Abstract
A total of 209 clinical isolates of Pseudomonas (193 Pseudomonas aeruginosa, 10 P. putida, 4 P. stutzeri, and 2 P. fluorescens isolates) with reduced susceptibilities to imipenem and/or ceftazidime were subjected to PCR assays with primers specific for blaIMP-1, blaIMP-2, blaVIM-1, and blaVIM-2 and sequence analysis to identify the metallo-β-lactamases (MBLs) prevalent among these organisms in Taiwan; and 21 isolates gave positive results. Five isolates including two P. putida and three P. stutzeri isolates were found to carry blaIMP-1, and six isolates including five P. putida and one P. stutzeri isolates harbored blaVIM-2. The remaining 10 isolates were P. aeruginosa, and all were found to carry a novel variant of blaVIM-2, designated blaVIM-3. There are only two nucleotide differences between blaVIM-2 and blaVIM-3, leading to two amino acid alterations. Our findings indicate that VIM-2 and its variant have become the most prevalent metalloenzymes in Pseudomonas in Taiwan. Southern hybridization with the blaVIM-2-, blaVIM-3-, and blaIMP-1 -specific probes revealed that only two VIM-2-producing P. putida isolates appeared to carry the MBL gene on plasmids. Pulsed-field gel electrophoresis showed that six VIM-3-producing P. aeruginosa isolates and two IMP-1-producing P. stutzeri isolates were genetically related, suggesting that the spread of these MBL genes in Taiwan could be due to clonal dissemination as well as genetic exchange between different clones.
Carbapenems are often used as antibiotics of last resort for the treatment of infections caused by gram-negative bacteria resistant to other β-lactam agents because of their broad spectra of activity and stability to hydrolysis by most β-lactamases including extended-spectrum β-lactamases (4, 12, 26, 27). The mechanisms of resistance to carbapenems in Pseudomonas aeruginosa often result from reduced levels of drug accumulation or increased levels of expression of pump efflux (8, 15, 17). The resistance is occasionally due to production of metallo-β-lactamases (MBLs), which can be either chromosomally encoded or plasmid mediated (8, 13, 22, 28, 32). IMP-1 was the first MBL identified in P. aeruginosa (28, 29). The enzyme has also been described in other nonfastidious, gram-negative nonfermenters and members of the family Enterobacteriaceae (9, 10, 20, 29). Several novel MBLs were identified recently, including VIM-1 from P. aeruginosa and IMP-2 from Acinetobacter baumannii in Italy (13, 25), VIM-2 from P. aeruginosa in France (22), and IMP-3 from Shigella flexneri in Japan (11). The spread of MBLs in gram-negative rods has been described in several other countries and is becoming an emerging threat (32; T. H. Koh, G. S. Babini, N. Woodford, L.-H. Sng, L. M. C. Hall, and D. M. Livermore, Letter, Lancet 353:2162, 1999; K. Lee, J. B. Lim, J. Yum, D. Yong, J. R. Choi, Y. Chong, J. M. Kim, and D. M. Livermore, Abstr. 40th Intersci. Conf. Antimicrob. Agents Chemother., abstr. 2003, p. 123, 2000). It remains unknown whether these MBLs have appeared in Taiwan. Thus, the purpose of the present study was to determine the IMP- and VIM-type MBLs prevalent among clinical isolates of the genus Pseudomonas in Taiwan.
MATERIALS AND METHODS
Bacterial strains.
The isolates studied were selected from clinical isolates of Pseudomonas spp. randomly collected between January 1997 and May 2000 at two university hospitals in Taiwan. Since susceptibilities to imipenem for IMP-1 producers covered a wide range (9, 10, 28, 29), all isolates that showed reduced susceptibilities to imipenem (inhibition zone diameter, <16 mm) and/or ceftazidime (inhibition zone diameter, <18 mm) on the basis of the results of the disk diffusion method (19) were examined. Thus, a total of 209 isolates, including 164 Pseudomonas isolates (154 P. aeruginosa, 4 P. stutzeri, 4 P. putida, and 2 P. fluorescens isolates) from the National Cheng Kung University Medical Center, a 900-bed teaching hospital in southern Taiwan, and 45 Pseudomonas isolates (39 P. aeruginosa and 6 P. putida isolates) from the National Taiwan University Hospital, a 1,800-bed medical center in northern Taiwan, were selected. All these isolates were recovered from different patients and were identified by conventional techniques (6) and/or with the API 20NE system (bioMérieux, Marcy l'Etoile, France).
Screening tests for detection of MBL producers.
The disk diffusion test for the screening of MBL producers was performed as described by Arakawa et al. (1). 2-Mercaptopropionic acid and 100 mM EDTA were used as MBL inhibitors.
