Abstract
A total of 113 blood culture isolates of Klebsiella pneumoniae from 10 hospitals in northern Taiwan were studied for SHV and TEM β-lactamase production. blaSHV was amplified from all isolates by PCR. TEM-type resistance, was found in 32 of the isolates and was of the TEM-1 type in all isolates. SHV-1, -2, -5, -11, and -12 and two novel enzymes were identified. These novel enzymes were designated SHV-25 and SHV-26 and had pIs of 7.5 and 7.6, respectively. Amino acid differences in comparison to the amino acid sequence of blaSHV-1 were found at positions T18A (ThrACC→AlaGCC), L35Q (LeuCTA→GluCAA), and M129V (MetATG→ValGTG) for SHV-25 and at position A187T (AlaGCC→ThrACC) for SHV-26. The results of substrate profiles and MIC determinations showed that the novel enzymes did not hydrolyze extended-spectrum cephalosporins, rendering the isolates susceptible to these agents. Inhibition profiles revealed that the 50% inhibitory concentration for SHV-26 was higher than those for SHV-1 and SHV-25, resulting in an intermediate resistance to amoxicillin-clavulanic acid. Forty-nine ribotypes were identified, suggesting that major clonal spread had not occurred in any of the hospitals. According to the amino acid sequence, SHV β-lactamases in Taiwan may basically be derived through stepwise mutation from SHV-1 or SHV-11 and further subdivided by four routes. The stepwise mutations initiated from SHV-1 or SHV-11 to SHV-2, SHV-5, and SHV-12 comprise the evolutionary change responsible for extended-spectrum β-lactamase (ESBL) production in Taiwan. The stepwise mutations that lead to a non-ESBL (SHV-25) and the β-lactamase (SHV-26) with reduced susceptibility to clavulanic acid are possibly derived from SHV-11 and SHV-1, respectively. The results suggest a stepwise evolution of SHV β-lactamases in Taiwan.
Extended-spectrum β-lactamases (ESBLs) are predominantly derived from plasmid-mediated TEM or SHV β-lactamases through a mutation or mutations that lead to one or more amino acid changes and that result in the alteration of the binding to and hydrolysis of specific substrates at the active site (6; G. A. Jacoby and K. Bush, http://www.lahey.org/studies /webt.htm). These enzymes, especially TEM and SHV, are most commonly found in Klebsiella pneumoniae and Escherichia coli and have been reported worldwide (6, 17, 19; Jacoby and Bush, http://www.lahey.org/studies/webt.htm). ESBLs have also recently been found in other members of the enteric bacteria (2, 8, 16). Most of these ESBLs are associated with nosocomial infections and treatment with antibiotics prior to the infection. Selective pressure has been suggested to enhance stepwise mutations in the nucleotides of classic β-lactamases (9).
One study observed that the SHV-type gene is ubiquitous in K. pneumoniae (1). The classic SHV-1 gene, which is normally encoded by plasmids of E. coli or other members of the family Enterobacteriaceae, is usually encoded by the chromosome in K. pneumoniae. It has been postulated that SHV-1 may have originated by separation from the chromosome of K. pneumoniae and extrachromosomal spread to other bacteria (1). This observation may be helpful in sketching the evolution of specific β-lactamases in certain localities. In the present report, we describe the molecular epidemiology of genes associated with TEM-type and SHV-type β-lactamases and two novel β-lactamases, SHV-25 and SHV-26, from K. pneumoniae isolates from 10 hospitals in northern Taiwan.
MATERIALS AND METHODS
Bacterial strains.
A total of 361 blood culture K. pneumoniae isolates from 10 hospitals located in northern Taiwan were collected in 1998 and 1999. Forty-three of 291 cefazolin-susceptible K. pneumoniae isolates were selected randomly due to a large number of cefazolin-susceptible isolates among the 10 hospitals. The remaining 70 isolates (for a total of 113 isolates) which were ceftazidime or cefotaxime intermediate or resistant were all included in the study.
Antimicrobial susceptibility testing.
The antimicrobial susceptibilities of the isolates were redetermined concomitantly by the disk diffusion and the agar dilution methods as described in National Committee for Clinical Laboratory Standards guidelines (12, 13). For susceptibility testing by the broth microdilution method, the following antimicrobial agents were obtained as standard reference powders of known potency for laboratory use: ampicillin and cephalothin from Sigma Chemical Co. (St. Louis, Mo.), clavulanic acid from SmithKline Beecham (Brockhans Park, United Kingdom), cefmetazole from Upjohn Co. (Kalamazoo, Mich.), imipenem from Merck Sharp & Dohme (West Point, Pa.), cefotaxime and gentamicin from Hoechst Marion Roussel (Frankfurt, Germany), ceftazidime from Glaxo Group Research Limited (Greenford, United Kingdom), aztreonam from Bristol-Myers Squibb Laboratories (Princeton, N.Y.), and ciprofloxacin from Bayer Co. (Leverkusen, Germany). All drugs were incorporated into Mueller-Hinton agar (Becton Dickinson Microbiology Systems, Sparks, Md.) in serial twofold concentrations from 0.03 to 128 μg/ml. Two control strains, E. coli ATCC 35218 and ATCC 25922, were included in each set of tests. The plates were incubated in ambient air at 35°C for 16 to 18 h. The MIC of each antimicrobial agent was defined as the lowest concentration which inhibited visible growth of the organism.
