Abstract
Utilization of ferrioxamines as sole sources of iron distinguishes Salmonella enterica serotypes Typhimurium and Enteritidis from a number of related species, including Escherichia coli. Ferrioxamine supplements have therefore been used in preenrichment and selection media to increase the bacterial growth rate while selectivity is maintained. We characterized the determinants involved in utilization of ferrioxamines B, E, and G by S. enterica serotype Typhimurium by performing siderophore cross-feeding bioassays. Transport of all three ferric siderophores across the outer membrane was dependent on the FoxA receptor encoded by the Fur-repressible foxA gene. However, only the transport of ferrioxamine G was dependent on the energy-transducing protein TonB, since growth stimulation of a tonB strain by ferrioxamines B and E was observed, albeit at lower efficiencies than in the parental strain. Transport across the inner membrane was dependent on the periplasmic binding protein-dependent ABC transporter complex comprising FhuBCD, as has been reported for other hydroxamate siderophores of enteric bacteria. The distribution of the foxA gene in the genus Salmonella, as indicated by DNA hybridization studies and correlated with the ability to utilize ferrioxamine E, was restricted to subspecies I, II, and IIIb, and this gene was absent from subspecies IIIa, IV, VI, and VII (formerly subspecies IV) and Salmonella bongori (formerly subspecies V). S. enterica serotype Typhimurium mutants with either a transposon insertion or a defined nonpolar frameshift (+2) mutation in the foxA gene were not able to utilize any of the three ferrioxamines tested. A strain carrying the nonpolar foxA mutation exhibited a significantly reduced ability to colonize rabbit ileal loops compared to the foxA+ parent. In addition, a foxA mutant was markedly attenuated in mice inoculated by either the intragastric or intravenous route. Mice inoculated with the foxA mutant were protected against subsequent challenge by the foxA+ parent strain.
The genus Salmonella shared an ancestor with Escherichia coli some 100 million to 160 million years ago (31) and has since become a pathogen in a wide range of warm- and cold-blooded vertebrate hosts. DNA has been either lost by deletion or introduced by bacteriophage- or plasmid-mediated horizontal transfer in both of these taxa since the divergence. The genetic material that is present in Salmonella enterica serotypes but absent from E. coli includes the Salmonella virulence plasmid, Salmonella pathogenicity island 1 (SPI1), SPI2, and smaller pathogenicity islets, such as the lpf operon and sifA (14). Other differences between the taxa involve genes whose products have a role in iron acquisition. For example, E. coli expresses the ferric dicitrate transport mechanism encoded by the fecABCD genes, but Salmonella spp. lack a comparable system (46). On the other hand, S. enterica expresses two outer membrane proteins, FepA and IroN, that mediate uptake of the catechol siderophore enterobactin, while E. coli possesses only the fepA gene (4). Moreover, many Salmonella strains are able to acquire iron complexed with ferrioxamines B, E, and G (22), hydroxamate siderophores that they do not themselves synthesize. In contrast, E. coli strains cannot efficiently utilize ferrioxamines (15).
Since ferrioxamines are potent growth factors for the common S. enterica serotypes but not for a number of closely related bacteria, they have been used as supplements in standard enrichment and selection procedures to increase the speed and sensitivity of detection of members of the genus Salmonella in a number of food products (19, 32, 38). For example, standard procedures used for detection of small numbers of organisms in eggs require preenrichment of mixtures of yolk and albumin at room temperature for 1 to 2 days. In contrast, preenrichment of artificially infected albumin in buffered peptone water supplemented with ferrioxamine E allowed detection of contaminating S. enterica serotype Typhimurium within 6 h (19, 36). Similarly, two to five cells of S. enterica serotype Enteritidis per 25 g of egg albumin were detected within 24 h when selective preenrichment in buffered peptone water containing ferrioxamine E was used (3).
Little is known about the mechanism of uptake of iron(III)-ferrioxamine complexes by Salmonella spp. Since the ferrioxamines have molecular weights of approximately 600, which is greater than the theoretical limit for passive diffusion through porins (27), it is likely that, like uptake of other siderophores of enteric bacteria, ferrioxamine uptake requires binding to a specific receptor protein, followed by TonB- and ExbBD-dependent transport across the outer membrane. Transport of siderophores through the inner membrane normally depends on a periplasmic binding protein-dependent ABC transporter complex. It is possible that ferrioxamine uptake by Salmonella spp. involves the FhuBCD inner membrane permease complex required for utilization of other hydroxamate siderophores (21). However, none of this has been demonstrated experimentally, nor has the ability of S. enterica serotypes other than serotypes Typhimurium and Enteritidis to utilize ferrioxamines as iron sources been determined. In this paper we describe experiments aimed at characterizing the mechanism of ferrioxamine transport by S. enterica serotype Typhimurium and at assessing the distribution of the ability to utilize ferrioxamine in the genus Salmonella. The results of a preliminary investigation of the role of the ferrioxamine receptor FoxA in experimental salmonellosis are also presented.
MATERIALS AND METHODS
Bacterial strains and plasmids.
The strains and plasmids used in this study are listed in Table 1. In addition, two previously characterized Salmonella strain collections were used for a hybridization analysis, the SARB collection consisting of 72 serotypes of subspecies I (7) and a collection consisting of representative strains of the other species and subspecies of the genus, including Salmonella bongori (35). Strains were cultured aerobically at 37°C as required. The ferrous iron chelator 2,2′-bipyridyl (Sigma) was added to media at the concentrations indicated below to impose iron limitation. Growth in liquid media was quantified by measuring the optical density at 620 nm. Bacteriophage P22 HT105/1 int− (39) was used for generalized transduction of markers between smooth strains of S. enterica. Transductants were routinely purified and made bacteriophage free by streaking them onto nonselective green indicator plates as described previously (23). All foxA+-foxA isogenic pairs of strains had equivalent growth characteristics in nutrient broth (NB) (Oxoid no. 2), as judged by the length of the lag phase, the maximal growth rate, and the climax cell density during the stationary phase.
TABLE 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Relevant characteristics | Source or reference |
|---|---|---|
| S. enterica serotype Typhimurium strains | ||
| ATCC 14028 | Wild type | ATCCa |
| AR1258 | ATCC 14028, entB | 44 |
| RK804 | AR1258, tonB | 18 |
| RK809 | AR1258, foxA::pMAP | This study |
| enb-7 | LT2, class II ent mutant | 33 |
| WR1024 | enb-7, fhuB::MudJ | Laboratory stock |
| SL1344 | hisG46 | 17 |
| SL1344/nr | SL1344, Nalr | This study |
| BK102 | SL1344/nr, foxA | This study |
| TML | Wild type | 12 |
| TML/nr | TML, Nalr | This study |
| RK102 | TML/nr, foxA | This study |
| E. coli SM10λpir | F−thi-1 thr-1 leuB6 supE44 tonA21 lacY1 recA::RP4-2-tc::Mu Kmr | R. Haigh |
| Plasmids | ||
| pUC18 | ColE1, bla | Laboratory stock |
| pRA17 | pUC18 carrying the 287-bp foxA fragment | This study |
| pRA5 | pUC18 carrying the foxA 5′ region | This study |
| pBluescriptSK | ColE1, bla | Laboratory stock |
| pRA19 | pBluescriptSK carrying the mutated 287-bp foxA fragment | This study |
| pRDH10 | sacRB cat oriR6K mobRP4, Tetr | R. Haigh |
| pRA21 | pRDH10 carrying the 287-bp foxA fragment | This study |
| pIRS618 | pSUKS1 carrying the serovar Typhimurium tonB gene | 45 |
| pMAP | bla oriR6K mobRP4 | 44 |
ATCC, American Type Culture Collection.
