Dear Editor,
In this Journal, Mensah et al., have demonstrated that the previous infection and vaccination protects from the severe disease post SARS-CoV-2 reinfection1. Although, a history of previous vaccination has been reported to reduce severity of the disease and mortality in COVID-19, it couldn't protect the individuals from getting a breakthrough infection. Many cases of breakthrough infection have been reported across the globe especially with the Delta variant.2 India was also severely affected with SARS-CoV-2 Delta variant during the second wave of pandemic during March-June 2021. Nationwide genomic surveillance of breakthrough cases in India revealed the predominance of Delta variant in causing the infection.2 Recently, enormous rise in SARS-CoV-2 cases was observed worldwide including India with an emergence of highly mutated Omicron variant. Apparently, the surge of reinfection was observed with Omicron as it largely evades the immune response from vaccination and past SARS-CoV-2 infections.3 Many incidences of SARS-CoV-2 reinfection post breakthrough infection have been reported from various countries.4 Here, we report a case with primary SARS-CoV-2 infection, breakthrough infection with Delta and reinfection with Omicron in a healthcare professional from India.
A 38 year health care professional, from Delhi presented with fever, severe bodyache and sore throat on 7th October 2020. On 9th October 2020, his nasopharyngeal and oropharyngeal (NP/OP) samples were found positive for SARS-CoV-2 (TRU-PCR®SARS-CoV-2 KIT, Ct value E gene-22, RdRp-21) by real-time reverse-transcription polymerase chain reaction (rRT-PCR). This was a primary SARS-CoV-2 infection of this case which occurred during the first wave of pandemic. The bodyaches and fever persisted for the next 4 days. Dry cough was present for a week. There were no complaints of breathlessness, vitals and SpO2 were normal. Post recovery, patient experienced symptoms of long COVID including generalized weakness for 2–3 weeks and insomnia for 2–3 months. The patient received first dose of Covishield vaccine (ChAdOx1 nCoV-19 manufactured by Serum Institute of India Limited, Pune) on 31st January 2021 and the second dose on 3rd March 2021.
After the period of over a year, the case developed bodyache and headache on November 2021. Subsequently, he tested positive for SARS-COV-2 by rRT-PCR (Ct value of 25). The bodyache persisted for 2–3 days, but case remained afebrile with no other symptoms. Recovery was uneventful except forgetfulness (“brain fog” of Covid-19). Serum sample was tested for total antibodies against SARS-CoV-2 on 01st December 2021 by VIDAS (VIDAS® SARS-COV-2, bioMérieux SA, France) and the titre was found to be 23 (test value (i) ≥1, positive). The case represented the breakthrough infection post primary SARS-CoV-2 infection and two dose of Covishield vaccination.
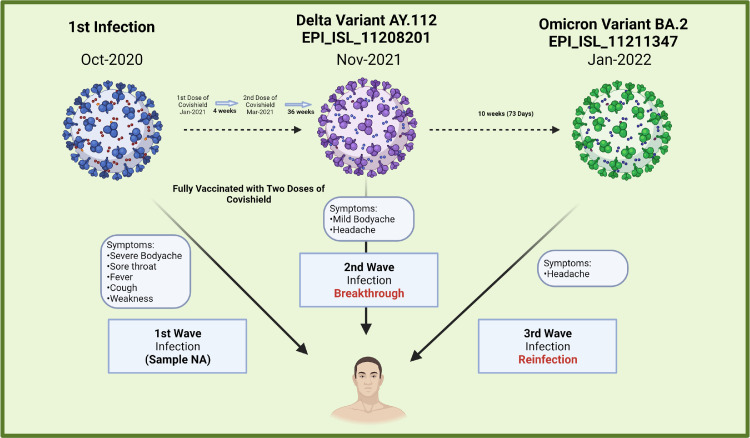
During the third wave of pandemic, the case developed headache on 24th January 2022 after being asymptomatic for a period of 2 months. The NP/OP specimens were again screened for SARS-CoV-2 by rRT-PCR and found to be positive (Ct value of E gene 27 and ORF 26).5 He was in home isolation for 7 days; the recovery was uneventful except for increased forgetfulness post reinfection. The case observed SARS-CoV-2 reinfection post the incidences of primary and breakthrough infection. During these three episodes of COVID-19 infection, the case didn't require hospitalization and received treatment for symptomatic relief under home isolation. The detailed events are depicted in Fig. 1 .
Fig. 1.
Timeline of the first, second and third infection: The clinical presentation of the three episodes, vaccination schedule and GISAID accession IDs of the two sequences are depicted.
