Abstract

The present article comprehensively examines six N′-(adamantan-2-ylidene)hydrazide derivatives using the Hirshfeld surface analysis, PIXEL energy for molecular dimers, lattice energies for crystal packing, and topological analysis for intramolecular and intermolecular interactions. The crystal structure of one of the N′-(adamantan-2-ylidene)hydrazide derivatives, namely, N′-(adamantan-2-ylidene)-5-bromothiophene-2-carbohydrazide 1, C15H17N2OSBr, has been determined and analyzed in detail along with five closely related structures. The molecular conformation of 1 is locked by an intramolecular C–S···N chalcogen bond as found in one of its closely related structure, namely, N′-(adamantan-2-ylidene)thiophene-2-carbohydrazide. Furthermore, a detailed potential energy surface scan analysis has been performed to highlight the importance of a chalcogen bond. Two of these compounds possess syn-orientation for amide units, whereas the corresponding moiety exhibits anti-conformations in the remaining four structures. The Hirshfeld surface and its decomposed fingerprint plots provide a qualitative picture of acyl substituent effects on the intermolecular interactions toward crystal packing of these six structures. Intermolecular interaction energies for dimers observed in these structures calculated by density functional theory (B97D3/def2-TZVP) and PIXEL (MP2/6-31G**) methods are comparable. This study also identifies that multiple hydrogen bonds, including N/C–H···O/N and C–H···π interactions, are collectively responsible for a self-assembled synthon. The nature and strength of these interactions have been studied using atoms in molecule topological analysis. The in vitro antiproliferative activity of compound 1 was assessed against five human tumor cell lines and showed marked antiproliferative activity.
1. Introduction
The adamantane cage is an essential building motif in several drugs.1−4 The chemotherapeutic potency of adamantane derivatives was initially discovered after developing amantadine5,6 and rimantadine7 as effective therapies against influenza A viral infections and tromantadine as an antiviral drug for the treatment of skin infection caused by herpes simplex virus.8 Antitumor activity was reported for some adamantane derivatives. Adaphostin,9,10 the synthetic retinoid CD437,11,12 adarotene (ST1926),13,14 and opaganib (ABC294640)15,16 are currently used as efficient therapies against different resistant cancers.
Continuing ongoing research efforts in the chemotherapeutic17−21 and structural characterization22−26 of adamantane-based derivatives, we describe herein the synthesis, structural characteristics, and antiproliferative activity of N′-(adamantan-2-ylidene)-5-bromothiophene-2-carbohydrazide derivative 1. The Cambridge Structural Database (CSD)27 search reveals five closely related structures for structure 1, and its closely related structures are compound 2 (csd refcode: YAJBES),28 compound 3 (csd refcode: ILESAV),29 compound 4 (csd refcode: OKOLOR),30 compound 5 (csd refocde: XEBTIJ),31 and compound 6 (csd refcode: NICCEH)32 shown in Figure 1. These related structures were used for comparative analysis to study the effect of the acyl substituents (R) on the crystal packing and to gain more insights into the nature and strength of different intermolecular interactions formed in these structures.
Figure 1.
Chemical structures of N′-(adamantan-2-ylidene)hydrazide derivatives.
Different types of hydrogen bonds, including N–H···O and N–H···N interactions, are found in biological macromolecules such as proteins and nucleic acids. The former hydrogen bond plays an important role in stabilizing various secondary structural elements such as helices and β-sheets in polypeptides, whereas both types of hydrogen bonds help stabilize base pair recognition in nucleic acids.33 In addition, the C–H···O interaction also participates in the stabilization of proteins and nucleic acids.34−36 Crystallography and computational tools have been used to describe supramolecular assembly governed by charge-assisted hydrogen bonds in complexes of amino acid–carboxylic acid model systems.37 Recently, authors have analyzed new types of supramolecular structures driven by salt bridge interactions and charge-assisted aromatic ring systems.38,39
In addition to the above interactions, the other types of intermolecular interactions, namely, C–H···O,40 C–H···S,41 C–H···Br,42 C–H···N,43 and C–H···C(π)44,45 interactions play vital roles in the self-assembly of organic molecules in the crystalline state and crystal engineering. The abovementioned interactions are formed in the N′-(adamantan-2-ylidene)hydrazide derivatives described in this work. Different theoretical tools, including Hirshfeld surface (HS) analysis, 2D-fingerprint (2D-FP) plots, PIXEL energy, and Bader’s atoms in molecular approach,46 were used to better understand the contribution of various intermolecular interactions to the crystal packing of these compounds. The role of nitrogen substituents on structural and packing features and vibrational behavior has been studied using adamantane–thiourea hybrid derivatives.47,48 There is a growing interest in the highly directional noncovalent interactions, that is, σ–hole and π–hole interactions. These interactions can be exploited in synthesis, catalysis, supramolecular chemistry, molecular recognition, and crystal engineering.49−52 The existence of an intramolecular S···N chalcogen bond in the N-acylhydrazone derivative and its effect on the molecular conformation has been studied utilizing molecular modeling and spectroscopic techniques.53 The present study also allowed us to investigate the role of an intramolecular chalcogen bond (C–S···N),54 and conformational preference of amide units exhibited in these compounds.
2. Results and Discussion
Compound 1 in this study can be obtained by combining 5-bromothiophene-2-carbohydrazide with 2-adamantanone. Furthermore, characterization of this compound was conducted by NMR (1H and 13C) spectroscopy (Figures S1 and S2, Supporting Information). A colorless block-shaped single crystal was obtained from ethanol solvent, and the crystal structure was determined. We also used five closely related structures of 1 to investigate the role of various substituents, namely, thiophene, 3-pyridyl, 4-pyridyl, phenyl, and methyl moieties in N′-(adamantan-2-ylidene)hydrazides on the molecular conformation, crystal packing, and intermolecular interactions. The intermolecular interactions within them have been studied using various theoretical tools. The N′-(adamantan-2-ylidene)hydrazides derivatives possess an adamantyl cage, hydrazine in the middle, and variable substituents as mentioned above. We also describe the antiproliferative properties of compound 1 against five different cancer cell lines in vitro.
2.1. General Molecular and Crystal Structural Description
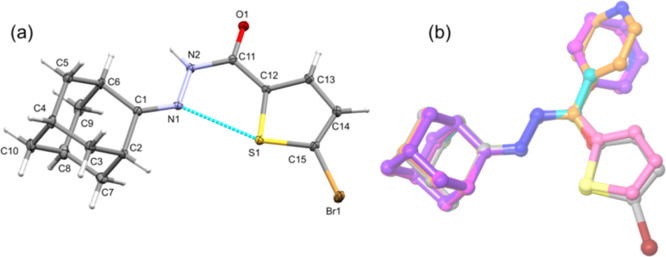
Compound 1 crystallizes in the monoclinic space group P21/c with a single molecule in the asymmetric unit. The crystal data and refinement parameters for 1 are summarized in Table 1 and the thermal ellipsoid representation of compound 1 is shown in Figure 2a. In 1, the central N=N–C unit and the thiophene ring are nearly coplanar, with the dihedral angle formed between them being 6.17°. However, the adamantane cage is twisted by 43.98° with respect to the mean plane of the thiophene ring. The six-membered rings constituting the adamantyl moiety have a typical chair conformation as previously observed. The bond lengths of C1=N1 [1.288 (1) Å], N1–N2 [1.383 (1) Å], and C11=O1 [1.243 (1) Å] bonds are comparable with those of related structures.28−32
Table 1. Crystal Data and Refinement Parameters for Compound 1.
| empirical formula | C15H17N2OSBr |
| formula weight | 353.27 |
| crystal system | monoclinic |
| space group | P21/c |
| a/Å | 12.0105(6) |
| b/Å | 9.8832(4) |
| c/Å | 12.8066(5) |
| α/deg | 90 |
| β/deg | 103.2030(10) |
| γ/deg | 90 |
| volume/Å3 | 1479.99(11) |
| Z | 4 |
| ρcalc g/cm3 | 1.585 |
| μ/mm–1 | 2.915 |
| F(000) | 720.0 |
| crystal size/mm3 | 0.47 × 0.21 × 0.18 |
| radiation | Mo Kα (λ = 0.71073) |
| 2Θ range for data collection/deg | 5.396 to 66.408 |
| index ranges | –18 ≤ h ≤ 16 |
| –15 ≤ k ≤ 14 | |
| –19 ≤ l ≤ 19 | |
| reflections collected | 20 031 |
| independent reflections | 5646 [Rint = 0.0263, Rσ = 0.0267] |
| data/restraints/parameters | 5646/0/186 |
| goodness-of-fit on F2 | 1.054 |
| final R indexes [I ≥ 2σ(I)] | R1 = 0.0251, wR2 = 0.0613 |
| final R indexes [all data] | R1 = 0.0292, wR2 = 0.0629 |
| largest diff. peak/hole/e Å–3 | 0.62/–0.82 |
| CCDC no. | 2123456 |
Figure 2.
(a) Thermal ellipsoids of compound 1 are drawn at the 50% probability level and an intramolecular C–S···N chalcogen bond is shown and (b) structural superimposition of 1 with its five closely related structures [color codes: 1 (gray), 2 (faded salmon), 3 (magenta), 4 (orange), 5 (violet), 6_mol A (yellow green), and 6_mol B (teal)].
