Abstract
Purpose:
In the field of medical diagnosis, deep learning-based computer-aided detection of diseases will reduce the burden of physicians in the diagnosis of diseases especially in the case of lung cancer nodule classification.
Materials and Methods:
A hybridized model which integrates deep features from Residual Neural Network using transfer learning and handcrafted features from the histogram of oriented gradients feature descriptor is proposed to classify the lung nodules as benign or malignant. The intrinsic convolutional neural network (CNN) features have been incorporated and they can resolve the drawbacks of handcrafted features that do not completely reflect the specific characteristics of a nodule. In the meantime, they also reduce the need for a large-scale annotated dataset for CNNs. For classifying malignant nodules and benign nodules, radial basis function support vector machine is used. The proposed hybridized model is evaluated on the LIDC-IDRI dataset.
Results:
It has achieved an accuracy of 97.53%, sensitivity of 98.62%, specificity of 96.88%, precision of 95.04%, F1 score of 0.9679, false-positive rate of 3.117%, and false-negative rate of 1.38% and has been compared with other state of the art techniques.
Conclusions:
The performance of the proposed hybridized feature-based classification technique is better than the deep features-based classification technique in lung nodule classification.
Keywords: Convolutional neural network, hybridized features, radial basis function support vector machine, residual neural network, transfer learning
INTRODUCTION
Cancer has become one of the leading causes of mortality in the whole world. It is one of the major public health concerns of every government.[1] The world is in midst of an unprecedented pandemic, that is, coronavirus disease (CoViD-19). However, death rates of cancer and AIDS, the number of deaths due to infectious diseases like the CoViD-19 would pale in comparison. In the year 2018 alone, 9.6 million deaths were due to various types of cancer.[2]
There are six common types of cancer such as prostate, colorectal, breast, stomach, cervical, and lung cancer. Lung cancer was thought to occur only in high per capita income countries. In the past decade, it has been identified as a global scourge.[3] Among the six most common cancer types worldwide mentioned above, lung cancer leads in terms of both incidence and mortality rate with an estimated 2.1 million new cases and deaths of 1.8 million people in the year 2018 alone.[4] Recent estimates point out that the lung cancer indents are about to increase by 38% and the total number of cases would rise to 2.89 million by 2030. The mortality due to the same would be a staggering 2.49 million (39% increase) by 2030.[5]
Computed tomography (CT) scans to detect lung cancer are taken in large volume and significant success has been achieved in reducing lung cancer mortality. The generated CT scan images are analyzed manually by the radiologist's slice by slice. Even though, the CT scans helped in the reduction of lung cancer-related deaths by 20%, the task of analyzing CT scans has created problems like human error. The task is time-consuming and is not economical.[6]
To overcome these difficulties, computer-aided diagnosis (CAD) has come to the front to analyze the large volume dataset. The application of CAD has improved the 5-year survival rate from 15% to over 70%.[7] Hence, the importance of CAD for the management of lung cancer disease has gained traction in the past decade.
Lung nodules are small masses of tissue appearing in the lung due to various reasons. They appear as opaque white objects on a CT image and the sizes vary from 3 mm to 30 mm. There can be benign lung nodules and malignant (cancerous) lung nodules. In few scans and subsequent analyses, a benign nodule can be classified as a cancerous lung nodule.[7] To avoid this error, an efficient classification scheme is the need of the hour. It is not easy to differentiate benign nodules and malignant nodules because both are having similar visual representations. Many CADs available in the literature are based on image processing and traditional machine learning techniques. Very recently, deep learning-based CADs are being introduced in most of the areas for detecting the abnormalities in the medical images.
Classifying benign nodules from malignant nodules is imperative in the analysis of lung cancer.[8] This can be carried out also by a biopsy or a positron emission tomography scan. Although many researches are available for differentiating pulmonary nodules, they depend on image processing-based segmentation and feature extraction techniques.[9,10] There are two broad categories in the classification of lung nodules. One category is the traditional classification where the features from the nodule images are calculated using different feature engineering techniques and these features are used to classify the lung nodules into benign and malignant. Another category is entirely independent of feature engineering by domain experts. Such method is based on deep learning where the deep learning algorithm itself learns the features from the given input images and classifies the nodules into benign and malignant. Among deep learning methodologies, CNN has been extensively used for extracting the features without manual intervention which are termed as deep features.
In the conventional CAD system, handcrafted features were computed from malignant and benign nodule images for the lung nodule classification using traditional classifiers such as linear discriminant analysis, artificial neural network, and support vector machine (SVM).[11,12,13,14,15] Deep learning architectures such as deep belief networks and CNN were able to classify the nodules more efficiently than the traditional classifiers which used handcrafted features.[16] A multi-crop convolutional neural network (MC-CNN) was developed to detect the malignancy of the nodules.[17] For improving the lung nodule classification, evolutionary algorithms were incorporated in CNN architecture.[18] Recently, the deep features were combined with the specific handcrafted features to improve the classification accuracy.[19,20,21,22,23,24,25]
Deep learning architecture has given better classification accuracies when compared with traditional handcrafted features. However, one inherent difficulty is in obtaining of the large volume of datasets to have significant results which is difficult in medical applications. It is imperative to note that while deep learning architecture is enough for detecting images in biometric systems, it may not be sufficient to outperform the handcrafted features in all cases where these methodologies are employed. For example, the handcrafted features are found to have better output in the cases of face and iris recognition whereas the deep features have outperformed the handcrafted features in fingerprint recognition systems. In the classification of lung nodules by handcrafted features, the accuracy is not better when compared with deep features-based classification. The classification scheme based on deep features may miss out few salient points if it is used alone as it is evident from the literature review. Hence, this research article presents an automatic system to assist clinicians in diagnosing lung nodules with the hybridized feature set where the experiments are carried out using the LIDC-IDRI dataset.
