Abstract
The mechanisms underlying memory loss associated with Alzheimer’s disease and related dementias (ADRD) remain unclear, and no effective treatments exist. Fundamental studies have shown that a set of transcriptional regulatory proteins of the nuclear receptor 4a (Nr4a) family serve as molecular switches for long-term memory. Here, we show that Nr4a proteins regulate the transcription of genes encoding chaperones that localize to the endoplasmic reticulum (ER). These chaperones fold and traffic plasticity-related proteins to the cell surface during long-lasting forms of synaptic plasticity and memory. Dysregulation of Nr4a transcription factors and ER chaperones is linked to ADRD, and overexpressing Nr4a1 or the chaperone Hspa5 ameliorates long-term memory deficits in a tau-based mouse model of ADRD, pointing toward innovative therapeutic approaches for treating memory loss. Our findings establish a unique molecular concept underlying long-term memory and provide insights into the mechanistic basis of cognitive deficits in dementia.
ER chaperones act as a long-term memory switch, and their dysregulation drives cognitive deficits in Alzheimer’s-related dementias.
INTRODUCTION
Impaired memory consolidation and the resulting long-term memory loss is an early symptom of Alzheimer’s disease and related dementias (ADRD) (1–3). Memory consolidation requires the transcription of new genes, in sophisticated spatial and temporal patterns, under the control of specific families of transcription factors (4–7). The largest class of transcription regulators in metazoans is composed of the nuclear receptor superfamily (8), which regulates diverse biological processes ranging from metabolism and reproduction to development and neuronal function. Among the several subclasses of nuclear receptors, the Nr4a subfamily [Nr4a1 (NUR77, NGF-IB), Nr4a2 (NURR1/HZF-3/NOT/RNR1), and Nr4a3 (NOR1)] has emerged as a critical mediator of long-term memory (4, 7). Notably, these ligand-independent “orphan” receptors are robustly up-regulated in the hippocampus within minutes after learning (3, 4). The learning-dependent expression of the Nr4a genes is regulated by histone acetylation, which is driven by recruitment of adenosine 3′,5′-monophosphate (cAMP) response element–binding protein (CREB)–binding protein (CBP) (9) to CREB response elements in the promoters of these genes (10, 11). Blocking the expression or inactivating the transactivation function of Nr4a factors is sufficient to impair long-term memory (4, 7) and synaptic plasticity (12), whereas transgenic or pharmacological activation enhances long-term memory (13, 14). Moreover, Nr4a function is impaired in brain disorders characterized by debilitating cognitive impairment, ranging from schizophrenia to Parkinson’s disease to ADRD (3, 4, 15, 16). However, despite the critical importance of the Nr4a subfamily, the effector genes that these transcription factors regulate in the hippocampus during memory consolidation have remained elusive. Here, we identify endoplasmic reticulum (ER) chaperone genes as downstream effector genes regulated by these transcription factors, and we establish a role for chaperone function in long-term memory and synaptic plasticity. We further demonstrate that Nr4a transcription factors and the downstream ER chaperones that they regulate are key mediators of the long-term memory loss associated with ADRD, providing candidate targets for the development of innovative therapeutic interventions.
RESULTS
Nr4a transcription factors regulate expression of genes encoding ER chaperones during memory consolidation
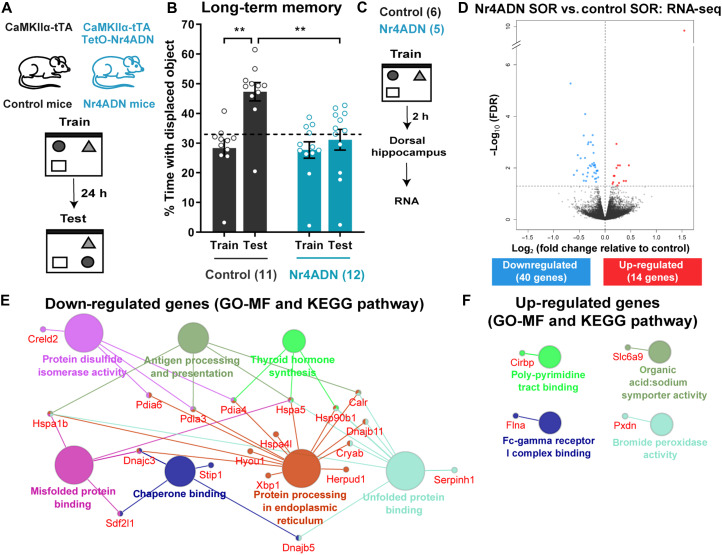
To identify effector genes regulated by the Nr4a subfamily during memory consolidation, we used transgenic mice that express a dominant-negative form of Nr4a1 (CaMKIIα-tTA TetO-Nr4ADN) in excitatory neurons such that the transcriptional activity of all three Nr4a family members is blocked in these cells (4). To assess hippocampus-dependent memory, we examined the performance of Nr4ADN mice in spatial object recognition (SOR), a task that depends on the preference of mice to explore a spatially displaced object (17). In a 24-hour test of long-term memory, the control mice (CaMKIIα-tTA), but not Nr4ADN mice (double transgenic: CaMKIIα-tTA, TetO-Nr4ADN), exhibited preference for the displaced object (Fig. 1, A and B). In contrast, in a 1-hour test of short-term memory, both the Nr4ADN and control mice showed a preference for the displaced object (fig. S1). Thus, Nr4ADN mice exhibit selective deficits in long-term spatial memory. To identify genes regulated by Nr4a transcription factors during memory consolidation, we trained Nr4ADN and control littermates in the SOR task and extracted total RNA from the dorsal hippocampus 2 hours after training (Fig. 1C). We chose this time point to identify effector genes targeted by the Nr4a subfamily of transcription factors, which are immediate early genes induced within minutes after training. This analysis revealed 54 differentially expressed genes (DEGs) (Fig. 1D and table S1) in Nr4ADN versus control mice, with 40 down-regulated and 14 up-regulated genes in Nr4ADN mice after learning (Fig. 1D).
Fig. 1. A multiprotein ER chaperone complex is downstream of Nr4a transcription factors during memory consolidation.
(A) Schematic depicting the SOR procedure. Mice expressing the tetracycline transactivator (tTA) protein under the CaMKIIα promoter (CaMKIIα-tTA: control mice) and littermates expressing both CaMKIIα-tTA and the dominant-negative mutant Nr4A1 under the control of TetO promoter (CaMKIIα-tTA, TetO-Nr4A dominant negative: Nr4ADN mice) were trained in SOR and then tested after 24 hours. (B) Preference for the displaced object (DO; dotted line marks 33% chance) during training and testing. Two-way ANOVA: significant main effect of genotype (F1,21 = 18.42, P = 0.0003) and significant main effect of sessions (F1,21 = 8.417, P = 0.0085). Sidak’s multiple comparisons test: **P = 0.0054 (control mice, train versus 24-hour test), **P = 0.0011 (control 24-hour test versus Nr4ADN 24-hour test). Control: n = 11 (3F); Nr4ADN: 12 (4F). (C) Schematic depiction of RNA-seq experiment. Nr4ADN and control male mice were trained in SOR and euthanized 2 hours after training. mRNA was harvested from the dorsal hippocampus and processed for the preparation of an RNA-seq library. (D) Volcano plot illustrating significance (y axis) and magnitude (x axis) of the down-regulation (blue) and up-regulation (red) of genes in Nr4ADN mice. (E and F) Functional groupings of network of enriched categories for genes whose differential expression [(E), down-regulation; (F), up-regulation] was significant, using the ClueGO and CluePedia plugins of the Cytoscape software. Gene Ontology (GO) terms include Molecular Functions (MF) and Kyoto Encyclopedia of Genes and Genomes (KEGG) and are represented as nodes (κ score level ≥ 0.4), with node size representing the significance of the term enrichment. Only the most significant term in each group is shown.
Enrichment network analysis was used to identify the pathways most represented among the down- and up-regulated genes. The down-regulated pathways included protein processing in the ER, chaperone binding, protein disulfide isomerase activity, and several other pathways related to protein folding in the ER (Fig. 1E). The up-regulated pathways included the poly-pyramidine tract binding pathway linked to Cirbp expression, an RNA binding protein associated with translational control (Fig. 1F). Protein-protein interaction (PPI) analysis of the down-regulated genes in Nr4ADN mice identified a significant cluster composed of nine chaperone proteins (Hspa5, Hsp90b1, Pdia3, Pdia4, Pdia6, Sdf2l1, Dnajb11, Hyou1, and Calr; fig. S2). All of these are components of a large ER multiprotein chaperone complex known to bind nascent proteins (18).
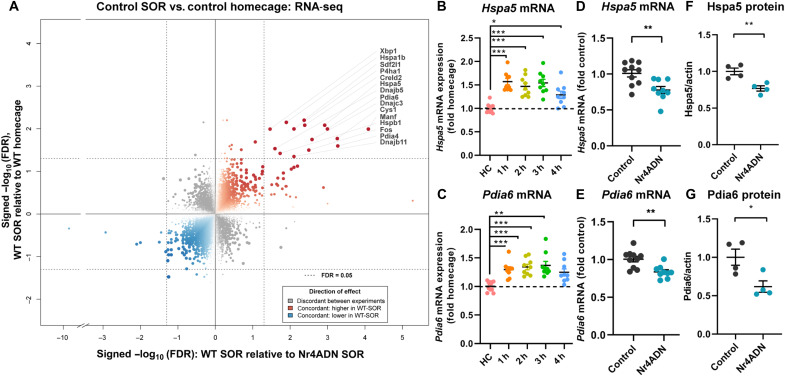
Next, we performed RNA sequencing (RNA-seq) using the dorsal hippocampus from control mice trained in SOR (SOR training +2 hours) or untrained mice [homecage (HC)]. RNA-seq analysis revealed that learning increased the expression of 42 genes and reduced expression of 9 genes (fig. S3 and table S2). Comparison of this gene expression data with the data from control and Nr4ADN mice after learning (Fig. 1D) identified 15 genes induced in control mice after learning that were down-regulated in Nr4ADN mice. These genes included ER chaperone genes Hspa5, Pdia4, Pdia6, Sdf2l1, and Dnajb11 (Fig. 2A). We next analyzed Nr4a1 occupancy on two of these candidate genes that are critical for protein folding (Hspa5 and Pdia6) (18) using data from a previously published Nr4a1 chromatin immunoprecipitation followed by sequencing (ChIP-seq) study (19). The promoters of the Hspa5 and Pdia6 genes were found to be enriched for Nr4a1 binding motifs (fig. S4), suggesting that this transcription factor directly regulates the expression of these genes. We further examined the expression profiles of these two candidate genes during the first 4 hours after learning in wild-type (C57BL/6J) mice, revealing that spatial learning induced the expression of both Hspa5 and Pdia6 genes (Fig. 2, B and C). Subsequent investigation of these two candidate genes in Nr4ADN mice confirmed that their regulation by the Nr4a proteins occurs only after learning (Fig. 2, D and E, and fig. S5). This regulation was not observed when expression of the Nr4ADN transgene was suppressed by treatment of the mice with doxycycline (fig. S6). These convergent data demonstrate that Nr4a transcription factors regulate the expression of a discrete set of ER chaperone genes during memory consolidation. Although these ER chaperones are known regulators of ER stress, Nr4ADN mice do not exhibit elevated levels of key ER stress markers (p-IRE1, ATF4, and ATF6) following learning (fig. S7), and we see that only a subset of genes linked to the unfolded protein response is regulated by Nr4a factors after learning.