PCR amplification and DNA sequencing.
Plasmids from clinical isolates were prepared by a rapid alkaline lysis procedure (31). PCR assays were performed to amplify the entire sequences of the blaIMP-1, blaIMP-2, blaVIM-1, and blaVIM-2 genes with the oligonucleotide primers listed in Table 1. PCR conditions for all these genes were 3 min at 94°C; 35 cycles of 1 min at 94°C, 1 min at 55°C, and 2 min at 72°C; and finally, 7 min at 72°C. The amplicons were purified with PCR Clean Up Kits (Roche Molecular Biochemicals, Mannheim, Germany) and were sequenced on an ABI PRISM 377 sequencer analyzer (Applied Biosystems, Foster City, Calif.) with the primers listed in Table 1.
TABLE 1.
Primers for PCR detection of MBL genes
| Gene | Use | Primers (5′ to 3′) | Positiona | Expected PCR product (bp) |
|---|---|---|---|---|
| blaIMP-1 | Amplification and sequencing | TGAGCAAGTTATCTGTATTC | 2 to 21 | 740 |
| TTAGTTGCTTGGTTTTGATG | 741 to 722 | |||
| Sequencing | CGGGCCTGGATAAAAAACTT | 465 to 446 | ||
| GCCACAAATTCATTTAGCGG | 394 to 413 | |||
| blaIMP-2 | Amplification | GGCAGTCGCCCTAAAACAAA | −37 to −18 | 737 |
| TAGTTACTTGGCTGTGATGG | 740 to 721 | |||
| blaVIM-1 | Amplification | TTATGGAGCAGCAACGATGT | −81 to −62 | 920 |
| CAAAAGTCCCGCTCCAACGA | 839 to 820 | |||
| blaVIM-2 | Amplification and sequencing | AAAGTTATGCCGCACTCACC | −34 to −15 | 865 |
| TGCAACTTCATGTTATGCCG | 831 to 812 | |||
| Sequencing | TCGACGGTGATGCGTACGTT | 415 to 396 | ||
| TTGATGTCCTTCGGGCGGCT | 368 to 387 |
Position number 1 for every MBL gene corresponds to the first base of the start codon.
Analytical IEF.
Crude preparations of β-lactamases were obtained by sonication (3) and were subjected to analytical isoelectric focusing (IEF) as described previously (18, 33). β-Lactamase activity was detected by overlaying the gels with 0.5 mM nitrocefin (Oxoid, Basingstoke, United Kingdom) in 50 mM HEPES (pH 7.5) supplemented with 2 mM ZnCl2 (25). Extracts from TEM-1-, TEM-10-, SHV-1-, CMY-1-, SHV-5-, CTX-M-3-, and CMY-2-producing strains were used as standards for pls of 5.4, 5.6, 7.6, 8.0, 8.2, 8.4, and 9.0, respectively. In addition, the pI was also calculated by reference to a calibration curve constructed from the relative mobility of the IEF markers (Amersham Pharmacia Biotech, Hong Kong, China).
Transfer of resistance.
Conjugation experiments were performed as described previously (23, 33) with streptomycin- and rifampin-resistant Escherichia coli C600 as the recipient (2). Tryptic soy agar plates supplemented with 500 μg of streptomycin (Sigma Chemical Company, St. Louis, Mo.) per ml or 64 μg of rifampin (Sigma) per ml and 10 μg of ceftazidime (Glaxo Group Research Ltd., Greenford, United Kingdom) per ml were used to select the transconjugants. To determine the sensitivity of the assay, direct transfer of the plasmid-encoded blaOXA-10 gene from a clinical isolate of P. aeruginosa into E. coli C600 was attempted, and transconjugant selection was performed on tryptic soy agar plates containing 100 μg of amoxicillin (SmithKline Beecham Pharmaceuticals, Surrey, United Kingdom) per ml and 64 μg of rifampin per ml.
Hybridization assays.
In order to confirm negative PCR results, colony hybridization was performed for all 209 isolates studies, as described elsewhere (7). The PCR products of blaVIM-2, blaVIM-3, and blaIMP-1 were labeled with [α-32P]dCTP (Amersham Pharmacia Bitotech) by the random priming technique with a commercial kit (GIBCO BRL, Life Technologies, Gaithersburg, Md.). The blaVIM-1- and blaIMP-2-specific probes were the PCR-generated amplicons from VIM-1-containing P. aeruginosa VR-143/97 (13) and IMP-2-containing A. baumannii AC-54/97 (25), respectively.
