Abstract
As apple fruits (Malus domestica) mature, they accumulate anthocyanins concomitantly with losing chlorophyll (Chl); however, the molecular pathways and events that coordinate Chl degradation and fruit coloration have not been elucidated. We showed previously that the transcription factor ETHYLENE RESPONSE FACTOR17 (MdERF17) modulates Chl degradation in apple fruit peels and that variation in the pattern of MdERF17 serine (Ser) residues is responsible for differences in its transcriptional regulatory activity. Here, we report that MdERF17 interacts with and is phosphorylated by MAP KINASE4 (MdMPK4-14G). Phosphorylation of MdERF17 at residue Thr67 by MdMPK4-14G is necessary for its transcriptional regulatory activity and its regulation of Chl degradation. We also show that MdERF17 mutants with different numbers of Ser repeat insertions exhibit altered phosphorylation profiles, with more repeats increasing its interaction with MdMPK4. MdMPK4-14G can be activated by exposure to darkness and is involved in the dark-induced degreening of fruit peels. We also demonstrate that greater phosphorylation of MdERF17 by MdMPK4-14G is responsible for the regulation of Chl degradation during light/dark transitions. Overall, our findings reveal the mechanism by which MdMPK4 controls fruit peel coloration.
Variation in the Ser repeat insertion in MdERF17 regulates fruit peel degreening through control of the degree of MdERF17 phosphorylation by MdMPK4 during light/dark transitions.
IN A NUTSHELL.
Background: One of the most important visual indicators of fruit ripening in apple is the color change caused by anthocyanin accumulation and chlorophyll degradation. We previously showed that the apple ethylene response factor (ERF) MdERF17 contributes to chlorophyll (Chl) degradation in the fruit peel via transactivation of chlorophyll degradation-related genes. MAPK cascades play an essential role in plant growth and development and are involved in the regulation of plant light responses. We also recently showed that apple MPK4 is activated by light and regulates anthocyanin accumulation by phosphorylating MYB1 in apple peel. We know relatively little about the molecular mechanisms coordinating the control of Chl loss and anthocyanin accumulation or whether they are coordinately regulated by external stimuli during ripening.
Question: As apple fruits mature, anthocyanin accumulation occurs concomitantly with loss of Chl; however, the molecular systems that bridge the two processes have not been elucidated. We wanted to test possible connections between diurnal rhythms and MAPK pathways via MdERF17 function during fruit coloration.
Findings: We found that MdMPK4-14G directly interacts with and phosphorylates MdERF17 in vitro and in vivo. MdMPK4-14G is essential for fruit peel degreening, since overexpression of a constitutively active form of MdMPK4-14G in fruit peel led to substantial Chl degradation, while the suppression of MdMPK4-14G displayed an opposite phenotype. The increased levels of phosphorylated MdERF17 at night are necessary for promoting Chl degradation in apple peel and variation in the Ser repeat number in ERF17 affects the accumulation of phosphorylated MdERF17. We established that MdMPK4s are involved in apple peel coloration and revealed the molecular mechanisms by which two MdMPK4 proteins regulate Chl breakdown and anthocyanin accumulation in response to light/dark transitions.
Next steps: Our work demonstrated that phosphorylation of MdERF17 by MdMPK4 regulates Chl degradation and that the ERF17 phosphorylation site recognized by MPK4-14G may be conserved in multiple species. It will be interesting to explore the range of conserved and independent regulatory mechanisms of Chl degradation in the fruit peel of different fruits.
Introduction
Fruit ripening is a complex developmental process involving the coordinated regulation of numerous metabolic pathways responsible for changes in color and firmness, aroma production, conversion of starch to sugars, and changes in acidity (Giovannoni et al., 2017). Ripening-associated color change, which is largely determined by the contents of flavonoids, carotenoids, and chlorophyll (Chl), is an important visual indicator of maturity that directly affects the economic value of fruit; it is also a key breeding target for developing new varieties (Giovannoni, 2004; Zhu et al., 2018). Thus, the genetic basis of fruit color variation and the regulatory systems controlling coloration are a major focus of research (Liu et al., 2015; Huang et al., 2018).
The anthocyanin biosynthetic pathway and associated regulatory genes have been characterized in many plant species, and several noncoding RNAs associated with anthocyanin accumulation have been identified (Espley et al., 2007; Gonzalez et al., 2008; An et al., 2012, 2015; Albert et al., 2014; Yoshida et al., 2015; Ma et al., 2021). Leaf senescence has been used as an experimental system to characterize the Chl degradation pathway and identify several Chl catabolic genes; these studies have revealed the importance of Chl catabolism in plant development as well as for mediating adaptation to environmental conditions (Schelbert et al., 2009; Hörtensteiner, 2013; Gao et al., 2016). NAM (no apical meristem), ATAF (Arabidopsis transcription activation factor), CUC (cup-shaped cotyledon) and WRKY transcription factors play key roles in controlling senescence-associated loss of Chl by regulating the expression of Chl degradation genes (Robatzek and Somssich, 2001; Balazadeh et al., 2010; Zhou et al., 2011; Mao et al., 2017). Common molecular pathways underlying Chl degradation in fruit ripening and leaf senescence have been identified (Harpaz-Saad et al., 2007; Müller et al., 2007; Hörtensteiner and Kräutler, 2011). In tomato (Solanum lycopersicum) fruit, the STAY-GREEN (SlSGR) gene is required for Chl degradation, and its silencing results in a stay-green phenotype (Barsan et al., 2010; Fujisawa et al., 2013). Studies in sweet orange (Citrus × sinensis) have indicated that the expression of the transcription factor gene ETHYLENE RESPONSE FACTOR13 (CitERF13) is induced during ripening; its overexpression in fruit peels accelerates degreening via the regulation of Pheophytin pheophorbide hydrolase (PPH) genes, which encode proteins that catabolize Chl (Yin et al., 2016). A recent study showed that tomato WRKY32 regulates fruit color by binding to W-box and W-box-like motifs in the regulatory promoter region of YELLOW-FRUITED TOMATO1, a putative component of the ethylene signaling pathway, WRKY32-knockdown tomato fruit display a yellow-fruited phenotype (Zhao et al., 2021). As apple fruit mature, anthocyanin accumulation is accompanied by the loss of Chl; however, the molecular mechanisms bridging the two processes have not been elucidated.
Anthocyanin accumulation and Chl degradation are regulated by phytohormonal and environmental cues (Shemer et al., 2008; Jaakola, 2013; Zhu et al., 2017; Yang et al., 2019). Fruit coloration is regulated at multiple levels by developmental programs and environmental stimuli, including light, which provides energy for fruit development and controls various physiological processes (Chen et al., 2004). Fluctuations in light levels play a key role in plant growth and development and coordinate metabolic and physiological processes associated with light/dark (diurnal) cycles (Harmer et al., 2000; Steed et al., 2021). For example, following light activation, photoreceptors induce the biosynthesis of Chl and anthocyanins (Kami et al., 2010). There is substantial evidence that the regulatory and structural genes required for anthocyanin biosynthesis are co-regulated and that their expression is enhanced by sunlight (Holton and Cornish, 1995). Other studies have shown that the transcription factor MdMYB1 enhances anthocyanin biosynthesis and that the expression of its encoding gene can be induced by light in apple skin (Takos et al., 2006; Allan et al., 2008). MdMYB1 can be degraded in the dark by the E3 ubiquitin ligase CONSTITUTIVE PHOTOMORPHOGENIC1 via the ubiquitin-dependent pathway (Li et al., 2012). The expression of most genes required for Chl biosynthesis, such as CHLH (a subunit of Mg-chelatase), HEMA1 (glutamyl-tRNA reductase), and GUN4 (GENOMES UNCOUPLED 4), is light-dependent and regulated by both diurnal and circadian rhythms (Matsumoto et al., 2004; Peter and Grimm, 2009). In contrast, exposure to dark can inhibit Chl biosynthesis and promote Chl degradation during leaf senescence, which is likely mediated by phytochrome signaling. The Chl contents of dark-treated PHYTOCHROME-INTERACTING FACTOR 4 (PIF4) overexpression lines decreased compared to wild-type plants because its encoded transcription factor PIF4 directly regulates the transcription of the Chl degradation regulatory gene NON-YELLOWING1 and substantially represses the expression of GOLDEN 2-LIKE2, which is involved in maintaining chloroplast activity, in Arabidopsis (Arabidopsis thaliana) (Song et al., 2014). However, the regulatory mechanism underlying the control of Chl loss in darkness, which results in fruit degreening, has not yet been fully characterized.
Environmental and developmental cues can be perceived by receptors and sensors and converted into physiological responses through various signal transduction pathways (Kholodenko, 2006), in which the evolutionarily conserved mitogen-activated protein kinase (MAPK) cascades play a critically important role (Rodriguez et al., 2010). Biochemical and genetic studies have revealed that MAPK signaling plays a role in developmental programs by mediating the phosphorylation of downstream targets, which contributes to transcriptional reprogramming (Komis et al., 2018). Arabidopsis MPK3, MPK4, and MPK6, as well as their orthologs in other plant species, are the most extensively studied (Lampard et al., 2008; Andreasson and Ellis, 2010; Yamada et al., 2017; Zhang et al., 2018). These three MAPK proteins are activated by light. Phosphorylation of MYB75 by MPK4 increases MYB75 stability, which is required for anthocyanin accumulation in response to light in Arabidopsis (Li et al., 2016a, 2016b). Although several studies have confirmed the importance of MAPK proteins in regulating plant development, few have been functionally characterized in the context of fruit development (Bergmann, 2004; Cho et al., 2008; Hettenhausen et al., 2012; Jia et al., 2016). We previously showed that the constitutive activation of MdMPK4 in transgenic apple fruit promotes light-induced anthocyanin production via MdMYB1 phosphorylation, suggesting that MAPK proteins affect fruit coloration (Yang et al., 2021). In apple, the most conspicuous phenotypic changes during fruit ripening are the color changes that occur as a consequence of the accumulation of anthocyanins and the breakdown of Chl (Hörtensteiner, 2006; Espley et al., 2009). The molecular mechanisms that coordinate these two processes have not yet been elucidated; it is also unclear whether these metabolic processes are simultaneously regulated by external stimuli during ripening. We also previously showed that apple ETHYLENE RESPONSE FACTOR17 (MdERF17) contributes to Chl degradation in the fruit peel; furthermore, we demonstrated that variation in the number of Ser residues in MdERF17 is responsible for differences in the transcriptional activity of MdERF17 that accounts for variation in Chl degradation rates in apple peels (Han et al., 2018).
In this study, we show that MdMPK4-14G directly interacts with and phosphorylates MdERF17 in vitro and in vivo and that MdMPK4-14G-mediated phosphorylation of MdERF17 contributes to fruit degreening by activating the expression of downstream genes. We also investigated the possible connection between diurnal rhythms and MAPK pathways via the function of MdERF17 during fruit development. Our results clarify the molecular mechanisms by which MdMPK4-14G promotes the breakdown of Chl in response to light/dark transitions.
Results
MdMPK4 interacts with the ERF transcription factor MdERF17
We recently showed that MdMPK4 interacts with and phosphorylate MdMYB1, which was essential for anthocyanin accumulation in response to light (Yang et al., 2021). MdMYB1 and MdERF17 also regulate anthocyanin accumulation and Chl degradation by mediating the shift in color from green to red during fruit ripening, and variation in the pattern of Ser residues in ERF17 is responsible for differences in its transcriptional regulatory activity (Allan et al., 2008; Han et al., 2018). To determine whether the MAPK pathway is involved in apple fruit degreening during fruit development, we evaluated phosphorylated MAPK proteins using phospho-p44/42 MAPK (Erk1/2) antibodies, as previously described (Boutrot et al., 2010), which identified three kinases with molecular weights of approximately 70, 50, and 42 kDa. We observed a correlation between the protein levels of potentially phosphorylated MPKs and the Chl degradation rate in apple fruit (“Gala”) peels (Supplemental Figure S1). In plants, MPK3, MPK4, and MPK6 have been extensively studied (Lampard et al., 2008; Andreasson and Ellis, 2010; Yamada et al., 2017; Zhang et al., 2018). We thus determined whether MdMPK3, MdMPK4, and/or MdMPK6 might interact with MdERF17 using bimolecular fluorescence complementation (BiFC) analysis. We previously designated MdERF17 variants containing three, six, and eight Ser repeats as ERF17-3S, ERF17-6S, and ERF17-8S, respectively(Han et al., 2018). Arabidopsis MPK3, MPK4, and MPK6 point mutants (MPK3D193G,E197A, MPK4D198G,E202A, and MPK6D218G,E222A) are constitutively active (CA) while maintaining their specificity toward their substrates (Berriri et al., 2012). We, therefore, introduced the corresponding mutations in apple MPK proteins which predicted to be CA MdMPK3E192G,E196A, MdMPK4D198G,D200G;E202A,E204A, and MdMPK6D224G,D227G;E228A, E231A(CA-MdMPK3, CA-MdMPK4, and CA-MdMPK6) to investigate substrate specificity. We previously identified two MdMPK3 genes (MdMPK3-03G, MD03G1108500 and MdMPK3-11G, MD11G1121500), two MdMPK4 genes (MdMPK4-06G, MD06G1089500 and MdMPK4-14G, MD14G1110400), and two MdMPK6 genes (MdMPK6-02G, MD02G1004000 and MdMPK6-15G, MD15G1147300) (Yang et al., 2021). We thus generated constructs encoding MdERF17 variants fused to the C-terminus of the yellow fluorescent protein (YFP) (MdERF17-3S-cYFP, MdERF17-8s-cYFP) and CA MdMPK3-03G, MdMPK3-11G MdMPK4-06G, MdMPK4-14G, MdMPK6-02G, and MdMPK6-15G fused to the N-terminus of YFP (CA-MdMPK3s, CA-MdMPK4s, or CA-MdMPK6s-nYFP), using both genes per kinase type. When we transiently co-infiltrated the MdERF17-cYFP and CA-MdMPK4s-nYFP constructs in Nicotiana benthamiana leaves, we observed fluorescence signal predominantly in the nucleus for all combinations (MdERF17-3S-cYFP/CA-MdMPK4-06G-nYFP, MdERF17-8S-cYFP/CA-MdMPK4-06G-nYFP, MdERF17-3S-cYFP/CA-MdMPK4-14G-nYFP, and MdERF17-8S-cYFP/CA-MdMPK4-14G-nYFP). In contrast, we detected no fluorescence for other combinations, although MPK3, MPK4, and MPK6 belong to the same kinase family. Therefore, cotransfection of the CA-MdMPK3-nYFP/CA-MdMPK6-nYFP and MdERF17-cYFP constructs also can be used as negative controls to confirm an interaction between MdMPK4 and MdERF17 (Figure 1A; Supplemental Figures S2–S4).