MATERIALS AND METHODS
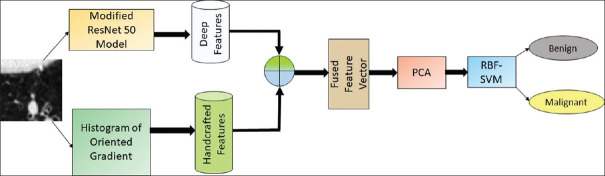
In this proposed approach, hybridization of deep features and handcrafted features are used to classify the lung nodules into benign and malignant. Figure 1 shows the schematic diagram of the proposed methodology. This system uses the modified ResNet50 model using the transfer learning technique for deep feature extraction and integrates the deep features with the traditional histogram of oriented gradient (HOG) features. Since the fused feature set is very large, training the machine learning classifier will be complex and take much time. To reduce the complexity and computation time, principal component analysis (PCA) has been introduced. PCA not only reduces the dimension of the data but also preserves the important information. For differentiating malignant nodules from benign nodules, radial basis function SVM (RBF-SVM) has been employed.
Figure 1.
Proposed methodology for lung nodule classification
Lung Image Database Consortium and Image Database Resource Initiative (LIDC-IDRI) dataset
LIDC/IDRI dataset[26] contains 1018 scans and each scan consists of CT images of the chest and an XML file which has annotations of 4 radiologists. In this XML file, the malignancy level for the nodules is specified. The malignancy rating of each nodule is represented in the range of 1–5. Based on the information given in the XML file, 2625 nodules have been extracted for this work with the help of pylidc library as given in Table 1 .[27] The nodules which got the malignancy rating 1 and 2 are termed as “highly unlikely for cancer” and 1136 nodules have been extracted; the nodules who got the malignancy rate 3 are termed as indeterminate nodules and 980 such nodules have been extracted. The nodules which got the malignancy rate 4 and 5 are termed as “highly likely for cancer” and 509 nodules have been extracted. The classification scheme employed in this work is based on bi-level classification. The nodules with malignancy ratings 1 and 2 are considered as benign nodules and the malignancy ratings 4 and 5 are considered as malignant nodules. The number of benign nodules and malignant nodules extracted from the database are 1136 and 509 respectively. These data are not sufficient to train the machine learning model to predict the classes. To increase the number of nodules, data augmentation technique is employed in this work. This technique will avoid the overfitting of the model. Using data augmentation techniques such as horizontal flipping and vertical flipping, the number of nodule images has been increased to 4544 benign nodules and 2036 malignant nodules [Table 2].
Table 1.
Malignancy rate for lung nodules in lung image database consortium - image database resource initiative
| Malignancy rate | Number of nodules | Nature of the nodule |
|---|---|---|
| 1, 2 | 1136 | Highly unlikely for cancer |
| 3 | 980 | Indeterminate |
| 4, 5 | 509 | Highly likely for cancer |
Table 2.
Dataset used for this work
| Type of nodule | Nodules extracted from database | Augmented nodules |
|---|---|---|
| Benign | 1136 | 4544 |
| Malignant | 509 | 2036 |
Preprocessing
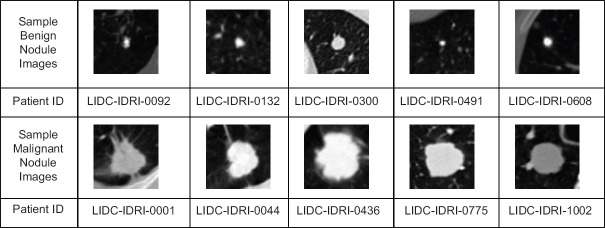
The lung CT images collected from the LIDC-IDRI dataset and each image has the matrix size of 512 × 512. The thoracic CT scan not only consists of lung parenchyma but also comprises of the image of sternum, rib, ascending aorta, superior vena cava, trachea, descending aorta, vertebra, thecal sac with the spinal cord. These extraneous information are not necessary for the experimentation. Thus, the aim of pre-processing is to extract the nodule region from the lung parenchyma. The centroid information of the nodule is available in the XML file and the region of interest of size 64 × 64 is cropped with respect to the centroid information. A sample of cropped lung nodule images from the dataset is shown in Figure 2.
Figure 2.
Sample benign and malignant nodule slices from LIDC-IDRI datase
Feature extraction
The images carry a lot of information and processing the entire information requires a huge necessity of memory and computation time. The aim of feature extraction is to extract the important properties or features from the input image which can differentiate one pattern from the rest of the patterns. During this feature extraction process, the irrelevant information will be eliminated without any significant loss in important information related to the input images.[28]
Handcrafted feature extraction using histogram of oriented gradient
HOG feature descriptor[29] is employed in the present work to extract the features related to the shape characteristics of lung nodules. The lobulated, spiculated, and ragged nodules are having more probability to be malignant whereas round, tentacular and polygonal-shaped nodules are having more probability to be benign. Wang et al. have reported that the HOG features are suitable for describing the shape and edge characteristics of the malignant and benign nodules.[30] In this work for calculating the HOG features, the orientation bin has been set to 9, the size of a cell is (8,8) and the number of cells in each block is (3,3).
