Abstract
The calponin family proteins in vertebrates, including calponin and transgelin (also known as SM22 or NP25), regulate actin-myosin interaction and actin filament stability and are involved in regulation of muscle contractility and cell migration. Related proteins are also present in invertebrates and fungi. Animals have multiple genes encoding calponin family proteins with variable molecular features, which are often expressed in the same tissues or cells. However, functional studies of this class of proteins have been reported only in limited species. Through database searches, I found that the calponin family proteins were diversified in animals by gene amplification and repeat expansion of calponin-like (CLIK) motifs, which function as actin-binding sequences. Transgelin-like proteins with a single CLIK motif are the most primitive type and present in fungi and animals. In many animals, additional calponin family proteins containing multiple CLIK motifs, as represented by vertebrate calponins with three CLIK motifs, are present. Interestingly, in several invertebrate species, there are uncharacterized calponin-related proteins with highly expanded repeats of CLIK motifs (up to 23 repeats in molluscs). These variable molecular features of the calponin family proteins may be results of evolutionary adaptation to a broad range of cell biological events.
Keywords: Actin, calponin, calponin-like motifs, muscle contraction, transgelin
The calponin family proteins regulate actin filament stability and actomyosin contractility
The calponin family proteins contain single or multiple calponin-like (CLIK) motifs that serve as actin-interacting sequences (Fu et al. 2000; Gimona et al. 2003; Goodman et al. 2003; Kranewitter et al. 2001; Mezgueldi et al. 1992) (Fig. 1A). Most of these proteins contain a single calponin-homology (CH) domain in the N-terminus (Fig. 1A), but this region does not bind to actin filaments (Galkin et al. 2006; Gimona and Mital 1998; Gimona and Winder 1998) and is absent from some of the proteins such as Caenorhabditis elegans UNC-87 (Goetinck and Waterston 1994a). Although CH domains are present in a variety of proteins, “actin-binding” CH domains are often present in tandem and have specific structural features that distinguish them from “non-actin-binding” CH domains (Gimona et al. 2002; Gimona and Winder 1998; Stradal et al. 1998; Yin et al. 2020). Here, the “calponin family” proteins are defined as proteins that contain at least one CLIK motif. Although most of the calponin family proteins contain only CH domains and CLIK motifs, some proteins contain other domains, such as an alligator protein (UniProt accession code: A0A151NRQ1) that has five immunoglobulin domains, two fibronectin type III domains, one CH and one CLIK. However, since such proteins are documented only in limited species, this article focuses on proteins consisting with only CH domains and CLIK motifs.
Figure 1.
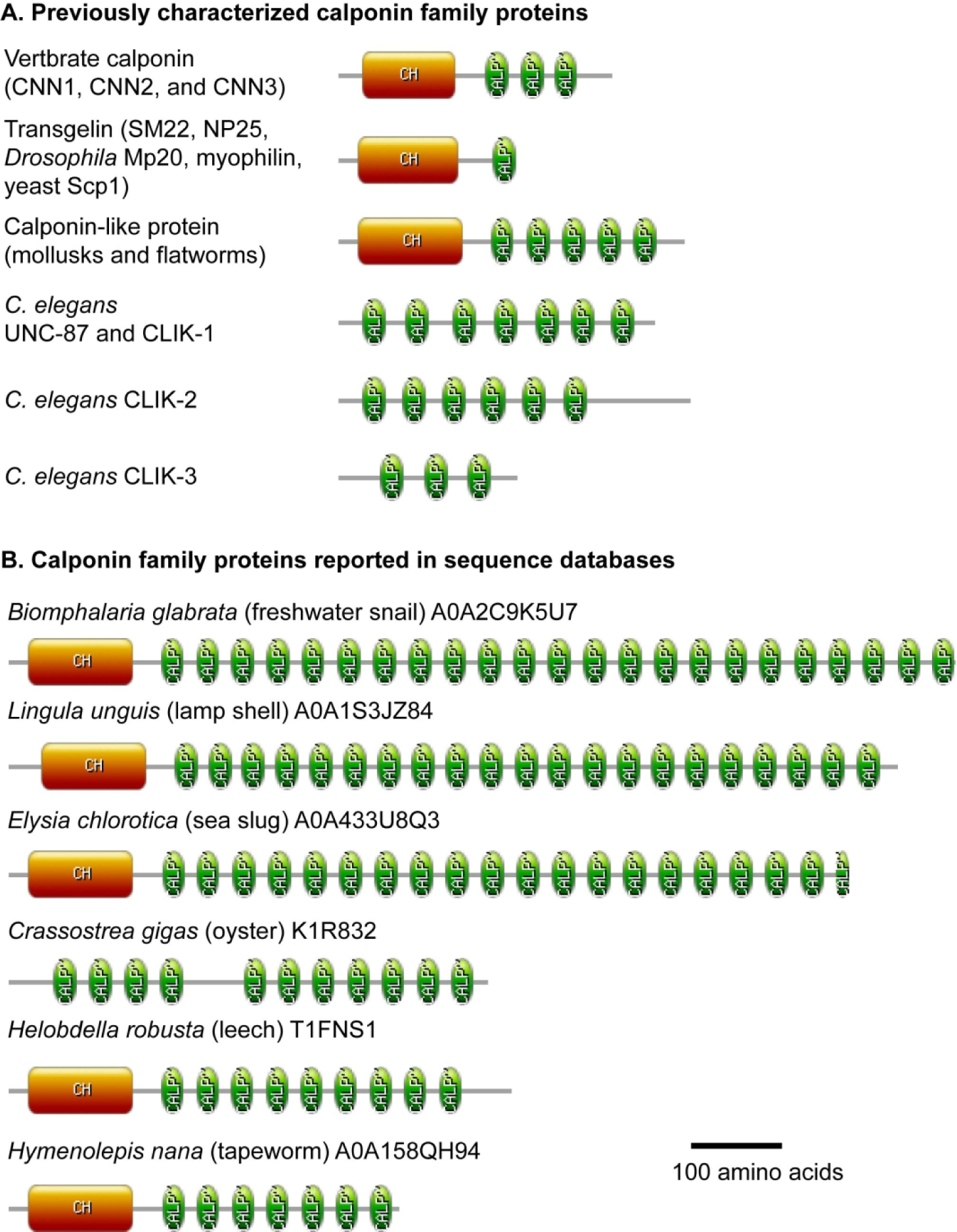
Schematic representation of structures of the calponin family proteins. Domain structures of previously characterized calponin family proteins (A) and related proteins reported only in sequence databases (B). Orange rounded rectangles are calponin homology (CH) domains, and green ellipses (indicated as CALP~) are CLIK motifs. Bar, 100 amino acids. Graphics were obtained from PROSITE (Sigrist et al. 2010) and annotated using Adobe Illustrator CS3.
