Abstract
Large scale screening of health care workers and the general population for asymptomatic COVID-19 infection requires modalities that are amenable to testing at scale while retaining acceptable levels of sensitivity and specificity.
This study evaluated a novel COVID-19 Direct-RT LAMP assay using saliva samples in asymptomatic individuals by comparison to RT-PCR. Additional studies were performed using VTM collected from routine diagnostic testing. Analytical sensitivity was determined for Direct RT-LAMP assay using the WHO International Standard. Finally, quantified results from RT-PCR testing of 9177 nose and throat swabs obtained from routine diagnostic testing were used to estimate the sensitivity of Direct RT-LAMP using the limit of detection curve obtained from the analytical sensitivity data.
Results from saliva testing demonstrated a sensitivity of 40.91% and a specificity of 100% for Direct RT-LAMP. The sensitivity and specificity for nose and throat swabs were 44.85% and 100% respectively. The 95% limit of detection (LOD) for Direct RT-LAMP was log 7.13 IU/ml (95% 6.9–7.5). The estimated sensitivity for Direct-RT LAMP based on the results of 9117 nose and throat swabs was 34% and 45% for saliva and VTM respectively.
The overall diagnostic sensitivity of Direct RT-LAMP was low compared to RT-PCR. Testing of nose and throat swabs and estimating the sensitivity based on a large cohort of clinical samples demonstrated similar results. This study highlights the importance of utilising the prospective collection of samples from the intended target population in the assessment of diagnostic sensitivity.
Keywords: Direct RT-LAMP, Real-time PCR, Saliva, COVID-19, SARS-CoV-2, Asymptomaitc, Screening
1. Introduction
It is now established that SARS-CoV-2 can be detected in asymptomatic individuals with estimates as high as 30.8% [1]. Throughout the pandemic there has been considerable concern around asymptomatic COVID-19 contributing to nosocomial infection. One screening pilot in a large UK teaching hospital found that 3% of staff had asymptomatic infection, of which 57% did not go on to develop symptoms [2]. There is also evidence to support the fact that asymptomatic infection in health care workers is associated with transmission in these settings [3,4]. There has therefore been a compelling case for COVID-19 screening in asymptomatic healthcare workers to reduce the risk of transmission to patients and in England this has been reflected in national guidance [5]. Large scale testing requires modalities that are amenable to testing at scale while retaining acceptable levels of sensitivity and specificity. Recent studies have shown that SARS-CoV-2 RNA can be detected in saliva, with viral loads peaking during the first week of symptoms [6]. Some studies have shown that viral loads in saliva are comparable to or higher than nasopharyngeal swabs [7,8] while others have observed lower viral loads [9]. The aim of this study was to evaluate a novel Direct-RT LAMP assay for the detection of SARS-CoV-2 in asymptomatic individuals using saliva samples. The study involved two phases; the first was conducted in the general population and was part of multicentre evaluation in collaboration with the UK Department of Health and Social Care (data from which has previous been published [10]) and the second focused on asymptomatic health care workers in secondary care as a continued evaluation of the performance of the assay.
2. Methods
2.1. Saliva sample collection and transport
Saliva samples were collected in two phases: phase one included asymptomatic members of the general population in the city centre of Salford (Greater Manchester, UK) between the 14th September and 15th October 2020, phase two included asymptomatic staff members at Manchester University NHS Foundation trust and the Christie NHS Foundation Trust between 20th November and the 22nd December 2020. Collection kits were provided, and samples were tested on the same day of sampling.
2.2. Nose and throat swab collection
An additional evaluation was made on the performance of Direct RT-LAMP with nose and throat swabs collected in Remel viral transport medium (VTM) (Thermo Fisher). The OptiGene IFU for Direct RT-LAMP lists nose and throat swabs collected in Virocult® media alongside saliva samples as an accepted sample type [11]. Specimens submitted for routine testing were collected over a period of three weeks, concurrent with phase 2 of the study. Previous positive patients were excluded from the assessment and then randomly selected before being unlinked and anonymised. Blinded testing was performed by a second operator within 8 h of receipt and before freezing.