The Etest (AB Biodisk, Solna, Sweden) was performed according to the manufacturer's instruction.
PCR amplification for detection of blaTEM and blaSHV.
The oligonucleotide primers used for PCR assays were as follows: 5′-ATAAAATTCTTGAAGACGAAA (primer A), 5′-GACAGTTACCAATGCTTAATCA (primer B), 5′-GGGTTATTCTTATTTGTCGC (primer C), and 5′-TTAGCGTTGCCAGTGCTC (primer D). Oligonucleotides were synthesized by GIBCO BRL (Grand Island, N.Y.). Primers A and B are known to be specific for blaTEM (7). Primers C and D are known to be specific for blaSHV (14).
Reactions were performed in a DNA Thermal Cycler (MJ Research Inc., Watertown, Mass.) in 50-μl reaction mixtures containing 2.5 U of Taq polymerase (Promega, Madison, Wis.), 1× buffer (consisting of 10 mM Tris-HCl [pH 8.3], 1.5 mM MgCl2, and 50 mM KCI), 0.01 μg of gelatin, each deoxynucleoside triphosphate at a concentration of 200 μM, and each oligonucleotide primer at a concentration of 2 μM. Thirty-five cycles with the following temperature profiles were performed for each reaction: 94°C, for 1 min, 58°C for 1 min, and 72°C for 1 min.
For direct DNA sequencing, PCR products were purified with Microspin S-300 HR PCR purification columns (Pharmacia, Uppsala, Sweden). Sequencing reactions were performed with consecutive primers specific for the blaTEM and blaSHV genes (9, 14) by the method of Sanger et al. (15). An automatic sequencer (model 377; ABI Prism; Perkin-Elmer, Norwalk, Conn.) was used.
Cloning of SHV-25 and SHV-26.
Entire SHV-25 and SHV-26 resistance genes were amplified by PCR with one set of primers. The primers used for PCR were as follows: (i) SHV-F (5′-GGGGAATTCTTATTTGTCGC) and (ii) SHV-R (5′-CAGAATTCGCTTAGCGTTGCCAGT) The PCR products were ligated with a phagemid vector (pPCR-Script CamSK+). This cloning vector includes a chloramphenicol resistance gene, a lac promoter for gene expression, T3 and T7 RNA polymerase promoters for in vitro production of RNA, and an f1 intergenic region for single-stranded DNA rescue. Ligated vectors were then transformed to calcium-treated competent E. coli DH5α cells by the ligation kit polishing protocol (Stratagene, La Jolla, Calif.). Tranformants were selected on a Mueller-Hinton agar plate containing 50 μg of ampicillin, 50 μg of chloramphenicol, and isopropyl-β-d-thiogalactopyranoside–5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside.
Isoelectric focusing.
Cells were harvested from 20-h brain heart infusion broth cultures by centrifugation, and the pellet was resuspended in 1 ml of phosphate buffer (10 mM; pH 7). The enzymes were released by two cycles of freezing (at −70°C) and thawing at room temperature and sonication for 5 min in a sonicator in ice-cold water. Isoelectric focusing was performed in an ampholine gel (pH 3.0 to 10.0; Pharmacia). Preparations from standard strains known to harbor TEM-1, SHV-1, and SHV-5 were used as standards. After isoelectric focusing, β-lactamases were detected by spreading nitrocefin (50 μg/ml) on the gel surface (11).
β-Lactamase substrate and inhibition profiles.
Crude β-lactamase extracts from the cloned SHV-1, SHV-25, and SHV-26 enzymes were used for substrate and inhibition assays (4). The assays were performed spectrophotometrically by measuring the change in absorbance at the appropriate wavelength for each substrate. The wavelengths were 240 nm for benzylpenicillin; 260 nm for cephaloridine, cephalothin, cefotaxime, and ceftazidime; and 500 nm for nitrocefin. β-Lactamase activity was determined in 1 ml of all substrates (except benzylpenicillin) at 0.01, 0.02, 0.04, 0.08, and 0.1 mM in 10 mM phosphate buffer (pH 7.0); benzylpenicillin was assayed at 0.1 mM, followed by concentrations of 0.2, 0.3, 0.4, 0.5, 0.6, 0.8, and 1 mM. One unit of activity was defined as the amount of enzyme that hydrolyzed 1 mol of benzylpenicillin per min per mg of total protein at room temperature. The rates of hydrolysis for each substrate were calculated relative to that of benzylpenicillin. Vmax and Km were determined from a linear plot of the hydrolysis rate for each substrate versus substrate concentration.
