Abstract
Background
The deep-sea may be regarded as a hostile living environment, due to low temperature, high hydrostatic pressure, and limited food and light. Isopods, a species-rich group of crustaceans, are widely distributed across different environments including the deep sea and as such are a useful model for studying adaptation, migration, and speciation. Similar to other deep-sea organisms, giant isopods have larger body size than their shallow water relatives and have large stomachs and fat bodies presumably to store organic reserves. In order to shed light on the genetic basis of these large crustaceans adapting to the oligotrophic environment of deep-sea, the high-quality genome of a deep-sea giant isopod Bathynomus jamesi was sequenced and assembled.
Results
B. jamesi has a large genome of 5.89 Gb, representing the largest sequenced crustacean genome to date. Its large genome size is mainly attributable to the remarkable proliferation of transposable elements (84%), which may enable high genome plasticity for adaptive evolution. Unlike its relatives with small body size, B. jamesi has expanded gene families related to pathways of thyroid and insulin hormone signaling that potentially contribute to its large body size. Transcriptomic analysis showed that some expanded gene families related to glycolysis and vesicular transport were specifically expressed in its digestive organs. In addition, comparative genomics and gene expression analyses in six tissues suggested that B. jamesi has inefficient lipid degradation, low basal metabolic rate, and bulk food storage, suggesting giant isopods adopt a more efficient mechanism of nutrient absorption, storage, and utilization to provide sustained energy supply for their large body size.
Conclusions
Taken together, the giant isopod genome may provide a valuable resource for understanding body size evolution and adaptation mechanisms of macrobenthic organisms to deep-sea environments.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12915-022-01302-6.
Keywords: Giant isopod, Deep-sea, Genome assembly, Body size evolution, Oligotrophic adaptation
Background
The deep-sea environment is characterized by darkness, low temperature, high hydrostatic pressure, and lack of food. Despite this hostile environment, a growing number of deep-dwelling animals have been identified in this ecosystem, including worms, mollusks, fish, crustaceans, and so on [1, 2]. Crustaceans are one of the dominant invertebrates inhabiting deep-sea environment, and among them, some macrobenthos (e.g., giant isopods and amphipods) are specifically attractive as their body sizes are significantly larger than their shallow-water relatives [3]. Decoding genomes of these deep-sea species helps us understanding their unique adaptive mechanisms [4–8], whereas deep-sea crustaceans, including macrobenthos, lack relevant genome information.
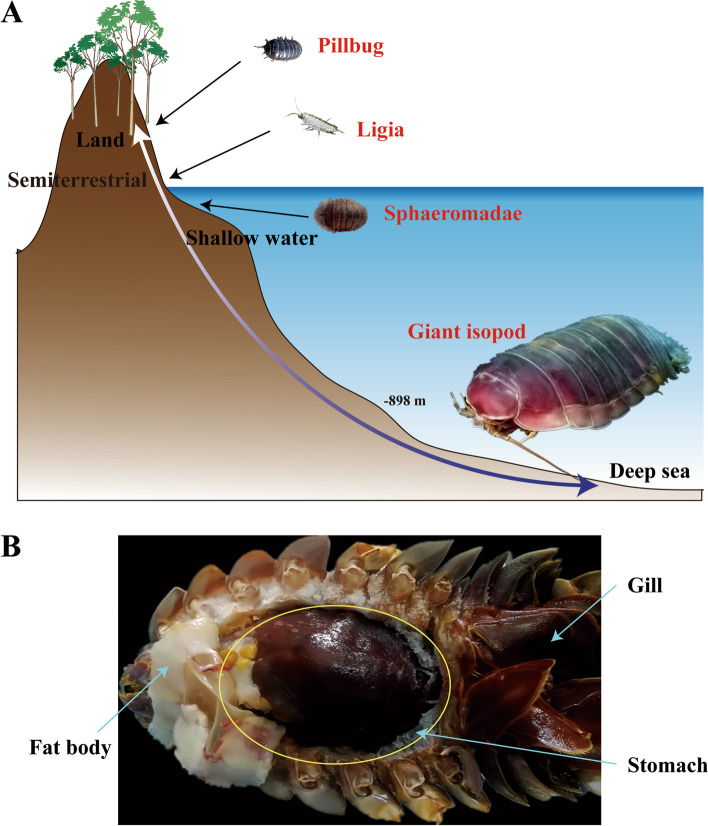
Isopods are a large group of crustaceans with more than 10,000 species described. So far, Isopoda is one of the limited groups widely distributed in various environments, as they have been found in all oceans at different depths (from intertidal zone to hadal zone), in fresh and brackish waters, and on land (Fig. 1A) [9]. Therefore, Isopoda is an ideal model for studying migration and speciation, especially for the shift from shallow water to deep sea, and from ocean to land. Notably, isopods are one of the most morphologically diverse groups of crustaceans. Its size ranges from 0.5 cm (dwarf species) to as big as 50 cm for giant isopods [10]. Consistent with the Cope-Bergmann's Rule, isopods from deep sea tend to be larger than their relatives in shallower waters [11]. As the largest extant animals on the planet are aquatic and many of them are deep-sea organisms, the impact of marine habitats and evolutionary adaptation on body size is mysterious and noteworthy [12]. Besides, body size has always been regarded as one of the most important quantitative traits in evolutionary scrutiny, which is strongly correlated with many physiological and fitness characters [13]. Thus, isopods provide an excellent model for studying the adaptive evolution of body size, whereas, even with a great number of species, only two isopods, Armadillidium vulgare and Armadillidium nasatum, have been sequenced so far, and they are both terrestrial [14, 15]. Genomics of marine isopods, especially deep-sea species, is far from being understood.
Fig. 1.
The distributions and phenotypes of isopods. A The distributions of various isopods from the land to deep-sea environments. B The morphology of the giant isopod, B. jamesi
Bathynomids (Crustacea: Isopoda: Cirolanidae) is regarded to be the “supergiant group” of isopods, which is well known for their big size [16, 17]. Bathynomids inhabit deep-sea benthic environment that are generally found on muddy bottoms at depths from 170 m to the dark of 2140 m [17, 18]. To adapt to the benthic environment, bathynomids adopt a burrowing behavior [19]. They have developed an oversized stomach, which can take approximately 2/3 of the whole body cavity when it fills up with food (Fig. 1B). In addition, midgut glands and adipocytes (collectively called “fat body”) are distributed throughout the body of bathynomids to store organic reserves [20]. Furthermore, bathynomids are well known for their extremely long fasting state (over 5 years), which should be the longest record to date [21]. Larger animals usually have greater absolute energy requirements [22]. However, the deep-sea conditions are harsh and food-limited, which seems to be unsuitable for the survival of giant animals [8, 23]. Therefore, a special efficient mechanism should be adopted by these supergiant isopods adapting to the deep-sea oligotrophic conditions.
Deep-sea expeditions provide excellent opportunities for us to uncover the unusual deep-sea creatures. During a recent expedition near Hainan Island in the northern South China Sea, a new deep-sea (a depth of 898 m) bathynomid species, Bathynomus jamesi Kou, Chen & Li, 2017, was collected and identified [24]. In this study, a high-quality genome assembly of B. jamesi was generated using PacBio sequencing technology. Analysis of the genomic characteristics identified potential factors related to the genome size evolution of B. jamesi. Based on the comparisons between the genomes of B. jamesi and its terrestrial relatives and other crustaceans, we have identified some expanded and positively selected gene families related to its body size evolution and deep-sea environment adaptation. This genome provides valuable resource for understanding evolutionary history of isopods and their deep-sea environmental adaptation mechanisms.
Results
Genome assembly and annotation
To estimate genome size of B. jamesi, a total of 235.25 Gb Illumina short reads were generated and utilized for genome survey analysis (Additional file 1 : Table S1). K-mer analysis indicated that the genome size of B. jamesi is approximately 5.24 Gb (Additional file 1: Fig. S1), which is larger than most crustacean genomes reported so far (generally < 2 Gb; Additional file 1: Table S2).
To assemble the genome of B. jamesi, 360.80 Gb PacBio long reads were generated and de novo assembled (Additional file 1: Table S1). The final assembly was 5.89 Gb in total length with a contig N50 length of 587.28 Kb, showing a higher continuity than the genomes of terrestrial isopod A. vulgare (contig N50 = 38.36 Kb) and many other crustaceans as well (Table 1) [14].
Table 1.
Summary of genome assembly and characteristics of B. jamesi and other three crustaceans
| Species | B. jamesi | A. vulgare | L. vannamei | E. sinensis |
|---|---|---|---|---|
| Genome size (bp) | 5,892,409,081 | 1,725,108,002 | 1,618,026,442 | 1,562,256,418 |
| Number of Contigs | 22,827 | 52,740 | 50,304 | 12,722 |
| Contig N50 (bp) | 587,279 | 38,359 | 57,650 | 26,045 |
| Contig N90 (bp) | 108,712 | 18,318 | 14,641 | 2,670 |
| Genome GC percent% | 37.28% | 29.15% | 35.68% | 46.39% |
| BUSCOs coverage (%) | 94.80% | 91.38% | 94.00% | 91.20% |
| Repeat percentage (%) | 85.32% | 69.54% | 49.39% | 45.30% |
| Gene number | 23,221 | 19,051 | 25,572 | 28,033 |
| Gene average length (bp) | 936 | 1259 | 1546 | 1078 |
| Exon number per gene | 4.18 | 4.93 | 5.94 | 3.26 |
| Exon average length (bp) | 223 | 181 | 260 | 330 |
| Intron average length (bp) | 3010 | 1872 | 1484 | 1602 |
This genome assembly displayed a high quality as assessed by the coverage of raw Illumina sequencing data (99.80%) and RNA-seq data (84.23%) (Additional file 1: Table S3). Besides, a high proportion of BUSCOs (94.98%) were covered by the B. jamesi genome, which is comparable to or better than many recent sequenced crustacean genomes (Table 1; Additional file 1: Fig. S2, Table S4) [14, 25–27].
A total of 23,221 protein-coding genes were predicted and annotated in the B. jamesi genome (Table 1; Additional file 1: Fig. S3). The average intron length of genes (3010 bp) was significantly longer than that of A. vulgare (1872 bp) and many other crustacean genomes with relative smaller genome sizes (Table 1). It is consistent with the view that intron size is positively correlated with genome size [28].
Repeats and genome size evolution
According to the Animal Genome Size Database (www.genomesize.com), C-value of isopods ranges from 1.71 to 8.82 pg, indicating there is a 5.2-fold variation of their genome sizes (Additional file 1: Table S5). B. jamesi has the largest genome (5.89 Gb) among sequenced crustacean genomes (Additional file 1: Table S2), which is about 3.4-fold larger than that of the A. vulgare genome (1.73 Gb) and approximately 49-fold of the clam shrimp Eulimnadia texana genome (0.12 Gb). Whole-genome duplication (WGD) has been identified to be one of the main factors causing genome expansion. However, only six syntenic blocks of paralogous genes were identified in the B. jamesi genome, which was far less than that of the horseshoe crab Tachypleus tridentatus (320 syntenic blocks), a species with WGD. In addition, Ks peak related to WGD and duplicated Hox gene cluster have not been identified in the B. jamesi genome (Additional file 1: Fig. S4). Therefore, B. jamesi appears to have not undergone WGD.