Figure 1.
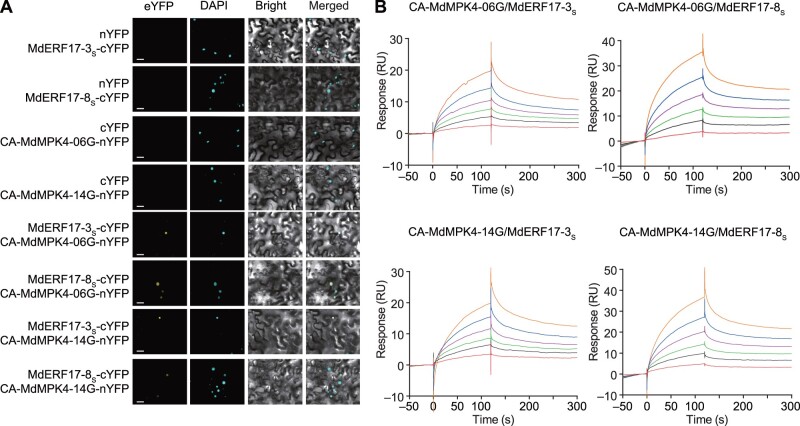
Interaction assays between MdERF17 and MdMPK4. A, BiFC assays to assess the interaction between MdERF17 and MdMPK4 in N. benthamiana leaves. The indicated constructs were transiently expressed in N. benthamiana leaves, using CA forms of the kinases. Co-expression of MdERF17-3S-cYFP, MdERF17-8S-cYFP, and nYFP or CA-MdMPK4-06G-nYFP, CA-MdMPK4-14G-nYFP, and cYFP were used as negative controls. Yellow fluorescence indicates a positive interaction, and DAPI staining was used to visualize the nuclei. Scale bars 20 µm. B, SPR binding profiles of MdERF17-3S and MdERF17-8S onto CA-MdMPK4-06G and CA-MdMPK4-14G. Various concentrations of recombinant MdERF17 protein were injected over the MdMPK4-immobilized sensor chip. The curves represent the concentrations of the injected MdERF17. From bottom to top: 0.094, 0.187, 0.375, 0.75, 1.5, and 3 µM were used for MdERF17-8S and 0.0625, 0.125, 0.25, 0.5, 1, and 2 µM were used for MdERF17-3S. RU, resonance units.
We also carried out surface plasmon resonance (SPR)-based interaction analyses to evaluate the binding affinity and generate kinetic information using recombinant glutatione S-transferase (GST)-MdERF17-3S, GST-MdERF17-8S, GST-CA-MdMPK4-06G, and GST-CA-MdMPK4-14G proteins. The resulting binding sensorgrams indicated that GST-CA-MdMPK4-06G and GST- CA-MdMPK4-14G show strong affinity for GST-MdERF17-3S and GST-MdERF17-8S (Figure 1B; Supplemental Figure S5). Overall, these results demonstrated that MdERF17 and MdMPK4 interact in vitro and in vivo and suggest that MdMPK4 might contribute to fruit degreening via post-transcriptional regulation of MdERF17.
MdMPK4 phosphorylates MdERF17 at Thr67
We conducted an in vitro kinase assay to examine whether MdMPK4 can phosphorylate MdERF17 using GST-tagged CA-MdMPK4-06G, CA-MdMPK4-14G, and MdERF17-8S. Immunoblotting analysis using anti-phosphoserine/threonine-specific antibodies revealed a band of the appropriate molecular weight for GST-CA-MdMPK4, confirming the protein kinase activity of CA-MdMPK4 and its autophosphorylation in vitro (Figure 2A). Similarly, we detected phosphorylation of MdERF17-8S by CA-MdMPK4-14G (Figure 2A). We confirmed the identities of the corresponding bands from Coomassie Brilliant Blue (CBB) staining by liquid chromatography–tandem mass spectrometry (LC–MS/MS) analysis (Supplemental Figure S6; Supplemental Table S1). The extent of phosphorylation by CA-MdMPK-06G was lower than that catalyzed by CA-MdMPK4-14G. We observed no phosphorylated band corresponding to MdERF17-8S (the negative control) in the absence of CA- MdMPK4.
Figure 2.
MdMPK4-14G directly phosphorylates MdERF17 in vitro and in vivo. A, In vitro phosphorylation analysis of MdERF17 by CA-MdMPK4-06G and CA-MdMPK4-14G. Upper panel, phosphorylated MdERF17, and autophosphorylation of MdMPK4 were detected using anti-phosphoserine/threonine. Bottom panel, recombinant MdMPK4 and MdERF17 proteins in CBB-stained gel. The MdMPK4 and MdERF17 proteins are indicated by arrows, and two intense MdERF17 protein bands are indicated with a red asterisk. B, Identification of the MdERF17 site phosphorylated by CA-MdMPK4-14G in vitro. One phosphopeptide peak (m/z 803.38) matching LGSYDpT67PEKAARAF originating from the AP2/ERF domain was detected from phosphorylated MdERF17, and Thr67 was identified as the phosphorylated residue by LC–MS/MS. C, In vitro analysis phosphorylation of wild-type and mutated MdERF17 by MdMPK4-14G. Recombinant MdERF17 and MdERF17T67A were incubated with CA-MdMPK4-14G. Upper, immunoblot with anti-phosphoserine/threonine antibody; bottom panel, CBB-stained gel. The MdMPK4 and MdERF17 proteins are indicated by arrows. D, The Thr67 residue in the AP2/ERF domain of MdERF17 is conserved in other plant species, but the number of Ser repeat insertions varies. E, Phosphorylation of MdERF17 at Thr67 by MdMPK4-14G in vivo. The indicated constructs were transiently infiltrated in N. benthamiana leaves. Phosphorylated MdERF17 was detected using anti-Phos-MdERF17 antibodies. MdERF17, MdERF17T67A, and MdMPK4 loading control were detected by immunoblot analysis using anti-Myc and anti-GFP antibodies. Wild-type plants were used as the negative controls. F and G, MdERF17 is phosphorylated by MdMPK4-14G in apple fruit calli. Apple calli transformation with OE-MdMPK4-14G and OE-CA-MdMPK4-14G was confirmed by RT-qPCR (F) and immunoblotting with anti-GFP antibodies (G). The levels of phosphorylated MdERF17 in apple calli were determined by immunoblotting with anti-Phos-MdERF17 antibodies. In (F), the expression levels in WT were set to 1; error bars indicate the standard deviation (sd) of three biological replicates. Asterisks indicate statistically significant differences (**P < 0.01) according to one-way ANOVA, followed by Tukey’s honestly significant difference test.
To further characterize the MdMPK4 phosphorylation sites on MdERF17-8S, we used LC–MS/MS to identify phosphopeptides from the in vitro kinase reactions described above. We detected the phosphorylation of Thr67 in a single phosphorylated peptide originating from the ERF/AP2 domain when incubated with CA-MdMPK4-14G, but not with CA-MdMPK4-06G (Figure 2B; Supplemental Table S2). To independently confirm the phosphorylation site of MdERF17-8S, we mutated Thr67 in MdERF17-8S to Ala to create the non-phosphorylatable mutant MdERF17T67A. We performed an in vitro kinase assay with MdERF17-8S and MdERF17T67A, which showed that the phosphorylation of by CA-MdMPK4-14G cannot phosphorylate MdERF17T67A, whereas CA-MdMPK4-14G strongly phosphorylated intact MdERF17 (Figure 2C), suggesting that Thr67 is a prominent CA-MdMPK4-14G target site. These results showed that CA-MdMPK4-14G can phosphorylate MdERF17-8S at Thr67; thus, we focused on MdMPK4-14G in our subsequent studies.
Thr67 is a highly conserved residue in MdERF17 orthologs from Arabidopsis, tobacco (Nicotiana tabacum), tomato, pear (Pyrus bretschneideri), and sweet orange; these orthologs are predicted to harbor varying numbers of Ser repeat insertions before the AP2/ERF domain. The ERF17 ortholog from peach (Prunus persica) did not contain the Thr67 phosphorylation site or Ser repeats, suggesting that the ERF17 phosphorylation site recognized by MPK4-14G may exist in some, but not all, species and that this might be related to the number of Ser repeats (Figure 2D).
To determine whether MdERF17 can be phosphorylated by CA-MdMPK4-14G in vivo, we raised polyclonal antibodies using a synthetic peptide harboring phosphorylated Thr67 (LGSYDpT67PEKAARAF). We established the specificity of these anti-Phos-MdERF17 antibodies by immunoblot analysis using CA-MdMPK4-14G, MdERF17-8S, and MdERF17- for in vitro kinase assays (Supplemental Figure S7). We also transiently infiltrated N. benthamiana leaves with the constructs MdERF17-Myc, MdERF17T67A-Myc, or MdERF17-Myc with CA-MdMPK4-14G-GFP or MdERF17T67A-Myc with CA-MdMPK4-14G-GFP and measured phosphorylated MdERF17 levels. We detected the tagged proteins by immunoblotting with anti-Myc and anti-GFP antibodies (Figure 2E) and used the anti-Phos-MdERF17 antibodies to examine MdERF17 phosphorylation in planta. We detected a phosphorylation signal corresponding to the molecular weight of MdERF17-Myc in leaves accumulating intact MdERF17-Myc that increases in intensity when co-expressed with CA-MdMPK4-14G-GFP; however, incubation of the protein extracts with lambda protein phosphatase (λPP) eliminated the signal corresponding to phospho-MdERF17. Conversely, we observed no phosphorylation signals in samples accumulating MdERF17T67A-Myc (Figure 2E). Furthermore, we expressed CA-MdMPK4-14G-GFP or MdMPK4-14G-GFP in apple calli. In both cases, we detected a strong band using the anti-Phos-MdERF17 antibodies that corresponds to the predicted molecular weight of MdERF17 (Figure 2, F and G), indicating that MdMPK4-14G phosphorylates MdERF17 in vivo. Collectively, these in vitro and in vivo experimental results suggest that MdMPK4-14G interacts with and phosphorylates MdERF17.
MdMPK4-14G activates the transcriptional activity of MdERF17
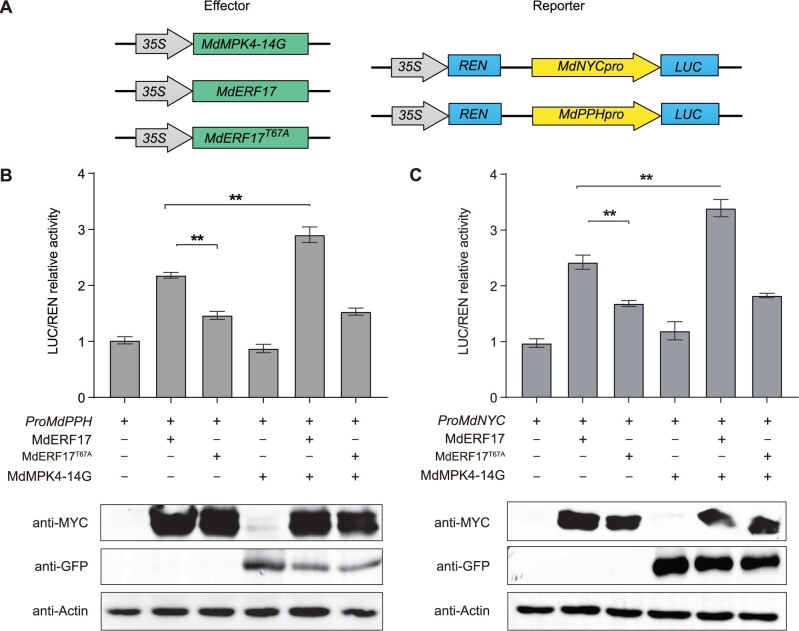
Our previous studies demonstrated that MdERF17 promotes Chl degradation by directly binding to the promoters of the Chl degradation-related genes MdPPH and NONYELLOW COLORING (MdNYC) (Han et al., 2018). To address whether MdERF17-mediated transactivation is controlled by MdMPK4-14G and requires phosphorylation at Thr67, we performed a firefly luciferase (LUC) complementation imaging assay in N. benthamiana leaves. Accordingly, we transiently co-infiltrated leaves with constructs encoding MdERF17, MdERF17T67A, and MdMPK4-14G effector constructs and driven by the cauliflower mosaic virus (CaMV) 35S promoter with different combinations of reporter constructs with LUC under the control of the MdNYC or MdPPH promoters (Figure 3A). The transient expression of MdMPK4-14G did not significantly affect relative LUC activity from either promoter construct when compared to the negative control without effector. However, transcriptional activity increased substantially when MdERF17 was co-expressed with the MdNYCpro and MdPPHpro reporters, which was consistent with our previous results (Han et al., 2018); co-expression of MdMPK4 and MdERF17 further induced MdERF17-activated MdNYC and MdPPH transcription. In contrast, expressing MdERF17T67A resulted in lower transcriptional output from the MdNYCpro and MdPPHpro reporters than MdERF17, and co-expression of MdERF17T67A and MdMPK4-14G did not raise LUC activity further, supporting the conclusion that MdMPK4-14G targets Thr67 to regulate MdERF17 activity (Figure 3, B and C). These results suggest that MdMPK4-14G affects the transcriptional regulatory activity of MdERF17 through phosphorylation at Thr67.
