Abstract
Molecular mechanisms of azole resistance in Candida albicans, including alterations in the target enzyme and increased efflux of drug, have been described, but the epidemiology of the resistance mechanisms has not been established. We have investigated the molecular mechanisms of resistance to azoles in C. albicans strains displaying high-level fluconazole resistance (MICs, ≥64 μg/ml) isolated from human immunodeficiency virus (HIV)-infected patients with oropharyngeal candidiasis. The levels of expression of genes encoding lanosterol 14α-demethylase (ERG11) and efflux transporters (MDR1 and CDR) implicated in azole resistance were monitored in matched sets of susceptible and resistant isolates. In addition, ERG11 genes were amplified by PCR, and their nucleotide sequences were determined in order to detect point mutations with a possible effect in the affinity for azoles. The analysis confirmed the multifactorial nature of azole resistance and the prevalence of these mechanisms of resistance in C. albicans clinical isolates exhibiting frank fluconazole resistance, with a predominance of overexpression of genes encoding efflux pumps, detected in 85% of all resistant isolates, being found. Alterations in the target enzyme, including functional amino acid substitutions and overexpression of the gene that encodes the enzyme, were detected in 65 and 35% of the isolates, respectively. Overall, multiple mechanisms of resistance were combined in 75% of the isolates displaying high-level fluconazole resistance. These results may help in the development of new strategies to overcome the problem of resistance as well as new treatments for this condition.
In Candida albicans fluconazole resistance is a multifactorial process mediated through multiple underlying mechanisms (4, 7, 8, 18, 34, 36, 42). Resistance can be the result of an alteration of the target enzyme, the cytochrome P-450 lanosterol 14α-demethylase (Erg11p), either by overexpression or as a result of point mutations in the gene that encodes it (ERG11) (3, 6, 7, 12, 14, 15, 16, 20, 33, 37, 41). The former creates the need for a higher intracellular azole concentration to complex all the enzyme molecules present in the cells, and the latter leads to amino acid substitutions, resulting in a decreased affinity for azole derivatives. A second major mechanism is failure of azole antifungal agents to accumulate inside the yeast cell as a consequence of enhanced drug efflux. This mechanism is mediated by two types of multidrug efflux transporters, the major facilitators (encoded by multidrug resistance genes) and those belonging to the ATP-binding cassette superfamily (ABC transporters, encoded by CDR genes). Upregulation of the CDR genes appears to confer resistance to multiple azoles, whereas upregulation of the MDR1 gene alone leads to fluconazole resistance exclusively (1, 18, 19, 22, 26, 30–32, 38–40).
These different molecular mechanisms implicated in the development of resistance to fluconazole have previously been described for a limited number of isolates by us and others by analyzing serial isolates from the same patient with decreasing susceptibility to the drug (18, 30, 40, 41; D. C. Calabrese, J. Bille, and D. Sanglard, Abstr. 5th Int. Meeting Candida Candidiasis, abstr. C55, p. 63, 1999). However, the relative prevalence of these mechanisms in clinical isolates displaying high-level fluconazole resistance is not known. In the study described here, we have evaluated the molecular mechanisms responsible for azole resistance in 20 C. albicans clinical isolates displaying high-level fluconazole resistance (MICs, ≥64 μg/ml) obtained from 12 different human immunodeficiency virus (HIV)-infected patients with oropharyngeal candidiasis (OPC).
(This work was partially presented at the 39th Interscience Conference on Antimicrobial Agents and Chemotherapy, San Francisco, Calif., 26 to 29 September 1999.)
MATERIALS AND METHODS
Isolates.
Clinical samples were obtained from HIV-infected patients enrolled in a prospective clinical study of OPC at the University of Texas Health Science Center at San Antonio and the Audie L. Murphy Division, South Texas Veterans Health Care System, San Antonio (27, 28). At the time of initial isolation, oral samples were plated on RPMI and CHROMagar Candida (CHROMagar Company, Paris, France) media with fluconazole (8 and 16 μg/ml) and without fluconazole to maximize detection of resistant yeasts, as described previously (24, 25). The identities of the clinical isolates were confirmed by standard biochemical and microbiological procedures, including assessment of carbohydrate assimilation patterns (API 20 C; Biomerieux, Marcy l'Etoile, France), germ tube formation in serum-containing medium, and the colors of the colonies in chromogenic medium (CHROMagar Candida). Initial fluconazole susceptibility testing was performed by an NCCLS methodology, and C. albicans isolates were considered resistant if the fluconazole MIC was ≥64 μg/ml (23, 29). Isolates were stored at room temperature as suspensions in sterile water and were subcultured onto plates containing Sabouraud dextrose agar 48 h prior to propagation in YEPD medium (2% yeast extract, 1% peptone, 2% glucose).
DNA-typing techniques for strain identification.
Strain identity was established by karyotyping, restriction fragment length polymorphism analysis, and fingerprinting analysis with the moderately repetitive Ca3 probe, as described before (17–19). The resulting banding patterns were analyzed visually and by using computer-assisted methods (Dendrom; Solltech Inc., Oakdale, Iowa) (35).
Antifungal drug susceptibility testing.
Testing of susceptibility to fluconazole (Pfizer Inc., Sandwich, United Kingdom), itraconazole (Janssen Pharmaceutica, Beerse, Belgium), voriconazole (Pfizer Inc.), posaconazole (SCH56592; Schering Plough, Kenilworth, N.J.), and amphotericin B (Bristol-Myers Squibb, Princeton, N.J.) was performed by an NCCLS methodology by a broth microdilution method (23, 29).
Northern blot analysis.
The different isolates were propagated in YEPD medium and harvested while growing in antifungal drug-free medium at the logarithmic phase at an approximate cell density of 7.5 × 107 cells/ml. Total RNA was obtained with the RNAeasy mini kit (Qiagen Inc., Valencia, Calif.) following the manufacturer's instructions. Equal amounts (approximately 5 μg) of RNA, as determined by A260 measurements, were separated by electrophoresis and subsequently transferred to nylon membranes (Nytran; Schleicher & Schuell, Keene, N.H.). Probes specific for the ERG11, MDR1, and CDR genes were purified from plasmids containing inserts of the respective genes, as described before (18). Probes specific for the CDR1 and CDR2 genes were prepared as described by Sanglard and colleagues by PCR amplification from plasmids containing these sequences (18, 32). All probes were labeled by random priming (Random Primers DNA Labeling System; Gibco-BRL, Gaithersburg, Md.), and hybridizations were performed with Rapid-hyb buffer (Amersham Life Science Inc., Arlington Heights, Ill.). After hybridization, the blots were washed by using high-stringency conditions and were exposed to autoradiography film (Kodak, Rochester, N.Y.) overnight at room temperature. The nylon membranes were probed sequentially with the different probes following stripping of the previously bound probe. For densitometric analysis, autoradiograms were scanned with the Adobe Photoshop program (Adobe Systems Inc., Mountain View, Calif.), and the signals were quantified with Dendron software (Solltech Inc.). Relative values were adjusted for differences in sample loading on the basis of quantification of 18S rRNA levels. A twofold increase in the densitometric values compared to the values obtained for the corresponding matched susceptible isolate was arbitrarily considered significant (upregulation).