The calponin family proteins (Fig. 1A) including calponin, transgelin (also known as SM22, NP25, Mp20, or myophilin), and UNC-87, bind to actin filaments and regulate actin filament stability and actomyosin contractility (Assinder et al. 2009; Carmichael et al. 1994; Dvorakova et al. 2014; Liu et al. 2020; Rozenblum and Gimona 2008; Winder and Walsh 1996). Calponin was originally isolated from chicken gizzard smooth muscle as a candidate regulator of contraction with a troponin-like function (Takahashi et al. 1986; Takahashi et al. 1988). In vitro studies demonstrated that calponin inhibits actomyosin ATPase. However, gene knockout of mouse calponin-1 (CNN1), that is predominantly expressed in smooth muscle, causes only modest alterations in smooth muscle contractility (Matthew et al. 2000; Takahashi et al. 2000). The interpretation of the phenotypes is complicated by the presence of multiple calponin isoforms and calponin-related proteins, which may have overlapping roles in muscle contractility. Mammals have three calponin genes (CNN1, CNN2, and CNN3) (Liu and Jin 2016) and three transgelin genes (TAGLN, TAGLN2, and TAGLN3) (Liu et al. 2020). Calponin-2 (CNN2) is also expressed in smooth muscle (Hossain et al. 2003), but either single deletion of CNN2 or double deletion of CNN1 and CNN2 in mice still causes only modest alterations in smooth muscle contractility (Feng et al. 2019). Transgelin-1/SM22α (TAGLN) is also predominantly expressed in smooth muscle (Lees-Miller et al. 1987), and TAGLN knockout mice show only mild defects in smooth muscle contractility (Gao et al. 2020). Whether calponin and transgelin have redundant roles in smooth muscle contraction remains unknown. Furthermore, the calponin family proteins are widely expressed in non-muscle cells, and genetic studies showed regulatory roles of calponin-2 (CNN2) in macrophage motility and phagocytosis (Huang et al. 2008) and essential roles of calponin-3 (CNN3) in brain development (Flemming et al. 2015; Junghans and Herzog 2018). Transgelin-2 (TAGLN2) is enriched in immune cells and regulates T-cell activation (Na et al. 2015) and phagocytosis in macrophages (Kim et al. 2017). Transgelin-3/NP25 (TAGLN3) is predominantly expressed in neurons (Ren et al. 1994), but its function is not understood. Perhaps, multiple calponin family proteins evolved in vertebrates to regulate broad actin-dependent events in both muscle and non-muscle cells. However, functional relationships among the calponin family proteins in vertebrates remain largely unclear.
The calponin family proteins are also present in non-vertebrates. Alignment of the CLIK motif sequences shows conserved sequences (Fig. 2A, Supplemental Fig. S1), suggesting common biochemical properties. However, functional studies of the calponin family proteins in non-vertebrate species are limited. In the nematode Caenorhabditis elegans, UNC-87 is a calponin-related protein with seven CLIK motifs (Fig. 1A, Fig. 2A) that regulates maintenance of sarcomeres in striated muscle (Goetinck and Waterston 1994a; Goetinck and Waterston 1994b; Yamashiro et al. 2007) and contractility of the somatic gonad (Ono et al. 2015). CLIK-1 is similar to UNC-87, and double knockdown of UNC-87 and CLIK-1 in C. elegans causes sterility due to severe cytoskeletal disorganization in the somatic gonad (Ono and Ono 2020), indicating that UNC-87 and CLIK-1 have overlapping roles in maintaining cytoskeletal integrity. In the fruit fly Drosophila melanogaster, Mp20 is a transgelin-like protein (Ayme-Southgate et al. 1989) that is involved in myoblast fusion (Bataille et al. 2010). Transgelin-like proteins are present in fungi [SCP1 in budding yeast (Goodman et al. 2003; Winder et al. 2003) and Stg1 in fission yeast (Nakano et al. 2005)] and involved in actin filament stability (Goodman et al. 2003; Winder et al. 2003) and endocytosis (Gheorghe et al. 2008). Thus, the calponin family proteins are conserved actin regulators, but how broadly these proteins are present among eukaryotes has not been clearly described in literature.