Figure 3.
MdMPK4-14G enhances the DNA-binding activity of MdERF17 by Thr67 phosphorylation. A, Schematic diagram of the effector constructs expressing MdMPK4-14G, MdERF17, and MdERF17T67A and the reporter vectors containing the MdPPH and MdNYC promoters. B and C, LUC/REN ratios after co-infiltration into N. benthamiana leaves with different reporter and effector construct combinations, calculated using a dual-LUC reporter assay system. LUC activity was normalized to REN activity, with LUC/REN ratios from the control without effector set to 1 (B and C, upper). Protein levels of MdERF17, MdERF17T67A and MdMPK4 in different combinations (B and C, lower). In (B) and (C), error bars indicate the SD of three biological replicates. Asterisks indicate statistically significant differences (**P < 0.01) according to Student’s t test.
Phosphorylation of MdERF17 promotes Chl degradation
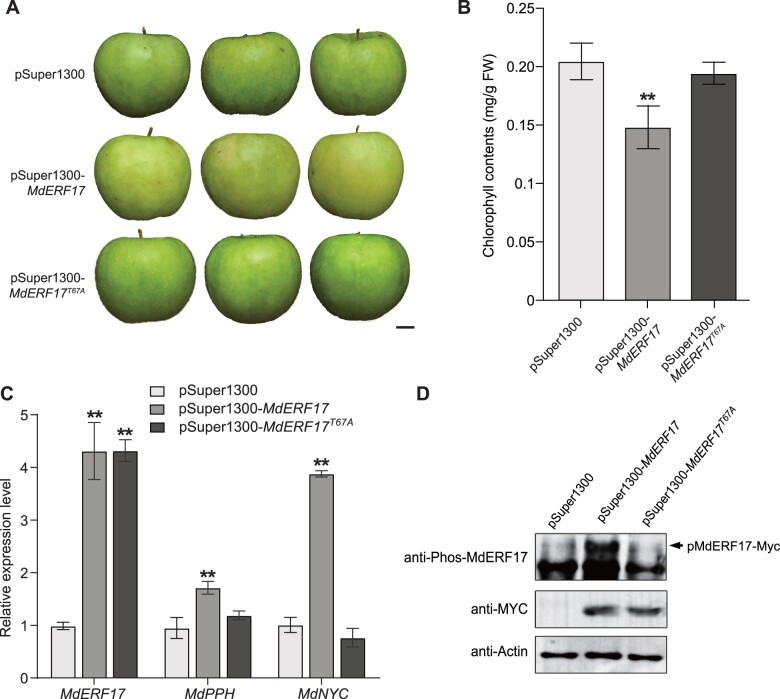
To assess the role of Thr67 phosphorylation in MdERF17 function in apple peels, we transiently overexpressed (OE) MdERF17 and MdERF17T67A driven by the constitutive 35S promoter in M. domestica “Granny Smith” fruit. We selected this cultivar because MdERF17 expression levels are undetectable in this background (Supplemental Figure S8, A and B), resulting in minimal interactions between endogenous MdERF17 and MdMPK4-14G (Han et al., 2018). MdERF17-overexpressing apple fruits exhibited degreening and reduced Chl contents relative to the empty vector controls. However, we observed no difference in Chl contents between MdERF17T67A transgenic fruit and the empty vector control (Figure 4, A and B). Although MdERF17 and MdERF17T67A transcript levels were similar in the transgenic fruit peels, MdNYC and MdPPH expression was high in MdERF17-overexpressing fruit, but not in MdERF17T67A-overexpressing fruit or control fruit (Figure 4C). Immunoblot analysis with anti-Phos-MdERF17 antibodies detected phosphorylation of MdERF17-Myc, but not of MdERF17T67A-Myc (Figure 4D), suggesting that Thr67 phosphorylation is necessary for MdERF17 to regulate Chl degradation in apple peels.
Figure 4.
MdMPK4-14G-mediated MdERF17 phosphorylation promotes Chl degradation. A, Phenotypes of vacuum-infiltrated apple fruit transformed with pSuper1300-MdERF17 or pSuper1300-MdERF17T67A were visualized 3 days after infiltration. Fruits infiltrated with an empty pSuper1300 vector were used as a control. Scale bar, 1 cm. B, Chl contents in the infiltrated apple fruits shown in (A). C, Relative expression levels of MdERF17 and the Chl degradation genes MdPPH and MdNYC in infiltrated apple fruit, as determined by RT-qPCR. The expression levels in the control were set to 1. D, Immunoblot analysis showing phosphorylated MdERF17 in infiltrated apple fruit. MdERF17 and MdERF17T67A were detected by immunoblotting with anti-Myc antibodies. Actin was used as a protein loading control. In (B) and (C), error bars indicate the SD of three biological replicates. Asterisks indicate statistically significant differences (**P < 0.01) according to one-way ANOVA, followed by Tukey’s honestly significant difference test.
Variation in the Ser repeat number in ERF17 alters its phosphorylation
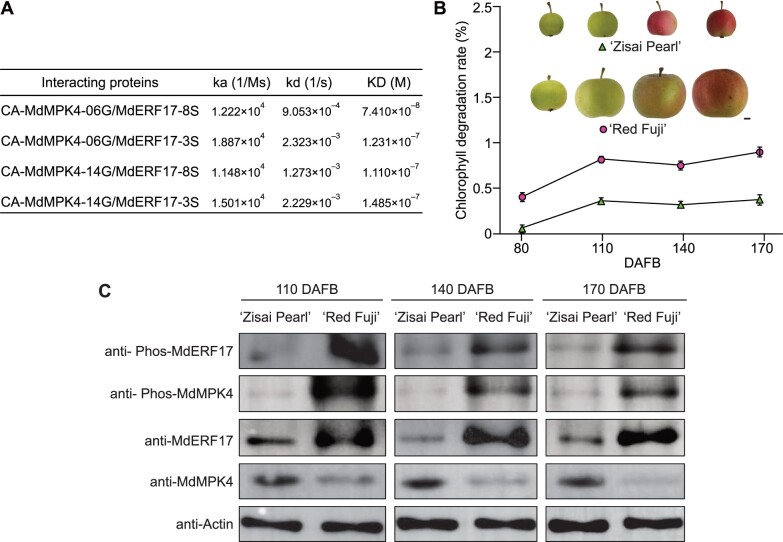
Given that the number of Ser repeats in ERF17 varies among Malus species, we hypothesized that this variation might affect the phosphorylation of MdERF17. An evaluation of the sensorgrams showed that MdERF17-8S has a lower KD value compared to MdERF17-3S, suggesting that MdERF17-8S has a higher affinity for CA-MdMPK4 proteins and that the increased Ser repeat number in MdERF17 enhances its interaction with MdMPK4 (Figure 5A). We previously showed that Malus asiatica “Zisai Pearl” is homozygous for a ERF17 variant encoding three Ser repeats, whereas M. domestica “Red Fuji” was heterozygous and encoded ERF17 variants with six or eight Ser repeats. The progeny from the “Zisai Pearl” × “Red Fuji” cross was ERF17-3S/ERF17-6S or ERF17-3S/ERF17-8S (Han et al., 2018). We confirmed the association between the ERF17 allele and the abundance of phosphorylated MdERF17 by analyzing the progeny from a cross between “Zisai Pearl” and “Red Fuji” and the parents. “Red Fuji” (ERF17-6S/ERF17-8S) had a higher Chl degradation rate than “Zisai Pearl” (ERF17-3S/ERF17-3S) (Figure 5B). Moreover, we observed weaker ERF17-phosphorylated bands in “Zisai Pearl” fruit compared to “Red Fuji” fruit (Figure 5C). Among the “Zisai Pearl” × “Red Fuji” hybrids, the ERF17-3S/ERF17-8S genotype accumulated more phosphorylated MdERF17 than the ERF17-3S/ERF17-6S genotype (Supplemental Figure S9, A–C). These results suggest that the number of Ser repeats in MdERF17 affects the accumulation of phosphorylated MdERF17 by affecting its ability to interact with and be phosphorylated by MdMPK4.
Figure 5.
Differences in the levels of phosphorylated MdERF17-8S and MdERF17-3S caused by Ser repeat number are associated with Chl degradation. A, Kinetic parameters for the interaction between MdERF17 and MdMPK4, as determined by SPR assays. ka: association rate. kd: dissociation rate. KD: kd/ka, dissociation equilibrium constant. B, Phenotypes and Chl degradation rate of “Zisai Pearl” and “Red Fuji” fruits at different developmental stages. Scale bar, 1 cm. C, Detection of MdERF17 and MdMPK4 phosphorylation and MdERF17 and MdMPK4 protein levels in the peels of “Zisai Pearl” and “Red Fuji” fruits from 110 to 170 DAFB using anti-Phos-MdERF17, anti-Phos-MdMPK4, anti-MdERF17, and anti-MdMPK4 antibodies. Actin was used as a protein loading control. In (B), error bars indicate the sd of three biological replicates.
Increased levels of phosphorylated MdERF17 at night are necessary for its function during fruit ripening
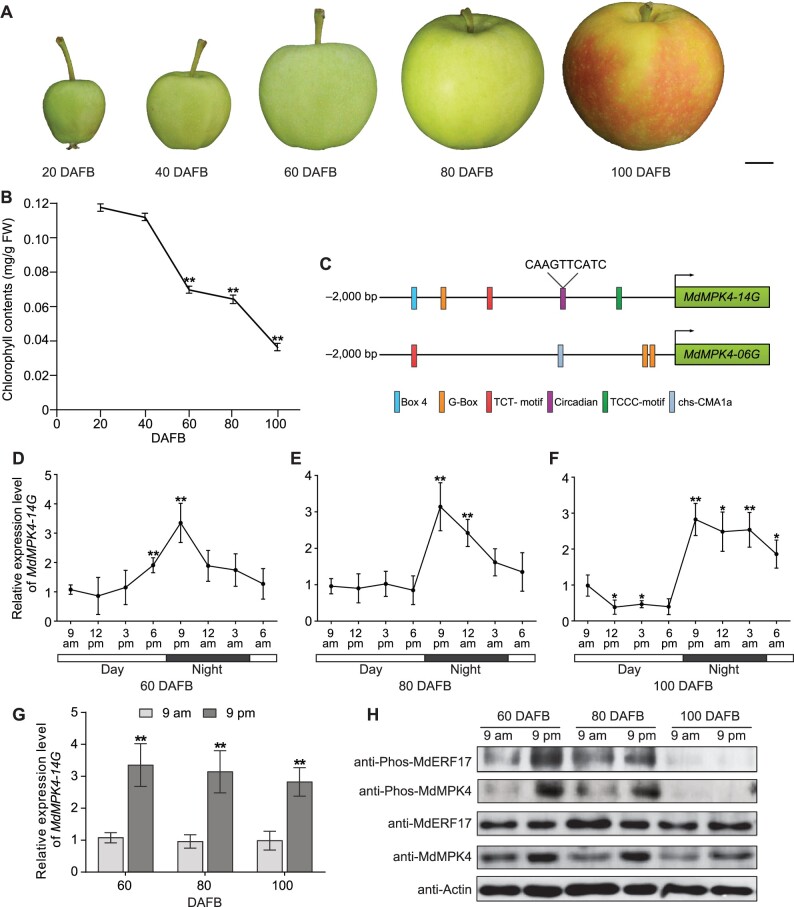
Based on the observation that MdMPK4-14G directly interacts with and phosphorylates MdERF17, we investigated the regulatory roles of MdMPK4-14G in apple peel degreening during ripening. Accordingly, we extracted total Chl from pericarp tissues at five stages of fruit development. Chl levels gradually decreased over the first two periods, 20 days after full bloom (DAFB) and 40 DAFB, before substantially declining in the subsequent 60 days to reach a very low level at the fruit-ripening stage (100 DAFB), suggesting that Chl degradation occurs mainly between 60 and 100 DAFB. We thus focused on this period, which coincided with the external color change from green to red, for subsequent analysis (Figure 6, A and B).
Figure 6.
MdMPK4-14G transcript and phosphorylation levels are higher in the dark during fruit development. Phenotypes (A) and Chl contents (B) of “Gala” fruits at different developmental stages. Scale bar, 1 cm. Asterisks indicate significant differences compared with Chl contents at 20 DAFB to show the Chl degradation period. C, Schematic illustration showing the distribution of putative circadian and light-responsive cis-elements present in the promoter regions of MdMPK4-14G and MdMPK4-06G, as predicted by the PLACE database. D–F, Time course analyses of endogenous MdMPK4-14G expression in apple peels, by RT-qPCR. Apple fruits at 60, 80, and 100 DAFB from plants grown under natural conditions were harvested in 3-h intervals throughout a 24-h period. The light period is indicated according to sunrise (represented by the white areas); darkness is represented by the dark areas. Asterisks indicate significant differences compared to the MdMPK4-14G expression levels at 9 am. G, MdMPK4-14G transcription levels change between 9 am and 9 pm from 60 to 100 DAFB. H, MdMPK4 and MdERF17 phosphorylation levels change between 9 am and 9 pm from 60 to 100 DAFB. Anti-MdMPK4 and anti-MdERF17 antibodies were used to detect endogenous MdMPK4 and MdERF17 proteins. Actin was used as a protein loading control. In (B) and (D–G), the expression levels at 9 am were set to 1; error bars indicate the SD of three biological replicates. Asterisks indicate statistically significant differences (*P<0.05; **P<0.01) according to Student’s t test.