The concentration of clavulanate required to inhibit enzymatic activity by 50% (IC50) was determined as follows. After 10 min of preincubation at room temperature of equal amounts of enzyme from each isolate with various amounts of clavulanate, enzyme hydrolytic activities were measured by using nitrocefin as the substrate.
Ribotyping.
Ribotyping was performed with the automated Riboprinter Microbial Characterization System (Qualicon, Wilmington, Del.) according to the manufacturer's instructions. Colonies were picked and loaded into the Riboprinter Microbial Characterization Unit (MCU). In the MCU, total DNA was digested with the EcoRI enzyme, separated by electrophoresis, and then transferred directly to nylon membranes. Ribopatterns were expressed by hybridization with a chemiluminescence-labeled DNA probe containing an rRNA operon (rrnB) from E. coli. The patterns were automatically imaged and stored in the MCUs computer. The positions of standard markers were used to correct for both lane-to-lane and membrane-to-membrane variations in band positions. The ribopattern for each isolate was compared to other patterns in the Riboprinter database. The system assigns isolates to a particular ribogroup on the basis of differences in band numbers, band position, and signal intensity at a given band position (3).
Nucleotide sequence accession numbers.
The sequences of the blaSHV-25 and blaSHV-26 genes have been deposited in GenBank and have been given GenBank accession numbers AF208796 and AF227204, respectively.
RESULTS
Antimicrobial susceptibility testing results.
All 113 K. pneumoniae isolates were uniformly resistant to ampicillin. Most of the isolates were resistant to cephalothin (62.0%), followed by ceftazidime (51.3%), cefotaxime (50.4%), aztreonam (48.7%), gentamicin (42.3%), and ciprofloxacin (20.4%). The rate of resistance to amoxicillin-clavulanic acid was low (5.3%). All isolates were sensitive to imipenem (Table 1).
TABLE 1.
Antimicrobial susceptibility testing results for 113 blood culture isolates of K. pneumoniae in northern Taiwan
| β-Lactamase (no. of isolates)a | MIC range (μg/ml)b
|
|||||||
|---|---|---|---|---|---|---|---|---|
| AMZ/CLA | CEF | ATM | CTX | CAZ | FOX | CIP | GEN | |
| SHV-1 (37) | 1–≥128 | 2–≥128 | ≤0.03–≥128 | ≤0.25–≥64 | ≤0.25–32 | 4–16 | 0.06–≥32 | 0.5–≥128 |
| SHV-11 (33) | 1–16 | 4–≥128 | ≤0.03–≥128 | ≤0.25–≥64 | ≤0.25–≥32 | 4–64 | ≤0.03–≥32 | 1–≥128 |
| SHV-2 (2) | 8 | ≥128 | 2–≥128 | 16–≥64 | ≥32 | 8 | 0.5–≥32 | 8–128 |
| SHV-5 (8) | 4–≥128 | ≥128 | ≤0.03–≥128 | 8–≥64 | 8–≥32 | 8–16 | 0.5–≥32 | 1–≥128 |
| SHV-12 (31) | 4–64 | ≥128 | 2–≥128 | 8–≥64 | 8–≥32 | 4–≥128 | ≤0.03–≥32 | 1–≥128 |
Type of SHV enzyme carried in the isolates. All isolates were resistant to ampicillin and susceptible to imipenem.
Abbreviations: AMZ/CLA, amoxicillin-clavulanate; CEF, cephalothin; ATM, aztreonam; CTX, cefotaxime; CAZ, ceftazidime; FOX, cefoxitin; CIP, ciprofloxacin; GEN, gentamicin. Susceptibility testing results for SHV-25 and SHV-26 carriers are presented in Table 2. The numbers of isolates, including SHV-25 and SHV-26 carriers, that were susceptible, intermediate, and resistant were as follows: for amoxicillin-clavulanate 82, 25, and 6, respectively; for cephalothin, 43, 0, and 70, respectively; for aztreonam, 56, 2, and 55, respectively; for cefotaxime, 53, 3, and 57, respectively; for ceftazidime, 55, 0, and 58, respectively; for cefoxitin, 80, 15, and 18, respectively; for ciprofloxacin, 79, 11, and 23, respectively; for gentamicin, 52, 13, and 48, respectively. The MICs at which 50 and 90% of isolates were inhibited were as follows for the eight antibiotics: 4 and 16 μg/ml, ≥128 and ≥128 μg/ml, 16 and ≥128 μg/ml, ≥64 and ≥64 μg/ml, ≥32 and ≥32 μg/ml, 4 and 128 μg/ml, 0.25 and ≥32 μg/ml, and 8 and ≥128 μg/ml, respectively.
