Abstract
A Staphylococcus aureus norA disruption mutant was created by allelic replacement. Exposure of this mutant to norfloxacin produced SA K1748, a derivative with raised fluoroquinolone MICs, found to be the result of a grlA mutation, and raised organic cation MICs. Ethidium and enoxacin uptake was identical in SA K1748 and its parent, but pre-exposure of SA K1748 to organic cations caused a reduction in ethidium uptake as a result of increased efflux. Altered ethidium uptake and efflux, as well as increased MICs of other organic cations, suggest that SA K1748 possesses a non-NorA multidrug efflux transporter that is inducible by its substrates.
NorA is a proton motive force (PMF)-dependent multidrug (MDR) efflux pump in Staphylococcus aureus (8). It is a member of the major facilitator superfamily and has 12 transmembrane-spanning segments (6, 8, 14, 17). Hydrophilic fluoroquinolones and monocationic organic compounds such as acriflavine, ethidium, and tetraphenylphosphonium bromide (TPP) are substrates of this pump (9). Interestingly, data available from the S. aureus genome sequencing projects suggest that NorA is not the only S. aureus MDR transporter; at least 10 regions encoding polypeptides having homology with NorA can be identified (1; data not shown). In this paper, we report the isolation of a mutant of S. aureus having a phenotype consistent with the expression of a non-NorA MDR efflux transporter.
PCR was employed to amplify norA from SA 8325-4, a standard laboratory strain of S. aureus, using primers designed so that the tetracycline resistance cassette of pT181 (tet) could be inserted within the gene (10, 15). To allow replication in S. aureus, pTS2, a temperature-sensitive S. aureus plasmid, was then introduced into the construct (4, 5). Allelic replacement in SA 8325-4 was carried out as described previously, producing a norA-disrupted derivative (SA K1712) (4; G. W. Kaatz, O. Hartford, L. O'Brien, S. M. Seo, M. Wahiduzzaman, and T. J. Foster, Abstr. 39th Intersci. Conf. Antimicrob. Agents Chemother., abstr. C810, p. 105, 1999).
MIC testing in the presence and absence of reserpine (20 μg/ml) was performed using a microdilution technique and cation-adjusted Mueller-Hinton II broth (CAMHB; BBL Microbiology Systems, Cockeysville, Md.) (12). Results were expressed as the geometric means of multiple (at least six) determinations.
SA K1712 was grown to an optical density at 660 nm (OD660) of ∼1 in CAMHB. Cells were concentrated and then exposed to norfloxacin incorporated into Mueller-Hinton agar at 10 times the MIC. After 48 h of incubation, surviving colonies, which appeared at a frequency of 7.4 × 10−9, were replica plated onto Mueller-Hinton agar containing ethidium at five times the MIC for SA K1712. Eight percent (3 of 37) of the norfloxacin-resistant strains were also ethidium resistant, and one (SA K1748) was chosen for further study.
The nucleotide sequence of the quinolone resistance-determining region of the grlA and gyrA genes of the study strains and that of the region of norA into which tet was inserted in SA K1712 were determined as described previously (2, 8, 18, 19).
Uptake studies were performed using whole cells. For [14C]enoxacin (Parke-Davis Pharmaceutical Research, Ann Arbor, Mich.; specific activity, 15.9 μCi/mg), a radiometric technique was employed as described elsewhere (9). Ethidium uptake was determined by using a modification of a fluorescence technique (13). Organisms were grown overnight in CAMHB with or without TPP at a concentration of 0.25 times the MIC of the strain being evaluated. A 4% (vol/vol) inoculum of this culture was added to 50 ml of the same medium as used for overnight growth. Cells were harvested at an OD660 of 0.7 to 0.8, washed in ice-cold CAMHB, and then resuspended in CAMHB to a final OD660 of 0.4 (∼1 × 109 CFU/ml). The suspension was warmed to 37°C, and ethidium (final concentration, 10 μg/ml) was added. The suspension was maintained at 37°C with agitation, and the fluorescence of aliquots was determined at frequent intervals (excitation wavelength, 530 nm; emission wavelength, 600 nm). The effect of carbonyl cyanide m-chlorophenylhydrazone (CCCP; final concentration, 100 μM), which disrupts the PMF, on drug uptake also was assessed.
For ethidium efflux, cells were grown overnight in CAMHB containing no additive or 0.25 times the MIC of acriflavine, benzalkonium chloride, ethidium, TPP, or norfloxacin. Organisms were diluted into the same medium as used for overnight growth, and at an OD660 of 0.7 to 0.8, ethidium loading of cells was accomplished by the addition of ethidium and CCCP (final concentrations, 10 μg/ml and 100 μM, respectively). After 20 min of incubation at room temperature, the OD660 was adjusted to 0.4 using fresh CAMHB containing ethidium and CCCP. Cells in 1 ml of this dilution were pelleted and then resuspended in fresh CAHMB with or without CCCP (100 μM), and the fluorescence of the suspension was monitored continuously (excitation and emission wavelengths, 530 and 600 nm, respectively).
