Summary
Ethical, environmental and health concerns around dairy products are driving a fast‐growing industry for plant‐based dairy alternatives, but undesirable flavours and textures in available products are limiting their uptake into the mainstream. The molecular processes initiated during fermentation by lactic acid bacteria in dairy products is well understood, such as proteolysis of caseins into peptides and amino acids, and the utilisation of carbohydrates to form lactic acid and exopolysaccharides. These processes are fundamental to developing the flavour and texture of fermented dairy products like cheese and yoghurt, yet how these processes work in plant‐based alternatives is poorly understood. With this knowledge, bespoke fermentative processes could be engineered for specific food qualities in plant‐based foods. This review will provide an overview of recent research that reveals how fermentation occurs in plant‐based milk, with a focus on how differences in plant proteins and carbohydrate structure affect how they undergo the fermentation process. The practical aspects of how this knowledge has been used to develop plant‐based cheeses and yoghurts is also discussed.
The mechanisms of fermentation by lactic acid bacteria are reviewed in relation to plant‐based dairy alternatives. Particular attention is paid to proteolytic and carbohydrate metabolism systems, and how these have been studied in plant‐based dairy alternative products is discussed.

Introduction
Fermentation of dairy milk by lactic acid bacteria is well documented, but how the molecular processes involved in fermentation occur in plant‐based milks has received little attention (Hayes et al., 2007; Mende et al., 2016; Ji et al., 2021). The increasing interest in plant‐based alternatives to products such as cheeses and yoghurts is hindered by difficulties in creating products with acceptable textures and flavours (Jaeger and Giacalone, 2021). Understanding how fermentation occurs in plant‐based dairy substitutes and how fermentation influences flavour and texture is key to developing products that are desirable to consumers wishing to switch to plant‐based alternatives.
Consumption of dairy products is declining in many Western countries, while plant‐based milk sales are increasing (Islam et al., 2021). Plant‐based milk now makes up 15% of the total milk industry, and other plant‐based dairy alternatives are also increasing in popularity (Good Food Institute, 2021). There is now a wide range of plant‐based dairy alternative products available, with markets for these products being in the millions or billions of dollars (Fig. 1). Increased awareness around the environmental impacts of dairy farming, ethical concerns, medical reasons such as lactose intolerance and cow’s milk allergy, and an increased perception of milk as being ‘unhealthy’ all contribute to consumers choosing plant‐based alternatives to milk and dairy products (McCarthy et al., 2017). Plant‐based milks use less energy in production and result in emission of less greenhouse gas per gram of protein than animal‐based milks (González et al., 2011), making them an attractive option for those wanting to reduce their carbon footprint. They are also a good alternative for those who are unable to consume cow’s milk for medical reasons as they do not contain lactose or other allergens present in cow’s milk, although some plant‐based milks such as soy or nut‐based milks contain other allergens. They tend to be lower in fat, especially saturated fat, so can be a good option for those looking for lower‐fat alternatives. Additionally, plant‐based milks can be produced so as to contain dietary fibre, adding nutritional benefits not found in dairy milk. On the other hand, current plant‐based milks are generally lower in protein, except soy, which is comparable with dairy milk, and plant‐based milks are commonly fortified with nutrients such as calcium and vitamin B12 to improve nutritional quality [for review see Mäkinen et al. (2016); Clegg et al. (2021)]. Plant‐based milks often contain less complete proteins, for example, cereals can be deficient in lysine, while legumes tend to be deficient in methionine and cysteine (Sim et al., 2021). Strategies to combat this, such as combining plant‐based milk types in a product to provide an overall complete amino acid profile, can be used. Plant proteins also tend to be more difficult to digest. This can be due to antinutritional factor compounds present such as protease inhibitors and non‐starch polysaccharides, as well as properties inherent to the protein structure such as cross‐linking, hydrophobicity and secondary structure elements (Sá et al., 2020).
Fig. 1.
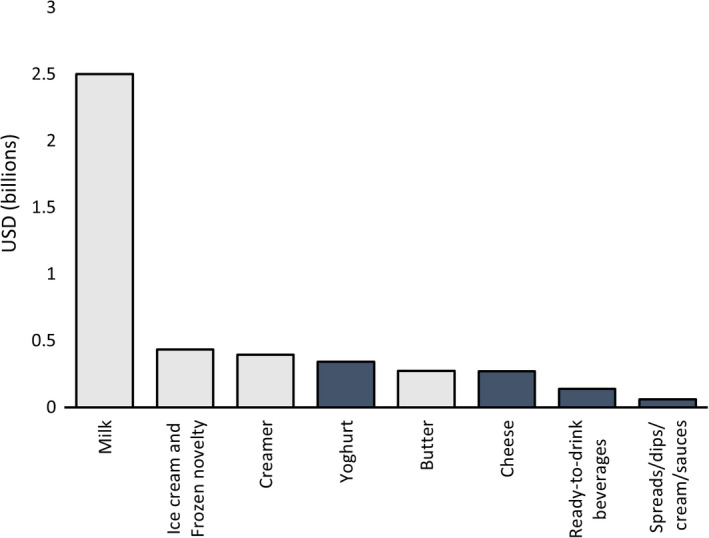
Sales in USD of different categories of plant‐based dairy alternatives in the USA in 2020. Categories in dark grey are those which can utilise fermentation. Adapted from Good Food Institute (2021).
Lactic acid bacteria (LAB) are the group of species responsible for fermentation in dairy‐based foods and have been used in food fermentation for centuries (Bernardeau et al., 2006). In these foods, fermentation can improve the flavour and texture of products. For example, the breakdown of proteins into amino acids and further into volatile organic compounds is one of the main sources of flavour compounds in cheese (Yvon and Rijnen, 2001; Smit et al., 2005), and the production of exopolysaccharides from sugars improves the texture of cheeses and yoghurts (Korcz and Varga, 2021). Fermentation has well‐documented health benefits, such as the introduction of bioactive small molecules including bioactive peptides and, in some cases, can improve the vitamin content (Şanlier et al., 2019). Fermentation can also extend product shelf‐life. LAB acidify the media, making it a less hospitable environment for other microbial organisms, and also produce a range of compounds which inhibit growth of other organisms such as organic acids, hydrogen peroxide and bacteriocins (Saranraj et al., 2013).
One reason for resistance to the adoption of plant‐based milks is the presence of undesirable ‘beany’ flavours and undesirable textures. These undesirable flavours can largely be attributed to aldehydes, primarily hexanal but also 3‐Z‐hexenal, as well as alcohols such as n‐hexanol, n‐pentanol and n‐heptanol, ketones such as ethyl vinyl ketone and furans such as n‐pentyl furan and 2‐(1‐pentenyl) furan (Rackis et al., 1979). Fermentation of plant‐based milks by a range of LAB is able to completely remove hexanal and decrease the concentration of other contributing volatile organic compounds in soy (Blagden and Gilliland, 2005). Antinutritional factors present in plants such as tannins, saponins, phytic acid, α‐galactosides and trypsin inhibitors are also broken down during fermentation by LAB, improving the nutritional quality of plant‐based products (Lopez et al., 2000; Adeyemo and Onilude, 2013). Therefore, fermentation and the development of fermented products could be an important pathway to improving the nutritional quality and acceptability of plant‐based milk products.
This review will discuss how the fermentation of plant proteins has been used to produce plant‐based dairy alternative products and highlight the gaps in understanding the molecular‐level processes that are occurring during fermentation. The current understanding of how molecular processes such as proteolysis and exopolysaccharide formation occur during the fermentation of dairy milk products will be discussed before relating this knowledge to plant‐based systems, in order to compare the differences between the two systems. This will help to demonstrate why the development of plant‐based dairy alternatives has faced difficulties in mimicking the flavour and texture profiles of dairy‐based products and will highlight potential avenues that could be exploited to improve plant‐based dairy products.
Fermentation processes
Proteolysis, lipolysis and carbohydrate metabolism are the major metabolic processes involved in the fermentation of dairy milk. Proteolysis results in the development of many of the major flavour compounds in hard cheeses (Fox et al., 1996; Smit et al., 2005). Carbohydrate metabolism produces lactic acid, which is the main driver of acidification of the product, diacetyl and acetaldehyde which are key flavour compounds, and exopolysaccharides, which are important for the development of flavour and texture, especially in yoghurts. Lipolysis is critical for flavour development in dairy cheeses, where lipases hydrolyse milk fats to release free fatty acids. These free fatty acids can be flavour compounds themselves and are more importantly precursors to flavour compounds such as methyl ketones, alcohols, lactones and esters [for review, see Jooyandeh et al. (2009)]. However, lipases mostly originate from microbial sources other than LAB and will not be discussed further in this review. For a summary of important fermentative characteristics of some commercially relevant LAB species in the fermented dairy industry, refer to Table 1.
Table 1.
Commercially relevant lactic acid bacteria species and their fermentation characteristics.
| Species name | Bacteria type | Fermentation type | Cell envelope proteinase present | Common commercial usage | Main role in dairy fermentation |
|---|---|---|---|---|---|
| Lactobacillus helveticus a | Thermophilic | Homofermentative | PrtH2 (common to all strains), PrtH, PrtH3, PrtH4 | Cheese (esp. mozzarella, Emmental, Grana Padano), fermented milk. | Most proteolytic LAB species commonly used in dairy fermentation |
| Streptococcus thermophilus b | Thermophilic | Homofermentative | PrtS | Yoghurt starter, cheese, fermented milk. | EPS production in yoghurt |
| Lactobacillus delbrueckii subsp. bulgaricus c | Thermophilic | Homofermentative | PrtB | Yoghurt starter, cheese, fermented milk. | Proteolysis in yoghurt |
| Lactococcus lactis subsp. cremoris d | Mesophilic | Homofermentative | PrtP | Cheese | Development of flavour compounds in cheese |
| Lactococcus lactis subsp. lactis d | Mesophilic | Homofermentative | PrtP | Cheese. | Fast acidification through conversion of lactose to lactic acid |
| Leuconostoc mesenteroides e | Mesophilic | Heterofermentative | n/a | Cheese (esp. Swiss styles, Roquefort) | Produces complex flavours and eye holes in cheese through production of CO2 and diacetyl. |
Proteolytic system
Due to the limited ability of LAB to synthesise amino acids, and the negligible quantities of free amino acids and available peptides present in dairy milk, LAB from milk require an effective proteolytic system (Fig. 2) in order to grow (Teusink and Molenaar, 2017). Cell envelope proteinases are the large, multidomain serine proteases that catalyse the first step of this process, where large proteins are broken down into peptides. Many different cell envelope proteinases have been described, varying in size, specificity for caseins, and domains present. These are designated as PrtX enzymes, where X corresponds to the specific cell envelope proteinase. Some species express the same cell envelope proteinase, for example, Lacticaseibacillus paracasei subsp. paracasei, Lactococcus lactis subsp. cremoris and Lc. lactis subsp. lactis all express PrtP. Lactobacillus helveticus expresses four different PrtH enzymes, and most other LAB express only one unique cell envelope proteinase (Table 1). These are secreted from the cell and anchored to the cell membrane, which ensures that proteolysis occurs close to the cell and allows efficient subsequent transportation. The catalytic domain contains the active site, composed of three amino acid residues (serine, histidine and aspartic acid) that are highly conserved across species (Hansen and Marcatili, 2020). The cell envelope proteinase from Lc. lactis subsp. lactis, PrtP, degrades caseins into peptides ranging from 4 to 30 amino acids, with no di‐ or tripeptides being detected, and only trace amounts of phenylalanine being detected in free amino acid analysis (Juillard et al., 1995). An oligopeptide permease (Opp) transport system then transports oligopeptides across the cell membrane, while an ion‐linked transporter (DtpT) and ABC transporter (Dpp) system transport any di‐ and tri‐peptides produced. Once transported into the cell, the oligopeptides are first broken down into smaller peptides by intracellular endopeptidases, aminopeptidases (PepN and PepC) and X‐prolyl dipeptidyl aminopeptidase (PepX). The generated smaller peptides are then further broken down by dipeptidases and tripeptidases into individual amino acids [for review of the proteolytic system, see Savijoki et al. (2006)]. These amino acids are then further broken down into volatile organic compounds, some of which are responsible for the characteristic flavours of dairy products. The specific compounds responsible for the flavour in several types of cheese are described in Table 2.