2.3. Sample analysis
2.3.1. RT-PCR
Saliva samples were inactivated by mixing 300 µl of sample with 300 µl Qiagen AL buffer (Qiagen, Hilden, Germany) and heated at 80 °C for 15 min. RT-PCR was split over 2 workstreams using CDC N1 and N2 primers and probes [12]. The first workstream used the Roche Flow automated sample protocol; 200 µl of sample/AL was extracted on a MagNAPure 96 MP96 platform and RT-PCR performed on a Lightcycler 480. The second workstream used nucleic acid extraction on the Qiagen QIAsymphony extracting 200 µl of sample and eluting into 60 µl followed by RT-PCR run on an Applied Biosystems 7500 Fast PCR system (Thermofisher). Both workstreams utilised the same RT-PCR assay, 5 µl of extracted RNA was added to a master mix composed of 5 µl of Applied Biosystems™ Taqpath™ 1-Step multiplex master mix no rox (Thermofisher), 1.5 µl of both N1 and N2 primer probe sets (Integrated DNA Technologies, USA) and 0.5 µl of Taqman® Human β2M endogenous control primer probe set (Thermofisher). Cycling parameters were the same for both the Lightcycler 480 and 7500 Fast PCR systems, 25 °C for 2 min, 55 °C for 15 min, 95 °C for 2 min and 45 cycles of 95 °C 3 secs and 55 °C for 30 secs. A nuclease free non template control (NTC) and SARS-CoV-2 whole genome RNA positive control (Vircell, Grenada, Spain) were included on each run. Confirmation of any positives with a Ct value ≥30 was performed using the Biofire® respiratory 2.1 panel, incorporating a SARS-CoV-2 target, (bioMérieux, Marcy-l'Étoile, France) according to the manufacturer's IFU. Any sample positive by the N1N2 assay but negative in the Biofire assay were reported as indeterminate.
To investigate the variability of Ct value caused by different gene targets, further testing was performed on nucleic acid extracts of positive samples using the genesig® SARS-CoV-2 winterplex assay (Primerdesign Ltd, Chandler's Ford, UK) according to the manufacturer's IFU. In brief 8 ul of previously extracted sample was added to master mix composed of 10 µl of onestep master mix and 2 µl of Orf 1a/b/S primer and probe mix followed by RT-PCR run on a Fast 7500 Fast PCR system with cycling parameters of 55 °C for 10 min, 95 °C for 2 min and 45 cycles of 95 °C 10 secs and 60 °C for 60 secs.
Samples for the nose and throat swab validation were tested using the Cobas SARS-CoV-2 assay (Roche, Basel, Switzerland) according to the manufacturer's instructions. The assay targets both Orf1ab and E genes, any single target or >Ct 35 positive results were confirmed using the Biofire SARS-CoV-2 assay.
2.3.2. Direct RT-LAMP
Samples were inactivated by mixing 50 µl of sample with 50 µl of RapiLyze buffer (OptiGene, Horsham, UK) in a 96 well MicroAmp Optical Reaction Plate (Thermofisher) and held at 98 °C for two minutes on an Applied Biosystems 9600 Thermal Cycler (Thermofisher). A negative extract control containing Rapilyze alone was included with each sample set.
2.3.3. Phase 1
During phase 1 of the study Direct SARS-CoV-2 RT-LAMP was performed according to the manufacturer's protocol v1.1. All Direct RT-LAMP tests were performed on inactivated saliva samples within 30 min post-inactivation. All samples were tested in singles. Master mix comprised 17.5 µl of Direct RT-LAMP master mix and 2.5 µl of 10X COVID 19 (OptiGene) to which 5 µl of inactivated sample was added and mixed. Direct RT-LAMP was performed on a Genie HT platform (OptiGene), using an amplification stage of 65 °C for 20 min followed by anneal curve analysis, 98 °C for 1 min then cooled to 80 °C at a rate of 0.05 °C/s. In addition to the Rapilyze extraction control, a nuclease free water NTC, a DNA CD-COV19–100 positive control (OptiGene) and an additional SARS-CoV-2 whole genome RNA positive control were included on each run. Samples were identified as positive automatically by the Genie HT platform through detection of increased fluorescence combined with an anneal peak within the defined range. All sample amplification plots were manually checked to ensure any potential signals were not missed.
2.3.4. Phase 2
Direct RT-LAMP in phase 2 of the study was performed using the Covid 19 DirectPlus RT-LAMP protocol. The modified DirectPlus RT-LAMP thermal protocol was provided by the manufacturer. The modification from phase 1 was an alteration to the interpretive software, directed on the final anneal analysis.