Deep feature extraction using convolutional neural network
Deep CNN (DCNN) is the end-to-end machine learning framework which does not need feature engineering of the input images. In natural image analysis, DCNNs have made a giant stride in tasks such as object recognition and image classification. If they are trained with less number of data, they cannot classify or recognize the given input with high accuracy.
There are two difficulties a researcher faces when he/she uses DCNNs in lung nodule classification. Firstly, the datasets available in the public domain like LIDC-IDRI for lung nodule classification task is very small in number when it is compared with millions of data available with the ImageNet dataset. The second difficulty is the subtlety of the nodule classification task as the differences between a benign and malignant nodules are not self-evident. The initial problem can be overcome by transferring the weights from pretrained CNNs which are trained for different applications to the problem at hand. This technique is referred to as transfer learning. The second difficulty can be taken into account by concatenating deep features and handcrafted features and creating hybridized features.
Residual Network (ResNet) is a DCNN model which is less complex than other models.[31] It is easier to optimize the residual network. When the network becomes deep, the accuracy of the network reaches saturation or it starts decreasing suddenly due to vanishing gradient problem. To eliminate the vanishing gradient problem, skip connections have been introduced in the ResNet architecture. The skip connections make a way to add the output from a previous layer to a later layer.[31]
The basic principle of the residual network is utilized in the lung nodule classification problem. The initial convolution layer has been modified to accept 64 x 64 size grayscale image input. Using the transfer learning concept, the pretrained weights are used at the starting point. The fully connected layers have been completely removed from the baseline ResNet50 model.
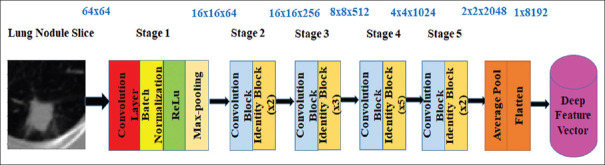
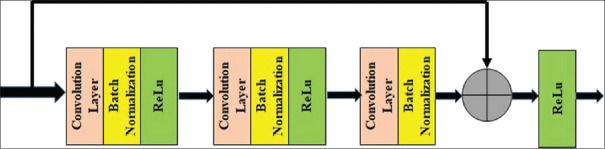
The modified architecture of ResNet50 is shown in Figure 3. This architecture consists of 5 stages. The first stage has a convolution layer, a batch normalization layer, a ReLu Layer and a max-pooling layer. In stage 2 to stage 5, convolution block and identity blocks are available. The convolution block has 3 convolutional layers and the identity block (ID block) has 3 convolutional layers. The identity block with the layers and skip connections are shown in Figure 4. The lung nodule image size used in this work is 64 × 64 and this image is given as input to stage 1. After stage 1, the image has been converted into feature map, the feature map of size 16 × 16 × 64 is obtained. Stage 2 gives the feature map of size 16 × 16 × 256. Stage 3, stage 4, and stage 5 give the feature map of size 8 × 8 × 512, 4 × 4 × 1024, and 2 × 2 × 2048, respectively. These feature maps are allowed to pass through the average pool layer and then flatten layer. A feature vector of size 1 × 8192 is obtained from the flatten layer. Thus, deep features are collected from all the training images.
Figure 3.
Modified ResNet50 model to extract deep features
Figure 4.
Skip connection in ResNet50
Hybridized features
The hybridized features (fh) have been formed by concatenating the deep ResNet50 features (fres) with the handcrafted HOG features (fhog). It is proposed to hybridize the two different features because one set of feature extraction methodology may overlook the significant results of the other methodology.[20] The hybridization methodology takes complete advantage of the powerful handcrafted features and the highest level DCNN features.[25] The features extracted by traditional feature descriptor and DCNN are complementary in nature and if concatenated, they give features that are combinations of both.[30]
Feature reduction using principal component analysis
PCA is one of the feature reduction methods. It will transform a large set of predictor variables into a smaller set of predictor variables but maintains most of the information present.[28] The hybridized feature set has a large number of features (predictor variables). If these features are applied to the classifier directly, it will lead to computational complexity and more computation time. To mitigate these problems, the PCA technique is employed in this work without losing much information.
Classification
Support vector classifier (SVC) is a highly admired machine learning algorithm for classification. It provides highly accurate classification. The SVC can handle non-linear data points using kernels. Linear kernel, polynomial kernel, and RBF kernel are common types used in SVC. Among these three types, the RBF kernel has been selected in the proposed methodology because of the attractive properties of RBF.
RBF kernel is invariant to translation and it is easy to tune this kernel because it has single parameter. Moreover, this kernel is isotropic.
The mathematical representation of the RBF kernel is given in equation 1.
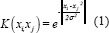
where σ is variance. It is a hyperparameter.
xi represents the support vector and xj represents data point.
xi − xj represents the Euclidean distance.
If the Euclidean distance between the support vector and data point is less, they are similar and the kernel value will be maximum for that data point. If the distance between the support vector and data point is more, they are dissimilar and the kernel value will be minimum for that data point. The maximum value of the kernel will be 1 and the minimum value of the kernel will be 0. It is important to find the optimal value for the parameter σ. This single parameter is tuned by employing Grid Search Cross-Validation approach. A significant advantage of using RBF SVM is less memory requirement because during training, it will store only the support vectors and not the entire data points.
Model evaluation
The performance of the proposed model has been evaluated by generating confusion matrix and receiver operating characteristic (ROC) curve. The structure of the confusion matrix for lung nodule classification is shown in Table 3. True negative (TN) infers that the benign nodule is correctly identified as benign nodule. True positive (TP) tells that the malignant nodule is correctly identified as malignant nodule. False-positive (FP) represents that the benign nodule is wrongly identified as malignant nodule. False-negative (FN) indicates that the malignant nodule is wrongly identified as benign nodule.