Fig. 2. A subset of the genes downstream of Nr4a is induced by learning.
(A) Quadrant plot based on total RNA-seq data. Genes induced by learning were identified on the basis of comparison of genes regulated in dorsal hippocampus of control male mice 2 hours after SOR (tTA+ Nr4ADN−, n = 2; tTA− Nr4ADN−, n = 2) and homecage control mice (tTA+ Nr4ADN−, n = 2; tTA− Nr4ADN−, n = 2). Quadrant plot comparing genes regulated by learning in control mice to genes regulated in Nr4ADN mice by learning. Genes up-regulated by SOR are down-regulated in Nr4ADN mice (labeled points). Size, opacity, and color intensity of each point reflect the minimum false discovery rate value for a gene between each experiment. (B and C) Expression of (B) Hspa5 and (C) Pdia6 mRNAs in C57BL/6J male mice trained in SOR and euthanized at the indicated times after training (1 hour: n = 9; 2 hours: n = 9; 3 hours: n = 9; 4 hours: n = 9), expressed as fold difference from that of mice handled only in the homecage (HC) (baseline controls, n = 10). One-way ANOVA: Hspa5: F4,41 = 13.00, P < 0.0001. Sidak’s multiple comparisons tests: ***P < 0.0001 (HC versus 1 hour), ***P < 0.0001 (HC versus 2 hours), ***P < 0.0001 (HC versus 3 hours), and *P = 0.0158 (HC versus 4 hours); Pdia6: F4,41 = 8.442, P < 0.0001. Sidak’s multiple comparisons tests: ***P = 0.0008 (HC versus 1 hour), ***P = 0.0001 (HC versus 2 hours), ***P < 0.0001 (HC versus 3 hours), and **P = 0.0054 (HC versus 4 hours). (D and E) Down-regulation of gene expression at 2 hours after SOR training in male Nr4ADN (n = 9) and control (n = 10) mice, as validated by qPCR. Unpaired t test: t17 = 3.305, **P = 0.0042 (Hspa5); t17 = 3.630, **P = 0.0021 (Pdia6). (F and G) Quantification of Western blot of lysates of synaptosomes isolated from the dorsal hippocampus of male Nr4ADN (n = 4) and control mice 2 hours after SOR training (n = 4). Unpaired t test: t6 = 4.011, **P = 0.0070 (Hspa5); t6 = 2.982, *P = 0.0246 (Pdia6).
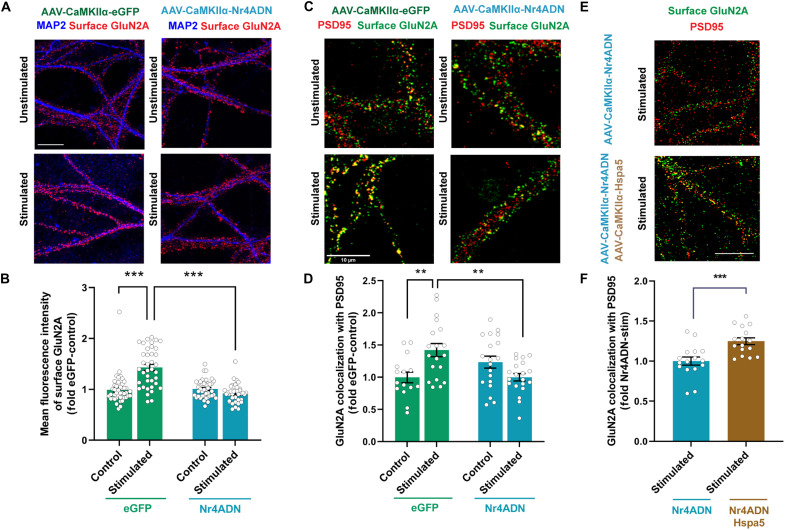
Chaperones, such as Hspa5 and Pdia6, are found in synaptosomes, where they facilitate the folding and assembly of nascent polypeptides, as well as the trafficking of proteins to the neuronal surface (20, 21). The synaptic abundance of Hspa5 and Pdia6 was significantly lower after SOR training in Nr4ADN mice compared to controls (Fig. 2, F and G, and fig. S8). Previous studies have demonstrated that Hspa5 plays a critical role in regulating the postsynaptic membrane delivery of the N-methyl-d-aspartate (NMDA) receptor subunit GluN2A in response to neuronal stimulation (20). Therefore, we next investigated whether Nr4a factors might regulate the surface expression of GluN2A. We expressed Nr4ADN [or enhanced green fluorescent protein (eGFP) as a control] in primary hippocampal neurons using a viral-based approach and assessed the distribution of GluN2A receptors following KCl-mediated neuronal depolarization. We found significant increases in surface levels of GluN2A in eGFP-transduced cells after neuronal depolarization (Fig. 3, A and B), whereas Nr4ADN-transduced cells failed to exhibit activity-induced trafficking of GluN2A (Fig. 3, A and B). We also found that Nr4ADN-expressing neurons show a significant decrease in activity-induced post-synaptic surface localization of GluN2A, as evidenced by the reduced colocalization of GluN2A with the postsynaptic density protein PSD95 (Fig. 3, C and D). Next, we investigated whether rescue of Hspa5 expression would be sufficient to increase post-synaptic localization of GluN2A in Nr4ADN-expressing neurons. Overexpression of Hspa5 increased PSD95 colocalization with GluN2A in Nr4ADN-expressing neurons (Fig. 3, E and F). Our findings demonstrate that the regulation of chaperone protein gene expression, such as Hspa5, by Nr4a transcription factors is critical for the folding and synaptic trafficking of receptor proteins that are key to synaptic plasticity.
Fig. 3. Hspa5 regulates activity-dependent surface trafficking of NMDA receptor GluN2A.
(A and B) Cultured neurons were transduced with eGFP or Nr4ADN on DIV 16 or 17 and stimulated with KCl before live staining. (A) Costaining for dendrites (MAP2 antibody) and surface GluN2A by immunofluorescence (IF) in transduced cells after KCl stimulation. Scale bar, 10 μm. (B) Quantification of surface staining for GluN2A in (A) as mean fluorescence intensity. Mixed-effect analysis: significant adeno-associated virus (AAV) construct × treatment interaction: F1,157 = 38.25, P < 0.0001. Sidak’s multiple comparisons test: ***P < 0.0001 (control eGFP versus stimulated eGFP) and ***P < 0.0001 (stimulated eGFP versus stimulated Nr4ADN). (C) Costaining for surface GluN2A and PSD95 by IF in transduced cells following KCl stimulation. Scale bar, 10 μm. (D) Quantification of surface GluN2A and PSD95 colocalization in (C). Mixed-effect analysis: significant AAV construct × treatment interaction: F1,69 = 14.57, P = 0.0003. Sidak’s multiple comparison tests: **P = 0.0026 (control eGFP versus stimulated eGFP) and **P = 0.0012 (stimulated eGFP versus stimulated Nr4ADN). (E) Cultured neurons were transduced with Nr4ADN or Nr4ADN + Hspa5 on DIV 16 or 17 and stimulated with KCl before live staining. Costaining for surface GluN2A and PSD95 by IF in transduced cells following KCl stimulation. Scale bar, 10 μm. (F) Quantification of surface GluN2A and PSD95 colocalization in (E). Unpaired t test: t30 = 3.727, ***P = 0.0008.
Restoring protein chaperone function prevents long-term memory and synaptic plasticity deficits in Nr4ADN mice
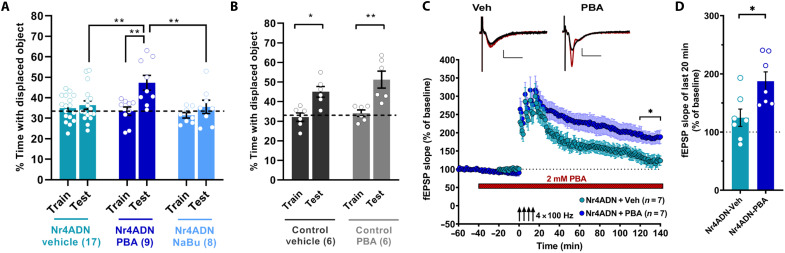
Chaperones fold nascent proteins into functional three-dimensional conformations (22), and their up-regulation after learning (Fig. 2) is essential for the activity-dependent processing and trafficking of key synaptic proteins (Fig. 3). Given the roles of chaperones in protein folding, we next performed experiments to determine whether the deficits in memory and synaptic plasticity observed in Nr4ADN mice are related to impairment of this process. First, we examined the effect of phenylbutyrate (PBA), a hydrophobic chemical chaperone, in Nr4ADN mice. PBA interacts with the exposed hydrophobic regions of nascent proteins to facilitate folding (23), partially reverses the mislocalization of proteins (24), and facilitates delivery of proteins that are critical for neuronal plasticity to the cell surface (25–27). PBA has shown promise in rescuing cognitive impairment in several mouse models of neurodegenerative diseases (28–31), and these neuroprotective effects have been attributed to its chaperone activity (28, 32). Notably, systemic delivery of a single dose of PBA to Nr4ADN mice immediately following SOR training rescued long-term memory deficits (Fig. 4A); the same treatment of control mice did not augment long-term memory (Fig. 4B). Because PBA functions as both a molecular chaperone and an inhibitor of histone deacetylases (HDACs), we confirmed that this rescue was not due to changes in expression of Hspa5 and Pdia6 in Nr4ADN mice (fig. S9). In addition, sodium butyrate (NaBu), an HDAC inhibitor that has no molecular chaperone activity (28), failed to rescue memory in Nr4ADN mice (Fig. 4A). This finding is consistent with our previous observation that broad HDAC inhibition is not sufficient to reverse the memory deficits in Nr4ADN mice (4).