2.3.5. Analytical sensitivity
Analytical sensitivity of the Direct RT-LAMP assay was determined using a dilution of the 1st WHO International Standard for SARS-CoV-2 RNA (NIBSC 20/146). The International standard was used to create doubling dilution series from an initial concentration of log 7.7 IU/ml down to log 4.99 IU/ml. Separate dilution series were performed in both SARS-CoV-2 RNA negative saliva and Sigma Virocult® VTM media (Medical Wire, Corsham, UK). Multiple replicates were tested at each dilution, two of the log 7.7 IU/ml dilution, 6 of the log 7.4 IU/ml dilution and 20 replicates for all other dilutions. All testing was performed using the Direct Plus RT-LAMP 20-minute thermal protocol.
3. Estimated proportion of clinical samples previously tested positive by the Roche Cobas SARS-Cov-2 assay that would be detected by Direct Lamp
Results for 9177 anonymised clinical samples tested for SARS-CoV-2 Orf1a/b using the Cobas® SARS-CoV-2 RT-PCR assay (Roche, Basel, Switzerland) were quantified by utilising a standard curve generated from a 10-fold dilution series of the NIBSC SARS-CoV-2 international standard. The estimated% LOD curve for the Direct LAMP assay determined by Probit analysis was then used to estimate the proportion of positive Cobas® SARS-CoV-2 RT-PCR assay samples that would be detected by Direct RT-LAMP.
3.1. Statistical analysis
The sensitivity, specificity and Probit analysis were performed in SPSS (IBM Corp. Released 2020. IBM SPSS Statistics for Windows, Version 27.0. Armonk, NY: IBM Corp) software to determine the 50% and 95% limit of detection (LOD) in IU/ml. Figs. 1 and 2 were produced in MedCalc (MedCalc Software Ltd, 2020).
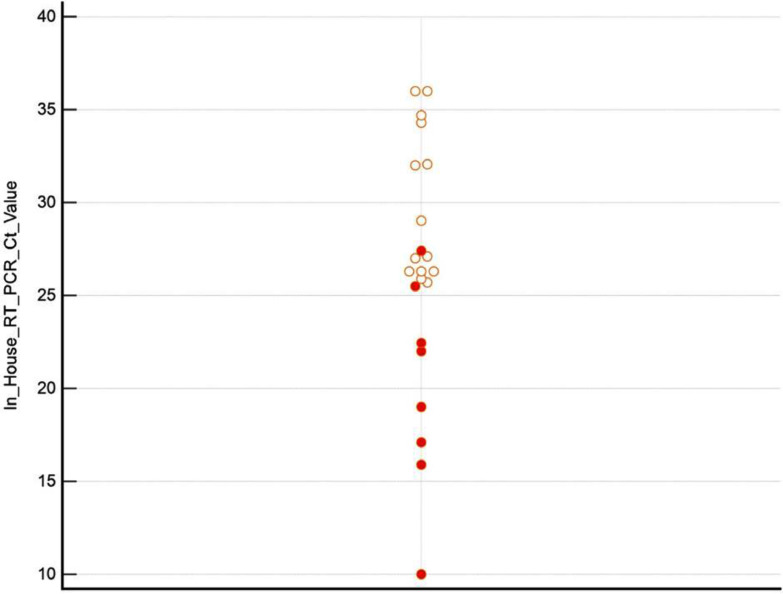
Fig. 1.
Dot distribution plot of the N1N2 assay Ct. values from confirmed RT-PCR positive saliva samples. Red circles were positive by RT-PCR and Direct RT-LAMP, clear circles are RT-PCR positive only.
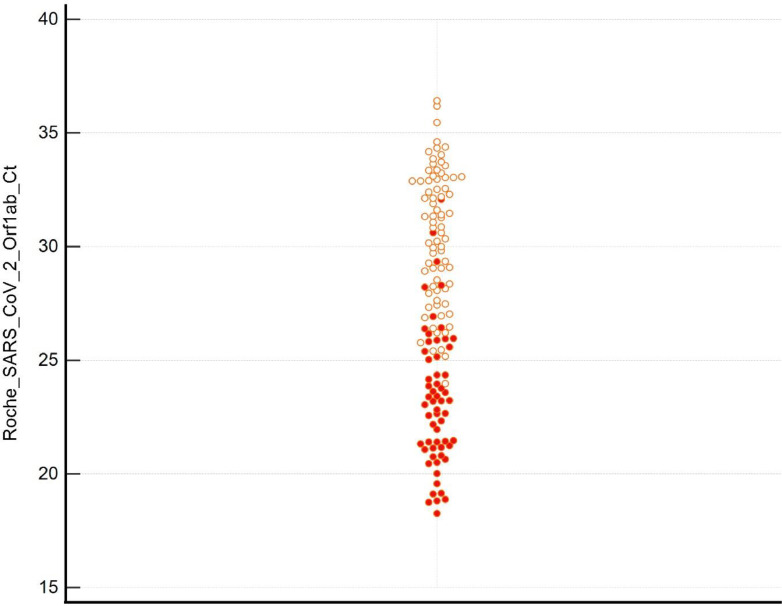
Fig. 2.