Fig. 2.

Generalised depiction of the proteolytic system of lactic acid bacteria. Protein is first broken down by cell envelope proteinase (CEP). Opp, Dpp and DtpT systems transport different sized peptides into the cell. PepX, PepN and PepC intracellular proteases catabolise larger peptides into di‐ and tripeptides. Dipeptidases (DP) and tripeptidases (TP) break di‐ and tripeptides down into free amino acids (FAAs). A wide range of catabolic enzymes then break the FAAs down into volatile organic compounds (VOCs).
Table 2.
Volatile organic compounds derived from amino acids which are important to the flavour of cheese. Adapted from Smit et al. (2005).
| Amino acid | Gouda | Cheddar | Camembert | Swiss |
|---|---|---|---|---|
| Leucine | 3‐Methylbutanal 3‐Methylbutanol 2‐Methylpropanol | 3‐Methylbutanal | 3‐Methylbutanal | 3‐Methylbutanal |
| Methionine | Methanethiol Dimethylsulphide Dimethyltrisulphide | Methional Methanethiol Dimethyldisulphide Dimethyltrisulphide | Methional Methanethiol Dimethylsulphide | Methional |
| Valine | ‐ | Isovaleric acid | ‐ | ‐ |
| Phenylalanine | ‐ | ‐ | Benzaldehyde Phenylacetaldehyde | ‐ |
Proteolysis in plant‐based dairy alternatives
The proteolytic systems of LAB and their function in dairy milk is well known [reviewed in Savijoki et al. (2006); Ji et al. (2021)]. In contrast, very little research has defined how these same systems work on the proteins of plant‐based foods. Research so far has focused primarily on soy (Aguirre et al., 2008; Boulay et al., 2020; Shirotani et al., 2021), as one of the most commonly used plant alternatives to dairy products until recently.
Differences in the structures of plant‐based and dairy‐based proteins mean that the abundance of research into proteolysis in dairy milk is not directly translatable to plant‐based milk. The major proteins in dairy milk are caseins, comprising around 80% of the protein in cow’s milk. Caseins are a family of flexible phosphoproteins comprising of αS1‐, αS2‐, β‐ and κ‐caseins. They form large micelles in milk due to the hydrophobic aggregation of the proteins, with negatively charged κ‐caseins forming the outside of the micelle (Wei and Yano, 2020). Prt enzymes are classified into two types based on their ability to breakdown caseins – PI enzymes mainly degrade ß‐caseins, while PIII are able to degrade αS1‐, ß‐ and κ‐caseins to a similar extent (Visser et al., 1986). PIII enzymes also have a broader specificity for ß‐casein than PI enzymes (Visser et al., 1991). The high abundance of proline residues in caseins leads to an open and disordered secondary structure, which makes them highly susceptible to hydrolysis by cell envelope proteinases (Savijoki et al., 2006). In contrast, most plant proteins are less accessible to proteases, due to being large, multimeric, globular proteins that fold into ordered, tightly packed structures. This is due to the disulphide bonds, hydrophobic effects, electrostatic forces, hydrogen bonding and van der Waals forces present (Sim et al., 2021). Table 3 illustrates some of the key differences between the casein proteins and some selected plant proteins to highlight how proteolysis is more easily facilitated in milk proteins than plant proteins. Carbonaro et al. (2012) investigated the relationship between the digestibility of proteins and their ß‐sheet content in legume, cereal, meat and milk proteins and found that digestibility had a strong inverse relationship to the ß‐sheet content. The authors hypothesised that this was due to the hydrophobic characteristics of these structures, which also make them prone to aggregation upon denaturation. As many plant proteins, especially globular proteins, have high ß‐sheet content (Table 3), this contributes to the low digestibility of these proteins. There have been no plant proteins described within the literature that possess flexible micellar structures analogous to casein, making it difficult to mimic the characteristics and structure of dairy products with plant‐based products (Sim et al., 2021).
Table 3.
Structure and level of hydrolysis by select plant and dairy proteins.
| Source | Protein | Protein type | % Unstructured | % ß‐sheet structure | Molecular weight (kDa) | % Proline residues | Hydrolysis by LAB |
|---|---|---|---|---|---|---|---|
| Dairy | αS1‐casein | Flexible | 33 a | 20 a | 23.6 b | 8.5 c | Some hydrolysis in hydrophilic regions d |
| αS2‐casein | Flexible | 9–34 a | 23–32 a | 25.2 b | 4.8 c | PIII hydrolysis only d | |
| ß‐casein | Flexible | 4–80+ a | 0–34 a | 24 b | 16.7 c | Extensive PI and PIII hydrolysis d | |
| κ‐casein | Flexible | 23–24 a | 35–40 a | 19 b | 11.8 c | Some hydrolysis by PI, PIII and PrtH enzymes d | |
| β−lacto globulin (whey) | Globular | 18–35 e | 49–57 e | 18.4 b | 4.9 c | Little hydrolysis d | |
| Soy | ß‐conglycinin | Globular | 14.4 f | 45.6 f | 150–200 b | 5.4 g | Reasonable hydrolysis of αS and αS' subunits, some hydrolysis of ß subunit h |
| Glycinin | Globular | 16.5 f | 47.3 f | 300–380 b | 4.3 g | Some hydrolysis of A subunit, very little hydrolysis of B subunit h | |
| Pea | Vicilin | Globular | ‐ | 30 f | 150 b | 4.4 g | Not tested |
| Maize | Zein | Prolamin | ‐ | ‐ | 19–24 b | 10.1 g | Not tested |
| Wheat | α/ß‐Gliadin | Prolamin | ‐ | ‐ | 30–45 i | 15–20 i | Some hydrolysis by 8 selected LAB species j , extensive hydrolysis by sourdough starters k . |
Casein bears several key similarities, however, with plant prolamins, the seed storage proteins of cereals, such as zein (found in maize) and gliadins (found in wheat). As demonstrated in Table 3, they are a similar molecular weight, possess high proportions of proline residues, are relatively disordered and spontaneously form large functional structures in their native biological systems. Some research suggests gliadin and gliadin‐derived peptides are suitable substrates for cell envelope proteinases. Pescuma et al. (2013) found that cell envelope proteinases from eight different species of LAB were able to hydrolyse gliadins from wheat. Gobbetti et al. (1996) found that a cell envelope proteinase isolated from Fructilactobacillus sanfranciscensis, one of the dominant LAB in sourdough fermentation, had improved proteolysis of gliadins compared with caseins, and that a dipeptidase and aminopeptidase isolated from F. sanfranciscensis had strong affinity for the hydrophobic peptides released from gliadins. However, the application of prolamin proteins in plant‐based milks may be limited, due to their role in triggering coeliac disease (Shan et al., 2002) and their low solubility in water (Shewry, 2019). Despite this, the similarity in some structural features of prolamins with caseins means these proteins and other prolamins from less common cereals deserve further investigation as potential substrates for cell envelope proteinases.
The ability of cell envelope proteinases to hydrolyse globular proteins has also been studied, mainly from soy. Proteolytic activity in soy milk was shown to be correlated to bacterial growth rate in soy milk for eight different LAB species (Donkor et al., 2007), and cell envelope proteinases from several LAB species were able to hydrolyse low‐molecular‐weight proteins from soy to varying extents (Pescuma et al., 2013). Emkani et al. (2021) demonstrated the evidence of proteolysis in pea extract after fermentation with LAB. Comparison of proteolysis by wild‐type LAB Streptococcus thermophilus compared with a mutant S. thermophilus in which the PrtS enzyme had been removed showed much more extensive proteolysis by the wild‐type, and population growth of the wild type was threefold greater (2.7x108 CFU/mL for the mutant compared with 7.6x108 CFU/mL for the wild type) (Boulay et al., 2020). The main proteins in soy are β‐conglycinin, a trimer formed of α‐, α’‐ and ß‐sub units, and glycinin, a hexamer with each subunit comprising an acidic and basic polypeptide, linked by a disulphide bond (Nishinari et al., 2014). LAB can hydrolyse these proteins to varying degrees depending on the strain. For most LAB strains, ß‐conglycinin is more effectively broken down than glycinin, with the α‐ and α’‐subunits generally being the most preferred substrate for the majority of strains tested, with an average of 46% and 38% of these subunits being hydrolysed, respectively, and the basic subunit of glycinin being the least degraded fraction for all strains with only an average of 13% of this subunit being hydrolysed (Aguirre et al., 2008). Identification of the peptides produced by different LAB strains showed that the specificity for soy proteins differs between LAB strains (Aguirre et al., 2014). The possible cleavage site in soybean proglycinin has been elucidated by X‐ray crystallography, showing that the protein subunits are mainly broken down via cleavage sites on the surface of the protein, likely due to accessibility (Shirotani et al., 2021). Further breakdown of peptides into amino acids and volatile organic compounds has been shown to occur in soy, with compounds related to leucine, isoleucine and phenylalanine/tyrosine being able to be detected after LAB fermentation of soy (Shirotani et al., 2021). These are known to be associated with the flavours of cheese (Yvon and Rijnen, 2001).
Milk proteins have been well studied as a source of bioactive peptides, which are often found in dairy products fermented with LAB (Rafiq et al., 2021). Bioactive peptides are peptides, generally 3–20 amino acids long, which have defined health benefits, such as positive impacts on the nervous, digestive, immune and cardiovascular systems or anticancer properties. Fermentation of plant proteins has also been demonstrated to produce bioactive peptides [for review, see Singh et al. (2014); Rizzello et al. (2016)]. For example, fermentation of soy with Lacticaseibacillus casei, Lactobacillus acidophilus, Lactobacillus Bulgaricus and S. thermophilus was found to produce antihypertensive peptides (Tsai et al., 2006); concentrations of lunasin, a peptide with anticancer, anti‐inflammatory and antioxidative properties, increased when wholemeal wheat, soybean, barley, amaranth and rye flours were fermented with lactic acid bacteria (Rizzello et al., 2012); and antioxidative peptides were produced from various cereal flours after fermentation with lactic acid bacteria (Coda et al., 2012b).