PCR for detection of blaTEM and blaSHV resistance genes and sequencing results.
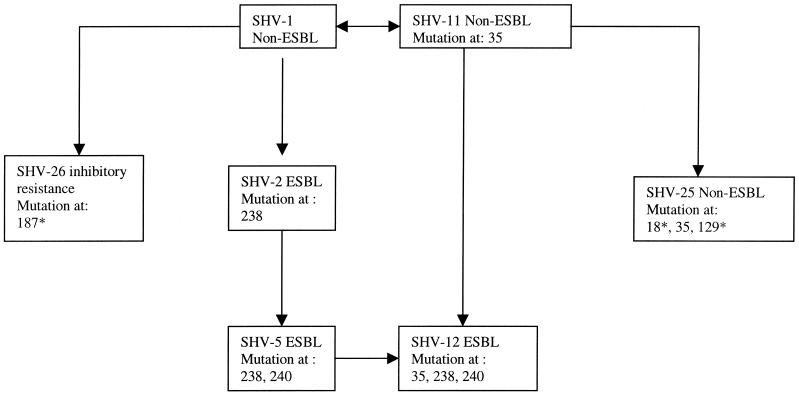
blaSHV amplification was achieved for all K. pneumoniae isolates. Thirty-two isolates were found to have coexisting TEM-type genes. Sequencing results revealed that all 32 TEM-type resistance genes were of the TEM-1 type. Among the SHV-type resistance genes, SHV-1, -2, -5, -11, and -12 and two novel SHV-type resistance genes were identified. Thirty-seven isolates carried the gene for SHV-1, 33 carried the gene for SHV-11, 2 carried the gene for SHV-2, 8 carried the gene for SHV-5, 31 carried the gene for SHV-12, and 1 each carried the genes for SHV-25 and SHV-26 (Table 1). The amino acid sequences of the two novel SHV-type β-lactamases differed from that of the classic SHV-1 β-lactamase, and the β-lactamases were provisionally designated SHV-25 and SHV-26. For SHV-25, the amino acids were changed at positions T18A (ThrACC→AlaGCC), L35Q (LeuCTA→GluCAA), and M129V (MetATG→ValGTG). For SHV-26, a single amino acid change at position A187T (AlaGCC→ThrACC) was found. Of the amino acid changes in SHV-25 and SHV-26, only the amino acid change at position 35 has been described previously, where this change was previously reported in SHV-11 and SHV-12 (6). On the basis of these results, a correlation tree for SHV-type resistance genes in Taiwan was drawn and the possible stepwise nucleotide mutations which caused the emergence of the ESBL producers was postulated (Fig. 1).
FIG. 1.
Correlation tree for SHV-type resistance genes in Taiwan. The asterisks next to the numbers indicate novel mutations.
Automated ribotyping.
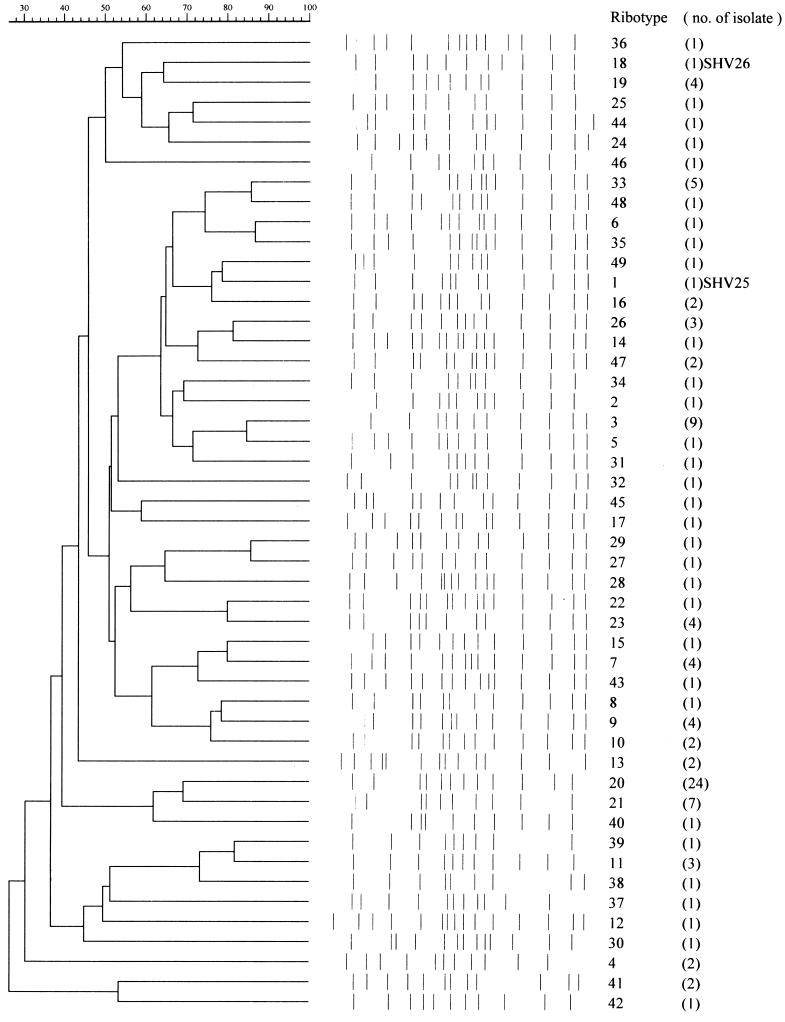
Forty-nine ribotypes were identified among the 113 isolates. The number of different ribotypes in each hospital ranged from 3 to 20. Since the isolates were randomly selected if isolates were sensitive to cephalothin, only a few isolates were selected from the hospital with the smaller number of ribotypes. A relatively large number of cefotaxime- or ceftazidime-resistant isolates was found in the hospital with the largest number of ribotypes. Ribotyping provided no evidence suggestive of clonal spread in any of the hospitals. Twenty-four of the 113 isolates were of ribotype 20, and these were distributed evenly among different hospitals. SHV-25 and SHV-26 carriers were in ribogroups 1 and 18, respectively. Only one strain of each of these ribotypes was found (Fig. 2).