Dot distribution plot of the N1N2 assay Ct. values from confirmed RT-PCR positive nose and throat swabs. Red circles represent specimens positive by both RT-PCR and Direct RT-LAMP, clear circles positive by RT-PCR only.
The estimated percentages of LOD for Direct RT-LAMP were obtained from a Probit model where the Direct RT-LAMP test result was the outcome and viral load (log10) was the predictor. These estimated percentages were represented as the LOD curves in Fig. 3 . The Probit model built from the Direct RT-LAMP test was then used to predict the probability of each Roche sample being tested positive in the Direct RT-LAMP test, using the predict function in R, version 4.0.3. The estimated sensitivity of the Direct RT-LAMP test was calculated as the sum of the predicted probabilities of the Roche samples divided by the sample size 9177.
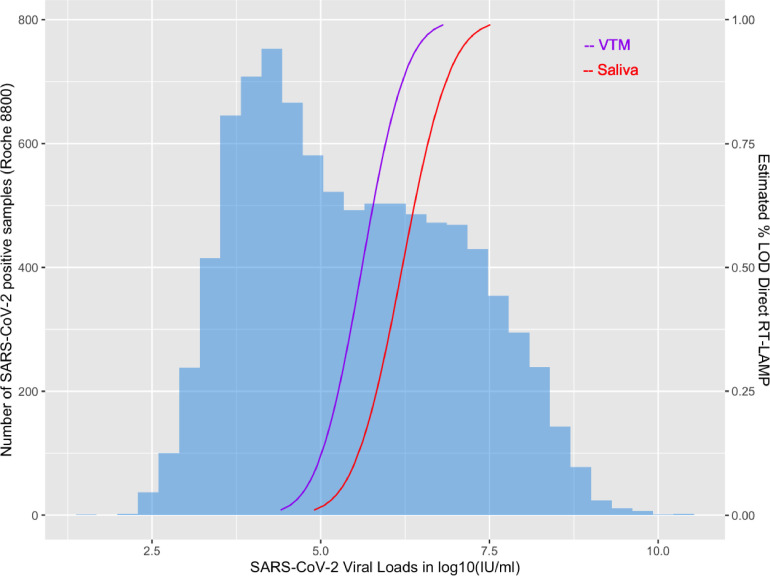
Fig. 3.
Distribution of viral loads from RT-PCR positive samples and% LOD curves for Direct RT-LAMP testing of saliva and VTM.
4. Results
A total of 1383 samples were collected during the two phases of the saliva sample study. The percentage of samples rejected due to issues with viscosity was 7%.
4.1. Analytical sensitivity of saliva samples
The results from testing multiple replicates of each dilution are listed in Table 1 . The 95% LOD for detection of SARS-CoV-2 RNA from saliva samples was log 7.13 IU/ml (95% 6.9–7.5), and for Virocult® VTM was log 6.46 IU/ml (95%CI 6.3–6.8).
Table 1.
Results for analytical sensitivity of Direct RT-LAMP. Table lists the concentration in IU/ml of a doubling dilution series performed both in saliva and Virocult medium, the total number of replicates tested for each dilution and the result at each dilution.