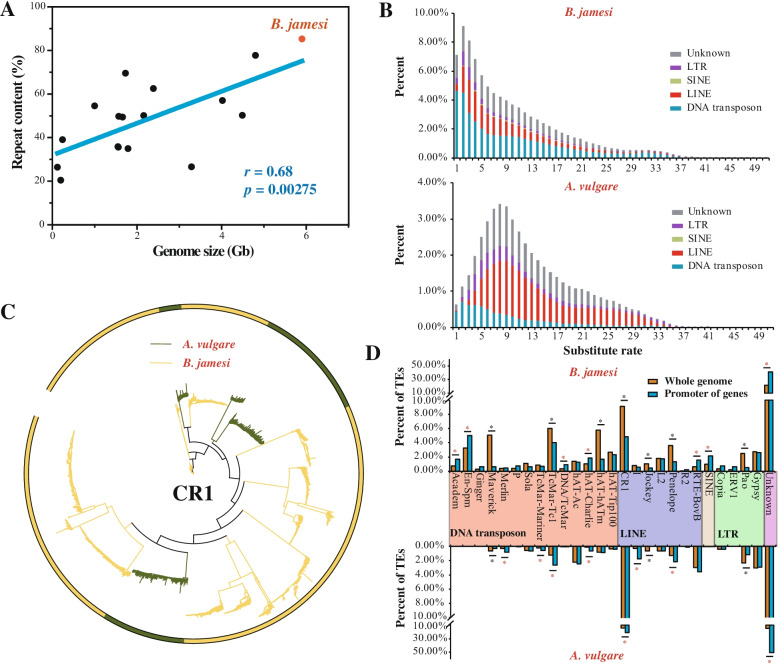
K-mer analysis indicated that 89.7% of the B. jamesi genome was composed of repetitive sequences. Consistently, based on the RepBase and a local repeat database that generated by RepeatModeler, a total of 5.03 Gb sequences (85.32%) were annotated as repeats, which were significantly more than those of any other crustaceans (generally < 60%, p < 0.05) (Table 2; Additional file 1: Table S2). A strong positive correlation between repeat content and genome size has been identified among crustacean genomes (r = 0.68, p = 0.00275, Pearson’s test) (Fig. 2A), suggesting that repeat proliferation might be the major driving force for the genome expansion of B. jamesi.
Table 2.
Comparison of the repeats among four crustaceans
| Repeats | B. jamesi | A. vulgare | L. vannamei | E. sinensis |
|---|---|---|---|---|
| Total length | 5.90 Gb | 1.73 Gb | 1.66 Gb | 1.56 Gb |
| Repeats | 85.32% | 69.54% | 49.39% | 35.57% |
| DNA | 35.99% | 7.08% | 9.33% | 2.30% |
| DNA/En-Spm | 3.28% | 0.00% | 6.39% | 0.82% |
| DNA/Maverick | 5.08% | 0.63% | 0.80% | 0.10% |
| DNA/Merlin | 0.37% | 0.28% | 0.00% | 0.01% |
| DNA/TcMar-Mariner | 0.87% | 0.21% | 0.06% | 0.00% |
| DNA/TcMar-Tc1 | 6.05% | 1.23% | 0.03% | 0.02% |
| DNA/hAT-Ac | 1.41% | 2.18% | 0.00% | 0.11% |
| DNA/hAT-Charlie | 1.04% | 0.11% | 1.00% | 0.09% |
| DNA/hAT-hATm | 5.77% | 0.81% | 0.00% | 0.00% |
| DNA/hAT-Tip100 | 2.67% | 0.36% | 0.00% | 0.00% |
| LINE | 19.36% | 20.24% | 2.82% | 9.72% |
| LINE/CR1 | 9.13% | 14.46% | 0.25% | 4.06% |
| LINE/Jockey | 1.06% | 0.63% | 0.06% | 0.05% |
| LINE/L2 | 1.80% | 0.62% | 0.35% | 0.36% |
| LINE/Penelope | 3.61% | 1.26% | 0.45% | 0.04% |
| LINE/RTE-BovB | 0.62% | 3.00% | 0.77% | 0.91% |
| SINE | 1.00% | 0.00% | 0.06% | 0.29% |
| LTR | 5.95% | 5.89% | 0.62% | 1.79% |
| LTR/ERV1 | 0.24% | 0.00% | 0.02% | 0.01% |
| LTR/Pao | 2.48% | 2.32% | 0.00% | 0.19% |
| LTR/Gypsy | 2.76% | 3.22% | 0.22% | 1.28% |
| Unknown | 21.97% | 14.87% | 3.42% | 10.39% |
| Satellite | 0.31% | 0.00% | 0.10% | 0.00% |
| Simple repeat | 0.65% | 18.08% | 23.93% | 6.90% |
| Low complexity | 0.01% | 3.57% | 9.49% | 2.04% |
Fig. 2.
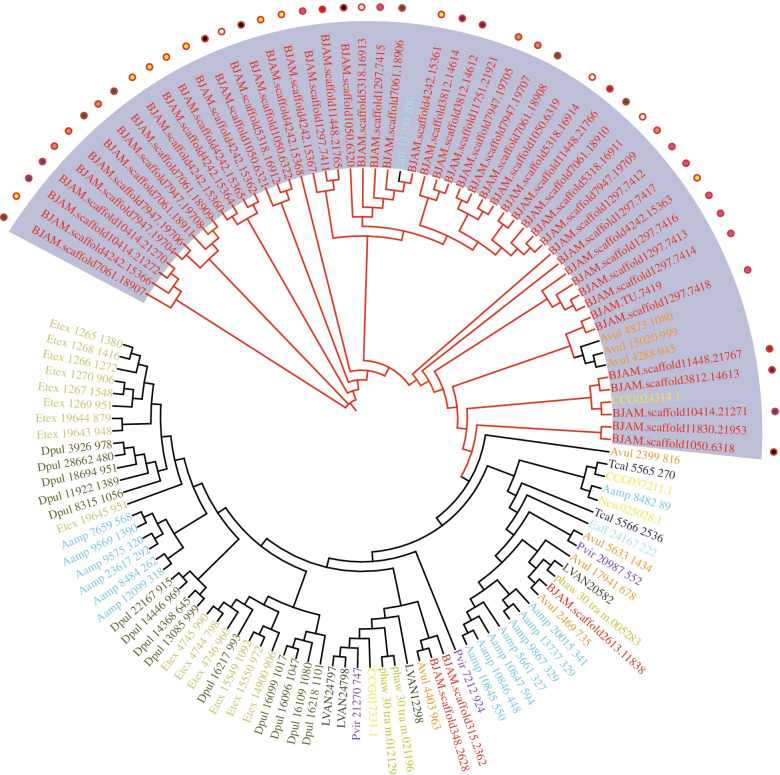
The evolution of transposable elements (TEs) and genome size. A The relationship between the genome size and repeat content. The repeat contents and genome sizes of the sequenced crustacean genomes were summarized in the Additional file 1: Table S2. The TE content and the genome size was positively correlated with the Pearson correlation r = 0.68 and p-value = 0.00275. B Kimura distance-based copy divergence analyses of TEs in the two isopod genomes, B. jamesi and A. vulgare. The graphs represent genome coverage for each TE superfamily in the different genomes analyzed. Clustering was performed according to their Kimura distances (K-value from 0 to 50). C Phylogenetic tree of the CR1 LINEs from B. jamesi (yellow) and A. vulgare (dark gray). D Enrichment analyses of TE families within gene promoters. The closest TE was calculated for each gene, and the content of the closest TEs were calculated and compared with that of the whole genome
Transposable elements (TEs) and simple sequence repeats (SSRs) accounted for 84.27% and 0.65% of the B. jamesi genome, respectively (Table 2). Different from B. jamesi, the A. vulgare genome contained significant less TEs (47.89%) and more SSRs (18.08%) that similar with the SSR-rich species [25]. TEs accounted for 98.77% of the total repeats of B. jamesi, indicating their prominent role in genome expansion. DNA transposons (35.99%), long interspersed nuclear elements (LINEs, 19.36%) and long terminal repeats (LTRs, 5.95%) were three major groups of TEs in the B. jamesi genome (Table 2). The proportions of LINEs and LTRs in the genome of B. jamesi was similar to its terrestrial relative A. vulgare. Among them, two typical LINEs (CR1 and Penelope) and two types of LTRs (Pao and Gypsy) showed apparent proliferation in the genomes of B. jamesi and A. vulgare. In contrast to LINE and LTR, DNA transposon was the most abundant TE (35.99%) in the B. jamesi genome, and its content was significantly higher than that of A. vulgare (7.08%, p < 0.05). Five types of DNA transposons, including TcMar-Tc1 (6.05%), hAT-hATm (5.77%), Maverick (5.08%), En-Spm (3.28%), and hAT-Tip100 (2.67%), are significantly expanded in the B. jamesi genome in comparison with A. vulgare (p < 0.05, Table 2).
To assess the evolutionary history of TE proliferation, we performed a divergence time estimation of TEs. More than 95% of TEs have a divergence rate of < 20%, indicating that most TEs in the B. jamesi genome are relatively young (Fig. 2B). The CR1-type LINE was the most abundant TE of both B. jamesi and A. vulgare, which accounted for 9.13% and 14.46% of the two genomes, respectively (Table 2). However, phylogenetic analysis of the total CR1-type LINEs of the two genomes indicated these TEs proliferated independently in the two isopods, rather than derived from their ancestor (Fig. 2C). In contrast to B. jamesi, CR1-type LINEs were relatively more ancient in A. vulgare with a divergence rate of > 20% (Additional file 1: Fig. S5). As the most abundant TEs (2.12 Gb) in the B. jamesi genome, DNA transposons were also proliferated in a recent time like CR1 (Additional file 1: Fig. S5). Therefore, DNA transposons and CR1-type LINEs are the two major types of TEs that contribute to the genome expansion of B. jamesi, and this proliferation event should have occurred in a relative recent time.
TE proliferation can promote genome plasticity by altering genome structure or regulating gene expression. Previous studies suggested that TEs enriched in the promoters of genes play important roles in regulating gene expressions in response to different stresses [29]. Thus, we next analyzed the gene-surrounding TEs to investigate their potential functions. Different from the previous report that TEs are usually enriched in upstream and downstream of genes immediately (within 2 Kb) [29], TEs in the genome of B. jamesi were uniformly distributed surrounding genes (up- or down-stream of 10 Kb), especially for LINEs, LTRs and Maverick of DNA transposons (Additional file 1: Fig. S6). Exceptionally, TcMar, En-Spm, and hAT of DNA transposon and SINEs showed a relative slight enrichment surrounding genes (within 2 Kb). When analyzing the neighboring TEs of total genes, it was interesting to find that although many types of TEs (e.g., Maverick, TcMar-Tc1, hAT-hATm, CR1, Penelope, and Pao) proliferated significantly in the B. jamesi genome, they were less distributed surrounding genes than in other genomic regions (p < 0.05, Fig. 2D). In contrast, some TEs with lower abundance were significantly enriched in the promoters of genes, including Academ, En-Spm, TcMar-Tigger, hAT-Charlie, RTE-BovB, and SINE. Therefore, unlike the findings of the previous study [29], our results suggest the significant proliferation of TEs should perform a more profound impact on the plasticity of the whole genome than on the architecture of protein-coding genes in B. jamesi.