Plasmids from the clinical isolates harboring the MBL genes were digested with BstEII (Roche Molecular Biochemicals) overnight. The digested DNA samples were electrophoresed on 0.8% agarose gels and subjected to Southern hybridization according to the original protocol (30) with the blaVIM-2-, blaVIM-3-, and blaIMP-1-specific probes prepared as described above.
Susceptibility tests.
The MICs of the antimicrobial agents were determined by the agar dilution method according to the guidelines of the National Committee for Clinical Laboratory Standards (19). The antimicrobial agents and their sources are as follows: aztreonam, Bristol-Myers Squibb, New Brunswick, N.J.; ceftazidime, Glaxo Group Research Ltd.; cefotaxime, Hoechst-Roussel Pharmaceuticals, Inc., Somerville, N.J.; imipenem, Merck Sharp & Dohme, West Point, Pa.; meropenem, Sumitomo Pharmaceuticals Ltd., Osaka, Japan; and piperacillin and tazobactam, Lederle Laboratories, Pearl River, N.Y.
PFGE analysis.
Genomic DNAs prepared by the procedure of Piggot et al. (21) were digested overnight with 10 U of SpeI (New England Biolabs, Beverly, Mass.) and subjected to pulsed-field gel electrophoresis (PFGE) as described elsewhere (5, 33). The samples were electrophoresed with the Pulsaphor plus system (Amersham Pharmacia Biotech) at 200 V for 30 h, with pulse times ranging from 5 to 30 s.
Nucleotide sequence accession number.
The nucleotide sequence of the novel MBL described in this paper has been assigned to the GenBank database under accession number AF300454.
RESULTS
Detection of MBL producers.
Among the 209 Pseudomonas isolates tested, 21 (10 P. aeruginosa, 7 P. putida, and 4 P. stutzeri isolates) potentiated the inhibitory zones between ceftazidime disks and EDTA disks or 2-mercaptopropionic acid disks, suggesting production of MBLs, the activities of which were inhibited by EDTA and 2-mercaptopropionic acid, by these isolates.
PCR amplification and DNA sequencing.
Of the 209 Pseudomonas isolates that were subjected to PCR assays, all 21 isolates positive by the screening test gave positive PCR results, while the remaining 188 isolates were negative. Sixteen isolates (10 P. aeruginosa, 5 P. putida, and 1 P. stutzeri isolates) were positive with the primers specific for blaVIM-2, and 5 (2 P. putida and 3 P. stutzeri isolates) were positive with the primers specific for blaIMP-1. Neither blaVIM-1 nor blaIMP-2 was amplified from any of the 209 isolates. The characteristics of these isolates are shown in Table 2. The sequence analysis revealed that 5 P. putida and 1 P. stutzeri isolates carried blaVIM-2 and that 2 P. putida and 3 P. stutzeri isolates harbored blaIMP-1. All 10 P. aeruginosa isolates were found to carry a variant blaVIM-2, designated blaVIM-3. The novel MBL gene differs from blaVIM-2 by replacement of a C with an A and an A with a G at nucleotides 178 and 443 of the structural gene, respectively, which results in Gly-to-Lys and Asn-to-Ser changes at amino acid positions 32 and 120 (24), respectively, according to the numbering for Bacillus cereus β-lactamase II (24).
TABLE 2.
Characteristics of the 21 Pseudomonas isolates carrying MBL genes
| Species and isolatea | Source | Collection date (yr.mo) | pI(s) of β-lactamase(s) | MBL | Location of MBL genesb | MIC (μ/ml)c
|
PFGE profile | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PIP | TZP | CTX | CAZ | ATM | IPM | MEM | |||||||
| P. aeruginosa | |||||||||||||
| NTU-23/99 | Urine | 1999.11 | 5.1, 8.0 | VIM-3 | Chromosome | 128 | 128 | >256 | >256 | 8 | >32 | 8 | A |
| NTU-26/99 | Blood | 1999.12 | 5.1, 5.6, 8.0 | VIM-3 | Chromosome | >256 | 64 | >256 | >256 | 1 | >32 | 8 | B |
| NTU-33/99 | Tonsil | 1999.12 | 5.1, 5.6, 8.3 | VIM-3 | Chromosome | 256 | 128 | >256 | >256 | 1 | >32 | 16 | C |
