Abstract
Histologic grading has been a simple and inexpensive method to assess tumor behavior and prognosis of invasive breast cancer grading, thereby identifying patients at risk for adverse outcomes, who may be eligible for (neo)adjuvant therapies. Histologic grading needs to be performed accurately, on properly fixed specimens, and by adequately trained dedicated pathologists that take the time to diligently follow the protocol methodology. In this paper, we review the history of histologic grading, describe the basics of grading, review prognostic value and reproducibility issues, compare performance of grading to gene expression profiles, and discuss how to move forward to improve reproducibility of grading by training, feedback and artificial intelligence algorithms, and special stains to better recognize mitoses. We conclude that histologic grading, when adequately carried out, remains to be of important prognostic value in breast cancer patients.
Keywords: Breast, Carcinoma, Histology, Grading
History of histologic grading
The importance of the histologic profile of invasive breast cancer in correlation with the disease course was first acknowledged by Von Hansemann in 1893. He presumed that tumors with a loss of differentiation, which he so-called anaplastic, had a greater tendency to metastasize, which he confirmed in 1902. In 1922, MacCarthy and Sistrunk described a correlation between post-mastectomy survival and the degree of differentiation, lymphocytic infiltration, and hyalinization of the tumor [1]. In 1925, Greenough was the first to describe a grading classification system, that, similar to the current classification, separated tumors into three grades of malignancy, based on tubular differentiation, the size of cells/nuclei, and hyperchromatism and mitosis [2]. Several other studies, which also took clinical staging into account, followed. Importantly, it was concluded that histologic grading was of value with regard to prognosis, yet, clinical staging was the most important factor [3].
Until the late 1950s, tumors were simply and only classified according their clinical stage, which does not take into account the accepted range of biological behavior of breast carcinomas. Bloom and Richardson observed at that time that clinical staging provided a useful guide, yet “it fails completely to indicate the likelihood of occult lymphatic and blood-born metastases being present in what appears to be an early case, nor the speed with which such metastases may develop” [4]. This prompted them to develop a technique of histologic grading, which they correlated with survival in a series of 1544 breast cancer patients [4]. In their classification system, tumors were allocated a score of 1–3 for each of three components, differentiation of tubule formation, pleomorphism, and “hyperchromatosis” or mitotic nuclei. A total score, derived from the summation of the three component scores, of 3–5 indicated a low-grade (I) tumor, scores 6–7 an intermediate (II) tumor, and scores of 8–9 a high-grade (III) tumor [4]. Importantly, Bloom and Richardson stated that the different grades were not different pathologic entities, and their 3 grades were based on arbitrarily divisions of a continuous scale of malignancy. They did not claim to have discovered a mathematically accurate grading classification, yet they emphasized that their point system was merely a useful aid in guiding prognosis [4]. Despite these compelling observations, histologic grading of breast cancer was not accepted as a routine procedure, mainly due to perceived reproducibility issues, until decades later [5].
The grading classification of Bloom and Richardson [4] was revised by Elston and Ellis in 1991, who use semiquantitative criteria to improve objectivity and reproducibility [5]. Tubular differentiation was based on evaluation of the percentage of tubule formation, hyperchromatic figures were excluded from assessment, and mitosis was counted using a defined field area. The degree of nuclear pleomorphism was scored according to more objective definitions based on comparison with normal cell types. Elston and Ellis demonstrated the relevance of histological grade in breast cancer and its strong correlation with prognosis, in a series of 1813 patients with primary operable disease who had been followed up for many years [5]. This Elston–Ellis modification of the Bloom and Richardson grading classification (also known as the Nottingham grading system (NGS)) has become globally used to guide management of invasive breast carcinoma [6–8].
Histologic grading: the basics
Histologic grade represents the degree of differentiation, which reflects the resemblance of tumor cells to normal breast cells. The NGS is a semiquantitative assessment of three morphological characteristics, being tubule/gland formation, nuclear pleomorphism, and mitotic frequency. The NGS is a simple and cheap method, which in principle can be performed in all breast cancer cases [9]. Furthermore, it merely requires appropriately prepared hematoxylin–eosin (HE)-stained tumor slides from optimally formalin-fixed paraffin-embedded tissue blocks by trained experienced pathologists, who are prepared to take the time to diligently follow the standard protocol.
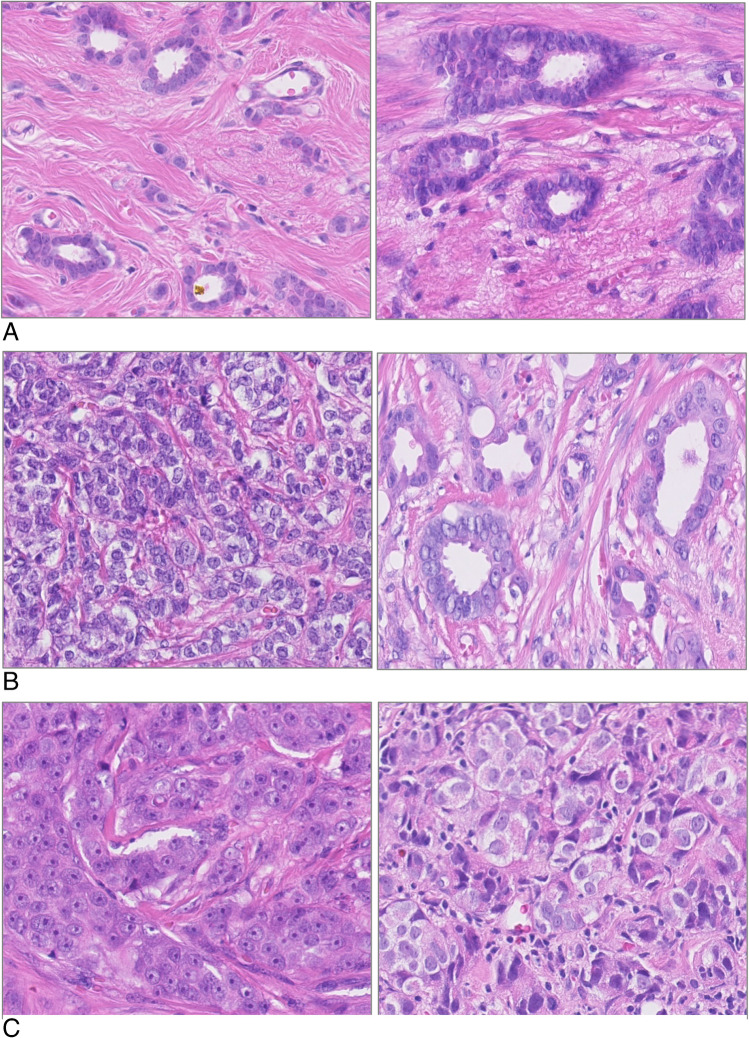
Grading itself is evaluated by a numerical scoring system of 1–3 per category (tubule formation, nuclear pleomorphism, mitotic count). Tubule/gland formation is classified according to the percentage of tubular or glandular acinar spaces (> 75% score 1, 10–75% score 2, < 10% score 3), where only those structures with clear central lumina enclosed by polarized cells are counted (Fig. 1A–C). The inside-out polarization features of micropapillary invasive carcinoma does not by itself count as tubule formation, although these cancers can have tubule formation on the inside of the micropapillary groups that counts as tubules.