Fig. 4. Chemical chaperone PBA reverses long-term memory and synaptic plasticity deficits in Nr4ADN mice.
(A) Nr4ADN mice were injected intraperitoneally with PBA (200 mg/kg, n = 9), sodium butyrate [NaBu; 200 mg/kg, n = 8 (2F)], or vehicle [n = 17 (2F)] immediately after SOR training and tested for long-term memory 24 hours later. Two-way ANOVA: significant treatment × session interaction (F2,31 = 4.207, P = 0.0242). Sidak’s multiple comparisons tests: **P = 0.0011 (Nr4ADN mice-PBA, train versus 24-hour test), **P = 0.0036 (Nr4ADN-PBA 24-hour test versus Nr4ADN-vehicle 24-hour test), and **P = 0.0084 (Nr4ADN-PBA 24-hour test versus Nr4ADN-NaBu 24-hour test). (B) Male control mice were injected intraperitoneally with PBA (200 mg/kg, n = 6) or vehicle (n = 6) immediately after completion of SOR training and tested for long-term memory 24 hours later. Two-way ANOVA: significant main effect of sessions: F1,10 = 33.46, P = 0.0002. Sidak’s multiple comparisons tests: *P = 0.0110 (control mice-vehicle, train versus test) and **P = 0.0018 (control mice-PBA, train versus test). (C and D) Effects of PBA on persistence of LTP. Expression of Nr4ADN attenuates persistence of LTP in hippocampal slices (Nr4ADN-veh), while bath treatment with 2 mM PBA rescues these LTP deficits (Nr4ADN-PBA) (two-way repeated-measures ANOVA, effect of PBA treatment: F1,12 = 8.125, P = 0.0146). The mean fEPSP slope over the last 20 min of the recordings was enhanced in PBA-treated slices compared to vehicle-treated slices (PBA-treated: 187.3 ± 16.2%, n = 7 slices, four mice; vehicle-treated: 124.6 ± 14.9%, n = 7 slices, five mice; unpaired t test, *P = 0.0146). Treatment with 2 mM PBA had no significant effect on the baseline responses (pre-drug baseline, 20 min: 100.1 ± 0.13%; post-drug pre-induction baseline, 20 min: 90.63 ± 6.6%; paired t test, P = 0.2094). The representative fEPSP traces shown are sampled at baseline (black) and at the end of the recording (red). Scale bar (2 mV), 10 ms. Error bars indicate SEM.
We previously showed that Nr4ADN mice exhibit deficits in a form of persistent, protein synthesis–dependent long-term potentiation (LTP) induced by repeated spaced high-frequency stimulation of the hippocampal CA1-Schaffer collateral synapses (12). Given that PBA treatment reversed long-term memory deficits in Nr4ADN mice, we examined its effects on this long-lasting form of LTP. Following 20 min of stable baseline field-excitatory postsynaptic potentials (fEPSPs) recordings, hippocampal slices from Nr4ADN mice were treated by bath application of 2 mM PBA [dissolved in the artificial cerebrospinal fluid (aCSF)]. After 40 min of PBA treatment, long-lasting LTP was induced using a spaced four-train stimulation protocol (four 100-Hz, 1-s trains separated by 5 min). Potentiation in slices from Nr4ADN mice decayed quickly, thus showing deficits in the persistence of LTP, as reported in our previous study (12). Treatment with PBA rescued the deficits in long-lasting LTP in Nr4ADN slices leading to persistently enhanced potentiation compared to the vehicle group (Fig. 4, C and D). At the concentration used, PBA did not have significant effects on the pre-induction baseline (Fig. 4C). These findings demonstrate that PBA treatment reverses the deficits in long-lasting synaptic plasticity and memory in Nr4ADN mice by promoting the folding of newly synthesized proteins.
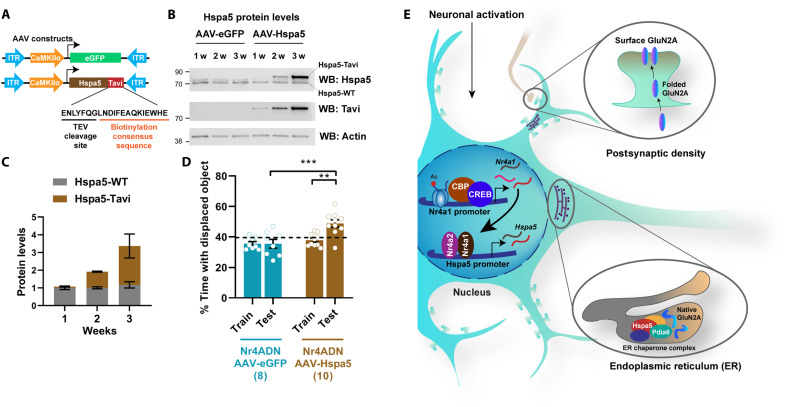
To define the specific role of the molecular chaperone Hspa5 in the memory deficits observed in Nr4ADN mice, we reinstated Hspa5 expression selectively in hippocampal excitatory neurons using a viral approach and performed behavioral studies 2 weeks after viral infusion (Fig. 5, A to C). The level of overexpression achieved was sufficient to reverse the long-term spatial memory deficits observed in Nr4ADN mice (Fig. 5D), supporting the idea that Hspa5 is downstream of Nr4a transcription factors (Fig. 5E). These findings suggest that the deficits in synaptic plasticity and long-term memory in Nr4ADN mice are due to disruption of a chaperone activity required for the folding of newly synthesized proteins. Overall, we conclude that the activity-induced regulation of ER chaperone Hspa5 by Nr4a transcription factors is essential for the native protein folding required for consolidation of long-term memory.
Fig. 5. The Nr4a proteins contribute to memory through downstream chaperone proteins.
(A) Schematic of viral constructs used to drive expression of Hspa5 in excitatory neurons of the dorsal hippocampus of male C57BL/6J mice. AAV9-CaMKIIα-eGFP served as vector control, and AAV9-CaMKIIα-Hspa5-Tavi was used to drive expression of Hspa5 in excitatory neurons. The Tavi-tag can be identified by an antibody against a consensus biotinylation sequence. (B) Western blot of synaptosomes, showing mild Hspa5-Tavi expression within 1 week of infusion, and expression approximately equal to that of endogenous Hspa5 within 2 weeks of infusion. (C) Quantitation of data in (B). (D) Long-term memory (24-hour) assessment of Nr4ADN mice infused with AAV-eGFP or AAV-Hspa5-Tavi into dorsal hippocampus. Two-way ANOVA: significant AAV type × session interaction: F1,16 = 6.985, P = 0.0177. Sidak’s multiple comparisons tests: **P = 0.0022 (AAV-Hspa5, train versus test), ***P = 0.0002 (AAV-Hspa5, 24-hour test versus AAV-eGFP, 24-hour test), while eGFP-infused Nr4ADN mice showed no preference toward the DO. AAV-Hspa5: n = 10 (4F) and AAV-eGFP: n = 8 (3F). (E) Schematic illustration of model wherein learning-induced expression of Nr4a1 drives Hspa5 expression to initiate protein folding in the ER that enables the expression of functional proteins at the synapse.
Activation of Nr4a1 or ER chaperone function ameliorates memory impairments in an ADRD mouse model
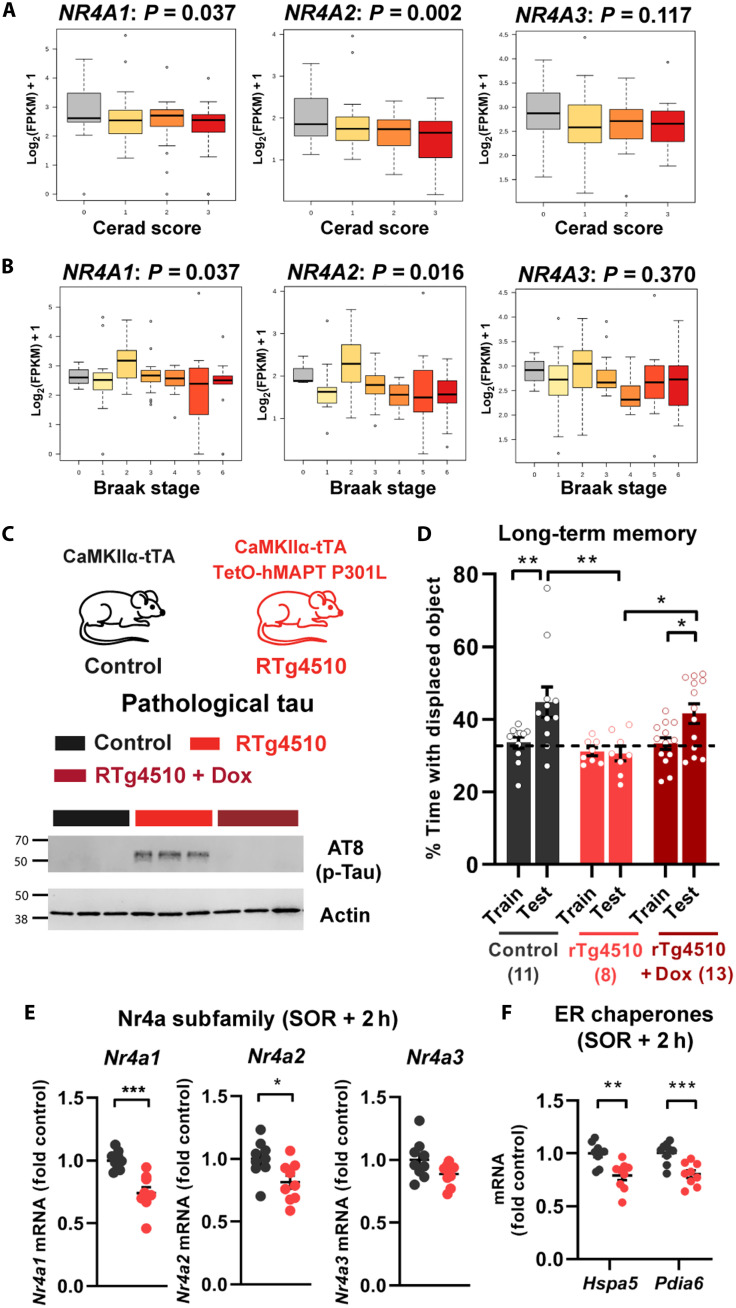
Nr4a transcription factors were previously implicated in Aβ aggregation and memory deficits (33). Therefore, we investigated the expression of Nr4a transcripts in the hippocampus of ADRD patients with increasing grades of pathology. Using a database of RNA-seq and pathological findings from postmortem brain tissue from patients and healthy controls in The Allen Brain Institute Study of Aging, Dementia, and Traumatic Brain Injury (34), we examined the relationship between hippocampal expression of NR4A subfamily members across increasing Cerad (Consortium to Establish a Registry for Alzheimer’s Disease) score, a neuropsychological assessment of the progression of AD, and Braak stage, a measure of the distribution and pathological burden of neurofibrillary tangles (NFTs). We found that levels of expression of NR4A1 and NR4A2 were negatively correlated with both the Cerad scores (Fig. 6A) and Braak stages (Fig. 6B), both measures of the severity of ADRD pathology, whereas NR4A3 was not significantly correlated with disease pathologies. Consistent with our findings, NR4A2 protein expression is reduced in postmortem AD hippocampus at Braak stage VI (33).