| NTU-34/99 | Urine | 1999.12 | 5.1, 8.0 | VIM-3 | Chromosome | 256 | 64 | >256 | >256 | 8 | >32 | 8 | A1 |
| NTU-39/00 | Urine | 2000.1 | 5.1, 8.0 | VIM-3 | Chromosome | 128 | 64 | >256 | >256 | 16 | >32 | >32 | A2 |
| NTU-43/00 | Sputum | 2000.1 | 5.1, 5.6, 8.3 | VIM-3 | Chromosome | 256 | 64 | >256 | >256 | 16 | 16 | 4 | D |
| NTU-79/00 | Sputum | 2000.4 | 5.1, 8.0 | VIM-3 | Chromosome | >256 | >256 | >256 | >256 | 2 | >32 | >32 | E |
| PB-59 | Blood | 1997.11 | 5.1, 8.0 | VIM-3 | Chromosome | 256 | 64 | >256 | >256 | 8 | >32 | 32 | A3 |
| PS-64/99 | Sputum | 1999.12 | 5.1, 8.0 | VIM-3 | Chromosome | 256 | 32 | >256 | >256 | 0.5 | 32 | >32 | A4 |
| PS-1205/00 | Sputum | 2000.5 | 5.1, 8.0 | VIM-3 | Chromosome | 128 | 32 | >256 | >256 | 2 | >32 | 8 | A5 |
| P. putida | |||||||||||||
| NTU-91/99 | Urine | 1999.4 | 5.0, 8.9 | VIM-2 | Plasmid | 128 | 64 | >256 | >256 | 32 | >32 | >32 | F |
| NTU-93/00 | Urine | 2000.3 | 5.0, 8.9 | VIM-2 | Plasmid | >256 | >256 | >256 | 64 | 32 | >32 | >32 | G |
| PS-579/00 | Urine | 2000.2 | 5.0, 8.2 | VIM-2 | Chromosome | 256 | 128 | >256 | 128 | 8 | >32 | >32 | H |
| PS-679/00 | Urine | 2000.2 | 5.0 | VIM-2 | Chromosome | 256 | 256 | >256 | 64 | 8 | >32 | >32 | I |
| PS-693/00 | Drainage tube | 2000.2 | 5.0, 8.2 | VIM-2 | Chromosome | 256 | 256 | >256 | 64 | 64 | >32 | >32 | J |
| NTU-92/99 | Urine | 1999.9 | 8.7 | IMP-1 | Chromosome | 8 | 4 | >256 | >256 | 128 | 0.5 | 0.25 | K |
| PB-166 | Blood | 1998.11 | 6.2, 8.7 | IMP-1 | Chromosome | 64 | 32 | >256 | >256 | 64 | 16 | >32 | L |
| P. stutzeri | |||||||||||||
| PB-24 | Blood | 1997.5 | 8.7, 8.9 | IMP-1 | Chromosome | 64 | 8 | >256 | >256 | 16 | 2 | 8 | M |
| PB-101 | Blood | 1998.2 | 6.9, 8.7 | IMP-1 | Chromosome | 64 | 32 | >256 | 256 | 0.25 | 32 | 32 | N |
| PB-207 | Blood | 1999.3 | 8.7, 8.9 | IMP-1 | Chromosome | 64 | 8 | >256 | >256 | 1 | 1 | 2 | M |
| PS-1153/00 | Urine | 2000.5 | 5.0 | VIM-2 | Chromosome | 64 | 32 | >256 | 64 | 16 | >32 | >32 | O |
Isolates with the prefix NTU were from the National Taiwan University Hospital, and those with the prefix PB or PS were from the National Cheng Kung University Medical Center.
Locations of the MBL genes were suggested by Southern hybridization of plasmid DNA preparations.
PIP, piperacillin; TZP, piperacillin-tazobactam; CTX, cefotaxime; CAZ, ceftazidime; ATM, aztreonam; IPM, imipenem; MEM, meropenem.
Hybridization assays and conjugation experiments.
The results of colony hybridization assays were consistent with those obtained by PCR. All 10 VIM-3 and 6 VIM-2 producers that were recognized by PCR hybridized with both the blaVIM-3- and the blaVIM-2-specific probes, and the 5 IMP-1 producers hybridized with the blaIMP-1-specific probe. All other isolates showed negative results with the probes specific for blaVIM-1, blaVIM-2, blaVIM-3, blaIMP-1, and blaIMP-2.
Conjugation experiments failed to demonstrate transfer of carbapenem resistance from any of the isolates. Amoxicillin resistance was transferred from the control strain to E. coli C600 at a low frequency (10−8 to 10−10 per donor cell). Plasmid DNAs were isolated from three P. aeruginosa isolates (isolates NTU-39/00, PS-64/99, and PB-59) and three P. putida isolates (isolates NTU-91/99, NTU-93/00, and PB-166). For all three P. aeruginosa isolates the same restriction pattern was generated by BstEII digestion. The restriction patterns of the three P. putida isolates were different, and the blaVIM-2-specific probe hybridized to approximately 9-kb DNA fragments for two VIM-2 producers. The probes specific for blaVIM-2, blaVIM-3, and blaIMP-1 did hybridize significantly with contaminated chromosomal DNAs for the remaining four VIM-2 producers and all VIM-3 and IMP-1 producers, respectively. Therefore, the MBLs in most of these isolates seem to be chromosomally located.