PCR amplification and sequencing.
The ERG11 genes encoding lanosterol l4α-demethylase from all isolates were amplified by PCR. Briefly, genomic DNA was extracted with YeaStar Genomic DNA (Zymo Research, Orange, Calif.) and was used as a template for amplification of ERG11 genes. PCR was carried out with high-fidelity Pwo DNA polymerase (Boehringer Mannheim, GmbH, Mannheim, Germany) with the following primers: 5′-GTT GAA ACT GTC ATT GAT GG (forward) and 5′-TCA GAA CAC TGA ATC GAA AG (reverse). Amplicons were purified with a QIAquick PCR purification kit (Qiagen Inc., Valencia, Calif.), and the nucleotide sequences for both strands were determined by primer elongation with an automated DNA sequencer (Applied Biosystems, Foster City, Calif.). Sequence data were compared to a published ERG11 sequence by using the BLAST program (2, 13).
Functional expression of C. albicans PCR-amplified ERG11 alleles in S. cerevisiae
Saccharomyces cerevisiae YKKB-13 (MATα ura3-52 lys2-801amber ade-101ochreI trp1-Δ63his3-Δ200 leu2-Δ1Δpdr5::TRP1), which is defective in the ATP-binding cassette transporter and which is therefore hypersusceptible to azole derivatives, was used for the expression of C. albicans ERG11 genes in YEp51 plasmids. YEp51 is a 2μm-based vector that contains a GAL10 promoter for inducible heterologous gene expression. S. cerevisiae YKKB-13 cannot grow on galactose, which is required for GAL10 induction, but it can grow on raffinose; therefore, both carbon sources were added to the same medium to ensure the simultaneous occurrence of growth of S. cerevisiae and induction of the GAL10 promoter. For the cloning of ERG11 genes from C. albicans isolates, the strategy developed by Sanglard et al. was followed (33). Briefly, the ERG11 genes were cloned from the genomic DNAs of the C. albicans isolates by PCR as described above. DNA was first extracted and used as a template for amplification of ERG11 alleles. PCR was carried out with high-fidelity Pwo DNA polymerase (Boehringer Mannheim) by using primers that span the entire ERG11 open reading frame flanked with BamHI and SalI restriction sites to allow the subcloning of amplified ERG11 fragments into YEp51 precut by the same enzymes (33). For each PCR with genomic DNA of a C. albicans isolate, at least 10 ERG11 expression plasmids were obtained. Plasmids were then transformed into S. cerevisiae YKKB-13 by a lithium acetate method in noninducing yeast nitrogen broth (YNB) medium with glucose as a carbon source. The expression of Erg11p was verified by growth of the Leu+ transformant in inducing YNB selective medium with galactose and raffinose as carbon sources. Then, disk diffusion assays with fluconazole were performed with S. cerevisiae transformants in raffinose-galactose YNB selective medium. The following considerations were taken into account when comparing the diameters of matched susceptible-resistant isolates with the controls: if in the disk assays the diameters between the isogenic susceptible and resistant isolates are similar between the isolates as well similar to that for the susceptible control strain, the mutation(s) present in the ERG11 genes from azole-resistant or azole-susceptible isolates do not alter the affinity of the target to fluconazole. If, on the contrary, the diameters are dissimilar compared to that for the susceptible control strain, the mutations found in both susceptible and resistant isolates play a role in the affinity of Erg11p for the azoles. When the diameters between the isogenic susceptible and resistant isolates are not identical in the disk assays, mutations in the ERG11 genes from azole-resistant isolates can be expected, and these could result in a difference of affinity of the target to azoles. The nucleotide sequences of the cloned C. albicans ERG11 alleles of interest were determined as described above. The mean disk diameters among resistant and susceptible isolates and a susceptible control were compared by a one-way analysis of variance. Differences were considered statistically significant when the P value was less than 0.05. The analyses were performed with SPSS software (version 6.12; SPSS, Chicago, Ill.).
RESULTS
Strain identification.
Since evaluation of molecular mechanisms of resistance requires the use of matched sets of susceptible and resistant isolates, DNA-typing techniques were used to assess strain isogenicity among a total of 20 highly resistant isolates (fluconazole MICs, ≥64 μg/ml) selected for analysis and their corresponding susceptible isolates recovered from 12 different HIV-infected patients. The high degree of relatedness among susceptible and resistant isolates obtained from the same patient was confirmed by all typing methods used (karyotyping, restriction fragment length polymorphism analysis, and Ca3 probe-based fingerprinting). Thus, susceptible and resistant isolates obtained from the same patient represented the same strain. Also, these experiments revealed that different patients harbored different C. albicans strains (data not shown).
Antifungal susceptibility testing.
The MICs of fluconazole, itraconazole, voriconazole, posaconazole, and amphotericin B for the different isolates are summarized in Table 1. Except for the isolate from patient 51, the initial isolate of the series for each patient was fluconazole susceptible (fluconazole MICs, ≤8 μg/ml); for the isolates from patient 51, the fluconazole MIC for the most susceptible isogenic isolate that was found was 16 μg/ml (susceptible, dose dependent). For each patient, the resistant isolates included in this study were selected on the basis of their high-level fluconazole resistance (fluconazole MICs, ≥64 μg/ml) and also after determination of their isogenicities with the corresponding susceptible isolates obtained from the same patient. By following the criteria established by Rex et al. (29) for the interpretive breakpoints for antifungal susceptibility testing for fluconazole and itraconazole against C. albicans, all the fluconazole-resistant isolates remained susceptible to itraconazole (itraconazole MICs, ≤1 μg/ml), although increases in the MICs were also observed compared to those for the susceptible isolates. Decreased susceptibility to itraconazole was detected in fluconazole-resistant isolates from patients 7, 9, 30, 43, 51, 59, and 64, with a 1 to 5 twofold dilution increase in the itraconazole MICs. In the case of voriconazole, elevated MICs were detected for fluconazole-resistant isolates from patients 7, 9, 14, 30, 42, 43, 51, 59, and 64, with up to a 6 twofold dilution increase in resistance. In the case of posaconazole, decreased susceptibility was also noted in fluconazole-resistant isolates from patients 7, 15, 43, 51, and 64, with up to 1 to 5 twofold dilution increases in resistance. Isolates with decreased susceptibilities to all four azole derivatives tested were detected in patients 7, 42, 43, 51, and 64. Considering the isolates that presented large decreases in their susceptibilities to the new azoles (isolates 2307, 3731, 2257, and 4380, for which MICs were 3 or more twofold dilutions higher than those for their susceptible counterparts), we could observe that the most frequent mechanism of resistance in these isolates was the overexpression of efflux pumps (predominantly, CDR-encoded pumps; see below). The differences in the amphotericin B MICs for the susceptible and the resistant isolates were small (in all cases they were within 1 twofold dilution), and all isolates remained susceptible to this agent.