Siderophore assays.
The abilities of bacterial strains to use siderophores as sources of iron were determined by performing cross-feeding tests. Vogel-Bonner medium supplemented with 2,2′-bipyridyl was used to test enterobactin-deficient strains (18). Enterobactin-producing strains were tested in bioassays by using egg white medium (EWM) consisting of nutrient agar base (25.6 g of NB per liter, 6 g of yeast extract per liter, 18 g of Na2HPO4 · 12H2O per liter, 6 g of KH2PO4 per liter, 44 g of Oxoid agar no. 1 per liter) to which fresh sterile egg white was added. The egg white was passed repeatedly through a sterile 1.2-mm-gauge syringe needle in order to mix it before use; 9 ml of the resulting preparation was added to 11 ml of nutrient agar base, which was sufficient for a single assay plate. Ferrioxamines B and E were gifts from H. P. Schnebli, Novartis Pharma Ltd., Basel, Switzerland; ferrioxamine G was prepared and purified as previously described (37); coprogen was a gift from G. Winkelmann, University of Tübingen, Tübingen, Germany; and enterobactin was prepared and purified as described by Young (48).
Recombinant DNA techniques.
Standard methods were used for isolation of chromosomal DNA, restriction endonuclease analysis, ligation, and transformation of plasmid DNA (24). Southern transfer by capillary action was performed as previously described (24). Labeling of DNA probes, hybridization, and immunological detection were performed by using Gene Images nonradioactive labeling and detection kits from Amersham. Hybridization was performed at 65°C in 5× SSC (1× SSC contains 8.77 g of NaCl per liter and 4.41 g of sodium citrate per liter, pH 7.0) containing 0.1% (wt/vol) sodium dodecyl sulfate, 5% (wt/vol) dextran sulfate, and 0.5% (wt/vol) blocking agent (supplied by the manufacturer). Filters were subsequently washed at high stringency with 0.1× SSC containing 0.1% sodium dodecyl sulfate. The plasmid DNA used for sequencing was isolated by using ion-exchange columns obtained from Qiagen.
Preparation of a foxA probe and attempts to clone the Salmonella foxA gene.
The foxA gene of Yersinia enterocolitica, which encodes a 77-kDa outer membrane protein involved in the transport of ferrioxamine B, has been cloned and characterized (1). We previously identified part of an open reading frame (GenBank accession no. U62282) in the genome of S. enterica serotype Typhimurium whose deduced amino acid sequence exhibits 45% identity with the amino acid sequence of the FoxA protein of Y. enterocolitica (44). Using this sequence, we designed two primers, FX1 (5′ AGGCGGATCCATCGGCGGC 3′) and FX2 (5′ ACGGGATCCAGATCACCGTCC 3′) (incorporating BamHI sites by including mismatching bases), which generated an approximately 300-bp PCR product that was subsequently cloned into the BamHI site of pUC18 to give a recombinant plasmid designated pRA17. Sequencing confirmed that the insert was a 287-bp subgenic fragment corresponding to base pairs 55 to 342 of the foxA coding sequence described previously (44).
Three plasmid libraries and four cosmid libraries of S. enterica (serotype Typhimurium, serotype Paratyphi, or serotype Typhi) chromosomal DNA were screened, either by hybridization with the subgenic foxA probe or by hybridization in conjunction with complementation in a strain unable to utilize ferrioxamine E as a sole source of iron. No recombinant plasmids or cosmids that contained the entire foxA gene were found in these libraries. However, one positively hybridizing clone (designated pRA5) from a pUC18 library of partially Sau3A-digested S. enterica serotype Typhimurium chromosomal DNA contained part of the foxA gene and the 5′ flanking sequence; thus, this clone allowed us to determine the regulatory region (GenBank accession no. AF060876).
Construction of defined nonpolar foxA mutants.
PCR primers FXM1 and FXM2 were designed by using the previously determined foxA sequence (44) to incorporate BglII restriction sites, as shown in Fig. 1B; these two primers were used along with primers P1L and P2L (which hybridized with sequences internal to the pUC18 vector sequence of pRA17), respectively, to amplify the foxA subgenic fragment borne by plasmid pRA17 as two separate PCR products that were approximately 190 and 230 bp long. These products were recovered, digested with BglII, ligated together, and cloned into pBluescriptSK at the XbaI-SstI sites to generate recombinant plasmid pRA19. Sequence analysis of the insert in pRA19 confirmed that a BglII cleavage site and a 2-bp insertion were introduced between base pairs 218 and 219 of the wild-type open reading frame (Fig. 1A). The +2 frameshift resulted in a termination codon 21 bases downstream of the insertion site.
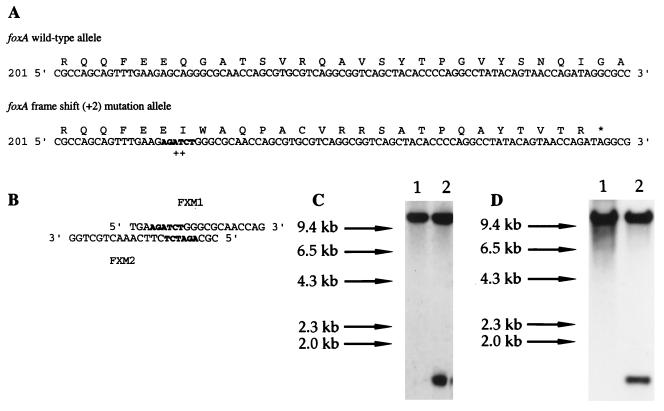
FIG. 1.
Construction of a frameshift mutation in the foxA gene of S. enterica serotype Typhimurium. (A) Comparison of the nucleotide sequences of the wild-type and mutated foxA alleles, as confirmed by sequence analysis. The mutation consists of a 2-bp (AT) insertion (indicated by plus signs) and a BglII restriction enzyme recognition sequence (indicated by boldface type). The mutation was introduced into a cloned subgenic foxA fragment as described in the text by using primers FXM1 and FXM2, which have mismatching base pairs that include the BglII recognition site (B). (C and D) The mutations were confirmed by Southern hybridization analysis in which we used a subgenic foxA fragment as the probe. (C) Analysis of BglII-digested chromosomal DNA of parental strain TML (lane 1) and the foxA derivative strain RK102 (lane 2). (D) Analysis of parental strain SL1344 (lane 1) and the foxA derivative strain BK102 (lane 2).