Two different orientations (syn and anti) for the amide unit have been observed in substituted N′-(adamantan-2-ylidene)hydrazide derivatives, including the crystal structure of compound 1 described in this work. In structures 3–6, the amide unit adopts an anti-conformation while the corresponding unit exhibits syn-orientation in compounds 1 and 2. The structural superimposition diagram (Figure 2b) reveals the different orientations of the amide unit. It also reveals that the 4-pyridyl ring (compound 4) rotates relatively larger than 3-pyridyl (compound 3) and phenyl (compound 5) rings. It should be noted that the syn-orientation for an amide unit in compounds 1 and 2 helps to form an intramolecular C–S···N chalcogen bond between the thiophene sulfur atom and one of the N atoms, which is not part of amide unit. In other compounds, there is no intramolecular interaction formed.
2.2. Intramolecular S···N Chalcogen Bonds and Syn–Anti-Conformations of the Amide Unit
The molecular conformation of compounds 1 and 2 is locked by an intramolecular C–S···N (C15–S1···N1) chalcogen bond, which stabilizes the syn-conformation of the amide unit (Figure 2a). The geometrical parameters for this chalcogen bond [compound 1: S···N = 2.787 (1) Å and ∠CSN = 163.06(1)° and compound 2 (major disordered component with an occupancy ratio of 0.833): S···N = 2.780 (1) Å and ∠CSN = 162.44(1)°] are comparable in these two structures. To understand the role of this chalcogen bond in stabilizing the molecular conformation and find the energy barrier between the syn- and anti-orientation of the amide unit, we have performed a relaxed potential surface scan around the C11–C12 (C5–C4 in 2) and N2–C11 (N1–C5 in 2) bonds of the structure of 1. The former bond describes the rotation of the thiophene ring, while the latter bond corresponds to the rotation of the amide unit.
Figure 3 shows that the syn-orientation has the least-energy conformer (N2–C11–C12–S1 = 0°) and the energy difference between syn- and anti-conformations is about 1.7 kcal mol–1 (0.7 kcal mol–1 for compound 2). The corresponding torsion angle in the X-ray structure of 1 is 8.24° (15.62° in 2). To investigate the importance of chalcogen bonds in stabilizing the molecular conformation, we have taken the first (syn; N2–C11–C12–S1 = 0°) and second (anti; N2–C11–C12–S1 = 180°) least-energy conformers of compound 1, and these conformers were subjected to structural optimization in the gas phase at the M062X-D3/cc-pVTZ level of theory. The optimized structures were also used to determine intramolecular non-bonding interactions using topological analysis. It is observed that the S1···N1 chalcogen bond stabilizes the syn-conformer, while C–H···N interaction stabilizes the anti-conformer as shown in Figure S3. The topological parameters for these interactions are summarized in Table S1. The dissociation energy for a chalcogen bond (4.5 kcal mol–1) is relatively stronger than C–H···N interaction (3.1 kcal mol–1). The topological analysis for X-ray geometry of 1 also reveals that the dissociation energy for a chalcogen bond is 4.5 kcal mol–1.
Figure 3.

Calculated potential energy [with the B3LYP/6-31+G(d,p) level of theory] for rotation around the C11–C12 (structure 1) and C5–C4 (structure 2) bonds.
The relaxed potential energy scan around the N2–C11 (N1–C5 in 2) bond in structure 1 indicates that syn-orientation has the least energy compared to anti-conformations for the amide unit. The energy difference between these two conformers is about 4.2 kcal mol–1 (Figure S4, Supporting Information). We noticed that the anti-conformer makes short H···H contacts between amide NH and adjacent H of the thiophene ring (2.17 Å) and between amine H and adjacent H atoms of the adamantyl moiety (1.90 Å). The former and latter short contacts are relatively longer in the crystal structures. The syn-conformer of the amide unit stabilized with an intramolecular C–H···N interaction (Figure S5, Supporting Information).
2.3. Hirshfeld Surface Analysis and 2D-FP Plots
HS analysis was performed to investigate the nature of the intermolecular interactions and their quantitative contributions to the crystal packing of 1 and its closely related N′-(adamantan-2-ylidene)hydrazide derivatives reported earlier. This analysis sheds light on how the contribution of different noncovalent interactions altered due to different acyl substituents. The close inter-contacts are observed in structures 1–6, as depicted in the HSs in Figure S6.
In compound 1, the intense red spots are associated with intermolecular N–H···O, two C–H···O, and C–H···C(π) interactions, which are shorter than vdW radii of respective interacting atoms. The amide unit has participated in generating a synthon formed by N–H···O and C–H···O interactions. One of the H atoms of the thiophene ring is involved in three-centered C–H···O/C interactions with carbonyl O and C atoms, respectively. A similar feature is observed for the former synthon, while the latter bifurcated C–H···O/C interactions disappear in the unbrominated thiophene-containing derivative (compound 2). However, the relatively less intense red spots show intermolecular C–H···S interaction formed between one of the H atoms of the adamantyl moiety and the S atom of the thiophene ring in 2.
When thiophene or bromothiophene is replaced with 3-pyridyl (compound 3), 4-pyridyl (compound 4), and phenyl (compound 5) substituents, the nature of close inter-contacts shows different features. However, compounds 3–5 display similar inter-contacts features on the HS. For instance, the wide and intense red spot is associated with an intermolecular N–H···N hydrogen bond. The relatively less intense spots appear for N–H···O and C–H···O/N/C interactions and a short C···N contact, and the intensity of the red spots is comparable for these interactions.
The derivative with a methyl substituent (compound 6) displays similar features as observed in compounds 1 and 2. Specifically, the large red spots are attributed to strong N–H···O hydrogen bonds in both molecules A and B of 6. Two small red spots adjacent to the N–H···O hydrogen bonds belong to intermolecular C–H···O interactions. The protons of the methyl and adamantyl groups have participated as donors for the C–H···O interactions, and the carbonyl oxygen has been involved as an acceptor for N–H···O and C–H···O interactions. Moreover, these three interactions are formed between molecules A and B, which are linked alternately. In addition, there are less intense red spots observed for the N–H···N interaction, in which the NH group of molecule A interacts with the N atom of molecule B, and this interaction links the molecules A and B alternately, generating a C(4) chain that runs parallel to the crystallographic a axis. The intensity and size of the red spots are associated with the N–H···N hydrogen bond, which is markedly different in 6 and compounds 3–5, and this feature has not been observed in compounds 1 and 2.
2D-FP plots were obtained from the HS analysis for structures 1–6 to investigate the distribution of different inter-contacts found in the crystal structures and how different substituents influence these contacts. In all six structures, the significant contributions arise from H···H contacts, and their contributions range from 42 to 73.1% (Figure S7, Supporting Information). The selected 2D-FP plots of compounds 1–6 are depicted in Figure 4.
Figure 4.
2D-FP plots of compounds 1–6. Values in parentheses show the percentage relative contributions of selected intermolecular interactions to the total HS area.
The 2D-FP plots of compounds 1 and 2 show several invariant features and differences in the distribution pattern for some inter-contacts. The Br-substitution in compound 1 reduces the contribution of H···H contacts by 14.6% compared to compound 2 with no Br atom. A slight reduction in the contribution is also observed for H···C, H···N, and H···O contacts in 1. These reductions are compensated by increasing H···Br interactions, which contribute about 16.5%. When a phenyl moiety replaces the thiophene ring, the contributions of H···H, H···C, H···N, and H···O contacts are slightly increased.
The H···H contacts appear as a single spike in compounds 1 and 2, and the shortest inter-contact is located at ∼2.2 Å (de + di), whereas the corresponding contacts show a blunt tip, and the closest distance located at the tip is 2.2 Å in compounds 3–5. We note that the shortest H···H contacts are observed beyond 2.3–2.4 Å in compound 6. The distribution of H···O contacts looks very similar in compounds 1, 2, and 6 and appears as sharp double spikes with the distance at the tip being 1.8–2.0 Å. The corresponding contacts in compounds 3–5 are relatively longer (2.3–2.4 Å), which suggests that these contacts could be weaker. The wing-like pattern represents H···C (or C–H···π) interactions, and these contacts are almost disappeared due to the absence of the aromatic ring (compound 6). However, these interactions make significant contribution to respective crystal packing in the remaining structures with an aromatic ring.
A pair of sharp spikes is observed for H···N contacts in pyridyl isomers (3 and 4), and the distance at the tip is 2.2 Å in both compounds. The corresponding distance is observed beyond 2.2 Å in the remaining structures. Moreover, the 4-pyridyl substituent reduces the contribution of H···N contacts by 7.7% and slightly increases the contribution of H···H (by 4.5%) and H···C (3.6%) compared to the 3-pyridyl isomer. The relative contribution of different contacts including H···H, H···C, H···O, and H···N is comparable in derivatives containing 4-pyridyl and phenyl substituents. These features indicate that the effect of the 4-pyridyl substituent is marginal compared to the 3-pyridyl substituent.
As shown in Figure 5, 30% of the total HS area occupied by the other interaction categories in compound 1 and the H···Br and H···S contacts make notable contribution among other types of contacts. The H···Br interaction contributes about 16.5% toward crystal packing of 1, and the shortest distance appears at 2.9 Å in the 2D-FP plot. At the same distance, H···S contacts are also located and contribute about 9.3% to the total HS area. The contribution of H···S contacts is comparable in compounds 1 and 2. However, the shortest distance of H···S (2.8 Å) is slightly shorter in compound 2 compared to compound 1. The nature of the intermolecular contacts and their quantitative contributions to the crystal packing were described in detail for six closely related adamantyl phenylthiourea derivatives.55 In this study, the authors found that the contribution of H···H contacts ranges from 34 to 72%, which is comparable to the present study. In 1–5, the contribution of H···C contacts is similar to that of adamantyl phenylthiourea derivatives. There are significant differences in the contribution of H···O contacts between the title compounds and the adamantyl phenylthiourea derivatives.