Figure 2.
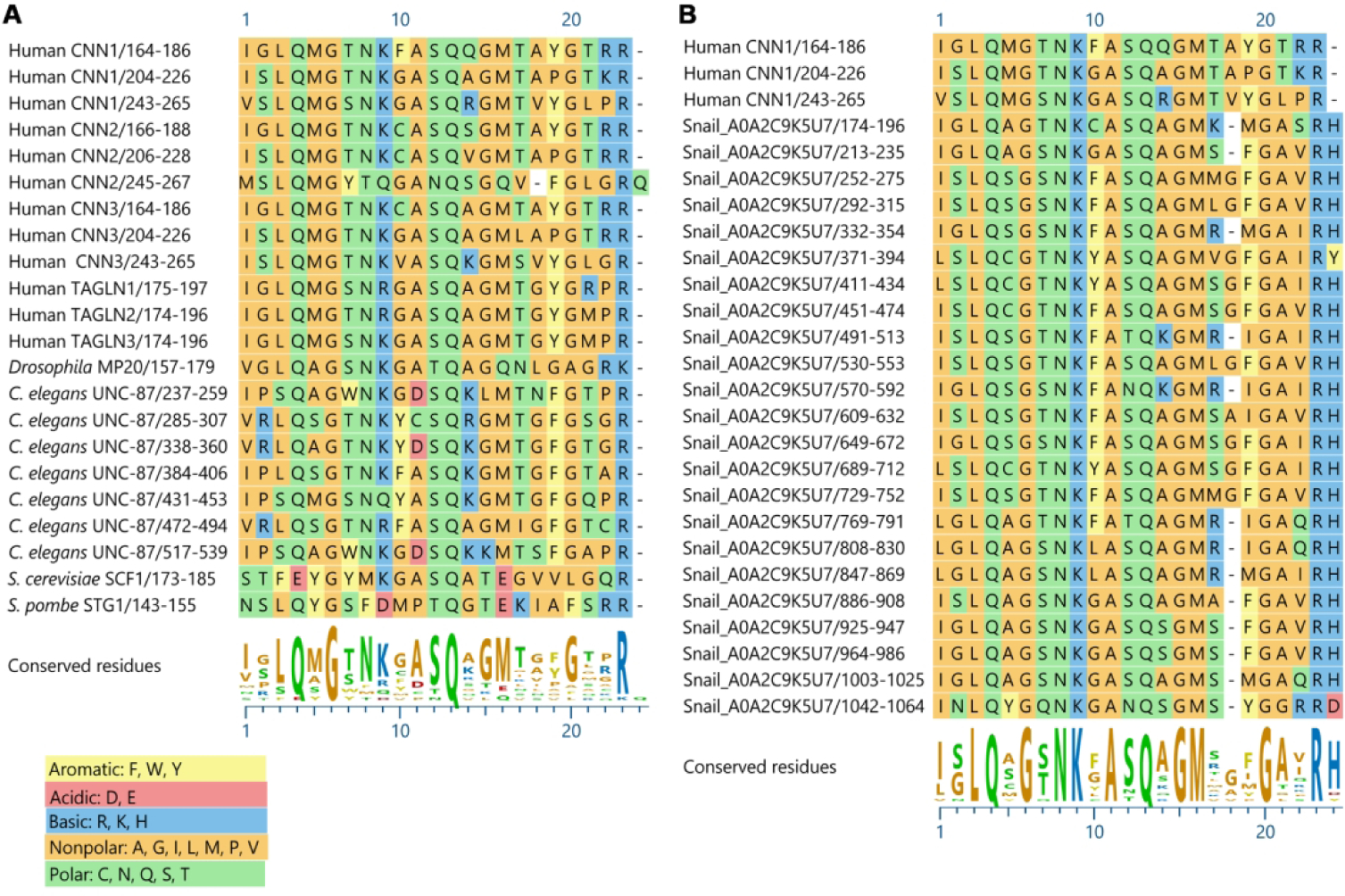
Alignment of CLIK motifs from representative calponin family proteins (A) and 23 CLIK motifs from the freshwater snail Biomphalaria glabrata A0A2C9K5U7, including three CLIK motifs from human calponin-1 (CNN1) for comparison (B). Amino acids are colored based on the chemical properties of side chains as indicated on the figure. The sequences were aligned with MUSCLE (Edgar 2004) using MegAlign Pro (DNASTAR, Inc). See Supplemental Figure S1 for alignment of 196 CLIK motifs.