To further elucidate the difference in MdERF17 phosphorylation caused by MdMPK4-06G and MdMPK4-14G, we analyzed 2,000 bp of the MdMPK4 promoter regions upstream from the translation start site using the PLACE database (https://www.dna.affrc.go.jp/PLACE/). Consistent with our previous study showing that several light-responsive elements are present in the two MdMPK4 promoter regions (Yang et al., 2021), we identified a “CAANNNNATC” cis-acting regulatory element involved in circadian control in the MdMPK4-14G promoter (Figure 6C) (Francisco et al., 2021). To test whether MdMPK4-14G expression was responsive to diurnal rhythms in apple peel during ripening, we measured MdMPK4-14G transcript levels in 3-h intervals over a 24-h period at 60, 80, and 100 DAFB. MdMPK4-14G transcript levels exhibited a diurnal expression pattern, with lower expression during the day and higher expression at night, reaching a peak at 9 pm (Figure 6, D–F). To test whether MdMPK4 protein stability might also show a diurnal pattern, we generated MdMPK4-specific antibodies and assessed protein abundance in the samples collected at 9 am and 9 pm at the different developmental stages. Immunoblot analysis indicated that the abundance of MdMPK4 protein is higher at night (9 pm) than during the daytime (9 am) and corresponded to the pattern seen in MdMAPK4-14G transcript levels (Figure 6, G and H). We also examined the changes in the phosphorylation status of MdMPK4 using anti-Phos-MdMPK4 antibodies and determined that MdMPK4 phosphorylation levels are correlated with protein abundance at 9 am and 9 pm in the different developmental stages. Furthermore, we observed higher levels of phosphorylation of MdERF17 by MdMPK4 in the dark than in the light, although MdERF17 protein abundance was similar at these two time points. This result suggested that MdERF17 phosphorylation is independent of protein levels and is modulated by light/dark cycles, possibly through MdMPK4 activity. A similar relationship between protein abundance and phosphorylation level was also reported for the EARLY FLOWERING 4 (ELF4) protein over a diurnal cycle (Choudhary et al., 2015). We noted that MdMPK4 and MdERF17 phosphorylation levels are low when fruits are fully ripe (at 100 DAFB) (Figure 6H). These results indicate that light/dark transitions and functional MdMPK4 are important in the control of MdERF17 phosphorylation and that a higher phosphorylation state of MdERF17 mediated by MdMPK4 at night might play a role in the regulation of Chl degradation.
MdMPK4-14G positively regulates fruit degreening under diurnal conditions
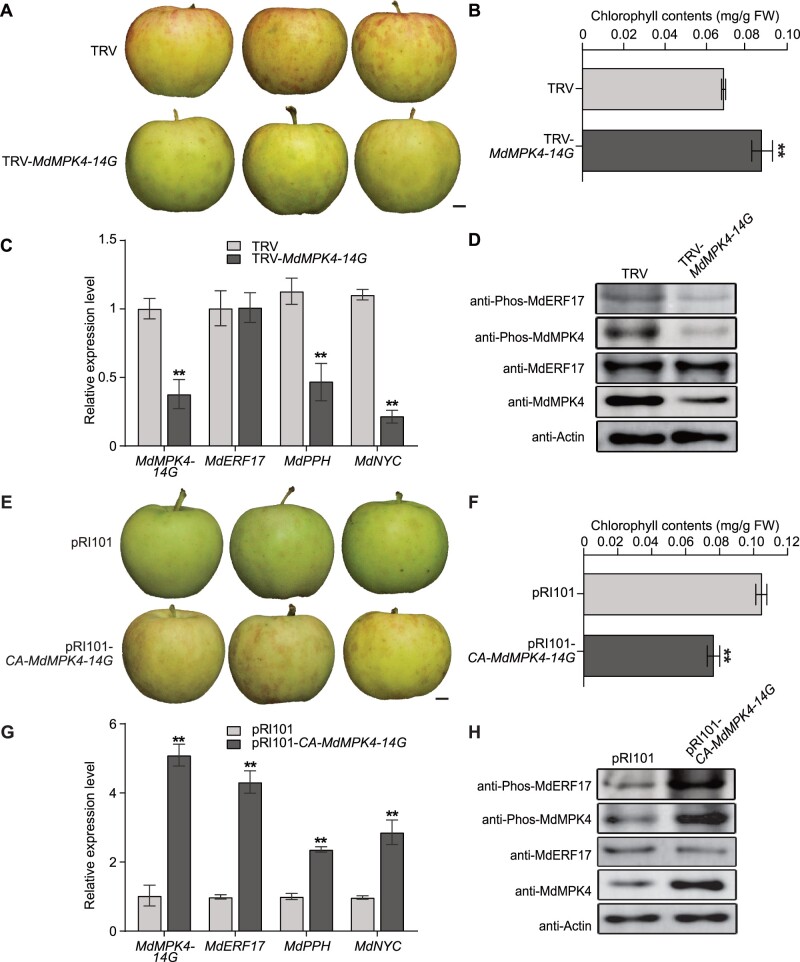
To elucidate the biological function of MdMPK4-14G in apple peel and investigate whether MdMPK4-14G is associated with the high level of phosphorylated MdERF17 at night, we conducted a transient expression assay in apple fruits. We generated a virus-induced gene silencing (VIGS) MdMPK4 construct (TRV-MdMPK4-14G), which we infiltrated into “Gala” apple fruits via Agrobacterium (Agrobacterium tumefaciens)-mediated transient transformation. We then stored the infiltrated apples in a phytotron with a 14-h light/10-h dark cycle in accordance with natural light conditions and collected samples during the day (9 am) and night (9 pm) to reflect the diurnal expression of MdMPK4-14G. Phosphorylated MdERF17 accumulated to similar levels in control fruits infiltrated with Agrobacterium containing the TRV empty vector and non-infiltrated fruits during ripening, while MdMPK4-14G silencing disrupted the dark-induced phosphorylation of MdERF17 in TRV-MdMPK4-14G fruits, indicating that dark-induced activation of MdERF17 depends on MdMPK4-14G in fruit peels (Supplemental Figure S10A). We therefore collected samples at 9 pm for subsequent assays. Compared to the TRV control, suppression of MdMPK4-14G resulted in delayed fruit degreening, which was accompanied by higher Chl contents and lower expression of the Chl degradation genes MdNYC and MdPPH (Figure 7, A–C). MdERF17 phosphorylation levels at 9 pm were much lower in MdMPK4-silenced fruit peels than in control fruit peels, although MdERF17 protein levels remained almost unchanged, indicating that phosphorylation by MdMPK4-14G does not affect MdERF17 stability (Figure 7D).
Figure 7.
Silencing or overexpression of MdMPK4-14G in “Gala” apple under 14-h light/10-h dark conditions. A, Phenotypes of TRV control and MdMPK4-14G-silenced (TRV-MdMPK4-14G) fruits. Scale bar, 1 cm. B, Chl contents in the peels of the TRV control and MdMPK4-14G-silenced fruits. C, Relative expression levels of MdMPK4-14G, MdERF17, and the Chl degradation genes MdPPH and MdNYC sampled at 9 pm in infiltrated apple fruits, as detected by RT-qPCR. D, Detection of MdERF17 and MdMPK4 phosphorylation and protein levels in infiltrated apple fruits sampled at 9 pm by immunoblot analysis with anti-MdERF17, anti-MdMPK4, anti-Phos-MdERF17 and anti-Phos-MdMPK4 antibodies. Actin was used as a protein loading control. E, Phenotypes of the pRI101 control and CA-MdMPK4-14G-overexpressing (pRI101-CA-MdMPK4-14G) fruits. Scale bar, 1 cm. F, Chl contents in the peels of the pRI101 control and CA-MdMPK4-14G-overexpressing fruits. G, Relative expression levels of MdMPK4-14G, MdERF17 and the Chl degradation genes MdPPH and MdNYC in infiltrated apple fruits sampled at 9 am, as detected by RT-qPCR. H, Detection of MdERF17 and MdMPK4 phosphorylation and protein levels in infiltrated apple fruits sampled at 9 am by immunoblot analysis with anti-MdERF17, anti-MdMPK4, anti-Phos-MdERF17, and anti-Phos-MdMPK4 antibodies. Actin was used as a protein loading control. In (B), (C), (F), and (G), the expression levels in the control were set to 1; error bars indicate the SD of three biological replicates. Asterisks indicate statistically significant differences (*P < 0.05, **P < 0.01) according to Student’s t test.
To further examine the relationship between MdMPK4-14G and Chl degradation, we transiently OE CA-MdMPK4-14G in “Gala” apple fruits and collected samples at different times of the day, as described above. Peels from OE-CA-MdMPK4-14G transgenic fruits exhibited higher levels of MdERF17 phosphorylation at 9 am compared to control fruit peels, but we observed no differences in phosphorylated MdERF17 between the transgenic fruit peels and the empty control at 9 pm (Supplemental Figure S10B). One possible explanation for these results is that dark-induced phosphorylation in the control already reach the maximal potential activity of MdMPK4-14G at night, thus MdMPK4-14G activity was only affected by overexpression during the day. The levels of phosphorylated MdERF17 were substantially higher in CA-MdMPK4-14G transgenic fruit peels than in the control at 9 am (Figure 7H). We also determined that overexpression of MdMPK4-14G in fruit peels leads to substantial Chl degradation compared to the control and that changes in MdNYC and MdPPH expression parallel the changes seen in Chl levels (Figure 7, E–G). Overall, these data indicate that MdMPK4-14G regulates fruit degreening under diurnal conditions.
To investigate whether MdMPK4-14G also contributes to Chl degradation in leaves, we generated stable transgenic N. benthamiana plants overexpressing MdMPK4-14G or CA-MdMPK4-14G. We isolated transgenic plants with higher MdMPK4-14G transcript levels and higher MdMPK4-14G abundance than in wild-type (Supplemental Figure S11, C and D). Compared to the wild-type plants, all transgenic plants exhibited an abnormal leaf color phenotype, with lower Chl contents under normal light conditions (Supplemental Figure S11, A and B). NbNYC and NbPPH expression were more highly expressed in MdMPK4-14G and CA-MdMPK4-14G transgenic plants than in wild-type plants (Supplemental Figure S11C), suggesting that the post-transcriptional regulation of MdERF17 by MdMPK4-14G also contributes to Chl degradation during leaf senescence.
MdMPK4-14G is involved in fruit degreening under dark conditions
Dark treatment can block Chl biosynthesis and promote its degradation during leaf senescence. Thus, we next investigated whether Chl degradation in the pericarp was induced by darkness, and whether the accumulation of phosphorylated MdMPK4-14G was directly regulated by dark signaling. To this end, we harvested “Gala” fruits at 80 DAFB (immature stage) and stored them in continuous darkness or constant light. We observed clear phenotypic differences between fruits maintained in dark or light conditions after 10 days, with a faster Chl degradation rate in the dark than in the light (Supplemental Figure S12, A and B). Reverse transcription-quantitative PCR (RT-qPCR) analyses indicated that MdERF17 expression is induced in darkness at this time point, which coincided with the increased expression of Chl degradation-related genes (Supplemental Figure S12C). Immunoblot analysis revealed that levels of phosphorylated MdERF17 and MdMPK4 are higher in the dark than in the light after 10 days of treatment (Supplemental Figure S12D). These results indicated that dark treatment promotes Chl degradation in fruit peels through the post-transcriptional regulation of MdERF17.