While the studies referenced above have shown that plant proteins can be broken down by LAB proteolytic systems, this has largely been demonstrated in soy and the understanding of the mechanistic basis for proteolysis is limited. Given that proteolysis is one of the key flavour and texture‐forming processes in fermentation, a better understanding of the process on a wider range of substrates would provide key insight into how to improve the fermentation of plant‐based foods into more commercially viable plant‐based dairy alternatives. ß‐Conglycinin, the soy protein most effectively hydrolysed by LAB (Aguirre et al., 2008) is a 7S globulin protein. Other 7S globulin proteins such as vicilins in pea may therefore be promising candidates for successful proteolysis by LAB, as they have similar structure and solubility. Prolamins from cereals are also an interesting candidate for which the proteolysis by LAB has not been investigated in the context of plant‐based dairy alternatives. Significant research into LAB proteolysis of gliadin proteins has been undertaken in sourdough research (Di Cagno et al., 2002; El Mecherfi et al., 2021; Reale et al., 2021). However, this research has often focused on screening sourdough strains for gluten degradation and will often not investigate the underlying hydrolytic mechanism. Notably, the degradation of prolamins by sourdough bacteria/fungi significantly increases the solubility of prolamins, overcoming the insolubility barrier of prolamins and improving their potential functionality (Ogilvie et al., 2021).
Understanding the specificity and accessibility of cell envelope proteinases towards plant proteins would be a valuable way to improve the structure of fermented products, while more research on the downstream metabolites, such as the volatile organic compounds produced, would help to improve the flavour of plant‐based dairy alternatives. Utilising LAB isolated from existing fermented vegetable sources, such as olives, sauerkraut, kimchi and tempeh, may also be a useful strategy to better understand the action of LAB proteolytic systems towards vegetable sources, as these proteolytic systems may be better adapted to plant proteins. For example, LAB strains isolated from pickled vegetables (Sáez et al., 2018) and tempeh (Lim et al., 2019) have demonstrated extracellular proteolytic activity. Use of these strains, or strains isolated from similar sources, to ferment plant‐based milk products may result in more effective proteolysis of plant proteins than the use of strains isolated from dairy sources as has commonly been used thus far.
Sugar metabolism
An important aspect of fermentation is the production of lactic acid from sugars. Lactic acid is produced from glucose, which in dairy milk comes from lactose. In the first step of lactic acid production, the sugar molecule is transported across the membrane and phosphorylated, then, in the case of disaccharides, hydrolysed into monosaccharide units. The glucose monomer is then broken down into lactic acid; in some LAB species the other monosaccharide (e.g., galactose from lactose) can also be metabolised (Stanley, 2003). The isomer of lactate produced varies between LAB species, depending on the levels of l‐lactate and d‐lactate dehydrogenase expressed by the LAB (Wang et al., 2021).
LAB that produce primarily lactic acid are described as being homofermentative, while those that also produce ethanol, acetic acid and/or carbon dioxide are described as heterofermentative. Homofermentative LAB can theoretically produce two moles of lactate for every mole of glucose. Most starter cultures in the dairy industry are homofermentative LAB, as the other by‐products from heterofermentative LAB can introduce undesirable flavours to the product. In homofermentative pathways, sugars are converted to pyruvate via the Embden–Meyerhoff–Parnas pathway, which is then converted to lactic acid by lactate dehydrogenase. Heterofermentative LAB produce only one lactic acid per glucose, as well as other by‐products via the phosphoketolase pathway (Mayo et al., 2010). This process is particularly important for the acidification of the product, as the production of lactic acid is responsible for the drop in pH seen during fermentation. This is important for the characteristic tart flavour associated with yoghurt and fermented drink products.
In a comparison of commercial plant‐based yoghurts (soy, coconut, cashew, almond and hemp) and commercial dairy yoghurt, Grasso et al. (2020) found that none of the plant‐based yoghurts had as high concentration of lactic acid as the dairy control did. The highest concentration was in soy yoghurt at 0.43/100 g, compared with 1.11/100 g in the dairy yoghurt. This suggests that the fermentation was more effective with the lactose substrate than the plant‐based substrates in these products. The plant‐based products mostly had acidity regulators added to decrease the pH to a similar level as was in the dairy control. The ability of different LAB species to convert sugars to lactic acid in plant substrates varies. Four strains of Lactobacillus acidophilus were able to grow with significant levels of lactic acid production in soy milk, and another four strains were able to grow after supplementation with additional sugar, while four strains of Lactobacillus delbrueckii subsp. bulgaricus were unable to effectively utilise sucrose and so exhibited poor growth in soy milk, even when supplemented with additional sugar (Wang et al., 1974).
Plant milks tend to contain high levels of α‐galactosides such as raffinose and stachyose, which are indigestible in the gut and are therefore passed into the lower intestine where they are digested by bacteria which contain the α‐galactosidase enzyme. This can result in gas which causes discomfort and flatulence, which can be a deterrent to consumption of plant‐based products. Some LAB contain the α‐galactosidase enzyme that is able to break these sugars down, so fermentation of plant‐based products can lead to a reduction α‐galactosides. A wide range of LAB species have been shown to have α‐galactosidase activity and the ability to break down raffinose and stachyose in soy (Mital and Steinkraus, 1975; Donkor et al., 2007; Hati et al., 2014) as well as in pea and bean flour (Duszkiewicz‐Reinhard et al., 1994). Much of the sugar present in plant‐based milks is in the form of raffinose and stachyose, so the presence of α‐galactosidases in LAB allows them to grow more effectively in a plant‐based medium. In the development of fermented plant‐based dairy alternatives, it is important to select LAB which can utilise the sugars present in plant‐based sources, and which express α‐galactosidase, for both consumer acceptance of the product and viability of the LAB growth.
Exopolysaccharide formation
Exopolysaccharides are generated by some LAB during the fermentation process and have been identified as key components to developing a desirable flavour and texture in fermented dairy products (Korcz and Varga, 2021). Exopolysaccharides may improve the texture of dairy products by interacting with other components of milk, such as proteins and fats, to form a large network throughout the substrate, acting as texturisers and thickeners to reduce syneresis and increase viscosity and stiffness (Mende et al., 2016). Exopolysaccharides impart desirable qualities onto dairy products at much lower concentrations than commercial polysaccharides that are commonly added during production (Ruas‐Madiedo et al., 2002). They are responsible for firm body, creaminess and a shiny surface in fermented dairy products, due to water‐binding and protein interaction (Korcz and Varga, 2021).
The structure and properties of exopolysaccharides can vary widely. Exopolysaccharides can be homopolysaccharides, consisting of only one type of monosaccharide unit, or heteropolysaccharides, consisting of two or more different monosaccharide units. Homopolysaccharides are relatively simple to synthesise and require little energy expenditure. They are usually synthesised from extracellular starch and sucrose, are generally large (>106 Da) and produced in large quantities (up to 10 g.L‐1). Heteropolysaccharides require a more complex and energy‐intensive synthesis, which can include charged groups, where phosphate or sulphate groups are bound to the monosaccharide subunits, or glucuronic acid is part of the subunit (Mende et al., 2016). They are synthesised from intracellular nucleotide precursors, are usually in the range of 104–106 Da and are produced in mg.L‐1 quantities (Abarquero et al., 2021). Exopolysaccharides can vary in the composition of the monosaccharide building blocks and in the degree of branching present. The majority of LAB in a dairy context synthesise uncharged heteropolysaccharides, usually consisting of some combination of glucose, galactose, N‐acetylgalactosamine and rhamnose (Mende et al., 2016).
Exopolysaccharides in plant‐based milk
Exopolysaccharide‐producing LAB strains have been used effectively in both low‐fat yoghurt and low‐fat cheese production, as the mouthfeel and creaminess imparted by the exopolysaccharides compensates for the loss of these characteristics due to the removal of fat (Güler‐Akin et al., 2009). Plant‐based milks tend to be much lower in fat than dairy milk (Chalupa‐Krebzdak et al., 2018), so the use of exopolysaccharide‐producing LAB strains is important in producing plant‐based products with a desirable texture profile.
Exopolysaccharide‐producing LAB strains are able to produce exopolysaccharides in plant‐based milks and in doing so increase the viscosity and decrease the syneresis of the product. Lactiplantibacillus plantarum 70810 and Lacticaseibacillus rhamnosus 6005 increased the exopolysaccharide content of fermented soy milk, resulting in an increase in viscosity compared with the control (Li et al., 2014). Fermentation of lupin milk was able to produce a yoghurt‐like product with an exopolysaccharide concentration comparable with dairy milk products. When the fermented lupin milk was ultra‐high temperature heated, it had a higher exopolysaccharide concentration, increased viscosity and lower susceptibility to syneresis compared with a pasteurised product (Hickisch et al., 2016a). Oat milk fermented with exopolysaccharide‐producing LAB strains showed greater viscosity in the resulting product than when fermented with non‐exopolysaccharide‐producing strains. The greatest viscosity was seen from oat milk containing glucose, which is able to be utilised in exopolysaccharide synthesis (Mårtensson et al., 2002).
Evidence suggests that many of the effects of exopolysaccharides on dairy products are caused by their interaction with proteins (Ayala‐Hernández et al., 2008, 2009). Exopolysaccharides increase the firmness of the casein network by interacting with the proteins and micelles. As the pH of milk drops during fermentation, neutrally and negatively charged exopolysaccharides interfere with coagulation of casein proteins, which change from negative to neutral or positively charged by the end of fermentation. This leads to a protein network with greater water retention ability and, therefore, decreased syneresis and increased viscosity (Ayala‐Hernández et al., 2009; Gentès et al., 2013; Zeidan et al., 2017). Fermentation of broad bean protein concentrate with two species of exopolysaccharide‐producing LAB increased the viscosity when enriched with sucrose, in a way that could not be mimicked by simply combining the protein concentrate with dextran and organic acids. A lack of proteolysis as determined by SDS‐PAGE and the requirement for sucrose confirmed that the texture relies on the in situ production of exopolysaccharides, and the interaction of these with the protein network (Xu et al., 2019). No other studies to our knowledge have investigated how the textures of fermented plant‐based dairy products are affected by the interaction between exopolysaccharides and plant proteins. Understanding how these interactions work would allow manipulation of composition to improve the texture of fermented plant‐based milk products.
Few studies have investigated the mechanism of exopolysaccharide production or their structure. While the carbohydrate content of dairy milk is made up almost exclusively of lactose, plant sources provide a much wider range of carbohydrate substrates that vary in their monosaccharide units, size, structure, digestibility, fermentability and solubility. They can exist as simple mono‐ or disaccharides (e.g., fructose, galactose, glucose, sucrose or maltose), as oligosaccharides (e.g., inulin or oligofructose) or as larger polysaccharides (e.g., dietary fibre) (McClements and Grossmann, 2021). Understanding how LAB utilise this wide range of carbohydrates would help to understand which sugar sources are best able to create exopolysaccharides to give the best improvement in texture for plant‐based products. The lack of research in this area is understandable, given that the knowledge of how exopolysaccharides contribute to texture in dairy products is also limited, due to the difficulties in separating, purifying and analysing these complex compounds (Mende et al., 2016), but this is a clear area where gains could be made in improving the texture of plant‐based dairy alternatives.