Fig. 1.
A Tubular differentiation score 1. B Tubular differentiation score 2. C Tubular differentiation score 3
Nuclear pleomorphism, describing the size and degree of variation in tumor cell nuclear size and shape, is scored in the least differentiated area of the tumor. It is assessed by examining the regularity of nuclear size and compared to the shape of normal epithelial cells in the surrounding tissue. Score 1 is allocated to tumor cells that are similar in size to normal epithelial cells, which show only minimal pleomorphism and whose nucleoli and chromatin pattern are inconspicuous at most (Fig. 2A). Tumor cells with nuclei that are 1.5–2 × larger than epithelial cells and with moderate pleomorphism and still inconspicuous nucleoli are given score 2 (Fig. 2B). In contrast, score 3 nuclei are more than 2 × larger in size, which vary considerably in size and which show vesicular chromatin and often prominent nucleoli (Fig. 2C).
Fig. 2.
A Nuclear pleomorphism score 1. B Nuclear pleomorphism score 2. C Nuclear pleomorphism score 3
Mitotic counting is performed in the most proliferative area of the tumor, usually the most solid area, typically at the periphery of the tumor. A score of 1–3 is based on the number of defined mitotic figures seen in a given tumor area or microscope field area, with cutoff points dependent on field area size assessed using the diameter of the high-power-field (HPF) (Table 1). Examples of well-defined mitotic figures can be found in Fig. 3.
Table 1.
Score thresholds for mitotic counts
| Field | Field | Mitotic count (score) | ||
|---|---|---|---|---|
| Diameter (mm) | Area (mm2) | 1 | 2 | 3 |
| 0.40 | 0.126 | ≤ 4 | 5–9 | ≥ 10 |
| 0.41 | 0.123 | ≤ 4 | 5–9 | ≥ 10 |
| 0.42 | 0.138 | ≤ 5 | 6–10 | ≥ 11 |
| 0.43 | 0.145 | ≤ 5 | 6–10 | ≥ 11 |
| 0.44 | 0.152 | ≤ 5 | 6–11 | ≥ 12 |
| 0.45 | 0.159 | ≤ 5 | 6–11 | ≥ 12 |
| 0.46 | 0.166 | ≤ 6 | 7–12 | ≥ 13 |
| 0.47 | 0.173 | ≤ 6 | 7–12 | ≥ 13 |
| 0.48 | 0.181 | ≤ 6 | 7–13 | ≥ 14 |
| 0.49 | 0.188 | ≤ 6 | 7–13 | ≥ 14 |
| 0.50 | 0.196 | ≤ 7 | 8–14 | ≥ 15 |
| 0.51 | 0.204 | ≤ 7 | 8–14 | ≥ 15 |
| 0.52 | 0.212 | ≤ 7 | 8–15 | ≥ 16 |
| 0.53 | 0.221 | ≤ 8 | 9–16 | ≥ 17 |
| 0.54 | 0.229 | ≤ 8 | 9–16 | ≥ 17 |
| 0.55 | 0.237 | ≤ 8 | 9–17 | ≥ 18 |
| 0.56 | 0.246 | ≤ 8 | 9–17 | ≥ 18 |
| 0.57 | 0.255 | ≤ 9 | 10–18 | ≥ 19 |
| 0.58 | 0.264 | ≤ 9 | 10–19 | ≥ 20 |
| 0.59 | 0.273 | ≤ 9 | 10–19 | ≥ 20 |
| 0.60 | 0.283 | ≤ 10 | 11–20 | ≥ 21 |
| 0.61 | 0.292 | ≤ 10 | 11–21 | ≥ 22 |
| 0.62 | 0.302 | ≤ 11 | 12–22 | ≥ 23 |
| 0.63 | 0.312 | ≤ 11 | 12–22 | ≥ 23 |
| 0.64 | 0.322 | ≤ 11 | 12–23 | ≥ 24 |
| 0.65 | 0.332 | ≤ 12 | 13–24 | ≥ 25 |
| 0.66 | 0.342 | ≤ 12 | 13–24 | ≥ 25 |
| 0.67 | 0.352 | ≤ 12 | 13–25 | ≥ 26 |
| 0.68 | 0.363 | ≤ 13 | 14–26 | ≥ 27 |
| 0.69 | 0.374 | ≤ 13 | 14–27 | ≥ 28 |
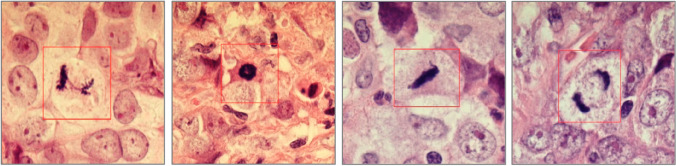
Fig. 3.
Mitotic figures
Overall, the three grade component values are summated, resulting in a total score of between 3 and 9 and then categorized into final grade. Scores 3–5 represent well-differentiated grade I tumors, scores 6–7 represent moderately differentiated grade II tumors, and scores 8–9 represent poorly differentiated grade III tumors.
Histologic grading: prognostic value
The NGS has shown to be of independent significance with regard to breast cancer prognosis [5, 7, 10–12]. In addition, histologic grade has been incorporated in prognostic index scores, of which the Nottingham prognostic index (NPI) [13] is regarded as the only index score that has been extensively validated and which retains its predicting ability in most independent populations [14, 15]. Within the NPI, histologic grade is combined with lymph node (LN) status and tumor size, where grade is considered equally important as lymph node status. In contrast, studies have suggested that histologic grade predicts tumor behavior more accurately than tumor size, which may be considered a more “time-dependent” factor [7, 8, 11, 13, 16].
Breast cancer is now detected at earlier stages by mammographic screening programs, thereby resulting in a greater proportion of both smaller [17, 18] and lymph node–negative tumors at diagnosis [19]. This furthermore increases the clinical contribution of histologic grade [7, 20].
The prognostic significance of histologic grading has been widely studied [5, 7, 10–12, 16, 21–29]. Henson et al. [29] included a spectacularly large number of 22,616 breast cancer cases. They showed similar prognosis for breast cancer patients with stage II/grade I disease and breast cancer patients with stage I/grade II disease [29]. Furthermore, they showed an excellent prognosis for small (< 2 cm) grade I tumors, even when they showed lymph node metastases at presentation. Therefore, Henderson et al. concluded that using histologic grade in conjunction with disease stage (consisting of tumor size and lymph node status) could improve outcome predictions [29].
These results were further supported by a somewhat more recent, retrospective series of 2219 operable breast cancer cases from a single institution by Rakha et al. [7]. Histologic grade proved to be associated with both breast cancer-specific and disease-free survival in the whole series, as well as within the specific subgroups of small tumors (T1a, T1b, T1c) and lymph node–negative and lymph node–positive tumors [7]. The latter has also been shown in other studies [7, 22, 24, 28, 30]. More importantly, the prognostic value of histologic grade was independent of tumor size and lymph node status [7]. Furthermore, it was shown that grade is complimentary and equivalent in impact magnitude to lymph node status, which is widely regarded as a major prognostic factor in breast cancer. For example, patients with grade II tumors and 1–3 positive lymph nodes had a better prognosis than patients with grade III tumors without any lymph node metastases [7]. Moreover, the Swedish two-county trial demonstrated that the independent prognostic effect of histologic grading on survival (as well as lymph node status and tumor size) is long-lasting (> 10 years) [26]. Of note, although grading was initially deemed applicable to Not Otherwise Specified (“ductal”) cancers, grading has been proven prognostically important across all histologic breast cancer types.