To introduce the +2 frameshift mutation into the bacterial chromosome by allelic exchange, the insert was recloned into the suicide plasmid vector pRDH10 at BamHI sites to generate plasmid pRA21. Plasmid pRDH10 contains oriR6K and so requires the protein product of λpir for replication (20); derivative plasmid pRA21 was therefore maintained in E. coli SM10λpir and introduced by conjugation into strains SL1344/nr and TML/nr (spontaneous nalidixic acid-resistant derivatives of S. enterica serotype Typhimurium strains SL1344 and TML, respectively), which were plated onto nutrient agar containing nalidixic acid (50 μg/ml) and chloramphenicol (20 μg/ml). Since the recipients lacked λpir, growth on this medium directly selected for merodiploids in which the plasmid (encoding chloramphenicol resistance) had integrated into the host chromosome by a single recombination event. To select for loss of the integrated suicide vector by a second homologous recombination event, merodiploid strains were grown in medium containing tryptone (10 g/liter), yeast extract (5 g/liter), and 5% (wt/vol) sucrose to the mid-log phase, and dilutions of the cultures were plated onto tryptone-yeast extract-sucrose agar to enrich for bacteria that had lost the sacRB gene of the vector plasmid pRDH10 (6, 40). Bacteria growing on this medium were tested for sensitivity to tetracycline (10 μg/ml) as an indicator of loss of the vector. They were then screened for the presence of a new BglII recognition site at base pairs 217 to 222 by PCR amplification of chromosomal DNA with primers FX1 and FX2 and subsequent digestion of the product with BglII. Resolved merodiploid derivatives of SL1344/nr and TML/nr carrying the +2 frameshift foxA mutation were designated BK102 and RK102, respectively. The presence of the mutation was confirmed by performing a Southern blot analysis of BglII-digested chromosomal DNA with a foxA-specific probe (Fig. 1C and D). A BglII >10-kb fragment in the DNA of each parental strain hybridized with the probe, while two fragments (1.5 and 10 kb) hybridized in the BglII-digested DNA of both putative foxA mutants.
Rabbit ileal loop anastomosis test.
Competitive colonization experiments to determine the effects of the foxA mutation were performed by using a modification of the rabbit ileal loop test with 2- to 3-kg specific-pathogen-free rabbits, as previously described (47). Each test loop (5 cm with 2-cm spacers) was inoculated by injecting 1 ml of a mixture (approximately 1:1) of RK102 (foxA) and the foxA+ parent strain TML (total inoculum, 107 or 108 CFU in phosphate-buffered saline). To rule out the possibility that the nalidixic acid resistance mutation in RK102 had any effect, a control experiment involving similar mixtures of TML and TML/nr was also performed. The negative and positive controls consisted of 1 ml of sterile phosphate-buffered saline and 1 mg of cholera toxin in 1 ml of phosphate-buffered saline, respectively. Terminal anesthesia was administered 18 h after infection, and the loops were surgically removed for analysis. All test loops contained a small amount (0.5 to 1 ml) of purulent exudate. The positive control loops (containing cholera toxin) contained 15 to 20 ml of accumulated fluid. No accumulated fluid or purulent exudate was present in the negative control loops. The proportion of nalidixic acid-resistant bacteria (TML/nr or RK102) in each test loop was determined by plating serial dilutions of lumen contents and homogenized loop tissue onto unsupplemented MacConkey agar (Oxoid) and onto MacConkey agar containing nalidixic acid (50 μg/ml). The ratios of nalidixic acid-sensitive bacteria to nalidixic acid-resistant bacteria recovered were expressed as log10 values for statistical analysis. The Student t test was used to determine whether the ratio for the bacteria recovered was significantly different from the ratio for the inoculum.
Mouse typhoid model.
Female BALB/c mice that were 6 to 8 weeks old and were housed under specific-pathogen-free conditions were used to determine the 50% lethal doses (LD50s) of bacterial strains by intragastric and intravenous inoculation. Bacteria grown overnight in NB without shaking at 37°C were harvested by centrifugation and resuspended in sterile phosphate-buffered saline. Serial 10-fold dilutions containing 103 to 109 CFU were prepared and used for intragastric inoculation into groups of five mice, and serial 10-fold dilutions containing 10 to 104 CFU were prepared and used for intravenous inoculation into other groups of five mice. Mortality was recorded 28 days after infection, and LD50s were calculated as described previously (34).
RESULTS
Role of TonB and FhuB in ferrioxamine-mediated iron uptake by Salmonella strains.
Siderophore cross-feeding tests on Vogel-Bonner medium supplemented with 2,2′-bipyridyl were used to assess the ability of the tonB strain RK804 to utilize the catechol siderophore enterobactin and the hydroxamate siderophores coprogen and ferrioxamines B, E, and G (Table 2). No growth stimulation of RK804 was observed around discs loaded with coprogen or enterobactin, confirming that functional TonB protein is required for high-affinity transport of these siderophores. Similarly, no growth stimulation of RK804 was observed around a disc loaded with ferrioxamine G. In contrast, however, there was significant growth stimulation of RK804 by ferrioxamines B and E, although the growth zones were somewhat smaller than those of the tonB+ parent strain AR1258. The growth of a derivative of strain RK804 harboring the recombinant plasmid pIRS618 (tonB+) was stimulated by all of the siderophores tested, and the growth zones were comparable in diameter to those of strain AR1258 (Table 2) but were less dense, which probably reflected a lower growth rate of the plasmid-containing strain.
TABLE 2.
Siderophore cross-feeding of various serotype Typhimurium strainsa
| Medium | Concn of 2,2′-bipyridyl (μM) | Strain | Relevant genotype | Results with the following siderophores:
|
||||
|---|---|---|---|---|---|---|---|---|
| Ferrioxamine B | Ferrioxamine E | Ferrioxamine G | Coprogen | Enterobactin | ||||
| Vogel-Bonnerb | 200 | AR1258 | entB | 28 | 29 | 26 | 28 | 30 |
| RK804 | entB tonB | 13 | 14 | 5 | 5 | 5 | ||
| RK804(pIRS618) | entB tonB+ | 26 | 25 | 23 | 28 | 30 | ||
| WR1024 | ent fhuB | 5 | 5 | 5 | 5 | 30 | ||
| RK809 | entB foxA | 8 | 5 | 5 | 28 | 30 | ||
| EWMc | 400 | SL1344/nr | Wild type | + | + | + | + | ND |
| BK102 | foxA | (+) | − | − | + | ND | ||
| TML/nr | Wild type | + | + | + | + | ND | ||
| RK102 | foxA | (+) | − | − | + | ND | ||
Approximately 106 cells of each strain tested were spread over the surface of Vogel-Bonner or EWM agar. Sterile filter paper discs (diameter, 5 mm), each impregnated with 1 μg of a siderophore, were placed on the agar surfaces.