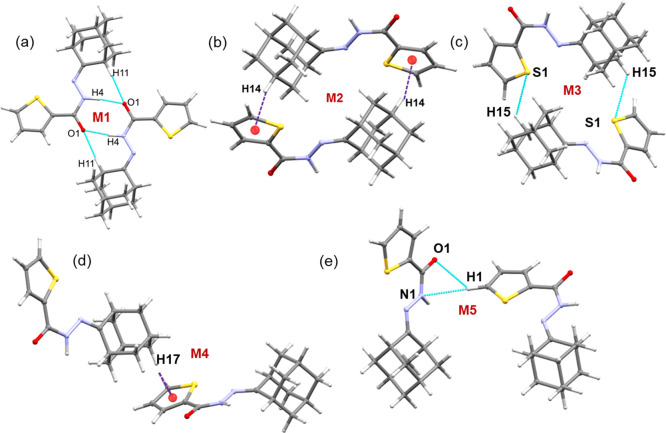
Figure 5.
Molecular dimers observed in the crystal structure of 1. (a) Strong M1 dimer is stabilized by N–H···O and C–H···O interactions, (b) three-centered (C–H···Br and C–H···S) interactions generate motif M2, (c) motif M3 stabilizes with C–H···O interaction, (d) adjacent M1 motifs are interlinked by motif M4, (e) motif M5 stabilizes with C–H···π interaction, and (f) supramolecular assembly built by alternate M1 and M6 motifs.
2.4. Molecular Dimers in the Crystal Structure of 1
The intermolecular interaction energies for the dimeric pairs observed in compounds 1–6 were calculated using the CLP-PIXEL code. This calculation revealed six dimeric motifs for compound 1 whose intermolecular interaction energies (Etot) range from −17.9 to −3.7 kcal mol–1 as summarized in Table 2. The dimeric motifs observed in compound 1 are illustrated in Figure 5. The total interaction energies of different molecular dimers are compared with those calculated by the counterpoise method using the B97D3/def2-TZVP level of approximation. In general, the energies of dimers calculated using two different approaches are comparable, except for dimers stabilized by C–H···π interactions.
Table 2. Intermolecular Interaction Geometries along with Energies (in kcal mol–1) for Various Dimers in 1–6 Obtained by the PIXEL and DFT Methodsa.
| PIXEL/MP2/6-31G** |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| dimer | CD | symmetry | important interactions | geometryb H···A (Å), ∠D–H···A (deg) | ECoul | Epol | Edisp | Erep | Etot | ΔEcp/B97D3/def2-TZVP |
| Compound 1 | ||||||||||
| M1 | 8.271 | –x + 1, –y + 1, –z + 2 | N2–H2···O1 | 1.93, 169 | –24.2 | –8.8 | –8.1 | 23.1 | –17.9 | –18.0 |
| C6–H6···O1 | 2.21, 156 | |||||||||
| M2 | 8.120 | x, −y + 1/2, z + 1/2 | C10–H10A···S1 | 2.91, 160 | –3.2 | –1.8 | –8.9 | 8.3 | –5.5 | –6.4 |
| C10–H10A···Br1 | 2.95, 132 | |||||||||
| M3 | 7.863 | –x + 1, y – 1/2, –z + 3/2 | C14–H14···O1 | 2.28, 163 | –3.5 | –1.5 | –5.9 | 5.8 | –5.2 | –6.4 |
| M4 | 7.742 | –x, y – 1/2, –z + 3/2 | C8–H8···Br1 | 3.04, 145 | –2.0 | –1.2 | –7.8 | 6.2 | –4.9 | –6.0 |
| M5 | 9.883 | x, y – 1, z | C4–H4···Cg1 | 2.85, 140 | –1.5 | –0.6 | –5.6 | 3.6 | –4.1 | –5.5 |
| M6 | 7.168 | –x, −y + 1, –z + 1 | C3–H3A···Br1 | 3.04, 144 | –1.1 | –0.5 | –4.6 | 2.6 | –3.7 | –4.6 |
| Compound 2 | ||||||||||
| M1 | 6.936 | –x + 1/2, –y + 1/2, –z + 1 | N1–H4···O1 | 1.84, 172 | –26.4 | –10.6 | –8.8 | 27.7 | –18.2 | –19.5 |
| C11–H11···O1 | 2.32, 142 | |||||||||
| M2 | 5.057 | –x + 1, –y + 1, –z + 1 | C13–H14···Cg1 | 2.53, 163 | –6.4 | –2.9 | –14.6 | 15.1 | –8.9 | –13.5 |
| M3 | 5.704 | –x + 1, y, −z + 1/2 | C14–H15···S1 | 2.95, 128 | –2.8 | –1.5 | –8.2 | 6.4 | –6.0 | –7.9 |
| M4 | 9.864 | x + 1/2, –y + 1/2, z + 1/2 | C15–H17···Cg1 | 2.61, 147 | –2.1 | –0.9 | –5.1 | 4.3 | –3.7 | –5.4 |
| M5 | 7.797 | x, −y + 1, z – 1/2 | C1–H1···O1 | 2.56, 132 | –1.0 | –0.8 | –3.1 | 2.3 | –2.6 | –3.0 |
| C1–H1···N1 | 2.68, 175 | |||||||||
| Compound 3 | ||||||||||
| M1 | 4.074 | –x + 1/2, –y + 3/2, z | N2–H19···N1 | 2.16, 160 | –15.9 | –7.5 | –17.0 | 22.8 | –17.6 | –18.6 |
| N2–H19···O1 | 2.49, 127 | |||||||||
| C1–H1···N1 | 2.43, 159 | |||||||||
| C1–H1···O1 | 2.40, 134 | |||||||||
| C13–H15···O1 | 2.34, 155 | |||||||||
| C9–H11···Cg2 | 2.85, 159 | |||||||||
| C13···N1 | 3.064 (1) | |||||||||
| M2 | 10.261 | –x + 3/2, y – 1/2, z – 1/2 | C4–H6··Cg2 | 2.70, 138 | –1.7 | –0.7 | –4.8 | 3.1 | –4.1 | –5.4 |
| Compound 4 | ||||||||||
| M1 | 4.056 | x – 1/2, –y + 3/2, z | N2–H19···N1 | 2.13, 162 | –17.0 | –8.1 | –17.8 | 24.6 | –18.3 | –18.6 |
| N2–H19···O1 | 2.54, 128 | |||||||||
| C1–H1···N1 | 2.38, 157 | |||||||||
| C1–H1··O1 | 2.39, 135 | |||||||||
| C13–H15···O1 | 2.39, 158 | |||||||||
| C4–H5···Cg3 | 2.88, 160 | |||||||||
| C13···N1 | 3.073 (1) | |||||||||
| M2 | 10.158 | –x + 1/2, y + 1/2, z + 1/2 | C8–H10··O1 | 2.65, 145 | –2.1 | –0.9 | –5.3 | 4.7 | –3.5 | –5.0 |
| C9–H12··Cg3 | 2.60, 142 | |||||||||
| Compound 5 | ||||||||||
| M1 | 4.046 | x – 1/2, y – 1, –z + 1/2 | N1–H1···N2 | 2.11, 161 | –16.7 | –8.2 | –17.4 | 25.5 | –16.9 | –18.7 |
| C9–H7···N2 | 2.47, 144 | |||||||||
| C9–H7···O1 | 2.25, 147 | |||||||||
| C3–H2···O1 | 2.32, 166 | |||||||||
| C3···N2 | 3.177 (1) | |||||||||
| M2 | 8.991 | –x + 1, y – 1/2, –z + 1/2 | C15–H17···Cg4 | 2.47, 164 | –2.8 | –1.2 | –6.1 | 5.8 | –4.3 | –6.1 |
| Compound 6 | ||||||||||
| M1B | 10.982 | –x + 1, –y + 1, –z + 1 | C24–H34···O2 | 2.34, 155 | –5.2 | –1.7 | –2.5 | 4.4 | –5.0 | –5.0 |
| M1AB | 4.851 | –x + 1, –y + 1, –z + 1 | N4–H33···N1 | 2.37, 127 | –15.4 | –6.7 | –12.5 | 18.1 | –16.5 | –16.7 |
| N4–H33···O1 | 2.03, 154 | |||||||||
| C20–H28···O1 | 2.28, 166 | |||||||||
| C24–H35···O1 | 2.45, 148 | |||||||||
| M2AB | 5.483 | –x, −y + 1, –z + 1 | N2–H15···N3 | 2.57, 123 | –14.1 | –5.7 | –8.6 | 14.1 | –14.1 | –14.4 |
| N2–H15···O2 | 1.97, 158 | |||||||||
| C8–H10···O2 | 2.33, 173 | |||||||||
| C12–H17···O2 | 2.51, 129 | |||||||||
| M3AB | 8.705 | x, y, z | C7–H9···O2 | 2.58, 135 | –1.4 | –1.0 | –2.7 | 1.9 | –3.2 | –3.4 |
Cg1, Cg2, Cg3, and Cg4 are centroids of thiophene, 3-pyridyl, 4-pyridyl, and phenyl rings and CD is the distance between the geometrical centres of the molecules.
Neutron values are given for all D–H···A interactions.