In addition, the prognostic role of histologic grading in specific subgroups, for whom the benefit of adjuvant chemotherapy is uncertain, like patients with low-volume lymph node metastases, or patients with ER-positive/lymph node–negative breast cancer, has also been established. For example, histologic grade is an independent prognostic factor in breast cancer patients with ER-positive disease, with [31] or without neoadjuvant endocrine therapy [32]. Furthermore, histologic grade has been shown to be one of the two remaining prognostic factors that was associated with relapse-free survival in a multivariate analyses of ER-positive/HER2-negative breast cancer patients [33].
As to the relative prognostic contribution of the three constituents of grade, several studies have shown that the mitotic count is the most important variable followed by nuclear atypia and then tubule formation [34–36].
Thus, as grade is an important and long-term prognostic factor across breast cancer subtypes, being equally important as lymph node status, more important than tumor size, and being of specific prognostic influence in different subgroups, it would be an omission to exclude histologic grade from clinical decision-making. Fortunately, histologic grade is currently widely incorporated in clinical breast cancer guidelines such as ASCO [37–39], NCCN [40], ESMO [41], UK NICE [42] based on PREDICT [43], and the St. Gallen Expert Panel [44].
Histologic grading: reproducibility issues
Although histologic grade has long known to be of prognostic value, its reproducibility has also been the subject of debate for decades. Firstly, the distribution of grade varies largely (i.e., up to 27%) between studies (Table 2) [7, 16, 22–24, 26, 27, 30, 35, 45–56]. However, these differences may partly be explained by the wide variety of patient cohorts (Table 2). For example, these cohorts vary in age, type of detection method (screening versus symptomatic, early, or advanced breast cancer), and type of tissue fixation.
Table 2.
Distribution of histologic grades in different invasive breast cancer studies
| Study | Number | Grade 1 | Grade 2 | Grade 3 |
|---|---|---|---|---|
| Elston, 1984 [45] | 625 | 17% | 37% | 46% |
| Davis et al., 1986 [46] | 1537 | 22% | 49% | 29% |
| Hopton et al., 1989 [47] | 874 | 29% | 46% | 25% |
| Le Doussal, et al., 1989 [35] | 1262 | 11% | 45% | 46% |
| Balslev et al., 1994 [48] | 9149 | 32% | 49% | 19% |
| Saimura et al., 1999 [22] | 741 | 19% | 37% | 44% |
| Reed et al., 2000 [30] | 613 | 25% | 41% | 35% |
| Simpson et al., 2000 [24] | 368 | 22% | 45% | 33% |
| Lundin et al., 2001 [23] | 1554 | 26% | 47% | 27% |
| Frkovic-Grazio and Bracko, 2002 [16] | 270 | 38% | 38% | 24% |
| Warwick et al., 2004 [26] | 1988 | 23% | 37% | 40% |
| Williams et al., 2006 [49] | 1058 | 20% | 46% | 34% |
| Rakha et al., 2008 [7] | 2219 | 18% | 36% | 46% |
| Thomas et al., 2009 [50] | 1650 | 26% | 45% | 29% |
| Blamey et al., 2010 [27] | 16,944 | 29% | 41% | 30% |
| Puig-Vives et al., 2013 [51] | 2122 | 20% | 47% | 33% |
| Seneviratne et al., 2015 [52] | 2146 | 25% | 52% | 23% |
| Sun et al., 2015 [53] | 1259 | 18% | 62% | 20% |
| Moller et al., 2016 [54] | 81,427 | 16% | 52% | 32% |
| Van Dooijeweert et al., 2019 [55] | 33,792 | 28% | 48% | 24% |
| Van Dooijeweert et al., 2020 [56] | 17,102 | 31% | 49% | 19% |
Secondly, inter- and intra-observer variation has been extensively reported, with a wide range Kappa values (0.43 and 0.85) which correlates to a range from “fair” to “almost perfect” agreement [9, 12, 25, 30, 57–69] (Table 3). A recent nationwide study in the Netherlands showed substantial variation in grading in daily clinical practice, both between pathology laboratories and between pathologists within individual laboratories [55]. Importantly, these differences were not explained by differences in case mix [55]. Subsequently, initiatives were launched to improve variation in grading. Feedback reports in which pathologists and laboratories were benchmarked against the nationwide average and their colleagues (all anonymized) were sent [56], and pathologists and residents were trained using e-learning [70]. Both initiatives resulted in a promising decrease in grading variation and may be implemented broadly in the field of pathology. Yet, it remains important to emphasize what Bloom and Richardson rightfully stated in 1957. The different grades are not different entities, they are not a fact, nor the truth, but they are (statistically) computed cutoff values in a spectrum of histopathologic features on a continuous scale of malignancy. With the clinical implications in mind [20], educating clinicians (like medical oncologists, radiotherapists, surgeons), who use histologic grade as an absolute cutoff, may be equally important to the improvement of histologic grading by pathologists. Furthermore, in the current era of shared decision-making, patients should perhaps be involved as well.
Table 3.
Inter- and intra-observer reproducibility studies on grading of invasive breast cancer
| Study | Cases | Readers | Inter-observer variation | Intra-observer variation |
|---|---|---|---|---|
| Theissig et al., 1990 [12] | 166 | 3 | Complete agreement 72.3%. Kappa 0.57 | - |
| Robbins et al., 1995 [57] | 50 | 5 | Complete agreement 83.3%. Kappa 0.73 | - |
| Frierson et al., 1995 [58] | 75 | 6 | Kappa 0.43 to 0.74 | - |
| Jacquemier et al., 1998 [59] | 24 | 21 | Complete agreement 69%. Kappa 0.53 | - |
| Sikka et al., 1999 [60] | 40 | 3 | Kappa 0.68 to 0.83 | - |
| Anderson et al., 2000 [25] | 52 | 2 | Kappa 0.54 | - |
| Boiesen et al., 2000 [9] | 93 | 7 | Kappa 0.54 | - |
| Reed et al., 2000 [30] | 613 | 2 | Kappa 0.69 | - |
| Page et al., 2001 [61] | 425 | 2 | Complete agreement 76% | - |
| Meyer et al., 2005 [62] | 7 | 49 | Kappa 0.50–0.59 | - |
| Chowdhury et al., 2006 [63] | 50 | 5 | Mean polychoric correlation 0.8 | - |
| Longacre et al., 2006 [64] | 35 | 13 | Kappa 0.5 to 0.7 | - |
| Ellis et al., 2006 [65] | 12 | 600 | Kappa 0.45 to 0.53 (figures after application of guidelines) | - |
| Bueno-de-Mesquita et al., 2010 [66] | 694 | 2 | Kappa 0.56 | |
| Postma et al., 2013 [67] | 310 | 2 | Kappa 0.80 | |
| Rabe et al., 2019 [68] | 100 | 6 | Kappa 0.58–0.85 | Mean Kappa 0.77 |
| Ginter et al., 2020 [69] | 143 | 6 | Kappa 0.50 | - |
Histologic grading: the way forward
Immunohistochemistry for proliferation markers
With the proliferation marker mitotic index being the most important constituent of grade, with observer variation in counting [34–36], one could argue that replacing mitotic index with a potentially more objective method based on immunohistochemical staining of proteins highlighting proliferating cells could help to reduce variation. Ki67 and phosphohistone H3 (PHH3) have shown most promise here. Ki67 is a protein expressed in all phases of the cell cycle, except for resting cells in the G0 phase. However, there is controversy with regard to its clinical utility in routine clinical management, due to variation in analytical practice [71–73] and the absence of consensus on cutoff values [74]. PHH3 is highly expressed in cells in the mitotic phase, and this proliferation marker has shown great promise, also with regard to reproducibility [75]. It may help to better identify mitotic cells and highlight the areas of highest proliferation and thereby increase reproducibility, especially in cases with sub-optimal fixation associated compromised morphology. More research on its clinical utility is necessary [75].