Comparative genomics
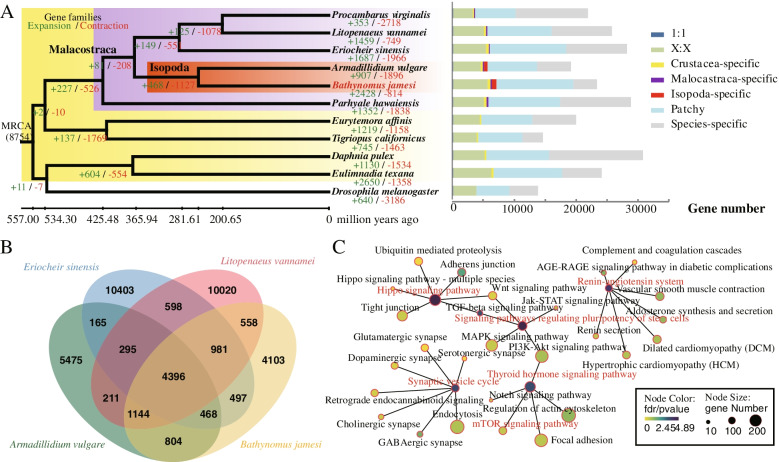
Based on 177 orthologous single-copy genes, a phylogenetic tree was constructed to confirm the placement of B. jamesi (Fig. 3A). As expected, the two isopods (B. jamesi and A. vulgare) were clustered together and then nested by the other four malacostraceans. Isopods were estimated to be diverged from their last common ancestor around 366 million years ago (Mya), which is a time of the Late Devonian-Epoch. The deep-sea isopod (B. jamesi) and the terrestrial isopod (A. vulgare) were estimated to divergent around 245 Mya, which is consistent with the fossil records of Oniscidea (219.6–358.9 Mya) [30]. Besides, fossil record showing that another deep-sea isopod Bathynomus giganteus has emerged as early as 160 Mya [2]. Therefore, the deep-sea bathynomids should be originated between 160 and 245 Mya.
Fig. 3.
Comparative genomes analyses of B. jamesi and its relatives. A Phylogenetic tree and divergence times of B. jamesi and other arthropods. The number of significantly expanded (+, green) and contracted (−, red) gene families is designated on each branch. B Number of gene families shared among four Malacostraca species shown as a Venn diagram. C KEGG enrichment analysis of the expanded gene families of B. jamesi. The enrichment analysis was performed by using the toolkit from Omicshare (https://www.omicshare.com/). The enriched KEGG terms was referred to the Additional file 1: Fig. S7
Comparative genomics analysis among 11 arthropod species identified 14,376 gene families. Among them, 418 gene families were isopod-specific (Fig. 3A), and 3683 gene families were specific in B. jamesi (Fig. 3B; Additional file 1: Table S6). Besides, a total of 274 significantly expanded gene families and 157 contracted families were identified in the B. jamesi genome (p < 0.05; Fig. 3A; Additional file 1: Table S7). These expanded gene families were functionally enriched in gene ontology (GO) terms related to membrane, peptidase activity, ion binding, proteolysis, and signal transduction (Additional file 1: Table S8). KEGG analysis significantly linked some of the expanded genes to Hippo signaling pathway, synaptic vesicle cycle, lipid metabolism (e.g., ether lipid metabolism and glycerophospholipid metabolism), and endocrine systems (e.g., renin-angiotensin system, insulin signaling pathway, and thyroid hormone signaling pathway) (Fig. 3C; Additional file 1: Fig. S7). The expansion of these gene families may reflect the adaptive evolution of B. jamesi to the deep-sea environment. Individual gene families related to body size evolution and deep-sea adaptation were discussed in greater depth in the later sections.
Strengthened pathways related to large body size
B. jamesi is a giant isopod with a body length of > 20 cm, which is significantly larger than its intertidal and terrestrial relatives, e.g., sea roaches and pill bugs (generally < 3 cm). Comparative genomics approach helps us discover the genetic characteristics associated with the body size evolution of giant isopods.
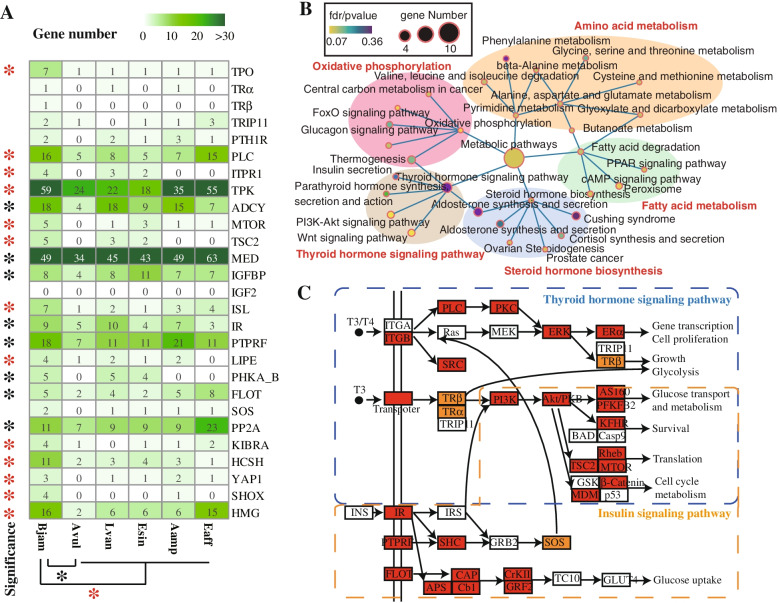
As indicated from the above comparative genomic analysis, the expanded gene families of B. jamesi were significantly enriched in the thyroid hormone signaling pathway (p = 3.95E−06) (Fig. 3C; Additional file 1: Fig. S7), which is an important pathway in regulating growth, development and metabolism [31]. Many gene families related to thyroid hormone (TH) synthesis and secretion were significantly expanded and tandemly duplicated in the B. jamesi genome (Fig. 4A), including phosphatidylinositol phospholipase C (PLC), inositol 1,4,5-triphosphate receptor type 1 (ITPR1), tyrosine-protein kinase (TPK), adenylate cyclase (ADCY), serine/threonine-protein kinase mTOR (MTOR), tuberous sclerosis 2 (TSC2), and mediator of RNA polymerase II transcription subunit (MED). TH signaling is regarded as a key modulator of fundamental biological processes that has been evolutionarily conserved in both vertebrate and invertebrate species. Thyroid peroxidase (TPO), thyroid hormone receptor α (TRα) and β (TRβ), and thyroid receptor-interacting protein 11 (TRIP11) are four key enzymes in TH biosynthesis and signaling transduction. Seven TPO genes, one TRα gene, one TRβ gene, and two TRIP11 genes were identified in the B. jamesi genome, indicating the presence of endogenous TH in this deep-sea organism. In contrast, only a single gene encoding TPO and TRIP11 was identified in the A. vulgare genome, with the lack of TRα and TRβ genes. Likewise, the loss of TRα and/or TRβ genes has also been found in other crustaceans (Fig. 4A). In addition to the gene family expansion, the thyroid hormone signaling pathway has been identified to be under strong positive selection (six positively selected genes, KEGG enrichment p value = 9.93E−03) (Fig. 4B). Thus, in contrast to other crustaceans, B. jamesi has a complete thyroid hormone signaling pathway, which has been strengthened in the case that many genes of this pathway were significantly expanded and positively selected (Fig. 4C).
Fig. 4.
Gene expansion in growth-related hormone signaling pathways. A Heatmap of the number of genes involved in thyroid and insulin hormone signaling pathways among six crustaceans. The included species are B. jamesi (Bjam), A. vulgare (Avul), A. amphitrite (Aamp), E. affinis (Eaff), L. vannamei (Lvan), and E. sinensis (Esin). The red star indicates the correspondent gene of B. jamesi is significantly more than that of the other five crustaceans, and the black star indicates correspondent gene of B. jamesi is significantly more than that of A. vulgare but similar to other crustaceans (p < 0.05). B KEGG enrichment of the positively selected genes in B. jamesi. C The strengthened thyroid and insulin hormone signaling pathways. Genes with red background indicates significantly expanded genes of B. jamesi in comparison to A. vulgare; Genes with orange background indicates these genes are present in B. jamesi but absent in A. vulgare
The insulin signaling is another hormone pathway that plays an important role in growth and development. In the B. jamesi genome, the insulin signaling pathway was also under significant enrichment of expanded gene families (p = 4.5E−02). A set of common genes involved in the insulin signaling pathway were identified to be tightly associated with body size evolution, including insulin growth factor (IGF), insulin-like growth factor-binding protein (IGFBP), insulin enhancer protein (ISL), and IGF receptor (IR) [32]. IGFs are normally bound to IGFBPs in great affinities that higher than IRs, and IGFBPs function as modulators of IGF availability and activity [33]. ISL is a LIM-homeodomain transcription factor involved in insulin secretion and metabolism, and also mediates glycolysis [34]. Like other crustaceans, IGF has not been identified in the B. jamesi genome, but IGFBP, ISL, and IR were all present in these crustaceans, indicating the presence of insulin signaling pathway. In contrast to A. vulgare, IGFBP complex acid labile subunit (IGFBP-ALS), ISL and IR were all significantly expanded in the B. jamesi genome (Fig. 4A). Apart from these genes, many other genes involved in this pathway were also expanded in B. jamesi, including receptor-type tyrosine-protein phosphatase F (PTPRF), hormone-sensitive lipase (LIPE), phosphorylase kinase alpha/beta subunit (PHKA_B), flotillin (FLOT), and MTOR (Fig. 4A). MTOR is the core gene of mTOR signaling pathway, which is also an important pathway in regulating animal growth and body size. This pathway locates at the downstream of the insulin signaling and thyroid hormone signaling pathway, and controls cell growth and metabolism in response to nutrients, growth factors, and cellular energy [35]. There are five genes encoding MTOR in the B. jamesi genome, which was significantly more than other crustaceans (Fig. 4A). In addition, there are four genes of the insulin signaling pathway (INPP5B, PRKCI, PRKAG, and RHEB) under positive selection in the B. jamesi genome (Fig. 4B). Therefore, similar to the thyroid hormone signaling pathway, the insulin signaling pathway of B. jamesi should also have been strengthened (Fig. 4C).
In addition to the two hormone signaling pathways, the Hippo signaling pathway was also significantly enriched by expanded gene families of B. jamesi (p = 3.44E−08), which may make some contributions to the huge stomach and fat body of B. jamesi, because this pathway is functional important in controlling organ size [36]. Compared with A. vulgare and other crustaceans, many key genes in the Hippo signaling pathway, including dachsous (HCSH, 11 members), Protein Kibra (KIBRA, 4 members), transcriptional coactivator YAP1 (YAP1, 3 members), and serine/threonine-protein phosphatase 2A (PP2A, 11 members), were significantly expanded in the B. jamesi genome (Fig. 4A). Besides these genes, many other genes involved in body size were also expanded in B. jamesi, including short-stature homeobox protein (SHOX) and high mobility group protein (HMG) (Fig. 4A). The deficiency of these two genes (SHOX and HMG) would result in dwarfism [37]. Taken together, these strengthened growth-related signaling pathways may make great contribution to the large body size of B. jamesi.