Analytical IEF.
The results of IEF are summarized in Table 2. Since VIM-type MBLs are acidic enzymes (13, 22) and all VIM-3-producing P. aeruginosa isolates contained one major band that focused at pH 5.1, the band of pl 5.1 was most likely contributed by VIM-3. The bands with pls of 5.0 and 8.7 were consistent with those of VIM-2 and IMP-1, respectively (16, 22). The bands with pls ranging from 8.0 to 8.9 were possibly contributed by AmpC-type β-lactamases of Pseudomonas (4).
Susceptibility tests.
The drug susceptibilities of the 21 isolates carrying MBL genes were determined and are shown in Table 2. All these isolates were resistant to cefotaxime and ceftazidime. The susceptibilities of these isolates to aztreonam, piperacillin, and carbapenems appeared to be diverse.
PFGE analysis.
PFGE revealed that six MBL-producing P. aeruginosa isolates were closely or possibly genetically related and that two IMP-1-producing P. stutzeri isolates were genetically indistinguishable (Table 2 and Fig. 1).
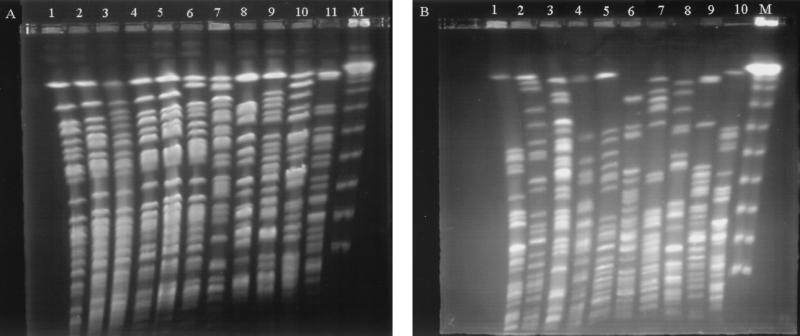
FIG. 1.
PFGE profiles of SpeI-digested genomic DNAs from 21 Pseudomonas isolates carrying IMP- or VIM-type MBL genes. (A) Lanes 1 to 9 and 11, VIM-3-producing P. aeruginosa isolates NTU-23/99, PS-1205/00, PB-59, NTU-34/99, NTU-39/00, PS-64/99, NTU-26/99, NTU-33/99, NTU-43/00, and NTU-79/00, respectively; lane 10, VIM-2-producing P. putida isolate PS-693/00; lane M, a bacteriophage lambda ladder. (B) Lanes 1, 4, 6, 9, and 10, IMP-1 producer isolates PB-207, NTU-92/99, PB-166, PB-101, and PB-24, respectively; lanes 2, 3, 5, 7, and 8, VIM-2 producers isolates PS-1153/00, NTU-91/99, NTU-93/99, PS-579/00, and PS-679/00, respectively; lane M, a bacteriophage lambda ladder.
DISCUSSION
In the present study, 21 isolates of Pseudomonas spp. were found to produce MBLs. Only 5 (23.8%) of them carried IMP-1, while 16 (76.2%) isolates harbored the VIM-2-related enzymes. These data indicate that the VIM-2-related enzymes are the most prevalent MBLs among clinical isolates of Pseudomonas spp. in Taiwan. The countrywide spread of the IMP-1 enzyme in gram-negative bacilli was reported in Japan (9, 10, 28, 29). Shortly after VIM-1 and VIM-2 were identified in Europe (13, 22), VIM-type MBLs were also found in Pseudomonas and Acinetobacter spp. in Korea (Lee et al., 40th ICAAC). These findings together indicate that both IMP- and VIM-related MBLs are widespread in eastern Asian countries. The novel enzyme, VIM-3, identified in the present study had only two amino acid differences from VIM-2 Whether the amino acid changes lead to differences in the biochemical characteristics of these two MBLs needs further investigation.
Acquired MBL genes in P. aeruginosa are often carried on plasmids and are usually nontransferable by conjugation, at least to E. coli (10, 11, 13, 22). In the present study, the MBL genes in our isolates could not be transferred to E. coli by conjugation experiments. Southern hybridization showed that only two P. putida isolates carried the blaVIM-2 gene on plasmids. All other isolates seemed to carry these MBL genes on their chromosomes. In addition, clonal spread was suggested by PFGE for six P. aeruginosa and two P. stutzeri isolates (Fig. 1). Since the blaVIM and blaIMP genes were found to be on mobile gene cassettes inserted into integrons (11, 13, 22, 25), our findings suggest that the spread of these MBL genes in Taiwan could be due to clonal dissemination as well as to movement of the resistance genes between different clones.