FIG. 2.
Ribotypes for 113 K. pneumoniae isolates. Similarity was calculated by the unweighted pair group method with arithmetic averages.
MICs for original isolates and cloned SHV-25 and SHV-26 carriers by Etest.
The original and cloned SHV-25 carriers were found to be ampicillin resistant. They were susceptible to amoxicillin-clavulanate, cephalothin, cefoxitin, cefotaxime, ceftazidime, aztreonam, ciprofloxacin, gentamicin, and imipenem. The original SHV-26 carrier isolate was resistant to ampicillin, cephalothin, cefoxitin, cefotaxime, ceftazidime, and aztreonam; had intermediate resistance to amoxicillin-clavulanate; and was susceptible to ciprofloxicin and imipenem. The cloned SHV-26 carrier retained its resistance to ampicillin and cephalothin and its intermediate resistance to amoxicillin-clavulanate. The cloned SHV-26 carrier was susceptible to cefoxitin, cefotaxime, ceftazidime, aztreonam, ciprofloxicin, and imipenem (Table 2).
TABLE 2.
MICs for original isolates and cloned SHV-25 and SHV-26 carriers by Etest
| Antibiotic | MIC (μg/ml)
|
|||||
|---|---|---|---|---|---|---|
| DH5α | Cloned SHV-1 carrier | Clinical SHV-25 carrier | Cloned SHV-25 carrier | Clinical SHV-26 carrier | Cloned SHV-26 carrier | |
| Ampicillin | 3 | ≥128 | ≥128 | ≥128 | ≥128 | ≥128 |
| Amoxicillin-clavulanate | 3 | 6 | 6 | 3 | 16 | 16 |
| Cephalothin | 0.19 | 6 | 4 | 6 | ≥256 | ≥256 |
| Aztreonam | 0.094 | 0.094 | 0.094 | 0.094 | ≥256 | 0.19 |
| Cefotaxime | ≤0.125 | ≤0.125 | ≤0.125 | ≤0.125 | ≥32 | ≤0.125 |
| Ceftazidime | ≤0.25 | ≤0.25 | 0.5 | ≤0.25 | ≥64 | 1 |
| Cefoxitin | 2 | 4 | 6 | 4 | 12 | 4 |
| Ciprofloxacin | 0.023 | 0.023 | 1.5 | 0.064 | 0.5 | 0.047 |
| Gentamicin | 0.094 | 0.125 | 24 | 1.5 | 32 | 0.19 |
| Imipenem | 0.25 | 0.25 | 0.19 | 0.25 | 0.19 | 0.25 |
Isoelectric focusing of original isolates and cloned SHV-25 and SHV-26 carrier.
Isoelectric focusing found in the SHV-25 carrier only one β-lactamase with a pI of 7.5. Three different β-lactamases with pIs of 5.4, 7.6, and >8.2 were identified in the original SHV-26 carrier. The β-lactamase with a pI of 7.6 was weakly produced, as shown by the slow appearance of a hydrolytic band at a pI of 7.6 compared to the bands for the other two enzymes. For the cloned SHV-26 carrier, a band with a pI of 7.6 was identified (Fig. 3).
FIG. 3.