Molecular mechanisms underpinning deep-sea oligotrophic adaptation
To adapt to the deep-sea oligotrophic environments, the mechanisms of food storage and utilization of giant isopods should have undergone strong selective pressure. In accordance, giant isopods have developed a huge stomach to store food and can survive from an extraordinary long fasting state (> 5 years) (Fig. 1B) [21].
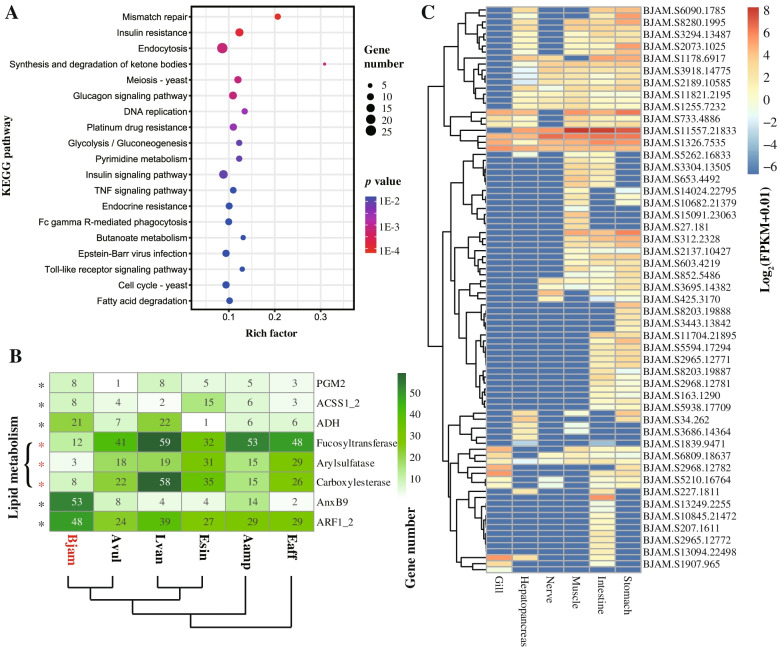
In order to identify potential genes related to nutrient storage, absorption, and utilization, RNA-seq analysis was performed on six tissues of B. jamesi. A total of 901 genes were identified to be specifically highly expressed in digestive organs, including stomach and intestine. These differently expressed genes enriched in the pathways of mismatch repair, insulin signaling and resistance, endocytosis, glycolysis, and so on (Fig. 5A). Glycolysis is an important metabolic process in which glucose is broken down to produce energy. Genes involved in the glycolysis pathway were mostly highly expressed in the stomach, intestine, and muscle of B. jamesi (Additional file 1: Fig. S8). Among them, phosphoglucomutase-2 (PGM2) is a transferase that plays an important role in carbohydrate metabolism of both glycogenolysis and glyconeogenesis [38]. Eight genes encoding PGM2 were identified in the B. jamesi genome, whereas only one PGM2 gene was found in the A. vulgare genome (Fig. 5B). Besides, these genes were tandemly duplicated on the scaffold281 and scaffold7261 of the B. jamesi genome, and they were mostly high expressed in stomach and intestine. Similar results were also identified in the genes encoding acetyl-CoA synthetase (ACSS1_2) and alcohol dehydrogenase (ADH), both of which participate in the TCA cycle for ATP production. A total of eight ACSS1_2s and 21 ADHs were identified in the B. jamesi genome, which were significantly more than that of A. vulgare (four ACSS1_2 and seven ADHs), and these genes were also highly expressed in the stomach and intestine. Therefore, B. jamesi may adopt an efficient mechanism of glycolysis to provide sufficient energy for its biological activities.
Fig. 5.
The differential gene expressions in six tissues of B. jamesi. A KEGG enrichment analysis of the highly expressed genes in stomach and intestine. The top 20 significantly enriched KEGG terms were displayed in the plot. B Heatmap of the number of genes involved in glycolysis, lipid metabolism and endocytosis among six crustaceans. The included species are B. jamesi (Bjam), A. vulgare (Avul), A. amphitrite (Aamp), E. affinis (Eaff), L. vannamei (Lvan), and E. sinensis (Esin). The red star indicates the correspondent gene of B. jamesi is significantly less than that of the other five crustaceans, and the black star indicates correspondent gene of B. jamesi is significantly more than that of the other five crustaceans (p < 0.05). C Expression level of the genes involved in the endocytosis of B. jamesi
Since giant isopods developed fat body to store organic reserves [20], the lipid synthesis and metabolism may under natural selection. However, out of our expectation, none of gene families related to lipid or fatty acid synthesis showed any signatures of expansion or under positive selection. Instead, some gene families related to lipid degradation were significantly contracted in the B. jamesi genome, including genes encoding fucosyltransferase, arylsulfatase, and carboxylesterase (Fig. 5B). These proteins are supposed to function in degrading glycolipids, sphingolipid and many esters. In addition, two genes related to fatty acid degradation (hcaD and echA) were under positive selection (Fig. 4B). Therefore, the lipid accumulation in the fat body should result from low efficiency of lipid degradation rather than high efficiency of lipid synthesis.
Beside energy production, the molecule transportation is also important for the absorption and utilization of food. Vesicular transport is an important process of transporting macromolecules through membrane, which has been identified to be under strong natural selection in deep-sea crustaceans [39]. Endocytosis is an essential process of vesicular transport, which actively transports molecules into cell by engulfing it with its membrane. The pathway of endocytosis was significantly enriched by differentially expressed genes (p = 1.8E-03), and a large number of them were specifically expressed in the stomach and intestine (Fig. 5C). Besides, some expanded gene families were identified to be involved in vesicular transport, and annexin B9 (AnxB9) was a representative one among them. AnxB9 is a functional protein involved in the formation of multivesicular bodies and regulation of protein trafficking, and even in stabilizing the endomembrane system during stress [40]. A total of 53 genes encoding AnxB9 were identified in the B. jamesi genome, which were significantly higher than in A. vulgare (eight genes) and other crustaceans (seven genes on average). These AnxB9 genes were mostly tandem duplicated in the B. jamesi genome (Fig. 6), and some of them were highly expressed in stomach, intestine, and muscle. Therefore, the expansion of gene families and their specific expression in digestive organs play an important role in the energy supply of giant isopod and help these organisms adapt to the oligotrophic conditions of the deep-sea environment.
Fig. 6.
Phylogenetic tree of the genes encoding AnxB9. The AnxB9 genes from various crustaceans were used for the tree construction, which labeled in various colors. A cluster of AnxB9 genes was specific expanded in the B. jamesi genome (gray background), and these genes were tandem duplicated in the genome. The circles with different colors indicate the genes located on different scaffolds
Discussion
With the development of deep-sea diving and genome-sequencing technologies, a growing number of deep-sea organisms have been discovered, collected, and identified, and their genomes have been sequenced [4–8]. Genome sequencing sheds lights on variable adaptive mechanisms of these species to the deep-sea environment. However, no deep-sea crustacean genome has been reported so far even though Crustacea comprises one of the dominant groups of deep-sea organisms. In this study, we reported the first deep-sea crustacean genome and analyzed its genome characteristics, which provides a valuable resource for studying the unique mechanisms by which crustaceans adapt to deep-sea ecosystems.
A large genome size is one of the most apparent characteristics shared by the genomes of B. jamesi and some other deep-sea species [5, 6]. Previous studies suggested that genome size tend to be larger in deeper conditions as driven by low temperature and high hydrostatic pressures from deep-sea environment [3, 41]. Indeed, besides B. jamesi, large genome size has also been detected in some other deep-sea crustaceans (e.g., amphipod Ampelisca macrocephola: ~ 63 Gb) and polar crustaceans (e.g., Antarctic krill Euphausia superba: ~ 42 Gb) [3, 42, 43]. However, this rule seems inapplicable for isopods because some shallow-water species (e.g., Sphaeroma destructor, 6.79 pg), terrestrial species (e.g., Oniscus asellus, 8.60 pg) and parasitic species (e.g., Nerocila munda, 8.82 pg) also have large genome sizes (Additional file 1: Table S5). Even among sympatric pairs of deep-sea species there is a large variation in their genome sizes, suggesting an absence of a dominant environment variable influencing genome size [3, 41]. However, as summarized in previous studies [44, 45], a rough generalization indicated that marine and terrestrial isopods have larger genomes overall versus freshwater species, and groundwater isopods have larger genomes than their surface-water relatives. Thus, it is likely that larger genomes are specific to certain families or genera or groups that inhabit similar conditions, e.g., deep-sea bathynomids. However, more evidences of the genome size studies, especially on the deep-sea taxa, is needed to support our hypothesis.
The selective pressure from deep-sea may be a candidate factor in shaping the large genomes of bathynomids. In addition to deep-sea habitat, there are many other factors showed possible relationships with genome size, including body size and life-history strategy [3, 41, 45]. Indeed, a relationship between genome size and body size has been identified in amphipods, but it is limited in giant organisms [3]. Thus, bathynomids may also apply a similar rule with giant amphipods because it is also a group of organisms displays size gigantism. Besides, the life-history strategy, including low basal metabolic rate (BMR), low mobility, and high longevity, are presumed to be positively correlated with genome size [44]. As expected, these traits have been commonly identified in deep-sea bathynomids [46]. Thus, the large genome size of bathynomids may tightly associate with their ecological habitats, body sizes and life-history strategies. Furthermore, the body size and life-history strategy of bathynomids also displayed close relationships with their deep-sea habitats. Firstly, their body sizes showed a positive correlation relationship with the depths of their habitats (Additional file 1: Fig. S9). According to the Bergmann’s rule, organisms inhabiting higher latitudes tend to have larger body sizes; this would, in turn, also be applicable to the deep-sea organisms [3, 47]. Indeed, many deep-sea crustaceans, including giant squids, giant sea spiders, giant isopods, and amphipods, are significantly larger than their shallow-water relatives [3]. Thus, the large body size of these crustaceans may be tightly associated with their deep-sea habitats. Besides, the low BMR is a well-known characteristic of gigantic animals living in places with limited feeding resources, such as the deep-sea oligotrophic environment [48]. Therefore, the factors driving genome size evolution of bathynomids appear to be complex, which may be the result of a combination of deep-sea habitat, body size and life-history strategy.