Consistent with the findings reported previously (9, 10, 28, 29), all our MBL-producing isolates were resistant to the broad-spectrum cephems ceftazidime and cefotaxime, while their susceptibilities to carbapenems, ureidopenicillins, and aztreonam were diverse (Table 2). Expression of MBLs could be cryptic or could be suppressed in the strains demonstrating low-level carbapenem resistance (10). The different permeability coefficients of each β-lactam may play an important role in resistance to each β-lactam in gram-negative bacteria that produce metalloenzymes (15, 17). Active efflux also acts as a contributing factor to β-lactam resistance in P. aeruginosa (14). Therefore, the lower MICs of piperacillin for some isolates could be due to their higher permeability coefficients or less efficient efflux pumps. Aztreonam is notable for its relatively low MICs for MBL-producing strains (13, 20, 22, 25). Therefore, reduced susceptibilities to aztreonam in a few isolates could be due to the presence of alternative resistance mechanisms. Isolate NTU-92/99, for which carbapenem MICs were notably low, had a relatively slow growth rate, although it possessed the biochemical and morphologic features characteristic of P. stutzeri. The slow growth rate and the difficulty in making suspensions might be partially responsible for the low carbapenem MICs for the isolate.
P. putida and P. stutzeri are environmental organisms and rarely cause human diseases (6). P. stutzeri has been reported to cause outbreaks of pseudobacteremia by contaminating antiseptics used to prepare skin (6). The medical records of the patients at the National Cheng Kung University Medical Center were reviewed. All these isolates were considered to be from the hospital environment. The three P. stutzeri and one P. putida isolates from blood samples and the P. putida isolate recovered from a drainage tube were of no clinical significance. They could originate from contaminated antiseptics, the patients' skin, blood culture bottles, or the drainage tube. The two P. putida and one P. stutzeri isolates from urine samples were all urinary catheterization related, and their clinical significance was not clear. Our findings suggest that the environmental organisms acquired the MBL genes under long-term selection pressure in the hospital environment.
Early detection of MBL-producing strains is important for clinicians for the selection of appropriate antimicrobial agents. PCR assays with primers specific for blaIMP-1 have been shown to be useful and reliable for detection of IMP-1 producers (29). Given that the numbers of metalloenzymes are increasing (11, 13, 22, 25), the use of multiple primers is needed. Since the PCR method is not cost-effective for use for daily testing in clinical laboratories, Arakawa et al. (1) have designed a simple disk diffusion test for screening for MBL-producing gram-negative rods; the test has a high degree of specificity and a high degree of sensitivity for the detection of IMP-1 producers. The results obtained by the screening test in the present study also correlated well with those obtained by the PCR method, indicating that the test is very useful for the detection not only of IMP-1-producing strains but also of VIM producers.
In conclusion, both IMP- and VIM-type MBLs have emerged in Pseudomonas spp. in Taiwan, and the VIM-2-related enzymes were the most prevalent types. The MBL genes were largely located on chromosomes, and their dissemination in Taiwan could be due to both mobilization of the resistance genes and clonal spread of the resistant strains.
ACKNOWLEDGMENTS
We kindly thank G. M. Rossolini, Dipartimento di Biologia Molecolare, Sezione Di Microbiologia, Università di Siena, Siena, Italy, for provision of IMP-2-containing A. baumannii strain AC-54/97 and VIM-1-containing P. aeruginosa strain VR-143/97.
This work was partially supported by grants NCKUH89-054 from National Cheng Kung University Hospital and NSC89-2320-B-006-149 from the National Science Council of the Republic of China.