CLIK motifs are defined as sequences of 23 residues (or up to 26 residues in some databases) (Fig. 2A, Supplemental Fig. S1), which were originally recognized as three repeats in chicken calponin (Takahashi and Nadal-Ginard 1991), and predicted to be intrinsically disordered (Rozenblum and Gimona 2008). The number of CLIK motifs are variable from one (transgelin) to seven (UNC-87) (Fig. 1A), and affinity to actin filaments is correlated with the number of CLIK motifs (Gimona et al. 2003; Kranewitter et al. 2001; Lener et al. 2004). However, a recent characterization of C. elegans CLIK-1 suggests that not all CLIK motifs have equivalent functions. CLIK-1 contains seven CLIK motifs and no CH domain, which resembles the structure of UNC-87 (Ono and Ono 2020) (Fig. 1A). Although UNC-87 binds to actin filaments and bundles them (Kranewitter et al. 2001; Yamashiro et al. 2007), CLIK-1 binds to actin filaments without bundling (Ono and Ono 2020). Both UNC-87 and CLIK-1 contain seven CLIK motifs, but UNC-87 binds to actin filaments with higher affinity than CLIK-1. Sequence comparison indicates that the three C-terminal CLIK motifs of CLIK-1 are less conserved than the rest of the CLIK motifs, suggesting that some of the CLIK motifs have different biochemical properties. Additionally, C. elegans has clik-2 (no CH and 5 CLIK motifs) (Fig. 1A), clik-3 (no CH and 3 CLIK motifs) (Fig. 1A), and two transgelin-like genes (cpn-1 and cpn-2). It should be noted that cpn-3 and cpn-4 have only one CH domain with no CLIK motif. clik-2 and clik-3 are involved in the regulation of pharyngeal muscle contraction (Wang et al. 2018), but functions of the four transgelin-like genes are unknown.
Diversification of calponin-related proteins: gene amplification and repeat expansion
To understand the presence and diversity of the calponin family proteins among eukaryotes, I searched databases for proteins that contain CLIK motifs and found an interesting pattern of molecular evolution of this class of proteins. The CLIK motif is designated as the “calponin family repeat” in databases [PF00402 in Pfam (Mistry et al. 2021); PDOC00808 in PROSITE (Sigrist et al. 2010)]. In the SMART database (Letunic et al. 2021), CLIK motifs are recognized only in fungi and metazoa. In other words, plants, algae, protozoa, and slime molds do not have a protein with a CLIK motif. Most of fungal species have a single copy of a transgelin-like gene, such as Scp1 in Saccharomyces cerevisiae (Goodman et al. 2003; Winder et al. 2003) and Stg1 in Schizosaccharomyces pombe (Nakano et al. 2005), but some species have two copies. The transgelin-like genes are also present throughout the metazoan species, suggesting that a transgelin-like protein is the most primitive form of the calponin family protein.
In addition to transgelin-like proteins, most of metazoan species have calponin family proteins with multiple CLIK motifs, as represented by vertebrate calponins (3 CLIK motifs) (Takahashi and Nadal-Ginard 1991), calponin-like proteins in flatworms and molluscs (5 CLIK motifs) (Matusovsky et al. 2017; Wang et al. 2016; Yang et al. 1999), and UNC-87 in C. elegans (7 CLIK motifs) (Goetinck and Waterston 1994a) (Fig. 1A). Interestingly, invertebrate species in several phyla have calponin-related proteins with highly expanded repeats of CLIK motifs (Fig. 1B). For example, calponin-related proteins in the freshwater snail Biomphalaria glabrata (UniProt accession number: A0A2C9K5U7), the brachiopod Lingula unguis (UniProt accession number: A0A1S3JZ84), and the sea slug Elysia chlorotica (UniProt accession number: A0A433U8Q3), contain 23, 21, and 20 CLIK motifs, respectively (Fig. 1B). The 23 CLIK motifs in the Biomphalaria snail protein are highly conserved and repetitive (Fig. 2B), suggesting conserved biochemical properties for actin-binding. Calponin-related proteins containing more than five CLIK motifs are found in the phyla Nematoda (up to 7 CLIK motifs), Mollusca (up to 23 CLIK motifs), Annelida (up to 9 CLIK motifs), Brachiopoda (up to 21 CLIK motifs), and Platyhelminthes (up to 7 CLIK motifs) (Table 1, Supplemental Table S1). Although most of these calponin-related proteins with expanded repeats of CLIK motifs are only reported in the databases and have not been functionally characterized, it is very likely that these proteins bind to actin and function as cytoskeletal regulators in a similar manner to UNC-87 and CLIK-1 in the nematode C. elegans, as suggested by highly conserved sequences of the CLIK motifs in these invertebrates (Fig. 2B, Supplemental Fig. S1). Intriguingly, these are soft-bodied invertebrates that are capable of highly flexible deformation of the body shape. It will be interesting if there is any functional relationship between the number of CLIK motifs and flexibility of the body shape.
Table 1.
Summary of the calponin family proteins in representative species.
| Kingdom Phylum | Species (common name) | Number of genesa | Maximum CLIK repeatsb | |
|---|---|---|---|---|
|
| ||||
| Animalia | Chordata | Homo sapiens (human) | 6 | 3 |
| Echinodermata | Strongylocentrotus purpuratus (sea urchin) 4 | 3 | ||
| Mollusca | Biomphalaria glabrata (snail) | 5 | 23 | |
| Brachiopoda | Lingula anatina (lamp shell) | 7 | 21 | |
| Annelida | Helobdella robusta (leech) | 7 | 9 | |
| Platyhelminthes | Hymenolepis nana (tapeworm) | 6 | 7 | |
| Arthropoda | Drosophila melanogaster (fruit fly) | 3 | 1 | |
| Nematoda | Caenorhabditis elegans (roundworm) | 6 | 7 | |
| Cnidaria | Nematostella vectensis (sea anemone) | 4 | 1 | |
| Porifera | Amphimedon queenslandica (sponge) | 3 | 1 | |
| Fungi | Ascomycota | Saccharomyces cerevisiae (yeast) | 1 | 1 |
Data are derived from WormBase (Harris et al. 2020) for C. elegans, WormBase ParaSite (Howe et al. 2017) for H. nana, MycoCosm (Grigoriev et al. 2014) for S. cerevisiae, and Ensembl (Howe et al. 2021) for all other species. See Supplemental Table S1 for accession numbers.