The deep-sea environmental stress can disrupt the epigenetic control of TEs leading to TE proliferation and increased genome size [49]. TE proliferation has long been considered to be the major cause of the large genome size of many species, which is also a consequence of the genome of B. jamesi and some other deep-sea species (e.g., the deep-sea clam Archivesica marissinica and tubeworm Paraescarpia echinospica) [5, 6]. Notably, B. jamesi has the highest content of TEs (84.27%) among sequenced crustacean genomes, and it is also significantly higher than that of A. marissinica (55.10%) and P. echinospica (42.20%), making its genome also larger than these two deep-sea species (1.52 Gb and 1.09 Gb, respectively). Therefore, in contrast to these two species, TEs of B. jamesi should be more active during the evolutionary history. Similar to the genomes of A. marissinica and P. echinospica, DNA transposons and LINEs were also the two major components of TEs in the B. jamesi genome, suggesting these TEs are adaptively selected and highly dynamic in deep-sea genomes. In consistent, our results suggested DNA transposons and LINEs were intensively proliferated in a relative recent time in the B. jamesi genome. The large amount of DNA insertion or deletion would result in a high genome plasticity [50]. As a consequence, the proliferation of TEs has profound effects on genome size, structure, stability and plasticity, and finally contributes to adaptive evolution [41, 51]. Therefore, the convergent proliferation of DNA transposons and LINEs in deep-sea species might play an important role in shaping highly plastic genomes and helping them adapt to the deep-sea environment. In addition, the highly dynamics of TEs may be beneficial for B. jamesi adapting deep-sea environment better.
The enormous size of giant isopods is a classic example of a phenomenon known as deep-sea gigantism. By analyzing the genome of B. jamesi, we get a glimpse of the molecular basis of its giant size. The growth-related hormone signaling pathways, including thyroid and insulin signaling pathways, play important roles in regulating growth and body size. Thyroid hormone influences growth in part by altering the secretion and effects of growth hormone. Growth hormone, in turn, mediates its effects by regulating the synthesis and secretion of insulin-like growth factor-I (IGF-I) [52]. The cooperation of these hormone pathways can cause gigantism when excess growth hormone leads to increased linear growth [53]. In this study, strengthened thyroid and insulin hormone signaling pathways were identified in the B. jamesi genome, which might functionally contribute to its gigantism. The mTOR pathway is the primary cell-autonomous nutrient sensor, while circulating hormones such as IGF-I and thyroid hormone are the main systemic regulators of growth and maturation in animals [31, 34, 35]. In addition, proper organ growth is also monitored and coordinated with whole-body growth through modulation of Hippo signaling [36]. Many genes participating in these pathways have been expanded in the B. jamesi genome, resulting for a strengthened network of growth-related pathways. The large body size of B. jamesi could be explained by the cooperation of these strengthened pathways.
According to previous studies, the body size has been identified to be strongly correlated with the BMR, duration of juvenile growth and longevity [12]. In consistent, our anatomic result showed that B. jamesi has a huge stomach to store food such that the fulfilled stomach occupied about 2/3 of the body. It also has a large amount of fat body spreading all over the body cavity to store organic reserves. Besides, it is likely that B. jamesi also has an extraordinary long fasting state as the starvation record (> 5 years) is kept by the congeneric species B. giganteus [21]. Furthermore, the low BMR, slow growth, high longevity have been observed in giant isopods, which may be a strategy for the survival of these large organisms [46]. Therefore, in correlation with large body size, B. jamesi has bulk food storage, low BMR, slower growth, and greater longevity.
Generally, organisms with large body sizes have greater absolute energy requirements [22], whereas food is definitely a limiting factor in the deep sea, for less food being available in deeper water and at greater distances from shore, which seems unsuitable for the survival of large animals. Thus, there is a debate about the adaptive evolution of deep-sea large animals and especially its possible causes. Large-sized body would be beneficial for competition and predation, whereas more energy will be costed to sustain the large-sized body. Kleiber’s Rule states that “larger animals are more efficient” [54, 55]. In the deep sea, the smaller surface area to volume ratio gives the giant isopods the advantage of conserving greater energy, with less energy lost to the surroundings through heat. Yet, the disadvantage of gigantism is that the absolute energy demand is large. It will be much more difficult to obtain adequate energy in the deep-sea environment, where food is usually an extremely scarce resource. To achieve a balance, it is likely that B. jamesi has adopted a sequence of survival strategies including low BMR, specialized energy storage organs, and efficient nutrient absorption and utilization. In this study, our results suggest B. jamesi have developed an efficient pathway for glycolysis and vesicular transport, which will contribute to its nutrient absorption and utilization. In addition, B. jamesi has low efficient of lipid degradation to support its lipid accumulation in fat body. Therefore, in contrast to small animals, an efficient mechanism of nutrient storage, absorption and utilization could be more important for the macrobenthos to adapt the deep-sea oligotrophic environment.
Conclusions
The genome of a deep-sea giant isopod B. jamesi was successfully assembled, representing the first high-quality genome of deep-sea crustaceans. Comparative genomic analyses provided new insights into the evolution of genome size and body size of animals and the adaptive mechanisms to the deep-sea extreme environments. The isopod genomes will shed lights on the habitat shift and evolution history of the crustaceans inhabiting deep-sea, shallow water, intertidal zone, and land. Furthermore, the genomic resources also provide powerful tools for broader studies on the ecology, evolutionary biology, and biological conservation of isopods.
Methods
Sampling and sequencing
The specimens of B. jamesi were collected by a deep-sea lander at a depth of 898 m near Hainan Island, in the northern South China Sea (17° 46.845′ N, 110° 38.217′ E). The specimens were identified as the species B. jamesi and kept in 75% ethanol and − 80 °C freezer [24]. The muscle of the legs of B. jamesi was collected for DNA extraction and genome sequencing. Total genomic DNA was extracted using TIANamp Marine Animal DNA Kits (Tiangen, Beijing, China) and used for Illumina and PacBio sequencing.
For Illumina sequencing, paired-end libraries with short insert size (350 bp) were constructed according to the instructions of the Illumina library preparation kit (Illumina, San Diego, USA). The constructed libraries were sequenced on an Illumina HiseqX-ten sequencing platform (Illumina, San Diego, USA). The raw sequencing reads were trimmed for quality subsequently using Trimmomatic v.0.35 (http://www.usadellab.org/cms/index.php?page=trimmomatic), and the retained clean reads were used for subsequent analyses.
For PacBio sequencing, genomic DNA was sheared to ~ 20 Kb, and the short fragments below the size of 10 Kb were filtered out using BluePippin (Sage Science, Beverly, USA). Filtered DNA was then used for the construction of the proprietary SMRTbell library using PacBio DNA Template Preparation Kit. SMRTbell libraries were used for single-molecule real time (SMRT) sequencing using the P6C5 sequencing chemistry (Pacific Biosciences, San Diego, USA), and then sequenced on the PacBio RSII sequencing platform (Pacific Biosciences, San Diego, USA).
RNA extraction and sequencing
In order to perform gene annotation and identification of tissue-specific expression genes, transcriptome sequencing was performed on six tissues of B. jamesi, namely gill, hepatopancreas, muscle, stomach, intestine, and nerve. According to the standard manufacturer’s protocol, total RNA was isolated and purified from each tissue using TRIzol extraction reagent (Thermo Fisher Scientific, Waltham, USA). RNA quality was determined by 1% agarose gel electrophoresis, and RNA concentration was assessed using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, USA). Transcriptome libraries were prepared according to the instructions of the TruSeq RNA Library Prep Kit (Illumina, San Diego, USA), and then sequenced on the Illumina HiSeq 2500 platform. The transcriptome reads were mapped to the genome using TopHat v1.2.1 [56]. Then, fragments per kilobase of transcript per million fragments mapped (FPKM) was calculated using Cufflinks v2.2.1 (http://cole-trapnell-lab.github.io/cufflinks/). The differential gene expression analysis was conducted using edgeR V3.10 [57].
Genome size estimation
Genome size of B. jamesi was estimated by K-mer analysis, which is widely used for the estimation of genome size and repeat content. Jellyfish was used to calculate K-mer frequencies based on the high-quality reads from the Illumina sequencing data [58]. A K-mer depth distribution was plotted and the peak depth could be identified. The genome size was estimated as the ratio of the total number of K-mers to the peak depth.
Genome assembly and quality assessment
The B. jamesi genome was de novo assembled based on PacBio subreads using FALCON pipeline (https://github.com/PacificBiosciences/FALCON/) with default parameters. The assembled sequences were then polished using Quiver (SMRT Analysis v2.3.0) based on the alignments of PacBio reads to the assembly. Besides, in order to make the genome assembly more accurate, five rounds of iterative error correction were performed using the aforementioned Illumina clean data.
To assess the quality of the genome assembly, Illumina sequencing reads were aligned to the genome using Bowtie2 and the genome coverage was calculated [59]. Besides, the unigenes obtained from the transcriptome data were mapped to the B. jamesi genome to assess the completeness of the gene regions. In addition, the sets of Benchmarking universal single-copy orthologs (BUSCO) was used to evaluate the completeness of the genome assembly (http://gitlab.com/ezlab/busco).
Repetitive sequence annotation
TEs in the B. jamesi genome were predicted by a combination of de novo-based and homology-based approaches. For TE annotation, both RepeatModeler and RepeatMasker were used to perform de novo identification [60]. RepeatMasker was used to identify transposable elements by aligning the genome assembly against the RepBase (RepBase21.04) and a local library generated by RepeatModeler with default parameters.
For phylogenetic analysis of TEs, MUSCLE v5 was used for generating multiple alignments of each cluster of TEs in a fast mode (-maxiters 2 -diags1) [61]. Based on the alignment results, the maximum likelihood (ML) method was used for phylogenetic tree construction with the parameters of “-n 1 -o tl -m 012345.” The visualization of the tree was performed on the iTOL (https://itol.embl.de/).
Protein-coding gene prediction and annotation
Protein-coding genes were predicted through the combination of de novo prediction, homology-based prediction and transcriptome-based prediction methods. For de novo prediction, the coding regions of the repeat-masked genome were predicted by Augustus v2.5.5 [62]. For homology-based prediction, protein-coding genes from Daphnia pulex, E. texana, Litopenaeus vannamei, Parhyale hawaiensis, Drosophila melanogaster, Bombyx mori, and Anopheles gambiae were downloaded from NCBI and mapped against the B. jamesi genome with Exonerate v2.2.0 (http://www.ebi.ac.uk/~guy/exonerate/). For transcriptome-based prediction, the transcriptome data was aligned to the B. jamesi genome using Tophat v2.1.1. Then, Cufflinks v2.2.1 was used to convert the transcripts to gene models [56]. Finally, all gene models predicted by above three methods were integrated into a non-redundant gene set through EvidenceModeler (EVM) v1.1.1 [63].
Functional annotation of the predicted genes was conducted by blasting against the NR and SwissProt databases using BLASTP program. Protein domains were annotated by mapping the genome to the InterPro and Pfam databases using InterProScan v5.50 and HMMER v3.3.1 [64, 65]. KEGG Automatic Annotation Server (KAAS) was used to annotate the pathways in which the genes might be involved through mapping against the KEGG database (https://www.genome.jp/kaas-bin/kaas_main). The GO classifications of the genes were extracted from the corresponding InterProScan or Pfam results (http://geneontology.org/docs/go-annotations/).
Gene family analyses
To understand the evolutionary dynamics of the genes, gene family clustering analysis was performed using the Markov clustering program OrthoFinder [66]. An all-to-all blast search was conducted on the protein-coding genes of 11 arthropods, including B. jamesi, A. vulgare, D. pulex, E. texana, Eurytemora affinis, L. vannamei, Eriocheir sinensis, Procambarus virginalis, P. hawaiensis, Tigriopus californicus, and D. melanogaster, using BLASTP program with a threshold value of E ≤ 1E−05.