TABLE 1.
Antifungal susceptibilities of C. albicans isolates
| Patient no. | Isolate no. | MIC (μg/ml) at 24 h/48 h
|
||||
|---|---|---|---|---|---|---|
| Fluconazole | Itraconazole | Voriconazole | Posaconazole | AmB | ||
| 7 | 412 | <0.125/0.5 | 0.015/0.015 | 0.015/0.03 | 0.015/0.015 | 0.06/0.125 |
| 2307 | 16/>64 | 0.25/0.5 | 0.5/0.5 | 0.25/0.25 | 0.25/0.25 | |
| 9 | 1002 | 0.25/0.25 | 0.015/0.015 | 0.03/0.03 | 0.015/0.015 | 0.06/0.125 |
| 2823 | >64/>64 | <0.015/0.06 | 0.125/0.125 | 0.03/0.06 | 0.125/0.25 | |
| 3795 | >64/>64 | 0.015/0.25 | 0.25/1 | <0.015/0.06 | 0.125/0.5 | |
| 14 | 580 | 1/4 | 0.06/0.06 | 0.03/0.03 | 0.015/0.015 | 0.06/0.125 |
| 2440 | 32/64 | 0.125/0.125 | 0.125/0.25 | 0.06/0.06 | 0.125/0.25 | |
| 2500 | 32/64 | <0.015/<0.015 | 0.125/0.25 | <0.015/0.03 | 0.125/0.25 | |
| 15 | 945 | 4/8 | 0.125/0.25 | 0.125/0.25 | 0.03/0.03 | 0.06/0.125 |
| 1619 | 32/64 | 0.125/0.25 | 0.25/0.5 | 0.125/0.125 | 0.125/0.25 | |
| 16 | 3107 | 2/4 | 0.03/0.03 | 0.125/0.5 | 0.03/0.03 | 0.06/0.125 |
| 3119 | 64/>64 | <0.015/<0.015 | 0.5/0.5 | <0.015/<0.015 | 0.125/0.25 | |
| 3120 | 32/>64 | 0.015/0.015 | 0.25/0.5 | <0.015/<0.015 | 0.125/0.25 | |
| 3184 | 64/64 | 0.06/0.06 | 1/1 | 0.06/0.06 | 0.25/0.5 | |
| 3281 | 32/64 | 0.06/0.06 | 0.5/1 | 0.03/0.06 | 0.125/0.5 | |
| 28 | 5044 | 4/4 | 0.125/0.125 | 0.125/0.5 | 0.06/0.06 | 0.06/0.125 |
| 5052 | 32/64 | 0.06/0.125 | 0.5/1 | 0.06/0.06 | 0.125/0.25 | |
| 30 | 5106 | 4/8 | 0.03/0.06 | 0.06/0.06 | 0.015/0.25 | 0.06/0.125 |
| 5108 | 32/64 | 0.06/0.25 | 0.5/0.5 | 0.06/0.125 | 0.125/0.5 | |
| 42 | 1691 | 0.25/0.25 | 0.015/0.015 | 0.03/0.03 | 0.015/0.015 | 0.06/0.125 |
| 3731 | >64/>64 | <0.015/0.125 | 0.25/0.5 | 0.06/0.125 | 0.125/0.25 | |
| 3733 | 64/64 | <0.015/0.125 | 0.25/0.5 | <0.015/0.03 | 0.125/0.25 | |
| 43 | 1649 | 0.25/0.5 | 0.015/0.015 | 0.03/0.03 | 0.015/0.015 | 0.06/0.125 |
| 3034 | >64/>64 | 0.5/0.5 | 0.5/2 | 0.25/0.5 | 0.25/0.5 | |
| 51 | 2274 | 16/16 | 0.06/0.06 | 0.125/0.125 | 0.015/0.03 | 0.125/0.25 |
| 2257 | 32/64 | 0.125/0.25 | 1/1 | 0.125/0.25 | 0.125/0.25 | |
| 2339 | 16/64 | 0.125/0.125 | 0.5/1 | 0.125/0.5 | 0.25/0.25 | |
| 59 | 3917 | 2/4 | 0.06/0.125 | 0.125/0.125 | 0.03/0.06 | 0.125/0.5 |
| 4617 | 32/64 | 0.125/0.25 | 0.25/1 | 0.125/0.125 | 0.125/0.25 | |
| 4639 | 64/>64 | 0.25/0.25 | 0.5/1 | 0.125/0.125 | 0.125/0.25 | |
| 64 | 4018 | 1/4 | 0.06/0.06 | 0.125/0.125 | 0.015/0.015 | 0.06/0.125 |
| 4380 | 32/64 | 0.06/0.25 | 0.5/1 | 0.06/0.125 | 0.125/0.25 | |
Levels of expression of ERG11, MDR1, and CDR genes in matched sets of susceptible and resistant C. albicans clinical isolates.
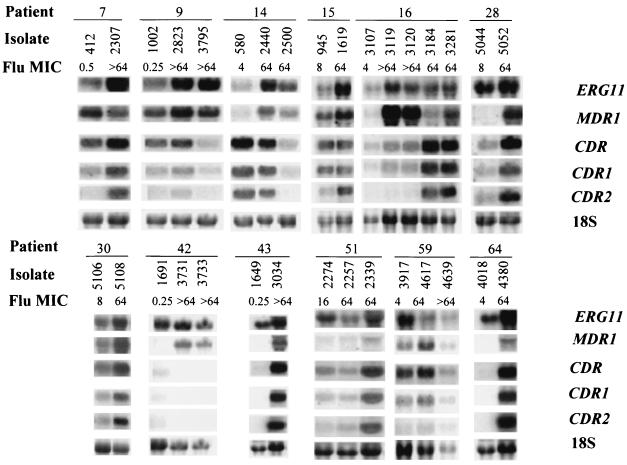
Total RNA extracted from the different isolates growing in YEPD medium in the absence of an antifungal drug was analyzed by a Northern blot technique with probes specific for the ERG11, MDR1, CDR1, and CDR2 genes and a probe that detects different members of the CDR gene family (18, 19, 22, 40). As shown in Fig. 1, overexpression of CDR genes was detected in 11 isolates (55%) from 10 patients (83%). In most instances, concomitant overexpression of CDR1 and CDR2 was observed. Upregulation of MDR1 was also observed in a total of 11 isolates (55%) from eight patients (67%). Upregulation of ERG11 genes was detected in seven isolates (35%) from five patients (42%).