Data are derived from PROSITE (Sigrist et al. 2010).
By contrast, species in many phyla have only transgelin-like proteins. Notably, the Arthropoda is one of those phyla (Table 1, Supplemental Table S1), despite the fact that arthropods have relatively complex body plans. Therefore, there is no apparent correlation between the complexity of the body plan and the number of CLIK motifs. For example, the genome of the fruit fly Drosophila melanogaster has three genes encoding transgelin-like proteins (Mp20, CG5023, and ChD64) but no genes encoding a protein with multiple CLIK motifs. Mp20 was identified as a muscle-specific protein (Ayme-Southgate et al. 1989), but ChD64 was identified as a nuclear protein that binds to a juvenile hormone-response element (Li et al. 2007). Function of CG5023 has not been characterized. Whether the three transgelin-like proteins have different functions remains unknown, but it is clear that Drosophila and other arthropods do not require a calponin-related protein with multiple CLIK motifs.
The number of genes encoding calponin family proteins (Table 1, Supplemental Table S1) suggests that this class of genes was amplified during animal evolution. However, I did not find strong correlation between the complexity of the body plans and the number of genes encoding calponin family proteins. Many invertebrates have more calponin-family genes than vertebrates. However, the composition of calponin-family genes in Drosophila is as simple as that in sponges (Table 1, Supplemental Table S1). As reported for vertebrate calponin family proteins, these genes are most likely expressed differently in various cell types and at developmental stages. Therefore, further mechanistic analyses will be required to uncover functional differences and/or redundancy among the calponin family proteins.
Perspectives
Variable numbers of CLIK motifs may be linked to the function of the calponin family proteins to stabilize actin filaments. A comparative study on the calponin family proteins with different number of CLIK motifs indicated that the number of CLIK motifs is correlated with their capability to stabilize actin filaments (Gimona et al. 2003; Lener et al. 2004). In cultured cells, transgelin-1/SM22α is associated with actin filaments and still allows dynamic cytoskeletal remodeling, whereas calponin is associated with actin filaments and inhibits cytoskeletal remodeling (Gimona et al. 2003). Although transgelins are implicated in actin filament stabilization (Gheorghe et al. 2008; Goodman et al. 2003; Na et al. 2015; Shapland et al. 1993), transgelin-associated actin filaments may still undergo turnover to support dynamic actin-dependent events such as cell migration (Liu et al. 2020). By contrast, vertebrate calponin (Lehman 1991) and C. elegans UNC-87 (Goetinck and Waterston 1994a) are components of muscle thin filaments, which require high stability during repeated muscle contraction and relaxation. The uncharacterized calponin family proteins with expanded repeats of CLIK motifs (Fig. 1B) may have even stronger properties to stabilize actin filaments.
Repeat expansion of actin-binding motifs is also present in other actin-binding proteins, including immunoglobulin domains in titin/connectin, kettin, and filamin (Nakamura et al. 2007; Otey et al. 2009), nebulin-like repeats (Bjorklund et al. 2010; Pappas et al. 2011), and Xin repeats (Pacholsky et al. 2004). There are several evolutionary advantages of having repeated actin-binding motifs. First, repeating actin-binding motifs can increase affinity with actin as described above. In the case of nebulin, the size of nebulin is important for actin stabilization and regulation of thin filament lengths (Castillo et al. 2009; Pappas et al. 2010). Second, repeating actin-binding motifs can make large molecules that can interact with many actin subunits without forming an oligomer. Structural studies suggest that CLIK motifs are disordered (Kozlowska et al. 2014; Li et al. 2008; Tarczewska et al. 2015). Therefore, repeated CLIK motifs should be flexible and extended to reach out to multiple actin subunits within a filament and/or multiple actin filaments to bundle them. Third, some of the actin-binding motifs can be functionally differentiated within a single molecule. Although both UNC-87 and CLIK-1 from C. elegans have seven CLIK motifs, they are different in actin affinity and actin bundling activity (Ono and Ono 2020), suggesting that not all CLIK motifs have the same properties. This may be important for differentiating functions of the calponin family proteins. Therefore, the variable molecular features may be results of evolutionary adaptation to different requirements in actin regulation.
In conclusion, the calponin family proteins evolved by gene amplification and repeat expansion of CLIK motifs. The database searches revealed previously unappreciated variety of this class of proteins, suggesting that some of them are adapted to regulate specific actin-dependent processes. Also, some of those may be involved in actin-independent processes such as signal transduction as demonstrated for mammalian calponin (Leinweber et al. 2000) and Drosophila ChD64 (Li et al. 2007). Further functional studies of the calponin family proteins should reveal broad biological significance of this class of proteins.
Supplementary Material
Acknowledgements
This work was supported by a grant from National Institutes of Health (R01 AR048615) to S. O. The author declares no conflict of interest.