Table 3.
Structure of the confusion matrix for lung nodule classification
| Benign | Malignant | |
|---|---|---|
| Actual class | ||
| Benign | TN | FP |
| Malignant | FN | TP |
| Predicted class |
TN: True negative, TP: True positive, FP: False positive, FN: False negative
Based on the values generated in the confusion matrix, different performance metrics such as Accuracy, Sensitivity, Specificity, Precision, F1 score, FP rate (FPR) and FN rate (FNR) are calculated.
For comparing the performance of the proposed model, different feature extraction and classifier combinations have been experimented. The handcrafted features extraction techniques such as Gray Level Co-occurrence Matrix (GLCM),[32] Local Binary Pattern (LBP)[33] and HOG[29] are used for extracting features. These feature sets are applied independently to train four different classifiers such as logistic regression, linear SVM, RBF SVM, and Random Forest. In the handcrafted feature-based experiments, 12 different combinations (models) have been analyzed. These models are listed in Table 4. To analyze the performance of deep features in lung nodule classification, VGG16, VGG19, and ResNet50 features are considered, and these features are used to train four different classifiers such as logistic regression, linear SVM, RBF SVM, and Random Forest. In the deep feature-based experiments, 12 different combinations (models) have been analyzed. These models are listed in Table 5. The proposed hybridized feature technique has been tested for 12 different combinations and those combinations (models) are listed in Table 6.
Table 4.
Performance measures of handcrafted features
| Model | Explanation | Accuracy (%) | Sensitivity (%) | Specificity (%) | Precision (%) | F1 score | FPR (%) | FNR (%) |
|---|---|---|---|---|---|---|---|---|
| Model 1 | GLCM + logistic regression | 57.31 | 85.18 | 40 | 46.44 | 0.6 | 60 | 14.8 |
| Model 2 | GLCM + linear SVM | 56.86 | 83.79 | 40.53 | 46 | 0.59 | 59 | 16 |
| Model 3 | GLCM + RBF SVM | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 4 | GLCM + random forest | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 5 | LBP + logistic regression | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 6 | LBP + linear SVM | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 7 | LBP + RBF SVM | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 8 | LBP + random forest | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 9 | HOG + logistic regression | 74 | 93.47 | 62 | 60 | 0.73 | 38 | 6.52 |
| Model 10 | HOG + linear SVM | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 11 | HOG + RBF SVM | 78 | 95.45 | 67 | 64 | 0.77 | 32.6 | 4.5 |
| Model 12 | HOG + random forest | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
FPR: False-positive rate, FNR: False-negative rate, GLCM: Gray level co-occurrence matrix, RBF: Radial basis function, SVM: Support vector machine, LBF: Local binary pattern, HOG: Histogram of oriented gradients
Table 5.
Performance measures of deep features
| Model | Explanation | Accuracy (%) | Sensitivity (%) | Specificity (%) | Precision (%) | F1 score | FPR (%) | FNR (%) |
|---|---|---|---|---|---|---|---|---|
| Model 13 | VGG16 + logistic regression | 58.36 | 63.24 | 55.4 | 46.24 | 0.534 | 44.6 | 36.76 |
| Model 14 | VGG16 + linear SVM | 62.24 | 0 | 100 | 0 | 0 | 0 | 100 |
| Model 15 | VGG16 + RBF-SVM | 82.985 | 83.79 | 82.49 | 74.39 | 0.788 | 17.5 | 16.2 |
| Model 16 | VGG16 + random forest | 77.84 | 82.21 | 75.18 | 66.77 | 0.737 | 24.8 | 17.78 |
| Model 17 | VGG19 + logistic regression | 74.22 | 80.04 | 70.69 | 62.31 | 0.701 | 29.3 | 19.96 |
| Model 18 | VGG19 + linear SVM | 76.42 | 76.68 | 76.26 | 66.21 | 0.711 | 23.74 | 23.32 |
| Model 19 | VGG19 + RBF-SVM | 83.06 | 90.12 | 78.78 | 72.04 | 0.8 | 21.22 | 9.88 |
| Model 20 | VGG19 + random forest | 80.15 | 84.58 | 77.46 | 69.48 | 0.763 | 22.54 | 15.4 |
| Model 21 | ResNet50 + logistic regression | 78.36 | 83 | 75.54 | 67.31 | 0.74 | 24.46 | 17 |
| Model 22 | ResNet50 + linear SVM | 79.03 | 87.15 | 74 | 67.12 | 0.758 | 25.89 | 12.85 |
| Model 23 | ResNet50 + RBF-SVM | 83.06 | 95.06 | 75.78 | 70.42 | 0.81 | 24.2 | 4.94 |
| Model 24 | ResNet50 + random forest | 80.3 | 87.15 | 79.14 | 68.9 | 0.77 | 23.86 | 12.85 |
FPR: False-positive rate, FNR: False-negative rate, RBF: Radial basis function, SVM: Support vector machine
Table 6.