The NP/OP specimens of the case collected during the second and third infection were analyzed with next generation sequencing using Illumina sequencing platform. The genomic characterization of the primary infection could not be performed because of the unavailability of clinical specimens. Briefly, RNA library was prepared, the preparation was carried out using the Illumina Coviseq protocol (Illumina®COVIDseq™, Illumina Inc, US). After amplification and purification, QubitTM dsDNA hs assay kit (Invitrogen, USA) was used to quantitate the libraries. Normalization of the libraries was carried out and it was further loaded on NextSeq 500/550 system. The output files were in bcl format, which were converted to fastq. The sequence analysis was done using CLC Genomics workbench version 22(CLC, Qiagen, Germany).6 The retrieved sequences were then mapped using the reference Wuhan sequence NC_045512. The phylogenetic tree was constructed using the MEGA software version 7 and the representative sequences downloaded from the GISAID (https://www.gisaid.org/) database. Variant analysis (50%) of the retrieved sequences was done using the CLC workbench.
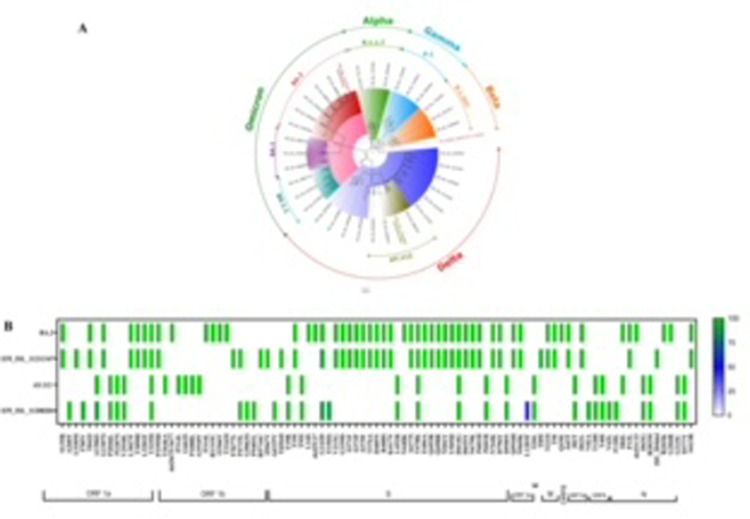
The genomic characterization of the clinical specimens demonstrated the secondary infection with AY.112 variant of Delta lineage (EPI_ISL_11208201) and third infection with BA.2 sub lineage of the Omicron (EPI_ISL_11211347) as per the Phylogenetic Assignment of Named Global Outbreak Lineages (PANGOLIN) system (https://github.com/hCoV-2019/pangolin) (Fig. 2 A). The GISAID clades of the two sequences of second and third infection were GK and GRA. The sequence from breakthrough sample when compared to the characteristic mutations of AY.112 (https://outbreak.info/) had absence of T3646A, P314L, G662S, P1000L and A1918V mutations in the ORF1ab regions and missing S84L mutation in the ORF8 gene (Fig. 2B). Additional mutations A260T, I785V, P4715L, G5063S, P5401L and A6319V in the ORF1ab genes, G142D, E156fs, L1203F in the Spike gene, V82A in ORF6, V26L and F120L in the ORF8 were present.
Fig. 2.
(A) MEGA-7 software was used to construct a neighbor-joining tree using a Tamura 3-parameter model with gamma distribution (0.5) and a bootstrap replication of 1000 cycles. The reference sequences of SARS-CoV-2 VOCs are shown along with the sequences obtained from specimens collected during breakthrough infection (EPI_ISL_11208201) marked with green color and Reinfection (EPI_ISL_11211347) marked with brown color. (B) Single nucleotide variation in the ORF 1a, ORF1 b, S, ORF 3a, E, M, ORF 6, ORF7ab, ORF 8 and N genes and its frequency in the two sequences of Breakthrough (EPI_ISL_11208201) and Reinfection (EPI_ISL_11211347). The X-axis is the SNVs and Y-axis represents the frequency.
The reinfection variant sequence, had characteristic mutations of BA.2 (https://outbreak.info/) except del 3675/3676 in ORF 1a, P314L, R1315C, I1566V and T2163I in ORF1b, L24S, del 25/27, N440K and Q19E in Spike region, S84L in ORF8 and del 31/33, R203K and G204R in Nucleocapsid (N) genes. The additional mutations seen were L530M in ORF 1a, F3677L, P4715L, R5716C, I5967V and T6564I in ORF1b, T89I in ORF 3a and 203–204 del in N gene (Fig. 2B). The mutations associated with immune evasion to antibody response elicited post infection or vaccination7 were found in the breakthrough infection in this case. Similarly, the mutations which have been attributed to increase affinity binding between Receptor binding domain and ACE2 and those linked to immune evasion in Omicron8 were found in the specimens collected during re-infection. The sequence analysis clearly demonstrates that the case had a breakthrough infection with Delta derivative (AY.112) and reinfection with the Omicron sub lineage BA.2. Although, the clinical specimens of the primary infection couldn't be characterized, the probability of B.1 as an infecting variant would be higher as infection occurred in October 2020 where emerging variants were not detected in India.