For the Vogel-Bonner medium bioassays, the sizes (in millimeters) of growth zones around siderophore discs were recorded after 18 h of incubation at 37°C; a value of 5 mm indicates that there was no growth stimulation.
In the EWM bioassays, growth was too diffuse to measure accurately as discrete zones, and so the results were recorded as follows: +, obvious growth; (+), limited growth around the disc; and −, no observable growth after 3 to 4 h of incubation at 37°C. ND, growth stimulation not determined.
To test whether a mutation in the fhuACDB operon abolished ferrioxamine utilization, strain WR1024 (fhuB::MudJ) was tested for growth stimulation in a cross-feeding test on Vogel-Bonner medium supplemented with 2,2′-bipyridyl (Table 2). No growth stimulation was observed around discs loaded with the hydroxamate siderophore coprogen or ferrioxamine B, E, or G, while growth stimulation comparable to that of the parental strain was observed, as expected, around discs loaded with the catechol compound enterobactin.
Distribution of a Y. enterocolitica foxA homolog within the genus Salmonella.
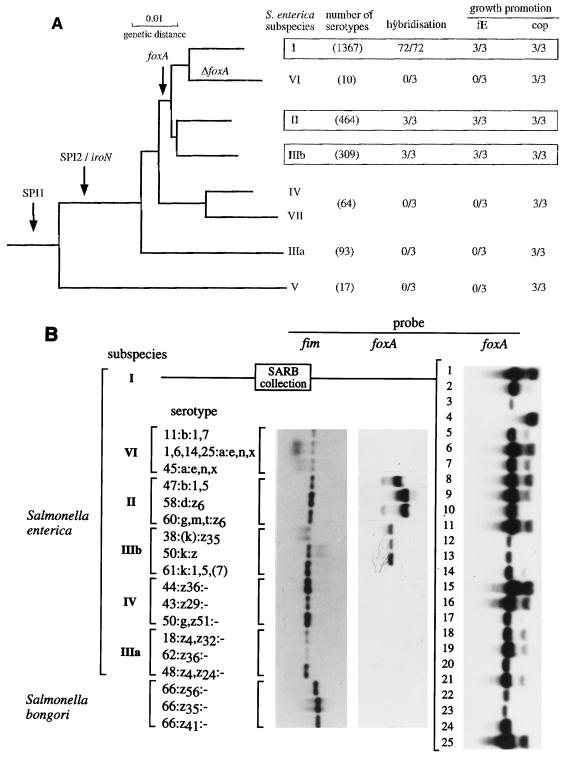
The requirement for FhuBCD and possibly TonB during uptake of iron complexed with ferrioxamines suggests that a receptor(s) for ferrioxamine binding is involved. We previously identified part of an open reading frame in the S. enterica serotype Typhimurium genome whose deduced amino acid sequence exhibited 45% identity with the amino acid sequence of the FoxA protein of Y. enterocolitica (44), an outer membrane protein involved in the transport of ferrioxamine B (1). Using the Salmonella sequence, we designed primers to generate a 287-bp PCR product (cloned in plasmid pRA17) corresponding to an intragenic fragment of the S. enterica serotype Typhimurium foxA gene. To determine how widespread the foxA homolog is among Salmonella isolates, chromosomal DNAs of representative strains were digested with EcoRI and analyzed by Southern blotting by using the insert fragment of plasmid pRA17 as the probe. All serotypes of S. enterica subspecies I, II, and IIIb were positive in this assay (Fig. 2B), while the DNA of strains belonging to subspecies IIIa, IV, VI, and VII and S. bongori did not hybridize with the foxA-specific probe. To test whether the distribution of foxA as determined by hybridization correlated with the ability to utilize ferrioxamines, growth stimulation of representative strains was assessed by performing bioassays on EWM. In each case, only isolates that were positive in hybridization experiments (isolates belonging to serotypes of subspecies I, II, and IIIb) were also positive in bioassays in which ferrioxamine E was the sole source of iron (Fig. 2A).
FIG. 2.
Distribution of the foxA gene and the ability to utilize ferrioxamine E as a sole source of iron among subspecies of the genus Salmonella. (A) Neighbor-joining dendrogram showing the relatedness of the various subspecies, based on variation in the combined coding sequences of five housekeeping genes of Salmonella spp. (8). The arrows indicate probable introduction by horizontal transfer of SPI1 and SPI2 and of the foxA gene; ΔfoxA indicates possible deletion of the foxA gene in the subspecies VI lineage. Note that Reeves et al. (35) did not differentiate S. enterica subspecies IV into subspecies IV and VII; therefore, the three serotypes tested were random examples of what was originally designated subspecies IV. The hybridization data are the number of strains that exhibited positive hybridization with the foxA-specific probe/total number of strains tested. The growth promotion data are the number of strains whose growth was promoted by ferrioxamine E (fE) or coprogen (cop) as a sole source of iron/total number of strains tested. (B) Southern blot analyses of chromosomal DNA of representative strains of each subspecies probed with a foxA-specific DNA fragment. For the SARB collection, only the first 25 of the 72 strains listed by Boyd et al. (7) were used, but all other isolates gave similar positive results with the foxA probe. For the representative serotypes of other subspecies, control hybridizations were performed by using a probe derived from the Salmonella fim operon (1) in order to validate negative foxA hybridizations.
Genetic analysis of the Salmonella foxA locus.
To confirm that the Salmonella DNA sequence homologous to Y. enterocolitica foxA plays a role in ferrioxamine uptake, a derivative of the enterobactin-deficient strain AR1258 (45) containing a pMAP insertion in the foxA homolog was constructed as previously described (44). This strain was designated RK809. The effect of the foxA::pMAP mutation was assessed by performing standard Vogel-Bonner medium diffusion plate bioassays. As noted above, growth of the foxA+ parent strain AR1258 was stimulated by coprogen, enterobactin, and ferrioxamines B, E, and G. Stimulation of the growth of the foxA::pMAP mutant RK809 by ferrioxamine B, however, was limited, and this strain did not grow at all in the presence of ferrioxamines E and G (Table 2), although it responded normally to coprogen and enterobactin. Growth assays in tryptone soy broth essentially confirmed the data obtained in the plate bioassays. Ferrioxamines B, E, and G supported growth of strain AR1258 (foxA+) in liquid culture (Fig. 3A) but not growth of the foxA mutant RK809 (Fig. 3B). These data suggest that the Salmonella FoxA protein acts as a receptor for all three ferrioxamines tested.