Another potentially important tool is the IHC-4 algorithm, which consists of a combination of ER-, PR-, HER2-, and Ki-67 status [76]. However, to date, this system has not been widely incorporated in breast cancer guidelines, mainly due to the above-mentioned lack of consensus on the clinical utility of Ki67-assessment.
Molecular profiling and gene expression profiling
In the past decade, molecular profiling and gene expression profiling (GEP) have emerged as new tools to predict tumor behavior. Molecular profiling studies showed that grade I, II, and III breast cancers are most likely different entities as they show specific molecular profiles at immunohistochemical, genomic, and transcriptomic levels, which further supports the relevance of histologic grading [77]. Furthermore, histologic grading has been shown to better correspond to the molecular profile of breast cancer than lymph node status and tumor size [78–80].
Although these “new” biomarker/molecular profile methods were launched with great excitement, it seems unlikely that molecular or gene expression profiling will substitute classic clinicopathologic variables. For ER-positive disease, for example, histologic grade remains an independent prognostic factor in multivariate models, even when molecular signatures are included [81, 82]. In addition, several studies have shown that the added value of GEPs to clinicopathologic variables (age, ER-status, lymph node status) in prognostic models may be limited and sometimes only equal to prognostic indices like the NPI [31, 83, 84].
Two well-known GEPs, Oncotype DX [81] and MammaPrint [85], are currently being used in daily clinical practice in some countries. However, it is also important to acknowledge that these tests are not accessible (i.e., $4.000 per Oncotype DX/per MammaPrint) nor applicable to every breast cancer patient [86–94]. Even more so, studies have shown that “simple” biomarkers like histologic grade and progesterone status can predict Oncotype DX scores, thereby saving the need for these expensive GEPs [95-100]. However, GEPs may be of added value in patients for whom the indication for adjuvant therapy remains doubtful based on classic biomarkers [85]. Lastly, molecular signatures and GEPs are not without flaws themselves, although they are generally considered to be more objective biomarkers [82]. For example, in the MammaPrint study, a change in the RNA extraction solution that was used to calculate the MammaPrint score led to a shift of genomic risk scores in > 150 patients [85]. In addition, similar to statistical cut-offs used for histologic grading, molecular tests and GEPs also depend on biostatistical approaches. Furthermore, intratumor heterogeneity has been found to affect the prognostic risk stratification by GEPs in early breast cancer [101]. Lastly, the results of GEPs and molecular profiles also depend on well-prepared tissue samples to begin with. Overall, molecular profiling and GEPs should not be seen as the new “holy grail” and will not substitute but rather complement classic clinicopathologic biomarkers, which in return need to be assessed adequately, by well-trained pathologists.
Artificial intelligence
Artificial intelligence methodology is currently finding considerable traction as a tool to aide pathologists. It is expected to be especially helpful with regard to reproducibility concerns that surround histologic grading in its current state. An example of this is the CAMELYON 16 challenge, which showed that some deep-learning algorithms achieved better diagnostic performance in detecting lymph node metastases in breast cancer patients than routine pathologists (under time pressure) and comparable diagnostic performance to expert pathologists (without any time constraints) [102]. Since then, promising results have been published, for example, on predicting tumor proliferation in breast cancer by deep-learning (TUPAC16 challenge) [103] and mitosis counting [104]. However, practical utility studies need to be performed [102, 103]. In addition, it is important to acknowledge that well-annotated (consensus based) datasets are required for the development of AI algorithms. A major pathology-led consortium with 46 partners from all fields of research and businesses, which aims to create a platform of Whole Slide Imaging (WSI) data to develop advanced AI algorithms (BIGPICTURE), may be very helpful here and will start in the near future. These advanced AI algorithms may be able to directly grade breast carcinoma themselves [101].
Conclusions
In conclusion, histologic grading is a simple and inexpensive method to assess tumor behavior and patient prognosis, thereby identifying patients at risk for adverse outcomes, who may be eligible for (neo)adjuvant therapies. However, histologic grading needs to be performed accurately, on properly fixed specimens, and by adequately trained dedicated pathologists that take the time to diligently follow the protocol methodology. Levels of inter-observer variation can and should still be improved. Feedback and training may be helpful tools to support this. In addition, artificial intelligence is very likely to be able to support pathologists in the near future. When accessible to patients, GEPs may complement classic pathology biomarkers in doubtful cases, rather than substitute them. Furthermore, the GEPs are costly and have flaws of their own. Hence, histologic grading, when adequately carried out, remains to be of important prognostic value in breast cancer patients.
Author contributions
This was an invited review agreed on the concept and jointly wrote the paper.
Availability of data and material
Not applicable.
Code availability
Not applicable.