Sequencing of SA K1712 revealed that tet was inserted into the coding region for the 11th transmembrane-spanning segment (data not shown), which should eliminate NorA function. This was found to be the case, as the MICs of NorA substrates for SA K1712 and a derivative of SA 8325-4 with norA deleted were identical, as were the enoxacin and ethidium uptake profiles (data not shown) (Kaatz et al., 39th ICAAC).
MICs are shown in Table 1. Inactivation of norA resulted in variably reduced MICs of all of the NorA substrates tested. The magnitudes of the MIC reductions were less than those reported by others for norA-disrupted strains (7, 21). The most likely explanation for these results is poor baseline expression of norA in SA 8325-4; experiments in our laboratory have shown this to be the case (data not shown). The minimal effect of norA inactivation on susceptibility to sparfloxacin is not surprising as this compound is known to be a poor NorA substrate (22). Interestingly, disruption of norA did not abolish the effect of reserpine on MICs; in its presence, modest (greater-than-twofold) reductions in the ethidium and benzalkonium chloride MICs were observed when SA K1712 was used. These data are consistent with the activity of one or more reserpine-sensitive non-NorA transporters, potentially MDR in nature, that recognize these compounds in this strain.
TABLE 1.
Susceptibilities of study strains to selected fluoroquinolones and organic cations
| Compound | MIC (μg/ml) in the absence (presence) of reserpinea for:
|
||
|---|---|---|---|
| SA 8325-4 | SA K1712 | SA K1748 | |
| Norfloxacin | 0.81 (0.41) | 0.35 (0.34) | 4.73 (2.80) |
| Enoxacin | 0.94 (0.60) | 0.83 (0.53) | 11.51 (8.84) |
| Ciprofloxacin | 0.30 (0.13) | 0.13 (0.10) | 0.83 (0.63) |
| Sparfloxacin | 0.14 (0.13) | 0.12 (0.13) | 0.24 (0.25) |
| Ethidium bromide | 2.76 (0.29) | 0.88 (0.29) | 10.70 (1.67) |
| Acriflavine | 7.48 (1.56) | 1.82 (1.25) | 3.25 (1.25) |
| Benzalkonium chloride | 1.10 (0.26) | 0.33 (0.11) | 0.64 (0.11) |
| TPP | 11.05 (4.42) | 4.00 (2.63) | 29.92 (6.25) |
Results are expressed as geometric means of multiple determinations.
Considerable increases in norfloxacin, enoxacin, ciprofloxacin, ethidium, and TPP MICs and smaller increases in those of acriflavine and benzalkonium chloride were observed for SA K1748, which was found to have a grlA mutation resulting in a Ser80→Phe substitution. The ethidium, benzalkonium chloride, and TPP MICs for SA K1748 were reduced more than fourfold in the presence of reserpine; a smaller effect, but still a more-than-twofold reduction, was observed for acriflavine. An even smaller effect (a less-than-twofold reduction) on norfloxacin, ciprofloxacin, and enoxacin MICs was observed; changes of this magnitude are typical of strains not overexpressing efflux pumps for these drugs (7, 8, 14). These data are consistent with increased activity of an organic cation pump in SA K1748.
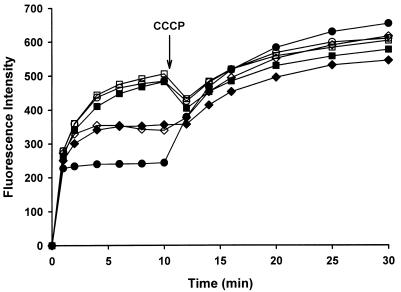
As shown in Fig. 1, disruption of norA resulted in increased ethidium uptake; the same was true for enoxacin (data not shown). No difference in ethidium uptake was observed between SA K1712 and SA K1748 unless the strains were grown in the presence of TPP at 0.25 times the MIC. Such exposure significantly reduced ethidium uptake, but had no effect on enoxacin uptake, in SA K1748 (data not shown). The addition of CCCP ultimately resulted in increased ethidium accumulation in all of the study strains, and this is consistent with disruption of the function of one or more PMF-dependent transporters.
FIG. 1.
Ethidium uptake by study strains. The open and closed symbols represent strains grown in CAMHB and CAMHB containing one-fourth of the appropriate MIC of TPP, respectively. Symbols: ◊ and ⧫, SA 8325-4; □ and ■, SA K1712; ○ and ●, SA K1748. CCCP (final concentration, 100 μM) was added at the indicated time. The data are means of duplicate experiments.