Applications
Fermentation of plant‐based milks with LAB can be used to make products analogous to fermented dairy products, such as cheese, yoghurt and fermented drinks. These can be produced from a wide range of plant‐based milks. Table 4 illustrates a detailed breakdown of types of plant‐based milks that have been used for fermented products that are either available commercially or have been investigated in the scientific literature. Plant‐based yoghurts tend to be made from a defined plant‐based milk or combination of milks and are almost always fermented. It is important that plant‐based yoghurts are fermented, as yoghurt is seen as an important source of probiotics, so the presence of live cultures in plant‐based yoghurt is a key aspect that makes them a ‘dairy yoghurt‐like’ product. Plant‐based cheeses, on the other hand, are not seen as an important source of probiotics, and cheese‐like products are very often made without cultures. It is also more difficult to define them as being made with a specific plant‐based milk, as the base ingredients are often plant oils and/or starches. This makes the plant‐based cheese making process less analogous to the dairy equivalent than yoghurt. Table 4 highlights the difference in prevalence of fermentation in the production of plant‐based cheese compared with yoghurt. There are many different fermented plant‐based yoghurts, based on a wide range of different plant‐based milks, both commercially sold and in the literature. In contrast, fermented cheese alternative products are scarce. Only a handful of fermented plant‐based cheese products can be found commercially, mostly based on nut milks, and a few more have been developed experimentally. The number of plant‐based cheese products that do not utilise fermentation is much greater, so there is clearly a large gap in current knowledge of how to successfully utilise fermentation of plant‐based milk for development of dairy‐free cheese alternatives.
Table 4.
Plant‐based milk types which have been used for experimental and commercial fermented yoghurts and cheeses.
| Plant | Commercial yoghurt brands | Experimental yoghurt references | Commercial cheese brands | Experimental cheese references |
|---|---|---|---|---|
| Coconut | Oui by Yoplait, Cocoyo, So Delicious | ‐ | ‐ | ‐ |
| Barley | Coda et al. (2012a) | ‐ | ‐ | ‐ |
| Emmer | ‐ | Coda et al. (2012a) | ‐ | ‐ |
| Maize | ‐ | ‐ | ‐ | ‐ |
| Millet | ‐ | Song et al. (2020) | ‐ | ‐ |
| Oat | Oatghurt, So delicious | Coda et al. (2012); Bernat et al. (2014); Brückner‐Gühmann et al. (2019b) | Miyoko’s (cheddar) | ‐ |
| Rice | Ricera (discontinued) | Cáceres et al. (2019); Magala et al. (2015); Uzuner et al. (2016) | ‐ | Nakamura et al. (2016) |
| Lupin | Luve | Jiménez‐Martínez et al. (2003); Hickisch et al. (2016a); Laaksonen et al. (2021) | ‐ | ‐ |
| Pea | Ripple (in development) | Klost and Drusch (2019) | ‐ | Sharma et al. (2018) |
| Peanut | ‐ | Isanga and Zhang (2009); Santos et al. (2014) | ||
| Chickpea | ‐ | ‐ | Miyoko’s (cheddar) | ‐ |
| Soy | Alpro, Silk, Sojade | Cheng et al. (1990); Ferragut et al. (2009); Yang et al. (2012) | ‐ | Li et al. (2013); Lorrungruang et al. (2014); Matias et al. (2014); Li et al. (2020) |
| Almond | Silk, Activia | Bernat et al. (2015) | Kite hill (spreadable cheese) | ‐ |
| Cashew | Forager project | ‐ | Savour (feta, ricotta), Vtopian (aged cheddar, brie, camembert), Treeline (aged cheddar), RIND (French‐style cheese) | Chen et al. (2020) |
| Quinoa | ‐ | Lorusso et al. (2018); Zannini et al. (2018) | ‐ | ‐ |
| Hemp | Sojade | ‐ | ‐ | ‐ |
| Potato | ‐ | Levy et al. (2021) | ‐ | ‐ |
| Tigernut | ‐ | Ogundipe et al. (2021) | ‐ | Mutiat et al. (2019) |
Current trends are moving consumers towards ‘clean‐label’ products. That is, products with fewer ingredients and ingredients that are perceived as ‘more natural’. Fermentation with LAB provides an advantage in this respect, as it can impart desirable properties into fermented products without the need to add ingredients to the label (Perpetuini et al., 2021). Breakdown of proteins into smaller peptides, free amino acids and volatile organic compounds can impart desirable flavours and potential health benefits into a product without the need for any additives, and exopolysaccharide formation by LAB can introduce thickening properties.
Yoghurt
There has been considerable development in plant‐based yoghurt production in recent years [for review see Boeck et al. (2021); Montemurro et al. (2021)]. As yoghurts are important sources of probiotics, much of the evaluation of plant‐based yoghurts has focused on the ability of LAB to grow to probiotic concentrations, and their viability in a plant‐based media, as well as the development of desirable flavours and textures. Most probiotic LAB are isolated from dairy milk, so the differences in available nutrients, pH and presence of antinutritional factors in plant‐based milk can all affect the viability of LAB species. Despite this, a wide range of plant‐based milks have been able to be successfully utilised as probiotic carriers (Rasika et al., 2021).
The most common strains of LAB for yoghurt production are a combination of S. thermophilus and L. delbrueckii subsp. bulgaricus, which exhibit a mutualistic relationship. Streptococcus thermophilus grows rapidly, with fast acid and carbon dioxide production that stimulates growth, but most strains exhibit low proteolytic activity. Lactobacillus delbrueckii subsp. bulgaricus strains tend to have much higher proteolytic activity, so produce peptides and free amino acids that S. thermophilus can utilise as a nitrogen source (Tian et al., 2018). This combination of LAB species has been utilised in the production of several experimental plant‐based yoghurts using oat (Brückner‐Gühmann et al., 2019b), potato (Levy et al., 2021), soy (Grygorczyk and Corredig, 2013), lupin (Jiménez‐Martínez et al., 2003; Laaksonen et al., 2021), pea (Klost and Drusch, 2019) and tigernut (Ogundipe et al., 2021) as a substrate, and in commercial yoghurts using soy, cashew, coconut and almond (Grasso et al., 2020) as a substrate. Brückner‐Gühmann et al. (2019b) reported a change in structure of oat protein concentrate, which they attributed to aggregation of proteins when fermented with S. thermophilus and L. delbrueckii subsp. bulgaricus, and some proteolysis, but no significant accumulation of free amino acids. These species were also used to produce an oat‐based yoghurt product with desirable flavour and texture attributes (Brückner‐Gühmann et al., 2019a). Jiménez‐Martínez et al. (2003) reported an acceptable sensory evaluation of lupin‐based products fermented with this combination of LAB species, while sensory evaluation of lupin‐based products by Laaksonen et al. (2021) showed that the product fermented with this combination of LAB species was the ‘most liked’ out of those tested.
The lower protein content and the difference in coagulation properties of the proteins means it can be difficult to obtain a similar consistency in plant‐based yoghurt compared with dairy yoghurt without the addition of thickening agents, especially at low pH (Sim et al., 2021), which affects the clean‐label image of these products. The use of exopolysaccharide‐producing LAB is one way that a similar texture can be achieved. However, in an analysis of 78 commercial plant‐based yoghurt products, 80% used additional thickening agents (Boeck et al., 2021). Exopolysaccharide‐producing LAB can impart similar properties to plant‐based yoghurt alternatives as added hydrocolloids, which improves the clean‐label image of the product (Korcz and Varga, 2021). Recent studies have investigated fermentation of quinoa milk with exopolysaccharide‐producing species of LAB. Fermentation with an EPS‐producing strain of Weissella confusa increased the viscosity of the product from 0.06 to 0.7 Pa/s and the water‐holding capacity from 63% to 98%, compared with fermentation with non‐EPS producing LAB strains which actually decreased both the viscosity and water‐holding capacity (Lorusso et al., 2018). Fermentation with Weissella cibaria resulted in a yoghurt product as viscous as dairy yoghurt (Zannini et al., 2018). The former attributed the textural properties to the formation of an exopolysaccharide‐protein network, whereas the latter observed extensive proteolysis of the protein matrix and instead attributed the increased viscosity to the production of high‐molecular‐weight exopolysaccharides.
The heat treatment method can affect the textural qualities of plant‐based yoghurt products. A more intensive heat treatment (UHT compared with pasteurisation) of a lupin‐based yoghurt reduced the time needed for LAB strains to ferment the product, improved the rheological properties and resulted in a greater concentration of exopolysaccharides (Hickisch et al., 2016a). Further research confirmed that the improved textural properties were due to denaturing of the proteins, allowing improved aggregation and network formation (Hickisch et al., 2016b). Almond milk treated with heat and pressure caused the proteins and oil droplets to aggregate into a continuous protein matrix, which promoted weak gel formation (Bernat et al., 2015).
Particle size also affects the quality of yoghurt products. Reducing particle size can reduce the grittiness and improve mouthfeel. Particle size is particularly relevant for plant‐based yoghurts, due to the tendency of plant proteins to aggregate, which has a negative impact on the syneresis and mouthfeel of the product. High‐pressure processing, where food is treated with a high pressure for a set period of time, was used to create plant‐based yoghurts from mung bean, chickpea, pea, lentil and broad bean and to produce gels with comparable viscosity and viscoelastic properties with commercial dairy yoghurts, without fermentation (Sim et al., 2020). High‐pressure homogenisation produces a uniform particle size by forcing the substrate through a narrow orifice. This technique was used in broad bean samples to cause a change in secondary structure of proteins and dissociated all protein aggregates >1 µm, leading to an increase in solubility from 35 to 99% (Yang et al., 2018).
Cheese
The range of plant‐based cheeses reported in the literature, especially those produced through fermentation by LAB, is considerably smaller than that of yoghurt. This is likely due to the greater challenges of producing a product from plant sources that can imitate the texture and flavour of cheese compared with yoghurt, and the fact that technology development has not focused on this area until recently.