Data availability statement
No sharable data were generated in this work.
References
- Assinder SJ, Stanton JA, Prasad PD. 2009. Transgelin: an actin-binding protein and tumour suppressor. Int J Biochem Cell Biol 41:482–486. [DOI] [PubMed] [Google Scholar]
- Ayme-Southgate A, Lasko P, French C, Pardue ML. 1989. Characterization of the gene for mp20: a Drosophila muscle protein that is not found in asynchronous oscillatory flight muscle. J Cell Biol 108:521–531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bataille L, Delon I, Da Ponte JP, Brown NH, Jagla K. 2010. Downstream of identity genes: muscle-type-specific regulation of the fusion process. Dev Cell 19:317–328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bjorklund AK, Light S, Sagit R, Elofsson A. 2010. Nebulin: a study of protein repeat evolution. J Mol Biol 402:38–51. [DOI] [PubMed] [Google Scholar]
- Carmichael JD, Winder SJ, Walsh MP, Kargacin GJ. 1994. Calponin and smooth muscle regulation. Can J Physiol Pharmacol 72:1415–1419. [DOI] [PubMed] [Google Scholar]
- Castillo A, Nowak R, Littlefield KP, Fowler VM, Littlefield RS. 2009. A nebulin ruler does not dictate thin filament lengths. Biophys J 96:1856–1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dvorakova M, Nenutil R, Bouchal P. 2014. Transgelins, cytoskeletal proteins implicated in different aspects of cancer development. Expert Rev Proteomics 11:149–165. [DOI] [PubMed] [Google Scholar]
- Edgar RC. 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng HZ, Wang H, Takahashi K, Jin JP. 2019. Double deletion of calponin 1 and calponin 2 in mice decreases systemic blood pressure with blunted length-tension response of aortic smooth muscle. J Mol Cell Cardiol 129:49–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flemming A, Huang QQ, Jin JP, Jumaa H, Herzog S. 2015. A conditional knockout mouse model reveals that calponin-3 is dispensable for early B cell development. PLoS One 10:e0128385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu Y, Liu HW, Forsythe SM, Kogut P, McConville JF, Halayko AJ, Camoretti-Mercado B, Solway J. 2000. Mutagenesis analysis of human SM22: characterization of actin binding. J Appl Physiol 89:1985–1990. [DOI] [PubMed] [Google Scholar]
- Galkin VE, Orlova A, Fattoum A, Walsh MP, Egelman EH. 2006. The CH-domain of calponin does not determine the modes of calponin binding to F-actin. J Mol Biol 359:478–485. [DOI] [PubMed] [Google Scholar]
- Gao Y, Liu G, Kong P, Song Y, Zhang D, Yin Y, Han M. 2020. Smooth muscle 22alpha deficiency impairs oxytocin-induced uterine contractility in mice at full-term pregnancy. Biochem Biophys Res Commun 529:884–889. [DOI] [PubMed] [Google Scholar]
- Gheorghe DM, Aghamohammadzadeh S, Smaczynska-de R II, Allwood EG, Winder SJ, Ayscough KR. 2008. Interactions between the yeast SM22 homologue Scp1 and actin demonstrate the importance of actin bundling in endocytosis. J Biol Chem 283:15037–15046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gimona M, Djinovic-Carugo K, Kranewitter WJ, Winder SJ. 2002. Functional plasticity of CH domains. FEBS Lett 513:98–106. [DOI] [PubMed] [Google Scholar]
- Gimona M, Kaverina I, Resch GP, Vignal E, Burgstaller G. 2003. Calponin repeats regulate actin filament stability and formation of podosomes in smooth muscle cells. Mol Biol Cell 14:2482–2491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gimona M, Mital R. 1998. The single CH domain of calponin is neither sufficient nor necessary for F-actin binding. J Cell Sci 111:1813–1821. [DOI] [PubMed] [Google Scholar]
- Gimona M, Winder SJ. 1998. Single calponin homology domains are not actin-binding domains. Curr Biol 8:R674–675. [DOI] [PubMed] [Google Scholar]
- Goetinck S, Waterston RH. 1994a. The Caenorhabditis elegans muscle-affecting gene unc-87 encodes a novel thin filament-associated protein. J Cell Biol 127:79–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goetinck S, Waterston RH. 1994b. The Caenorhabditis elegans UNC-87 protein is essential for maintenance, but not assembly, of bodywall muscle. J Cell Biol 127:71–78. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodman A, Goode BL, Matsudaira P, Fink GR. 2003. The Saccharomyces cerevisiae calponin/transgelin homolog Scp1 functions with fimbrin to regulate stability and organization of the actin cytoskeleton. Mol Biol Cell 14:2617–2629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grigoriev IV, Nikitin R, Haridas S, Kuo A, Ohm R, Otillar R, Riley R, Salamov A, Zhao X, Korzeniewski F and others. 2014. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res 42:D699–704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris TW, Arnaboldi V, Cain S, Chan J, Chen WJ, Cho J, Davis P, Gao S, Grove CA, Kishore R and others. 2020. WormBase: a modern Model Organism Information Resource. Nucleic Acids Res 48:D762–D767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hossain MM, Hwang DY, Huang QQ, Sasaki Y, Jin JP. 2003. Developmentally regulated expression of calponin isoforms and the effect of h2-calponin on cell proliferation. Am J Physiol Cell Physiol 284:C156–167. [DOI] [PubMed] [Google Scholar]