Declarations
Ethics approval
Not applicable, no use of data or tissue, neither human participants, nor animals.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Sistrunk WE, MacCarty WC. Life expectancy following radical amputation for carcinoma of the breast: a clinical and pathologic study of 218 cases. Ann Surg. 1922;75(1):61–69. [PMC free article] [PubMed] [Google Scholar]
- 2.Greenough RB. Varying degrees of malignancy in cancer of the breast. Can Res. 1925;9(4):453–463. [Google Scholar]
- 3.Wolff B. Histological grading in carcinoma of the breast. Br J Cancer. 1966;20(1):36–40. doi: 10.1038/bjc.1966.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bloom HJ, Richardson WW. Histological grading and prognosis in breast cancer: a study of 1409 cases of which 359 have been followed for 15 years. Br J Cancer. 1957;11(3):359–377. doi: 10.1038/bjc.1957.43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 1991;19(5):403–410. doi: 10.1111/j.1365-2559.1991.tb00229.x. [DOI] [PubMed] [Google Scholar]
- 6.Curigliano G, Burstein HJ, Winer EP, et al. De-escalating and escalating treatments for early-stage breast cancer: the St. Gallen International Expert Consensus Conference on the Primary Therapy of Early Breast Cancer 2017. Ann Oncol. 2017;28(8):1700–1712. doi: 10.1093/annonc/mdx308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Rakha EA, El-Sayed ME, Lee AH, et al. Prognostic significance of Nottingham histologic grade in invasive breast carcinoma. J Clin Oncol. 2008;26(19):3153–3158. doi: 10.1200/JCO.2007.15.5986. [DOI] [PubMed] [Google Scholar]
- 8.Rakha EA, Reis-Filho JS, Baehner F, et al. Breast cancer prognostic classification in the molecular era: the role of histological grade. Breast Cancer Res. 2010;12(4):207. doi: 10.1186/bcr2607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Boiesen P, Bendahl PO, Anagnostaki L, et al. Histologic grading in breast cancer--reproducibility between seven pathologic departments. South Sweden Breast Cancer Group. Acta Oncol. 2000;39(1):41–45. doi: 10.1080/028418600430950. [DOI] [PubMed] [Google Scholar]
- 10.Pereira H, Pinder SE, Sibbering DM, et al. Pathological prognostic factors in breast cancer. IV: Should you be a typer or a grader? A comparative study of two histological prognostic features in operable breast carcinoma. Histopathology. 1995;27(3):219–226. doi: 10.1111/j.1365-2559.1995.tb00213.x. [DOI] [PubMed] [Google Scholar]
- 11.Sundquist M, Thorstenson S, Brudin L, et al. Applying the Nottingham prognostic index to a Swedish breast cancer population. South East Swedish Breast Cancer Study Group. Breast Cancer Res Treat. 1999;53(1):1–8. doi: 10.1023/a:1006052115874. [DOI] [PubMed] [Google Scholar]
- 12.Theissig F, Kunze KD, Haroske G, et al. Histological grading of breast cancer. Interobserver, reproducibility and prognostic significance. Pathol Res Pract. 1990;186(6):732–736. doi: 10.1016/S0344-0338(11)80263-3. [DOI] [PubMed] [Google Scholar]
- 13.Galea MH, Blamey RW, Elston CE, et al. The Nottingham prognostic index in primary breast cancer. Breast Cancer Res Treat. 1992;22(3):207–219. doi: 10.1007/BF01840834. [DOI] [PubMed] [Google Scholar]
- 14.Phung MT, Tin Tin S, Elwood JM. Prognostic models for breast cancer: a systematic review. BMC Cancer. 2019;19(1):230. doi: 10.1186/s12885-019-5442-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Clark GM. Do we really need prognostic factors for breast cancer? Breast Cancer Res Treat. 1994;30(2):117–126. doi: 10.1007/BF00666054. [DOI] [PubMed] [Google Scholar]
- 16.Frkovic-Grazio S and Bracko M (2002) Long term prognostic value of Nottingham histological grade and its components in early (pT1N0M0) breast carcinoma. J Clin Pathol 55(2):88-92. 10.1136/jcp.55.2.88 [Right doi not to be found in PubMed at this moment] [DOI] [PMC free article] [PubMed]
- 17.Anttinen J, Kautiainen H, Kuopio T. Role of mammography screening as a predictor of survival in postmenopausal breast cancer patients. Br J Cancer. 2006;94(1):147–151. doi: 10.1038/sj.bjc.6602895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shen Y, Yang Y, Inoue LY, et al. Role of detection method in predicting breast cancer survival: analysis of randomized screening trials. J Natl Cancer Inst. 2005;97(16):1195–1203. doi: 10.1093/jnci/dji239. [DOI] [PubMed] [Google Scholar]
- 19.Hanrahan EO, Valero V, Gonzalez-Angulo AM, Hortobagyi GN, et al. Prognosis and management of patients with node-negative invasive breast carcinoma that is 1 cm or smaller in size (stage 1; T1a, bN0M0): a review of the literature. J Clin Oncol. 2006;24(13):2113–2122. doi: 10.1200/JCO.2005.02.8035. [DOI] [PubMed] [Google Scholar]
- 20.van Dooijeweert C, Baas IO, Deckers IAG, et al (2021) The increasing importance of histologic grading in tailoring adjuvant systemic therapy in 30,843 breast cancer patients. Breast Cancer Res Treat. 10.1007/s10549-021-06098-7 [DOI] [PMC free article] [PubMed]
- 21.Kollias J, Murphy CA, Elston CW, et al. The prognosis of small primary breast cancers. Eur J Cancer. 1999;35(6):908–912. doi: 10.1016/s0959-8049(99)00056-8. [DOI] [PubMed] [Google Scholar]
- 22.Saimura M, Fukutomi T, Tsuda H et al (1999) Prognosis of a series of 763 consecutive node-negative invasive breast cancer patients without adjuvant therapy: analysis of clinicopathological prognostic factor. J Surg Oncol 71(2):101–105. 10.1002/(sici)1096-9098(199906)71:2%3c101::aid-jso8%3e3.0.co;2-g [DOI] [PubMed]
- 23.Lundin J, Lundin M, Holli K, et al. Omission of histologic grading from clinical decision making may result in overuse of adjuvant therapies in breast cancer: results from a nationwide study. J Clin Oncol. 2001;19(1):28–36. doi: 10.1200/JCO.2001.19.1.28. [DOI] [PubMed] [Google Scholar]
- 24.Simpson JF, Gray R, Dressler LG, et al. Prognostic value of histologic grade and proliferative activity in axillary node-positive breast cancer: results from the Eastern Cooperative Oncology Group Companion Study, EST 4189. J Clin Oncol. 2000;18(10):2059–2069. doi: 10.1200/JCO.2000.18.10.2059. [DOI] [PubMed] [Google Scholar]
- 25.Anderson TJ, Alexander FE, Lamb J, et al. Pathology characteristics that optimize outcome prediction of a breast screening trial. Br J Cancer. 2000;83(4):487–492. doi: 10.1054/bjoc.2000.1286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Warwick J, Tabàr L, Vitak B, et al. Time-dependent effects on survival in breast carcinoma: results of 20 years of follow-up from the Swedish two-county study. Cancer. 2004;100(7):1331–1336. doi: 10.1002/cncr.20140. [DOI] [PubMed] [Google Scholar]
- 27.Blamey RW, Hornmark-Stenstam B, Ball G, et al. ONCOPOOL - a European database for 16,944 cases of breast cancer. Eur J Cancer. 2010;46(1):56–71. doi: 10.1016/j.ejca.2009.09.009. [DOI] [PubMed] [Google Scholar]