Expansion and contraction of the gene families among these 11 species were determined. Based on the clustering results calculated by OrthoFinder and cladogram of these 11 species, gene gain and loss analysis was conducted by CAFE 5 [67]. The expansion and contraction of each gene family was examined by comparing cluster size differences between the ancestor and each of the current species. A random birth and death process model was used to identify gene gain and loss along each lineage of the RAxML tree.
Phylogenetic analysis
According to the results of gene family clustering, 177 single-copy orthologous genes were selected for phylogenetic tree construction. For each ortholog group, the amino acid sequence alignment was conducted using MUSCLE v5 with the default settings [61]. The 177 protein alignments were merged to form a super alignment matrix. Then, the ML method was used for phylogenetic tree construction under the PROTGAMMAJTT model using RAxML [68]. ML phylogeny and branch lengths were obtained by RAxML with 1000 bootstrap replicates. The divergence time estimation was conducted by combining programs of r8s and RAxML [69]. Fossil-derived timescales and evolutionary history were obtained from TIMETREE (http://timetree.org/).
Whole-genome duplication analysis
To infer WGD events in B. jamesi, we performed a series of analyses on the B. jamesi genome, including intrachromosome synteny block identification, Hox gene cluster comparison, and synonymous substitution (Ks) distribution analysis. To identify the synteny blocks, an all-against-all BLASTP method (E value < 1E−5) was used to detect paralogous genes in the B. jamesi genome, as well as the genomes of A. vulgare, Daphnia magna, T. tridentatus, and L. vannamei. Synteny blocks with at least five collinear homologous genes were detected using MCScanX software [70] with the following standard parameters: MATCH_SCORE: 50, MATCH_SIZE: 20, GAP_PENALTY: -1, OVERLAP_WINDOW: 5, E_VALUE: 1e-05, and MAX GAPS: 25. Genes were further classified by duplicate gene-classifier in MCScanX. For the Ks distribution analysis, the Ks values of the blocks from intraspecies were calculated using the HKY model [71, 72]. The Hox gene cluster, which contains at least nine highly conserved Hox genes, was identified in seven arthropod genomes, including a species with WGD, T. tridentatus [73].
Positive selection analysis
The nonsynonymous/synonymous substitution ratio (ω = dN/dS) was calculated to identify positively selected genes. The alignment of the orthologous genes was conducted with MUSCLE, and the stop codon and gaps in the alignment were removed. PAML was used to predict positively selected genes using the branch model [74, 75]. For the branch model test, positive selection was indicated when a significant difference between the alternative and null models was observed using the likelihood ratio test (LRT). The null model assumes that ω is constant (ω = 1), whereas the alternative model allows ω to vary among different branches.
Supplementary Information
Additional file 1: Table S1. Statistics of genome sequencing data of B. jamesi. Table S2. Genome size and repeat contents of the sequenced crustacean genomes. Table S3. Summary of the Illumina sequencing reads coverage on the assembly genome. Table S4. Core gene estimation for B. jamesi assembly. Table S5. Summary of isopod genome sizes. Table S6. Summary of the orthologous gene clusters analyzed in 11 species. Table S7. The gene family analysis results using CAFE. Table S8. GO enrichment of the expanded gene families of B. jamesi. Figure S1. K-mer distribution of the B. jamesi genome sequences. Figure S2. The core gene coverage of crustacean genomes. Figure S3. A Venn diagram of the statistics of the functional annotation. Figure S4. Whole-genome duplication analysis of B. jamesi. Figure S5. Age distribution of major expanded TEs in the two isopod genomes. Figure S6. The TE distribution surrounding genes. Figure S7. KEGG enrichment of the expanded gene families of B. jamesi. Figure S8. Expression level of the genes involved in the glycolysis of B. jamesi. Figure S9. The distribution ranges and max body lengths of the species of genus Bathynomus.
Acknowledgements
We acknowledge the support from Oceanographic Data Center, IOCAS.
Abbreviations
- TEs
Transposable elements
- SSRs
Simple sequence repeats
- LINEs
Long interspersed nuclear elements
- LTRs
Long terminal repeats
- Mya
Million years ago
- GO
Gene ontology
- CAMs
Cell adhesion molecules
- PLC
Phosphatidylinositol phospholipase C
- ITPR1
Inositol 1,4,5-triphosphate receptor type 1
- TPK
Tyrosine-protein kinase
- ADCY
Adenylate cyclase
- MTOR
Serine/threonine-protein kinase mTOR
- TSC2
Tuberous sclerosis 2
- MED
Mediator of RNA polymerase II transcription subunit
- TH
Thyroid hormone
- TPO
Thyroid peroxidase
- TRα
Thyroid hormone receptor α
- TRβ
Thyroid hormone receptor β
- TRIP11
Thyroid receptor-interacting protein 11
- IGF
Insulin growth factor
- IGFBP
Insulin-like growth factor-binding protein
- ISL
Insulin enhancer protein
- IR
IGF receptor
- IGFBP-ALS
Insulin-like growth factor-binding protein complex acid labile subunit
- PTPRF
Receptor-type tyrosine-protein phosphatase F
- LIPE
Hormone-sensitive lipase
- PHKA_B
Phosphorylase kinase alpha/beta subunit
- FLOT
Flotillin
- HCSH
Dachsous
- KIBRA
Protein Kibra
- YAP1
Transcriptional coactivator YAP1
- PP2A
Serine/threonine-protein phosphatase 2A
- SHOX
Short-stature homeobox protein
- HMG
High mobility group protein
- PGM2
Phosphoglucomutase-2
- ACSS1_2
Acetyl-CoA synthetase
- ADH
Alcohol dehydrogenase
- AnxB9
Annexin B9
- BMR
Basal metabolic rate
- SMRT
Single-molecule real time
- FPKM
Fragments per kilobase of transcript per million fragments mapped
- BUSCO
Benchmarking universal single-copy orthologs
- ML
Maximum likelihood
- KAAS
KEGG Automatic Annotation Server
- LRT
Likelihood ratio test
Authors’ contributions
F.L., X.L., J.X., J.Y., Q.K., and X.Z. initiated, managed, and drove the genome sequencing project. Q.K., X.Z. and J.Y. collected the animal material. J.Y., X.Z., and Y.S. prepared DNA sequencing and analysis. J.Y., Y.S., and C.L. performed genome assembly, gene annotation, genome structure analyses, and phylogenetic analyses. X.Z. and J.Y. conducted transcriptome sequencing and analysis. S.L., Y.Y., C.Z., and S.J conducted the genetics analysis. C.L., J.Y., and Y.S. submitted the genome data. J.Y., X.Z., and Q.K. wrote the manuscript and additional supplementary files. F. L., J.X., X.L., S.L., Y.Y., and S.J revised the manuscript. All authors read and approved the final manuscript.
Funding
We acknowledge financial support from the National Key Research & Development Program of China (2018YFD0900404 and 2018YFD0900103), the Natural Science Foundation of China (42176105, 31830100, 31972782, and 41876167), and the China Agriculture Research system-48 (CARS-48).
Availability of data and materials
All PacBio long-read sequencing data are available in the NCBI SRA database under accession ID of SRR16962112-SRR16962114. The genome assembly is available in the NCBI under Bioproject ID PRJNA776076 [76]. The genome assembly, predicted genes, repeats, and all raw sequencing data of genome and transcriptome are also available on the database at the link (username: sph; password: sph@8786@326): http://210.72.156.40/download/sphaeromadae/. In order to perform Protein-coding gene annotation and comparative genomics analysis, the genomes of 14 arthropods were downloaded from NCBI, including A. vulgare (PRJNA501402) [14], D. pulex (PRJNA12756) [77], E. texana (PRJNA352082) [78], E. affinis (PRJNA423276) [79], L. vannamei (PRJNA438564) [27], E. sinensis (PRJNA238496) [26], P. virginalis (PRJNA356499) [80], P. hawaiensis (PRJNA306836) [81], T. californicus (PRJNA237968) [82], D. magna (PRJNA490418) [83], T. tridentatus (PRJNA510236) [4], D. melanogaster (PRJNA559813) [84, 85], B. mori (PRJDB4947) [86], and A. gambiae (PRJNA20301) [75].
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Jianbo Yuan, Xiaojun Zhang and Qi Kou contributed equally to this work.
Contributor Information
Jianhai Xiang, Email: jhxiang@qdio.ac.cn.
Xinzheng Li, Email: lixzh@qdio.ac.cn.
Fuhua Li, Email: fhli@qdio.ac.cn.
References
- 1.Dover CLV. The ecology of deep-sea hydrothermal vents. USA: Princeton Univ Press; 2000.
- 2.Shen YJ, Kou Q, Zhong ZX, Li XZ, He LS, He SP, Gan XN. The first complete mitogenome of the South China deep-sea giant isopod Bathynomus sp (Crustacea: Isopoda: Cirolanidae) allows insights into the early mitogenomic evolution of isopods. Ecol Evol. 2017;7(6):1869–1881. doi: 10.1002/ece3.2737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ritchie H, Jamieson AJ, Piertney SB. Genome size variation in deep-sea amphipods. R Soc Open Sci. 2017;4(9):170862. [DOI] [PMC free article] [PubMed]
- 4.Liu RY, Wang K, Liu J, Xu WJ, Zhou Y, Zhu CL, Wu BS, Li YX, Wang W, He SP, et al. De novo genome assembly of limpet Bathyacmaea lactea (Gastropoda: Pectinodontidae): the first reference genome of a deep-sea gastropod endemic to cold seeps. Genome Biol Evol. 2020;12(6):905–910. doi: 10.1093/gbe/evaa100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sun YA, Sun J, Yang Y, Lan Y, Ip JCH, Wong WC, Kwan YH, Zhang YJ, Han Z, Qiu JW, et al. Genomic signatures supporting the symbiosis and formation of chitinous tube in the deep-sea tubeworm Paraescarpia echinospica. Mol Biol Evol. 2021;38(10):4116–4134. doi: 10.1093/molbev/msab203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ip JC, Xu T, Sun J, Li R, Chen C, Lan Y, Han Z, Zhang H, Wei J, Wang H, et al. Host-endosymbiont genome integration in a deep-sea chemosymbiotic clam. Mol Biol Evol. 2021;38(2):502–518. doi: 10.1093/molbev/msaa241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sun J, Zhang Y, Xu T, Zhang Y, Mu HW, Zhang YJ, et al. Adaptation to deep-sea chemosynthetic environments as revealed by mussel genomes. Nat Ecol Evol. 2017;1(5):0121. [DOI] [PubMed]
- 8.Wang K, Shen YJ, Yang YZ, Gan XN, Liu GC, Hu K, Li YX, Gao ZM, Zhu L, Yan GY, et al. Morphology and genome of a snailfish from the Mariana Trench provide insights into deep-sea adaptation. Nat Ecol Evol. 2019;3(5):823–833. doi: 10.1038/s41559-019-0864-8. [DOI] [PubMed] [Google Scholar]
- 9.Hartebrodt L. The biology, ecology, and societal importance of marine isopods. Encyclopedi World's Biomes. 2020;4:567–72.