Fig. 6. Down-regulation of Nr4a factors and their effector genes is linked to ADRD pathology.
(A and B) Expression profiles of NR4A1, NR4A2, and NR4A3 in the hippocampus, from the Allen Brain Institute Study of Aging, Dementia, and Traumatic Brain Injury, correlated with (A) Cerad scores, which reflect the density of neuritic plaques, and (B) Braak stages, which reflect the severity of neurofibrillary tangles. (C) Schematic depiction of control (CaMKIIα-tTA) or rTg4510 (CaMKIIα-tTA and TetO-hMAPT P301L) mice and Western blots of tau phosphorylation (AT8, phosphorylation at both Ser202 and Thr205) in the dorsal hippocampus in the presence or absence of doxycycline (Dox). (D) Long-term memory in rTg4510 and control mice at 4 months of age, following training in SOR. Two-way ANOVA: significant main effect of genotype/treatment (F2,29 = 4.792, P = 0.0159) and significant main effect of sessions (F1,29 = 9.221, P = 0.0050). Sidak’s multiple comparisons test: **P = 0.0096 (control mice, train versus test), *P = 0.0439 (rTg4510-Dox mice, train versus test), **P = 0.0016 (rTg4510 mice, 24-hour test versus control mice, 24-hour test), and *P = 0.0140 (rTg4510 mice, 24-hour test versus rTg4510 Dox, 24-hour test). Control, n = 11 (3F); rTg4510, n = 8 (4F); and rTg4510 + Dox, n = 13 (7F). (E and F) rTg4510 and control mice (n = 9 per group) were trained in SOR, and 2 hours later, the dorsal hippocampus was collected and processed for RNA extraction and the analysis of gene expression. (E) Nr4a subfamily gene expression: unpaired t test: Nr4a1: t16 = 4.878, ***P = 0.0002; Nr4a2: t16 = 2.621, *P = 0.0185. (F) Hspa5 and Pdia6 gene expression: unpaired t test: Hspa5: t16 = 3.692, **P = 0.0020; Pdia6: t16 = 4.177, ***P = 0.0007.
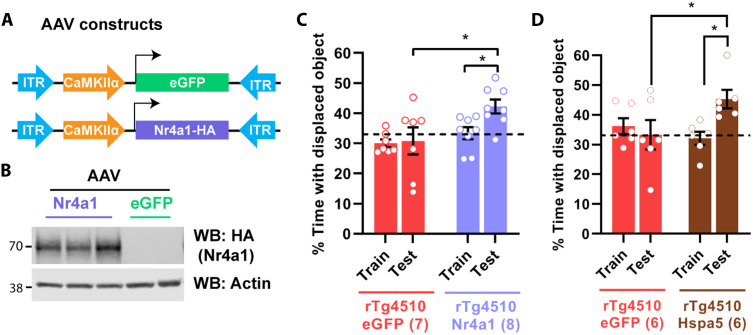
Cognitive impairment is a significant feature of ADRD, and we hypothesized that NR4A down-regulation might be related to the compromise in cognitive abilities. To examine this in a mouse model of ADRD, we used the rTg4510 mouse, which overexpresses mutant human tau (tau P301L) exclusively in excitatory neurons (35, 36). These mice develop tangle-like inclusions (35, 36) and show pathological hyperphosphorylation of tau proteins (AT8) in the dorsal hippocampus starting at 3 to 4 months of age (Fig. 6C). These mice have deficits in spatial learning in the Morris water maze, contextual fear conditioning (35–38), and long-term spatial memory in the SOR task (Fig. 6D). Doxycycline treatment prevents these memory deficits (Fig. 6D), demonstrating that it is the expression of the mutant tau transgene and not the transgene insertion site (39) that drives the tauopathy-like phenotype. As in the case of the human postmortem data, the expression of both Nr4a1 and Nr4a2 was down-regulated in the dorsal hippocampus of rTg4510 mice after SOR training (Fig. 6E). These findings validate the appropriateness of using this mouse line as a model for Nr4a dysregulation. Furthermore, we found that Hspa5 and Pdia6 were down-regulated in the dorsal hippocampus of rTg4510 mice after SOR training (Fig. 6F). To determine the extent to which Nr4a transcription factors contribute to the memory impairment seen in rTg4510 mice, we overexpressed Nr4a1 in the dorsal hippocampus of adult mice (Fig. 7, A and B). This reversed the deficits in long-term spatial memory normally observed in rTg4510 mice (Fig. 7C). Last, to determine the role of Hspa5 chaperone in ADRD-associated memory impairment, we overexpressed Hspa5 in the dorsal hippocampus of rTg4510 mice. Strikingly, Hspa5 overexpression ameliorated long-term memory deficits in rTg4510 mice (Fig. 7D). These findings link the function of the Nr4a family of transcription factors to the cognitive deficits associated with neurodegenerative disorders, and they suggest that targeting the Nr4a family and their downstream effector genes would be beneficial in the treatment of memory deficits associated with ADRD.
Fig. 7. Restoration of Nr4a1 or ER chaperone function reverses memory deficits in a mouse model of ADRD.
(A) Schematic depiction of AAV constructs used to drive expression of Nr4a1 in excitatory neurons in the dorsal hippocampus of 3-month-old rTg4510 mice. (B) Western blot showing expression of virally transduced Nr4a1-HA in dorsal hippocampus 4 weeks following infusion. (C) rTg4510 mice 3 months of age were infused with AAV9-CaMKIIα-Nr4a1-HA or control vector (AAV9-CaMKIIα-eGFP), and 4 weeks later, they were trained in SOR. Long-term memory was tested 24 hours after the training session. Two-way ANOVA: significant main effect of AAV-type infusion (F1,13 = 6.597, P = 0.0234). Sidak’s multiple comparison tests: *P = 0.0127 (AAV-Nr4a1, 24-hour test versus AAV-eGFP, 24-hour test) and *P = 0.0489 (AAV-Nr4a1 train versus AAV-Nr4a1 24-hour test). AAV-Nr4A1: n = 8; AAV-eGFP: n = 7. (D) rTg4510 mice 3.5 months of age were infused with AAV9CaMKIIα-Hspa5-Tavi or control vector (AAV9-CaMKIIα-eGFP), and 2 weeks later, they were trained in SOR. Long-term memory was tested 24 hours after the training session. Two-way ANOVA: significant session × AAV interaction, F1,10 = 8.767, P = 0.0143. Sidak’s multiple comparison tests: *P = 0.0419 (AAV-eGFP test versus AAV-Hspa5 test) and *P = 0.0419 (AAV-Hspa5 train versus AAV-Hspa5 test). AAV-eGFP: n = 6 (3F); AAV-Hspa5: n = 6 (3F).
DISCUSSION
Here, we show that Nr4a transcription factors act during memory consolidation to drive the expression of genes encoding chaperones that are part of a multiprotein complex within the ER, thereby facilitating folding of the proteins into their functional conformations (18). This study provides functional evidence that these chaperones are involved in synaptic plasticity and long-term memory. The results demonstrating that long-term memory can be reversed in Nr4ADN mice by either application of the chemical chaperone PBA or overexpression of Hspa5 reveal that the protein folding machinery plays a critical role in memory consolidation. Our work in hippocampal neurons identifies the synaptic membrane protein GluN2A as a candidate target protein whose surface trafficking is regulated by Nr4a1-driven expression of Hspa5. Nr4a1 was previously shown to be involved in regulating dendritic spine density (40), consistent with the hypothesis that the target genes of this transcription factor affect synaptic structure and function. Our identification of ER chaperones as effector genes during memory consolidation provides a novel link between the induction of gene expression and protein synthesis, which are hallmarks of memory consolidation and the synaptic plasticity that leads to modification of neural circuits and behavioral alterations.
Our study extends these fundamental findings on the molecular mechanisms of memory, advancing our understanding of memory loss associated with neurodegenerative disorders by identifying changes in expression of the Nr4a family of transcription factors in both human AD brains and a mouse model of ADRD. Tau transgenic models and human tauopathy data exhibit widespread loss of heterochromatin (41), impaired chromatin remodeling, and nuclear lamina formation (42). Levels of the lysine acetyltransferase CBP are reduced in THY-Tau22 mutant mice (3) and in human AD patient samples (43). CBP and histone acetylation regulate the expression of Nr4a family genes, and consistent with our findings that overexpression of Nr4a1 reverses memory deficits in tau mutant mice, both the pharmacological activation of CBP (3, 44, 45) and the inhibition of HDAC activity restore memory in several mouse models of ADRD (46–48). Although the exact mechanisms underlying the transcriptional alterations in neurodegenerative disorders remain to be identified, they represent attractive targets for the development of drugs to ameliorate cognitive deficits, which are a debilitating aspect of ADRDs. Our finding that overexpression of Nr4a1 or Hspa5 chaperone reverses memory loss in a tau-based model of ADRD supports this as an innovative therapeutic approach.
The work described here identifies ER chaperone proteins as critical molecular regulators of memory storage. ER chaperones have been studied mainly for their roles in ER stress and the unfolded protein response; their role in memory consolidation is underexplored. Our work here links a subset of these ER chaperones, including Hspa5 and protein disulfide isomerases, to protein folding and trafficking within critical time windows during memory consolidation, laying the groundwork for future experiments to identify additional downstream targets of these ER chaperones, with promises of a more complete understanding of the fundamental molecular mechanisms of memory consolidation that go awry in neurodegenerative disorders.
MATERIALS AND METHODS
Data reporting
No statistical methods were used to predetermine sample size.