- 28.Mirza AN, Mirza NQ, Vlastos G, et al. Prognostic factors in node-negative breast cancer: a review of studies with sample size more than 200 and follow-up more than 5 years. Ann Surg. 2002;235(1):10–26. doi: 10.1097/00000658-200201000-00003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Henson DE, Ries L, Freedman S et al (1991) Relationship among outcome, stage of disease, and histologic grade for 22,616 cases of breast cancer. The basis for a prognostic index. Cancer 68(10):2142–2149. 10.1002/1097-0142(19911115)68:10%3c2142::aid-cncr2820681010%3e3.0.co;2-d [DOI] [PubMed]
- 30.Reed W, Hannisdal E, Boehler PJ et al (2000) The prognostic value of p53 and c-erb B-2 immunostaining is overrated for patients with lymph node negative breast carcinoma: a multivariate analysis of prognostic factors in 613 patients with a follow-up of 14–30 years. Cancer 88(4):804–813. 10.1002/(sici)1097-0142(20000215)88:4%3c804::aid-cncr11%3e3.0.co;2-y [DOI] [PubMed]
- 31.Wirapati P, Sotiriou C, Kunkel S, et al. Meta-analysis of gene expression profiles in breast cancer: toward a unified understanding of breast cancer subtyping and prognosis signatures. Breast Cancer Res. 2008;10(4):R65. doi: 10.1186/bcr2124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ellis MJ, Tao Y, Luo J, et al. Outcome prediction for estrogen receptor-positive breast cancer based on postneoadjuvant endocrine therapy tumor characteristics. J Natl Cancer Inst. 2008;100(19):1380–1388. doi: 10.1093/jnci/djn309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Desmedt C, Haibe-Kains B, Wirapati P, et al. Biological processes associated with breast cancer clinical outcome depend on the molecular subtypes. Clin Cancer Res. 2008;14(16):5158–5165. doi: 10.1158/1078-0432.CCR-07-4756. [DOI] [PubMed] [Google Scholar]
- 34.Genestie C, Zafrani B, Asselain B, et al. Comparison of the prognostic value of Scarff-Bloom-Richardson and Nottingham histological grades in a series of 825 cases of breast cancer: major importance of the mitotic count as a component of both grading systems. Anticancer Res. 1998;18(1b):571–576. [PubMed] [Google Scholar]
- 35.Le Doussal V, Tubiana-Hulin M, Friedman S et al (1989) Prognostic value of histologic grade nuclear components of Scarff-Bloom-Richardson (SBR). An improved score modification based on a multivariate analysis of 1262 invasive ductal breast carcinomas. Cancer 64(9):1914–1921. 10.1002/1097-0142(19891101)64:9%3c1914::aid-cncr2820640926%3e3.0.co;2-g [DOI] [PubMed]
- 36.Baak JP, van Diest PJ, Voorhorst FJ, et al. Prospective multicenter validation of the independent prognostic value of the mitotic activity index in lymph node-negative breast cancer patients younger than 55 years. J Clin Oncol. 2005;23(25):5993–6001. doi: 10.1200/JCO.2005.05.511. [DOI] [PubMed] [Google Scholar]
- 37.Andre F, Ismaila N, Henry NL, et al. Use of biomarkers to guide decisions on adjuvant systemic therapy for women with early-stage invasive breast cancer: ASCO clinical practice guideline update-integration of results from TAILORx. J Clin Oncol. 2019;37(22):1956–1964. doi: 10.1200/JCO.19.00945. [DOI] [PubMed] [Google Scholar]
- 38.Henry NL, Somerfield MR, Abramson VG, et al. Role of patient and disease factors in adjuvant systemic therapy decision making for early-stage, operable breast cancer: American Society of Clinical Oncology Endorsement of Cancer Care Ontario Guideline Recommendations. J Clin Oncol. 2016;34(19):2303–2311. doi: 10.1200/JCO.2015.65.8609. [DOI] [PubMed] [Google Scholar]
- 39.Henry NL, Somerfield MR, Abramson VG, et al. Role of patient and disease factors in adjuvant systemic therapy decision making for early-stage, operable breast cancer: update of the ASCO endorsement of the Cancer Care Ontario Guideline. J Clin Oncol. 2019;37(22):1965–1977. doi: 10.1200/JCO.19.00948. [DOI] [PubMed] [Google Scholar]
- 40.Goetz MP, Gradishar WJ, Anderson BO. NCCN Guidelines Insights: Breast Cancer, Version 3.2018. 2019. J Natl Compr Canc Netw. 2019;17(2):118–126. doi: 10.6004/jnccn.2019.0009. [DOI] [PubMed] [Google Scholar]
- 41.Cardoso F, Kyriakides S, Ohno S, et al. Early breast cancer: ESMO clinical practice guidelines for diagnosis, treatment and follow-up†. Ann Oncol. 2019;30(8):1194–1220. doi: 10.1093/annonc/mdz173. [DOI] [PubMed] [Google Scholar]
- 42.(NICE) N.I.f.H.a.C.E. NICE guideline [NG101]- Early and locally advanced breast cancer: diagnosis and management. 18 July 2018 Available from: https://www.nice.org.uk/guidance/ng101
- 43.Cambridge Breast Unit (2021) U.o.C.D.o.O.a.t.U.s.E.C.I.a.R.C.E. Predict breast cancer. Available from: https://breast.predict.nhs.uk/
- 44.Burstein HJ, Curigliano G, Loibl S, et al. Estimating the benefits of therapy for early stage breast cancer: the St Gallen International Consensus Guidelines for the primary therapy of early breast cancer 2019. Ann Oncol. 2019;30(10):1541–1557. doi: 10.1093/annonc/mdz235. [DOI] [PubMed] [Google Scholar]
- 45.Elston CW. The assessment of histological differentiation in breast cancer. Aust N Z J Surg. 1984;54(1):11–15. doi: 10.1111/j.1445-2197.1984.tb06677.x. [DOI] [PubMed] [Google Scholar]
- 46.Davis BW, Gelber RD, Goldhirsch A et al (1986) Prognostic significance of tumor grade in clinical trials of adjuvant therapy for breast cancer with axillary lymph node metastasis. Cancer 58(12):2662–2670. 10.1002/1097-0142(19861215)58:12%3c2662::aid-cncr2820581219%3e3.0.co;2-y [DOI] [PubMed]
- 47.Hopton DS, Thorogood J, Clayden AD, et al. Observer variation in histological grading of breast cancer. Eur J Surg Oncol. 1989;15(1):21–23. [PubMed] [Google Scholar]
- 48.Balslev I, Axelsson CK, Zedeler K, et al. The Nottingham prognostic index applied to 9,149 patients from the studies of the Danish Breast Cancer Cooperative Group (DBCG) Breast Cancer Res Treat. 1994;32(3):281–290. doi: 10.1007/BF00666005. [DOI] [PubMed] [Google Scholar]
- 49.Williams C, Brunskill S, Altman D et al (2006) Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy. Health Technol Assess 10(34):iii–iv, ix–xi, 1–204. 10.3310/hta10340 [DOI] [PubMed]
- 50.Thomas JS, Kerr GR, Jack WJ, et al. Histological grading of invasive breast carcinoma–a simplification of existing methods in a large conservation series with long-term follow-up. Histopathology. 2009;55(6):724–731. doi: 10.1111/j.1365-2559.2009.03429.x. [DOI] [PubMed] [Google Scholar]
- 51.Puig-Vives M, Sánchez MJ, Sánchez-Cantalejo J, et al. Distribution and prognosis of molecular breast cancer subtypes defined by immunohistochemical biomarkers in a Spanish population-based study. Gynecol Oncol. 2013;130(3):609–614. doi: 10.1016/j.ygyno.2013.05.039. [DOI] [PubMed] [Google Scholar]