The inversion-related molecules form the most stable dimer (motif M1) via intermolecular N–H···O and C–H···O hydrogen bonds, in which the carbonyl oxygen atom (O1) is involved as an acceptor for both interactions (Figure 5a). The role of these interactions is to connect the molecules in the adjacent layers to build columnar packing along the crystallographic ac plane. As expected, this dimeric motif is predominantly electrostatic in nature, with an 80% contribution toward the stabilization.
Motif M2 is stabilized by intermolecular C–H···S and C–H···Br interactions, in which one of the protons (atom H10A) of the adamantyl moiety is involved as a donor for both of these interactions (Figure 5b). These interactions link the molecules to generate a chain that runs parallel to the c axis. Furthermore, the electrostatic and dispersion energies contribute about 36 and 64% toward the stabilization of dimer M2.
One of the H atoms of the bromothiophene ring is involved in an intermolecular C–H···O interaction with the carbonyl O atom. This interaction stabilizes the molecular dimer M3 and links the molecules into a chain parallel to the crystallographic b axis (Figure 5c). The PIXEL energy analysis reveals that the component of electrostatic energy contributes about 46% toward the stabilization of this dimer. The dispersive nature (71%) of dimer M4 is caused by an intermolecular C–H···Br interaction, in which one of the adamantyl H atoms is involved in the interaction. Furthermore, motif M4 bridges the adjacent dimers of motif M1, and these two motifs are arranged alternately to form a chain (Figure 5d).
Additionally, M5 and M6 motifs are dispersive in nature, contributing 73–74% toward stabilizing these motifs. The former motif M5 is stabilized by an intermolecular C–H···π interaction, and adamantyl and thiophene rings are engaged in this interaction (Figure 5e). The latter motif M6 is formed by an intermolecular C–H···Br interaction to generate a ring as shown in Figure 5f. Furthermore, motifs M6 and M1 are also arranged alternately in the crystal structure.
2.5. Molecular Dimers in Related Structures
The CLP-PIXEL calculation was carried out for five closely related structures of 1 in order to identify the energetically significant dimers found in the solid state. The intermolecular interaction energy and its various energy components are summarized in Table 2. Structure 2 has five molecular dimers with intermolecular interaction energies ranging from −18.2 to −2.6 kcal mol–1. As shown in Figure 6, these dimers stabilize by intermolecular N–H···O and C–H···O/N/S/π interactions, which are similar to compound 1 except C–H···Br and C–H···N interactions. The intermolecular interaction energy of dimer M1 in structures 1 and 2 is nearly the same (Figure 6a). The electrostatic energy contributes about 81% toward the stabilization of dimer M1 in this structure. For the stabilization of remaining dimers (M2–M5; Figure 6b–e), the electrostatic and dispersion energies contribute in the range of 34–39 and 61–66%, respectively.
Figure 6.
Molecular dimers observed in the crystal structure of 2. (a) Strongest M1 dimer is stabilized by N–H···O [R22(8) ring] and C–H···O [R22(14) ring] interactions, (b) intermolecular C–H···π interaction stabilizes the molecular dimer M2, (c) weak C–H···S interaction provides stability to the dimer M3, (d) motif M4 stabilizes with an intermolecular C–H···π interaction, and (e) bifurcated C–H···O/N interactions stabilize motif M5.
In the remaining compounds (3–6), the most stable dimer (motif M1) in these structures stabilized by more number of interactions, including N–H···O/N and C–H···O/N/π interactions, could be due to the trans-conformation of the amide unit in these structures. In compounds 1 and 2, the amide unit exhibits the cis-conformation. The molecular dimers observed in structures 3–6 are illustrated in Figure 7.
Figure 7.
Dimers in the crystal structures of 3–5 and a small red sphere represents centroid of the aromatic ring in these structures. Dimers in these structures are labeled.
3-Pyridyl (compound 3) and 4-pyridyl (compound 4) derivatives have only two molecular dimers in each in the solid state. In both compounds, the most stable dimer (M1) is stabilized by N–H···N/O and C–H···N/O/π interactions and a C···N short contact [3.064–3.073 (1) Å]. A short C···N contact (3.177 Å) is also present in dimer M1 of structure 5. The intermolecular interaction energy (−17.6 and −18.3 kcal mol–1) for this dimer is comparable in these two structures. The second most stable dimer (M2) is stabilized primarily by an intermolecular C–H···π interaction. The H atom of the adamantyl unit and pyridine ring has participated in C–H···π interaction in 3 and 4 as a donor and an acceptor, respectively. In compound 4, dimer M2 is further stabilized by an intermolecular C–H···O interaction. Again, the intermolecular interaction energy for the M2 dimer in both compounds is comparable.
The phenyl derivative 5 also has two molecular dimers similar to compounds 3 and 4. The stability of the M1 dimer in 5 is slightly weaker than the M1 dimers in 3 and 4. The presence of additional N–H···O and C–H···π interaction in structures 3 and 4 could be a reason for more stability, and no such C–H···π is present in dimer M1 of compound 5. Another common feature observed in 3–5 is that the dimer M2 is stabilized by an intermolecular C–H···π interaction, in which the aromatic ring acts as an acceptor in all three compounds. Furthermore, the strength of this dimer in these compounds is very similar. For the stabilization of M1 dimers in 3–5, the contribution of electrostatic energy is nearly the same (58–59%). However, the contribution of electrostatic energy is significantly reduced (∼20%) compared to that on structures 1 and 2.
The methyl derivative 6 was crystallized in the monoclinic system with the P21/c space group, and the asymmetric unit contains two crystallographically independent molecules (A and B). In the asymmetric unit, molecules A and B interact via intermolecular C–H···O interaction (motif M3AB). The dimeric motifs observed in this structure are depicted in Figure 8. The PIXEL energy analysis suggests that no direct interaction is formed between the A molecule and its symmetry-related partners. However, an intermolecular C–H···O interaction is formed between molecule B and its symmetry-related equivalents (motif M1B), generating a cyclic dimer [R22(8)]. Three molecular dimers are formed between A and B molecules and their symmetry equivalents (motifs M1AB to M3AB). The first two most stable dimers (M1AB and M2AB) are stabilized by N–H···N/O and C–H···O interactions, which are similar to those observed in compounds 3–5. However, the strength of these dimers is slightly weaker compared to other structures. It is noted that intermolecular C–H···N interaction is not observed in these dimers. However, compounds 2–5 have C–H···N interaction, which provides additional stabilization to dimer M1 in these structures.
Figure 8.
Dimers in the crystal structure of 6. Dimers in this structure are labeled.
2.6. Lattice Energies
The total lattice energies (Etot) for crystal structure 1 and its closely related analogues 2–6 were calculated and are summarized in Table 3. This table shows that the contribution of electrostatic energy (ECoul + Epol) and dispersion energy (Edisp) components toward the stabilization of crystal structures is comparable. The contribution of dispersion energy is 51 to 57%, while the contribution of electrostatic energy is 43 to 49%. From the total stabilization energy, we can see that the stabilization energy is slightly higher (4.4 kcal mol–1) due to Br substitution compared to that of compound 2, in which the H atom replaces the Br atom. Similarly, we noted the stabilization energy difference (6.8 kcal mol–1) between 3-pyridyl (compound 3) and 4-pyridyl (compound 4) substituents and higher stabilization energy (−76.6 kcal mol–1) for compound 4. The total stabilization energy (−75.9 kcal mol–1) for phenyl (compound 5) is comparable with that of compound 4. Compared to the other five compounds, the lower stabilization (−59.0 kcal mol–1) is observed for the methyl derivative (compound 6).
Table 3. Lattice Energy (in kcal mol–1).
| compound code | Coulombic | polarization | dispersion | repulsion | cell dipole | total |
|---|---|---|---|---|---|---|
| 1 | –22.6 | –8.9 | –42.1 | 39.0 | 0.0 | –34.7 |
| ILESAV | –21.6 | –11.2 | –36.6 | 32.5 | –0.5 | –37.2 |
| OKOLOR | –23.4 | –12.1 | –40.2 | 38.8 | –0.9 | –37.8 |
| XEBTIJ | –23.2 | –11.7 | –41.0 | 39.8 | 0.0 | –36.1 |
| YAJBES | –22.4 | –9.5 | –37.3 | 38.1 | 0.0 | –31.1 |
| NICCEH | –19.9 | –9.2 | –29.8 | 27.4 | 0.0 | –31.6 |
2.7. Topological Analysis
The topological
and energetic properties of the noncovalent interactions at the bond
critical points (BCPs) in the observed dimers of the crystal structures
of 1–6 were analyzed by comparing selected topological
properties, including the electron density [ρ(r)], the Laplacian of electron density [∇2ρ(r)], the potential electronic energy density [V(r)], the kinetic electronic energy density [G(r)], the total electronic energy density
[H(r) = V(r) + G(r)], the | | value, and the dissociation energy value
(De) proposed by Espinosa et al.(56)
| value, and the dissociation energy value
(De) proposed by Espinosa et al.(56)
In 1, the intermolecular N–H···O and C–H···O/S/Br/π interactions have been involved in the stabilization of observed molecular dimers (Table S2 and Figure S8, Supporting Information). According to Gatti’s assignment, all these interactions are classified as closed-shell interactions using the values of |−V(r)/G(r)| < 1, H(r) > 0, and the positive value of Laplacian of electron density [∇2ρ(r) > 0].57 Among the observed intermolecular interactions, the N–H···O interaction is more vital as evident from the dissociation energy value (De = 6.7 kcal mol–1). The corresponding value is in the range of 2.7–3.1 kcal mol–1 for C–H···O interactions. The strength of C–H···S and two of the C–H···Br interactions (M2 and M4) is comparable, and the dissociation energy value is in the range of 1.0–1.3 kcal mol–1. In addition, the De value is equal for one of the C–H···Br interactions and C–H···π interaction observed in dimers M6 and M5, respectively. Furthermore, we used the KP-4 rule to differentiate between hydrogen bond and van der Waals interactions.58 This analysis suggests that all the interactions mentioned above show hydrogen bonding character, except C–H···π interaction (dimer M5). The C–H···π interaction displays van der Waals character as the bond path (Rij) value is longer (3.483 Å).