| Quantity |
Total No. Tested | Direct RT-LAMP |
||||
|---|---|---|---|---|---|---|
| SARS-CoV-2 RNA |
Saliva |
Virocult |
||||
| IU/ml | Log IU/ml | Pos | Neg | Pos | Neg | |
| 50,118,723 | 7.70 | 2 | 2 | 0 | 2 | 0 |
| 25,059,362 | 7.40 | 6 | 6 | 0 | 6 | 0 |
| 12,529,681 | 7.10 | 20 | 19 | 1 | 20 | 0 |
| 6,264,840 | 6.80 | 20 | 18 | 2 | 20 | 0 |
| 3,132,420 | 6.50 | 20 | 13 | 7 | 20 | 0 |
| 1,566,210 | 6.19 | 20 | 9 | 11 | 18 | 2 |
| 783,105 | 5.89 | 20 | 5 | 15 | 12 | 8 |
| 391,553 | 5.59 | 20 | 4 | 16 | 7 | 13 |
| 195,776 | 5.29 | 20 | 0 | 20 | 6 | 14 |
| 97,888 | 4.99 | 20 | 1 | 19 | 4 | 16 |
4.2. Diagnostic sensitivity and specificity saliva samples
A total of 22 samples were positive by RT-PCR (Table 2 ). Nine samples were positive by Direct RT-LAMP, with an overall sensitivity of 40.91% and a specificity of 100% (Table 3 ). In phase 1, two RT-PCR positive samples were characterised as negative by the Direct RT-LAMP instrument but showed strong amplification below Tp 10:00 and produced peaks with anneal temp analysis around the correct temperature and so were recorded as positive.
Table 2.
Summary of results for Direct RT-LAMP testing of saliva samples collected from asymptomatic individuals during phase 1 and 2. Samples were deemed invalid if the total volume was too low or the consistency was too viscous to pipette.
| Phase | Samples | RT-PCR Positive | RT-PCR Negative | RT-PCR Indeterminate | Direct LAMP Positive | Invalid | % Invalid |
|---|---|---|---|---|---|---|---|
| 1 | 1383 | 15 | 1285 | 1 | 7 | 82 | 6 |
| 2 | 929 | 7 | 831 | 5 | 1 | 86 | 9 |
| Total | 2312 | 22 | 2116 | 5 | 8 | 168 | 7 |
Table 3.
Sensitivity and Specificity of Direct RT-LAMP in saliva collected from asymptomatic individuals relative to reference N1N2 RT-PCR assay. Only positive and negative results from reference RT-PCR were included in the analysis, indeterminate results were excluded. The sensitivity and specificity results for phase 1 and phase 2 are listed separately with the overall results in the final row.
| RT-PCR |
Sensitivity%* | Specificity%* | ||||
|---|---|---|---|---|---|---|
| Pos | Neg | |||||
| Direct RT-LAMP | Phase 1 | Pos | 7 | 0 | 46.67 (21.27–73.41) | 100 (99.71–100) |
| Neg | 8 | 1285 | ||||
| Phase 2 | Pos | 2 | 0 | 28.57 (3.67–70.96) | 100 (99.56–100) | |
| Neg | 5 | 831 | ||||
| Overall | Pos | 9 | 0 | 40.91 (20.71–63.65) | 100 (99.83–100) | |
| Neg | 13 | 2116 | ||||
SARS Cov-2 samples positive by the N1/N2 PCR assay were confirmed in 21 of the 22 specimens by the Wintreplex assay (Table 4 ), with a single sample with a Ct of 34.7 recorded as negative. The Ct values produced by both assays were similar, with a mean difference in Ct of 0.9.
Table 4.
Comparative results from testing saliva samples using N1N2, Direct RT-LAMP and Winterplex assays.
| Study Phase | N1N2 Assay |
Direct RT-LAMP |
Winterplex Orf1a/b |
|||
|---|---|---|---|---|---|---|
| Result | Ct. value | Result | TP Value | Result | Ct. value | |
| 1 | P | 10.0 | P* | P | 11.0 | |
| 1 | P | 36.0 | N | P | 36.3 | |
| 1 | P | 17.1 | P | 06:17 | P | 19.6 |
| 1 | P | 22.0 | P | 09:49 | P | 21.1 |
| 1 | P | 15.9 | P | 09:21 | P | 16.3 |
| 1 | P | 34.7 | N | N | ||
| 1 | P | 26.3 | N | P | 27.3 | |
| 1 | P | 25.5 | P* | P | 27.6 | |
| 1 | P | 34.3 | N | P | 34.9 | |
| 1 | P | 27.1 | N | P | 32.5 | |
| 1 | P | 32.1 | N | P | 34.4 | |
| 1 | P | 32.0 | N | P | 31.1 | |
| 1 | P | 29.0 | N | INS | INS | |
| 1 | P | 27.4 | P | 11:19 | INS | INS |
| 1 | P | 22.4 | P | 10:17 | INS | INS |
| 2 | P | 19.0 | P | 10:37 | P | 19.2 |
| 2 | P | 27.0 | N | P | 27.8 | |
| 2 | P | 36.0 | N | P | 35.6 | |
| 2 | P | 26.3 | N | P | 25.5 | |
| 2 | P | 25.9 | N | P | 25.8 | |
| 2 | P | 25.7 | N | P | 27.0 | |
| 2 | P | 26.3 | P⁎⁎ | 14:00 | P | 27.5 |
P, positive result: N, negative result: INS, insufficient, Ct., Cycle Threshold and TP, Time Positivity.