Mouse lines
Nr4ADN mice
Adult Nr4ADN mice were 2 to 4 months old at the time of behavioral or biochemical experiments. They were maintained on a C57BL/6J background and harbored both the CaMKIIα-tTA and Tet-O-Nr4ADN transgenes (4). Incorporation of the CaMKIIα-tTA transgene into chromosome 12 causes a 508.12-kb deletion that affects five genes: Vipr2, Wdr60, D430020J02Rik, Ncapg2, and Ptprn2. To account for any effects of the deleted genes on memory or gene expression (49), age-matched CaMKIIα-tTA–expressing littermates were used as controls throughout the study.
rTg4510 mice
These mice were 3 to 4 months old at the time of behavioral or biochemical experiments. They were maintained on a C57BL/6J background and harbored two transgenes: CaMKIIα-driven tTA and TetO-driven human tau P301L. Age-matched CaMKIIα-driven tTA-expressing littermates were used as controls.
C57BL/6J mice
Adult male mice purchased from The Jackson Laboratory were 2 to 4 months of age during behavioral or biochemical experiments. All mice had free access to food and water; lights were maintained on a 12-hour light/12-hour dark cycle.
To suppress TetO-driven expression of Nr4ADN or human tauP301L transgenes, Nr4ADN or rTg4510 mice were placed on a diet containing doxycycline (200 mg/kg; Bio-Serv) from weaning until behavioral experiments. All behavioral testing was performed during the light cycle between Zeitgeber time (ZT) 0 to 2. For all behavioral and biochemical experiments, mice were randomly assigned to groups, housed individually for 7 days before experiments, and handled for 2 min per day for 5 days. All experiments were conducted according to U.S. National Institutes of Health (NIH) guidelines for animal care and use and were approved by the Institutional Animal Care and Use Committee of the University of Iowa, Iowa.
Drugs
Sodium PBA (Sigma-Aldrich) and NaBu (Sigma-Aldrich) were dissolved in saline. For electrophysiology experiments, the dose of PBA was chosen on the basis of the range of median inhibitory concentration (IC50) values reported in the published literature (50). For behavioral experiments, mice were injected with PBA or NaBu intraperitoneally at a dose of 200 mg/kg, immediately after SOR training. Control mice were injected with vehicle (0.9% sterile saline).
Adeno-associated virus constructs
AAV2.9-CaMKIIα-eGFP, AAV2.9-CaMKIIα-Nr4A1-HA, AAV2.9-CaMKIIα-Hspa5-Tavi, AAV2.2-CaMKIIα-Nr4ADN, and AAV2.2-CaMKIIα-eGFP were purchased from VectorBuilder (VectorBuilder Inc.).
Stereotactic surgeries
Mice were anesthetized using isoflurane and kept on a warm heated pad throughout the stereotactic surgery procedure. Meloxicam was injected as analgesics (51). Viral infusion was performed using a 33-gauge beveled needle [World Precision Instruments (WPI)] attached to a 10-μl Nanofil syringe controlled by a microsyringe pump (UMP3; WPI). The coordinates for dorsal hippocampus were as follows: anteroposterior, −1.9 mm; mediolateral, ±1.5 mm; and 1.5 mm below bregma. The needle was lowered to the site of injection over the course of 5 min and remained at the target for 1 min before injection was initiated (0.2 μl per min). Each hippocampus was injected with approximately 1 μl of the relevant constructs. After injection was completed, the needle remained at the site for one additional minute and then slowly removed over a 5-min period. A small amount of bone wax (Lukens) was then used to close the drill holes, and the incision was closed with sutures.
SOR task
All animals were housed individually for 1 week before behavioral experiments were initiated. Age-matched littermates were used. Mice were handled for 2 min per day for five consecutive days before the behavioral task. All spatial memory tasks were conducted between ZT0 and ZT2. Briefly, mice were habituated in the open-field arena for 6 min during the habituation session, followed by three 6-min training sessions in the same open field containing three different glass objects. The intertrial interval was 3 min, during which the mice were returned to their homecage, and the objects and arena were cleaned with 70% ethanol. An internal spatial cue (vertical black lines printed on a white paper 18 cm by 12 cm in size) was attached to one wall of the open field to allow the mice to locate each object relative to the spatial cue during free exploration of the arena. After either 1 or 24 hours following the training sessions, mice were brought back to the open field in which the location of one of the objects was displaced to another spatial location. Time spent exploring the displaced object (DO) and the nondisplaced objects (NDO) during the 6-min test session was recorded. The exploration was hand-scored by an experimenter blinded to the genotype or treatment. Animals were assigned to the arenas randomly, without use of any randomization software. An object was scored as “explored” if it was sniffed or touched, or the face was in close proximity (within 1 cm) to the object, as described previously (52).
Electrophysiology
Nr4ADN male mice 2 to 3 months of age were used. Mice were euthanized by cervical dislocation, and the brain was quickly dissected into cold artificial cerebrospinal fluid (aCSF), which was continuously bubbled with carbogen (95% O2, 5% CO2). The isolation of hippocampi and preparation of acute hippocampal slices were performed as described (53). Transverse acute hippocampal slices of 400-μm thickness were prepared from both hippocampi using a manual McIlwain slicer (Stoelting). The slices were quickly transferred onto a net insert in an interface recording chamber (Fine Science Tools, Foster City, CA) and left to equilibrate to a humidified carbogen atmosphere at 28°C for at least 2 to 3 hours before recordings were initiated. The slices were perfused at 1 ml/min with oxygenated aCSF throughout the experiments. The aCSF used for both the dissection and recordings was composed of 124 mM NaCl, 4.4 mM KCl, 1 mM NaH2PO4, 2.5 mM CaCl2.2H2O, 1.3 mM MgSO4.7H2O, 26.2 mM NaHCO3, and 10 mM d-glucose (pH ~7.4) when equilibrated with carbogen. fEPSPs were recorded in the CA1 stratum radiatum by stimulating Schaffer collaterals with a monopolar, lacquer-coated stainless steel electrode (~5-megohm resistance, A-M Systems, #571000) and recording with an aCSF-filled glass microelectrode (2- to 5-megohm resistance). In all experiments, test stimulation was a biphasic, constant-current pulse (100-μs duration) delivered every minute at a stimulation intensity that evoked ~40% of the maximal fEPSP amplitude, as determined by an input-output curve (stimulation intensity versus fEPSP amplitude). Also, a stable baseline was recorded for at least 20 min before LTP was induced or drug was applied. LTP was induced by a spaced four-train stimulation protocol consisting of four 100-Hz, 1-s trains delivered at 5-min intervals at the test stimulus intensity. For each experiment, PBA solution (2 mM) was prepared fresh by dissolving in aCSF, and it was applied to the bath and protected from light. The solution was recirculated after 30 min of initial application. In the electrophysiological data presented, n represents the number of slices. Data were acquired using Clampex 10 and Axon Digidata 1440 digitizer (Molecular Devices, Union City, CA) at 20 kHz and were low-pass–filtered at 2 kHz with a four-pole Bessel filter. Data analysis was performed using the Clampfit 10 software (Molecular Devices, Union City, CA). Data were plotted, and statistical analyses were performed using the GraphPad Prism 8 software. For each slice, the fEPSP slopes were normalized against the average slope over the 20-min baseline (pre-drug baseline in the PBA-treated slices). Data are presented as means ± SEM. LTP persistence was assessed by comparing the final 20-min recordings for the vehicle and treatment groups using a two-way repeated-measures analysis of variance (ANOVA). The mean fEPSP slope of the final 20-min recordings for each group was compared using two-tailed, unpaired t test. Statistical significance was set at P < 0.05.
Isolation of whole-cell extracts and synaptosomal fractions
For whole-cell lysate preparation, flash-frozen dorsal hippocampal tissue was homogenized mechanically in 300 μl of ice-cold radioimmunoprecipitation assay (RIPA) buffer (Sigma-Aldrich) supplemented with 0.2% Triton X-100 (Sigma-Aldrich) and Protease and Phosphatase Inhibitor Cocktail (1:100; Thermo Fisher Scientific). The lysate was kept on ice for 30 min, following which they were centrifuged at 10,000g for 15 min at 4°C. The pellet was discarded, and the supernatant (whole-cell lysate) was collected for Western blot analysis. For synaptosomal extraction, hippocampal tissue was mechanically homogenized in Syn-PER Reagent (Thermo Fisher Scientific) containing Halt Protease and Phosphatase Inhibitor Cocktail (1:100; Thermo Fisher Scientific). The homogenate was centrifuged at 1200g for 10 min at 4°C, after which the pellet (nuclear fraction) was discarded and the supernatant was centrifuged again at 15,000g for 25 min at 4°C. This pellet (synaptosomal fraction) was resuspended in RIPA buffer (Sigma-Aldrich) containing Halt Protease and Phosphatase Inhibitor Cocktail (1:100) and 0.2% Triton X-100 (Sigma-Aldrich).
Western blot analysis
Protein extracts were transferred to polyvinylidene difluoride membranes as previously described (3). Membranes were blocked with Odyssey Blocking Buffer in tris-buffered saline (LI-COR) and incubated overnight at 4°C with the following primary antibodies: Hspa5 (1:2000; Proteintech, 11587-1-AP), Pdia6 (1:2000; Abcam, ab11432), Biotin Ligase Epitope Tag (for Tavi-tag detection, 1:1000; Abcam, ab106159), PSD95 (1:5000; Thermo Fisher Scientific, 6G6-1C9), hemagglutinin (HA) (1:2000; Millipore Sigma), phospho-tau AT8 (1:5000; BioLegend, 806503), actin (1:10,000; Thermo Fisher Scientific), ATF4 (1:500; Thermo Fisher Scientific, PA5-27576), ATF6 (1:500; Thermo Fisher Scientific, PA5-114886), and phospho-IRE1α (1:500; Thermo Fisher Scientific, PA5-85738). Membranes were washed and incubated with appropriate IRDye immunoglobulin G (IgG) secondary antibodies, including anti-rabbit IRDye 800LT (1:5000; LI-COR) and anti-mouse IRDye 680CW (LI-COR). Images were acquired using the Odyssey Infrared Imaging System (LI-COR). Quantification of Western blot bands was performed using Image Studio Lite v.5.2 (LI-COR).