Most dairy cheese is manufactured through curdling of milk, often with the use of proteases such as chymosin from animal or microbial‐derived rennet, or acidification and then subsequent fermentation of the curds by LAB (Visser, 1993). The structure of plant proteins is very different to that of caseins, and rennet does not induce curdling in plant‐based milks, so a range of other curdling and coagulation methods, or alternative methods moving away from those used in dairy, must be employed (Jeewanthi and Paik, 2018). Plant‐based cheeses can broadly be split into those that utilise fermentation and those that do not. Those that do not utilise fermentation are usually not based on plant milks, due to their low fat and protein content, but are instead based on oils (e.g., coconut or palm oil) and starches (e.g., potato or tapioca starch). These bases impart a cheese‐like fat content and structure to the product and are able to somewhat mimic the texture and meltability of cheese. The oil ingredients help the product to mimic the meltability of cheese but not the stretch and flow, while the starches provide some amount of stretch to the product (Mattice and Marangoni, 2020). This type of plant‐based cheese is the most readily available commercially, with 80% of plant‐based cheeses in the UK being based on coconut or palm oil (Saraco and Blaxland, 2020). However, very little evaluation of these types of cheeses can be found in the literature. Fermented plant‐based cheeses use LAB to ferment the plant base until a desired texture is achieved. Among fermented plant‐based cheese products, many commercial products are cashew nut‐based, while soy is the base of most literature‐reported fermented cheeses, and many studies have been undertaken to attempt to improve the quality attributes of soy cheeses (Jeewanthi and Paik, 2018). It is possible that manufacturers avoid soy due to allergies and negative perceptions of soy due to issues around genetic modification, presence of phytoestrogens and perception of unpleasant taste (Schyver and Smith, 2005).
The meltability and stretchability of cheese is caused by the weakening of non‐covalent casein‐casein interactions, which is particularly difficult to mimic with plant‐based proteins (Mattice and Marangoni, 2020), and commercial cheeses that are currently available do not score well in sensory studies or have comparable meltability (Saraco and Blaxland, 2020). Advances in improving the meltability and stretchability of plant‐based cheeses have been made using zein as the plant protein, which has a much higher proline content and a much less tightly packed structure than other plant proteins and forms a non‐covalently linked network in aqueous solutions. A 30% zein content cheese‐like product displayed comparable rheological, textural and melt‐stretch properties with a dairy cheddar control. The sensory properties of this product have not yet been tested to determine whether this is a viable product, but early work looks promising (Mattice and Marangoni, 2020).
The literature on sensory evaluation has been very limited for different plant‐based cheese products. In all studies that utilised fermentation, the likeability of the fermented product was better than that of the unfermented product, mostly attributed to the reduction of gritty mouthfeel and beany flavours. However, none had a likeability rating equal to or better than the dairy control, where this was tested (Short et al., 2021). This clearly demonstrates that further research is needed to develop plant‐based cheese products that can achieve similar or better likeability scores to dairy‐based products. Until this is achieved, plant‐based products will be limited in their potential market. Many different techniques have been employed to improve the acceptability rating of soy‐based cheeses. Adejuyitan et al. (2014) reported a Nigerian cheese product based on soy and coconut milk and found that increasing the percentage of coconut milk improved the overall acceptability of the product. In a study by Li et al. (2013), the soy cheese product that combined fermentation with the addition of glucono‐δ‐lactone received the best score. Combined fermentation by LAB and Geotrichum candidum also increased positive texture attributes of a soy cheese product, although this was not compared with a dairy control or an LAB‐only soy product (Li et al., 2020). Chumchuere et al. (2000) developed a fermented soy cheese product of which the hedonic score was only acceptable once the product was fried. Research into development of a cream cheese‐like product from soy found that fermentation produced diacetyl, to which the cream cheese flavour was attributed. Diacetyl concentration increased upon the addition of sodium acetate or sodium citrate to the fermentation (Hofmann and Marshall, 1985).
Despite the wide range of fermented cashew nut cheese products commercially available, there is a clear gap in the literature analysing these products. Chen et al. (2020) developed a cashew‐based cheese and analysed the microbial and nutritional content but did not describe the sensory perception or textural qualities. Oyeyinka et al. (2019) measured the consumer acceptability of cheeses with different percentages of soy and cashew milk and found that increasing the percentage of cashew milk decreased the acceptability rating. However, this was not a fermented product. It would also be valuable to understand how the sensory rating of fermented commercially available products compares with non‐fermented products, as they are very different products attempting to fill the same market niche.
There is limited research into cheeses produced with other plant milk bases. Nakamura et al. (2016) developed a cheese based on rice milk. However, none of the products achieved a very high overall acceptability rating (29–42%). The products that obtained the most similar consistency to dairy cheese all used gelatine as a coagulating agent that, as an animal‐derived ingredient, is unlikely to be appropriate for many of the target market for this product. Sharma et al. (2018) developed a fermented peanut‐based cheese spread but did not analyse the sensory or textural properties of this product.
Proteolysis and the eventual breakdown of proteins into free amino acids and volatile organic compounds is one of the most important processes for flavour production in cheese manufacture (Yvon and Rijnen, 2001; Smit et al., 2005). Analysis of compounds produced during plant‐based cheese production in the literature is limited. Shirotani et al. (2021) observed leucine, isoleucine and phenylalanine or tyrosine catabolised into 3‐methylbutanal, 2‐methylbutanal and benzaldehyde, respectively, which are known cheese flavour compounds, in fermentation of soy protein isolate with L. helveticus strains. Further study into flavour compounds released during the fermentation of plant‐based milks would contribute to the understanding of how to improve the flavour of plant‐based cheeses.
Unlike yoghurt, which necessarily contains two microbial species for fermentation (S. thermophilus and L. delbrueckii) and potentially a small number of additional species for enhanced flavour, texture or nutritional qualities, cheese can utilise a wide range of microbial biodiversity. This becomes particularly important in aged, artisanal and traditional cheese types, where a wide range of varying microbial species is integral to the development of unique and interesting flavours. For example, analysis of Livanjski cheese, a traditional cheese from Bosnia and Herzegovina identified a total of 159 viable isolates across 11 cheese samples (Vladimír et al., 2020). Very little is known about the importance of microbial biodiversity in plant‐based cheeses, especially since many products currently available do not even utilise fermentation (Grossmann and McClements, 2021). An understanding of the role played by different starter cultures in plant‐based cheese production will help to produce cheese alternatives with more complex and developed flavour profiles.
Although there have been numerous studies on the development of plant‐based cheeses and there are a wide range of products available commercially, it is clear that there is still a lot of work to be done in developing products that will be viable alternatives for the majority of consumers. Understanding the molecular processes occurring during fermentation in plant‐based food may hold the key to developing new and improved plant‐based cheese products.
Conclusions and future focus
Although the systems that LAB use to ferment dairy products have been well studied, there has been very little research to date on how these processes work in plant‐based foods. There is evidence that LAB do effectively ferment plant‐based milks, as has been shown many times in the development of various plant‐based cheeses and yoghurts, but this has for the most part been demonstrated only through the acidification, bacterial count and textural and sensory changes of the substrate. That is, the outcome of the fermentation has been well documented but what is occurring on a molecular level has not. The downstream metabolites of proteolysis are responsible for much of the flavour formation in fermented dairy products, especially cheese, and bioactive peptides produced can result in health benefits. Carbohydrate metabolism allows for the characteristic acidification of the product through production of lactic acid, and the formation of exopolysaccharides that through their interaction with the protein network helps to develop desirable textures. These mechanisms are complex and interconnected. How they work in dairy milk systems is very different to how they work in plant‐based systems, due to the inherent differences in the structure of the substrates. Future work needs to focus on better understanding the underlying processes occurring during fermentation in plant‐based foods and, in particular, how the proteins and sugars specific to different types of plant‐based milk serve as substrates for the proteolytic and glycolytic systems within LAB. Better understanding of the interactions between networks and the break down products from different plant‐based milks will help to develop improvements in the texture and flavour in dairy alternative products, through developing processes to help optimise desirable outcomes.
In addition, studying other plant‐based milk types in addition to soy, where most studies to date have focused, would be advantageous to determine which plant substrates undergo fermentation in a way that results in a product most acceptable to consumers. This review has focused on plant‐based dairy alternatives, but better understanding fermentation in plant‐based products more generally could help to develop a whole range of new plant‐based foods, whether these are designed to mimic animal products or not. Sensory attributes are the most important factor when buying food, more so than price or perceived healthiness (Glanz et al., 1998), so developing plant‐based products with desirable sensory attributes is one of the biggest barriers to the acceptance of these products on the market. Achieving an acceptable sensory profile may be possible through either developing products that highlight natural flavours of the plants that consumers approve of or through developing products that most closely resemble the animal products with which consumers are more familiar. For either approach to be successful, improving our understanding of LAB fermentation processes on a molecular level will be one of the key factors for success. Overcoming the barriers for non‐users to adopt plant‐based dairy products is likely to be challenging, as resistance can come from a range of sources and is not necessarily based on the actual sensory properties of the product. Emotional, conceptual, situational, attitudinal and behavioural aspects can also play a role, as can neophobia (Jaeger and Giacalone, 2021). It will be important to understand all the factors that need to be overcome to encourage adoption of plant‐based products into the mainstream, but having a product with desirable sensory qualities is undeniably an important first step.
Conflict of interest
The authors declare no conflict of interest.
Acknowledgements
A.H. is funded by a Food Transitions 2050 scholarship from the University of Canterbury. We would like to thank Dr. Kevin Sutton for his insight and discussions in the early stages of the research area and Dr. Olivia Ogilvie for her expertise and contribution to the wheat protein sections.
Microbial Biotechnology (2022) 15(5), 1404–1421
Funding information
Ministry of Business, Innovation and Employment Smart Ideas (UOCX1706); The New Zealand Royal Society Marsden Fund (UOC1506); Biomolecular Interactions Centre (University of Canterbury).