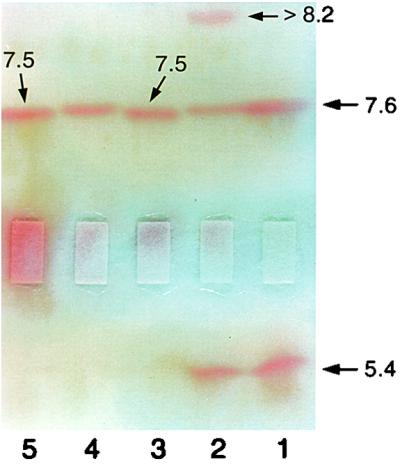
Isoelectric focusing of SHV-25 and SHV-26 carriers and the E. coli strains into which SHV-25 and SHV-26 were cloned. Lane 1, TEM-1 and SHV-1 standard; lane 2, SHV-26 carrier; lane 3, SHV-25 carrier; lane 4, cloned SHV-26 carrier; lane 5, cloned SHV-25 carrier. The numbers indicate pI values.
Substrates and inhibition profiles for SHV-25 and SHV-26.
SHV-1, SHV-25, and SHV-26 had very similar substrate profiles, and none of them hydrolyzed cefotaxime or ceftazidime (Table 3). All of them had relatively slower hydrolysis rates (Vmaxs) for cephaloridine and cephalothin compared to those for penicillin G. SHV-25 had a lower relatively Km than those of SHV-1 and SHV-26. The IC50s for SHV-1 and SHV-25 were three times lower than that for SHV-26 (Table 3).
TABLE 3.
Substrates and inhibition profiles for SHV-25 and SHV-26
| Substrate or inhibitora | SHV-1
|
IC50 (μg/ml)c | SHV-25
|
IC50 (μg/ml) | SHV-26
|
IC50 (μg/ml) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vmaxb | Km | Relative Vmax (%) | Relative Vmax/Km | Vmax | Km | Relative Vmax (%) | Relative Vmax/Km | Vmax | Km | Relative Vmax (%) | Relative Vmax/Km | ||||
| Clavulanic acid | 31 ± 0.28 | 26 ± 0.04 | 96 ± 0.41 | ||||||||||||
| PenG | 0.93 ± 0.01 | 53.3 ± 0.35 | 100 | 100 | 51.0 ± 0.59 | 297 ± 0.37 | 100 | 100 | 14.0 ± 0.16 | 512 ± 1.23 | 100 | 100 | |||
| CEF | 0.03 ± 0.004 | 21.2 ± 0.15 | 3.3 | 8.33 | 1.12 ± 0.07 | 3.72 ± 0.1 | 2.19 | 175.2 | 0.51 ± 0.09 | 81.5 ± 1.3 | 3.65 | 22.9 | |||
| LOR | 0.39 ± 0.03 | 147 ± 0.47 | 41.9 | 15.2 | 19.0 ± 0.46 | 63.3 ± 2.53 | 37.2 | 175.1 | 3.10 ± 0.05 | 118 ± 0.5 | 22.2 | 96 | |||
| CTX | <0.01 | NDd | ND | ND | <0.01 | ND | ND | ND | <0.01 | ND | ND | ND | |||
| CAZ | <0.01 | ND | ND | ND | <0.01 | ND | ND | ND | <0.01 | ND | ND | ND | |||
Abbreviations: PenG, penicillin G; CEF, cephalothin; LOR, cephaloridine; CTX, cefotaxime; CAZ, ceftazidime.
Nanomoles of substrate hydrolyzed per minute per milligram of protein.
Inhibitory concentration of clavulanic acid required to reduce nitrocefin hydrolytic activity by 50%.
ND, not determined.
DISCUSSION
In the present study, we have tried to analyze a type of resistance gene that is located in the chromosome of K. pneumoniae. Since the SHV type of resistance has been postulated to be present in all K. pneumoniae strains (1), studies with this type of resistance gene may allow more precise delineation of the correlation or, possibly, the evolution of different SHV-related mutations among all isolates.
MIC determinations showed that all of the isolates were resistant to ampicillin. On the other hand, 72.6% of the isolates were susceptible to amoxicillin-clavulanate, indicating that resistance was inhibited by a β-lactamase inhibitor. Twenty-five (22.1%) isolates were partially inhibited by amoxicillin-clavulanate, and only 6 (5.3%) isolates were not. Among 70 cephalothin-resistant isolates, >48% were resistant to aztreonam, cefotaxime, or ceftazidime. However, most of these isolates were shown to be sensitive to cefoxitin, indicating that the production of β-lactamases among these isolates had no effect on susceptibility to the cephamycin class of antibiotics. Although imipenem resistance due to the presence of a combination of an outer membrane protein and AmpC has previously been reported in Spain (10), imipenem is still a drug to which isolates were susceptible in this study.