Performance analysis of hybridized features in lung nodule classification
| Model | Explanation | Accuracy (%) | Sensitivity (%) | Specificity (%) | Precision (%) | F1 score | FPR (%) | FNR (%) |
|---|---|---|---|---|---|---|---|---|
| Model 25 | VGG16 + HOG + logistic regression | 78.66 | 80.83 | 77.34 | 68.39 | 0.74 | 22.66 | 19.16 |
| Model 26 | VGG16 + HOG + linear SVM | 78.43 | 82 | 76.25 | 67.69 | 0.74 | 23.74 | 17.98 |
| Model 27 | VGG16 + HOG + RBF-SVM | 82.46 | 54.35 | 99.5 | 98.56 | 0.7 | 0.005 | 45.6 |
| Model 28 | VGG16 + HOG + random forest | 73.88 | 31.22 | 99.76 | 98.75 | 0.47 | 0.002 | 68.77 |
| Model 29 | VGG19 + HOG + logistic regression | 76.56 | 87.15 | 70.14 | 63.9 | 0.74 | 29.85 | 12.84 |
| Model 30 | VGG19 + HOG + linear SVM | 76.49 | 85.38 | 71 | 79.37 | 0.82 | 28.89 | 14.89 |
| Model 31 | VGG19 + HOG + RBF-SVM | 93.28 | 89.5 | 95.6 | 92.43 | 0.91 | 4.44 | 10.5 |
| Model 32 | VGG19 + HOG + random forest | 73.65 | 30.63 | 99.76 | 98.72 | 0.47 | 0.002 | 69.37 |
| Model 33 | ResNet50 + HOG + logistic regression | 82.9 | 89.72 | 78.77 | 71.94 | 0.79 | 21.2 | 10.27 |
| Model 34 | ResNet50 + HOG + linear SVM | 79.6 | 95.8 | 69.78 | 65.8 | 0.78 | 30.2 | 4 |
| Model 35 | ResNet50 + HOG + random forest | 88.13 | 84.78 | 90.16 | 83.95 | 0.84 | 9.8 | 15.2 |
| Model 36 | ResNet50 + HOG + RBF-SVM | 97.53 | 98.62 | 96.88 | 95.04 | 0.97 | 3.12 | 1.38 |
FPR: False-positive rate, FNR: False-negative rate, RBF: Radial basis function, SVM: Support vector machine, HOG: Histogram of oriented gradients
RESULTS AND DISCUSSION
The experiments have been carried out using NVIDIA Titan RTX GPU. For training the classifiers, 80% of data (benign nodule = 3635; malignant nodule = 1629) is used for training and 20% of data (benign nodule = 1629; malignant nodule = 407) is used for testing. The hyper-parameters of all the classifiers are tuned by the Gridsearch Cross Validation method.
Analysis of handcrafted features
Under handcrafted features-based lung nodule classification experiments, out of 12 models (Model 1 to Model 12), four models such as HOG with Logistic Regression (Model 9), HOG with Linear SVM (Model 10), GLCM with Logistic Regression (Model 1) and GLCM with Linear SVM (Model 2) can differentiate the benign lung nodules from malignant lung nodules. The rest of the models have become underfitting and are not able to identify the malignant class. The performance measures obtained through the experiments for these models are listed in Table 4. From Table 4, it is understood that even though many models got 62.24% of accuracy, they do not identify any malignant nodule. They have misclassified all the nodules as benign nodules and this led to maximum FNR (100%). It is noted that GLCM features misclassified benign nodules as malignant nodules. Therefore, for GLCM features, Model 1 and 2 have produced high FPR. Among the three feature descriptors, HOG features are better than GLCM and LBP features because they have produced less FNR. Model 11 (HOG+RBF-SVM) performed better in lung nodule classification which has got 78% of accuracy, 95.45% of sensitivity, 67% of specificity, 64% of precision, 0.77 of F1 score, 32.6% of FPR and 4.5% of FNR. However, in medical image analysis, both FNR and FPR should be low. The FPR of Model 11 is very high. It is observed that the handcrafted features are not able to classify the lung nodules more accurately.
Analysis of deep features
From the deep features-based experimentations, it is observed that ResNet50 + RBF-SVM (Model 23) has performed better when compared to other models (Model 13–24). The performance metrics of deep features in lung nodule classification is given in Table 5. Model 23 has achieved the accuracy of 83.06%, sensitivity of 95.06%, specificity of 75.28%, the precision of 70.42%, F1 score of 0.81, FPR of 24.2%, and FNR of 4.94%. It is noted that the accuracy of deep features is better than handcrafted features but FPR is not reduced much. To reduce FPR, the hybridized features and classification combinations are tested.
Analysis of hybridized features
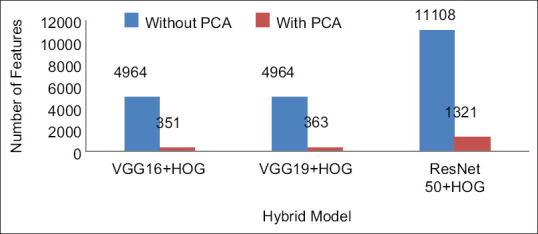
The dimension of the hybridized feature set is large. From the HOG feature descriptor, 2916 features are computed. The number of deep features from VGG16, VGG19, and ResNet50 models are 2048, 2048, and 8192, respectively. After feature concatenation, the dimension of the hybrid feature set is very large, and it is shown in Figure 5. If the classifiers are trained with such large dataset, the computation time would be large. To preserve the important features and reduce the computation time, PCA is employed. The number of hybridized features is reduced with the help of PCA and the reduced number of features is shown in Figure 5. Due to the reduced feature set, the computation time for classification has been reduced and it is shown in Figure 6.
Figure 5.
Feature reduction using principal component analysis
Figure 6.