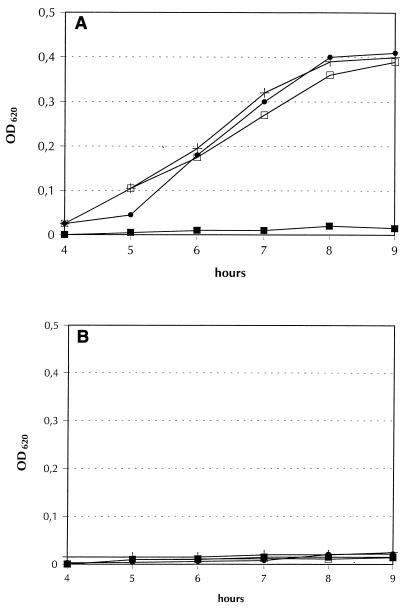
FIG. 3.
Growth of S. enterica serotype Typhimurium strain AR1258 (entB) (A) and its foxA derivative RK809 (B) in tryptone soy broth that was supplemented with 200 μM 2,2′-bipyridyl and ferrioxamine B (●), E (+), or G (□) (at a concentration of 1 μg/ml) or not supplemented (■). Optical densities at 620 nm (OD620) were determined at intervals during incubation with aeration by shaking at 37°C. The data are from a representative experiment; we performed at least five experiments in which similar results were obtained.
An alternative possibility is that the inability of RK809 to use ferrioxamines is due to polar effects of the pMAP insertion on downstream genes. To eliminate this possibility, we attempted to clone the entire foxA gene in order to test its ability to complement the insertion mutation but were not successful. Instead, we constructed a defined, nonpolar +2 frameshift mutation in the foxA gene as described above and tested the effect by performing diffusion plate bioassays on EWM supplemented with 2,2′-bipyridyl (Table 2). The phenotype of mutant strains BK102 and RK102 with regard to ferrioxamine uptake was essentially the same as that observed for strain RK809 carrying the potentially polar foxA mutation, confirming that FoxA acts as a receptor for all three ferrioxamines tested.
Sequence analysis of the Salmonella foxA locus.
For reasons that we cannot explain, it was not possible to clone the entire Salmonella foxA gene, and thus the nucleotide sequence of the gene remains incomplete. However, hybridization screening of libraries (in which an intragenic foxA fragment was used as the probe) did yield clones containing part of foxA that enabled us to add to the previously reported sequence (44). In particular, plasmid pRA5 contained a 2,227-bp insert encoding part of foxA and a previously unidentified open reading frame approximately 100 bp upstream of foxA. Sequence analysis of this insert fragment indicated that the ATG start codon of the Salmonella foxA gene is preceded by a putative ribosome binding site (AATAAA at positions 1098 to 1103) and a putative Fur box (GGTAATAATTCTTATTTAC at positions 1078 to 1097) that is identical at 12 of 19 bases to the consensus sequence (9, 13, 41). The Salmonella foxA coding sequence determined so far encodes a deduced amino acid sequence containing 374 residues that exhibits 40.7% identity with the amino acid sequence of the Y. enterocolitica FoxA protein and significant similarity to the amino acid sequences of a number of other TonB-dependent outer membrane receptor proteins. It was predicted that a signal peptide identified by using the SignalP program (26) would be cleaved between residues 30 and 31. TonB box I and TonB box III were identified at residues 6 to 14 and 111 to 140, respectively, of the mature protein on the basis of a comparison with other previously identified TonB box sequences (1). A comparison with other proteins suggested that TonB box II is probably encoded by part of foxA not present in pRA5.
Effect of foxA on the colonization of rabbit ileal loops.
Competitive colonization in the rabbit ileal loop anastomosis test (43, 47) was used to determine the potential role of the Salmonella FoxA protein in virulence. Approximately 1:1 mixtures of pairs of strains were inoculated into test intestinal loops, and a significant variation from equivalence among bacterial mixtures recovered from the lumen contents or associated with the lumen walls was considered an indication that there were differences in the colonizing abilities of the two strains. There was no significant difference in the recovery of strains TML and TML/nr, which ruled out the possibility that the mutation causing nalidixic acid resistance had any effect in the latter strain. However, recovery of the foxA mutant RK102, a derivative of TML/nr, was significantly less than recovery of strain TML both in the lumen contents and associated with intestinal loop tissue (Fig. 4), strongly suggesting that FoxA plays a role in intestinal colonization by Salmonella strains. Addition of 1 mg of ferrioxamine E per ml to mixed inocula did not have a significant effect on the competitive indices observed (data not shown).
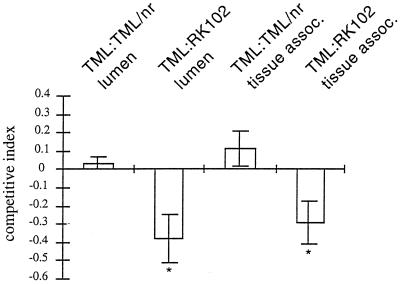
FIG. 4.
Comparison of colonization of rabbit ileal loops by S. enterica serovar Typhimurium strain TML with colonization of rabbit ileal loops by derivative strains TML/nr and RK102 (foxA). We performed competitive colonization experiments involving wild-type strain S. enterica TML and either the nalidixic acid-resistant derivative strain TML/nr or the nalidixic acid-resistant foxA mutant strain RK102 by using a total of 10 ileal loops in two rabbits. Approximately 5-cm-long loops were inoculated with 1:1 mixtures containing 107 or 108 cells. After 28 h the proportion of nalidixic acid-resistant coliform bacteria present in the lumen and associated with the gut tissue was determined. The proportion of nalidixic acid-resistant coliform bacteria recovered was expressed as a log10 value (competitive index) and was subjected to a statistical analysis in which the Student t test was used. The asterisks indicate competitive indices significantly different (P > 0.01) from the input ratio (log10 1 = 0).
Mouse virulence of foxA+ and foxA strains.
To determine if a mutation in foxA affects the ability of S. enterica serotype Typhimurium to cause lethal infections in mice, LD50s were determined after intragastric inoculation. The LD50 of strain SL1344/nr was calculated to be 105 CFU, which was approximately equal to the LD50 reported previously for the nalidixic acid-sensitive parent strain SL1344 (45). In contrast, the LD50 of the foxA mutant strain BK102 (a derivative of SL1344/nr) was >109 CFU, and no deaths occurred up to 28 days postinoculation at any inoculum dose.
Five mice from each group intragastrically inoculated with 103, 104, or 105 CFU and four mice from each group inoculated with 106, 107, 108, or 109 CFU of strain BK102 were challenged with a single intragastric dose containing approximately 5 × 108 CFU of the fully virulent strain SL1344/nr in 0.2 ml of phosphate-buffered saline on day 28 after the initial inoculation. The challenge dose, which was greater than the LD50, would have resulted in the death of naive mice within 8 days. The number of deaths in each group was recorded 10 days after the challenge (Table 3). There were few or no survivors in the groups originally inoculated with 103, 104, or 105 CFU of strain BK102, indicating that there was little or no protection. However, there was significant protection in groups of mice originally inoculated with >105 CFU of strain BK102.