References
- Abarquero, D. , Renes, E. , Fresno, J.M. , and Tornadijo, M.E. (2021) Study of exopolysaccharides from lactic acid bacteria and their industrial applications: a review. Int J Food Sci Technol 57: 16–26. doi: 10.1111/ijfs.15227. [DOI] [Google Scholar]
- Adejuyitan, J.A. , Olanipekun, B.F. , and Moyinwin, O.A. (2014) Production and evaluation of cheese‐like product from the blend of soy milk and coconut milk. Arch Appl Sci Res 6: 263–266. [Google Scholar]
- Adeyemo, S.M. , and Onilude, A.A. (2013) Enzymatic reduction of anti‐nutritional factors in fermenting soybeans by Lactobacillus plantarum isolates from fermenting cereals. Niger Food J 31: 84–90. [Google Scholar]
- Aguirre, L. , Garro, M.S. , and Savoy de Giori, G. (2008) Enzymatic hydrolysis of soybean protein using lactic acid bacteria. Food Chem 111: 976–982. [Google Scholar]
- Aguirre, L. , Hebert, E.M. , Garro, M.S. , and Savoy de Giori, G. (2014) Proteolytic activity of Lactobacillus strains on soybean proteins. LWT ‐ Food Sci Technol 59: 780–785. [Google Scholar]
- Ayala‐Hernández, I. , Hassan, A. , Goff, H.D. , Mira de Orduña, R. , and Corredig, M. (2008) Production, isolation and characterization of exopolysaccharides produced by Lactococcus lactis subsp. cremoris JFR1 and their interaction with milk proteins: Effect of pH and media composition. Int Dairy J 18: 1109–1118. [Google Scholar]
- Ayala‐Hernández, I. , Hassan, A.N. , Goff, H.D. , and Corredig, M. (2009) Effect of protein supplementation on the rheological characteristics of milk permeates fermented with exopolysaccharide‐producing Lactococcus lactis subsp. cremoris . Food Hydrocoll 23: 1299–1304. [Google Scholar]
- Bernardeau, M. , Guguen, M. , and Vernoux, J.P. (2006) Beneficial lactobacilli in food and feed: Long‐term use, biodiversity and proposals for specific and realistic safety assessments. FEMS Microbiol Rev 30: 487–513. [DOI] [PubMed] [Google Scholar]
- Bernat, N. , Cháfer, M. , González‐Martínez, C. , Rodríguez‐García, J. , and Chiralt, A. (2014) Optimisation of oat milk formulation to obtain fermented derivatives by using probiotic Lactobacillus reuteri microorganisms. Food Sci Technol Int 21: 145–157. [DOI] [PubMed] [Google Scholar]
- Bernat, N. , Chafera, M. , Chiralt, A. , and Gonzalez‐Martınez, C. (2015) Probiotic fermented almond “milk” as an alternative to cow‐milk yoghurt. Int J Food Stud 4: 201–211. [Google Scholar]
- Bhattacharjee, C. , Saha, S. , Biswas, A. , Kundu, M. , Ghosh, L. , and Das, K. (2005) Structural changes of ß‐lactoglobulin during thermal unfolding and refolding — An FT‐IR and circular dichroism study. Protein J 24: 27–35. [DOI] [PubMed] [Google Scholar]
- Blagden, T.D. , and Gilliland, S.E. (2005) Reduction of levels of volatile components associated with the “beany” flavor in soymilk by Lactobacilli and Streptococci . J Food Sci 70: 186–189. [Google Scholar]
- Boeck, T. , Sahin, A.W. , Zannini, E. , and Arendt, E.K. (2021) Nutritional properties and health aspects of pulses and their use in plant‐based yogurt alternatives. Compr Rev Food Sci Food Saf 20: 3858–3880. [DOI] [PubMed] [Google Scholar]
- Boulay, M. , Al Haddad, M. , and Rul, F. (2020) Streptococcus thermophilus growth in soya milk: Sucrose consumption, nitrogen metabolism, soya protein hydrolysis and role of the cell‐wall protease PrtS. Int J Food Microbiol 335: 108903. [DOI] [PubMed] [Google Scholar]
- Brückner‐Gühmann, M. , Banovic, M. , and Drusch, S. (2019a) Towards an increased plant protein intake: Rheological properties, sensory perception and consumer acceptability of lactic acid fermented, oat‐based gels. Food Hydrocoll 96: 201–208. [Google Scholar]
- Brückner‐Gühmann, M. , Vasil'eva, E. , Culetu, A. , Duta, D. , Sozer, N. , and Drusch, S. (2019b) Oat protein concentrate as alternative ingredient for non‐dairy yoghurt‐type product. J Sci Food Agric 99: 5852–5857. [DOI] [PubMed] [Google Scholar]
- Cáceres, P.J. , Peñas, E. , Martínez‐Villaluenga, C. , García‐Mora, P. , and Frías, J. (2019) Development of a multifunctional yogurt‐like product from germinated brown rice. LWT 99: 306–312. [Google Scholar]
- Carbonaro, M. , Maselli, P. , and Nucara, A. (2012) Relationship between digestibility and secondary structure of raw and thermally treated legume proteins: a Fourier transform infrared (FT‐IR) spectroscopic study. Amino Acids 43: 911–921. [DOI] [PubMed] [Google Scholar]
- Chalupa‐Krebzdak, S. , Long, C.J. , and Bohrer, B.M. (2018) Nutrient density and nutritional value of milk and plant‐based milk alternatives. Int Dairy J 87: 84–92. [Google Scholar]
- Chen, J.M. , Al, K.F. , Craven, L.J. , Seney, S. , Coons, M. , McCormick, H. , et al. (2020) Nutritional, microbial, and allergenic changes during the fermentation of cashew ‘cheese’ product using a quinoa‐based rejuvelac starter culture. Nutrients 12: 648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng, Y.J. , Thompson, L.D. , and Brittin, H.C. (1990) Sogurt, a yogurt‐like soybean product: development and properties. J Food Sci 55: 1178–1179. [Google Scholar]
- Chumchuere, S. , MacDougall, D.B. , and Robinson, R.K. (2000) Production and properties of a semi‐hard cheese made from soya milk. Int J Food Sci Tech 35: 577–581. [Google Scholar]
- Clegg, M.E. , Tarrado Ribes, A. , Reynolds, R. , Kliem, K. , and Stergiadis, S. (2021) A comparative assessment of the nutritional composition of dairy and plant‐based dairy alternatives available for sale in the UK and the implications for consumers’ dietary intakes. Food Res Int 148: 110586. [DOI] [PubMed] [Google Scholar]
- Coda, R. , Lanera, A. , Trani, A. , Gobbetti, M. , and Di Cagno, R. (2012a) Yogurt‐like beverages made of a mixture of cereals, soy and grape must: Microbiology, texture, nutritional and sensory properties. Int J Food Microbiol 155: 120–127. [DOI] [PubMed] [Google Scholar]
- Coda, R. , Rizzello Carlo, G. , Pinto, D. , and Gobbetti, M. (2012b) Selected lactic acid bacteria synthesize antioxidant peptides during sourdough fermentation of cereal flours. Appl Environ Microbiol 78: 1087–1096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Cagno, R. , De Angelis, M. , Lavermicocca, P. , De Vincenzi, M. , Giovannini, C. , Faccia, M. , and Gobbetti, M. (2002) Proteolysis by sourdough lactic acid bacteria: effects on wheat flour protein fractions and gliadin peptides involved in human cereal intolerance. Appl Environ Microbiol 68: 623–633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donkor, O.N. , Henriksson, A. , Vasiljevic, T. , and Shah, N.P. (2007) α‐Galactosidase and proteolytic activities of selected probiotic and dairy cultures in fermented soymilk. Food Chem 104: 10–20. [Google Scholar]
- Duszkiewicz‐Reinhard, W. , Gujska, E. , and Khan, K. (1994) Reduction of stachyose in legume flours by lactic acid bacteria. J Food Sci 59: 115–117. [Google Scholar]
- El Mecherfi, K.‐E. , Lupi, R. , Cherkaoui, M. , Albuquerque, M.A.C. , Todorov, S.D. , Tranquet, O. , et al. (2021) Fermentation of gluten by Lactococcus lactis LLGKC18 reduces its antigenicity and allergenicity. Probiotics Antimicrob Proteins. doi: 10.1007/s12602-021-09808-1. [DOI] [PubMed] [Google Scholar]
- Emkani, M. , Oliete, B. , and Saurel, R. (2021) Pea protein extraction assisted by lactic fermentation: Impact on protein profile and thermal properties. Foods 10: 549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferragut, V. , Cruz, N.S. , Trujillo, A. , Guamis, B. , and Capellas, M. (2009) Physical characteristics during storage of soy yogurt made from ultra‐high pressure homogenized soymilk. J Food Eng 92: 63–69. [Google Scholar]
- Fox, P.F. , Wallace, J.M. , Morgan, S. , Lynch, C.M. , Niland, E.J. , and Tobin, J. (1996) Acceleration of cheese ripening. Antonie Van Leeuwenhoek 70: 271–297. [DOI] [PubMed] [Google Scholar]
- Fukushima, D. (1991) Structures of plant storage proteins and their functions. Food Rev Int 7: 353–381. [Google Scholar]
- Gentès, M.‐C. , St‐Gelais, D. , and Turgeon, S.L. (2013) Exopolysaccharide–milk protein interactions in a dairy model system simulating yoghurt conditions. Dairy Sci Technol 93: 255–271. [Google Scholar]
- Glanz, K. , Basil, M. , Maibach, E. , Goldberg, J. , and Snyder, D.A.N. (1998) Why Americans eat what they do: Taste, nutrition, cost, convenience, and weight control concerns as influences on food consumption. J Am Diet Assoc 98: 1118–1126. [DOI] [PubMed] [Google Scholar]
- Gobbetti, M. , Smacchi, E. , and Corsetti, A. (1996) The proteolytic system of Lactobacillus sanfrancisco CB1: Purification and characterization of a proteinase, a dipeptidase, and an aminopeptidase. Appl Environ Microbiol 62: 3220–3226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- González, A.D. , Frostell, B. , and Carlsson‐Kanyama, A. (2011) Protein efficiency per unit energy and per unit greenhouse gas emissions: Potential contribution of diet choices to climate change mitigation. Food Policy 36: 562–570. [Google Scholar]
- Good Food Institute (2021) U.S. retail market data for the plant‐based industry [WWW document]. URL https://gfi.org/marketresearch/ [Google Scholar]
- Grasso, N. , Alonso‐Miravalles, L. , and O’Mahony, J.A. (2020) Composition, physicochemical and sensorial properties of commercial plant‐based yogurts. Foods 9: 252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grossmann, L. , and McClements, D.J. (2021) The science of plant‐based foods: approaches to create nutritious and sustainable plant‐based cheese analogs. Trends Food Sci Technol 118: 207–229. [Google Scholar]
- Grygorczyk, A. , and Corredig, M. (2013) Acid induced gelation of soymilk, comparison between gels prepared with lactic acid bacteria and glucono‐δ‐lactone. Food Chem 141: 1716–1721. [DOI] [PubMed] [Google Scholar]
- Güler‐Akin, M.B. , Serdar Akin, M. , and Korkmaz, A. (2009) Influence of different exopolysaccharide‐producing strains on the physicochemical, sensory and syneresis characteristics of reduced‐fat stirred yoghurt. Int J Dairy Technol 62: 422–430. [Google Scholar]