- 52.Seneviratne S, Lawrenson R, Scott N, et al. Breast cancer biology and ethnic disparities in breast cancer mortality in New Zealand: a cohort study. PLoS ONE. 2015;10(4):e0123523. doi: 10.1371/journal.pone.0123523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Sun J, Chen C, Wei W, et al. Associations and indications of Ki67 expression with clinicopathological parameters and molecular subtypes in invasive breast cancer: a population-based study. Oncol Lett. 2015;10(3):1741–1748. doi: 10.3892/ol.2015.3461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Møller H, Henson K, Lüchtenborg M. Short-term breast cancer survival in relation to ethnicity, stage, grade and receptor status: national cohort study in England. Br J Cancer. 2016;115(11):1408–1415. doi: 10.1038/bjc.2016.335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.van Dooijeweert C, van Diest PJ, Willems SM, et al. Significant inter- and intra-laboratory variation in grading of invasive breast cancer: a nationwide study of 33,043 patients in the Netherlands. Int J Cancer. 2019 doi: 10.1002/ijc.32330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.van Dooijeweert C, van Diest PJ, Baas IO, et al (2020) Variation in breast cancer grading: the effect of creating awareness through laboratory-specific and pathologist-specific feedback reports in 16 734 patients with breast cancer. J Clin Pathol jclinpath-2019-206362. 10.1136/jclinpath-2019-206362 [DOI] [PubMed]
- 57.Robbins P, Pinder S, de Klerk N. Histological grading of breast carcinomas: a study of interobserver agreement. Hum Pathol. 1995;26(8):873–879. doi: 10.1016/0046-8177(95)90010-1. [DOI] [PubMed] [Google Scholar]
- 58.Frierson HF, Jr, Wolber RA, Berean KW, et al. Interobserver reproducibility of the Nottingham modification of the Bloom and Richardson histologic grading scheme for infiltrating ductal carcinoma. Am J Clin Pathol. 1995;103(2):195–198. doi: 10.1093/ajcp/103.2.195. [DOI] [PubMed] [Google Scholar]
- 59.Jacquemier J and Charpin C (1998) [Reproducibility of histoprognostic grades of invasive breast cancer]. Ann Pathol 18(5):385-90. Article in French [PubMed]
- 60.Sikka M, Agarwal S, Bhatia A. Interobserver agreement of the Nottingham histologic grading scheme for infiltrating duct carcinoma breast. Indian J Cancer. 1999;36(2–4):149–153. [PubMed] [Google Scholar]
- 61.Page DL, Gray R, Allred DC, et al. Prediction of node-negative breast cancer outcome by histologic grading and S-phase analysis by flow cytometry: an Eastern Cooperative Oncology Group Study (2192) Am J Clin Oncol. 2001;24(1):10–18. doi: 10.1097/00000421-200102000-00002. [DOI] [PubMed] [Google Scholar]
- 62.Meyer JS, Alvarez C, Milikowski C, et al. Breast carcinoma malignancy grading by Bloom-Richardson system vs proliferation index: reproducibility of grade and advantages of proliferation index. Mod Pathol. 2005;18(8):1067–1078. doi: 10.1038/modpathol.3800388. [DOI] [PubMed] [Google Scholar]
- 63.Chowdhury N, Pai MR, Lobo FD, et al. Interobserver variation in breast cancer grading: a statistical modeling approach. Anal Quant Cytol Histol. 2006;28(4):213–218. [PubMed] [Google Scholar]
- 64.Longacre TA, Ennis M, Quenneville LA, Bane AL, et al. Interobserver agreement and reproducibility in classification of invasive breast carcinoma: an NCI breast cancer family registry study. Mod Pathol. 2006;19(2):195–207. doi: 10.1038/modpathol.3800496. [DOI] [PubMed] [Google Scholar]
- 65.Ellis IO, Coleman D, Wells C, et al. Impact of a national external quality assessment scheme for breast pathology in the UK. J Clin Pathol. 2006;59(2):138–145. doi: 10.1136/jcp.2004.025551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Bueno-de-Mesquita JM, Nuyten DS, Wesseling J, et al. The impact of inter-observer variation in pathological assessment of node-negative breast cancer on clinical risk assessment and patient selection for adjuvant systemic treatment. Ann Oncol. 2010;21(1):40–47. doi: 10.1093/annonc/mdp273. [DOI] [PubMed] [Google Scholar]
- 67.Postma EL, Verkooijen HM, van Diest PJ et al (2013) Discrepancy between routine and expert pathologists’ assessment of non-palpable breast cancer and its impact on locoregional and systemic treatment. Eur J Pharmacol 717(1–3):31–35. 10.1016/j.ejphar.2012.12.033 [DOI] [PubMed]
- 68.Rabe K, Snir OL, Bossuyt V, et al. Interobserver variability in breast carcinoma grading results in prognostic stage differences. Hum Pathol. 2019;94:51–57. doi: 10.1016/j.humpath.2019.09.006. [DOI] [PubMed] [Google Scholar]
- 69.Ginter PS, Idress R, D’Alfonso TM, et al (2020) Histologic grading of breast carcinoma: a multi-institution study of interobserver variation using virtual microscopy. Mod Pathol. 10.1038/s41379-020-00698-2 [DOI] [PMC free article] [PubMed]
- 70.van Dooijeweert C, Deckers IAG, de Ruiter EJ, et al. The effect of an e-learning module on grading variation of (pre)malignant breast lesions. Mod Pathol. 2020 doi: 10.1038/s41379-020-0556-6. [DOI] [PubMed] [Google Scholar]
- 71.Focke CM, Decker T, van Diest PJ. Intratumoral heterogeneity of Ki67 expression in early breast cancers exceeds variability between individual tumours. Histopathology. 2016;69(5):849–861. doi: 10.1111/his.13007. [DOI] [PubMed] [Google Scholar]
- 72.Focke CM, van Diest PJ, Decker T. St Gallen 2015 subtyping of luminal breast cancers: impact of different Ki67-based proliferation assessment methods. Breast Cancer Res Treat. 2016;159(2):257–263. doi: 10.1007/s10549-016-3950-5. [DOI] [PubMed] [Google Scholar]
- 73.Focke CM, Bürger H, van Diest PJ, et al. Interlaboratory variability of Ki67 staining in breast cancer. Eur J Cancer. 2017;84:219–227. doi: 10.1016/j.ejca.2017.07.041. [DOI] [PubMed] [Google Scholar]
- 74.Penault-Llorca F, Radosevic-Robin N. Ki67 assessment in breast cancer: an update. Pathology. 2017;49(2):166–171. doi: 10.1016/j.pathol.2016.11.006. [DOI] [PubMed] [Google Scholar]
- 75.van Steenhoven JEC, Kuijer A, Kornegoor R, et al. Assessment of tumour proliferation by use of the mitotic activity index, and Ki67 and phosphohistone H3 expression, in early-stage luminal breast cancer. Histopathology. 2020;77(4):579–587. doi: 10.1111/his.14185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Cuzick J, Dowsett M, Pineda S, et al. Prognostic value of a combined estrogen receptor, progesterone receptor, Ki-67, and human epidermal growth factor receptor 2 immunohistochemical score and comparison with the Genomic Health recurrence score in early breast cancer. J Clin Oncol. 2011;29(32):4273–4278. doi: 10.1200/JCO.2010.31.2835. [DOI] [PubMed] [Google Scholar]
- 77.Natrajan R, Lambros MB, Geyer FC, et al. Loss of 16q in high grade breast cancer is associated with estrogen receptor status: evidence for progression in tumors with a luminal phenotype? Genes Chromosomes Cancer. 2009;48(4):351–365. doi: 10.1002/gcc.20646. [DOI] [PubMed] [Google Scholar]