TABLE 3.
Protection of mice previously inoculated with strain BK102 against subsequent challenge with S. enterica serotype Typhimurium strain SL1344/nr
| BK102 inoculum (CFU)a | No. of survivors/no. of mice testedb |
|---|---|
| 109 | 3/4 |
| 108 | 3/4 |
| 107 | 4/4 |
| 106 | 3/4 |
| 105 | 0/5 |
| 104 | 2/5 |
| 103 | 0/5 |
The BK102 inoculum was administered 28 days prior to challenge with strain SL1344/nr.
Number of mice surviving 10 days after inoculation with 5 × 108 CFU of SL1344/nr/number of mice tested.
LD50s were also determined following intravenous inoculation of mice; consistent with previous reports (45), the LD50 of the parent strain SL1344/nr was <10 CFU, but none of the mice inoculated with 104 CFU of the foxA strain died, indicating that the LD50 of BK102 was >104 CFU.
DISCUSSION
The foxA gene encoding the Salmonella ferrioxamine receptor was first identified as a cloned subgenic fragment having a deduced amino acid sequence that was 45% identical to the amino acid sequence of the specific outer membrane ferrioxamine receptor of Y. enterocolitica (44). Phenotypic analysis of foxA mutants carrying either a pMAP insertion (potentially polar) mutation or a defined nonpolar frameshift mutation demonstrated that the FoxA protein is required for utilization of all three ferrioxamine molecules tested. Moreover, using the subgenic foxA fragment as a probe, we showed that the foxA gene is not ubiquitous among Salmonella serotypes but is limited to subspecies I, II, and IIIb. The presence of foxA correlated with growth stimulation by ferrioxamine E in bioassays; strains that were negative as determined by hybridization were also negative in bioassays performed with ferrioxamine E. Although this observation may suggest that inclusion of ferrioxamines as medium supplements would not be useful in all cases, it is important to note that the three subspecies that express foxA account for >99% of the clinical isolates and 2,140 of 2,324 of the serotypes described so far. We therefore recommend that ferrioxamine E should be included in preenrichment and selection protocols for rapid identification of the vast majority of clinically significant serotypes of the genus Salmonella in food and feed.
If it is assumed that subspecies I, II, and IIIb contain a greater diversity of serotypes because they have adapted more successfully to the pathogenic lifestyle, then it is possible that foxA (or a closely linked determinant) may be selectively advantageous for Salmonella pathogenicity. It has been suggested that limited phylogenetic distribution is characteristic of introduction of a genetic determinant into a genome by horizontal gene transfer (28). Thus, ancestral Salmonella serotypes apparently became more successful by acquiring genetic determinants, such as SPI1 and SPI2, by bacteriophage- or plasmid-mediated transfer from other organisms. SPI1 is present in virtually all Salmonella serotypes but not in E. coli and so was probably acquired shortly after the taxa diverged from their common ancestor. SPI1 encodes determinants required for invasion of epithelial cells (10, 25), a basic requirement for Salmonella pathogenesis. Similarly, since SPI2 is present in all serotypes of S. enterica but not in S. bongori, it probably entered the S. enterica lineage shortly after S. enterica and S. bongori split from their common ancestor (29). Acquisition of SPI2, which is required for systemic infection (16, 30, 42), resulted in a more successful lineage, as judged by adaptive radiation into a greater range of hosts, including both cold- and warm-blooded animals, and a greater diversity of serotypes. It is tempting to speculate that introduction of foxA, which probably occurred after the split leading to subspecies I, II, IIIb, and VI on the one hand and subspecies IV and VII on the other, may have contributed to the further radiation of serotypes belonging to subspecies I, II, and IIIb. Fortuitous loss of the foxA gene by deletion, as in subspecies VI, could lead to underrepresentation of serotypes of a subspecies if the selective advantage is lost concomitantly.
As is typical for siderophores of enteric bacteria, utilization of ferrioxamine G by Salmonella strains is completely dependent on the energy-transducing inner membrane protein TonB for transport across the outer membrane following interaction with FoxA. In contrast, and very unusually, utilization of ferrioxamines B and E occurs in the absence of functional TonB protein, albeit at lower efficiency. One of the few other examples of TonB-independent iron supply in Salmonella strains is the iron supply mediated by the enterobactin precursor 2,3-dihydroxybenzoic acid (18); in this case it is believed that the small size of the molecule compared to other siderophores allows it to diffuse freely through the outer membrane porins. The molecular sizes of ferrioxamines B, E, and G, however, are not significantly different from one another, and so the mechanism by which ferrioxamines B and E traverse the outer membrane in the absence of TonB is not clear. This mechanism appears to depend on the FoxA receptor, however, since no leaky phenotype was observed in any strain containing a mutation in the foxA gene; this essentially eliminates the possibility that passive diffusion through porins occurs. Transport of ferrioxamines B, E, and G across the inner membrane involves the periplasmic binding protein-dependent permease comprising the FhuBCD proteins, which is the same route as the route for other hydroxamate siderophores in E. coli and Salmonella strains (21).