In 2, the N–H···O interaction shows intermediate bonding character between shared and closed-shell interaction according to Gatti’s assignment [∇2ρ(r) > 1 and H(r) < 0 and |−V(r)/G(r)| > 1]. The dissociation energy value for this hydrogen bond is 9.3 kcal mol–1. The remaining interactions observed in different dimers of 2 are closed-shell interactions (Table S2 and Figure S9, Supporting Information). The strength of the C–H···S hydrogen bond is the same in structures 1 and 2. The dissociation energy for the C–H···N interaction is 1.2 kcal mol–1 and slightly weaker as compared to that of structures 3 and 4.
In structures 3–5, the strength of N–H···N interaction is more than two times stronger than N–H···O interaction. The De value for the N–H···N interaction is in the range of 4.0–4.6 kcal mol–1 (Table S3 and Figures S10–S12, Supporting Information). In addition, the strength of C–H···O interactions found in dimer M1 in these structures is comparable. It should be noted that the C···N interaction is slightly stronger than some of the C–H···O and C–H···π interactions. Structure 6 demonstrates contrasting strengths between N–H···N and N–H···O interactions compared to structures 3–5 (Table S3 and Figure S13). In 6, the N–H···O interaction is stronger than N–H···N interaction. It is worth noting that the De value for N–H···O interaction is comparable to that of structure 1. Similarly, the De value for C–H···O interactions is comparable in all six structures described in this study. Furthermore, all the noncovalent interactions found in these structures are closed-shell-type interactions based on Gatti’s assignment, except for N–H···O hydrogen bonds in structure 2. Furthermore, it is noted that the N–H···O interactions show the trend of exponential decay (with R2 > 0.99) in the magnitudes of electron density [ρ(r)], Laplacian of electron density [∇2ρ(r)], and the dissociation energy with the increasing length of the bond path (Rij). A similar trend is also noted for N–H···N interactions observed in these six structures. The C–H···O/N interactions also follow the similar exponential decay with the R2 > 0.94 for the above pairs.
2.8. In Vitro Antiproliferative Activity
The in vitro antiproliferative activity of the title compound 1 was assessed against five human tumor cell lines, namely, PC-3 (human prostate cancer), HEPG-2 (hepatocellular carcinoma), HCT-116 (colorectal carcinoma), MCF-7 (mammary gland breast cancer), and HeLa (epithelioid carcinoma) using 3-[4,5-dimethylthiazoyl-2-yl]-2,5-diphenyltetrazolium bromide assay.59,60 The results of in vitro antiproliferative activity of compound 1 and the anticancer drug doxorubicin61 are shown in Table 4.
Table 4. In Vitro Antiproliferative Activity of Compound 1 and the Anticancer Drug Doxorubicin against Human Prostate Cancer (PC-3), Hepatocellular Carcinoma (HEPG-2), Colorectal Carcinoma (HCT-116), Mammary Gland Breast Cancer (MCF-7), and Epithelioid Carcinoma (HeLa) Cell Lines.
| IC50 (μM)a |
|||||
|---|---|---|---|---|---|
| PC-3 | HEPG-2 | HCT-116 | MCF-7 | HeLa | |
| compound 1 | 41.75 ± 2.9 | 16.38 ± 1.5 | 29.55 ± 2.2 | 9.32 ± 0.7 | 7.18 ± 0.5 |
| doxorubicin | 8.87 ± 0.6 | 4.50 ± 0.2 | 5.23 ± 0.3 | 4.17 ± 0.2 | 5.57 ± 0.4 |
IC50 values presented as the mean ± SD of three separate determinations, p < 0.05.
The antiproliferative activity results showed that compound 1 displayed potent antiproliferative activity (IC50 < 10 μM) against MCF-7 and HeLa cell lines with IC50 9.32 and 7.18 μM, respectively, moderate activity (IC50 16.38 μM) against HEPG-2 cell lines, and lower activity against PC-3 and HCT-116 cell lines with IC50 41.75 and 29.55 μM, respectively. Based on the antiproliferative activity of compound 1, it would be considered as a promising anticancer drug candidate for further investigations.
3. Conclusions
In conclusion, we present the synthesis of the N′-(adamantan-2-ylidene)-5-bromothiophene-2-carbohydrazide compound exhibiting antiproliferative activity. The structure and energetics of molecular dimers in six closely related N′-(adamantan-2-ylidene)hydrazide derivatives have been described. Two of the compounds in this series possess an intramolecular C–S···N-type chalcogen bond. The potential energy surface scan and topological analysis revealed the importance of this σ–hole interaction in fixing the molecular conformation. HS analysis has been used to study the effect of the R group on the contribution of intermolecular interactions toward crystal packing. PIXEL energy analysis revealed that the strong dimer formed in all six structures stabilized primarily by strong N–H···O and N–H···N interactions. Additional stabilization comes from weak C–H···O/N/π interactions. Topological analysis of intermolecular interactions at their BCPs revealed that all of the interactions are closed-shell interactions, except N–H···O hydrogen bonds in one of the structures discussed in this study. This study is expected to be essential for understanding the supramolecular self-assembly driven by multiple hydrogen bonds and conformational preferences (syn and anti) of amide units present in these derivatives, which could help understand the antiproliferative activity.
4. Materials and Methods
4.1. Synthesis and Crystallization
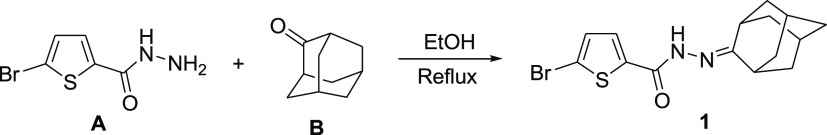
Compound 1 was synthesized via condensation of 5-bromothiophene-2-carbohydrazide A with 2-adamantanone B (Scheme 1).
Scheme 1. Synthetic Pathway for Compound 1.
A mixture of 5-bromothiophene-2-carbohydrazide (1.1 g, 5.0 mmol) and 2-adamantanone (0.75 g, 5.0 mmol) in ethanol (10 mL) was heated under reflux with stirring for 3 h. On cooling, the precipitated crystalline solid was filtered, dried, and recrystallized from ethanol to yield 1.62 g (92%) of the title compound 1 (C15H17BrN2OS) as colorless block crystals. mp: 224–226 °C. 1H NMR (CDCl3, 500.13 MHz): δ 1.87–2.11 (m, 14H, adamantane-H), 7.10 (d, 1H, thiophene-H, J = 4.0 Hz), 7.88 (d, 1H, thiophene-H, J = 4.0 Hz), 10.05 (s, 1H, NH). 13C NMR (CDCl3, 125.76 MHz): δ 27.74, 31.03, 37.82, 39.24, 162.31 (adamantane-C), 123.62, 128.94, 133.80, 134.66 (thiophene-C), 164.47 (C=O).
4.2. Single-Crystal X-ray Diffraction
X-ray intensity data for compound 1 were collected on a Bruker APEX-II CCD diffractometer at room temperature (296 K) using Mo Kα radiation. The absorptions were corrected by the SADABS multi-scan method (SADABS, Bruker, 2014). The structure was solved using the SIR-2011 program,62 and the structural refinement was performed with the SHELXL 2018/3 program.63 The position of the amide NH atom was located from a difference Fourier map and refined freely. All the remaining H atoms were placed in geometrically idealized positions with C–H = 0.93–98 Å and were constrained to ride on their parent atoms with Uiso(H) = 1.2Ueq(C). During the final refinement, the reflections 0 1 1 and −6 0 6 were omitted due to [error/esd] > 10. The final refined structure was checked using PLATON.64,65 The crystal packing and molecular dimers were drawn using the program MERCURY.66
4.3. Computational Details
The HS analysis and 2D-FP plots for structures 1–6 were obtained with the CrystalExplorer-17.5 program.67
The lattice energies and intermolecular interaction energies for dimers formed in crystal structures of 1–6 were calculated using the PIXELC code implemented in the CLP program.68,69 The electron density of the molecules was obtained at the MP2/6-31G** level of theory for the PIXELC calculation. All the density functional theory (DFT) calculations were performed with the Gaussian 09 program.70 For compound 1, the relaxed potential energy surface scan was performed for the torsion angle N2–C11–C12–S1 with the B3LYP/6-31+G(d,p) level of theory with an increase of 10° starting from −180 to 180°. The complexation energies for dimers observed in the crystal structures were also calculated at the B97D3/def2-TZVP level of theory,71,72 incorporating Grimme’s empirical dispersion correction term (D3).73 The complexation energies (ΔEcp) were also corrected for basis set superposition errors using the counterpoise method.74
The theoretical charge density analysis was performed for the dimers (in their X-ray geometries with normalized H positions and at the M06-2X-D3/cc-pVTZ level of theory) using the AIMALL package.75 Selected topological parameters such as electron density, ρ(r), Laplacian of the electron density, ∇2ρ(r), potential electronic energy density V(r), kinetic electronic energy density, G(r), and total electronic energy density [H(r) = V(r) + G(r)] were used at their BCPs to characterize the nature and strength of intermolecular interactions. The dissociation energy (De) for interaction was calculated using the EML empirical scheme, that is, De = −0.5 × V(r), to assess the strength of the interaction.56 The noncovalent bonding nature and character (HB and vdW interactions) were characterized using first four criteria of Koch–Popelier58 as described in our earlier work.22
Acknowledgments
This work was funded by the Deanship of Scientific Research at Princess Nourah bint Abdulrahman University through the Research groups Program (grant no. RGP-1442-0010-5).