- 10.Ono A, Tada T, Kose H. 2017 31st Ieee International Conference on Advanced Information Networking and Applications Workshops (Ieee Waina 2017) 2017. Research on giant isopod concerning the importance of biodiversity and its publicity by using ICT; pp. 449–454. [Google Scholar]
- 11.Hunt G, Roy K. Climate change, body size evolution, and Cope's Rule in deep-sea ostracodes. Proc Natl Acad Sci U S A. 2006;103(5):1347–1352. doi: 10.1073/pnas.0510550103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Weber JA, Park SG, Luria V, Jeon S, Kim HM, Jeon Y, Bhak Y, Jun JH, Kim SW, Hong WH, et al. The whale shark genome reveals how genomic and physiological properties scale with body size. Proc Natl Acad Sci U S A. 2020;117(34):20662–20671. doi: 10.1073/pnas.1922576117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Blanckenhorn WU. The evolution of body size: what keeps organisms small? Q Rev Biol. 2000;75(4):385–407. doi: 10.1086/393620. [DOI] [PubMed] [Google Scholar]
- 14.Chebbi MA, Becking T, Moumen B, Giraud I, Gilbert C, Peccoud J, Cordaux R. The genome of Armadillidium vulgare (Crustacea, Isopoda) provides insights into sex chromosome evolution in the context of cytoplasmic sex determination. Mol Biol Evol. 2019;36(4):727–741. doi: 10.1093/molbev/msz010. [DOI] [PubMed] [Google Scholar]
- 15.Becking T, Chebbi MA, Giraud I, Moumen B, Laverre T, Caubet Y, et al. Sex chromosomes control vertical transmission of feminizing Wolbachia symbionts in an isopod. PLoS Biol. 2019;17(10):e3000438. [DOI] [PMC free article] [PubMed]
- 16.Brionesfourzan P, Lozanoalvarez E. Aspects of the biology of the giant isopod Bathynomus-Giganteus Edwards, A.Milne, 1879 (Flabellifera, Cirolanidae), Off the Yucatan Peninsula. J Crustacean Biol. 1991;11(3):375–385. doi: 10.2307/1548464. [DOI] [Google Scholar]
- 17.Sankar R, Rajkumar M, Sun J, Gopalakrishnan A, Vasanthan TM, Ananthan G, Trilles JP. First record of three giant marine Bathynomids (Crustacea, Isopoda, Cirolanidae) from India. Acta Oceanol Sin. 2011;30(1):113–117. doi: 10.1007/s13131-011-0097-4. [DOI] [Google Scholar]
- 18.Cocke BT. Deep-sea isopods in aquaria. Tropical Fish Hobbyist. 1986;35:48–52. [Google Scholar]
- 19.Matsui T, Moriyama T, Kato R. Burrow plasticity in the deep-sea isopod Bathynomus doederleini (Crustacea: Isopoda: Cirolanidae) Zoolog Sci. 2011;28(12):863–868. doi: 10.2108/zsj.28.863. [DOI] [PubMed] [Google Scholar]
- 20.Biesiot PM, Wang SY, Perry HM, Trigg C. Organic reserves in the midgut gland and fat body of the giant deep-sea isopod Bathynomus giganteus. J Crustacean Biol. 1999;19(3):450–458. doi: 10.2307/1549253. [DOI] [Google Scholar]
- 21.Ginn F, Beisel U, Barua M. Flourishing with awkward creatures: togetherness, vulnerability, killing. Environ Humanit. 2014;4(1):113–123. doi: 10.1215/22011919-3614953. [DOI] [Google Scholar]
- 22.Clauss M, Frey R, Kiefer B, Lechner-Doll M, Loehlein W, Polster C, Rossner GE, Streich WJ. The maximum attainable body size of herbivorous mammals: morphophysiological constraints on foregut, and adaptations of hindgut fermenters. Oecologia. 2003;136(1):14–27. doi: 10.1007/s00442-003-1254-z. [DOI] [PubMed] [Google Scholar]
- 23.Martins E, Queiroz A, Santos RS, Bettencourt R. Finding immune gene expression differences induced by marine bacterial pathogens in the Deep-sea hydrothermal vent mussel Bathymodiolus azoricus. Biogeosciences. 2013;10(11):7279–7291. doi: 10.5194/bg-10-7279-2013. [DOI] [Google Scholar]
- 24.Kou Q, Chen J, Li X, He L, Wang Y. New species of the giant deep-sea isopod genus Bathynomus (Crustacea, Isopoda, Cirolanidae) from Hainan Island, South China Sea. Integrative zoology. 2017;12(4):283–291. doi: 10.1111/1749-4877.12256. [DOI] [PubMed] [Google Scholar]
- 25.Yuan JB, Zhang XJ, Wang M, Sun YM, Liu CZ, Li SH, et al. Simple sequence repeats drive genome plasticity and promote adaptive evolution in penaeid shrimp. Commun Biol. 2021;4(1):186. [DOI] [PMC free article] [PubMed]
- 26.Cui ZX, Liu Y, Yuan JB, Zhang XJ, Ventura T, Ma KY, et al. The Chinese mitten crab genome provides insights into adaptive plasticity and developmental regulation. Nat Commun. 2021;12(1):2395. [DOI] [PMC free article] [PubMed]
- 27.Zhang XJ, Yuan JB, Sun YM, Li SH, Gao Y, Yu Y, et al. Penaeid shrimp genome provides insights into benthic adaptation and frequent molting. Nat Commun. 2019;10:356. [DOI] [PMC free article] [PubMed]
- 28.Wendel JF, Cronn RC, Alvarez I, Liu B, Small RL, Senchina DS. Intron size and genome size in plants. Mol Biol Evol. 2002;19(12):2346–2352. doi: 10.1093/oxfordjournals.molbev.a004062. [DOI] [PubMed] [Google Scholar]
- 29.Wicker T, Gundlach H, Spannagl M, Uauy C, Borrill P, Ramirez-Gonzalez RH, et al. Impact of transposable elements on genome structure and evolution in bread wheat. Genome Biol. 2018;19:103. [DOI] [PMC free article] [PubMed]
- 30.Lins LSF, Ho SYW, Lo N. An evolutionary timescale for terrestrial isopods and a lack of molecular support for the monophyly of Oniscidea (Crustacea: Isopoda) Org Divers Evol. 2017;17(4):813–820. doi: 10.1007/s13127-017-0346-2. [DOI] [Google Scholar]
- 31.Mourouzis I, Lavecchia AM, Xinaris C. Thyroid hormone signalling: from the dawn of life to the bedside. J Mol Evol. 2020;88(1):88–103. doi: 10.1007/s00239-019-09908-1. [DOI] [PubMed] [Google Scholar]
- 32.Bouwman AC, Daetwyler HD, Chamberlain AJ, Ponce CH, Sargolzaei M, Schenkel FS, Sahana G, Govignon-Gion A, Boitard S, Dolezal M, et al. Meta-analysis of genome-wide association studies for cattle stature identifies common genes that regulate body size in mammals. Nat Genet. 2018;50(3):362–367. doi: 10.1038/s41588-018-0056-5. [DOI] [PubMed] [Google Scholar]
- 33.Hwa V, Oh Y, Rosenfeld RG. The insulin-like growth factor-binding protein (IGFBP) superfamily. Endocr Rev. 1999;20(6):761–787. doi: 10.1210/edrv.20.6.0382. [DOI] [PubMed] [Google Scholar]
- 34.Guo T, Bai YH, Cheng XJ, Han HB, Du H, Hu Y, Jia SQ, Xing XF, Ji JF. Insulin gene enhancer protein 1 mediates glycolysis and tumorigenesis of gastric cancer through regulating glucose transporter 4. Cancer Commun. 2021;41(3):258–272. doi: 10.1002/cac2.12141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Saxton RA, Sabatini DM. mTOR signaling in growth, metabolism, and disease (vol 168, pg 960, 2017) Cell. 2017;169(2):362. doi: 10.1016/j.cell.2017.03.035. [DOI] [PubMed] [Google Scholar]
- 36.Hayashi S, Yokoyama H, Tamura K. Roles of Hippo signaling pathway in size control of organ regeneration. Dev Growth Differ. 2015;57(4):341–351. doi: 10.1111/dgd.12212. [DOI] [PubMed] [Google Scholar]
- 37.Chung J, Zhang X, Collins B, Sper RB, Gleason K, Simpson S, Koh S, Sommer J, Flowers WL, Petters RM, et al. High mobility group A2 (HMGA2) deficiency in pigs leads to dwarfism, abnormal fetal resource allocation, and cryptorchidism. Proc Natl Acad Sci U S A. 2018;115(21):5420–5425. doi: 10.1073/pnas.1721630115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Morava E. Galactose supplementation in phosphoglucomutase-1 deficiency; review and outlook for a novel treatable CDG. Mol Genet Metab. 2014;112(4):275–279. doi: 10.1016/j.ymgme.2014.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yuan JB, Zhang XJ, Gao Y, Zhang XX, Liu CZ, Xiang JH, Li FH. Adaptation and molecular evidence for convergence in decapod crustaceans from deep-sea hydrothermal vent environments. Mol Ecol. 2020;29(20):3954–3969. doi: 10.1111/mec.15610. [DOI] [PubMed] [Google Scholar]
- 40.Tjota M, Lee S-K, Juan W, Williams JA, Khanna MR, Thomas GH. Annexin B9 binds to βH-spectrin and is required for multivesicular body function in Drosophila. J Cell Sci. 2011;124(17):2914–2926. doi: 10.1242/jcs.078667. [DOI] [PubMed] [Google Scholar]
- 41.Chenais B, Caruso A, Hiard S, Casse N. The impact of transposable elements on eukaryotic genomes: from genome size increase to genetic adaptation to stressful environments. Gene. 2012;509(1):7–15. doi: 10.1016/j.gene.2012.07.042. [DOI] [PubMed] [Google Scholar]
- 42.Rees DJ, Dufresne F, Glemet H, Belzile C. Amphipod genome sizes: first estimates for Arctic species reveal genomic giants. Genome. 2007;50(2):151–158. doi: 10.1139/G06-155. [DOI] [PubMed] [Google Scholar]
- 43.Huang YT, Bian C, Liu ZQ, Wang LL, Xue CH, Huang HL, et al. The first genome survey of the Antarctic Krill (Euphausia superba) provides a valuable genetic resource for polar biomedical research. Mar Drugs. 2020;18(4):185. [DOI] [PMC free article] [PubMed]
- 44.Lefebure T, Morvan C, Malard F, Francois C, Konecny-Dupre L, Gueguen L, Weiss-Gayet M, Seguin-Orlando A, Ermini L, Sarkissian C, et al. Less effective selection leads to larger genomes. Genome Res. 2017;27(6):1016–1028. doi: 10.1101/gr.212589.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Jeffery NW. Genome size diversity and evolution in the Crustacea. Canada: University of Guelph; 2015. A Thesis Presented to The Faculty of Graduate Studies
- 46.Boos H, Scalco ACS, Araujo PB. Biological and ecological traits of Bathynomus giganteus and Bathynomus miyarei (Crustacea: Isopoda): contribution to the conservation of deep-sea in southern Brazil. Aquatic Conserv Ma Freshwater Ecosyst. 2021;31(3):1–11.