FIG. 1.
Northern blots of total RNA from clinical C. albicans isolates analyzed with radiolabeled probes specific for ERG11, MDR1, CDR, CDR1, and CDR2. Hybridizations were performed as described in Materials and Methods. The bottom rows show the amounts of 18S rRNA used to standardize and calibrate signal levels according to lane loading parameters. Flu, fluconazole.
Mutations in ERG11 genes from C. albicans isolates resistant to azole antifungal agents.
ERG11 genes obtained from the genomic DNAs of all C. albicans isolates were amplified by PCR with high-fidelity Pwo DNA polymerase. Fragments of the expected length (1.6 kb) were obtained in each case. In order to identify the point mutations present in the ERG11 genes of the resistant isolates, we obtained their sequences. All the sequences contained at least one cryptic nucleotide variation compared to the published sequence of ERG11 (13; data not shown). No variation that led to an amino acid substitution was found in three resistant isolates. In addition, all ERG11 genes from the other 17 azole-resistant isolates contained one or more nucleotide variations that led to amino acid substitutions in the protein sequence. Point mutations in ERG11 genes that resulted in 13 different amino acid substitutions were detected (Table 2). To demonstrate that the identified amino acid substitutions in ERG11 from fluconazole-resistant strains could confer resistance to antifungal drugs in an intact yeast cell, the ERG11 genes from fluconazole-resistant and matched susceptible isolates were expressed in S. cerevisiae. Disk diffusion assays with fluconazole were performed with 10 S. cerevisiae transformants obtained from each C. albicans clinical isolate. Each transformant was subjected to a fluconazole disk diffusion assay on raffinose- and galactose-containing YNB agar. Diameters of inhibition were recorded for each transformant, and the results are presented in Table 3. The decrease in the diameter of inhibition reflects the fact that alterations in ERG11 proteins, which translate into a lower level of susceptibility, had occurred. In two cases (isolate 2307 from patient 7 and isolate 2500 from patient 14), no differences in diameters of inhibition were observed compared to those for the isogenic susceptible strain as well as fluconazole-susceptible S. cerevisiae transformant YKKB-13 used as a control. Two different point mutations that led to amino acid substitutions K128T and V437I were found in these isolates. In the case of the K128T amino acid substitution, it appeared to be present in both the susceptible and the resistant isolate. Neither amino acid substitution (K128T or V437I) altered the affinity of Erg11p for fluconazole, and therefore, these substitutions are not associated with azole resistance. Other investigators have previously indicated that neither point mutation is linked to the azole antifungal agent resistance phenotype. In one case (isolate 5108 from patient 30), a decrease in the diameter of inhibition was observed in fluconazole-resistant transformant YKKB-13 compared to that for the fluconazole-susceptible transformant. The G464S substitution not present in the susceptible transformant was detected. In other cases (isolates 3107 and 3119 from patient 16; isolates 2274, 2257, and 2339 from patient 51; and isolates 5044 and 5052 from patient 28), no differences in diameters of inhibition were recorded for yeasts expressing ERG11 genes compared with those for the fluconazole-susceptible and -resistant isolates from a given patient, although a decrease in the diameter of inhibition was observed compared to that for fluconazole-susceptible S. cerevisiae YKKB-13 transformed with a susceptible control. In these isolates, three amino acid substitutions linked to a phenotype of less susceptibility were found: Y132F, S405F, and D446N. In two other cases (isolate 1619 from patient 15 and isolate 3731 from patient 42), two distinct diameters of inhibition were measured for YKKB-13 transformants expressing the ERG11 alleles. This fact is consistent with the diploidy of C. albicans; each allele of the genomic ERG11 loci of these two isolates was amplified by PCR, and one of these alleles encodes an altered protein that decreases the susceptibility of the S. cerevisiae strain expressing the corresponding allele. In these isolates, five amino acid substitutions were found: D116E, G450E, G307S, F126L, and K143R. For three other isolates (isolates 3917, 4617, and 4639 from patient 59), two distinct diameters of inhibition were measured for YKKB-13 expressing the ERG11 alleles from the susceptible isolate (in one case, no mutation was observed and the other transformant presented the F449S amino acid substitution). In the case of the resistant isolates, two substitutions were detected: F449S and T229A. Globally, 11 amino acid substitutions were found to be associated with a resistance phenotype: D116E, G450E, G307S, Y132F, D446N, G464S, F126L, K143R, S405F, F449S, and T229A. Of these, G307S and D446N have not been yet reported by other laboratories (3, 6, 11, 12, 16, 20, 33). On the other hand, two amino acid substitutions, K128T and V437I, were confirmed to not participate in azole resistance, in agreement with previous reports (6, 33).
TABLE 2.
