WI, Hill CP. 2015. Binding of substrates to the central pore of the Vps4 ATPase is autoinhibited by the microtubule interacting and trafficking (MIT) domain and activated by MIT interacting motifs (MIMs). J Biol Chem 290:13490–13499. 10.1074/jbc.M115.642355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Han H, Hill CP. 2019. Structure and mechanism of the ESCRT pathway AAA+ ATPase Vps4. Biochem Soc Trans 47:37–45. 10.1042/BST20180260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Davies BA, Azmi IF, Payne J, Shestakova A, Horazdovsky BF, Babst M, Katzmann DJ. 2010. Coordination of substrate binding and ATP hydrolysis in Vps4-mediated ESCRT-III disassembly. Mol Biol Cell 21:3396–3408. 10.1091/mbc.E10-06-0512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Kharkwal H, Furgiuele SS, Smith CG, Wilson DW. 2015. Herpes simplex virus capsid-organelle association in the absence of the large tegument protein UL36p. J Virol 89:11372–11382. 10.1128/JVI.01893-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Lee GE, Murray JW, Wolkoff AW, Wilson DW. 2006. Reconstitution of herpes simplex virus microtubule-dependent trafficking in vitro. J Virol 80:4264–4275. 10.1128/JVI.80.9.4264-4275.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Hashemi SH, Li JY, Faigle R, Dahlstrom A. 2003. Adrenergic differentiation and SSR2a receptor expression in CAD-cells cultured in serum-free medium. Neurochem Int 42:9–17. 10.1016/s0197-0186(02)00065-7. [DOI] [PubMed] [Google Scholar]
- 66.DuRaine G, Wisner TW, Johnson DC. 2020. Characterization of the herpes simplex virus (HSV) tegument proteins that bind to gE/gI and US9, which promote assembly of HSV and transport into neuronal axons. J Virol 94:e01113-20. 10.1128/JVI.01113-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.DuRaine G, Wisner TW, Howard P, Johnson DC. 2018. Kinesin-1 proteins KIF5A, -5B, and -5C promote anterograde transport of herpes simplex virus enveloped virions in axons. J Virol 92:e01269-18. 10.1128/JVI.01269-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Diwaker D, Murray JW, Barnes J, Wolkoff AW, Wilson DW. 2020. Deletion of the pseudorabies virus gE/gI-US9p complex disrupts kinesin KIF1A and KIF5C recruitment during egress, and alters the properties of microtubule-dependent transport in vitro. PLoS Pathog 16:e1008597. 10.1371/journal.ppat.1008597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Arii J, Watanabe M, Maeda F, Tokai-Nishizumi N, Chihara T, Miura M, Maruzuru Y, Koyanagi N, Kato A, Kawaguchi Y. 2018. ESCRT-III mediates budding across the inner nuclear membrane and regulates its integrity. Nat Commun 9:3379. 10.1038/s41467-018-05889-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Snyder A, Polcicova K, Johnson DC. 2008. Herpes simplex virus gE/gI and US9 proteins promote transport of both capsids and virion glycoproteins in neuronal axons. J Virol 82:10613–10624. 10.1128/JVI.01241-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Schoneberg J, Pavlin MR, Yan S, Righini M, Lee IH, Carlson LA, Bahrami AH, Goldman DH, Ren X, Hummer G, Bustamante C, Hurley JH. 2018. ATP-dependent force generation and membrane scission by ESCRT-III and Vps4. Science 362:1423–1428. 10.1126/science.aat1839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Su M, Guo EZ, Ding X, Li Y, Tarrasch JT, Brooks CL, III, Xu Z, Skiniotis G. 2017. Mechanism of Vps4 hexamer function revealed by cryo-EM. Sci Adv 3:e1700325. 10.1126/sciadv.1700325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Gumy LF, Katrukha EA, Grigoriev I, Jaarsma D, Kapitein LC, Akhmanova A, Hoogenraad CC. 2017. MAP2 defines a pre-axonal filtering zone to regulate KIF1- versus KIF5-dependent cargo transport in sensory neurons. Neuron 94:347–362. 10.1016/j.neuron.2017.03.046. [DOI] [PubMed] [Google Scholar]
- 74.Leterrier C. 2016. The axon initial segment, 50 years later: a nexus for neuronal organization and function. Curr Top Membr 77:185–233. 10.1016/bs.ctm.2015.10.005. [DOI] [PubMed] [Google Scholar]
- 75.Leterrier C. 2018. The axon initial segment: an updated viewpoint. J Neurosci 38:2135–2145. 10.1523/JNEUROSCI.1922-17.2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Taylor MP, Kramer T, Lyman MG, Kratchmarov R, Enquist LW. 2012. Visualization of an alphaherpesvirus membrane protein that is essential for anterograde axonal spread of infection in neurons. mBio 3:e00063-12. 10.1128/mBio.00063-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Tanneti NS, Federspiel JD, Cristea IM, Enquist LW. 2020. The axonal sorting activity of pseudorabies virus Us9 protein depends on the state of neuronal maturation. PLoS Pathog 16:e1008861. 10.1371/journal.ppat.1008861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Osten P, Khatri L, Perez JL, Kohr G, Giese G, Daly C, Schulz TW, Wensky A, Lee LM, Ziff EB. 2000. Mutagenesis reveals a role for ABP/GRIP binding to GluR2 in synaptic surface accumulation of the AMPA receptor. Neuron 27:313–325. 10.1016/s0896-6273(00)00039-8. [DOI] [PubMed] [Google Scholar]
- 79.Richards AL, Sollars PJ, Smith GA. 2016. New tools to convert bacterial artificial chromosomes to a self-excising design and their application to a herpes simplex virus type 1 infectious clone. BMC Biotechnol 16:64. 10.1186/s12896-016-0295-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Smith GA, Enquist LW. 2000. A self-recombining bacterial artificial chromosome and its application for analysis of herpesvirus pathogenesis. Proc Natl Acad Sci USA 97:4873–4878. 10.1073/pnas.080502497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Bohannon KP, Sollars PJ, Pickard GE, Smith GA. 2012. Fusion of a fluorescent protein to the pUL25 minor capsid protein of pseudorabies virus allows live-cell capsid imaging with negligible impact on infection. J Gen Virol 93:124–129. 10.1099/vir.0.036145-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Taylor GM, Hanson PI, Kielian M. 2007. Ubiquitin depletion and dominant-negative VPS4 inhibit rhabdovirus budding without affecting alphavirus budding. J Virol 81:13631–13639. 10.1128/JVI.01688-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A. 2012. Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682. 10.1038/nmeth.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Schneider CA, Rasband WS, Eliceiri KW. 2012. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]