- Hansen, E.B. , and Marcatili, P. (2020) Modeled structure of the cell envelope proteinase of Lactococcus lactis . Front Bioeng Biotechnol 8: 1476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hati, S. , Vij, S. , Mandal, S. , Malik, R.K. , Kumari, V. , and Khetra, Y. (2014) α‐Galactosidase activity and oligosaccharides utilization by Lactobacilli during fermentation of soy milk. J Food Process Preserv 38: 1065–1071. [Google Scholar]
- Hayes, M. , Ross, R.P. , Fitzgerald, G.F. , and Stanton, C. (2007) Putting microbes to work: dairy fermentation, cell factories and bioactive peptides. Part I: Overview. Biotechnol J 2: 426–434. [DOI] [PubMed] [Google Scholar]
- Hemme, D. , and Foucaud‐Scheunemann, C. (2004) Leuconostoc, characteristics, use in dairy technology and prospects in functional foods. Int Dairy J 14: 467–494. [Google Scholar]
- Hickisch, A. , Beer, R. , Vogel, R.F. , and Toelstede, S. (2016a) Influence of lupin‐based milk alternative heat treatment and exopolysaccharide‐producing lactic acid bacteria on the physical characteristics of lupin‐based yogurt alternatives. Food Res Int 84: 180–188. [DOI] [PubMed] [Google Scholar]
- Hickisch, A. , Bindl, K. , Vogel, R.F. , and Toelstede, S. (2016b) Thermal treatment of lupin‐based milk alternatives – Impact on lupin proteins and the network of respective lupin‐based yogurt alternatives. Food Res Int 89: 850–859. [DOI] [PubMed] [Google Scholar]
- Hofmann, C.J. , and Marshall, W.E. (1985) Lactic fermentation of ground soybean for use in imitation cream cheese products. J Food Sci 50: 325–329. [Google Scholar]
- Horne, D.S. (2002) Casein structure, self‐assembly and gelation. Curr Opin Colloid Interface Sci 7: 456–461. [Google Scholar]
- Isanga, J. , and Zhang, G. (2009) Production and evaluation of some physicochemical parameters of peanut milk yoghurt. LWT‐Food Sci Technol 42: 1132–1138. [Google Scholar]
- Islam, N. , Shafiee, M. , and Vatanparast, H. (2021) Trends in the consumption of conventional dairy milk and plant‐based beverages and their contribution to nutrient intake among Canadians. J Hum Nutr Diet 34: 1022–1034. doi: 10.1111/jhn.12910. [DOI] [PubMed] [Google Scholar]
- Iyer, R. , Tomar, S.K. , Uma Maheswari, T. , and Singh, R. (2010) Streptococcus thermophilus strains: multifunctional lactic acid bacteria. Int Dairy J 20: 133–141. [Google Scholar]
- Jaeger, S.R. , and Giacalone, D. (2021) Barriers to consumption of plant‐based beverages: A comparison of product users and non‐users on emotional, conceptual, situational, conative and psychographic variables. Food Res Int 144: 110363. [DOI] [PubMed] [Google Scholar]
- Jeewanthi, R.K.C. , and Paik, H.‐D. (2018) Modifications of nutritional, structural, and sensory characteristics of non‐dairy soy cheese analogs to improve their quality attributes. J Food Sci Technol 55: 4384–4394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji, D. , Ma, J. , Xu, M. , and Agyei, D. (2021) Cell‐envelope proteinases from lactic acid bacteria: biochemical features and biotechnological applications. Compr Rev Food Sci Food Saf 20: 369–400. [DOI] [PubMed] [Google Scholar]
- Jiménez‐Martínez, C. , Hernández‐Sánchez, H. , and Dávila‐Ortiz, G. (2003) Production of a yogurt‐like product from Lupinus campestris seeds. J Sci Food Agric 83: 515–522. [Google Scholar]
- Jooyandeh, H. , Kaur, A. , and Minhas, K.S. (2009) Lipases in dairy industry: a review. J Food Sci Technol 46: 181–189. [Google Scholar]
- Juillard, V. , Laan, H. , Kunji, E.R. , Jeronimus‐Stratingh, C.M. , Bruins, A.P. , and Konings, W.N. (1995) The extracellular PI‐type proteinase of Lactococcus lactis hydrolyzes beta‐casein into more than one hundred different oligopeptides. J Bacteriol 177: 3472–3478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klost, M. , and Drusch, S. (2019) Structure formation and rheological properties of pea protein‐based gels. Food Hydrocoll 94: 622–630. [Google Scholar]
- Korcz, E. , and Varga, L. (2021) Exopolysaccharides from lactic acid bacteria: techno‐functional application in the food industry. Trends Food Sci Technol 110: 375–384. [Google Scholar]
- Laaksonen, O. , Kahala, M. , Marsol‐Vall, A. , Blasco, L. , Järvenpää, E. , Rosenvald, S. , et al. (2021) Impact of lactic acid fermentation on sensory and chemical quality of dairy analogues prepared from lupine (Lupinus angustifolius L.) seeds. Food Chem 346: 128852. [DOI] [PubMed] [Google Scholar]
- Levy, R. , Okun, Z. , Davidovich‐Pinhas, M. , and Shpigelman, A. (2021) Utilization of high‐pressure homogenization of potato protein isolate for the production of dairy‐free yogurt‐like fermented product. Food Hydrocoll 113: 106442. [Google Scholar]
- Li, C. , Li, W. , Chen, X. , Feng, M. , Rui, X. , Jiang, M. , and Dong, M. (2014) Microbiological, physicochemical and rheological properties of fermented soymilk produced with exopolysaccharide (EPS) producing lactic acid bacteria strains. LWT ‐ Food Sci Technol 57: 477–485. [Google Scholar]
- Li, Q. , Xia, Y. , Zhou, L. , and Xie, J. (2013) Evaluation of the rheological, textural, microstructural and sensory properties of soy cheese spreads. Food Bioprod Process 91: 429–439. [Google Scholar]
- Li, Y. , Zhang, X. , Yang, J.‐J. , Ma, X.‐Y. , Jia, X.‐D. , Du, P. , and Li, A.‐L. (2020) Influence of the addition of Geotrichum candidum on the microbial, chemical, textural, and sensory features of soft soy cheese. J Food Process Preserv 44: e14823. [Google Scholar]
- Lim, Y.H. , Foo, H.L. , Loh, T.C. , Mohamad, R. , and Abdullah, N. (2019) Comparative studies of versatile extracellular proteolytic activities of lactic acid bacteria and their potential for extracellular amino acid productions as feed supplements. J Anim Sci Biotechnol 10: 15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lopez, H.W. , Ouvry, A. , Bervas, E. , Guy, C. , Messager, A. , Demigne, C. , and Remesy, C. (2000) Strains of lactic acid bacteria isolated from sour doughs degrade phytic acid and improve calcium and magnesium solubility from whole wheat flour. J Agric Food Chem 48: 2281–2285. [DOI] [PubMed] [Google Scholar]
- Lorrungruang, C. , Sinma, K. , Pantagrud, P. , Wannasirisuk, S. , Mahabandha, K. , and Khucharoenphaisan, K. (2014) Red cheese production from Soymilk by Monascus purpureus and Lactobacillus casei . J Appl Sci 14: 2819–2824. [Google Scholar]
- Lorusso, A. , Coda, R. , Montemurro, M. , and Rizzello, C.G. (2018) Use of selected lactic acid bacteria and quinoa flour for manufacturing novel yogurt‐like beverages. Foods 7: 51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Magala, M. , Kohajdová, Z. , Karovičová, J. , Greifová, M. , and Hojerová, J. (2015) Application of lactic acid bacteria for production of fermented beverages based on rice flour. Czech J Food Sci 33: 458–563. [Google Scholar]
- Mäkinen, O.E. , Wanhalinna, V. , Zannini, E. , and Arendt, E.K. (2016) Foods for special dietary needs: Non‐dairy plant‐based milk substitutes and fermented dairy‐type products. Crit Rev Food Sci Nutr 56: 339–349. [DOI] [PubMed] [Google Scholar]
- Mårtensson, O. , Öste, R. , and Holst, O. (2002) Texture promoting capacity and EPS formation by lactic acid bacteria in three different oat‐based non‐dairy media. Eur Food Res Technol 214: 232–236. [Google Scholar]
- Matias, N.S. , Bedani, R. , Castro, I.A. , and Saad, S.M.I. (2014) A probiotic soy‐based innovative product as an alternative to petit‐suisse cheese. LWT ‐ Food Sci Technol 59: 411–417. [Google Scholar]
- Mattice, K.D. , and Marangoni, A.G. (2020) Physical properties of plant‐based cheese products produced with zein. Food Hydrocoll 105: 105746. [Google Scholar]
- Mayo, B. , Aleksandrzak‐Piekarczyk, T. , Fernández, M. , Kowalczyk, M. , Álvarez‐Martín, P. , and Bardowski, J. (2010) Updates in the metabolism of lactic acid bacteria. In Biotechnology of Lactic Acid Bacteria. New Jersey: Blackwell Publishing, pp. 3–33. [Google Scholar]
- McCarthy, K.S. , Parker, M. , Ameerally, A. , Drake, S.L. , and Drake, M.A. (2017) Drivers of choice for fluid milk versus plant‐based alternatives: what are consumer perceptions of fluid milk? J Dairy Sci 100: 6125–6138. [DOI] [PubMed] [Google Scholar]
- McClements, D.J. , and Grossmann, L. (2021) The science of plant‐based foods: constructing next‐generation meat, fish, milk, and egg analogs. Compr Rev Food Sci Food Saf 20: 4049–4100. [DOI] [PubMed] [Google Scholar]
- Mende, S. , Rohm, H. , and Jaros, D. (2016) Influence of exopolysaccharides on the structure, texture, stability and sensory properties of yoghurt and related products. Int Dairy J 52: 57–71. [Google Scholar]
- Mital, B.K. , and Steinkraus, K.H. (1975) Utilisation of oligosaccharides by lactic acid bacteria during fermentation of soy milk. J Food Sci 40: 114–118. [Google Scholar]
- Montemurro, M. , Pontonio, E. , Coda, R. , and Rizzello, C.G. (2021) Plant‐based alternatives to yogurt: state‐of‐the‐art and perspectives of new biotechnological challenges. Foods 10: 316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mutiat, B. , Oyeyinka, S. , Kolawole, F. , Joseph, J. , and Olajobi, G. (2019) Chemical composition and sensory properties of soy‐tiger nut cheese. Ceylon J Sci 48: 353. [Google Scholar]
- Nakamura, M. , Kitamura, Y. , and Kokawa, M. (2016) Development of a cheese‐type food using rice milk. Food Sci Technol Res 22: 605–609. [Google Scholar]
- Nishinari, K. , Fang, Y. , Guo, S. , and Phillips, G.O. (2014) Soy proteins: a review on composition, aggregation and emulsification. Food Hydrocoll 39: 301–318. [Google Scholar]
- O’Regan, J. , Ennis, M. P. , and Mulvihill, D. M. (2009) Chapter 13: Milk proteins. In Handbook of Hydrocolloids, 2nd edn. Phillips, G. O. , Williams, P. A. (eds). Sawston: Woodhead Publishing, pp. 298–358. [Google Scholar]
- Ogilvie, O.J. , Gerrard, J.A. , Roberts, S. , Sutton, K.H. , Larsen, N. , and Domigan, L.J. (2021) A case study of the response of immunogenic gluten peptides to sourdough proteolysis. Nutrients 13: 1906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogundipe, O.O. , Fasogbon, B.M. , Ogundipe, F.O. , Oredope, O. , and Amaezenanbu, R.U. (2021) Nutritional composition of non‐dairy yogurt from sprouted tigernut tubers. J Food Process Preserv 45: e15884. [Google Scholar]