RNA extraction, cDNA synthesis, and quantitative real-time reverse transcription polymerase chain reaction
Dorsal hippocampi were dissected and immediately stored at −80°C in RNAlater solution (Ambion) for later isolation of total RNA. For RNA extraction, Qiazol (Qiagen) was added to the hippocampal tissues and they were homogenized using stainless steel beads (Qiagen). Chloroform was then added to the homogenates, and the samples were centrifuged at 12,000g at room temperature for 15 min. RNA was precipitated from the aqueous phase using ethanol and then cleaned using an RNeasy kit (Qiagen). RNA was eluted in nuclease-free water and treated with DNase (Qiagen) at room temperature for 25 min. The cleaned RNA was precipitated in ethanol, sodium acetate (pH 5.2), and glycogen overnight at −20°C. RNA samples were centrifuged at top speed at room temperature for 20 min. The precipitates were further washed with 70% ethanol and centrifuged at top speed for 5 min. The RNA precipitates were dried and resuspended in nuclease-free water, and concentrations were estimated using a NanoDrop (Thermo Fisher Scientific). Complementary DNAs (cDNAs) were prepared from 1 μg of RNA using the SuperScript IV First-Strand Synthesis System (Ambion). Real-time reverse transcription polymerase chain reactions (RT-PCRs) were performed in a 384-well optical reaction plate with optical adhesive covers (Life Technologies). Each reaction was composed of 2.25 μl of cDNA (2 ng/μl), 2.5 μl of Fast SYBR Green Master Mix (Thermo Fisher Scientific), and 0.25 μl of primer mix (IDT). A minimum of three technical replicates per reaction was performed on the QuantStudio 7 Flex Real-Time PCR System (Applied Biosystems, Life Technologies). Data were normalized to housekeeping genes (Tubulin, Pgk1, and B2m), and 2(−ΔΔCt) method was used for gene expression analysis.
RNA library preparation, sequencing, and analysis
RNA libraries were prepared at the Iowa Institute of Human Genetics (IIHG), Genomics Division, using the Illumina TruSeq Stranded Total RNA with Ribo-Zero gold sample preparation kit (Illumina Inc., San Diego, CA). The KAPA Illumina Library Quantification Kit (KAPA Biosystems, Wilmington, MA) was used to measure library concentrations. Pooled libraries were sequenced on Illumina HiSeq4000 sequencers with 150-bp paired-end chemistry (Illumina) at the IIHG core. RNA-seq data were processed with the bcbio-nextgen pipeline (https://github.com/bcbio/bcbio-nextgen, version 1.1.4). The pipeline uses STAR (54) to align reads to the mm10 genome build (GENCODE release M10, Ensembl 89 annotation) and quantifies expression at the gene level with featureCounts (55). All further analyses were performed using R (56). For gene-level count data, the R package EDASeq (57) was used to adjust for GC content effects (full quantile normalization) and account for sequencing depth (upper quartile normalization) (figs. 10 and 12). Latent sources of variation in expression levels were assessed and accounted for using RUVSeq (RUVr mode using all features) (figs. 11 and 13) (58). Appropriate choice of the RUVSeq parameter k was guided through inspection of P value distributions, relative log expression (RLE) plots, and principal components analysis (PCA) plots. Specifically, the smallest value k was chosen where the P value distribution showed an expected peak below 0.05, RLE plots were evenly distributed and zero-centered, and PCA plots demonstrated replicate sample clustering in the first three principal components (59). Differential expression analysis was conducted using the edgeR quasi-likelihood pipeline (60–62). Codes to reproduce the RNA-seq analysis are available at https://github.com/ethanbahl/chatterjee2021_nr4a.
Gene Ontology and pathway enrichment analyses of DEGs
The ClueGO (63) and CluePedia plug-ins of the Cytoscape 3.7.5 software (64) were used in “Functional analysis” mode, using the default parameters for analyzing gene ontology, molecular function, and KEGG pathways in networks for DEGs. The names of significant DEG were pasted into the “Load Marker List” of ClueGO, and the organism “Mus Musculus [10090]” was selected.
Construction of PPI networks
The protein-protein interactive network was constructed using STRING (65) (version 11.0), which uses the STRING database (http://string-db.org/) (66). The PPI network was constructed to identify the interactions between proteins encoded by down-regulated DEGs based on experimental data. The DEG names were pasted into “STRING protein query.” Active interaction sources, including text mining, experiments, databases, coexpression, neighborhood, gene fusion, and co-occurrence, were applied, and highest interaction score confidence (0.900) was selected to construct the PPI networks. Full network was constructed, where the edges indicate both functional and physical protein associations.
ChIP-seq analysis
Using the SRA toolkit fastq-dump, raw ChIP-seq data from Liu et al. (19) [Gene Expression Omnibus (GEO) accession code GSE96969] were downloaded, including Nr4a1-HA (SRR6788331), IgG-control (SRR6788333), and input DNA (SRR6788332). The raw reads were trimmed using Trimmomatic version 0.36 with the parameters ILLUMINACLIP:2:30:15 LEADING:30 TRAILING:30 MINLEN:23, and quality inspection was conducted using FastQC version 0.11.5. The trimmed reads from all datasets were aligned using BWA version 0.7.15 with the algorithm mem and default parameters. Signal density files in BedGraph format were generated using BEDTools genomecov version 2.26.0 with default parameters and then converted in uniform 10-nucleotide bin WIG files for further normalization steps. Peak calling was done using MACS2 version 2.1.2 with a q-value cutoff of 0.01 and default parameters. Each signal density file corresponding to a dataset was scaled such that the total sum of the signal over the mouse mm10 genome was equivalent to 1 million reads of 100 nucleotides. The signal of the input dataset was then subtracted from its corresponding immunoprecipitation dataset to generate the “sclWT-ctrl” files. The normalized WIG files were then encoded in bigWig format using the Kent utilities and visualized using the Integrative Genomics Viewer with the mouse mm10 reference genome.
Primary neuronal cultures and adeno-associated virus transduction
Hippocampi from mice (C57BL/6J) were used to generate primary neuronal cultures as previously described (67). Briefly, hippocampi from postnatal P0/P1 pups were dissected, trypsinized (0.25%), and triturated using a fire-polished glass Pasteur pipette to prepare a single-cell suspension. Cells were then plated on poly-l-lysine (1 mg/ml)–coated four-well glass-bottom dishes (Cellvis) at optimal density (150 to 200 cells/mm2) and maintained in Neurobasal medium (Gibco) containing B27 Supplement (Gibco), in an incubator with 5% CO2 and at 37°C. At days in vitro (DIV) 16 to 17, neurons were transduced with AAV2.2-CaMKIIα-Nr4ADN, AAV2.9-CamKIIa-Hspa5-Tavi, and AAV2.2-CaMKIIα-eGFP (titer of concentrated viral stock was 1 × 1013 to 2 × 1013 genome copies/ml) in a 1:1000 dilution of Neurobasal medium containing B27 Supplement. At 8 to 10 hours following transduction, half of the existing medium was replenished with fresh Neurobasal medium containing B27 Supplement. Cultures were typically maintained until DIV 23 to 25 before experiments commenced.
KCl stimulation and surface labeling of GluN2A
At DIV 23 to 25, neurons were incubated in low KCl–Hepes-buffered saline (290 mOsm) [110 mM NaCl, 5.4 mM KCl, 1.8 mM CaCl2, 0.8 mM MgCl2, 10 mM d-glucose, and 10 mM Hepes-NaOH (pH 7.4)] for 60 min. Thereafter, neurons were stimulated for another 60 min with high KCl-HBS (same as low KCl-HBS, except for 55 mM NaCl and 60 mM KCl). The high KCl-HBS was washed off, and live neurons were then immunolabeled with N-terminal NMDAR2A antibody (1:25; Thermo Fisher Scientific) in low KCl-HBS for 30 min to exclusively stain the surface GluN2A receptors. Following the antibody incubation, the cells were washed twice with phosphate-buffered saline containing Mg2+and Ca2+ (PBS-MC; 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 2 mM KH2PO4, 1 mM MgCl2, and 0.1 mM CaCl2). Cells were then fixed in PBS-MC containing 2% paraformaldehyde and 2% sucrose for 15 min at 37°C, washed thrice in PBS-MC at room temperature, and blocked with PBS-MC containing 2% bovine serum albumin for 60 min at room temperature. Cells were incubated with Alexa Fluor 647–conjugated goat anti-rabbit secondary antibody (1:200; Invitrogen) at room temperature for 90 min in blocking solution. Neurons were then permeabilized with PBS-MC containing 0.1% Triton X-100 at room temperature for 5 min and incubated with blocking solution for 60 min and thereafter with MAP2 antibody (1:1000; Sigma-Aldrich) for 8 to 10 hours at 4°C. For PSD95 immunocytochemistry, permeabilized neurons were incubated with PSD95 antibody (1:4000; Enzo Life Sciences) for 8 to 10 hours at 4°C. Cells were washed thrice in PBS-MC and incubated with Alexa Fluor 546–conjugated goat anti-mouse secondary antibody (1:200; Invitrogen) at room temperature for 60 min. Last, cells were washed three times with PBS-MC at room temperature and preserved in PBS-MC for future imaging.
Confocal imaging and image analysis
Cultured neurons after completion of the experiments were imaged using an Olympus FV3000 confocal microscope with a 100× numerical aperture = 1.45 oil immersion objective at 1024 × 1024 pixel resolution. High-magnification images were captured using 3× optical zoom. All images (8 bit) were acquired with identical settings for laser power, detector gain, and pinhole diameter for each experiment and between experiments. Images from the different channels were stacked and projected at maximum intensity using ImageJ (NIH). Mean fluorescence intensity of surface GluN2A and the colocalization between surface GluN2A and PSD95 puncta were assessed using plugins in ImageJ.
Analysis of human AD
Using RNA sequencing data from the “Aging, Dementia, and TBI Study” (34) provided by the Allen Institute for Brain Science, we fit linear models between the RNA integrity number (RIN)–corrected and log2-transformed hippocampal expression levels of NR4A family members and the individual’s Cerad score, a semiquantitative estimate of neuritic plaque density (34).
Statistics
Behavioral and biochemical data were analyzed using paired or unpaired two-tailed t tests and either one-way or two-way ANOVAs (in some cases, with repeated measures as the within-subject variable). Sidak’s tests were used for post hoc analyses, where needed. Differences were considered statistically significant when P < 0.05. As indicated for each figure panel, all data are plotted in either bar graphs, in which symbols represent each data point, or dot plots, where each symbol represents an individual data point. Graphs were plotted as means ± SEM.
Acknowledgments
We thank the Iowa Institute of Human Genetics (IIHG) core for RNA-seq library preparation and sequencing. We thank the Neural Circuits and Behavior Core in the Iowa Neuroscience Institute for use of facilities. We thank R. Merrill and S. Strack for help with optimization of cell culture experiments and L. Lyons, T. Nickl-Jockschat, J. Weiner, and K. Campbell for comments on the manuscript. We also thank C. Cosme, S. Dahlke, and S. Ghodsi for technical assistance.