A serum sample collected post 8 days reinfection was tested by Anti-SARS-CoV-2 human IgG ELISAs namely, whole virion inactivated antigen, receptor Binding Domain of Spike Protein (S1-RBD) specific IgG and Nucleocapsid (N) protein IgG capture ELISA as described earlier.9IgG antibody titre was found to be 1:400, 1:1300 and 1:80 for whole antigen, S1-RBD and N protein ELISA. Although the case had a breakthrough infection followed by a reinfection, the antibody titres were lower than expected.
With all these findings, we confirm the case of a breakthrough infection with Delta despite the hybrid immunity followed by a reinfection with Omicron post breakthrough infection. A recent study has shown neutralization for Omicron variant being reduced to 25.9 fold after a previous delta infection.8 It is reported that individuals with a prior infection with delta variant, remain susceptible to a repeat infection with Omicron.3 , 8 The findings of the study prove the immune evasion potential of the Omicron post natural infection and vaccination.
The fight against COVID-19 is far from over. Although, vaccine boosters or third dose have shown an augmentation in the immune response against Omicron variant,8 it has been found to wane over time. In such situation, non pharmacological interventions in the form of mask, hand hygiene and infection control remain the most reliable weapons to curb the direct transmission of SARS-CoV-2.
Ethical statement
The study was approved by the Institutional Biosafety Committee and Institutional Human Ethics Committee of ICMR-National Institute of Virology, Pune, India under the project titled “Assessment of immunological responses in breakthrough cases of SARS- CoV-2 in post COVID-19 vaccinated group” [No. NIV/IEC/May/2021/D-10 dated May 20, 2021]. The informed written consent was taken from the case under this study.
Funding
Financial support was provided by the Indian Council of Medical Research (ICMR), New Delhi at ICMR-National Institute of Virology, Pune under intramural funding ‘COVID-19′.
Conflicts of Interest
Authors do not have a conflict of interest among themselves.
Acknowledgment
Authors gratefully acknowledge the encouragement and support extended by Prof. (Dr.) Priya Abraham, Director, ICMR-NIV, Pune.
The authors would like to gratefully acknowledge the staff of ICMR-NIV, Pune including Mrs. Triparna Majumdar, Mrs. Savita Patil, Dr Rajlaxmi Jain, Miss. Pranita Gawande, Miss. Jyoti R Yemul, Dr. Gajanan Sapkal and Dr. Gururaj R. Deshpande for extending the excellent support.
References
- 1.Mensah A.A., Lacy J., Stowe J., Seghezzo G., Sachdeva R., Simmons R., et al. Disease severity during SARS-COV-2 reinfection: a nationwide study. J Infect. 2022;84(4):542–550. doi: 10.1016/j.jinf.2022.01.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Singh U.B., Rophina M., Chaudhry R., Senthivel V., Bala K., Bhoyar R.C, et al. Variants of concern responsible for SARS-CoV-2 vaccine breakthrough infections from India. J Med Virol. 2021:1–5. doi: 10.1002/jmv.27461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Altarawneh H.N., Chemaitelly H., Hasan M.R., Ayoub H.H., Qassim S., AlMukdad S., He D. Protection against the omicron variant from previous SARS-CoV-2 infection. N Engl J Med. 2022;386(13):1288–1290. doi: 10.1056/NEJMc2200133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Gao L., Mu X., Jiao Y.M., Wang F.S. Breakthrough Infection of SARS-CoV-2: An Emerging Challenge That Is Threatening Our World. Infectious Diseases & Immunity. 2021:E001. doi: 10.1097/ID9.0000000000000027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Choudhary M.L., Vipat V., Jadhav S., Basu A., Cherian S., Abraham P., et al. Development of in vitro transcribed RNA as positive control for laboratory diagnosis of SARS-CoV-2 in India. Ind J Med Res. 2020;151(2–3):251. doi: 10.4103/ijmr.IJMR_671_20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Thangaraj J.W.V., Yadav P.D., Kumar C.P.G., Shete A., Nyayanit D.A., Rani D.S., et al. Predominance of delta variant among COVID-19 vaccinated and unvaccinated individuals, India, May 2021. J Infect. 2022;84:113–115. doi: 10.1016/j.jinf.2021.08.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zhang M., Liang Y., Yu D., Du B., Cheng W., Li L., et al. A systematic review of vaccine breakthrough infections by SARS-CoV-2 Delta variant. Int. J. Biol. Sci. 2022;(18):889–900. doi: 10.7150/ijbs.68973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dejnirattisai W., Huo J., Zhou D., Zahradnı´k J., Supasa P., Liu C., et al. SARS-CoV-2 Omicron-B.1.1.529 leads to widespread escape from neutralizing antibody responses. Cell. 2022;185 doi: 10.1016/j.cell.2021.12.046. 467–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Deshpande G., Kaduskar O., Deshpande K., Bhatt V., Yadav P.D., Gurav Y., et al. Longitudinal clinic-serological analysis of anti-nucleocapsid and anti-receptor binding domain of spike protein antibodies against SARS-CoV-2. Int J Infect Dis. 2021;112:103–110. doi: 10.1016/j.ijid.2021.09.024. [DOI] [PMC free article] [PubMed] [Google Scholar]