Two results from Direct RT-LAMP were listed as negative by the Optigene HT instrument but showed strong amplification and an anneal peak at the correct temperature, these samples were considered positive for sensitivity/specificity calculations.
Was initially negative when the first Direct Plus RT-LAMP 14:00 protocol was used, was positive on repeat using 20:00 protocol.
The sensitivity of Direct RT-LAMP is closely related to the Ct value produced by the RT-PCR assay, as seen in Fig. 1. Sensitivity was 100% on samples with a Ct <25, representing samples with a high viral burden. Sensitivity drops off rapidly above this level as the viral load declines.
4.3. Diagnostic sensitivity/specificity nose and throat swabs
Results from testing nose and throat swabs by Direct RT-LAMP are listed in Table 5 . The overall sensitivity in nose and throat swabs was similar to saliva at 44.85% (95%CI 36.32 – 53.61%) with a specificity of 100%. Fig. 2 shows the spread of Ct values producing true positive and false negative results by Direct RT-LAMP. The same pattern of high sensitivity at Ct values ≤25 and a reduced sensitivity ≥25 is displayed.
Table 5.
Sensitivity and Specificity of Direct RT-LAMP using swabs received in Remell VTM collected from asymptomatic and symptomatic individuals relative to reference Cobas® SARS-CoV-2 RT-PCR assay.
| RT-PCR |
Sensitivity%* | Specificity%* | |||
|---|---|---|---|---|---|
| Pos | Neg | ||||
| Direct RT-LAMP | Pos | 61 | 0 | 44.85 (36.32–53.61) | 100 (97.63–100) |
| Neg | 75 | 154 | |||
4.4. Estimated proportion of clinical samples previously tested positive by the Roche Cobas SARS-CoV-2 assay that would be detected by Direct Lamp
Fig. 3 provides a graphical representation of the estimated proportion of RT-PCR positive samples that would not be detected by Direct RT-LAMP by projecting the estimated% LOD curve of the OptiGene Direct RT-LAMP onto the distribution of positive RT-PCR samples. Direct RT-LAMP in salvia and VTM showed estimated sensitivities of 34% and 45% respectively, predicting that 6061 saliva and 5083 VTM samples would not have been detected by Direct RT-LAMP.
5. Discussion
The overall diagnostic sensitivity of Direct RT-LAMP for detection of SARS-CoV-2 directly from saliva samples was 40.91% combined with a specificity of 100%. Direct RT-LAMP failed to detect most samples that were positive above Ct 25 by RT-PCR. These results were obtained prospectively from combined data across two phases of asymptomatic testing in Manchester. Therefore, this study provides a representative insight into the diagnostic sensitivity of the OptiGene LAMP assay as a function of the range of viral loads present in an asymptomatic population.
This study was limited by a small number of positive samples due to the relatively low prevalence at the time. To address this, randomly selected, first-time positive nose and throat swabs from routine diagnostic testing were re-tested with the Direct RT-LAMP assay. The results showed a moderately increased sensitivity of 44.85% with an upper confidence level limit of 53.61%. The slightly increased sensitivity in comparison to saliva samples suggests that the VTM did not inhibit the LAMP reaction.
Probit analysis from the analytical sensitivity study demonstrated an estimated 95% limit of detection of log 7.13 IU/ml for Direct RT-LAMP from saliva samples and log 6.46 IU/ml from Virocult® samples. Other studies using RT-LAMP technology have also recorded analytical sensitivities of between log 5 – 5.8 log c/ml [13,14] however, the DHSC report for OptiGene Direct RT-LAMP states an analytical sensitivity of 1000 c/ml [15]. Contributing factors to the lower analytical sensitivity determined in our study include the use of whole inactivated virus rather than RNA, real saliva or VTM as the dilution matrix and the large numbers of replicates around the end point of the assay enabling a 95% Probit calculation to define the LOD These factors result in a more accurate representation of the LOD in clinical samples.