Our study shows agreement with the conclusion that SHV-type resistance genes are ubiquitous in K. pneumoniae (1). blaSHV was amplified from all isolates, and seven different types of SHV β-lactamases were identified, including two novel β-lactamases, SHV-25 and SHV-26. blaTEM was amplified from 32 isolates, and all of these isolates carried the TEM-1 resistance gene. According to their amino acid sequences, SHV β-lactamases in Taiwan may basically come from SHV-1 or SHV-11 and are further subdivided by four possible routes. Through stepwise mutation, SHV-1 may form SHV-2 (mutation at amino acid 238) and then SHV-5 (mutations at amino acids 238 and 240). SHV-12 (mutations at amino acids 35, 238, and 240) may possibly come from either SHV-5 or SHV-11 (Fig. 1). These stepwise mutations appear to comprise the evolutionary history for SHV-type ESBL producers in Taiwan. On the other hand, a non-ESBL (SHV-25) and a β-lactamase that results in reduced susceptibility to clavulanic acid (SHV-26) are possibly derived from SHV-11 and SHV-1. Ribotyping revealed that there were a total 49 different ribotypes, suggesting a high degree of genetic polymorphism in our collection. Relationships between isolates could be established in only a few instances. The bacteria with two novel β-lactamases were distributed into two particular ribotypes. No other isolates were found in these ribotypes (Fig. 2). Clinically, the patients with a K. pneumoniae SHV-25 carrier (a patient had underlying cirrhosis of the liver with ascites) and a K. pneumoniae SHV-26 carrier (a patient who was bedridden due to an old cerebral infarction and diabetes mellitus) were under treatment with gentamicin and/or cefazolin. The patient infected with the SHV-26 carrier died shortly after the treatment was begun (data not shown). The discovery of SHV-26, a β-lactamase with reduced susceptibility to clavulanic acid that renders bacteria intermediately resistant to an inhibitor consisting of a β-lactam, may augur the development of β-lactamase inhibitor resistance apart from ESBL inhibitor resistance.
Genetically, comparison of the sequence of blaSHV-25 with that of blaSHV-1 revealed three different mutations in SHV-25, at positions 18, 35, and 129 (Jacoby and Bush, http://www.lahey.org./studies/webt/htm). The mutation at position 35 has been described in SHV-11 and has no influence on substrate and inhibitor profiles, the pI value, or the MIC (6). A previous study has shown that the side chain of β-lactam antibiotics is connected to Asn-132 via a hydrogen bond and therefore stabilizes the catalytically important conserved Ser-Asp-Asn (SDN) loop from positions 130 to 132 (5). Thus, the modification at position 129, which is closer to SDN, may result in a lowering of the Km and a change of the pI to 7.5. The influence of the modification at position 18 of the signal peptide in leading the β-lactamase to the periplasm, where it resides, remains unclear. However, our kinetics data showed that relatively low Km values have been detected for cephalothin and cephaloridine. Theoretically, the MIC will be raised when a low Km is achieved (6). The MICs obtained in the present study indicate that the cephalothin MIC for isolates with SHV-25 is similar to those for isolates with SHV-1. A previous study showed that a nucleotide substitution, I8F, in the signal sequence in SHV-7 led to slightly increased cephalosporin MICs, suggesting the more efficient transfer of the enzyme precursor into the periplasmic space. Whether the presence of a modification at position 18 in SHV-25, which is within the signal sequence, leads the β-lactamase to a location other than periplasm is unknown. Further confirmation by site-directed mutagenesis should be conducted. For SHV-26, the threefold increase in the clavulanic acid IC50 compared to those for SHV-1 and SHV-25 resulted in a higher amoxicillin-clavulanate MIC for SHV-26. Mutation at position 187 may involve the interaction with clavulanic acid.
In conclusion, we have described two novel β-lactamases, one of which is a non-ESBL and one of which is an enzyme from an isolate with reduced susceptibility to clavulanic acid. The prevalence of these enzymes is rare in northern Taiwan. Although the results do not provide information on the mutation rate, the results obtained in the present study suggest the routes of evolution of SHV β-lactamase genes among isolates in northern Taiwan. Antibiotic selective pressure may explain the development of ESBL-type resistance as well as inhibitor-resistant β-lactam resistance.
ACKNOWLEDGMENTS
This work was sponsored by grants from the National Science Council (grants NSC89-2314-B-016-032 and NSC89-2314-B-016-014) and, partly, by the National Health Research Institute of Taiwan.