- 47.Timofeev SF. Bergmann's principle and deep-water gigantism in marine crustaceans. Biol Bull. 2001;28(6):646–650. doi: 10.1023/A:1012336823275. [DOI] [PubMed] [Google Scholar]
- 48.Vermeij GJ. Gigantism and its implications for the history of life. Plos One. 2016;11(1):e0146092. doi: 10.1371/journal.pone.0146092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Brown A, Thatje S. Explaining bathymetric diversity patterns in marine benthic invertebrates and demersal fishes: physiological contributions to adaptation of life at depth. Biol Rev. 2014;89(2):406–426. doi: 10.1111/brv.12061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Todd RT, Wikoff TD, Forche A, Selmecki A. Genome plasticity in Candida albicans is driven by long repeat sequences. Elife. 2019;8:e45954. [DOI] [PMC free article] [PubMed]
- 51.Schrader L, Schmitz J. The impact of transposable elements in adaptive evolution. Mol Ecol. 2019;28(6):1537–1549. doi: 10.1111/mec.14794. [DOI] [PubMed] [Google Scholar]
- 52.Wolf M, Ingbar SH, Moses AC. Thyroid-hormone and growth-hormone interact to regulate insulin-like growth factor-i messenger ribonucleic-acid and circulating levels in the rat. Endocrinology. 1989;125(6):2905–2914. doi: 10.1210/endo-125-6-2905. [DOI] [PubMed] [Google Scholar]
- 53.Hannah-Shmouni F, Trivellin G, Stratakis CA. Genetics of gigantism and acromegaly. Growth Horm IGF Res. 2016;30-31:37–41. doi: 10.1016/j.ghir.2016.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Rau AR. Biological scaling and physics. J Biosci. 2002;27(5):475–478. doi: 10.1007/BF02705043. [DOI] [PubMed] [Google Scholar]
- 55.Thommen A, Werner S, Frank O, Philipp J, Knittelfelder O, Quek Y, et al. Body size-dependent energy storage causes Kleiber’s law scaling of the metabolic rate in planarians. Elife. 2019;8:e38187. [DOI] [PMC free article] [PubMed]
- 56.Trapnell C, Pachter L, Lsalzberg S. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25(9):1105. doi: 10.1093/bioinformatics/btp120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–140. doi: 10.1093/bioinformatics/btp616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Marcais G, Kingsford C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics. 2011;27(6):764–770. doi: 10.1093/bioinformatics/btr011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9(4):357–359. doi: 10.1038/nmeth.1923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Tarailo-Graovac M, Chen N. Current protocols in bioinformatics / editoral board, Andreas D Baxevanis [et al] 2009. Using RepeatMasker to identify repetitive elements in genomic sequences. [DOI] [PubMed] [Google Scholar]
- 61.Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792–1797. doi: 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Stanke M, Steinkamp R, Waack S, Morgenstern B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 2004;32(Web Server issue):W309–W312. doi: 10.1093/nar/gkh379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Haas B, Salzberg S, Zhu W, Pertea M, Allen J, Orvis J, White O, Buell C, Wortman J. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 2008;9(1):R7. doi: 10.1186/gb-2008-9-1-r7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Zdobnov EM, Apweiler R. InterProScan--an integration platform for the signature-recognition methods in InterPro. Bioinformatics. 2001;17(9):847–848. doi: 10.1093/bioinformatics/17.9.847. [DOI] [PubMed] [Google Scholar]
- 65.Prakash A, Jeffryes M, Bateman A, Finn RD. Current protocols in bioinformatics / editoral board, Andreas D Baxevanis [et al] 2017. The HMMER Web Server for Protein Sequence Similarity Search. [DOI] [PubMed] [Google Scholar]
- 66.Emms DM, Kelly S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 2015;16:157. doi: 10.1186/s13059-015-0721-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Mendes FK, Vanderpool D, Fulton B, Hahn MW. CAFE 5 models variation in evolutionary rates among gene families. Bioinformatics. 2020;36(22-23):5516–5518. doi: 10.1093/bioinformatics/btaa1022. [DOI] [PubMed] [Google Scholar]
- 68.Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30(9):1312–1313. doi: 10.1093/bioinformatics/btu033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Yang Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci : CABIOS. 1997;13(5):555–556. doi: 10.1093/bioinformatics/13.5.555. [DOI] [PubMed] [Google Scholar]
- 70.Wang Y, Tang H, Debarry JD, Tan X, Li J, Wang X, Lee TH, Jin H, Marler B, Guo H, et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012;40(7):e49. doi: 10.1093/nar/gkr1293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Hasegawa M, Kishino H, Yano TA. Dating of the human ape splitting by a molecular clock of mitochondrial-DNA. J Mol Evol. 1985;22(2):160–174. doi: 10.1007/BF02101694. [DOI] [PubMed] [Google Scholar]
- 72.Schwager EE, Sharma PP, Clarke T, Leite DJ, Wierschin T, Pechmann M, et al. The house spider genome reveals an ancient whole-genome duplication during arachnid evolution. BMC Biol. 2017;15:62. [DOI] [PMC free article] [PubMed]
- 73.Gong L, Fan G, Ren Y, Chen Y, Qiu Q, Liu L, Qin Y, Liu B, Jiang L, Li H, et al. Chromosomal level reference genome of Tachypleus tridentatus provides insights into evolution and adaptation of horseshoe crabs. Mol Ecol Resour. 2019;19(3):744–756. doi: 10.1111/1755-0998.12988. [DOI] [PubMed] [Google Scholar]
- 74.Zhang J, Nielsen R, Yang Z. Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol Biol Evol. 2005;22(12):2472–2479. doi: 10.1093/molbev/msi237. [DOI] [PubMed] [Google Scholar]
- 75.Mongin E, Louis C, Holt RA, Birney E, Collins FH. The Anopheles gambiae genome: an update. Trends Parasitol. 2004;20(2):49–52. doi: 10.1016/j.pt.2003.11.003. [DOI] [PubMed] [Google Scholar]
- 76.Bathynomus jamesi isolate:JY-2021 Genome sequencing. NCBI accession: PRJNA776076. (2021). https://www.ncbi.nlm.nih.gov/bioproject/PRJNA776076
- 77.Colbourne JK, Pfrender ME, Gilbert D, Thomas WK, Tucker A, Oakley TH, Tokishita S, Aerts A, Arnold GJ, Basu MK, et al. The ecoresponsive genome of Daphnia pulex. Science. 2011;331(6017):555–561. doi: 10.1126/science.1197761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Baldwin-Brown JG, Weeks SC, Long AD. A new standard for crustacean genomes: the highly contiguous, annotated genome assembly of the clam shrimp eulimnadia texana reveals HOX gene order and identifies the sex chromosome. Genome Biol Evol. 2018;10(1):143–156. doi: 10.1093/gbe/evx280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Choi BS, Kim DH, Kim MS, Park JC, Lee YH, Kim HJ, Jeong CB, Hagiwara A, Souissi S, Lee JS. The genome of the European estuarine calanoid copepod Eurytemora affinis: potential use in molecular ecotoxicology. Mar Pollut Bull. 2021;166:112190. doi: 10.1016/j.marpolbul.2021.112190. [DOI] [PubMed] [Google Scholar]
- 80.Gutekunst J, Andriantsoa R, Falckenhayn C, Hanna K, Stein W, Rasamy J, et al. Clonal genome evolution and rapid invasive spread of the marbled crayfish. Nat Ecol Evol. 2018;2(3):567–73. [DOI] [PubMed]
- 81.Kao D, Lai AG, Stamataki E, Rosic S, Konstantinides N, Jarvis E, et al. The genome of the crustacean Parhyale hawaiensis, a model for animal development, regeneration, immunity and lignocellulose digestion. Elife. 2016;5:e20062. [DOI] [PMC free article] [PubMed]
- 82.Barreto FS, Watson ET, Lima TG, Willett CS, Edmands S, Li W, Burton RS. Genomic signatures of mitonuclear coevolution across populations of Tigriopus californicus. Nat Ecol Evol. 2018;2(8):1250–1257. doi: 10.1038/s41559-018-0588-1. [DOI] [PubMed] [Google Scholar]
- 83.Lee BY, Choi BS, Kim MS, Park JC, Jeong CB, Han J, Lee JS. The genome of the freshwater water flea Daphnia magna: a potential use for freshwater molecular ecotoxicology. Aquat Toxicol. 2019;210:69–84. doi: 10.1016/j.aquatox.2019.02.009. [DOI] [PubMed] [Google Scholar]
- 84.Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, et al. The genome sequence of Drosophila melanogaster. Science. 2000;287(5461):2185–95. [DOI] [PubMed]
- 85.Kim BY, Wang JR, Miller DE, Barmina O, Delaney E, Thompson A, et al. Highly contiguous assemblies of 101 drosophilid genomes. Elife. 2021;10:e66405. [DOI] [PMC free article] [PubMed]
- 86.International Silkworm Genome C The genome of a lepidopteran model insect, the silkworm Bombyx mori. Insect Biochem Mol Biol. 2008;38(12):1036–1045. doi: 10.1016/j.ibmb.2008.11.004. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Table S1. Statistics of genome sequencing data of B. jamesi. Table S2. Genome size and repeat contents of the sequenced crustacean genomes. Table S3. Summary of the Illumina sequencing reads coverage on the assembly genome. Table S4. Core gene estimation for B. jamesi assembly. Table S5. Summary of isopod genome sizes. Table S6. Summary of the orthologous gene clusters analyzed in 11 species. Table S7. The gene family analysis results using CAFE. Table S8. GO enrichment of the expanded gene families of B. jamesi. Figure S1. K-mer distribution of the B. jamesi genome sequences. Figure S2. The core gene coverage of crustacean genomes. Figure S3. A Venn diagram of the statistics of the functional annotation. Figure S4. Whole-genome duplication analysis of B. jamesi. Figure S5. Age distribution of major expanded TEs in the two isopod genomes. Figure S6. The TE distribution surrounding genes. Figure S7. KEGG enrichment of the expanded gene families of B. jamesi. Figure S8. Expression level of the genes involved in the glycolysis of B. jamesi. Figure S9. The distribution ranges and max body lengths of the species of genus Bathynomus.
Data Availability Statement
All PacBio long-read sequencing data are available in the NCBI SRA database under accession ID of SRR16962112-SRR16962114. The genome assembly is available in the NCBI under Bioproject ID PRJNA776076 [76]. The genome assembly, predicted genes, repeats, and all raw sequencing data of genome and transcriptome are also available on the database at the link (username: sph; password: sph@8786@326): http://210.72.156.40/download/sphaeromadae/. In order to perform Protein-coding gene annotation and comparative genomics analysis, the genomes of 14 arthropods were downloaded from NCBI, including A. vulgare (PRJNA501402) [14], D. pulex (PRJNA12756) [77], E. texana (PRJNA352082) [78], E. affinis (PRJNA423276) [79], L. vannamei (PRJNA438564) [27], E. sinensis (PRJNA238496) [26], P. virginalis (PRJNA356499) [80], P. hawaiensis (PRJNA306836) [81], T. californicus (PRJNA237968) [82], D. magna (PRJNA490418) [83], T. tridentatus (PRJNA510236) [4], D. melanogaster (PRJNA559813) [84, 85], B. mori (PRJDB4947) [86], and A. gambiae (PRJNA20301) [75].