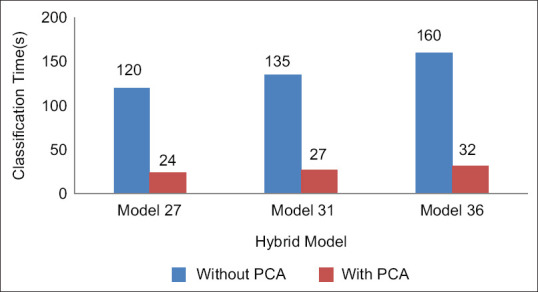
Computation time for hybrid models
After including HOG features with deep features, the performance of the classifiers has been improved in most of the cases as it is evident from Table 6. If the specificity of Model 27 and Model 28 is considered, the values are 99.5% and 99.76%, respectively, but they cannot be considered as the best model for lung nodule classification because their sensitivity values are 54.35% and 31.22% respectively. Hence they are not able to detect the malignant nodules and their FNR is very high, i.e. 45.6% and 68.77%, respectively.
Model 19 is a deep feature model where VGG19 features are analyzed using RBF-SVM classifier. This model has produced the accuracy of 83.06%, sensitivity of 90.12%, specificity of 78.78%, precision of 72.04%, F1 score of 0.8, FPR of 21.22% and FNR of 9.88%. Model 31 is a hybridized model where the hybridized features (VGG19 + HOG) are analyzed with RBF-SVM classifier which has produced the accuracy of 93.28%, sensitivity of 89.5%, specificity of 95.6%, precision of 92.43%, F1 score of 0.91, FPR of 4.44%, and FNR of 10.5%.
When Model 19 and Model 31 are compared, it has been inferred that after including HOG with VGG19 features the accuracy of Model 31 has got increased by 10.22% and FPR has got reduced by 16.78%.
The proposed Model 36 (ResNet50 + HOG with RBF-SVM classifier) has performed very well when compared to all other models (TP = 401; TN = 1581; FP = 51; FN = 6). The performance of Model 36 has outperformed in all the metrics with the accuracy of 97.53%, sensitivity of 98.62%, specificity of 96.88%, precision of 95.04%, F1 score of 0.97, FPR of 3.12%, and FNR of 1.38%. When Model 36 is compared with Model 23 (ResNet50 with RBF-SVM Classifier), it is found that the accuracy, sensitivity, specificity, precision and F1 score of Model 35 got improved by 14.47%, 3.56%, 21%, 24.62, and 0.16%, respectively. Moreover, FPR and FNR have got reduced by 21.08% and 6.58%, respectively. It is observed that the proposed model (model 36) can classify malignant nodules and benign nodules efficiently. The ROC curve of the proposed model is shown in Figure 7.
Figure 7.
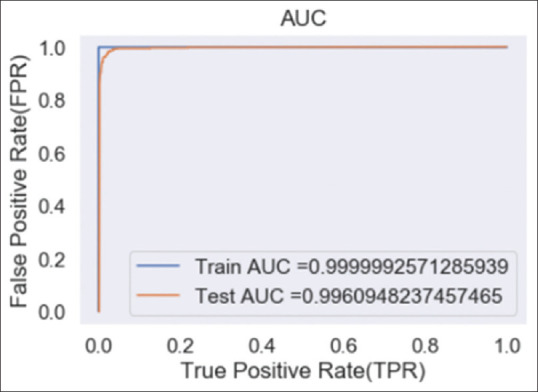
Receiver operating characteristic of the proposed hybrid model
Table 7 gives the comparison of the proposed method with other state-of-the-art methods. It can be observed that the proposed methodology has outperformed in lung nodule classification when compared to the related works. Li et al. have employed a hybridization technique which integrates handcrafted features such as intensity, geometric features and texture features with MC-CNN features and reported 88.58% of accuracy, 82.6% of sensitivity, 91.82% of specificity, 8.28% of FPR and 17.4% of FNR.[25]
Table 7.
Comparison of performance metrics with state-of-the-art methods
| Related works | Accuracy (%) | Sensitivity (%) | Specificity (%) | Precision (%) | F1 score | FPR (%) | FNR (%) | AUC |
|---|---|---|---|---|---|---|---|---|
| Proposed approach | 97.53 | 98.62 | 96.88 | 95.04 | 0.97 | 3.12 | 1.38 | 0.996 |
| Li et al.[25] | 88.58 | 82.60 | 91.82 | - | - | 8.28 | 17.4 | - |
| Wang et al.[30] | 91.75 | - | - | - | - | - | - | 0.970 |
| Nibali et al.[34] | 89.9. | 91.07 | 88.64 | 89.35 | - | - | - | 0.946 |
| da Nóbrega et al.[35] | 88.41 | 85.38 | - | 73.48 | 0.79 | - | - | 0.932 |
| Xie et al.[19] | 87.74 | 81.11 | 89.67 | - | - | - | - | 0.945 |
| Shen et al.[17] | 87.14 | 77 | 93 | - | - | - | - | 0.93 |
| de Carvalho et al.[36] | 92.63 | 90.7 | 93.47 | - | - | - | - | 0.934 |
| Kumar et al.[37] | 75.01 | 83.35 | - | - | - | - | - | - |
| Han et al.[15] | - | - | - | - | - | - | - | 0.927 |
| Dhara et al.[7] | - | 82.89 | 80.73 | - | - | - | - | 0.882 |
| Hussein et al.[38] | 91.26 | - | - | - | - | - | - | - |
FPR: False-positive rate, FNR: False-negative rate, AUC: Area under the curve
The proposed method is less complex than the method given by Shulong Li and gives improvement in accuracy by 8.95%, sensitivity by 16.02%, specificity by 4.8%. The FPR and FNR values are very less when compared to Shulong Li's approach. Wang et al. have fused LBP and HOG features with features from multichannel CNN's features.[30] The accuracy of Wang et al. is lesser than the proposed model by 5.78%. Antonio et al. have used topology-based phylogenetic diversity index and CNN for lung nodule classification. Their approach has given good results because segmentation of nodules has been done for the input images.[36] However in the proposed methodology without segmentation of nodules, the features have been extracted from the 2D nodule patches and the results are better than the methodology given by Antonio et al. with 4.9% improvement in accuracy, 7.92% improvement in sensitivity, and 3.41% improvement in specificity. The proposed methodology promises an agreeable balance between sensitivity and specificity and it implies that it is well balanced in classifying the malignant and benign nodules.