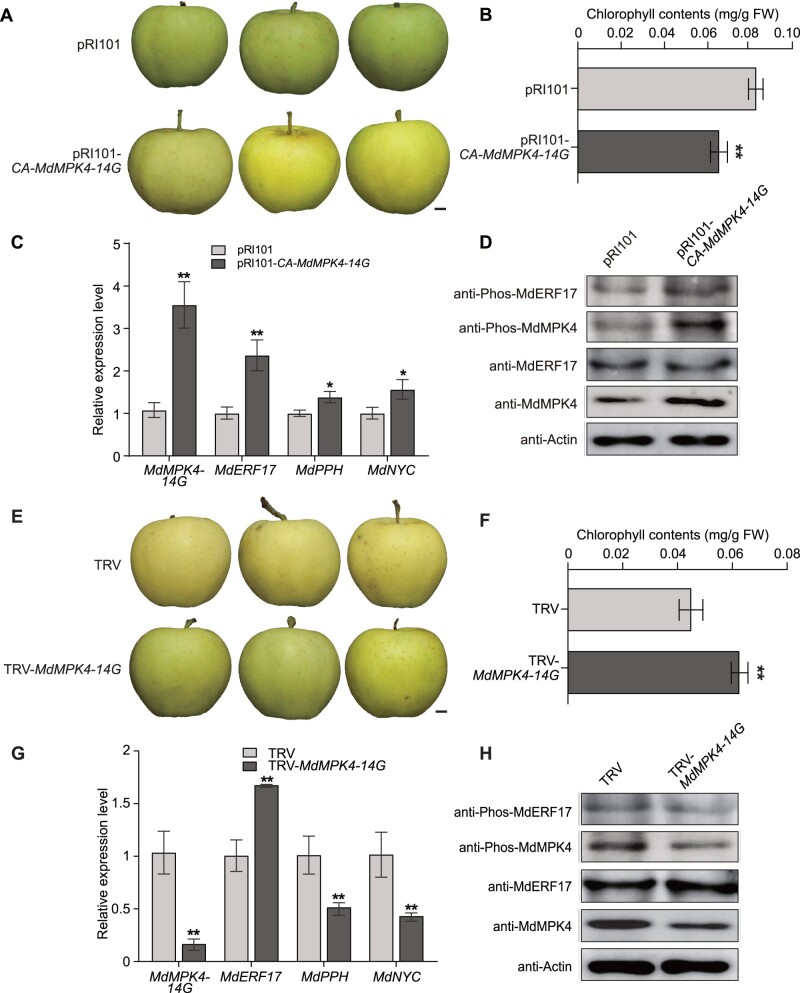
Because MdERF17 phosphorylation by MdMPK4-14G is induced by darkness, we determined whether MdMPK4-14G was involved in the regulation of Chl degradation under constant darkness. We stored “Gala” fruits transformed with the OE-CA-MdMPK4-14G or TRV-MdMPK4-14G constructs via Agrobacterium-mediated infiltration in a phytotron under constant darkness for 3 days (for overexpression) or 5 days (for silencing). We observed that overexpression of CA-MdMPK4-14G enhances the accumulation of Chl and increases the expression of MdNYC and MdPPH in transiently CA-MdMPK4-14G-overexpressing fruit peels (Figure 8, B and C). In contrast, MdMPK4-14G silencing resulted in a decrease in Chl contents and the repression of MdNYC and MdPPH expression (Figure 8, F and G). To determine whether MdERF17 phosphorylation by MdMPK4-14G occurred under continuous dark conditions, we assessed the abundance of phosphorylated MdERF17 in CA-MdMPK4-14G-overexpressing or MdMPK4-14G-silenced fruit peels. Immunoblot analysis using anti-Phos-MdERF17 antibodies demonstrated that the relative levels of phosphorylated MdERF17 increase when MdMPK4-14G was OE, while we observed the opposite pattern when MdMPK4-14G was silenced (Figure 8, D and H). These data suggest that MdERF17 phosphorylation by MdMPK4-14G promotes Chl degradation under dark conditions.
Figure 8.
Overexpression or silencing of MdMPK4-14G in “Gala” apple under continuous dark conditions. A, Phenotypes of the pRI101 control and CA-MdMPK4-14G-overexpressing (pRI101-CA-MdMPK4-14G) fruits. Scale bar, 1 cm. B, Chl contents in the peels of the pRI101 control and CA-MdMPK4-14G-overexpressing fruits. C, Relative expression levels of MdMPK4-14G, MdERF17, and the Chl degradation genes MdPPH and MdNYC in infiltrated apple fruits, as detected by RT-qPCR. D, Detection of MdERF17 and MdMPK4 phosphorylation and protein levels in infiltrated apple fruits. Samples were collected from the peels of the pRI101 control and CA-MdMPK4-14G-overexpressing fruits and then were subjected to immunoblot analysis with anti-MdERF17, anti-MdMPK4, anti-Phos-MdERF17, and anti-Phos-MdMPK4 antibodies. Actin was used as a protein loading control. E, Phenotypes of the TRV control and MdMPK4-14G-silenced (TRV-MdMPK4-14G) fruits. Scale bar, 1 cm. F, Chl contents in the peels of the TRV control and MdMPK4-14G-silenced fruits. G, Relative expression levels of MdMPK4-14G, MdERF17, and the Chl degradation genes MdPPH and MdNYC in infiltrated apple fruits, as detected by RT-qPCR. H, Detection of MdERF17 and MdMPK4 phosphorylation and protein levels in infiltrated apple fruits. Samples were collected from the peels of the TRV control and MdMPK4-14G-silenced fruits and then were subjected to immunoblot analysis with anti-MdERF17, anti-MdMPK4, anti-Phos-MdERF17, and anti-Phos-MdMPK4 antibodies. Actin was used as a protein loading control. In (B), (C), (F), and (G), the expression levels in the control were set to 1; error bars indicate the SD of three biological replicates. Asterisks indicate statistically significant differences (*P<0.05; **P<0.01) according to Student’s t test.
Discussion
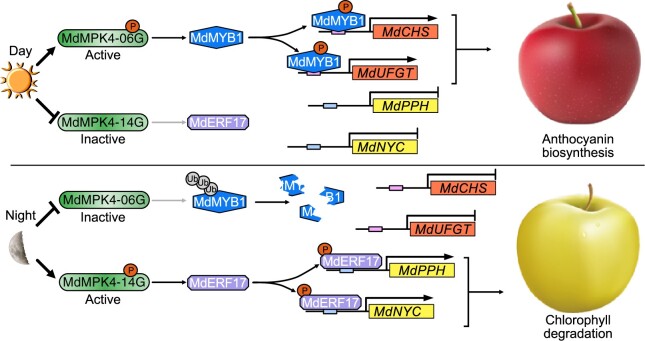
Based on the data presented in this study, we propose a model for the regulation of apple peel coloration. During the fruit coloration stage, light in the daytime activates MdMPK4-06G, which phosphorylates MdMYB1, thereby stabilizing MdMYB1 and promoting its function in upregulating the expression of genes in the anthocyanin biosynthesis pathway in the fruit peel. At night, MdMPK4-14G is activated due to its high abundance, which results in MdERF17 phosphorylation, thereby promoting fruit degreening through the activation of downstream genes. The results of our study demonstrate that the transcriptional regulation of MdMPK4-14G and the abundance of MdMPK4-14G follow a diurnal rhythm and that MdMPK4 phosphorylates and activates MdERF17 under dark conditions, which is consistent with the rapid degradation of Chl in the fruit peel. These findings reveal the molecular mechanisms by which two MdMPK4 proteins regulate Chl breakdown and anthocyanin accumulation in response to light/dark transitions (Figure 9).
Figure 9.
Model of Chl degradation controlled by MdMPK4-14G-mediated phosphorylation of MdERF17 in apple fruits. In the dark, MdMYB1 is ubiquitinated and degraded by the 26S proteasome pathway. In contrast, MdERF17 is phosphorylated by dark-activated MdMPK4-14G, which results in high MdERF17 transcriptional activity and the subsequent induction of the expression of target genes, such as MdPPH and MdNYC, thus promoting Chl degradation. Upon light exposure, MdMYB1 is stabilized by MdMPK4-06G-mediated phosphorylation, and activated MdMYB1 binds to its target gene promoters to promote anthocyanin accumulation. MdERF17 phosphorylation is reduced by lower MdMPK4-14G expression and prevented from binding to its target genes for Chl degradation.
We previously showed that different numbers of Ser repeats within the MdERF17 affect the basal transcriptional regulatory activity of MdERF17 and that the Ser repeat number is a key factor in the binding of MdERF17 to the promoters of Chl degradation genes (Han et al., 2018). Our results showed that repeat Ser insertions can enhance transcriptional regulatory activity; however, the mechanistic basis by which Ser repeat number affects the transcriptional regulatory activity of MdERF17 remains unclear. Phosphorylation may modify protein structure and/or conformation, thereby activating or inhibiting enzymatic activity, altering its interactions with other molecules (e.g. proteins, DNA, and RNA), affecting its subcellular localization or modifying its stability. Our SPR study on recombinant proteins showed that the increased Ser repeat number in MdERF17 enhances its interaction with MdMPK4. We performed analyses in plants (parents and progeny) with varying numbers of Ser residues to characterize the relationship between Ser residues and phosphorylation accumulation. Overall, we showed that the number of Ser repeats affects the transcriptional regulatory activity of ERF17 through phosphorylation of the Thr67 residue. We further showed that differences in the number of Ser repeats in ERF17 affect the color of fruit, which is affected by phosphorylated MdMPK4. This finding may prove useful for genetic marker-assisted breeding.
Several lines of evidence suggest that MAPK cascades are involved in the regulation of plant light responses, including observations that the transcription of several Arabidopsis genes involved in MAPK cascades is regulated by light. For example, the Arabidopsis MKK3-MPK6 cascade is activated in response to blue light in a MYC2-dependent manner and regulates the expression of HY5, GBF1, and SPA1 (Sethi et al., 2014). Similarly, the high light-induced expression of the copper/zinc superoxide dismutase genes CSD1 and CSD2 is mediated by MKK5 (Xing et al., 2013). MKK3 acts as a negative regulator of MAPKs in darkness and in light-induced MAPK activation during dark/light transitions (Lee, 2015). MPK4 is activated by light treatment and participates in light-induced anthocyanin accumulation through phosphorylation of MYB75, thereby increasing MYB75 stability in Arabidopsis (Li et al., 2016a, 2016b). Our recent study showed that apple MPK4 phosphorylates MYB1 to enhance light-induced anthocyanin accumulation in fruit peels (Yang et al., 2021). Although phosphorylation is known to be crucial for plant light responses by promoting transcription factor stability, the kinases that phosphorylate regulatory transcription factors involved in light signaling are not themselves associated with light/dark transitions. Anthocyanin biosynthesis is enhanced by sunlight, and dark treatment can promote the degradation of Chl. We showed that MdMPK4s act as molecular bridges in these processes during apple fruit ripening by linking Chl degradation to color changes of peels. Our study indicates that light/dark transitions and MdMPK4-14G are important for controlling MdERF17 phosphorylation and that a greater abundance of phosphorylated MdERF17 mediated by MdMPK4-14G at night contributes to the regulation of Chl degradation in apple peels.
The families of plant MAPK components have expanded during evolution, and different combinations of components accomplish particular functions (Lampard et al., 2008; Sethi et al., 2014; Li et al., 2016a, 2016b; Chen et al., 2021). The apple genome harbors two MdMPK4 genes, MdMPK4-06G and MdMPK4-14G, as a consequence of genome duplication, and these two genes share 96% sequence identity. Given that light-induced phosphorylation of transcription factors occurs mostly at Ser/Thr sites (Hardtke et al., 2000; Ni et al., 2013), we hypothesized that the levels of anthocyanin pigments are regulated by ERF17 phosphorylation in response to light/dark stimuli. We previously showed that MdMPK4-06G expression is more responsive to light treatment than MdMPK4-14G in apple fruit (Yang et al., 2021) and that the MdMPK4-14G promoter contains a unique “CAANNNNATC” cis-acting regulatory element involved in circadian control. We showed here that MdMPK4-14G exhibits a diurnal expression profile, wherein its expression was lower during the day and higher at night and that increasing levels of phosphorylated MdERF17 at night are necessary for its function during fruit ripening. As a result, cooperation between MdMPK4-06 and MdMPK4-14G increases fruit coloration during the day and enhances Chl degradation at night. A recent whole-genome duplication (WGD) event might have played a key role in shaping the genome of the domesticated apple (Velasco et al., 2010), and successive WGDs have generated more genes, which provide an opportunity for functional innovations to arise via dosage effects, neofunctionalization, or subfunctionalization (Blanc and Wolfe, 2004; Pontes et al., 2004; Madlung et al., 2005; Ma et al., 2021; Cusack and Wolfe, 2007; Freeling, 2009; Bekaert et al., 2011; Hudson et al., 2011). Our results indicate that the subfunctionalization of MdMPK4 genes involved in the regulation of fruit degreening and peel coloration might make fruits more attractive for frugivores and thus increase seed dispersal.
We previously showed that tomato fruits are red and that SolERF017 in this species contains five Ser repeats. However, S. pennellii fruits are green, and SolERF017 in this species encodes four Ser repeats (Han et al., 2018). There is allelic variation in ERF17 in other species, as well as variation in the number of Ser residues before the AP2/ERF domain in Arabidopsis, tobacco, tomato, orange, and pear, suggesting that the ERF17 phosphorylation sites recognized by MPK4-14G may be conserved in multiple species and possibly related to the Ser repeats located before the AP2/ERF domain. However, such regulatory systems vary among taxa, and the overall regulatory landscape is likely more complex. For example, the Thr67 phosphorylation site and Ser repeats are missing in peach. An allele of the STAY-GREEN (SGR) gene, which has been shown in other species to encode a key enzyme for Chl degradation (Matsuda et al., 2016; Shimoda et al., 2016), was recently shown to simultaneously regulate carotenoid biosynthesis and Chl degradation in citrus (Zhu et al., 2021). Thus, the range of conserved and independent regulatory mechanisms for Chl degradation in the fruit peel of different fruits has not yet been fully characterized and requires further investigation.
Materials and methods
Plant materials
Apple (Malus domestica cv. “Gala”) fruits were collected from the Northern suburb farm (Beijing, China). Fruits were harvested at 20, 40, 60, 80, and 100 DAFB. Eighty-DAFB “Gala” fruits were subjected to different light treatments by placing the fruit in continuous light or constant darkness for 10 days. The parent cultivars Malus asiatica “Zisai Pearl” and M. domestica “Red Fuji” and their progeny were used to compare the protein levels of different MdERF17 genotypes. Fruits were harvested at 50, 80, 110, 140, and 170 DAFB. Fruit peels were collected and frozen in liquid nitrogen for later analysis, which was conducted in three biological replicates. “Gala” fruits, apple fruit calli, and N. benthamiana plants were used for Agrobacterium (A. tumefaciens)-mediated transformation. The apple calli were subcultured in Murashig and Skoog medium containing 1.5 mg L−1 2,4-dichlorophenoxyacetic acid (2,4-D) and 0.4 mg L−1 6-benzylaminopurine (6-BA) at 24°C in the dark. N. benthamiana plants were cultivated in a growth chamber at 23°C ± 1°C and 50% humidity (100 µmol m−2 s−1 with fluorescent lamps) with a 16-h light/8-h dark cycle.