- 78.Yu K, Lee CH, Tan PH, et al. A molecular signature of the Nottingham prognostic index in breast cancer. Cancer Res. 2004;64(9):2962–2968. doi: 10.1158/0008-5472.can-03-2430. [DOI] [PubMed] [Google Scholar]
- 79.Lu X, Lu X, Wang ZC, et al. Predicting features of breast cancer with gene expression patterns. Breast Cancer Res Treat. 2008;108(2):191–201. doi: 10.1007/s10549-007-9596-6. [DOI] [PubMed] [Google Scholar]
- 80.Groenendijk FH, Jager A, Cardoso F, et al. A nationwide registry-based cohort study of the MammaPrint genomic risk classifier in invasive breast cancer. Breast. 2018;38:125–131. doi: 10.1016/j.breast.2017.12.015. [DOI] [PubMed] [Google Scholar]
- 81.Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004;351(27):2817–2826. doi: 10.1056/NEJMoa041588. [DOI] [PubMed] [Google Scholar]
- 82.Weigelt B, Baehner FL, Reis-Filho JS. The contribution of gene expression profiling to breast cancer classification, prognostication and prediction: a retrospective of the last decade. J Pathol. 2010;220(2):263–280. doi: 10.1002/path.2648. [DOI] [PubMed] [Google Scholar]
- 83.Dunkler D, Michiels S, Schemper M. Gene expression profiling: does it add predictive accuracy to clinical characteristics in cancer prognosis? Eur J Cancer. 2007;43(4):745–751. doi: 10.1016/j.ejca.2006.11.018. [DOI] [PubMed] [Google Scholar]
- 84.Edén P, Ritz C, Rose C et al (2004) “Good Old” clinical markers have similar power in breast cancer prognosis as microarray gene expression profilers. Eur J Cancer 40(12):1837–1841. 10.1016/j.ejca.2004.02.025 [DOI] [PubMed]
- 85.Cardoso F, van’t Veer LJ, Bogaerts J, et al. 70-gene signature as an aid to treatment decisions in early-stage breast cancer. New England J Med. 2016;375(8):717–729. doi: 10.1056/NEJMoa1602253. [DOI] [PubMed] [Google Scholar]
- 86.Jasem J, Amini A, Rabinovitch R, et al. 21-gene recurrence score assay as a predictor of adjuvant chemotherapy administration for early-stage breast cancer: an analysis of use, therapeutic implications, and disparity profile. J Clin Oncol. 2016;34(17):1995–2002. doi: 10.1200/JCO.2015.65.0887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Orucevic A, Heidel RE, Bell JL. Utilization and impact of 21-gene recurrence score assay for breast cancer in clinical practice across the United States: lessons learned from the 2010 to 2012 National Cancer Data Base analysis. Breast Cancer Res Treat. 2016;157(3):427–435. doi: 10.1007/s10549-016-3833-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Roberts MC, Kurian AW, Petkov VI. Uptake of the 21-gene assay among women with node-positive, hormone receptor-positive breast cancer. J Natl Compr Canc Netw. 2019;17(6):662–668. doi: 10.6004/jnccn.2018.7266. [DOI] [PubMed] [Google Scholar]
- 89.Davis BA, Aminawung JA, Abu-Khalaf MM, et al. Racial and ethnic disparities in oncotype DX test receipt in a statewide population-based study. J Natl Compr Canc Netw. 2017;15(3):346–354. doi: 10.6004/jnccn.2017.0034. [DOI] [PubMed] [Google Scholar]
- 90.Konecny G, Pauletti G, Pegram M, et al Quantitative association between HER-2/neu and steroid hormone receptors in hormone receptor-positive primary breast cancer. (0027–8874 (Print)). 10.1093/jnci/95.2.142 [DOI] [PubMed]
- 91.Press DJ, Ibraheem A, Dolan ME, Goss KH, et al. Racial disparities in omission of oncotype DX but no racial disparities in chemotherapy receipt following completed oncotype DX test results. Breast Cancer Res Treat. 2018;168(1):207–220. doi: 10.1007/s10549-017-4587-8. [DOI] [PubMed] [Google Scholar]
- 92.Weldon CB, Trosman JR, Gradishar WJ, et al. Barriers to the use of personalized medicine in breast cancer. Journal of Oncology Practice. 2012;8(4):e24–e31. doi: 10.1200/JOP.2011.000448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Ricks-Santi LJ, McDonald JT. Low utility of oncotype DX(R) in the clinic. Cancer Med. 2017;6(3):501–507. doi: 10.1002/cam4.837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Zhang L, Hsieh MC, Petkov V, et al. Trend and survival benefit of oncotype DX use among female hormone receptor-positive breast cancer patients in 17 SEER registries, 2004–2015. Breast Cancer Res Treat. 2020;180(2):491–501. doi: 10.1007/s10549-020-05557-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Allison KH, Kandalaft PL, Sitlani CM, et al. Routine pathologic parameters can predict oncotype DXTM recurrence scores in subsets of ER positive patients: who does not always need testing? Breast Cancer Res Treat. 2012;131(2):413–424. doi: 10.1007/s10549-011-1416-3. [DOI] [PubMed] [Google Scholar]
- 96.Huang JL, Kizy S, Marmor S, et al. Tumor grade and progesterone receptor status predict 21-gene recurrence score in early stage invasive breast carcinoma. Breast Cancer Res Treat. 2018;172(3):671–677. doi: 10.1007/s10549-018-4955-z. [DOI] [PubMed] [Google Scholar]
- 97.Orucevic A, Bell JL, King M, et al. Nomogram update based on TAILORx clinical trial results - oncotype DX breast cancer recurrence score can be predicted using clinicopathologic data. Breast. 2019;46:116–125. doi: 10.1016/j.breast.2019.05.006. [DOI] [PubMed] [Google Scholar]
- 98.McNabb AP, Heidel RE, Orucevic A, et al (2019) Breast 46:116–125. 10.1016/j.breast.2019.05.006 [DOI] [PubMed]
- 99.Wu SG, Zhang WW, Wang J, et al. Progesterone receptor status and tumor grade predict the 21-gene recurrence score of invasive lobular breast cancer. Biomark Med. 2019;13(12):1005–1012. doi: 10.2217/bmm-2019-0209. [DOI] [PubMed] [Google Scholar]
- 100.Yoo SH, Kim TY, Kim M, et al (2019) Development of a nomogram to predict the recurrence score of 21-gene prediction assay in hormone receptor–positive early breast cancer. Clin Breast Cancer. 10.1016/j.clbc.2019.07.010 [DOI] [PubMed]
- 101.Gyanchandani R, Lin Y, Lin HM, et al. Intratumor heterogeneity affects gene expression profile test prognostic risk stratification in early breast cancer. Clin Cancer Res. 2016;22(21):5362–5369. doi: 10.1158/1078-0432.CCR-15-2889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.EhteshamiBejnordi B, Veta M, Johannes van Diest P, et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA. 2017;318(22):2199–2210. doi: 10.1001/jama.2017.14585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Veta M, Heng YJ, Stathonikos N, et al. Predicting breast tumor proliferation from whole-slide images: The TUPAC16 challenge. Med Image Anal. 2019;54:111–121. doi: 10.1016/j.media.2019.02.012. [DOI] [PubMed] [Google Scholar]
- 104.Veta M, van Diest PJ, Jiwa M, et al. Mitosis counting in breast cancer: object-level interobserver agreement and comparison to an automatic method. PLoS ONE. 2016;11(8):e0161286. doi: 10.1371/journal.pone.0161286. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.
Not applicable.