REFERENCES
- 1.Babini G S, Livermore D M. Are SHV beta-lactamases universal in Klebsiella pneumoniae? Antimicrob Agents Chemother. 2000;44:2230. doi: 10.1128/aac.44.8.2230-2230.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Blahova J, Kralikova K, Krcmery V, Sr, Schafer V. Extended-spectrum beta-lactamase-producing strains of Enterobacter cloacae transferring resistance to cefotaxime and ceftazidime. J Chemother. 1999;11:97–102. doi: 10.1179/joc.1999.11.2.97. [DOI] [PubMed] [Google Scholar]
- 3.Brisse S, Milatovic D, Fluit A C, Kusters K, Toelstra A, Verhoef J, Schmitz F J. Molecular surveillance of European quinolone-resistant clinical isolates of Pseudomonas aeruginosa and Acinetobacter spp. using automated ribotyping. J Clin Microbiol. 2000;38:3636–3645. doi: 10.1128/jcm.38.10.3636-3645.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bush K, Sykes R B. Methodology for the study of beta-lactamases. Antimicrob Agents Chemother. 1986;30:6–10. doi: 10.1128/aac.30.1.6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Knox J R. Extended-spectrum and inhibitor-resistant TEM-type beta-lactamases: mutations, specificity, and three-dimensional structure. Antimicrob Agents Chemother. 1995;39:2593–2601. doi: 10.1128/aac.39.12.2593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Livermore D M. β-Lactamases in laboratory and clinical resistance. Clin Microbiol Rev. 1995;8:557–584. doi: 10.1128/cmr.8.4.557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mabilat C, Goussard S. PCR detection and identification of genes for extended-spectrum β-lactamases. In: Persing D H, Smith T F, Tenover F C, White T J, editors. Diagnostic molecular microbiology: principles and applications. Washington, D.C.: American Society for Microbiology; 1993. pp. 553–563. [Google Scholar]
- 8.Marchandin H, Carriere C, Sirot D, Pierre H J, Darbas H. TEM-24 produced by four different species of Enterobacteriaceae, including Providencia rettgeri, in a single patient. Antimicrob Agents Chemother. 1999;43:2069–2073. doi: 10.1128/aac.43.8.2069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Martinez J L, Baquero F. Mutation frequencies and antibiotic resistance. Antimicrob Agents Chemother. 2000;44:1771–1777. doi: 10.1128/aac.44.7.1771-1777.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Martinez-Martinez L, Pascual A, Hernandez-Alles S, Alvarez-Diaz D, Suarez A I, Tran J, Benedi V J, Jacoby G A. Roles of beta-lactamases and porins in activities of carbapenems and cephalosporins against Klebsiella pneumoniae. Antimicrob Agents Chemother. 1999;43:1669–1673. doi: 10.1128/aac.43.7.1669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Matthew M, Harris A M. Identification of beta-lactamases by analytical isoelectric focusing: correlation with bacterial taxonomy. J Gen Microbiol. 1976;94:55–67. doi: 10.1099/00221287-94-1-55. [DOI] [PubMed] [Google Scholar]
- 12.National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial disk susceptibility tests. Approved standard M2–A6. Wayne, Pa: National Committee for Clinical Laboratory Standards; 1997. [Google Scholar]
- 13.National Committee for Clinical Laboratory Standards. Performance standards for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. Approved standard M7–A4. Wayne, Pa: National Committee for Clinical Laboratory Standards; 1997. [Google Scholar]
- 14.Rasheed J K, Jay C, Metchock B, Berkowitz F, Weigel L, Crellin J, Steward C, Hill B, Medeiros A A, Tenover F C. Evolution of extended-spectrum beta-lactam resistance (SHV-8) in a strain of Escherichia coli during multiple episodes of bacteremia. Antimicrob Agents Chemother. 1997;41:647–653. doi: 10.1128/aac.41.3.647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sanger F, Nicklen S, Coulson A R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA. 1977;74:5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Silva J, Aguilar C, Becerra Z, Lopez-Antunano F, Garcia R. Extended-spectrum beta-lactamases in clinical isolates of enterobacteria in Mexico. Microb Drug Resist. 1999;5:189–193. doi: 10.1089/mdr.1999.5.189. [DOI] [PubMed] [Google Scholar]
- 17.Siu L K, Lu P L, Hsueh P R, Lin F M, Chang S C, Luh K T, Ho M, Lee C Y. Bacteremia due to extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae in a pediatric oncology ward: clinical features and identification of different plasmids carrying both SHV-5 and TEM-1 genes. J Clin Microbiol. 1999;37:4020–4027. doi: 10.1128/jcm.37.12.4020-4027.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Stobberingh E E, Arends J, Hoogkamp-Korstanje J A, Goessens W H, Visser M R, Buiting A G, Debets-Ossenkopp Y J, van Ketel R J, van Ogtrop M L, Sabbe L J, Voorn G P, Winter H L, van Zeijl J H. Occurrence of extended-spectrum beta-lactamases (ESBL) in Dutch hospitals. Infection. 1999;27:348–354. doi: 10.1007/s150100050041. [DOI] [PubMed] [Google Scholar]