Nucleotide and amino acid substitutions in ERG11 genes from C. albicans isolates
| Patient | Isolate | Fluconazole MIC (μg/ml) | Substitutiona
|
||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D116E | F126L | K128T | Y132F | K143R | T229A | G307S | S405F | V437I | D446N | F449S | G450E | G464S | |||
| 7 | 412 | 0.5 | A383C | ||||||||||||
| 2307 | >64 | A383C | |||||||||||||
| 9 | 1002 | 0.25 | |||||||||||||
| 2823 | >64 | ||||||||||||||
| 3795 | >64 | T376C | A428G | ||||||||||||
| 14 | 580 | 4 | |||||||||||||
| 2440 | 64 | G1309A | |||||||||||||
| 2500 | 64 | G1309A | |||||||||||||
| 15 | 945 | 8 | T348A | G1349A | |||||||||||
| 1619 | 64 | T348A | G919A | G1349A | |||||||||||
| 16 | 3107 | 4 | A395T | ||||||||||||
| 3119 | >64 | A395T | |||||||||||||
| 3120 | >64 | A395T | |||||||||||||
| 3184 | 64 | A395T | |||||||||||||
| 3281 | 64 | A395T | |||||||||||||
| 28 | 5044 | 4 | G1309A | G1336A | |||||||||||
| 5052 | 64 | G1309A | G1336A | ||||||||||||
| 30 | 5106 | 8 | G1309A | ||||||||||||
| 5108 | 64 | G1309A | G1390A | ||||||||||||
| 42 | 1691 | 0.25 | A383C | ||||||||||||
| 3731 | >64 | T376C | A428G | ||||||||||||
| 3733 | 64 | T376C | A428G | ||||||||||||
| 43 | 1649 | 0.5 | |||||||||||||
| 3034 | >64 | ||||||||||||||
| 51 | 2274 | 16 | C1214T | ||||||||||||
| 2257 | 64 | C1214T | |||||||||||||
| 2339 | 64 | C1214T | |||||||||||||
| 59 | 3917 | 4 | T1346C | ||||||||||||
| 4617 | 64 | A685G | T1346C | ||||||||||||
| 4639 | >64 | A685G | T1346C | ||||||||||||
| 64 | 4018 | 4 | |||||||||||||
| 4380 | 64 | G1309A | |||||||||||||
The base numbers are with respect to the first ATG codon of ERG11.
TABLE 3.
Fluconazole susceptibilities of S. cerevisiae strains expressing C. albicans ERG11 genes, as determined by zone of inhibition
| Patient no. | C. albicans isolatea | Mean ± SD zone of inhibition (mm)b | ERG11 allele expressed | Amino acid substitution(s) |
|---|---|---|---|---|
| 7 | 412 (S) | 43.75 ± 1.12 | 412 | K128T |
| 2307 (R) | 43.08 ± 0.90 | 2307 | K128T | |
| 14 | 1632 (S) | 42.80 ± 1.93 | 580 | None |
| 1640 (R) | 44.18 ± 0.75 | 1640 | V437I | |
| 15 | 945 (S) | 32.50 ± 0.70c | 945 | D116E, G450E |
| 1619 (R) | 0c | 1619-1 | D116E, G450E, G307S | |
| 32.80 ± 1.35c | 1619-2 | D116E, G450E | ||
| 16 | 3107 (S) | 14.50 ± 1.24c | 3107 | Y132F |
| 3119 (R) | 15.57 ± 3.94c | 3119 | Y132F | |
| 28 | 5044 (S) | 36.81 ± 0.91c | 5044 | V437I, D446N |
| 5052 (R) | 37.56 ± 2.00c | 5052 | V437I, D446N | |
| 30 | 5106 (S) | 43.37 ± 1.50 | 5106 | V437I |
| 5108 (R) | 30.58 ± 0.79c | 5108 | V437I, G464S | |
| 42 | 1691 (S) | 42.87 ± 0.83 | 1691 | K128T |
| 3731 (R) | 0c | 3731-1 | F126L, K143R | |
| 22.33 ± 0.52c | 3731-2 | K143R | ||
| 51 | 2274 (S) | 31.55 ± 1.85c | 2274 | S405F |
| 2257 (R) | 33.16 ± 1.83c | 2257 | S405F | |
| 2339 (R) | 33.07 ± 1.64c | 2339 | S405F | |
| 59 | 3917 (S) | 42.62 ± 1.50 | 3917-1 | None |
| 32.62 ± 1.19c | 3917-2 | F449S | ||
| 4617 (R) | 21.05 ± 0.87c | 4617 | F449S, T229A | |
| 4639 (R) | 24.37 ± 1.60c | 4639 | F449S, T229A |
S, susceptible; R, resistant.
Mean diameters were obtained by measuring the zones of inhibition produced by 10 independent S. cerevisiae Leu+ transformants. The mean diameter for YKKB-13 transformed with YEp51 alone was 63 mm. The mean diameter for YKKB-13 transformed with a susceptible control was 43 mm. The mean diameter for YKKB-13 transformed with a resistant control was 0 mm.
P < 0.05.
Multifactorial nature of resistance: combinations of different molecular mechanisms are responsible for azole resistance in a majority of isolates displaying high-level fluconazole resistance.
In five isolates (25%) from five patients (41.6%), concomitant upregulation of the CDR and multidrug resistance genes was noted. In three isolates (15%) from two patients (10%), upregulation of ERG11 appeared to be associated with upregulation of MDR1; and in three isolates (15%) from two patients, (10%) they were detected together with CDR gene upregulation. Point mutations in ERG11 genes with an effect on the affinity of the enzyme for the azoles were observed in 13 isolates (65%) from seven patients (58.3%). In two isolates (10%) from two patients (16.6%), ERG11 upregulation was detected simultaneously with point mutations in their ERG11 genes. In 11 isolates (55%), point mutations in ERG11 genes appeared to be combined with the upregulation of efflux pumps; more precisely, in 7 isolates it appeared to be associated with upregulation of MDR1 genes and in another 7 isolates it appeared to be associated with upregulation of CDR genes, with both efflux pumps combined appearing in 3 isolates. See Table 4 for a compendium of the amino acid substitutions and gene overexpression in each of the isolates displaying high-level fluconazole resistance compared to matched susceptible isolates.
TABLE 4.
Summary of amino acid substitutions in Erg11p and gene overexpression in fluconazole-resistant C. albicans isolates compared to matched susceptible isolates
| Patient | Isolate | Fluconazole MIC (μg/ml) | Amino acid substitutions in Erg11p | Gene overexpression |
|---|---|---|---|---|
| 7 | 412 | 0.5 | K128T | |
| 2307 | >64 | K128T | ERG11, CDR, CDR1, and CDR2 | |
| 9 | 1002 | 0.25 | ||
| 2823 | >64 | ERG11, MDR1, and CDR2 ERG11 | ||
| 3795 | >64 | |||
| 14 | 580 | 4 | ||
| 2440 | 64 | V437I | V437I, ERG11, and MDR1 | |
| 2500 | 64 | V437I | V437I, ERG11, and MDR1 | |
| 15 | 945 | 8 | D116E and G450E | |
| 1619 | 64 | D116E and G450E | ERG11 and CDR2 | |
| 16 | 3107 | 4 | Y132F | |
| 3119 | >64 | Y132F | MDR1 | |
| 3120 | >64 | Y132F | MDR1 | |
| 3184 | 64 | Y132F | CDR | |
| 3281 | 64 | Y132F | CDR | |
| 28 | 5044 | 4 | V437I and D446N | |
| 5052 | 64 | V437I and D446N | MDR1, CDR, CDR1, and CDR2 | |
| 30 | 5106 | 8 | V437I | |
| 5108 | 64 | V437I and G464S | ERG11, MDRI, CDR, and CDR1 | |
| 42 | 1691 | 0.25 | K128T | |
| 3731 | >64 | F126L and K143R | MDR1 | |
| 3733 | 64 | F126L and K143R | MDR1 | |
| 43 | 1649 | 0.5 | ||
| 3034 | >64 | MDR1, CDR, CDR1, and CDR2 | ||
| 51 | 2274 | 16 | S405F | |
| 2257 | 64 | S405F | ||
| 2339 | 64 | S405F | CDR, CDR1, and CDR2 | |
| 59 | 3917 | 4 | F449S | |
| 4617 | 64 | F449S and T229A | MDR1, CDR, and CDR1 | |
| 4639 | >64 | F449S and T229A | ||
| 64 | 4018 | 4 | ||
| 4380 | 64 | V437I | CDR, CDR1, and CDR2 |
DISCUSSION
The multiplicity of mechanisms of resistance to azole antifungal agents represents a set of biological tools that enables yeast cells to develop resistance by using different combinations of mechanisms. A limited number of studies support the role of these mechanisms in the development of C. albicans resistance in a small number of clinical isolates (34, 42). The aim of the present study was to assess the prevalence of specific mechanisms of resistance in matched sets of susceptible and resistant C. albicans isolates recovered from HIV-infected patients with OPC monitored longitudinally while under treatment with fluconazole.