REFERENCES
- 1.Arakawa Y, Shibata N, Shibayama K, Kurokawa H, Yagi T, Fugiwara H, Goto M. Convenient test for screening metallo-β-lactamase-producing gram-negative bacteria by using thio compounds. J Clin Microbiol. 2000;38:40–43. doi: 10.1128/jcm.38.1.40-43.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bachmann B J, Low K B. Linkage map of Escherichia coli K-12, edition 6. Microbiol Rev. 1980;44:1451–1456. doi: 10.1128/mr.44.1.1-56.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bauernfeind A, Grimm H, Schweighart S. A new plasmidic cefotaximase in a clinical isolate of Escherichia coli. Infection. 1990;18:294–298. doi: 10.1007/BF01647010. [DOI] [PubMed] [Google Scholar]
- 4.Bush K, Jacoby G A, Medeiros A A. A functional classification scheme for β-lactamases and its correlation with molecular structure. Antimicrob Agents Chemother. 1995;39:1211–1233. doi: 10.1128/aac.39.6.1211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chu G, Vollrath D, Davis R W. Separation of large DNA molecules by contour-clamped homogeneous electric fields. Science. 1986;234:1582–1585. doi: 10.1126/science.3538420. [DOI] [PubMed] [Google Scholar]
- 6.Gilligan P H, Whittier S. Pseudomonas and Burkholderia. In: Murray P R, Baron E J, Pfaller M A, Tenover F C, Yolken R H, editors. Manual of clinical microbiology. 7th ed. Washington, D.C.: American Society for Microbiology; 1999. pp. 517–525. [Google Scholar]
- 7.Grunstein M, Hogness D S. Colony hybridization: a method for the isolation of cloned DNAs that contain a specific gene. Proc Natl Acad Sci USA. 1975;72:3961–3965. doi: 10.1073/pnas.72.10.3961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hancock R E W. Resistance mechanisms in Pseudomonas aeruginosa and other nonfermentative gram-negative bacteria. Clin Infect Dis. 1998;27(Suppl. 1):S93–S99. doi: 10.1086/514909. [DOI] [PubMed] [Google Scholar]
- 9.Hirakata Y, Izumikawa K, Yamaguchi T, Takemura H, Tanaka H, Yoshida R, Matsuda J, Nakano M, Tomono K, Maesaki S, Kaku M, Yamada Y, Kamihira S, Kohno S. Rapid detection and evaluation of clinical characteristics of emerging multiple-drug-resistant gram-negative rods carrying the metallo-β-lactamase gene blaIMP. Antimicrob Agents Chemother. 1998;42:2006–2011. doi: 10.1128/aac.42.8.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ito H, Arakawa Y, Ohsuka S, Wacharotayankun R, Kato N, Ohta M. Plasmid-mediated dissemination of the metallo-β-lactamase gene blaIMP among clinical isolated strains of Serratia marcescens. Antimicrob Agents Chemother. 1995;39:824–829. doi: 10.1128/aac.39.4.824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Iyobe S, Kusadokoro H, Ozaki J, Matsumura N, Minami S, Haruta S, Sawai T, O'Hara K. Amino acid substitutions in a variant of IMP-1 metallo-β-lactamase. Antimicrob Agents Chemother. 2000;44:2023–2027. doi: 10.1128/aac.44.8.2023-2027.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Jacoby G A, Medeiros A A. More extended-spectrum β-lactamases. Antimicrob Agents Chemother. 1991;35:1697–1704. doi: 10.1128/aac.35.9.1697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lauretti L, Riccio M L, Mazzariol A, Cornaglia G, Amicosante G, Fontana R, Rossolini G M. Cloning and characterization of blaVIM, a new integron-borne metallo-β-lactamase gene from a Pseudomonas aeruginosa clinical isolate. Antimicrob Agents Chemother. 1999;43:1584–1590. doi: 10.1128/aac.43.7.1584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Li X Z, Ma D, Livermore D M, Nikaido H. Role of efflux pump(s) in intrinsic resistance of Pseudomonas aeruginosa: active efflux as a contributing factor to β-lactam resistance. Antimicrob Agents Chemother. 1994;38:1742–1752. doi: 10.1128/aac.38.8.1742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Livermore D M. Interplay of impermeability and chromosomal β-lactamase activity in imipenem-resistant Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1992;36:2046–2048. doi: 10.1128/aac.36.9.2046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Marumo K, Takeda A, Nakamura Y, Nakaya K. Purification and characterization of metallo-β-lactamase from Serratia marcescens. Microbiol Immunol. 1995;39:27–33. doi: 10.1111/j.1348-0421.1995.tb02164.x. [DOI] [PubMed] [Google Scholar]
- 17.Matsumura N, Minami S, Watanabe Y, Iyobe S, Mitsuhashi S. Role of permeability in the activities of β-lactams against gram-negative bacteria which produce a group 3 β-lactamase. Antimicrob Agents Chemother. 1999;43:2084–2086. doi: 10.1128/aac.43.8.2084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Matthew M, Harris M, Marshall M J, Rose G W. The use of analytical isoelectric focusing for detection and identification of β-lactamases. J Gen Microbiol. 1975;88:169–178. doi: 10.1099/00221287-88-1-169. [DOI] [PubMed] [Google Scholar]