It is not clear why Salmonella strains express a receptor for siderophores that they do not synthesize. We assume that ferrioxamines are not generally available in the vertebrate hosts, except possibly in the intestinal lumen, where they may be secreted by members of the normal gut flora. However, although some members of the family Enterobacteriaceae do indeed synthesize ferrioxamines as their principal siderophores (37), there is no direct evidence that ferrioxamines are secreted by gut commensal species. Nevertheless, the role of FoxA was investigated in the context of experimental infections by S. enterica serotype Typhimurium in rabbit ileal loops and in the murine typhoid model. The former model has been reported to be a good model for enteric infection, which allows measurement of intestinal colonization and fluid secretion, while the latter model is widely used as a model for typhoid fever. In each case, foxA+ and defined foxA derivatives of strains known to respond well in either model were used (TML in rabbits [12] and SL1344 in mice [17]). We observed that there was a small but significant decrease in the ability of the foxA mutant of TML to compete with the wild-type strain during colonization of rabbit ileal loops. Addition of ferrioxamine E with the inoculum did not affect the competitive index. This may suggest that sufficient ferrioxamine for maximal growth was already present in the loops, perhaps due to secretion by commensal bacterial species. However, it was not possible to demonstrate the presence of ferrioxamines in bioassays with appropriate indicator strains (data not shown). A more likely possibility is that the selective advantage of FoxA is not due simply to its function as a receptor for the transport of ferrioxamines as sources of iron. This possibility is strongly supported by the data obtained from the mouse typhoid model, in which a disease that occurs predominantly in the tissues comprising the reticuloendothelial system would not be expected to be abrogated by a mutation in genes involved in the early colonization stages of infection. Attenuation of the foxA mutants following both oral inoculation and intravenous inoculation was surprising and cannot be explained simply on the basis of the inability of these mutants to utilize ferrioxamines as sources of iron. The increases in LD50s (>10,000-fold) were in fact considerably greater than the increases in LD50s observed for mutations in the lpf and inv genes (10- to 50-fold with oral inoculation and no effect with parenteral inoculation), both of which have effects during the early stages of infection, including invasion of the intestinal mucosa (5, 11). Furthermore, a strain lacking a functional TonB protein was mildly attenuated in experimental infections of mice (there was an approximately fivefold increase in the LD50), but only with oral inoculation, and bacteria were recovered in significantly smaller numbers from the Peyer’s patches, the normal focus of S. enterica serotype Typhimurium infections during the early stages of infection in the murine host (45). Although the reason for the significantly reduced virulence of foxA mutants is obscure, a number of possibilities may be considered. First, since the entire foxA gene could not be cloned and thus it was not possible to complement the frameshift mutation in virulence assays, indirect effects of the mutation (i.e., effects other than loss of the foxA function) cannot be eliminated (for example, expression of a truncated form of FoxA, which may compromise the integrity of the outer membrane). Perhaps more likely is the possibility that FoxA has an essential role in addition to its role in the uptake of ferrioxamines, just as, for example, OmpC has been implicated in bacterial association and internalization by macrophages (25a). The expression of foxA is derepressed under iron-limiting conditions (44); it is therefore probable that in the interstitial medium and blood serum where Salmonella cells initially interact with macrophages, the presence of transferrin results in high levels of FoxA in the outer membrane. Defining the precise role of FoxA in these circumstances requires further analysis; experiments to determine the mechanism of attenuation in foxA mutants and the ability of these mutants to protect against challenge with a fully virulent foxA+ strain are in progress in our laboratories.
ACKNOWLEDGMENTS
This work was supported by the British-German Academic Research Collaboration (ARC) Programme of the British Council and the Deutscher Akademischer Austauschdienst (ARC project 354 awarded to P.H.W. and R.R.) and by Medical Research Council Collaborative Studentship CS 93 15 awarded to R.A.K. in association with Medeva Group Research.
We thank R. Haigh for the gift of plasmid pRDH10 and E. coli SM10λpir, H. P. Schnebli for providing ferrioxamines B and E, and G. Winkelmann for providing coprogen.
REFERENCES
- 1.Bäumler A, Hantke K. Ferrioxamine uptake in Yersinia enterocolitica: characterization of the receptor protein FoxA. Mol Microbiol. 1992;6:1309–1321. doi: 10.1111/j.1365-2958.1992.tb00852.x. [DOI] [PubMed] [Google Scholar]
- 2.Bäumler A J, Gilde A J, Tsolis R M, van der Velden A W, Ahmer B M M, Heffron F. Contribution of horizontal gene transfer and deletion events to development of distinctive patterns of fimbrial operons during the evolution of Salmonella serotypes. J Bacteriol. 1997;179:317–322. doi: 10.1128/jb.179.2.317-322.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bäumler A J, Heffron F, Reissbrodt R. Rapid detection of Salmonella enterica with primers specific for iroB. J Clin Microbiol. 1997;35:1224–1230. doi: 10.1128/jcm.35.5.1224-1230.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bäumler A J, Norris T L, Lasco T, Voigt W, Reissbrodt R, Rabsch W, Heffron F. IroN, a novel outer membrane siderophore receptor characteristic of Salmonella enterica. J Bacteriol. 1998;180:1446–1453. doi: 10.1128/jb.180.6.1446-1453.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bäumler A J, Tsolis R M, Heffron F. The lpf fimbrial operon mediates adhesion to murine Peyer’s patches. Proc Natl Acad Sci USA. 1996;93:279–283. doi: 10.1073/pnas.93.1.279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Blomfield I C, Vaughn V, Rest R F, Eisenstein B I. Allelic exchange in Escherichia coli using the Bacillus subtilis sacB gene and a temperature-sensitive pSC101 replicon. Mol Microbiol. 1991;5:1447–1457. doi: 10.1111/j.1365-2958.1991.tb00791.x. [DOI] [PubMed] [Google Scholar]
- 7.Boyd E F, Wang F-S, Beltran P, Plock S A, Nelson K, Selander R K. Salmonella reference collection B (SARB): strains of 37 serovars of subspecies I. J Gen Microbiol. 1993;139:1125–1132. doi: 10.1099/00221287-139-6-1125. [DOI] [PubMed] [Google Scholar]
- 8.Boyd E F, Wang F-S, Whittam T S, Selander R K. Molecular genetic relationship of the salmonellae. Appl Environ Microbiol. 1996;62:804–808. doi: 10.1128/aem.62.3.804-808.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Brickman T J, Ozenberger B B, McIntosh M A. Regulation of divergent transcription from the iron responsive fepB-entC promoter-operator regions in Escherichia coli. J Mol Biol. 1990;216:883–895. doi: 10.1016/0022-2836(90)90229-F. [DOI] [PubMed] [Google Scholar]