The majority of the fluconazole-resistant isolates also showed decreased levels of susceptibility to the various azole compounds tested: itraconazole, voriconazole, and posaconazole. In the case of voriconazole, this could be explained by the fact that voriconazole shows properties similar to those of fluconazole with respect to its capacity to be a substrate for multidrug efflux transporters and to respond to ERG11 mutations, as has recently been shown by Sanglard et al. (D. Sanglard, F. Ischer, and J. Bille, Abstr. 40th Intersci. Conf. Antimicrob. Agents Chemother., abstr. 1711, p. 393, 2000). Confirming previous results, decreased susceptibility to multiple azole derivatives was mainly associated with overexpression of CDR genes (18, 30, 40), but it was also associated with the presence of specific point mutations in the ERG11 gene. Importantly, all isolates remained susceptible to amphotericin B, indicating the lack of cross-resistance to polyene antifungal agents.
The most frequent molecular mechanism of azole resistance was the upregulation of efflux pumps. Thus, overexpression of the CDR and MDR1 genes was detected in isolates from 83 and 67% of the patients, respectively. Upregulation of ERG11 genes was detected in isolates from 42% of the patients. In 86% of the isolates it to be appeared associated with the upregulation of MDR1 and CDR genes, and in 28% of the isolates it appeared to be associated with point mutations in their ERG11 genes.
PCR amplification and sequencing of the ERG11 genes encoding lanosterol 14α-demethylase showed 13 nucleotide changes that led to amino acid substitutions in the enzymes of the resistant isolates. Using the technique developed by Sanglard et al. (33), we could demonstrate that 11 of these point mutations were linked to increases in the MICs of fluconazole when the alleles carrying these mutations were expressed in S. cerevisiae. Overall, point mutations in ERG11 genes with an effect on the affinity of the enzyme for the azoles were observed in the isolates from 58% of the patients. In 55% of the isolates it appeared to be combined with upregulation of efflux pumps. While nine mutations were described previously, two were novel (G307S and D446N). The fact that many of the mutations described here were also found independently by others in the ERG11 genes from other isolates obtained in different geographic locations illustrates that there may be preferential amino acid positions able to confer a phenotype of resistance to fluconazole and other azole derivatives. These mutations repeatedly identified by different groups may represent “hot spots” for the development of azole resistance (3, 6, 16, 21, 33, 37). Remarkably, most of these substitutions are present in domains that are highly conserved in lanosterol 14α-demethylases across fungi, suggesting the importance of these residues for the maintenance of function through evolution. According to molecular modeling of the C. albicans lanosterol 14α-demethylase, these regions correspond to important functional domains of the enzyme in its interaction with the heme moiety at its active site and at another region believed to play a role in the entry of the substrate in the substrate pocket (5, 10). Three other conclusions can be drawn from the nucleotide sequences obtained: (i) allelic differences are present in the ERG11 gene for some of the substitutions identified; (ii) multiple isolates obtained from the same patient at different intervals exhibited the same or very similar polymorphisms, indicating a high degree of relatedness; and (iii) differences in nucleotide sequences among strains obtained from different patients indicate heterogeneity in the C. albicans population.
In summary, we have performed a study to evaluate the prevalence of molecular mechanisms of azole resistance in C. albicans strains displaying high-level fluconazole resistance isolated from a cohort of HIV-infected patients who presented with OPC while on treatment with fluconazole. The results obtained showed that the resistance to fluconazole and other azoles is the result of a combination of different molecular mechanisms, with the predominating mechanism being the overexpression of efflux transporters (ABC transporters and major facilitators), alone or in combination with overexpression of the target enzyme and the presence of point mutations in such enzymes that alter the interaction between the azole antifungal agents and the enzyme. These results may help in the development of new strategies to overcome the problem of resistance as well as new treatments for this condition (9, 34).
ACKNOWLEDGMENTS
This work was supported by a grant from Pfizer Inc. and Public Health Service grants 5 R01 DE11381 (to T.F.P.), 1 R29 AI42401 (to J.L.L.-R.), and M01-RR-01346 for the Frederic C. Bartter General Clinical Research Center. S.P. acknowledges the receipt of a NATO postdoctoral fellowship. M.M. was supported by a research supplement to support underrepresented minorities (grant 3 R01 DE11381-04A2S2, to T.F.P.). R.A.S. was supported by the Prematriculation Program, Medical Hispanic Center of Excellence, UTHSCSA. D.S. was supported by grant 3100-055901.98/1 from the Swiss National Foundation. Chromogenic medium was provided by the CHROMagar Company.
We thank the Fungus Testing Laboratory at UTHSCSA for performing antifungal susceptibility testing.