- Howe KL, Achuthan P, Allen J, Allen J, Alvarez-Jarreta J, Amode MR, Armean IM, Azov AG, Bennett R, Bhai J and others. 2021. Ensembl 2021. Nucleic Acids Res 49:D884–D891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howe KL, Bolt BJ, Shafie M, Kersey P, Berriman M. 2017. WormBase ParaSite - a comprehensive resource for helminth genomics. Mol Biochem Parasitol 215:2–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang QQ, Hossain MM, Wu K, Parai K, Pope RM, Jin JP. 2008. Role of H2-calponin in regulating macrophage motility and phagocytosis. J Biol Chem 283:25887–25899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Junghans D, Herzog S. 2018. Cnn3 regulates neural tube morphogenesis and neuronal stem cell properties. FEBS J 285:325–338. [DOI] [PubMed] [Google Scholar]
- Kim HR, Lee HS, Lee KS, Jung ID, Kwon MS, Kim CH, Kim SM, Yoon MH, Park YM, Lee SM and others. 2017. An essential role for TAGLN2 in phagocytosis of lipopolysaccharide-activated macrophages. Sci Rep 7:8731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozlowska M, Tarczewska A, Jakob M, Szpotkowski K, Wojtas M, Rymarczyk G, Ozyhar A. 2014. Calponin-like Chd64 is partly disordered. PLoS One 9:e96809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kranewitter WJ, Ylanne J, Gimona M. 2001. UNC-87 is an actin-bundling protein. J Biol Chem 276:6306–6312. [DOI] [PubMed] [Google Scholar]
- Lees-Miller JP, Heeley DH, Smillie LB, Kay CM. 1987. Isolation and characterization of an abundant and novel 22-kDa protein (SM22) from chicken gizzard smooth muscle. J Biol Chem 262:2988–2993. [PubMed] [Google Scholar]
- Lehman W 1991. Calponin and the composition of smooth muscle thin filaments. J Muscle Res Cell Motil 12:221–224. [DOI] [PubMed] [Google Scholar]
- Leinweber B, Parissenti AM, Gallant C, Gangopadhyay SS, Kirwan-Rhude A, Leavis PC, Morgan KG. 2000. Regulation of protein kinase C by the cytoskeletal protein calponin. J Biol Chem 275:40329–40336. [DOI] [PubMed] [Google Scholar]
- Lener T, Burgstaller G, Gimona M. 2004. The role of calponin in the gene profile of metastatic cells: inhibition of metastatic cell motility by multiple calponin repeats. FEBS Lett 556:221–226. [DOI] [PubMed] [Google Scholar]
- Letunic I, Khedkar S, Bork P. 2021. SMART: recent updates, new developments and status in 2020. Nucleic Acids Res 49:D458–D460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li M, Li S, Lou Z, Liao X, Zhao X, Meng Z, Bartlam M, Rao Z. 2008. Crystal structure of human transgelin. J Struct Biol 162:229–236. [DOI] [PubMed] [Google Scholar]
- Li Y, Zhang Z, Robinson GE, Palli SR. 2007. Identification and characterization of a juvenile hormone response element and its binding proteins. J Biol Chem 282:37605–37617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J, Zhang Y, Li Q, Wang Y. 2020. Transgelins: cytoskeletal associated proteins implicated in the metastasis of colorectal cancer. Front Cell Dev Biol 8:573859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu R, Jin JP. 2016. Calponin isoforms CNN1, CNN2 and CNN3: Regulators for actin cytoskeleton functions in smooth muscle and non-muscle cells. Gene 585:143–153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matthew JD, Khromov AS, McDuffie MJ, Somlyo AV, Somlyo AP, Taniguchi S, Takahashi K. 2000. Contractile properties and proteins of smooth muscles of a calponin knockout mouse. J Physiol 529 Pt 3:811–824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matusovsky OS, Dobrzhanskaya AV, Pankova VV, Kiselev KV, Girich UV, Shelud’ko NS. 2017. Crenomytilus grayanus 40kDa calponin-like protein: cDNA cloning, sequence analysis, tissue expression, and post-translational modifications. Comp Biochem Physiol Part D Genomics Proteomics 22:98–108. [DOI] [PubMed] [Google Scholar]
- Mezgueldi M, Fattoum A, Derancourt J, Kassab R. 1992. Mapping of the functional domains in the amino-terminal region of calponin. J Biol Chem 267:15943–15951. [PubMed] [Google Scholar]
- Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, Tosatto SCE, Paladin L, Raj S, Richardson LJ and others. 2021. Pfam: The protein families database in 2021. Nucleic Acids Res 49:D412–D419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Na BR, Kim HR, Piragyte I, Oh HM, Kwon MS, Akber U, Lee HS, Park DS, Song WK, Park ZY and others. 2015. TAGLN2 regulates T cell activation by stabilizing the actin cytoskeleton at the immunological synapse. J Cell Biol 209:143–162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura F, Osborn TM, Hartemink CA, Hartwig JH, Stossel TP. 2007. Structural basis of filamin A functions. J Cell Biol 179:1011–1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakano K, Bunai F, Numata O. 2005. Stg 1 is a novel SM22/transgelin-like actin-modulating protein in fission yeast. FEBS Lett 579:6311–6316. [DOI] [PubMed] [Google Scholar]