- 19.National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial susceptibility testing; ninth informational supplement. M100–S9. Wayne, Pa: National Committee for Clinical Laboratory Standards; 1999. [Google Scholar]
- 20.Osano E, Arakawa Y, Wacharotayankun R, Ohta M, Horii T, Ito H, Yoshimura F, Kato N. Molecular characterization of an enterobacterial metallo-β-lactamase found in a clinical isolates of Serratia marcescens that shows imipenem resistance. Antimicrob Agents Chemother. 1994;38:71–78. doi: 10.1128/aac.38.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Piggot P J, Amjad M, Wu J J, Sandoval H, Castro J. Genetic and physical maps of Bacillus subtilis 168. In: Harwood C R, Cutting S M, editors. Molecular biology methods for bacillus. West Sussex, England: John Wiley & Sons Ltd.; 1990. pp. 493–543. [Google Scholar]
- 22.Poirel L, Naas T, Nicolas D, Collet L, Bellais S, Cavallo J D, Nordmann P. Characterization of VIM-2, a carbapenem-hydrolyzing metallo-β-lactamase and its plasmid- and integron-borne gene from a Pseudomonas aeruginosa clinical isolate in France. Antimicrob Agents Chemother. 2000;44:891–897. doi: 10.1128/aac.44.4.891-897.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Provence D L, Curtiss R., III . Gene transfer in gram-negative bacteria. In: Gerhardt P, Murray R G E, Wood W A, Krieg N R, editors. Methods for general and molecular bacteriology. Washington, D.C.: American Society for Microbiology; 1994. pp. 319–347. [Google Scholar]
- 24.Rasmussen B A, Bush K. Carbapenem-hydrolyzing β-lactamases. Antimicrob Agents Chemother. 1997;41:223–232. doi: 10.1128/aac.41.2.223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Riccio M L, Franceschini N, Boschi L, Caravelli B, Cornaglia G, Fontana R, Amicosante G, Rossolini G M. Characterization of the metallo-β-lactamase determinant of Acinetobacter baumannii AC-54/97 reveals the existence of blaIMP allelic variants carried by gene cassettes of different phylogeny. Antimicrob Agents Chemother. 2000;44:1229–1235. doi: 10.1128/aac.44.5.1229-1235.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sanders C C. Chromosomal cephalosporinases responsible for multiple resistance to newer β-lactam antibiotics. Annu Rev Microbiol. 1987;41:573–593. doi: 10.1146/annurev.mi.41.100187.003041. [DOI] [PubMed] [Google Scholar]
- 27.Schiappa D A, Hayden M K, Matushek M G, Hashemi F N, Sullivan J, Smith K Y, Miyashiro D, Quinn J P, Weinstein R A, Trenholme G M. Ceftazidime-resistant Klebsiella pneumoniae and Escherichia coli bloodstream infection: a case-control and molecular epidemiologic investigation. J Infect Dis. 1996;174:529–536. doi: 10.1093/infdis/174.3.529. [DOI] [PubMed] [Google Scholar]
- 28.Senda K, Arakawa Y, Nakashima K, Ito H, Ichiyama S, Shimokata K, Kato N, Ohta M. Multifocal outbreaks of metallo-β-lactamase-producing Pseudomonas aeruginosa resistant to broad-spectrum-β-lactams, including carbapenems. Antimicrob Agents Chemother. 1996;40:349–353. doi: 10.1128/aac.40.2.349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Senda K, Arakawa Y, Ichiyama S, Nakashima K, Ito H, Ohsuka S, Shimokata K, Kato N, Ohta M. PCR detection of metallo-β-lactamase gene (blaIMP) in gram-negative rods resistant to broad-spectrum β-lactams. J Clin Microbiol. 1996;34:2909–2913. doi: 10.1128/jcm.34.12.2909-2913.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Southern E M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975;98:503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- 31.Takahashi S, Nagano Y. Rapid procedure for isolation of plasmid DNA and application to epidemiological analysis. J Clin Microbiol. 1984;20:608–613. doi: 10.1128/jcm.20.4.608-613.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Tsakris A, Pournaras S, Woodford N, Palepou M-F I, Babini G S, Douboyas J, Livermore D M. Outbreak of infections caused by Pseudomonas aeruginosa producing VIM-1 carbapenemase in Greece. J Clin Microbiol. 2000;38:1290–1292. doi: 10.1128/jcm.38.3.1290-1292.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yan J J, Wu S M, Tsai S H, Wu J J, Su I J. Prevalence of SHV-12 among clinical isolates of Klebsiella pneumoniae producing extended-spectrum β-lactamase and identification of a novel AmpC enzyme (CMY-8) in southern Taiwan. Antimicrob Agents Chemother. 2000;44:1438–1442. doi: 10.1128/aac.44.6.1438-1442.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]