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsomega.2c00159.
1H and 13C NMR spectra for compound 1, potential energy surface scan, topological parameters for intramolecular and intermolecular interactions, and molecular graphs for intramolecular and intermolecular interactions (PDF)
The authors declare no competing financial interest.
Supplementary Material
References
- Spilovska K.; Zemek F.; Korabecny J.; Nepovimova E.; Soukup O.; Windisch M.; Kuca K. Adamantane—A Lead Structure for Drugs in Clinical Practice. Curr. Med. Chem. 2016, 23, 3245–3266. 10.2174/0929867323666160525114026. [DOI] [PubMed] [Google Scholar]
- Wanka L.; Iqbal K.; Schreiner P. R. The Lipophilic Bullet Hits the Targets: Medicinal Chemistry of Adamantane Derivatives. Chem. Rev. 2013, 113, 3516–3604. 10.1021/cr100264t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J.; Obando D.; Liao V.; Lifa T.; Codd R. The Many Faces of the Adamantyl Group in Drug Design. Eur. J. Med. Chem. 2011, 46, 1949–1963. 10.1016/j.ejmech.2011.01.047. [DOI] [PubMed] [Google Scholar]
- Lamoureux G.; Artavia G. Use of the Adamantane Structure in Medicinal Chemistry. Curr. Med. Chem. 2010, 17, 2967–2978. 10.2174/092986710792065027. [DOI] [PubMed] [Google Scholar]
- Wendel H. A.; Snyder M. T.; Pell S. Trial of Amantadine in Epidemic Influenza. Clin. Pharmacol. Ther. 1966, 7, 38–43. 10.1002/cpt19667138. [DOI] [PubMed] [Google Scholar]
- Vernier V.; Harmon J. B.; Stump J. M.; Lynes T. E.; Marvel J. P.; Smith D. H. The Toxicologic and Pharmacologic Properties of Amantadine Hydrochloride. Toxicol. Appl. Pharmacol. 1969, 15, 642–665. 10.1016/0041-008X(69)90066-0. [DOI] [PubMed] [Google Scholar]
- Rabinovich S.; Baldini J. T.; Bannister R. Treatment of Influenza. The Therapeutic Efficacy of Rimantadine HC1 in a Naturally Occurring Influenza A2 Outbreak. Am. J. Med. Sci. 1969, 257, 328–335. 10.1097/00000441-196905000-00005. [DOI] [PubMed] [Google Scholar]
- Rosenthal K. S.; Sokol M. S.; Ingram R. L.; Subramanian R.; Fort R. C. Tromantadine: Inhibitor of Early and Late Events in Herpes Simplex Virus Replication. Antimicrob. Agents Chemother. 1982, 22, 1031–1036. 10.1128/aac.22.6.1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Long J.; Manchandia T.; Ban K.; Gao S.; Miller C.; Chandra J. Adaphostin Cytoxicity in Glioblastoma Cells Is ROS-Dependent and Is Accompanied by Upregulation of Heme Oxygenase-1. Cancer Chemother. Pharmacol. 2007, 59, 527–535. 10.1007/s00280-006-0295-5. [DOI] [PubMed] [Google Scholar]
- Li M.; Wang H.; Hill D. L.; Stinson S.; Veley K.; Grossi I.; Peggins J.; Covey J. M.; Zhang R. Preclinical Pharmacology of the Novel Antitumor Agent Adaphostin, a Tyrphostin Analog That Inhibits Bcr/Abl. Cancer Chemother. Pharmacol. 2006, 57, 607–614. 10.1007/s00280-005-0094-4. [DOI] [PubMed] [Google Scholar]
- Holmes W. F.; Soprano D. R.; Soprano K. J. Comparison of the Mechanism of Induction of Apoptosis in Ovarian Carcinoma Cells by the Conformationally Restricted Synthetic Retinoids CD437 and 4-HPR. J. Cell. Biochem. 2003, 89, 262–278. 10.1002/jcb.10505. [DOI] [PubMed] [Google Scholar]
- Han T.; Goralski M.; Capota E.; Padrick S. B.; Kim J.; Xie Y.; Nijhawan D. The Antitumor Toxin CD437 Is a Direct Inhibitor of DNA Polymerase α. Nat. Chem. Biol. 2016, 12, 511–515. 10.1038/nchembio.2082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorenzo P.; Alvarez R.; Ortiz M. A.; Alvarez S.; Piedrafita F. J.; de Lera Á. R. Inhibition of IkappaB Kinase-Beta and Anticancer Activities of Novel Chalcone Adamantyl Arotinoids. J. Med. Chem. 2008, 51, 5431–5440. 10.1021/jm800285f. [DOI] [PubMed] [Google Scholar]
- Nasr R. R.; Hmadi R. A.; El-Eit R. M.; Iskandarani A. N.; Jabbour M. N.; Zaatari G. S.; Mahon F.-X.; Pisano C. C. P.; Darwiche N. D. ST1926, an Orally Active Synthetic Retinoid, Induces Apoptosis in Chronic Myeloid Leukemia Cells and Prolongs Survival in a Murine Model. Int. J. Cancer 2015, 137, 698–709. 10.1002/ijc.29407. [DOI] [PubMed] [Google Scholar]
- Yang J.; Yang C.; Zhang S.; Mei Z.; Shi M.; Sun S.; Shi L.; Wang Z.; Wang Y.; Li Z.; Xie C. ABC294640, a Sphingosine Kinase 2 Inhibitor, Enhances the Antitumor Effects of TRAIL in Non-Small Cell Lung Cancer. Cancer Biol. Ther. 2015, 16, 1194–1204. 10.1080/15384047.2015.1056944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dai L.; Smith C. D.; Foroozesh M.; Miele L.; Qin Z. The Sphingosine Kinase 2 Inhibitor ABC294640 Displays Anti-Non-Small Cell Lung Cancer Activities in Vitro and in Vivo. Int. J. Cancer 2018, 142, 2153–2162. 10.1002/ijc.31234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- El-Emam A. A.; Al-Deeb O. A.; Al-Omar M.; Lehmann J. Synthesis, Antimicrobial, and Anti-HIV-1 Activity of Certain 5-(1-Adamantyl)-2-Substituted Thio-1,3,4-Oxadiazoles and 5-(1-Adamantyl)-3-Substituted Aminomethyl-1,3,4-Oxadiazoline-2-Thiones. Bioorg. Med. Chem. 2004, 12, 5107–5113. 10.1016/j.bmc.2004.07.033. [DOI] [PubMed] [Google Scholar]
- Al-Wahaibi L.; Hassan H.; Abo-Kamar A.; Ghabbour H.; El-Emam A. Adamantane-Isothiourea Hybrid Derivatives: Synthesis, Characterization, In Vitro Antimicrobial, and In Vivo Hypoglycemic Activities. Molecules 2017, 22, 710. 10.3390/molecules22050710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- El-Emam A. A.; Al-Tamimi A.-M. S.; Al-Omar M. A.; Alrashood K. A.; Habib E. E. Synthesis and Antimicrobial Activity of Novel 5-(1-Adamantyl)-2-Aminomethyl-4-Substituted-1,2,4-Triazoline-3-Thiones. Eur. J. Med. Chem. 2013, 68, 96–102. 10.1016/j.ejmech.2013.07.024. [DOI] [PubMed] [Google Scholar]
- Al-Mutairi A. A.; Al-Alshaikh M. A.; Al-Omary F. A. M.; Hassan H. M.; El-Mahdy A. M.; El-Emam A. A. Synthesis, Antimicrobial, and Anti-Proliferative Activities of Novel 4-(Adamantan-1-Yl)-1-Arylidene-3-Thiosemicarbazides, 4-Arylmethyl N′-(Adamantan-1-yl)Piperidine-1-Carbothioimidates, and Related Derivatives. Molecules 2019, 24, 4308. 10.3390/molecules24234308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hassan H. M.; Al-Wahaibi L. H.; Shehatou G. S.; El-Emam A. A. Adamantane-Linked Isothiourea Derivatives Suppress the Growth of Experimental Hepatocellular Carcinoma via Inhibition of TLR4-MyD88-NF-ΚB Signaling. Am. J. Cancer Res. 2021, 11, 350–369. [PMC free article] [PubMed] [Google Scholar]