- Ono K, Obinata T, Yamashiro S, Liu Z, Ono S. 2015. UNC-87 isoforms, Caenorhabditis elegans calponin-related proteins, interact with both actin and myosin and regulate actomyosin contractility. Molecular Biology of the Cell 26:1687–1698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ono S, Ono K. 2020. Two Caenorhabditis elegans calponin-related proteins have overlapping functions that maintain cytoskeletal integrity and are essential for reproduction. J Biol Chem 295:12014–12027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otey CA, Dixon R, Stack C, Goicoechea SM. 2009. Cytoplasmic Ig-domain proteins: cytoskeletal regulators with a role in human disease. Cell Motil Cytoskeleton 66:618–634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pacholsky D, Vakeel P, Himmel M, Lowe T, Stradal T, Rottner K, Furst DO, van der Ven PF. 2004. Xin repeats define a novel actin-binding motif. J Cell Sci 117:5257–5268. [DOI] [PubMed] [Google Scholar]
- Pappas CT, Bliss KT, Zieseniss A, Gregorio CC. 2011. The Nebulin family: an actin support group. Trends Cell Biol 21:29–37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pappas CT, Krieg PA, Gregorio CC. 2010. Nebulin regulates actin filament lengths by a stabilization mechanism. J Cell Biol 189:859–870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren WZ, Ng GY, Wang RX, Wu PH, O’Dowd BF, Osmond DH, George SR, Liew CC. 1994. The identification of NP25: a novel protein that is differentially expressed by neuronal subpopulations. Brain Res Mol Brain Res 22:173–185. [DOI] [PubMed] [Google Scholar]
- Rozenblum GT, Gimona M. 2008. Calponins: adaptable modular regulators of the actin cytoskeleton. Int J Biochem Cell Biol 40:1990–1995. [DOI] [PubMed] [Google Scholar]
- Shapland C, Hsuan JJ, Totty NF, Lawson D. 1993. Purification and properties of transgelin: a transformation and shape change sensitive actin-gelling protein. J Cell Biol 121:1065–1073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sigrist CJ, Cerutti L, de Castro E, Langendijk-Genevaux PS, Bulliard V, Bairoch A, Hulo N. 2010. PROSITE, a protein domain database for functional characterization and annotation. Nucleic Acids Res 38:D161–166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stradal T, Kranewitter W, Winder SJ, Gimona M. 1998. CH domains revisited. FEBS Lett 431:134–137. [DOI] [PubMed] [Google Scholar]
- Takahashi K, Hiwada K, Kokubu T. 1986. Isolation and characterization of a 34,000-dalton calmodulin- and F-actin-binding protein from chicken gizzard smooth muscle. Biochem Biophys Res Commun 141:20–26. [DOI] [PubMed] [Google Scholar]
- Takahashi K, Hiwada K, Kokubu T. 1988. Vascular smooth muscle calponin. A novel troponin T-like protein. Hypertension 11:620–626. [DOI] [PubMed] [Google Scholar]
- Takahashi K, Nadal-Ginard B. 1991. Molecular cloning and sequence analysis of smooth muscle calponin. J Biol Chem 266:13284–13288. [PubMed] [Google Scholar]
- Takahashi K, Yoshimoto R, Fuchibe K, Fujishige A, Mitsui-Saito M, Hori M, Ozaki H, Yamamura H, Awata N, Taniguchi S and others. 2000. Regulation of shortening velocity by calponin in intact contracting smooth muscles. Biochem Biophys Res Commun 279:150–157. [DOI] [PubMed] [Google Scholar]
- Tarczewska A, Kozlowska M, Dobryszycki P, Kaus-Drobek M, Dadlez M, Ozyhar A. 2015. Insight into the unfolding properties of Chd64, a small, single domain protein with a globular core and disordered tails. PLoS One 10:e0137074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H, Park H, Liu J, Sternberg PW. 2018. An efficient genome editing strategy to generate putative null mutants in Caenorhabditis elegans using CRISPR/Cas9. G3 (Bethesda) 8:3607–3616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J, Gao J, Xie J, Zheng X, Yan Y, Li S, Xie L, Zhang R. 2016. Cloning and mineralization-related functions of the calponin gene in Chlamys farreri. Comp Biochem Physiol B Biochem Mol Biol 201:53–58. [DOI] [PubMed] [Google Scholar]
- Winder SJ, Jess T, Ayscough KR. 2003. SCP1 encodes an actin-bundling protein in yeast. Biochem J 375:287–295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winder SJ, Walsh MP. 1996. Calponin. Curr Top Cell Regul 34:33–61. [DOI] [PubMed] [Google Scholar]
- Yamashiro S, Gimona M, Ono S. 2007. UNC-87, a calponin-related protein in C. elegans, antagonizes ADF/cofilin-mediated actin filament dynamics. J Cell Sci 120:3022–3033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang W, Zheng YZ, Jones MK, McManus DP. 1999. Molecular characterization of a calponin-like protein from Schistosoma japonicum. Mol Biochem Parasitol 98:225–237. [DOI] [PubMed] [Google Scholar]
- Yin LM, Schnoor M, Jun CD. 2020. Structural characteristics, binding partners and related diseases of the calponin homology (CH) domain. Front Cell Dev Biol 8:342. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
No sharable data were generated in this work.