REFERENCES
- 1.Albertson G D, Niimi M, Cannon R D, Jenkinson H F. Multiple efflux mechanisms are involved in Candida albicans fluconazole resistance. Antimicrob Agents Chemother. 1996;40:2835–2841. doi: 10.1128/aac.40.12.2835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Altschul S F, Madden T L, Schaffer A A, Zhang J, Zhang Z, Miller W, Lipman D J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Asai K, Tsuchimori N, Okonogi K, Perfect J R, Gotoh O, Yoshida Y. Formation of azole-resistant Candida albicans by mutation of sterol 14-demethylase P450. Antimicrob Agents Chemother. 1999;43:1163–1169. doi: 10.1128/aac.43.5.1163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Balzi E, Goffeau A. Genetics and biochemistry of yeast multidrug resistance. Biochim Biophys Acta. 1994;1187:152–162. doi: 10.1016/0005-2728(94)90102-3. [DOI] [PubMed] [Google Scholar]
- 5.Boscott P E, Grant G H. Modeling cytochrome P450 14 alpha demethylase (Candida albicans) from P450cam. J Mol Graph. 1994;12:185–192. doi: 10.1016/0263-7855(94)80086-3. [DOI] [PubMed] [Google Scholar]
- 6.Favre B, Didmon M, Ryder N S. Multiple amino acid substitutions in lanosterol 14α-demethylase contribute to azole resistance in Candida albicans. Microbiology. 1999;145:2715–2725. doi: 10.1099/00221287-145-10-2715. [DOI] [PubMed] [Google Scholar]
- 7.Franz R, Kelly S L, Lamb D C, Kelly D E, Ruhnke M, Morschhäuser J. Multiple molecular mechanisms contribute to a stepwise development of fluconazole resistance in clinical Candida albicans strains. Antimicrob Agents Chemother. 1998;42:3065–3072. doi: 10.1128/aac.42.12.3065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Franz R, Ruhnke M, Morschhäuser J. Molecular aspects of fluconazole resistance development in Candida albicans. Mycoses. 1999;42:453–458. doi: 10.1046/j.1439-0507.1999.00498.x. [DOI] [PubMed] [Google Scholar]
- 9.Georgepapadakou N H, Walsh T J. Antifungal agents: chemotherapeutic targets and immunologic strategies. Antimicrob Agents Chemother. 1996;40:279–291. doi: 10.1128/aac.40.2.279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ji H, Zhang W, Zhou Y, Zhang M, Zu J, Song Y, Lu J, Zhu J. A three-dimensional model of lanosterol 14α-demethylase of Candida albicans and its interaction with azole antifungals. J Med Chem. 2000;43:2493–2505. doi: 10.1021/jm990589g. [DOI] [PubMed] [Google Scholar]
- 11.Kelly S, Lamb D C, Kelly D E. Y132.H substitution in Candida albicans sterol 14α-demethylase confers fluconazole resistance by preventing binding to haem. FEMS Microbiol Lett. 1999;180:171–175. doi: 10.1111/j.1574-6968.1999.tb08792.x. [DOI] [PubMed] [Google Scholar]
- 12.Kelly S, Lamb D C, Loeffler J, Einsele H, Kelly D E. The G464S amino acid substitution in Candida albicans sterol 14α-demethylase causes fluconazole resistance in the clinic through reduced affinity. Biochem Biophys Res Commun. 1999;262:174–179. doi: 10.1006/bbrc.1999.1136. [DOI] [PubMed] [Google Scholar]
- 13.Lai M H, Kirsch D R. Nucleotide sequence of cytochrome P450 LIA1 (lanosterol 14α-demethylase) from Candida albicans. Nucleic Acids Res. 1989;17:804. doi: 10.1093/nar/17.2.804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lamb D C, Kelly D E, Schunck W H, Shyadehi A Z, Akhtar M, Lowe D J, Baldwin B D, Kelly S L. The mutation T315A in Candida albicans sterol 14α-demethylase causes reduced enzyme activity and fluconazole-resistance through reduced affinity. J Biol Chem. 1997;272:5682–5688. doi: 10.1074/jbc.272.9.5682. [DOI] [PubMed] [Google Scholar]
- 15.Lamb D C, Kelly D E, White T C, Kelly S L. The R467R amino acid substitution in Candida albicans sterol 14α-demethylase caused drug resistance through reduced affinity. Antimicrob Agents Chemother. 2000;44:63–67. doi: 10.1128/aac.44.1.63-67.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Löffler J, Kelly S L, Hebart H, Schumacher U, Lass-Flörl C, Einsele H. Molecular analysis of cyp51 from fluconazole-resistant Candida albicans strains. FEMS Microbiol Lett. 1997;151:263–268. doi: 10.1111/j.1574-6968.1997.tb12580.x. [DOI] [PubMed] [Google Scholar]
- 17.López-Ribot J L, McAtee R K, Kirkpatrick W R, Patterson T F. Comparison of DNA-based typing methods to assess genetic diversity and relatedness among Candida albicans clinical isolates. Rev Iberoam Micol. 2000;17:49–54. [PubMed] [Google Scholar]
- 18.Lopez-Ribot J L, McAtee R K, Lee L N, Kirkpatrick W R, White T C, Sanglard D, Patterson T F. Distinct patterns of gene expression associated with development of fluconazole resistance in serial Candida albicans isolates from human immunodeficiency virus-infected patients with oropharyngeal candidiasis. Antimicrob Agents Chemother. 1998;42:2932–2937. doi: 10.1128/aac.42.11.2932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lopez-Ribot J L, McAtee R K, Perea S, Kirkpatrick W R, Rinaldi M G, Patterson T F. Multiple resistant phenotypes of Candida albicans coexist during episodes of oropharyngeal candidiasis in human immunodeficiency virus-infected patients. Antimicrob Agents Chemother. 1999;43:1621–1630. doi: 10.1128/aac.43.7.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Manavathu E K, Kallakuri S, Arganoza M T, Vazquez J A. Amino acid variations of cytochrome P-450 lanosterol 14α-demethylase (CYP51A1) from fluconazole-resistant clinical isolates of Candida albicans. Rev Iberoam Micol. 1999;16:198–203. [PubMed] [Google Scholar]
- 21.Marichal P, Koymans L, Willemsens S, Bellens D, Verhasselt P, Luyten W, Borges M, Ramaekers F C, Odds F C, Vanden Bossche H. Contribution of mutations in the cytochrome P450 14α-demethylase (Erg11, Cyp51p) to azole resistance in Candida albicans. Microbiology. 1999;145:2701–2713. doi: 10.1099/00221287-145-10-2701. [DOI] [PubMed] [Google Scholar]
- 22.Marr K A, Lyons C N, Rustad T, Bowden R A, White T C. Rapid, transient fluconazole resistance in Candida albicans is associated with increased mRNA levels of CDR. Antimicrob Agents Chemother. 1998;42:2584–2589. doi: 10.1128/aac.42.10.2584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.National Committee for Clinical Laboratory Standards. Reference method for broth dilution antifungal susceptibility testing of yeast's: approved standard. NCCLS document M27-A. Wayne, Pa: National Committee for Clinical Laboratory Standards; 1997. [Google Scholar]