CONCLUSIONS
This research work proposes a deep hybridized model to classify the lung nodules into two different categories: malignant nodule and benign nodule. In clinical routine, there are a few complications in the classification of lung nodules due to visual representation of these nodules may appear similar. The proposed methodology enjoys the benefits of both ResNet50 based deep features and the handcrafted HOG features. Due to the hybridization of the features, the classifier can differentiate the malignant nodules and benign nodules with high accuracy of 97.53%, sensitivity of 98.62%, specificity of 96.88%, precision of 95.04%, F1 score of 0.97, FPR of 3.12%, and FNR of 1.38%. In addition, our proposed approach has been compared to handcrafted feature-based lung nodule classification and deep feature-based lung nodule classification models. The proposed approach outperforms well when it is compared to different deep learning models used in lung nodule classification. The future scope of this work will be focused on the hybridization of EfficientNet with HOG features.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
REFERENCES
- 1.Siegel R, Miller K, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30. doi: 10.3322/caac.21590. [DOI] [PubMed] [Google Scholar]
- 2.Wild CP, Weiderpass E, Stewart BW. World Cancer Report 2020. Lyon, France: International Agency for Research on Cancer, World Health Organization; 2020. [Google Scholar]
- 3.Kanavos P. The rising burden of cancer in the developing world. Ann Oncol. 2006;17(Suppl 8):viii15–23. doi: 10.1093/annonc/mdl983. [DOI] [PubMed] [Google Scholar]
- 4.Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 5.Saab MM, Kilty C, Noonan B, FitzGerald S, Collins A, Lyng Á, et al. Public health messaging and strategies to promote “SWIFT” lung cancer detection: A qualitative study among high-risk individuals. J Cancer Educ. 2020 doi: 10.1007/s13187-020-01916-w. Ahead of print. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Abraham J. Reduced lung cancer mortality with low-dose computed tomographic screening. Community Oncol. 2011;8:441–2. doi: 10.1056/NEJMoa1102873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dhara AK, Mukhopadhyay S, Dutta A, Garg M, Khandelwal N. A combination of shape and texture features for classification of pulmonary nodules in lung CT images. J Digit Imaging. 2016;29:466–75. doi: 10.1007/s10278-015-9857-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.McNitt-Gray MF, Hart EM, Wyckoff N, Sayre JW, Goldin JG, Aberle DR. A pattern classification approach to characterizing solitary pulmonary nodules imaged on high resolution CT: Preliminary results. Med Phys. 1999;26:880–8. doi: 10.1118/1.598603. [DOI] [PubMed] [Google Scholar]
- 9.Khan SA, Kenza K, Nazir M, Usman M. Proficient lung nodule detection and classification using machine learning techniques. J Intell Fuzzy Syst. 2015;28:905–17. [Google Scholar]
- 10.Suzuki K, Li F, Sone S, Doi K. Computer-aided diagnostic scheme for distinction between benign and malignant nodules in thoracic low-dose CT by use of massive training artificial neural network. IEEE Trans Med Imaging. 2005;24:1138–50. doi: 10.1109/TMI.2005.852048. [DOI] [PubMed] [Google Scholar]
- 11.Armato SG, 3rd, Altman MB, Wilkie J, Sone S, Li F, Doi K, et al. Automated lung nodule classification following automated nodule detection on CT: A serial approach. Med Phys. 2003;30:1188–97. doi: 10.1118/1.1573210. [DOI] [PubMed] [Google Scholar]
- 12.Way TW, Hadjiiski LM, Sahiner B, Chan HP, Cascade PN, Kazerooni EA, et al. Computer-aided diagnosis of pulmonary nodules on CT scans: Segmentation and classification using 3D active contours. Med Phys. 2006;33:2323–37. doi: 10.1118/1.2207129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Matsuki Y, Nakamura K, Watanabe H, Aoki T, Nakata H, Katsuragawa S, et al. Usefulness of an artificial neural network for differentiating benign from malignant pulmonary nodules on high-resolution CT: Evaluation with receiver operating characteristic analysis. AJR Am J Roentgenol. 2002;178:657–63. doi: 10.2214/ajr.178.3.1780657. [DOI] [PubMed] [Google Scholar]
- 14.Kuruvilla J, Gunavathi K. Lung cancer classification using neural networks for CT images. Comput Methods Programs Biomed. 2014;113:202–9. doi: 10.1016/j.cmpb.2013.10.011. [DOI] [PubMed] [Google Scholar]
- 15.Han F, Wang H, Zhang G, Han H, Song B, Li L, et al. Texture feature analysis for computer-aided diagnosis on pulmonary nodules. J Digit Imaging. 2015;28:99–115. doi: 10.1007/s10278-014-9718-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hua KL, Hsu CH, Hidayati SC, Cheng WH, Chen YJ. Computer-aided classification of lung nodules on computed tomography images via deep learning technique. Onco Targets Ther. 2015;8:2015–22. doi: 10.2147/OTT.S80733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Shen W, Zhou M, Yang F, Yu D, Dong D, Yang C, et al. Multi-crop convolutional neural networks for lung nodule malignancy suspiciousness classification. Pattern Recognit. 2017;61:663–73. [Google Scholar]