Funding: This work was supported by NIH grant R01 MH 087463 to T.A., NIH grant K99 AG 068306 and the Nellie Ball Trust to S.C., The Gary & LaDonna Wicklund Research Fund for Cognitive Memory Disorders to T.A., and The University of Iowa Hawkeye Intellectual and Developmental Disabilities Research Center (HAWK-IDDRC) P50 HD103556 to T.A. and Lane Strathearn. T.A. is also supported by the Roy J. Carver Chair in Neuroscience.
Author contributions: S.C. and T.A. conceived the study. S.C. and T.A. designed the experiments with input from J.M. and K.P.G. S.C. performed the behavioral tasks, stereotactic surgeries, and molecular biology experiments. E.B. and Y.V. performed the bioinformatic analysis. U.M., J.D.L., A.L.Y., and E.N.W. performed biochemical experiments and analyzed behavioral results. U.M. performed cell culture experiments. M.S.S. performed electrophysiological experiments. S.C. and T.A. interpreted the results and wrote the article.
Competing interests: The authors declare that they have no competing interests.
Data and materials availability: All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials. All the uncropped Western blots and raw data are also provided. The RNA-seq data have been deposited in the National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO) and are accessible through GEO Series accession number GSE167566. The code for analyses and figures related to RNA-seq data can be accessed through GitHub (https://github.com/ethanbahl/chatterjee2021_nr4a). The data and codes can also be accessed through Zenodo (https://doi.org/10.5281/zenodo.5879784).
Supplementary Materials
This PDF file includes:
Figs. S1 to S13
Tables S1 to S3
REFERENCES AND NOTES
- 1.Roy D. S., Arons A., Mitchell T. I., Pignatelli M., Ryan T. J., Tonegawa S., Memory retrieval by activating engram cells in mouse models of early Alzheimer’s disease. Nature 531, 508–512 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Weston P. S. J., Nicholas J. M., Henley S. M. D., Liang Y., Macpherson K., Donnachie E., Schott J. M., Rossor M. N., Crutch S. J., Butler C. R., Zeman A. Z., Fox N. C., Accelerated long-term forgetting in presymptomatic autosomal dominant Alzheimer’s disease: A cross-sectional study. Lancet Neurol. 17, 123–132 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chatterjee S., Cassel R., Schneider-Anthony A., Merienne K., Cosquer B., Tzeplaeff L., Halder Sinha S., Kumar M., Chaturbedy P., Eswaramoorthy M., le Gras S., Keime C., Bousiges O., Dutar P., Petsophonsakul P., Rampon C., Cassel J. C., Buée L., Blum D., Kundu T. K., Boutillier A. L., Reinstating plasticity and memory in a tauopathy mouse model with an acetyltransferase activator. EMBO Mol. Med. 10, e8587 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Hawk J. D., Bookout A. L., Poplawski S. G., Bridi M., Rao A. J., Sulewski M. E., Kroener B. T., Manglesdorf D. J., Abel T., NR4A nuclear receptors support memory enhancement by histone deacetylase inhibitors. J. Clin. Invest. 122, 3593–3602 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Marco A., Meharena H. S., Dileep V., Raju R. M., Davila-Velderrain J., Zhang A. L., Adaikkan C., Young J. Z., Gao F., Kellis M., Tsai L. H., Mapping the epigenomic and transcriptomic interplay during memory formation and recall in the hippocampal engram ensemble. Nat. Neurosci. 23, 1606–1617 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Alberini C. M., Kandel E. R., The regulation of transcription in memory consolidation. Cold Spring Harb. Perspect. Biol. 7, a021741 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.McNulty S. E., Barrett R. M., Vogel-Ciernia A., Malvaez M., Hernandez N., Davatolhagh M. F., Matheos D. P., Schiffman A., Wood M. A., Differential roles for Nr4a1 and Nr4a2 in object location vs. object recognition long-term memory. Learn Mem. 19, 588–592 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bookout A. L., Jeong Y., Downes M., Yu R. T., Evans R. M., Mangelsdorf D. J., Anatomical profiling of nuclear receptor expression reveals a hierarchical transcriptional network. Cell 126, 789–799 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Chatterjee S., Angelakos C. C., Bahl E., Hawk J. D., Gaine M. E., Poplawski S. G., Schneider-Anthony A., Yadav M., Porcari G. S., Cassel J. C., Giese K. P., Michaelson J. J., Lyons L. C., Boutillier A. L., Abel T., The CBP KIX domain regulates long-term memory and circadian activity. BMC Biol. 18, 155 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Vecsey C. G., Hawk J. D., Lattal K. M., Stein J. M., Fabian S. A., Attner M. A., Cabrera S. M., McDonough C. B., Brindle P. K., Abel T., Wood M. A., Histone deacetylase inhibitors enhance memory and synaptic plasticity via CREB:CBP-dependent transcriptional activation. J. Neurosci. 27, 6128–6140 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Carpenter M. D., Hu Q., Bond A. M., Lombroso S. I., Czarnecki K. S., Lim C. J., Song H., Wimmer M. E., Pierce R. C., Heller E. A., Nr4a1 suppresses cocaine-induced behavior via epigenetic regulation of homeostatic target genes. Nat. Commun. 11, 504 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bridi M. S., Abel T., The NR4A orphan nuclear receptors mediate transcription-dependent hippocampal synaptic plasticity. Neurobiol. Learn. Mem. 105, 151–158 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kwapis J. L., Alaghband Y., López A. J., Long J. M., Li X., Shu G., Bodinayake K. K., Matheos D. P., Rapp P. R., Wood M. A., HDAC3-mediated repression of the Nr4a family contributes to age-related impairments in long-term memory. J. Neurosci. 39, 4999–5009 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chatterjee S., Walsh E. N., Yan A. L., Giese K. P., Safe S., Abel T., Pharmacological activation of Nr4a rescues age-associated memory decline. Neurobiol. Aging 85, 140–144 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Skerrett R., Malm T., Landreth G., Nuclear receptors in neurodegenerative diseases. Neurobiol. Dis. 72 ( Pt. A), 104–116 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jeon S. G., Yoo A., Chun D. W., Hong S. B., Chung H., Kim J. I., Moon M., The critical role of Nurr1 as a mediator and therapeutic target in Alzheimer’s disease-related pathogenesis. Aging Dis. 11, 705–724 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Park A. J., Havekes R., Fu X., Hansen R., Tudor J. C., Peixoto L., Li Z., Wu Y. C., Poplawski S. G., Baraban J. M., Abel T., Learning induces the translin/trax RNase complex to express activin receptors for persistent memory. eLife 6, e27872 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Meunier L., Usherwood Y. K., Chung K. T., Hendershot L. M., A subset of chaperones and folding enzymes form multiprotein complexes in endoplasmic reticulum to bind nascent proteins. Mol. Biol. Cell 13, 4456–4469 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Liu X., Wang Y., Lu H., Li J., Yan X., Xiao M., Hao J., Alekseev A., Khong H., Chen T., Huang R., Wu J., Zhao Q., Wu Q., Xu S., Wang X., Jin W., Yu S., Wang Y., Wei L., Wang A., Zhong B., Ni L., Liu X., Nurieva R., Ye L., Tian Q., Bian X. W., Dong C., Genome-wide analysis identifies NR4A1 as a key mediator of T cell dysfunction. Nature 567, 525–529 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zhang X. M., Yan X. Y., Zhang B., Yang Q., Ye M., Cao W., Qiang W. B., Zhu L. J., du Y. L., Xu X. X., Wang J. S., Xu F., Lu W., Qiu S., Yang W., Luo J. H., Activity-induced synaptic delivery of the GluN2A-containing NMDA receptor is dependent on endoplasmic reticulum chaperone Bip and involved in fear memory. Cell Res. 25, 818–836 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wang J., Lee J., Liem D., Ping P., HSPA5 gene encoding Hsp70 chaperone BiP in the endoplasmic reticulum. Gene 618, 14–23 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hartl F. U., Bracher A., Hayer-Hartl M., Molecular chaperones in protein folding and proteostasis. Nature 475, 324–332 (2011). [DOI] [PubMed] [Google Scholar]
- 23.Cortez L., Sim V., The therapeutic potential of chemical chaperones in protein folding diseases. Prion 8, 197–202 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Perlmutter D. H., Chemical chaperones: A pharmacological strategy for disorders of protein folding and trafficking. Pediatr. Res. 52, 832–836 (2002). [DOI] [PubMed] [Google Scholar]
- 25.Rubenstein R. C., Egan M. E., Zeitlin P. L., In vitro pharmacologic restoration of CFTR-mediated chloride transport with sodium 4-phenylbutyrate in cystic fibrosis epithelial cells containing delta F508-CFTR. J. Clin. Invest. 100, 2457–2465 (1997). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Burrows J. A., Willis L. K., Perlmutter D. H., Chemical chaperones mediate increased secretion of mutant alpha 1-antitrypsin (α1-AT) Z: A potential pharmacological strategy for prevention of liver injury and emphysema in alpha 1-AT deficiency. Proc. Natl. Acad. Sci. U.S.A. 97, 1796–1801 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Andersen E., Chollet M. E., Baroni M., Pinotti M., Bernardi F., Skarpen E., Sandset P. M., Skretting G., The effect of the chemical chaperone 4-phenylbutyrate on secretion and activity of the p.Q160R missense variant of coagulation factor FVII. Cell Biosci. 9, 69 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cuadrado-Tejedor M., Ricobaraza A. L., Torrijo R., Franco R., Garcia-Osta A., Phenylbutyrate is a multifaceted drug that exerts neuroprotective effects and reverses the Alzheimer’s disease-like phenotype of a commonly used mouse model. Curr. Pharm. Des. 19, 5076–5084 (2013). [DOI] [PubMed] [Google Scholar]
- 29.Ricobaraza A., Cuadrado-Tejedor M., Marco S., Perez-Otano I., Garcia-Osta A., Phenylbutyrate rescues dendritic spine loss associated with memory deficits in a mouse model of Alzheimer disease. Hippocampus 22, 1040–1050 (2012). [DOI] [PubMed] [Google Scholar]
- 30.Ricobaraza A., Cuadrado-Tejedor M., Pérez-Mediavilla A., Frechilla D., del Río J., García-Osta A., Phenylbutyrate ameliorates cognitive deficit and reduces tau pathology in an Alzheimer’s disease mouse model. Neuropsychopharmacology 34, 1721–1732 (2009). [DOI] [PubMed] [Google Scholar]