The limit of detection established for the Direct LAMP was used to predict the proportion of a large cohort of real-world positive clinical samples that would not have been detected by Direct RT-LAMP. The results show that the majority had RNA levels below the 50% LOD for Direct RT-LAMP providing further evidence supporting the diagnostic sensitivities observed in our study. Although there are limitations to this prediction, the robust determination of LOD using simulated saliva samples and the calibration of the clinical samples to the International Standard provides as accurate as possible a prediction using retrospective data.
Previous studies using OptiGene RT-LAMP reagents have shown sensitivities ranging from 34% [16] to 67% [17] . Two studies performed by the UK Department of Health and Social Care (DHSC) [15,18] used the same assay as this study. The assay performance characteristics from this study were consistent with the performance in those reports, showing reliable detection of RT-PCR positive samples with a Ct value <25, but failure to reliably detect samples with Ct values >25. However, the results from this study differ significantly regarding the observed diagnostic sensitivity. The technical validation report [18] for Direct RT LAMP lists a diagnostic sensitivity of 80% (CI 72–85%) from a mixture of 158 positive clinical and spiked saliva samples and 72% from a mixture of 173 positive clinical and spiked swab samples. The follow up rapid evaluation report [15] lists a diagnostic sensitivity of 79% (CI 73–84) from 226 positive saliva samples (including 59 spiked saliva) and a sensitivity of 70% from 199 positive clinical swab samples. For both studies, a range of high and low viral loads were selected for assessment of the diagnostic sensitivity rather than prospective collection from a target population. The sensitivity from clinical saliva samples (excluding spiked samples) is not clear from either report. The manufacturer IFU does include options for testing samples in duplicate which may increase the sensitivity of the assay at the cost of throughput but the DHSC report does not specify when this was used.
To rule out any possibility that the gene used in our saliva RT-PCR assay was a contributing factor to the observed sensitivity, we compared Ct values from positive samples obtained by the in-house N1/N2 PCR with the Winterplex assay, which targets the same ORF 1ab gene used in the Direct RT-LAMP assay. There was good concordance between the Ct values of both assays with a mean difference of only 0.90, indicating minimal variation in sensitivity. We therefore conclude that either assay could be used as a comparator for Direct LAMP.
The use of saliva samples offers some advantages for ease of collection and our study has shown a similar sensitivity to nose and throat swabs. However, direct testing of samples without nucleic extraction presents technical challenges for enzymatic amplification of RNA by RT-PCR or LAMP methods which may be inhibited by clinical samples. Saliva samples are heterogenous and our experience was that saliva specimens may often be highly viscous making the use of automated sample processing extremely difficult. MHRA TPP guidance for point of care [19] and laboratory based [20] assays recommends the use of an internal control but the OptiGene assay does not currently include this, making it impossible to identify the presence of inhibitory factors in the sample.
The overall diagnostic sensitivity of Direct RT-LAMP in this real-world prospective study was low compared to RT-PCR. Testing of a larger set of randomly collected nose and throat swabs confirmed a similar sensitivity with a similar range of viral loads. The analytical sensitivity data and projection on to the viral loads observed in a large cohort of clinical samples tested throughout the COVID-19 pandemic is also consistent with the low diagnostic sensitivity that was observed. This study highlights the importance of utilising the prospective collection of samples from the intended target population in the assessment of diagnostic sensitivity.
Declarations of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
This work was partially funded by the UK Department of Health and Social Care as part of a national evaluation of Direct RT LAMP for the testing of saliva samples from asymptomatic individuals.