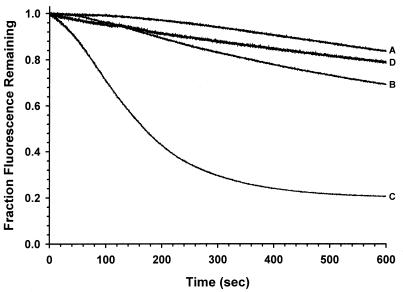
Compared to SA K1748 grown in CAMHB without additives, growth in this medium containing TPP resulted in significantly increased ethidium efflux; CCCP reversed this effect (Fig. 2; 2.5-fold additional efflux compared to the uninduced strain). Growth in the presence of acriflavine, benzalkonium chloride, and ethidium resulted in similarly enhanced efflux (data not shown). Acriflavine and benzalkonium chloride appear to be better inducers than they are substrates, as both compounds significantly increased ethidium efflux despite their MICs being raised only twofold compared to those for the parent strain (Table 1). Somewhat surprisingly, exposure to norfloxacin also augmented ethidium efflux, but by only ∼10% compared to that of the uninduced strain (data not shown).
FIG. 2.
Efflux of ethidium by study strains. A, SA K1712; B, SA K1748; C, SA K1748 induced with TPP; D, SA K1748 induced with TPP and exposed to CCCP (final concentration, 100 μM). The data are means of duplicate experiments.
The grlA mutation identified in SA K1748 explains, at least in part, its increased fluoroquinolone MICs. It has been shown that a single topoisomerase mutation generally results in a modest decrease in fluoroquinolone susceptibility (2, 3). The increases in norfloxacin and enoxacin MICs observed for SA K1748 were considerable and suggest the possibility that additional fluoroquinolone resistance mutations are present in this strain. However, reduced uptake, at least with respect to enoxacin, is not a contributing factor in this resistance.
Our data are consistent with the presence of at least one mutation in SA K1748 that enhances the activity of a non-NorA, CCCP-sensitive MDR efflux system upon exposure to its substrates. Substrate inducibility suggests the presence of a regulatory mutation(s); we have described a similar phenomenon with respect to norA in another S. aureus strain (9). The substrate profile of SA K1748 is reminiscent of that conferred by Qac proteins, which are transporters that recognize selected organic cations (16, 20). All of the qac determinants identified to date are plasmid based; no chromosomal equivalent has been identified.
The fact that norfloxacin exposure supplied the selective pressure for the emergence of the mutation(s) present in SA K1748 is intriguing, as is the fact that it appears to induce (albeit poorly) the MDR efflux system of SA K1748. Fluoroquinolones do have mutagenic potential, and this may explain the emergence of an MDR pump regulatory mutation that may be present in SA K1748 (11). The fact that norfloxacin is not a substrate for the MDR system of SA K1748 but still induces it suggests the possibility that it does so at the level of a global regulator of MDR pump expression. This may not be a direct interaction but may be mediated by stress-responsive proteins. It is necessary to identify the gene(s) involved in the SA K1748 MDR system, as well as those involved in the regulation of this system and that of norA, to investigate this possibility.
Acknowledgments
This work was supported by VA research funds.