Chl measurements
Chl was extracted from fresh fruit peels in 80% (v/v) acetone. Total Chl contents were quantified spectrophotometrically at 645 and 663 nm as previously described (Porra et al., 1989).
RNA extraction, cDNA synthesis, and expression analysis
Total RNA was extracted using a modified cetyltrimethylammonium bromide method (Gasic et al., 2004). First-strand cDNA was synthesized from 2 µg of total RNA using the TRUEscript RT Kit (Aidlab, Beijing, China). Gene expression was measured by RT-qPCR analysis on an ABI QuantStudioTM 6 Flex system (Applied Biosystems Inc., Foster City, CA, USA); the Premix Ex Taq kit (Probe qPCR) (TaKaRa, Kyoto, Japan) was used for MdMPK4, and the SensiFASTTM SYBR Lo-ROX Kit (Bioline, London, UK) was used for the other genes. The thermal cycling conditions consisted of an initial denaturation step of 2 min at 95°C, followed by 40 cycles of 95°C for 15 s, 60°C for 30 s, and 72°C for 30 s. Relative expression values were normalized to MdACTIN (MD12G1140800) using the 2 (–ΔΔCt) method (Livak and Schmittgen, 2001). The gene-specific primers are listed in Supplemental Table S3.
Antibody production
The synthetic peptides SGEPSERNDVKYKG and FMTEYVVTRWYRAP, corresponding to MdERF17 and MdMPK4, respectively, were conjugated with keyhole limpet hemocyanin. Two rabbits were immunized with each antigen, and test bleeds were performed 7 days after each boost using an enzyme-linked immunosorbent assay to assess the immune response. Polyclonal antibodies were purified from rabbit serum using affinity chromatography, which detected the purified GST-tagged MdERF17 or MdMPK4 protein. Meanwhile, MdMPK4 proteins were identified by LC–MS/MS analysis. Polyclonal rabbits antibodies were also raised against phospho-peptides encompassing the MdMPK4 Thr202/Tyr204 phosphorylation site (FMTEYVVTRWYRAP) and the MdERF17 Thr67 phosphorylation site (LGSYDTPEKAARAF). Purification with the phosphorylated peptides and non-phosphorylated peptides (cross absorption) was performed to isolate antibodies that specifically recognize the phosphorylated form.
Protein extraction and immunoblot analysis
Total proteins were extracted from apple calli using the Plant Protein Extraction Kit (CWbiotech, Beijing, China). For apple fruit peels, a phenol extraction method was used that is optimized for recalcitrant plant tissues or organs such as peels (Faurobert et al., 2007). Briefly, apple fruit pericarp was frozen in liquid nitrogen after harvest and ground into a fine powder using precooled steel cylinders. The ground tissue was suspended in an equal volume of extraction buffer (500 mM Tris–HCl, 50 mM EDTA, 700 mM sucrose, 100 mM KCl, pH 8.0) and phenol (Solarbio, Beijing, China). The extraction buffer was added to the phenol solution for repeated extraction before precipitation with precipitation solution (0.1 M ammonium acetate in cold methanol) and dissolution in 8 M urea. The proteins were separated by 10% (w/v) sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS–PAGE) and transferred to a polyvinylidene fluoride (PVDF) membrane (Millipore, Burlington, MA, USA; 0.45 µm; IPVH00010). Anti-Myc (Abmart, Shanghai, China; M20002; diluted 1:5,000 in Tris-buffered saline with 0.1% Tween-20 detergent [TBST]), anti-GFP (Abmart; M20004; diluted 1:5,000 in TBST), anti-HA (Abmart; M20003; diluted 1:5,000 in TBST), anti-GST (Abcam, Cambridge, MA, USA; ab92; diluted 1:2,000 in TBST), anti-Actin (Abmart; M20009; diluted 1:2,000 in TBST), anti-MdMPK4 (Beijing QiWei YiCheng Tech Co., Ltd, Beijing, China; diluted 1:2,000 in TBST), anti-Phos-MdMPK4 (Beijing QiWei YiCheng Tech Co., Ltd; diluted 1:2,000 in TBST), anti-MdERF17 (GenScript, Piscataway, NJ, USA; diluted 1:500 in TBST), anti-Phospho-p44/42 MAPK (Erk1/2) (Thr202/Tyr204) (Cell Signaling Technology, Danvers, MA, USA; 4370; diluted 1:2,000 in TBST), and anti-Phos-MdERF17 (GenScript; diluted 1:500 in TBST) were used as primary antibodies. Peroxidase-conjugated goat anti-rabbit (Abmart; M21002) or goat anti-mouse (Abmart; M21001) antibodies were used as second antibodies at 1:4,000 dilution in TBST. Membranes were developed with an eECL Western blot kit (CWbiotech, Beijing, China), and the signal was detected by chemiluminescence imaging (Amersham Imager 680). Main figures show one representative experiment; replicates are provided as Supplemental File S1.
Protein production and purification
For recombinant protein production, the MdMPK4D198G,D200G;E202A,E204A (CA) coding sequence was amplified and introduced into the pGEX4T-2 vector (Yang et al., 2021). MdERF17- was generated by site-directed mutagenesis PCR and confirmed by sequencing. The full-length coding sequences of MdERF17-8S and MdERF17- were inserted into the pGEX6P-1 vector to generate GST-tag fusion proteins. The primers used are listed in Supplemental Table S3. All constructs were transformed into Escherichia coli strain BL21, and cells were induced with 0.3-mM isopropyl-β-d-thiogalactopyranoside overnight at 16°C. The GST-tagged recombinant proteins were purified using glutathione-sepharose resin (CWbiotech, Beijing, China) according to the manufacturer’s instructions.
BIFC and SPR analysis
The coding sequences of the CA variants MdMPK3-03GE192G;E196A, MdMPK3-11GE192G;E196A, MdMPK6-02GD224G;E228A, MdMPK6-15GD227G;E231A, MdMPK4-06GD198G;E202A, MdMPK4-14GD200G;E204A, MdERF17-3S, and MdERF17-8S were cloned in-frame with that of YFP fragments in the pSPYNE and pSPYCE vectors to generate the CA-MdMPK3-03G-nYFP, CA-MdMPK3-11G-nYFP, CA-MdMPK6-02G-nYFP, CA-MdMPK6-15G-nYFP, CA-MdMPK4-06G-nYFP, CA-MdMPK4-14G-nYFP, MdERF17-3S-cYFP, and MdERF17-8S-cYFP constructs (Hu et al., 2020). The constructs were introduced into Agrobacterium strain GV3101 and transiently co-infiltrated into N. benthamiana leaves. After infiltration, plants were grown for 48 h before observation. YFP fluorescence was visualized using a confocal laser scanning microscope (Olympus FV3000, Tokyo, Japan). The accumulation of fusion proteins was detected by immunoblot analysis with anti-HA and anti-MYC antibodies, respectively.
In vitro SPR assays were performed as previously described (Han et al., 2018). Briefly, SPR measurements were carried out using different concentrations (0.094, 0.187, 0.375, 0.75, 1.5, and 3 μM) of purified recombinant MdERF17-8S and MdERF17-3S (0.0625, 0.125, 0.25, 0.5, 1, and 2 μM) proteins on a Biacore T200 platform with CA-MdMPK4 proteins immobilized on CM5 sensor chips (GE Healthcare) by amine coupling. KD values between CA-MdMPK4 and MdERF17 were calculated using Scrubber2-T200 software (BioLogic Software Pty Ltd.) with a steady-state affinity model.
In vitro phosphorylation and LC–MS/MS analysis
In vitro kinase assays were performed as previously described (Sasabe et al., 2011). Briefly, recombinant GST-MdERF17-8S, or the mutant version GST-MdERF17-, was incubated with activated GST-CA-MdMPK4 in kinase reaction buffer (25 mM Tris–HCl pH 7.5, 10 mM MgCl2, 1 mM CaCl2, 1 mM DTT, 1 mM ATP) at 30°C for 40 min. The reaction was stopped by adding 4 × SDS loading buffer and boiling at 100°C for 5 min. Reaction products were subjected to electrophoresis on 10% (w/v) SDS–PAGE, followed by either CBB staining prior to LC–MS/MS analysis (Beijing Biotech-Pack Scientific) or immunoblotting with anti-phosphoserine/threonine antibodies (ECM, PP2551; 1:2,000 dilution).
The Coomassie-stained bands corresponding to purified GST-MdERF17 based on its the predicted molecular weight were excised from the SDS–PAGE gel and treated with trypsin, and the GST-MdERF17 protein phosphorylated by GST-CA-MdMPK4 in vitro was subjected to standard in-solution chymotryptic digestion. After enzymatic digestion, surfactants were removed by adding extraction solution (5% [v/v] trifluoroacetic acid and 50% [v/v] acetonitrile and 45% [v/v] ddH2O). The extracted peptides were lyophilized to near dryness and then resuspended in 10 μL of 0.1% (v/v) formic acid prior to LC–MS/MS analysis, which was performed using a Q-Exactive mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA) with a nanoelectrospray ion source was equipped with a nanoflow HPLC instrument (EASY-nLC1200 system, Thermo Fisher Scientific). Peptides were separated on a 150 μm × 150 mm in-house made column (1.9 μm, 100 Å, Dr Maisch GmbH, Germany) at a flow rate of 600 nL/min while running a 6%–95% acetonitrile gradient and were sprayed directly into the mass spectrometer. The top 20 most intense peptide ions from the preview scan in the Orbitrap were acquired. The phosphopeptides were identified and quantified based on the extracted ion chromatogram peak area using MaxQuant (1.6.2.10) with carbamidomethylation as a fixed modification and phosphorylation and oxidation as variable modifications.
In vivo phosphorylation assay
Full-length MdERF17 and MdERF17T67A coding sequences were cloned into the pSuper1300 vector to generate the overexpression plasmids pSuper1300-MdERF17-Myc and pSuper1300-MdERF17T67A-Myc (Liang et al., 2020). The MdMPK4-14GD200G;E204A coding sequence was ligated into the pSuper1300 vector to generate the construct pSuper1300-CA-MdMPK4-14G-GFP. Agrobacterium GV3101 cultures containing different combinations of these constructs were transiently co-infiltrated into N. benthamiana leaves, with wild-type plants used as a negative control. After 20 h post infiltration, proteins were extracted from the infiltrated leaves and resuspended in extraction buffer at 4°C (50 mM Tris–HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1% [v/v] Triton X-100, 20% [v/v] glycerol, 1 × Complete protease inhibitor cocktail [Sigma-Aldrich, St Louis, MO, USA] and PhosSTOP [Roche, Basel, Switzerland]). λPP was purchased from New England Biolabs, and treatments were conducted by adding λPP with reaction buffer to total proteins as per the manufacturer’s instructions. MdERF17, MdERF17T67A, and MdMPK4-14G proteins were detected by immunoblot analysis using anti-Myc (Abmart, M20002; diluted 1:5,000 in TBST) or anti-GFP (Abmart, M20004; diluted 1:5,000 in TBST) antibodies. In planta phosphorylated MdERF17 levels were determined by immunoblotting using anti-Phos-MdERF17 antibodies (GenScript; 1:500 dilution).
Dual-LUC reporter assay
A 1,500-bp promoter region upstream of the translation start site of MdNYC and MdPPH were amplified by PCR and cloned into the pGreenII0800-LUC vector to generate the reporter constructs (Liang et al., 2020). The 35Spro:MdMPK4-14G, 35Spro:MdERF17, and 35Spro:MdERF17T67A constructs were used as effectors, which were co-infiltrated with their respective reporters into N. benthamiana leaves by Agrobacterium-mediated transient infiltration. The Renilla LUC gene (REN) under the control of the 35S promoter in the pGreenII 0800-LUC vector was used as the internal control. After 48-h infiltration, the infiltrated areas of N. benthamiana leaves were ground to a fine powder in liquid nitrogen and resuspended in 1 × Passive Lysis Buffer. The LUC/REN ratios were calculated using the Dual-Luciferase Reporter Assay System according to the manufacturer’s instructions (Promega, Madison, WI, USA) on a Glo-Max 20/20 luminometer (Promega).
Agrobacterium-mediated transient transformation
To generate overexpression constructs, the coding sequences of CA-MdMPK4-14G, MdERF17, and MdERF17T67A were amplified and inserted into the pRI101 or pSuper1300 vectors to obtain the pRI101-CA-MdMPK4-14G, pSuper1300-MdERF17-Myc, and pSuper1300-MdERF17T67A-Myc constructs (Li et al., 2016a, 2016b; Liang et al., 2020). To reduce MdMPK4 transcript levels in apple fruit, VIGS-mediated gene silencing was performed by inserting a partial coding sequence specific to MdMPK4-14G into the pTRV2 virus vector to generate the TRV-MdMPK4-14G construct. Agrobacterium strain GV3101 cells containing the indicated plasmids were collected by centrifugation at 5,000 rpm for 10 min and resuspended in infiltration buffer (10 mM MES, 10-mM MgCl2, 100-μM acetosyringone, pH 5.6). The pTRV1 and pTRV2 cultures were mixed in a 1:1 (v/v) ratio before infiltration. Agrobacterium harboring the pRI101-CA-MdMPK4-14G or TRV-MdMPK4-14G constructs was subsequently transformed into “Gala” fruit at 75 and 90 DAFB, respectively, and the pSuper1300-MdERF17-Myc and pSuper1300-MdERF17T67A-Myc overexpression vectors were transformed into 150-DAFB “Granny Smith” fruit by immersing fruit in the bacterial suspension and infiltrating under a vacuum of –80 kPa for 2 min twice. The infiltrated fruits were then placed in a phytotron at 24°C under 14-h light/10-h dark or 24-h constant dark conditions and subsequently used for phenotypic examination and RNA and protein extraction. For the 14-h light/10-h dark treatment, the illumination started at 5:30 am and stopped at 7:30 pm, which was consistent with external sunrise and sunset times, respectively.