- Oyeyinka, A.T. , Odukoya, J.O. , and Adebayo, Y.S. (2019) Nutritional composition and consumer acceptability of cheese analog from soy and cashew nut milk. J Food Process Preserv 43: e14285. [Google Scholar]
- Perpetuini, G. , Chuenchomrat, P. , Pereyron, V. , Haure, M. , Lorn, D.A. , Quan, L.‐H. , et al. (2021) Microorganisms, the ultimate tool for clean label foods? Inventions 6: 31. [Google Scholar]
- Pescuma, M. , Espeche Turbay, M.B. , Mozzi, F. , Font de Valdez, G. , Savoy de Giori, G. , and Hebert, E.M. (2013) Diversity in proteinase specificity of thermophilic lactobacilli as revealed by hydrolysis of dairy and vegetable proteins. Appl Microbiol Biotechnol 97: 7831–7844. [DOI] [PubMed] [Google Scholar]
- Rackis, J.J. , Sessa, D.J. , and Honig, D.H. (1979) Flavor problems of vegetable food proteins. J Am Oil Chem Soc 56: 262–271. [Google Scholar]
- Rafiq, S. , Gulzar, N. , Sameen, A. , Huma, N. , Hayat, I. , and Ijaz, R. (2021) Functional role of bioactive peptides with special reference to cheeses. Int J Dairy Technol 74: 1–16. [Google Scholar]
- Rasika, D.M.D. , Vidanarachchi, J.K. , Rocha, R.S. , Balthazar, C.F. , Cruz, A.G. , Sant’Ana, A.S. , and Ranadheera, C.S. (2021) Plant‐based milk substitutes as emerging probiotic carriers. Curr Opin Food Sci 38: 8–20. [Google Scholar]
- Reale, A. , Di Stasio, L. , Di Renzo, T. , De Caro, S. , Ferranti, P. , Picariello, G. , et al. (2021) Bacteria do it better! Proteomics suggests the molecular basis for improved digestibility of sourdough products. Food Chem 359: 129955. [DOI] [PubMed] [Google Scholar]
- Rizzello, C.G. , Nionelli, L. , Coda, R. , and Gobbetti, M. (2012) Synthesis of the cancer preventive peptide lunasin by lactic acid bacteria during sourdough fermentation. Nutr Cancer 64: 111–120. [DOI] [PubMed] [Google Scholar]
- Rizzello, C.G. , Tagliazucchi, D. , Babini, E. , Sefora Rutella, G. , Taneyo Saa, D.L. , and Gianotti, A. (2016) Bioactive peptides from vegetable food matrices: research trends and novel biotechnologies for synthesis and recovery. J Funct Foods 27: 549–569. [Google Scholar]
- Ruas‐Madiedo, P. , Hugenholtz, J. , and Zoon, P. (2002) An overview of the functionality of exopolysaccharides produced by lactic acid bacteria. Int Dairy J 12: 163–171. [Google Scholar]
- Sá, A.G.A. , Moreno, Y.M.F. , and Carciofi, B.A.M. (2020) Food processing for the improvement of plant proteins digestibility. Crit Rev Food Sci Nutr 60: 3367–3386. [DOI] [PubMed] [Google Scholar]
- Sadat‐Mekmene, L. , Genay, M. , Atlan, D. , Lortal, S. , and Gagnaire, V. (2011) Original features of cell‐envelope proteinases of Lactobacillus helveticus . A review. Int J Food Microbiol 146: 1–13. [DOI] [PubMed] [Google Scholar]
- Sáez, G.D. , Flomenbaum, L. , and Zárate, G. (2018) Lactic acid bacteria from Argentinean fermented foods: isolation and characterization for their potential use as starters for fermentation of vegetables. Food Technol Biotechnol 56: 398–410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Şanlier, N. , Gökcen, B.B. , and Sezgin, A.C. (2019) Health benefits of fermented foods. Crit Rev Food Sci Nutr 59: 506–527. [DOI] [PubMed] [Google Scholar]
- Saraco, M. , and Blaxland, J. (2020) Dairy‐free imitation cheese: is further development required? Br Food J 122: 3727–3740. [Google Scholar]
- Saranraj, P. , Naidu, M.A. , and Sivasakthivelan, P. (2013) Lactic acid bacteria and its antimicrobial properties: a review. Int J Pharm Biol Arch 4: 1124–1133. [Google Scholar]
- Savijoki, K. , Ingmer, H. , and Varmanen, P. (2006) Proteolytic systems of lactic acid bacteria. Appl Microbiol Biotechnol 71: 394–406. [DOI] [PubMed] [Google Scholar]
- Schyver, T. , and Smith, C. (2005) Reported attitudes and beliefs toward soy food consumption of soy consumers versus nonconsumers in natural foods or mainstream grocery stores. J Nutr Educ Behav 37: 292–299. [DOI] [PubMed] [Google Scholar]
- Shan, L.U. , Molberg, Ø. , Parrot, I. , Hausch, F. , Filiz, F. , Gray, G.M. , et al. (2002) Structural basis for gluten intolerance in celiac sprue. Science 297: 2275–2279. [DOI] [PubMed] [Google Scholar]
- Sharma, P. , Sharma, D. , and Amin, A. (2018) Development of a functional fermented peanut‐based cheese analog using probiotic bacteria. BioTechnologia 99: 435–441. [Google Scholar]
- Shevkani, K. , Singh, N. , Chen, Y. , Kaur, A. , and Yu, L. (2019) Pulse proteins: secondary structure, functionality and applications. J Food Sci Technol 56: 2787–2798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shewry, P. (2019) What is gluten—why is it special? Front Nutr 6: 101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shirotani, N. , Bygvraa Hougaard, A. , Lametsch, R. , Agerlin Petersen, M. , Rattray, F.P. , and Ipsen, R. (2021) Proteolytic activity of selected commercial Lactobacillus helveticus strains on soy protein isolates. Food Chem 340: 128152. [DOI] [PubMed] [Google Scholar]
- Short, E.C. , Kinchla, A.J. , and Nolden, A.A. (2021) Plant‐based cheeses: a systematic review of sensory evaluation studies and strategies to increase consumer acceptance. Foods 10: 725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sim, S.Y.J. , Hua, X.Y. , and Henry, C.J. (2020) A novel approach to structure plant‐based yogurts using high pressure processing. Foods 9: 1126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sim, S.Y.J. , Srv, A. , Chiang, J.H. , and Henry, C.J. (2021) Plant proteins for future foods: a roadmap. Foods 10: 1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh, B.P. , Vij, S. , and Hati, S. (2014) Functional significance of bioactive peptides derived from soybean. Peptides 54: 171–179. [DOI] [PubMed] [Google Scholar]
- Smit, G. , Smit, B.A. , and Engels, W.J.M. (2005) Flavour formation by lactic acid bacteria and biochemical flavour profiling of cheese products. FEMS Microbiol Rev 29: 591–610. [DOI] [PubMed] [Google Scholar]
- Song, X. , Sun, X. , Ban, Q. , Cheng, J. , Zhang, S. , and Guo, M. (2020) Gelation and microstructural properties of a millet‐based yogurt‐like product using polymerized whey protein and xanthan gum as thickening agents. J Food Sci 85: 3927–3933. [DOI] [PubMed] [Google Scholar]
- Stanley, G. (2003) Cheeses | starter cultures employed in cheese‐making. In Encyclopedia of Food Sciences and Nutrition, 2nd edn. Caballero, B. (ed.), Oxford: Academic Press, pp. 1051–1056. [Google Scholar]
- Teusink, B. , and Molenaar, D. (2017) Systems biology of lactic acid bacteria: for food and thought. Curr Opin Syst Biol 6: 7–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian, H. , Li, B. , Evivie, S.E. , Sarker, S.K. , Chowdhury, S. , Lu, J. , et al. (2018) Technological and genomic analysis of roles of the cell‐envelope protease PrtS in yoghurt starter development. Int J Mol Sci 19: 1068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsai, J.S. , Lin, Y.S. , Pan, B.S. , and Chen, T.J. (2006) Antihypertensive peptides and γ‐aminobutyric acid from prozyme 6 facilitated lactic acid bacteria fermentation of soymilk. Proc Biochem 41: 1282–1288. [Google Scholar]
- Uzuner, A. E. , Kinik, O. , Korel, F. , Yildiz, G. , and Yerlikaya, O. (2016) Usage of rice milk in probiotic yoghurt production. Carpathian J Food Sci Technol 8: 5–25. [Google Scholar]
- Visser, S. (1993) Proteolytic enzymes and their relation to cheese ripening and flavor: an overview. J Dairy Sci 76: 329–350. [Google Scholar]
- Visser, S. , Exterkate, F.A. , Slangen, C.J. , and de Veer, G.J.C.M. (1986) Comparative study of action of cell wall proteinases from various strains of Streptococcus cremoris; on bovine αs1‐, β‐, and κ‐casein. Appl Environ Microbiol 52: 1162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Visser, S. , Robben, A.J.P.M. , and Slangen, C.J. (1991) Specificity of a cell‐envelope‐located proteinase (PIII‐type) from Lactococcus lactis subsp. cremoris AM1 in its action on bovine β‐casein. Appl Microbiol Biotechnol 35: 477–483. [DOI] [PubMed] [Google Scholar]
- Vladimír, D. , Miloslava, K. , Markéta, M. , Jaroslava, H. , and Petr, R. (2020) Microbial diversity of Livanjski cheese with the emphasis on lactic acid bacteria based on culture‐dependent and sequencing method. Int J Dairy Technol 73: 202–214. [Google Scholar]
- Wang, H.L. , Kraidej, L. , and Hesseltine, C.W. (1974) Lactic acid fermentation of soybean milk. J Milk Food Technol 37: 71–73. [Google Scholar]
- Wang, Y. , Wu, J. , Lv, M. , Shao, Z. , Hungwe, M. , Wang, J. , et al. (2021) Metabolism characteristics of lactic acid bacteria and the expanding applications in food industry. Front Bioeng Biotechnol 9: 378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei, F. , and Yano, H. (2020) Development of “new” bread and cheese. Processes 8: 1541. [Google Scholar]
- Xu, Y. , Coda, R. , Holopainen‐Mantila, U. , Laitila, A. , Katina, K. , and Tenkanen, M. (2019) Impact of in situ produced exopolysaccharides on rheology and texture of fava bean protein concentrate. Food Res Int 115: 191–199. [DOI] [PubMed] [Google Scholar]
- Yang, J. , Liu, G. , Zeng, H. , and Chen, L. (2018) Effects of high pressure homogenization on faba bean protein aggregation in relation to solubility and interfacial properties. Food Hydrocoll 83: 275–286. [Google Scholar]
- Yang, M. , Fu, J. , and Li, L. (2012) Rheological characteristics and microstructure of probiotic soy yogurt prepared from germinated soybeans. Food Technol Biotechnol 50: 73. [Google Scholar]
- Yvon, M. , and Rijnen, L. (2001) Cheese flavour formation by amino acid catabolism. Int Dairy J 11: 185–201. [Google Scholar]
- Zannini, E. , Jeske, S. , Lynch, K.M. , and Arendt, E.K. (2018) Development of novel quinoa‐based yoghurt fermented with dextran producer Weissella cibaria MG1. Int J Food Microbiol 268: 19–26. [DOI] [PubMed] [Google Scholar]
- Zeidan, A.A. , Poulsen, V.K. , Janzen, T. , Buldo, P. , Derkx, P.M.F. , Øregaard, G. , and Neves, A.R. (2017) Polysaccharide production by lactic acid bacteria: from genes to industrial applications. FEMS Microbiol Rev 41: S168–S200. [DOI] [PubMed] [Google Scholar]