- 31.Wiley J. C., Pettan-Brewer C., Ladiges W. C., Phenylbutyric acid reduces amyloid plaques and rescues cognitive behavior in AD transgenic mice. Aging Cell 10, 418–428 (2011). [DOI] [PubMed] [Google Scholar]
- 32.Mimori S., Ohtaka H., Koshikawa Y., Kawada K., Kaneko M., Okuma Y., Nomura Y., Murakami Y., Hamana H., 4-Phenylbutyric acid protects against neuronal cell death by primarily acting as a chemical chaperone rather than histone deacetylase inhibitor. Bioorg. Med. Chem. Lett. 23, 6015–6018 (2013). [DOI] [PubMed] [Google Scholar]
- 33.Moon M., Jung E. S., Jeon S. G., Cha M. Y., Jang Y., Kim W., Lopes C., Mook-Jung I., Kim K. S., Nurr1 (NR4A2) regulates Alzheimer’s disease-related pathogenesis and cognitive function in the 5XFAD mouse model. Aging Cell 18, e12866 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Miller J. A., Guillozet-Bongaarts A., Gibbons L. E., Postupna N., Renz A., Beller A. E., Sunkin S. M., Ng L., Rose S. E., Smith K. A., Szafer A., Barber C., Bertagnolli D., Bickley K., Brouner K., Caldejon S., Chapin M., Chua M. L., Coleman N. M., Cudaback E., Cuhaciyan C., Dalley R. A., Dee N., Desta T., Dolbeare T. A., Dotson N. I., Fisher M., Gaudreault N., Gee G., Gilbert T. L., Goldy J., Griffin F., Habel C., Haradon Z., Hejazinia N., Hellstern L. L., Horvath S., Howard K., Howard R., Johal J., Jorstad N. L., Josephsen S. R., Kuan C. L., Lai F., Lee E., Lee F., Lemon T., Li X., Marshall D. A., Melchor J., Mukherjee S., Nyhus J., Pendergraft J., Potekhina L., Rha E. Y., Rice S., Rosen D., Sapru A., Schantz A., Shen E., Sherfield E., Shi S., Sodt A. J., Thatra N., Tieu M., Wilson A. M., Montine T. J., Larson E. B., Bernard A., Crane P. K., Ellenbogen R. G., Keene C. D., Lein E., Neuropathological and transcriptomic characteristics of the aged brain. eLife 6, e31126 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Santacruz K., Lewis J., Spires T., Paulson J., Kotilinek L., Ingelsson M., Guimaraes A., DeTure M., Ramsden M., McGowan E., Forster C., Yue M., Orne J., Janus C., Mariash A., Kuskowski M., Hyman B., Hutton M., Ashe K. H., Tau suppression in a neurodegenerative mouse model improves memory function. Science 309, 476–481 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ramsden M., Kotilinek L., Forster C., Paulson J., McGowan E., SantaCruz K., Guimaraes A., Yue M., Lewis J., Carlson G., Hutton M., Ashe K. H., Age-dependent neurofibrillary tangle formation, neuron loss, and memory impairment in a mouse model of human tauopathy (P301L). J. Neurosci. 25, 10637–10647 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Yue M., Hanna A., Wilson J., Roder H., Janus C., Sex difference in pathology and memory decline in rTg4510 mouse model of tauopathy. Neurobiol. Aging 32, 590–603 (2011). [DOI] [PubMed] [Google Scholar]
- 38.Cook C., Dunmore J. H., Murray M. E., Scheffel K., Shukoor N., Tong J., Castanedes-Casey M., Phillips V., Rousseau L., Penuliar M. S., Kurti A., Dickson D. W., Petrucelli L., Fryer J. D., Severe amygdala dysfunction in a MAPT transgenic mouse model of frontotemporal dementia. Neurobiol. Aging 35, 1769–1777 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gamache J., Benzow K., Forster C., Kemper L., Hlynialuk C., Furrow E., Ashe K. H., Koob M. D., Factors other than hTau overexpression that contribute to tauopathy-like phenotype in rTg4510 mice. Nat. Commun. 10, 2479 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Chen Y., Wang Y., Ertürk A., Kallop D., Jiang Z., Weimer R. M., Kaminker J., Sheng M., Activity-induced Nr4a1 regulates spine density and distribution pattern of excitatory synapses in pyramidal neurons. Neuron 83, 431–443 (2014). [DOI] [PubMed] [Google Scholar]
- 41.Frost B., Hemberg M., Lewis J., Feany M. B., Tau promotes neurodegeneration through global chromatin relaxation. Nat. Neurosci. 17, 357–366 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Montalbano M., McAllen S., Puangmalai N., Sengupta U., Bhatt N., Johnson O. D., Kharas M. G., Kayed R., RNA-binding proteins Musashi and tau soluble aggregates initiate nuclear dysfunction. Nat. Commun. 11, 4305 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Schueller E., Paiva I., Blanc F., Wang X. L., Cassel J. C., Boutillier A. L., Bousiges O., Dysregulation of histone acetylation pathways in hippocampus and frontal cortex of Alzheimer’s disease patients. Eur. Neuropsychopharmacol. 33, 101–116 (2020). [DOI] [PubMed] [Google Scholar]
- 44.Schneider A., Chatterjee S., Bousiges O., Selvi B. R., Swaminathan A., Cassel R., Blanc F., Kundu T. K., Boutillier A. L., Acetyltransferases (HATs) as targets for neurological therapeutics. Neurotherapeutics 10, 568–588 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chatterjee S., Mizar P., Cassel R., Neidl R., Selvi B. R., Mohankrishna D. V., Vedamurthy B. M., Schneider A., Bousiges O., Mathis C., Cassel J. C., Eswaramoorthy M., Kundu T. K., Boutillier A. L., A novel activator of CBP/p300 acetyltransferases promotes neurogenesis and extends memory duration in adult mice. J. Neurosci. 33, 10698–10712 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Benito E., Urbanke H., Ramachandran B., Barth J., Halder R., Awasthi A., Jain G., Capece V., Burkhardt S., Navarro-Sala M., Nagarajan S., Schütz A. L., Johnsen S. A., Bonn S., Lührmann R., Dean C., Fischer A., HDAC inhibitor-dependent transcriptome and memory reinstatement in cognitive decline models. J. Clin. Invest. 125, 3572–3584 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Janczura K. J., Volmar C. H., Sartor G. C., Rao S. J., Ricciardi N. R., Lambert G., Brothers S. P., Wahlestedt C., Inhibition of HDAC3 reverses Alzheimer’s disease-related pathologies in vitro and in the 3xTg-AD mouse model. Proc. Natl. Acad. Sci. U.S.A. 115, E11148–E11157 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 48.Graff J., Rei D., Guan J.-S., Wang W.-Y., Seon J., Hennig K. M., Nieland T. J. F., Fass D. M., Kao P. F., Kahn M., Su S. C., Samiei A., Joseph N., Haggarty S. J., Delalle I., Tsai L.-H., An epigenetic blockade of cognitive functions in the neurodegenerating brain. Nature 483, 222–226 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Goodwin L. O., Splinter E., Davis T. L., Urban R., He H., Braun R. E., Chesler E. J., Kumar V., van Min M., Ndukum J., Philip V. M., Reinholdt L. G., Svenson K., White J. K., Sasner M., Lutz C., Murray S. A., Large-scale discovery of mouse transgenic integration sites reveals frequent structural variation and insertional mutagenesis. Genome Res. 29, 494–505 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Engelhard H. H., Homer R. J., Duncan H. A., Rozental J., Inhibitory effects of phenylbutyrate on the proliferation, morphology, migration and invasiveness of malignant glioma cells. J. Neurooncol. 37, 97–108 (1998). [DOI] [PubMed] [Google Scholar]
- 51.Havekes R., Bruinenberg V. M., Tudor J. C., Ferri S. L., Baumann A., Meerlo P., Abel T., Transiently increasing cAMP levels selectively in hippocampal excitatory neurons during sleep deprivation prevents memory deficits caused by sleep loss. J. Neurosci. 34, 15715–15721 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Vogel-Ciernia A., Wood M. A., Examining object location and object recognition memory in mice. Curr. Protoc. Neurosci. 69, 8.31.1–8.31.17 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Shetty M. S., Sharma M., Hui N. S., Dasgupta A., Gopinadhan S., Sajikumar S., Investigation of synaptic tagging/capture and cross-capture using acute hippocampal slices from rodents. J. Vis. Exp., 53008 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Dobin A., Davis C. A., Schlesinger F., Drenkow J., Zaleski C., Jha S., Batut P., Chaisson M., Gingeras T. R., STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Liao Y., Smyth G. K., Shi W., featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014). [DOI] [PubMed] [Google Scholar]
- 56.R. C. Team, R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019). [Google Scholar]
- 57.Risso D., Schwartz K., Sherlock G., Dudoit S., GC-content normalization for RNA-Seq data. BMC Bioinformatics 12, 480 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Risso D., Ngai J., Speed T. P., Dudoit S., Normalization of RNA-seq data using factor analysis of control genes or samples. Nat. Biotechnol. 32, 896–902 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Peixoto L., Risso D., Poplawski S. G., Wimmer M. E., Speed T. P., Wood M. A., Abel T., How data analysis affects power, reproducibility and biological insight of RNA-seq studies in complex datasets. Nucleic Acids Res. 43, 7664–7674 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Robinson M. D., McCarthy D. J., Smyth G. K., edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.McCarthy D. J., Chen Y., Smyth G. K., Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Chen Y., Lun A. T., Smyth G. K., From reads to genes to pathways: Differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Res. 5, 1438 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Bindea G., Mlecnik B., Hackl H., Charoentong P., Tosolini M., Kirilovsky A., Fridman W. H., Pagès F., Trajanoski Z., Galon J., ClueGO: A cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Shannon P., Markiel A., Ozier O., Baliga N. S., Wang J. T., Ramage D., Amin N., Schwikowski B., Ideker T., Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Doncheva N. T., Morris J. H., Gorodkin J., Jensen L. J., Cytoscape StringApp: Network analysis and visualization of proteomics data. J. Proteome Res. 18, 623–632 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Szklarczyk D., Franceschini A., Wyder S., Forslund K., Heller D., Huerta-Cepas J., Simonovic M., Roth A., Santos A., Tsafou K. P., Kuhn M., Bork P., Jensen L. J., von Mering C., STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Kaech S., Banker G., Culturing hippocampal neurons. Nat. Protoc. 1, 2406–2415 (2006). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figs. S1 to S13
Tables S1 to S3