Construction of transgenic tobacco plants
The coding sequences of MdMPK4-14G and CA-MdMPK4-14G were inserted into the pCAMBIA2301 vector harboring the CaMV 35S promoter to generate the 35S:MdMPK4-14G and 35S: CA-MdMPK4-14G overexpression plasmids (Zhang et al., 2017). These plasmids were transformed into N. benthamiana plants to construct OE-MdMPK4-14G and OE-CA-MdMPK4-14G using the leaf disc method (Horsch et al., 1985); a medium containing kanamycin (30 μg mL−1) was then used to select transformants. T3 homozygous lines expressing high levels of MdMPK4 were used for experiments.
Generation of transgenic apple “Orin” calli
The full-length coding sequences of MdMPK4-14G and CA-MdMPK4-14G were amplified using the primers listed in Supplemental Table S3 and cloned into the pSuper1300 expression vector to generate the pSuper1300-MdMPK4-14G-GFP and pSuper1300-CA-MdMPK4-14G-GFP overexpression plasmids. Subsequently, the resulting vectors were transformed into “Orin” calli as previously described (Li et al., 2021).
Statistical analyses
Data were analyzed using one-way ANOVA followed by Tukey’s honestly significant difference test for multiple comparisons or Student’s t test for two independent samples (*P < 0.05 and **P < 0.01). SPSS Statistics version 22.0, GraphPad Prism version 8.0, and Microsoft Excel 2019 were used for data analyses. A summary of all analyses is given in Supplemental Data Set S1.
Accession numbers
Sequence data from this article can be found in the GenBank/EMBL libraries under the following accession numbers: ERF17-3S (MF498883), ERF17-8S (MF498882), MdMPK4-06G (XP_028960537), and MdMPK4-14G (XP_028948944).
The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD030037.
Supplemental data
The following materials are available in the online version of this article.
Supplemental Figure S1. Immunoblot analysis with Phospho-p44/42 MAPK (Erk1/2) antibodies to analyze the relationship between MAPK activity and fruit degreening during fruit ripening.
Supplemental Figure S2. BiFC assays show that MdERF17 does not interact with MdMPK3 or MdMPK6 in N. benthamiana leaves.
Supplemental Figure S3. Protein abundance of MdERF17, MdMPK3 and MdMPK6 fusion proteins detected by immunoblot analysis in BIFC assays.
Supplemental Figure S4. Protein abundance of MdERF17 and MdMPK4 fusion proteins detected by immunoblot analysis in BIFC assays.
Supplemental Figure S5. SPR analysis of the interaction between MdMPK4 and GST, which was used as negative control.
Supplemental Figure S6. LC–MS/MS analysis of MdERF17 protein bands separated by SDS-PAGE.
Supplemental Figure S7. Analysis of the specificity of anti-Phos-MdERF17 antibodies using CA-MdMPK4, MdERF17-8S, and MdERF17- with an in vitro kinase assay.
Supplemental Figure S8. Detection of MdERF17 transcripts in peels of “Granny Smith” and “Gala” fruit.
Supplemental Figure S9. Analysis of Chl degradation rate and protein abundance in “Zisai Pearl” × “Red Fuji” F1 hybrids.
Supplemental Figure S10. MdMPK4-14G affects phosphorylated MdERF17 under 14-h light/10-h dark conditions.
Supplemental Figure S11. MdMPK4-14G contributes to Chl degradation in N. benthamiana leaves.
Supplemental Figure S12. Effects of light and dark treatments on fruit peel degreening.
Supplemental Table S1. Analysis of MdERF17 protein bands extracted from an SDS–PAGE gel by LC–MS/MS.
Supplemental Table S2. MdERF17 phosphopeptide information.
Supplemental Table S3. List of primers used in this study.
Supplemental Data Set S1. ANOVA and Student's t-test Tables.
Supplemental File S1. Uncropped immunoblots and replicate experiments.
Supplementary Material
Acknowledgments
We thank PlantScribe (www.plantscribe.com) for editing this manuscript.
Funding
The work was supported by the National Key R&D Program of China (2018YFD1000200), the National Natural Science Foundation of China (no. 31872941, 32072543), the 111 Project (B17043), and the 2115 Talent Development Program of China Agricultural University.
Conflict of interest statement. The authors declare that they have no conflict of interest.
Contributor Information
Shuai Wang, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Ting Wang, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Qiqi Li, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Chen Xu, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Ji Tian, Plant Science and Technology College, Beijing University of Agriculture, Beijing, China.
Yi Wang, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Xinzhong Zhang, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Xuefeng Xu, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Zhenhai Han, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
Ting Wu, College of Horticulture, State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China.
T.W. and Z.H. conceived and designed the research. S.W., T.W., Q.L., and C.X. conducted the experiments. J.T., Y.W., X.Z., and X.X. contributed reagents and analytical tools. T.W. and S.W. wrote the manuscript. All authors read and approved the manuscript.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (https://academic.oup.com/plcell) is Ting Wu (wuting@cau.edu.cn).
References
- Albert NW, Davies KM, Lewis DH, Zhang H, Montefiori M, Brendolise C, Boase MR, Ngo H, Jameson PE, Schwinn KE (2014) A conserved network of transcriptional activators and repressors regulates anthocyanin pigmentation in Eudicots. Plant Cell 26: 962–980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Allan AC, Hellens RP, Laing WA (2008) MYB transcription factors that colour our fruit. Trends Plant Sci 13: 99–102 [DOI] [PubMed] [Google Scholar]
- An X, Tian Y, Chen K, Liu X, Liu D, Xie X, Cheng C, Cong P, Hao Y (2015) MdMYB9 and MdMYB11 are involved in the regulation of the JA-induced biosynthesis of anthocyanin and proanthocyanidin in apples. Plant Cell Physiol 56: 650–662 [DOI] [PubMed] [Google Scholar]
- An X, Tian Y, Chen K, Wang X, Hao Y (2012) The apple WD40 protein MdTTG1 interacts with bHLH but not MYB proteins to regulate anthocyanin accumulation. J Plant Physiol 169: 710–717 [DOI] [PubMed] [Google Scholar]
- Andreasson E, Ellis B (2010) Convergence and specificity in the Arabidopsis MAPK nexus. Trends Plant Sci 15: 106–113 [DOI] [PubMed] [Google Scholar]
- Balazadeh S, Siddiqui H, Allu AD, Matallana-Ramirez LP, Caldana C, Mehrnia M, Zanor M, Köhler B, Mueller-Roeber B (2010) A gene regulatory network controlled by the NAC transcription factor ANAC092/AtNAC2/ORE1 during salt-promoted senescence. Plant J 62: 250–264 [DOI] [PubMed] [Google Scholar]
- Barsan C, Sanchez-Bel P, Rombaldi C, Egea I, Rossignol M, Kuntz M, Zouine M, Latché A, Bouzayen M, Pech J (2010) Characteristics of the tomato chromoplast revealed by proteomic analysis. J Exp Bot 61: 2413–2431 [DOI] [PubMed] [Google Scholar]
- Bekaert M, Edger PP, Pires JC, Conant GC (2011) Two-phase resolution of polyploidy in the Arabidopsis metabolic network gives rise to relative and absolute dosage constraints. Plant Cell 23: 1719–1728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bergmann DC, Lukowitz W, Somerville CR (2004) Stomatal development and pattern controlled by a MAPKK kinase. Science 304: 1494–1497 [DOI] [PubMed] [Google Scholar]
- Berriri S, Garcia AV, Dit Frey NF, Rozhon W, Pateyron S, Leonhardt N, Montillet J, Leung J, Hirt H, Colcombet J (2012) Constitutively active mitogen-activated protein kinase versions reveal functions of Arabidopsis MPK4 in pathogen defense signaling. Plant Cell 24: 4281–4293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blanc G, Wolfe KH (2004) Functional divergence of duplicated genes formed by polyploidy during Arabidopsis evolution. Plant Cell 16: 1679–1691 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boutrot F, Segonzac C, Chang KN, Qiao H, Ecker JR, Zipfel C, Rathjen JP (2010) Direct transcriptional control of the Arabidopsis immune receptor FLS2 by the ethylene-dependent transcription factors EIN3 and EIL1. Proc Natl Acad Sci USA 107: 14502–14507 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen M, Chory J, Fankhauser C (2004) Light signal transduction in higher plants. Annu Rev Genet 38: 87–117 [DOI] [PubMed] [Google Scholar]
- Chen M, Ni L, Chen J, Sun M, Qin C, Zhang G, Zhang A, Jiang M (2021) Rice calcium/calmodulin-dependent protein kinase directly phosphorylates a mitogen-activated protein kinase kinase to regulate abscisic acid responses. Plant Cell 33: 1790–1812 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho SK, Larue CT, Chevalier D, Wang H, Jinn T, Zhang S, Walker JC (2008) Regulation of floral organ abscission in Arabidopsis thaliana. Proc Natl Acad Sci USA 105: 15629–15634 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choudhary MK, Nomura Y, Wang L, Nakagami H, Somers DE (2015) Quantitative circadian phosphoproteomic analysis of Arabidopsis reveals extensive clock control of key components in physiological, metabolic, and signaling pathways. Mol Cell Proteomics 14: 2243–2260 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cusack BP, Wolfe KH (2007) When gene marriages don’t work out: divorce by subfunctionalization. Trends Genet 23: 270–272 [DOI] [PubMed] [Google Scholar]
- Espley RV, , HellensRP, , Putterill J,, StevensonDE, , Kutty-Amma S,, Allan AC (2007) Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 49: 414–427 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Espley RV, Brendolise C, Chagné D, Kutty-Amma S, Green S, Volz R, Putterill J, Schouten HJ, Gardiner SE, Hellens RP, et al. (2009) Multiple repeats of a promoter segment causes transcription factor autoregulation in red apples. Plant Cell 21: 168–183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faurobert M, Pelpoir E, Chaib J (2007) Phenol extraction of proteins for proteomic studies of recalcitrant plant tissues. Methods Mol Biol 355: 9–14 [DOI] [PubMed] [Google Scholar]
- Francisco M, Kliebenstein DJ, Rodríguez VM, Soengas P, Abilleira R, Cartea ME (2021) Fine mapping identifies NAD-ME1 as a candidate underlying a major locus controlling temporal variation in primary and specialized metabolism in Arabidopsis. Plant J 106: 454–467 [DOI] [PubMed] [Google Scholar]
- Freeling M (2009) Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental, or by transposition. Annu Rev Plant Biol 60: 433–453 [DOI] [PubMed] [Google Scholar]
- Fujisawa M, Nakano T, Shima Y, Ito Y (2013) A large-scale identification of direct targets of the tomato MADS Box transcription factor RIPENING INHIBITOR reveals the regulation of fruit ripening. Plant Cell 25: 371–386 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao S, Gao J, Zhu X, Song Y, Li Z, Ren G, Zhou X, Kuai B (2016) ABF2, ABF3, and ABF4 promote ABA-mediated chlorophyll degradation and leaf senescence by transcriptional activation of chlorophyll catabolic genes and senescence-associated genes in Arabidopsis. Mol Plant 9: 1272–1285 [DOI] [PubMed] [Google Scholar]
- Gasic K, Hernandez A, Korban SS (2004) RNA extraction from different apple tissues rich in polyphenols and polysaccharides for cDNA library construction. Plant Mol Biol Rep 22: 437–438 [Google Scholar]
- Giovannoni J, Nguyen C, Ampofo B, Zhong S, Fei Z (2017) The Epigenome and transcriptional dynamics of fruit ripening. Annu Rev Plant Biol 68: 61–84 [DOI] [PubMed] [Google Scholar]
- Giovannoni JJ (2004) Genetic regulation of fruit development and ripening. Plant Cell 16: S170–S180 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53: 814–827 [DOI] [PubMed] [Google Scholar]
- Han Z, Hu Y, Lv Y, Rose JKC, Sun Y, Shen F, Wang Y, Zhang X, Xu X, Wu T, et al. (2018) Natural variation underlies differences in ETHYLENE RESPONSE FACTOR17 activity in fruit peel degreening. Plant Physiol 176: 2292–2304 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hardtke CS, Gohda K, Osterlund MT, Oyama T, Okada K, Deng XW (2000) HY5 stability and activity in Arabidopsis is regulated by phosphorylation in its COP1 binding domain. EMBO J 19: 4997–5006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harmer SL, Hogenesch JB, Straume M, Chang HS, Han B, Zhu T, Wang X, Kreps JA, Kay SA (2000) Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science 290: 2110–2113 [DOI] [PubMed] [Google Scholar]