References
- 1.Hoehl S., et al. Evidence of SARS-CoV-2 infection in returning travelers from Wuhan, China. N. Engl. J. Med. 2020;382:1278. doi: 10.1056/NEJMc2001899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rivett L., et al. Screening of healthcare workers for SARS-CoV-2 highlights the role of asymptomatic carriage in COVID-19 transmission. Elife. 2020;9 doi: 10.7554/eLife.58728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Arons M.M., et al. Presymptomatic SARS-CoV-2 infections and transmission in a skilled nursing facility. N. Engl. J. Med. 2020;382:2081–2090. doi: 10.1056/NEJMoa2008457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ellingford J.M., et al. Genomic and healthcare dynamics of nosocomial sars-cov-2 transmission. Elife. 2021;10 doi: 10.7554/eLife.65453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.NICE . COVID-19 Rapid guideline: Haematopoietic Stem Cell Transplantation. 2020. Overview | COVID-19 rapid guideline: haematopoietic stem cell transplantation | Guidance | NICE. [Google Scholar]
- 6.Zhu, J., Guo, J., Xu, Y. & Chen, X. Viral dynamics of SARS-CoV-2 in saliva from infected patients. (2020) doi:10.1016/j.jinf.2020.06.059. [DOI] [PMC free article] [PubMed]
- 7.Iwasaki S., et al. Comparison of SARS-CoV-2 detection in nasopharyngeal swab and saliva. J. Infect. 2020;81:e145. doi: 10.1016/j.jinf.2020.05.071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wyllie A.L., et al. Saliva is more sensitive for SARS-CoV-2 detection in COVID-19 patients than nasopharyngeal swabs. Camila Odio 8. 2020;3:12. [Google Scholar]
- 9.Williams E., Bond K., Zhang B., Putland M., Williamson D.A. Saliva as a Noninvasive Specimen for Detection of SARS-CoV-2. J. Clin. Microbiol. 2020;58 doi: 10.1128/JCM.00776-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kidd S.P., et al. RT-LAMP has high accuracy for detecting SARS-CoV-2 in saliva and naso/oropharyngeal swabs from asymptomatic and symptomatic individuals. medRxiv. 2021 doi: 10.1101/2021.06.28.21259398. 2021.06.28.21259398. [DOI] [Google Scholar]
- 11.OptiGene. OptiGene Limited Instructions For Use COVID-19_RNA RT-LAMP KIT-500. http://www.optigene.co.uk/wp-content/uploads/2021/06/IFU_RNA_v1.4.pdf (21AD).
- 12.Real-time RT-PCR Primers and Probes for COVID-19 | CDC. https://www.cdc.gov/coronavirus/2019-ncov/lab/rt-pcr-panel-primer-probes.html.
- 13.Mautner L., et al. Rapid point-of-care detection of SARS-CoV-2 using reverse transcription loop-mediated isothermal amplification (RT-LAMP) Virol. J. 2020;17:1–14. doi: 10.1186/s12985-020-01435-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dudley D.M., et al. Optimizing direct RT-LAMP to detect transmissible SARS-CoV-2 from primary nasopharyngeal swab samples. PLoS ONE. 2020;15 doi: 10.1371/journal.pone.0244882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Department of Health and Social Care. Rapid evaluation of OptiGene RT-LAMP assay (direct and RNA formats) - GOV.UK. https://www.gov.uk/government/publications/rapid-evaluation-of-optigene-rt-lamp-assay-direct-and-rna-formats/rapid-evaluation-of-optigene-rt-lamp-assay-direct-and-rna-formats (2020).
- 16.Wilson-Davies E.S.W., et al. Concerning the OptiGene Direct LAMP assay, and it's use in at-risk groups and hospital staff. J. Infect. 2021;82:282. doi: 10.1016/j.jinf.2021.01.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Fowler V.L., et al. A highly effective reverse-transcription loop-mediated isothermal amplification (RT-LAMP) assay for the rapid detection of SARS-CoV-2 infection. J. Infect. 2021;82:117–125. doi: 10.1016/j.jinf.2020.10.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Department of Health and Social Care. Technical Validation of OptiGene RT LAMP Assay (Direct and RNA Formats). https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/941765/technical-validation-report-Optigene-Direct-RT-LAMP-swab.pdf (2020).
- 19.Medicines and Healthcare products Regulatory Agency. Target Product Profile: point of Care SARS-CoV-2 detection tests - GOV.UK. https://www.gov.uk/government/publications/how-tests-and-testing-kits-for-coronavirus-covid-19-work/target-product-profile-point-of-care-sars-cov-2-detection-tests (2021).
- 20.Medicines and Healthcare products Regulatory Agency. Target Product Profile: laboratory-Based SARS-CoV-2 Viral Detection tests - GOV.UK. https://www.gov.uk/government/publications/how-tests-and-testing-kits-for-coronavirus-covid-19-work/target-product-profile-laboratory-based-sars-cov-2-viral-detection-tests (2021).