- El-Emam A. A.; Saveeth Kumar E.; Janani K.; Al-Wahaibi L. H.; Blacque O.; El-Awady M. I.; Al-Shaalan N. H.; Percino M. J.; Thamotharan S. Quantitative Assessment of the Nature of Noncovalent Interactions in N-Substituted-5-(Adamantan-1-yl)-1,3,4-Thiadiazole-2-Amines: Insights from Crystallographic and QTAIM Analysis. RSC Adv. 2020, 10, 9840–9853. 10.1039/D0RA00733A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al-Wahaibi L. H.; Joubert J.; Blacque O.; Al-Shaalan N. H.; El-Emam A. A. Crystal Structure, Hirshfeld Surface Analysis and DFT Studies of 5-(Adamantan-1-yl)-3-[(4-Chlorobenzyl)Sulfanyl]-4-Methyl-4H-1,2,4-Triazole, a Potential 11β-HSD1 Inhibitor. Sci. Rep. 2019, 9, 19745. 10.1038/s41598-019-56331-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al-Wahaibi L. H.; Grandhi D. S.; Tawfik S. S.; Al-Shaalan N. H.; Elmorsy M. A.; El-Emam A. A.; Percino M. J.; Thamotharan S. Probing the Effect of Halogen Substituents (Br, Cl, and F) on the Non-Covalent Interactions in 1-(Adamantan-1-yl)-3-Arylthiourea Derivatives: A Theoretical Study. ACS Omega 2021, 6, 4816–4830. 10.1021/acsomega.0c05793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al-Wahaibi L. H.; Sujay S.; Muthu G. G.; El-Emam A. A.; Venkataramanan N. S.; Al-Omary F. A. M.; Ghabbour H. A.; Percino J.; Thamotharan S. Theoretical Investigations of Two Adamantane Derivatives: A Combined X-Ray, DFT, QTAIM Analysis and Molecular Docking. J. Mol. Struct. 2018, 1159, 233–245. 10.1016/j.molstruc.2018.01.064. [DOI] [Google Scholar]
- Al-Omary F. A. M.; Chowdary Gude N.; Al-Rasheed L. S.; Alkahtani H. N.; Hassan H. M.; Al-Abdullah E. S.; El-Emam A. A.; Percino M. J.; Thamotharan S. X-Ray and Theoretical Investigation of (Z)-3-(Adamantan-1-yl)-1-(Phenyl or 3-Chlorophenyl)-S-(4-Bromobenzyl)Isothioureas: An Exploration Involving Weak Non-Covalent Interactions, Chemotherapeutic Activities and QM/MM Binding Energy. J. Biomol. Struct. Dyn. 2020, 1–16. 10.1080/07391102.2020.1840443. [DOI] [PubMed] [Google Scholar]
- Groom C. R.; Bruno I. J.; Lightfoot M. P.; Ward S. C. The Cambridge Structural Database. Acta Crystallogr., Sect. B: Struct. Sci., Cryst. Eng. Mater. 2016, 72, 171–179. 10.1107/S2052520616003954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kadi A. A.; Alanzi A. M.; El-Emam A. A.; Ng S. W.; Tiekink E. R. T. N’-(Adamantan-2-ylidene)Thiophene-2-Carbohydrazide. Acta Crystallogr., Sect. E: Struct. Rep. Online 2011, 67, o3127. 10.1107/s1600536811044758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al-Tamimi A.-M. S.; Ghabbour H. A.; El-Emam A. A. Crystal Structure of N′-(Adamantan-2-ylidene)Pyridine-3-Carbohydrazide, C16H19N3O. Z. Kristallogr.—New Cryst. Struct. 2016, 231, 573–575. 10.1515/ncrs-2015-0194. [DOI] [Google Scholar]
- Al-Abdullah E. S.; Ghabbour H. A.; Al-Jabal M. M.; Fun H.-K.; El-Emam A. A. Crystal Structure of N′-(Adamantan-2-ylidene)-Isonicotinohydrazide, C16H19N3O. Z. Kristallogr.—New Cryst. Struct. 2016, 231, 273–275. 10.1515/ncrs-2015-0118. [DOI] [Google Scholar]
- Almutairi M. S.; El-Emam A. A.; El-Brollosy N. R.; Said-Abdelbaky M.; García-Granda S. N’-(Adamantan-2-ylidene)Benzohydrazide. Acta Crystallogr., Sect. E: Struct. Rep. Online 2012, 68, o2247–o2248. 10.1107/S1600536812028644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hashmi A. S. K.; Grundl M. A.; Nass A. R.; Naumann F.; Bats J. W.; Bolte M. Photochemical Synthesis of Prochiral Dialkyl 3,3-Dialkylcyclopropene-1,2-Dicarboxylates with Facial Shielding Substituents and Related Substrates. Eur. J. Org. Chem. 2001, 2001, 4705–4732. 10.1002/1099-0690(200112)2001:24<4705::aid-ejoc4705>3.0.co;2-j. [DOI] [Google Scholar]
- Jeffrey G. A.; Saenger W.. Hydrogen Bonding in Biological Structures; Springer Science & Business Media, 2012. [Google Scholar]
- Derewenda Z. S.; Lee L.; Derewenda U. The Occurrence of C–H···O Hydrogen Bonds in Proteins. J. Mol. Biol. 1995, 252, 248–262. 10.1006/jmbi.1995.0492. [DOI] [PubMed] [Google Scholar]
- Manikandan K.; Ramakumar S. The Occurrence of C–H···O Hydrogen Bonds in α-Helices and Helix Termini in Globular Proteins. Proteins: Struct., Funct., Bioinf. 2004, 56, 768–781. 10.1002/prot.20152. [DOI] [PubMed] [Google Scholar]
- Leonard G. A.; McAuley-Hecht K.; Brown T.; Hunter W. N. Do C-H...O Hydrogen Bonds Contribute to the Stability of Nucleic Acid Base Pairs?. Acta Crystallogr., Sect. D: Struct. Biol. 1995, 51, 136–139. 10.1107/S0907444994004713. [DOI] [PubMed] [Google Scholar]
- Sudarvizhi V.; Balakrishnan T.; Percino M. J.; Stoeckli-Evans H.; Thamotharan S. Evaluation of Charge Assisted Hydrogen Bonds in L-(S)-Lysinium L-(S)-Mandelate Dihydrate and L-(S)-Alanine L-(S)-Mandelic Acid Complexes: Inputs from Hirshfeld Surface, PIXEL Energy and QTAIM Analysis. J. Mol. Struct. 2020, 1220, 128701. 10.1016/j.molstruc.2020.128701. [DOI] [Google Scholar]
- Seth S. K.; Bauzá A.; Frontera A. Bipolar Behaviour of Salt-Bridges: A Combined Theoretical and Crystallographic Study. New J. Chem. 2018, 42, 12134–12142. 10.1039/C8NJ02194E. [DOI] [Google Scholar]
- Tripathi S.; Islam S.; Seth S. K.; Bauzá A.; Frontera A.; Mukhopadhyay S. Supramolecular Assemblies Involving Salt Bridges: DFT and X-Ray Evidence of Bipolarity. CrystEngComm 2020, 22, 8171–8181. 10.1039/D0CE01356K. [DOI] [Google Scholar]
- Steiner T.; Desiraju G. R. Distinction between the Weak Hydrogen Bond and the van Der Waals Interaction. Chem. Commun. 1998, 8, 891–892. 10.1039/A708099I. [DOI] [Google Scholar]
- Ghosh S.; Chopra P.; Wategaonkar S. C–H···S Interaction Exhibits All the Characteristics of Conventional Hydrogen Bonds. Phys. Chem. Chem. Phys. 2020, 22, 17482–17493. 10.1039/D0CP01508C. [DOI] [PubMed] [Google Scholar]
- Safin D. A.; Babashkina M. G.; Robeyns K.; Garcia Y. C–H···Br–C vs. C–Br···Br–C vs. C–Br···N Bonding in Molecular Self-Assembly of Pyridine-Containing Dyes. RSC Adv. 2016, 6, 53669–53678. 10.1039/C6RA10094E. [DOI] [Google Scholar]
- Bosch E. Role of Sp-C–H---N Hydrogen Bonding in Crystal Engineering. Cryst. Growth Des. 2010, 10, 3808–3813. 10.1021/cg100707y. [DOI] [Google Scholar]
- Salonen L. M.; Ellermann M.; Diederich F. Aromatic Rings in Chemical and Biological Recognition: Energetics and Structures. Angew. Chem., Int. Ed. 2011, 50, 4808–4842. 10.1002/anie.201007560. [DOI] [PubMed] [Google Scholar]
- Nishio M. The CH/π Hydrogen Bond in Chemistry. Conformation, Supramolecules, Optical Resolution and Interactions Involving Carbohydrates. Phys. Chem. Chem. Phys. 2011, 13, 13873–13900. 10.1039/C1CP20404A. [DOI] [PubMed] [Google Scholar]
- Bader R. F. W.Atoms in Molecules: A Quantum Theory; Oxford University Press: USA, 1994. [Google Scholar]
- Saeed A.; Erben M. F.; Bolte M. Synthesis, Structural and Vibrational Properties of 1-(Adamantane-1-Carbonyl)-3-Halophenyl Thioureas. Spectrochim. Acta, Part A 2013, 102, 408–413. 10.1016/j.saa.2012.10.043. [DOI] [PubMed] [Google Scholar]
- Saeed A.; Flörke U.; Erben M. F. The Role of Substituents in the Molecular and Crystal Structure of 1-(Adamantane-1-Carbonyl)-3-(Mono)- and 3,3-(Di) Substituted Thioureas. J. Mol. Struct. 2014, 1065–1066, 150–159. 10.1016/j.molstruc.2014.03.002. [DOI] [Google Scholar]
- Seth S. K.; Bauzá A.; Mahmoudi G.; Stilinović V.; López-Torres E.; Zaragoza G.; Keramidas A. D.; Frontera A. On the Importance of Pb···X (X = O, N, S, Br) Tetrel Bonding Interactions in a Series of Tetra- and Hexa-Coordinated Pb(Ii) Compounds. CrystEngComm 2018, 20, 5033–5044. 10.1039/C8CE00919H. [DOI] [Google Scholar]
- Bauzá A.; Seth S. K.; Frontera A. Tetrel Bonding Interactions at Work: Impact on Tin and Lead Coordination Compounds. Coord. Chem. Rev. 2019, 384, 107–125. 10.1016/j.ccr.2019.01.003. [DOI] [Google Scholar]