- 24.Patterson T F, Kirkpatrick W R, Revankar S G, McAtee R K, Fothergill A W, McCarthy D I, Rinaldi M G. Comparative evaluation of macrodilution and chromogenic agar screening for determining fluconazole susceptibility of Candida albicans. J Clin Microbiol. 1996;34:3237–3239. doi: 10.1128/jcm.34.12.3237-3239.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Patterson T F, Revankar S G, Kirkpatrick W R, Dib O, Fothergill A W, Redding S W, Sutton D A, Rinaldi M G. Simple method for detecting fluconazole-resistant yeasts with chromogenic agar J. Clin Microbiol. 1996;34:1794–1797. doi: 10.1128/jcm.34.7.1794-1797.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Prasad R, De W P, Goffeau A, Balzi E. Molecular cloning and characterization of a novel gene of Candida albicans, CDR1, conferring multiple resistance to drugs and antifungals. Curr Genet. 1995;27:320–329. doi: 10.1007/BF00352101. [DOI] [PubMed] [Google Scholar]
- 27.Revankar S G, Dib O P, Kirkpatrick W R, McAtee R K, Fothergill A W, Rinaldi M G, Redding S W, Patterson T F. Clinical evaluation and microbiology of oropharyngeal infection due to fluconazole-resistant Candida in human immunodeficiency virus-infected patients. Clin Infect Dis. 1998;26:960–963. doi: 10.1086/513950. [DOI] [PubMed] [Google Scholar]
- 28.Revankar S G, Kirkpatrick W R, McAtee R K, Dib O P, Fothergill A W, Redding S W, McGough D A, Rinaldi M G, Patterson T F. Detection and significance of fluconazole resistance in oropharyngeal candidiasis in HIV-infected patients. J Infect Dis. 1996;174:821–827. doi: 10.1093/infdis/174.4.821. [DOI] [PubMed] [Google Scholar]
- 29.Rex J H, Pfaller M A, Galgiani J N, Bartlett M S, Espinel-Ingroff A, Ghannoum M A, Lancaster M, Odds F C, Rinaldi M G, Walsh T J, Berry A L. Development of interpretive breakpoints for antifungal susceptibility testing: conceptual framework and analysis of in vitro-in vivo correlation data for fluconazole, itraconazole, and Candida infections. Clin Infect Dis. 1997;24:235–247. doi: 10.1093/clinids/24.2.235. [DOI] [PubMed] [Google Scholar]
- 30.Sanglard D, Kuchler K, Ischer F, Pagani J L, Monod M, Bille J. Mechanisms of resistance to azole antifungal agents in Candida albicans isolates from AIDS patients involve specific multidrug transporters. Antimicrob Agents Chemother. 1995;39:2378–2386. doi: 10.1128/aac.39.11.2378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sanglard D, Ischer F, Monod M, Bille J. Susceptibilities of Candida albicans multidrug transporter mutants to various antifungal agents and other metabolic inhibitors. Antimicrob Agents Chemother. 1996;40:2300–2305. doi: 10.1128/aac.40.10.2300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sanglard D, Ischer F, Monod M, Bille J. Cloning of Candida albicans genes conferring resistance to azole antifungal agents: characterization of CDR2, a new multidrug ABC transporter gene. Microbiology. 1997;143:405–416. doi: 10.1099/00221287-143-2-405. [DOI] [PubMed] [Google Scholar]
- 33.Sanglard D, Ischer F, Koymans L, Bille J. Amino acid substitutions in the cytochrome P-450 lanosterol 14α-demethylase (CYP51A1) from azole-resistant Candida albicans clinical isolates contribute to resistance to azole antifungal agents. Antimicrob Agents Chemother. 1998;42:241–253. doi: 10.1128/aac.42.2.241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sanglard D, Ischer F, Calabrese D, de Micheli M, Bille J. Multiple resistance mechanisms to azole antifungals in yeast clinical isolates. Drug Resistance Updates. 1998;1:255–265. doi: 10.1016/s1368-7646(98)80006-x. [DOI] [PubMed] [Google Scholar]
- 35.Schmid J, Voss E, Soll D R. Computer-assisted methods for assessing strain relatedness in Candida albicans by fingerprinting with the moderately repetitive sequence Ca3. J Clin Microbiol. 1990;28:1236–1243. doi: 10.1128/jcm.28.6.1236-1243.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Vanden Bossche H, Marichal P, Odds F C. Molecular mechanisms of drug resistance in fungi. Trends Microbiol. 1994;2:393–400. doi: 10.1016/0966-842x(94)90618-1. [DOI] [PubMed] [Google Scholar]
- 37.Vanden Bossche H, Marichal P, Gorrens J, Bellens D, Moereels H, Janssen P A J. Mutation in cytochrome P450-dependent 14α-demethylase results in decreased affinity for azole antifungals. Biochem Soc Trans. 1990;18:56–59. doi: 10.1042/bst0180056. [DOI] [PubMed] [Google Scholar]
- 38.Venkateswarlu K D, Denning W, Manning N J, Kelly S L. Resistance to fluconazole in Candida albicans from AIDS patients correlated with reduced intracellular accumulation of drug. FEMS Microbiol Lett. 1995;131:337–341. doi: 10.1111/j.1574-6968.1995.tb07797.x. [DOI] [PubMed] [Google Scholar]
- 39.Walsh T J, Gonzalez C E, Piscitelli S, Bacher J D, Peter J, Torres R, Shetti D, Katsov V, Kligys D, Lyman C A. Correlation between in vitro and in vivo antifungal activities in experimental fluconazole-resistant oropharyngeal and esophageal candidiasis. Antimicrob Agents Chemother. 2000;38:2369–2373. doi: 10.1128/jcm.38.6.2369-2373.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.White T C. Increased mRNA levels of ERG16, CDR, and MDR1 correlate with increases in azole resistance in Candida albicans isolates from a patient infected with human immunodeficiency virus. Antimicrob Agents Chemother. 1997;41:1482–1487. doi: 10.1128/aac.41.7.1482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.White T C. The presence of an R467K amino acid substitution and loss of allelic variation correlate with an azole-resistant lanosterol 14α-demethylase in Candida albicans. Antimicrob Agents Chemother. 1997;41:1488–1494. doi: 10.1128/aac.41.7.1488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.White T C, Marr K A, Bowden R A. Clinical, cellular, and molecular factors that contribute to antifungal drug resistance. Clin Microbiol Rev. 1998;11:382–402. doi: 10.1128/cmr.11.2.382. [DOI] [PMC free article] [PubMed] [Google Scholar]