REFERENCES
- 1.Altschul S F, Madden T L, Shaffer A A, Zhang J, Zhang Z, Miller W, Lipman D J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ferrero L, Cameron B, Manse B, Lagneaux D, Crouzet J, Famechon A, Blanche F. Cloning and primary structure of Staphylococcus aureus DNA topoisomerase IV: a primary target of fluoroquinolones. Mol Microbiol. 1994;13:641–653. doi: 10.1111/j.1365-2958.1994.tb00458.x. [DOI] [PubMed] [Google Scholar]
- 3.Ferrero L, Cameron B, Crouzet J. Analysis of gyrA and grlA mutations in stepwise-selected ciprofloxacin-resistant mutants of Staphylococcus aureus. Antimicrob Agents Chemother. 1995;39:1554–1558. doi: 10.1128/aac.39.7.1554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Foster T J. Molecular genetic analysis of staphylococcal virulence. Methods Microbiol. 1998;27:433–454. [Google Scholar]
- 5.Greene C, McDevitt D, Francois P, Vaudaux P E, Lew D P, Foster T J. Adhesion properties of mutants of Staphylococcus aureus defective in fibronectin-binding proteins and studies on the expression of fnb genes. Mol Microbiol. 1995;17:1143–1152. doi: 10.1111/j.1365-2958.1995.mmi_17061143.x. [DOI] [PubMed] [Google Scholar]
- 6.Griffith J K, Baker M E, Rouch D A, Page M G P, Skurray R A, Paulsen I T, Chater K F, Baldwin S A, Henderson P F J. Membrane transport proteins: implications of sequence comparisons. Curr Opin Cell Biol. 1992;4:684–695. doi: 10.1016/0955-0674(92)90090-y. [DOI] [PubMed] [Google Scholar]
- 7.Hsieh P-C, Siegel S A, Rogers B, Davis D, Lewis K. Bacteria lacking a multidrug pump: a sensitive tool for drug discovery. Proc Natl Acad Sci USA. 1998;95:6602–6606. doi: 10.1073/pnas.95.12.6602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kaatz G W, Seo S M, Ruble C A. Efflux-mediated fluoroquinolone resistance in Staphylococcus aureus. Antimicrob Agents Chemother. 1993;37:1086–1094. doi: 10.1128/aac.37.5.1086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kaatz G W, Seo S M. Inducible NorA-mediated multidrug resistance in Staphylococcus aureus. Antimicrob Agents Chemother. 1995;39:2650–2655. doi: 10.1128/aac.39.12.2650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Khan S A, Novick R P. Complete nucleotide sequence of pT181, a tetracycline-resistance plasmid from Staphylococcus aureus. Plasmid. 1983;10:251–259. doi: 10.1016/0147-619x(83)90039-2. [DOI] [PubMed] [Google Scholar]
- 11.Mamber S W, Kolek B, Brookshire K W, Bonner D P, Fung-Tomc J. Activity of quinolones in the Ames Salmonella TA102 mutagenicity test and other bacterial genotoxocity assays. Antimicrob Agents Chemother. 1993;37:213–217. doi: 10.1128/aac.37.2.213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.National Committee for Clinical Laboratory Standards. Approved standard M7-A4. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. 4th ed. Villanova, Pa: National Committee for Clinical Laboratory Standards; 1997. [Google Scholar]
- 13.Neyfakh A A, Bidnenko V E, Chen L B. Efflux-mediated multidrug resistance in Bacillus subtilis: similarities and dissimilarities with the mammalian system. Proc Natl Acad Sci USA. 1991;88:4781–4785. doi: 10.1073/pnas.88.11.4781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ng E Y, Trucksis M, Hooper D C. Quinolone resistance mediated by norA: physiologic characterization and relationship to flqB, a quinolone resistance locus on the Staphylococcus aureus chromosome. Antimicrob Agents Chemother. 1994;38:1345–1355. doi: 10.1128/aac.38.6.1345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Novick R P. Properties of a cryptic high-frequency transducing phage in Staphylococcus aureus. Virology. 1963;33:155–166. doi: 10.1016/0042-6822(67)90105-5. [DOI] [PubMed] [Google Scholar]
- 16.Paulsen I T, Brown M H, Dunstan S J, Skurray R A. Molecular characterization of the staphylococcal multidrug resistance export protein QacC. J Bacteriol. 1995;177:2827–2833. doi: 10.1128/jb.177.10.2827-2833.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Paulsen I T, Brown M H, Skurray R A. Proton-dependent multidrug efflux systems. Microbiol Rev. 1996;60:575–608. doi: 10.1128/mr.60.4.575-608.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sanger F, Nicklen S, Coulson A R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA. 1977;74:5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sreedharan S, Oram M, Jensen B, Peterson L R, Fisher L M. DNA gyrase gyrA mutations in ciprofloxacin-resistant strains of Staphylococcus aureus: close similarity with quinolone resistance mutations in Escherichia coli. J Bacteriol. 1990;172:7260–7262. doi: 10.1128/jb.172.12.7260-7262.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tennent J M, Lyon B R, Midgley M, Jones I G, Purewal A S, Skurray R A. Physical and biochemical characterization of the qacA gene encoding antiseptic and disinfectant resistance in Staphylococcus aureus. J Gen Microbiol. 1989;135:1–10. doi: 10.1099/00221287-135-1-1. [DOI] [PubMed] [Google Scholar]
- 21.Yamada H, Kurose-Hamada S, Fukuda Y, Mitsuyama J, Takahata M, Minami S, Wantanabe Y, Narita H. Quinolone susceptibility of norA-disrupted Staphylococcus aureus. Antimicrob Agents Chemother. 1997;41:2308–2309. doi: 10.1128/aac.41.10.2308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yoshida H, Bogaki M, Nakamura S, Ubukata K, Konno M. Nucleotide sequence and characterization of the Staphylococcus aureus norA gene, which confers resistance to quinolones. J Bacteriol. 1990;172:6942–6949. doi: 10.1128/jb.172.12.6942-6949.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]