- 18.da Silva G, da Silva Neto O, Silva A, de Paiva A, Gattass M. Lung nodules diagnosis based on evolutionary convolutional neural network. Multimed Tools Appl. 2017;76:19039–55. [Google Scholar]
- 19.Xie Y, Zhang J, Xia Y, Fulham M, Zhang Y. Fusing texture, shape and deep model-learned information at decision level for automated classification of lung nodules on chest CT. Inf Fusion. 2018;42:102–10. [Google Scholar]
- 20.Nanni L, Ghidoni S, Brahnam S. Handcrafted vs.non-handcrafted features for computer vision classification. Pattern Recognit. 2017;71:158–72. [Google Scholar]
- 21.Nguyen DT, Pham TD, Baek NR, Park KR. Combining deep and handcrafted image features for presentation attack detection in face recognition systems using visible-light camera sensors. Sensors (Basel) 2018;18:699. doi: 10.3390/s18030699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Georgescu M, Ionescu R, Popescu M. Local learning with deep and handcrafted features for facial expression recognition. IEEE Access. 2019;7:64827–36. [Google Scholar]
- 23.Li X, Shen L, Shen M, Qiu C. Integrating handcrafted and deep features for optical coherence tomography based retinal disease classification. IEEE Access. 2019;7:33771–7. [Google Scholar]
- 24.Paul R, Hawkins SH, Balagurunathan Y, Schabath MB, Gillies RJ, Hall LO, et al. Deep feature transfer learning in combination with traditional features predicts survival among patients with lung adenocarcinoma. Tomography. 2016;2:388–95. doi: 10.18383/j.tom.2016.00211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li S, Xu P, Li B, Chen L, Zhou Z, Hao H, et al. Predicting lung nodule malignancies by combining deep convolutional neural network and handcrafted features. Phys Med Biol. 2019;64:175012. doi: 10.1088/1361-6560/ab326a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.LIDC-IDRI – The Cancer Imaging Archive (TCIA) Public Access-Cancer Imaging Archive Wiki; 2021. Doi.org. [Last accessed on 2021 Mar 14]. Available from: http://doi.org/10.7937/K9/TCIA.2015.LO9QL9SX .
- 27.Hancock MC, Magnan JF. Lung nodule malignancy classification using only radiologist-quantified image features as inputs to statistical learning algorithms: Probing the Lung Image Database Consortium dataset with two statistical learning methods. J Med Imaging (Bellingham) 2016;3:044504. doi: 10.1117/1.JMI.3.4.044504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bruntha PM, Alex Pandian I, Sam Abraham S. Classification of lung nodule using hybridized deep feature technique. J Inf Technol Manage. 2020;12:109–28. [Google Scholar]
- 29.Dalal N, Triggs B. IEEE Computer Society Conference on Computer Vision and Pattern Recognition. IEEE; 2005. Histograms of Oriented Gradients for Human Detection; pp. 886–93. [Google Scholar]
- 30.Wang H, Zhao T, Li LC, Pan H, Liu W, Gao H, et al. A hybrid CNN feature model for pulmonary nodule malignancy risk differentiation. J Xray Sci Technol. 2018;26:171–87. doi: 10.3233/XST-17302. [DOI] [PubMed] [Google Scholar]
- 31.He K, Zhang X, Ren S, Sun J. IEEE Conference on Computer Vision and Pattern Recognition. IEEE; 2016. Deep Residual Learning for Image Recognition; pp. 770–8. [Google Scholar]
- 32.Haralick R, Shanmugam K, Dinstein I. Textural features for image classification. IEEE Trans Syst Man Cybern. 1973;SMC-3:610–21. [Google Scholar]
- 33.Ojala T, Pietikäinen M, Harwood D. A comparative study of texture measures with classification based on featured distributions. Pattern Recognit. 1996;29:51–9. [Google Scholar]
- 34.Nibali A, He Z, Wollersheim D. Pulmonary nodule classification with deep residual networks. Int J Comput Assist Radiol Surg. 2017;12:1799–808. doi: 10.1007/s11548-017-1605-6. [DOI] [PubMed] [Google Scholar]
- 35.da Nóbrega R, Rebouças Filho P, Rodrigues M, da Silva S, Dourado Júnior C, de Albuquerque V. Lung nodule malignancy classification in chest computed tomography images using transfer learning and convolutional neural networks. Neural Comput Appl. 2018;32:11065–82. [Google Scholar]
- 36.de Carvalho Filho A, Silva A, de Paiva A, Nunes R, Gattass M. Classification of patterns of benignity and malignancy based on CT using topology-based phylogenetic diversity index and convolutional neural network. Pattern Recognit. 2018;81:200–12. [Google Scholar]
- 37.Kumar D, Wong A, Clausi D. 12th Conference on Computer and Robot Vision. IEEE; 2015. Lung Nodule Classification Using Deep Features in CT Images; pp. 133–8. [Google Scholar]
- 38.Hussein S, Cao K, Song Q, Bagci U. International Conference on Information Processing in Medical Imaging. Cham: Springer; 2017. Risk Stratification of Lung Nodules Using 3D CNN-Based Multi-Task Learning; pp. 249–60. [Google Scholar]