- Mahmudov K. T.; Kopylovich M. N.; Guedes da Silva M. F. C.; Pombeiro A. J. L. Chalcogen Bonding in Synthesis, Catalysis and Design of Materials. Dalton Trans. 2017, 46, 10121–10138. 10.1039/C7DT01685A. [DOI] [PubMed] [Google Scholar]
- Khan I.; Panini P.; Khan S. U.-D.; Rana U. A.; Andleeb H.; Chopra D.; Hameed S.; Simpson J. Exploiting the Role of Molecular Electrostatic Potential, Deformation Density, Topology, and Energetics in the Characterization of S···N and Cl···N Supramolecular Motifs in Crystalline Triazolothiadiazoles. Cryst. Growth Des. 2016, 16, 1371–1386. 10.1021/acs.cgd.5b01499. [DOI] [Google Scholar]
- Pinheiro P. d. S. M.; Rodrigues D. A.; Alves M. A.; Tinoco L. W.; Ferreira G. B.; de Sant’Anna C. M. R.; Fraga C. A. M. Theoretical and Experimental Characterization of 1,4-N···S σ-Hole Intramolecular Interactions in Bioactive N-Acylhydrazone Derivatives. New J. Chem. 2018, 42, 497–505. 10.1039/C7NJ03543H. [DOI] [Google Scholar]
- Aakeroy C. B.; Bryce D. L.; Desiraju G. R.; Frontera A.; Legon A. C.; Nicotra F.; Rissanen K.; Scheiner S.; Terraneo G.; Metrangolo P.; Resnati G. Definition of the Chalcogen Bond (IUPAC Recommendations 2019). Pure Appl. Chem. 2019, 91, 1889–1892. 10.1515/pac-2018-0713. [DOI] [Google Scholar]
- Saeed A.; Bolte M.; Erben M. F.; Pérez H. Intermolecular Interactions in Crystalline 1-(Adamantane-1-Carbonyl)-3-Substituted Thioureas with Hirshfeld Surface Analysis. CrystEngComm 2015, 17, 7551–7563. 10.1039/C5CE01373A. [DOI] [Google Scholar]
- Espinosa E.; Molins E.; Lecomte C. Hydrogen Bond Strengths Revealed by Topological Analyses of Experimentally Observed Electron Densities. Chem. Phys. Lett. 1998, 285, 170–173. 10.1016/S0009-2614(98)00036-0. [DOI] [Google Scholar]
- Gatti C. Chemical Bonding in Crystals: New Directions. Z. Kristallogr.—Cryst. Mater. 2005, 220, 399–457. 10.1524/zkri.220.5.399.65073. [DOI] [Google Scholar]
- Koch U.; Popelier P. L. A. Characterization of C-H-O Hydrogen Bonds on the Basis of the Charge Density. J. Phys. Chem. 1995, 99, 9747–9754. 10.1021/j100024a016. [DOI] [Google Scholar]
- Mosmann T. Rapid Colorimetric Assay for Cellular Growth and Survival: Application to Proliferation and Cytotoxicity Assays. J. Immunol. Methods 1983, 65, 55–63. 10.1016/0022-1759(83)90303-4. [DOI] [PubMed] [Google Scholar]
- Berridge M. V.; Tan A. S. Characterization of the Cellular Reduction of 3-(4,5-Dimethylthiazol-2-yl)-2,5-Diphenyltetrazolium Bromide (MTT): Subcellular Localization, Substrate Dependence, and Involvement of Mitochondrial Electron Transport in MTT Reduction. Arch. Biochem. Biophys. 1993, 303, 474–482. 10.1006/abbi.1993.1311. [DOI] [PubMed] [Google Scholar]
- Tacar O.; Sriamornsak P.; Dass C. R. Doxorubicin: An Update on Anticancer Molecular Action, Toxicity and Novel Drug Delivery Systems. J. Pharm. Pharmacol. 2012, 65, 157–170. 10.1111/j.2042-7158.2012.01567.x. [DOI] [PubMed] [Google Scholar]
- Burla M. C.; Caliandro R.; Camalli M.; Carrozzini B.; Cascarano G. L.; Giacovazzo C.; Mallamo M.; Mazzone A.; Polidori G.; Spagna R. SIR2011: A New Package for Crystal Structure Determination and Refinement. J. Appl. Crystallogr. 2012, 45, 357–361. 10.1107/S0021889812001124. [DOI] [Google Scholar]
- Sheldrick G. M. Crystal Structure Refinement with SHELXL. Acta Crystallogr., Sect. C: Struct. Chem. 2015, 71, 3–8. 10.1107/S2053229614024218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spek A. L. Structure Validation in Chemical Crystallography. Acta Crystallogr., Sect. D: Struct. Biol. 2009, 65, 148–155. 10.1107/S090744490804362X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spek A. L. CheckCIF Validation ALERTS: What They Mean and How to Respond. Acta Crystallogr., Sect. E: Struct. Rep. Online 2020, 76, 1–11. 10.1107/S2056989019016244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macrae C. F.; Sovago I.; Cottrell S. J.; Galek P. T. A.; McCabe P.; Pidcock E.; Platings M.; Shields G. P.; Stevens J. S.; Towler M.; Wood P. A. Mercury 4.0: From Visualization to Analysis, Design and Prediction. J. Appl. Crystallogr. 2020, 53, 226–235. 10.1107/S1600576719014092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spackman P. R.; Turner M. J.; McKinnon J. J.; Wolff S. K.; Grimwood D. J.; Jayatilaka D.; Spackman M. A. CrystalExplorer. A Program for Hirshfeld Surface Analysis, Visualization and Quantitative Analysis of Molecular Crystals. J. Appl. Crystallogr. 2021, 54, 1006–1011. 10.1107/S1600576721002910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gavezzotti A. Efficient Computer Modeling of Organic Materials. The Atom-Atom, Coulomb-London-Pauli (AA-CLP) Model for Intermolecular Electrostatic-Polarization, Dispersion and Repulsion Energies. New J. Chem. 2011, 35, 1360–1368. 10.1039/C0NJ00982B. [DOI] [Google Scholar]
- Gavezzotti A. Calculation of Lattice Energies of Organic Crystals: The PIXEL Integration Method in Comparison with More Traditional Methods. Z. Kristallogr.—Cryst. Mater. 2005, 220, 499–510. 10.1524/zkri.220.5.499.65063. [DOI] [Google Scholar]
- Frisch M. J.; Trucks G. W.; Schlegel H. B.; Scuseria G. E.; Robb M. A.; Cheeseman J. R.; Scalmani G.; Barone V.; Mennucci B.; Petersson G. A.; Nakatsuji H.; Caricato M.; Li X.; Hratchian H. P.; Izmaylov A. F.; Bloino J.; Zheng G.; Sonnenberg J. L.; Hada M.; Ehara M.; Toyota K.; Fukuda R.; Hasegawa J.; Ishida M.; Nakajima T.; Honda Y.; Kitao O.; Nakai H.; Vreven T.; Montgomery J. A. Jr.; Peralta J. E.; Ogliaro F.; Bearpark M. J.; Heyd J.; Brothers E. N.; Kudin K. N.; Staroverov V. N.; Kobayashi R.; Normand J.; Raghavachari K.; Rendell A. P.; Burant J. C.; Iyengar S. S.; Tomasi J.; Cossi M.; Rega N.; Millam N. J.; Klene M.; Knox J. E.; Cross J. B.; Bakken V.; Adamo C.; Jaramillo J.; Gomperts R.; Stratmann R. E.; Yazyev O.; Austin A. J.; Cammi R.; Pomelli C.; Ochterski J. W.; Martin R. L.; Morokuma K.; Zakrzewski V. G.; Voth G. A.; Salvador P.; Dannenberg J. J.; Dapprich S.; Daniels A. D.; Farkas Ö.; Foresman J. B.; Ortiz J. V.; Cioslowski J.; Fox D. J.. Gaussian 09, Revision D.01; Gaussian, Inc.: Wallingford, CT, USA, 2013.
- Becke A. D. Density-Functional Thermochemistry. V. Systematic Optimization of Exchange-Correlation Functionals. J. Chem. Phys. 1997, 107, 8554–8560. 10.1063/1.475007. [DOI] [Google Scholar]
- Schmider H. L.; Becke A. D. Optimized Density Functionals from the Extended G2 Test Set. J. Chem. Phys. 1998, 108, 9624–9631. 10.1063/1.476438. [DOI] [Google Scholar]
- Grimme S.; Antony J.; Ehrlich S.; Krieg H. A Consistent and Accurate Ab Initio Parametrization of Density Functional Dispersion Correction (DFT-D) for the 94 Elements H-Pu. J. Chem. Phys. 2010, 132, 154104. 10.1063/1.3382344. [DOI] [PubMed] [Google Scholar]
- Boys S. F.; Bernardi F. The Calculation of Small Molecular Interactions by the Differences of Separate Total Energies. Some Procedures with Reduced Errors. Mol. Phys. 1970, 19, 553–566. 10.1080/00268977000101561. [DOI] [Google Scholar]
- Keith T. A.AIMAll, Ver. 19.02.13; TK Gristmill Software: Overland Park, KS, 2019.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.