- Harpaz-Saad S, Azoulay T, Arazi T, Ben-Yaakov E, Mett A, Shiboleth YM, Hörtensteiner S, Gidoni D, Gal-On A, Goldschmidt EE, et al. (2007) Chlorophyllase is a rate-limiting enzyme in chlorophyll catabolism and is posttranslationally regulated. Plant Cell 19: 1007–1022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hettenhausen C, Baldwin IT, Wu J (2012) Silencing MPK4 in Nicotiana attenuata enhances photosynthesis and seed production but compromises abscisic acid-induced stomatal closure and guard cell-mediated resistance to Pseudomonas syringae pv tomato DC3000. Plant Physiol 158: 759–776 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7: 1071–1083 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horsch RB, Fry JE, Hoffmann NL, Eichholtz D, Rogers SA, Fraley RT (1985) A simple and general method for transferring genes into plants. Science 227: 1229–1231 [DOI] [PubMed] [Google Scholar]
- Hörtensteiner S (2006) Chlorophyll degradation during senescence. Annu Rev Plant Biol 57: 55–77 [DOI] [PubMed] [Google Scholar]
- Hörtensteiner S (2013) Update on the biochemistry of chlorophyll breakdown. Plant Mol Biol 82: 505–517 [DOI] [PubMed] [Google Scholar]
- Hörtensteiner S, Kräutler B (2011) Chlorophyll breakdown in higher plants. Biochim Biophys Acta 1807: 977–988 [DOI] [PubMed] [Google Scholar]
- Hu Y, Han Z, Sun Y, Wang S, Wang T, Wang Y, Xu K, Zhang X, Xu X, Han Z, et al. (2020) ERF4 affects fruit firmness through TPL4 by reducing ethylene production. Plant J 103: 937–950 [DOI] [PubMed] [Google Scholar]
- Huang D, Wang X, Tang Z, Yuan Y, Xu Y, He J, Jiang X, Peng S, Li L, Butelli E, et al. (2018) Subfunctionalization of the Ruby2-Ruby1 gene cluster during the domestication of citrus. Nat Plants 4: 930–941 [DOI] [PubMed] [Google Scholar]
- Hudson CM, Puckett EE, Bekaert M, Pires JC, Conant GC (2011) Selection for higher gene copy number after different types of plant gene duplications. Genome Biol Evol 3: 1369–1380 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaakola L (2013) New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci 18: 477–483 [DOI] [PubMed] [Google Scholar]
- Jia W, Li B, Li S, Liang Y, Wu X, Ma M, Wang J, Gao J, Cai Y, Zhang Y, et al. (2016) Mitogen-activated protein kinase cascade MKK7-MPK6 plays important roles in plant development and regulates shoot branching by phosphorylating PIN1 in Arabidopsis. PLoS Biol 14: e1002550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kami C, Lorrain S, Hornitschek P, Fankhauser C (2010) Light-regulated plant growth and development. Curr Top Dev Biol 91: 29–66 [DOI] [PubMed] [Google Scholar]
- Kholodenko BN (2006) Cell-signalling dynamics in time and space. Nat Rev Mol Cell Bio 7: 165–176 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komis G, Šamajová O, Ove Ka M, Šamaj J (2018) Cell and developmental biology of plant mitogen-activated protein kinases. Annu Rev Plant Biol 69: 237–265 [DOI] [PubMed] [Google Scholar]
- Lampard GR, Macalister CA, Bergmann DC (2008) Arabidopsis stomatal initiation is controlled by MAPK-mediated regulation of the bHLH SPEECHLESS. Science 322: 1113–1116 [DOI] [PubMed] [Google Scholar]
- Lee H (2015) Mitogen-activated protein kinase kinase 3 is required for regulation during dark-light transition. Mol Cells 38: 651–656 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li D, Sun Q, Zhang G, Zhai L, Li K, Feng Y, Wu T, Zhang X, Xu X, Wang Y, et al. (2021) MxMPK6-2-bHLH104 interaction is involved in reactive oxygen species signaling in response to iron deficiency in apple rootstock. J Exp Bot 72: 1919–1932 [DOI] [PubMed] [Google Scholar]
- Li S, Wang W, Gao J, Yin K, Wang R, Wang C, Petersen M, Mundy J, Qiu J (2016a) MYB75 phosphorylation by MPK4 is required for light-induced anthocyanin accumulation in Arabidopsis. Plant Cell 28: 2866–2883 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li T, Jiang Z, Zhang L, Tan D, Wei Y, Yuan H, Li T, Wang A (2016b) Apple (Malus domestica) MdERF2 negatively affects ethylene biosynthesis during fruit ripening by suppressing MdACS1 transcription. Plant J 88: 735–748 [DOI] [PubMed] [Google Scholar]
- Li Y, Mao K, Zhao C, Zhao X, Zhang H, Shu H, Hao Y (2012) MdCOP1 ubiquitin E3 ligases interact with MdMYB1 to regulate light-induced anthocyanin biosynthesis and red fruit coloration in apple. Plant Physiol 160: 1011–1022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang Y, Jiang C, Liu Y, Gao Y, Lu J, Aiwaili P, Fei Z, Jiang C, Hong B, Ma C, et al. (2020) Auxin regulates sucrose transport to repress petal abscission in Rose (Rosa hybrida). Plant Cell 32: 3485–3499 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L, Shao Z, Zhang M, Wang Q (2015) Regulation of carotenoid metabolism in tomato. Mol Plant 8: 28–39 [DOI] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25: 402–408 [DOI] [PubMed] [Google Scholar]
- Ma H,, Yang T,, Li Y,, Zhang J,, Wu T,, Song T,, Yao Y,, TianJ (2021) . The long noncoding RNA MdLNC499 bridges MdWRKY1 and MdERF109 function to regulate early-stage light-induced anthocyanin accumulation in apple fruit. Plant Cell 33: 3309–3330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madlung A, Tyagi AP, Watson B, Jiang H, Kagochi T, Doerge RW, Martienssen R, Comai L (2005) Genomic changes in synthetic Arabidopsis polyploids. Plant J 41: 221–230 [DOI] [PubMed] [Google Scholar]
- Mao C, Lu S, Lv B, Zhang B, Shen J, He J, Luo L, Xi D, Chen X, Ming F (2017) A Rice NAC transcription factor promotes leaf senescence via ABA biosynthesis. Plant Physiol 174: 1747–1763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuda K, Shimoda Y, Tanaka A, Ito H (2016) Chlorophyll a is a favorable substrate for Chlamydomonas Mg-dechelatase encoded by STAY-GREEN. Plant Physiol Bioch 109: 365–373 [DOI] [PubMed] [Google Scholar]
- Matsumoto F, Obayashi T, Sasaki-Sekimoto Y, Ohta H, Takamiya K, Masuda T (2004) Gene expression profiling of the tetrapyrrole metabolic pathway in Arabidopsis with a mini-array system. Plant Physiol 135: 2379–2391 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller T, Ulrich M, Ongania KH, Kräutler B (2007) Colorless tetrapyrrolic chlorophyll catabolites found in ripening fruit are effective antioxidants. Angew Chem Int Ed 46: 8699–8702 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ni W, Xu S, Chalkley RJ, Pham TND, Guan S, Maltby DA, Burlingame AL, Wang Z, Quail PH (2013) Multisite light-induced phosphorylation of the transcription factor PIF3 is necessary for both its rapid degradation and concomitant negative feedback modulation of photoreceptor phyB levels in Arabidopsis. Plant Cell 25: 2679–2698 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peter E, Grimm B (2009) GUN4 is required for posttranslational control of plant tetrapyrrole biosynthesis. Mol Plant 2: 1198–1210 [DOI] [PubMed] [Google Scholar]
- Pontes O, Neves N, Silva M, Lewis MS, Madlung A, Comai L, Viegas W, Pikaard CS (2004) Chromosomal locus rearrangements are a rapid response to formation of the allotetraploid Arabidopsis suecica genome. Proc Natl Acad Sci USA 101: 18240–18245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Porra RJ, Thompson WA, Kriedemann PE (1989) Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim Biophys Acta 975: 384–394 [Google Scholar]
- Robatzek S, Somssich IE (2001) A new member of the Arabidopsis WRKY transcription factor family, AtWRKY6, is associated with both senescence- and defence-related processes. Plant J 28: 123–133 [DOI] [PubMed] [Google Scholar]
- Rodriguez MCS,, Petersen M,, Mundy J (2010) Mitogen-activated protein kinase signaling in plants. Ann Rev Plant Biol 61: 621–649 [DOI] [PubMed] [Google Scholar]
- Sasabe M, Boudolf V, Veylder LD, Inze D, Genschik P, Machida Y (2011) Phosphorylation of a mitotic kinesin-like protein and a MAPKKK by cyclin-dependent kinases (CDKs) is involved in the transition to cytokinesis in plants. Proc Natl Acad Sci USA 108: 17844–17849 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schelbert S, Aubry S, Burla B, Agne B, Kessler F, Krupinska K, Hortensteiner S (2009) Pheophytin pheophorbide hydrolase (pheophytinase) is involved in chlorophyll breakdown during leaf senescence in Arabidopsis. Plant Cell 21: 767–785 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sethi V, Raghuram B, Sinha AK, Chattopadhyay S (2014) A mitogen-activated protein kinase cascade module, MKK3-MPK6 and MYC2, is involved in blue light-mediated seedling development in Arabidopsis. Plant Cell 26: 3343–3357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shemer TA, Harpaz-Saad S, Belausov E, Lovat N, Krokhin O, Spicer V, Standing KG, Goldschmidt EE, Eyal Y (2008) Citrus chlorophyllase dynamics at ethylene-induced fruit color-break: a study of chlorophyllase expression, posttranslational processing kinetics, and in situ intracellular localization. Plant Physiol 148: 108–118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimoda Y, Ito H, Tanaka A (2016) Arabidopsis STAY-GREEN, mendel’s green cotyledon gene, encodes magnesium-dechelatase. Plant Cell 28: 2147–2160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song Y, Yang C, Gao S, Zhang W, Li L, Kuai B (2014) Age-triggered and dark-induced leaf senescence require the bHLH transcription factors PIF3, 4, and 5. Mol Plant 7: 1776–1787 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steed G, Ramirez DC, Hannah MA, Webb AAR (2021) Chronoculture, harnessing the circadian clock to improve crop yield and sustainability. Science 372: c9141. [DOI] [PubMed] [Google Scholar]
- Takos AM, Jaffé FW, Jacob SR, Bogs J, Robinson SP, Walker AR (2006) Light-induced expression of a MYB gene regulates anthocyanin biosynthesis in red apples. Plant Physiol 142: 1216–1232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar SK, Troggio M, Pruss D, et al. (2010) The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42: 833–839 [DOI] [PubMed] [Google Scholar]
- Xing Y, Cao Q, Zhang Q, Qin L, Jia W, Zhang J (2013) MKK5 regulates high light-induced gene expression of Cu/Zn superoxide dismutase 1 and 2 in Arabidopsis. Plant Cell Physiol 54: 1217–1227 [DOI] [PubMed] [Google Scholar]
- Yamada K, Yamaguchi K, Yoshimura S, Terauchi A, Kawasaki T (2017) Conservation of chitin-induced MAPK signaling pathways in rice and Arabidopsis. Plant Cell Physiol 58: 993–1002 [DOI] [PubMed] [Google Scholar]
- Yang T, Ma H, Li Y, Zhang Y, Zhang J, Wu T, Song T, Yao Y, Tian J (2021) Apple MPK4 mediates phosphorylation of MYB1 to enhance light‐induced anthocyanin accumulation. Plant J 106: 1728–1745 [DOI] [PubMed] [Google Scholar]
- Yang T, Ma H, Zhang J, Wu T, Song T, Tian J, Yao Y (2019) Systematic identification of long noncoding RNAs expressed during light-induced anthocyanin accumulation in apple fruit. Plant J 100: 572–590 [DOI] [PubMed] [Google Scholar]
- Yin X, Xie X, Xia X, Yu J, Ferguson IB, Giovannoni JJ, Chen K (2016) Involvement of an ethylene response factor in chlorophyll degradation during citrus fruit degreening. Plant J 86: 403–412 [DOI] [PubMed] [Google Scholar]
- Yoshida K, Ma D, Constabel CP (2015) The MYB182 protein down-regulates proanthocyanidin and anthocyanin biosynthesis in poplar by repressing both structural and regulatory flavonoid genes. Plant Physiol 167: 693–710 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang M, Su J, Yan Z, Xu J, Zhang S (2018) Conveying endogenous and exogenous signals: MAPK cascades in plant growth and defense. Curr Opin Plant Biol 45: 1–10 [DOI] [PubMed] [Google Scholar]
- Zhang Z, Li J, Li F, Liu H, Yang W, Chong K, Xu Y (2017) OsMAPK3 phosphorylates OsbHLH002/OsICE1 and inhibits Its ubiquitination to activate OsTPP1 and enhances rice chilling tolerance. Dev Cell 43: 731–743 [DOI] [PubMed] [Google Scholar]
- Zhao W, Li Y, Fan S, Wen T, Wang M, Zhang L, Zhao L (2021) The transcription factor WRKY32 affects tomato fruit colour by regulating YELLOW FRUITED-TOMATO 1, a core component of ethylene signal transduction. J Exp Bot 72: 4269–4282 [DOI] [PubMed] [Google Scholar]
- Zhou X, Jiang Y, Yu D (2011) WRKY22 transcription factor mediates dark-induced leaf senescence in Arabidopsis. Mol Cells 31: 303–313 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu G,, Wang S,, Huang Z,, Zhang S,, Liao Q,, Zhang C,, Lin T,, Qin M,, Peng M,, Yang C,et al. Rewiring of the Fruit Metabolome in Tomato Breeding. Cell 172: 249–261.e12 [DOI] [PubMed] [Google Scholar]
- Zhu K, Zheng X, Ye J, Huang Y, Chen H, Mei X, Xie Z, Cao L, Zeng Y, Larkin RM, et al. (2021) Regulation of carotenoid and chlorophyll pools in hesperidia, anatomically unique fruits found only in Citrus. Plant Physiol 187: 829–845 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu X, Chen J, Qiu K, Kuai B (2017) Phytohormone and light regulation of chlorophyll degradation. Front Plant Sci 8: 1911. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.