- 10.Galán J E. Molecular-genetic bases of Salmonella entry into host cells. Mol Microbiol. 1996;20:263–271. doi: 10.1111/j.1365-2958.1996.tb02615.x. [DOI] [PubMed] [Google Scholar]
- 11.Galán J E, Curtiss R., III Cloning and molecular characterization of genes whose products allow Salmonella typhimurium to penetrate tissue culture cells. Proc Natl Acad Sci USA. 1989;86:6383–6387. doi: 10.1073/pnas.86.16.6383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gianella R A, Broitman S A, Zamcheck N. Salmonella enteritis. I. Role of reduced gastric secretion in pathogenesis. Am J Dig Dis. 1971;16:1000–1006. doi: 10.1007/BF02235012. [DOI] [PubMed] [Google Scholar]
- 13.Griggs D W, Konisky J. Mechanism for iron regulated transcription for the Escherichia coli cir gene: metal-dependent binding of the Fur protein to the promoters. J Bacteriol. 1989;171:1048–1052. doi: 10.1128/jb.171.2.1048-1054.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Groisman E A, Ochman H. How salmonella became a pathogen. Trends Microbiol. 1997;5:343–349. doi: 10.1016/S0966-842X(97)01099-8. [DOI] [PubMed] [Google Scholar]
- 15.Hantke K. Identification of an iron uptake system specific for coprogen and rhodotorulic acid in Escherichia coli K12. Mol Gen Genet. 1983;191:301–306. doi: 10.1007/BF00334830. [DOI] [PubMed] [Google Scholar]
- 16.Hensel M, Shea J E, Gleeson A C, Jones M D, Dalton E, Holden D W. Simultaneous identification of bacterial virulence genes by negative selection. Science. 1995;269:400–403. doi: 10.1126/science.7618105. [DOI] [PubMed] [Google Scholar]
- 17.Hoiseth S K, Stocker B A. Aromatic-dependent Salmonella typhimurium are non-virulent and effective as live vaccines. Nature. 1981;291:238–239. doi: 10.1038/291238a0. [DOI] [PubMed] [Google Scholar]
- 18.Kingsley R, Rabsch W, Roberts M, Reissbrodt R, Williams P H. TonB-dependent iron supply in Salmonella by α-ketoacids and α-hydroxyacids. FEMS Microbiol Lett. 1996;140:65–70. doi: 10.1111/j.1574-6968.1996.tb08316.x. [DOI] [PubMed] [Google Scholar]
- 19.Kingsley R, Rabsch W, Stephens P, Roberts M, Reissbrodt R, Williams P H. Iron supplying systems of Salmonella in diagnostics, epidemiology and infection. FEMS Immunol Med Microbiol. 1995;11:257–264. doi: 10.1111/j.1574-695X.1995.tb00154.x. [DOI] [PubMed] [Google Scholar]
- 20.Kolter R, Inuzuka M, Helinski D R. Trans-complementation-dependent replication of a low molecular weight origin fragment from plasmid R6K. Cell. 1978;15:1199–1208. doi: 10.1016/0092-8674(78)90046-6. [DOI] [PubMed] [Google Scholar]
- 21.Koster W. Iron(III) hydroxamate transport across the cytoplasmic membrane in Escherichia coli. Biol Metals. 1991;4:23–32. doi: 10.1007/BF01135553. [DOI] [PubMed] [Google Scholar]
- 22.Luckey M, Pollack J R, Wayne R, Ames B N, Neilands J B. Iron uptake in Salmonella typhimurium: utilization of exogenous siderophores as iron carriers. J Bacteriol. 1972;111:731–738. doi: 10.1128/jb.111.3.731-738.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Maloy S R, Stewart V J, Taylor R K. Experiments in bacterial pathogenesis. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1994. [Google Scholar]
- 24.Maniatis T, Sambrook J, Fritsch E F. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 25.Mills V. Tissue culture invasion—fact or fiction. Trends Microbiol. 1995;3:69–71. doi: 10.1016/s0966-842x(00)88878-2. [DOI] [PubMed] [Google Scholar]
- 25a.Negm R S, Pistole T G. Abstracts of the 96th General Meeting of the American Society for Microbiology 1996. Washington, D.C: American Society for Microbiology; 1996. OmpC::Tn10 mutagenesis of Salmonella typhimurium and its effect on the adherence to macrophages; p. 258. [Google Scholar]
- 26.Nielsen H, Engelbrecht J, Brunak S, von Heijne G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 1997;10:1–6. doi: 10.1093/protein/10.1.1. [DOI] [PubMed] [Google Scholar]
- 27.Nikaido H. Outer membrane. In: Neidhardt F C, Curtiss III R, Ingraham J L, Lin E C C, Low K B, Megasanik B, Reznikoff W S, Riley M, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella: cellular and molecular biology. Vol. 1. Washington, D.C: American Society for Microbiology; 1996. pp. 29–47. [Google Scholar]
- 28.Ochman H, Bergthorsson U. Genome evolution in enteric bacteria. Curr Opin Genet Dev. 1995;5:734–738. doi: 10.1016/0959-437x(95)80005-p. [DOI] [PubMed] [Google Scholar]
- 29.Ochman H, Groisman E A. Distribution of pathogenicity islands in Salmonella spp. Infect Immun. 1996;64:5410–5412. doi: 10.1128/iai.64.12.5410-5412.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ochman H, Soncini F C, Solomon F, Groisman E A. Identification of a pathogenicity island required for Salmonella survival in host-cells. Proc Natl Acad Sci USA. 1996;93:7800–7804. doi: 10.1073/pnas.93.15.7800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ochman H, Wilson A C. Evolution in bacteria: evidence for a universal substitution rate in cellular genomes. J Mol Evol. 1987;26:74–86. doi: 10.1007/BF02111283. [DOI] [PubMed] [Google Scholar]
- 32.Pless P, Reissbrodt R. Improvement of Salmonella detection on motility enrichment media by ferrioxamine E-supplementation of pre-enrichment culture. Int J Food Microbiol. 1995;27:147–159. doi: 10.1016/0168-1605(94)00160-8. [DOI] [PubMed] [Google Scholar]
- 33.Pollack J R, Ames B N, Neilands J B. Iron transport in Salmonella typhimurium: mutants blocked in the biosynthesis of enterobactin. J Bacteriol. 1970;104:635–639. doi: 10.1128/jb.104.2.635-639.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Reed L J, Muench H. A simple method for estimating fifty percent endpoints. Am J Hyg. 1938;27:493–497. [Google Scholar]
- 35.Reeves M W, Evins G M, Heiba A A, Plikaytis B D, Farmer J J., III Clonal nature of Salmonella typhi and its genetic relatedness to other salmonellae as shown by multilocus enzyme electrophoresis, and proposal of Salmonella bongori comb. nov. J Clin Microbiol. 1989;27:313–320. doi: 10.1128/jcm.27.2.313-320.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Reissbrodt R, Rabsch W. Selective pre-enrichment of Salmonella from eggs by siderophore supplements. Zentralbl Bakteriol Parasitenkd Infektionskr Hyg Abt 1 Orig. 1993;279:344–353. doi: 10.1016/s0934-8840(11)80367-1. [DOI] [PubMed] [Google Scholar]
- 37.Reissbrodt R, Rabsch W, Chapeanrouge A, Jung G, Winkelmann G. Isolation and identification of ferrioxamines E and G in Hafnia alvei. Biol Metals. 1990;3:54–60. [Google Scholar]
- 38.Reissbrodt R, Vielitz E, Kormann E, Rabsch W, Kuhn H. Ferrioxamine E-supplemented pre-enrichment and enrichment media improve various isolation methods for Salmonella. Int J Food Microbiol. 1996;29:81–91. doi: 10.1016/0168-1605(95)00024-0. [DOI] [PubMed] [Google Scholar]
- 39.Schmeiger H. Phage P22 mutants with increased or decreased transduction abilities. Mol Gen Genet. 1972;119:75–81. doi: 10.1007/BF00270447. [DOI] [PubMed] [Google Scholar]
- 40.Selbitschka W, Niemann S, Pühler A. Construction of gene replacement vectors for Gram negative bacteria using a genetically modified sacRB gene as a positive selection marker. Appl Microbiol Biotechnol. 1993;38:615–618. [Google Scholar]
- 41.Shea C M, McIntosh M A. Nucleotide sequence and genetic organization of the ferric enterobactin transport system: homology to other periplasmic binding protein-dependent systems in Escherichia coli. Mol Microbiol. 1991;5:1415–1428. doi: 10.1111/j.1365-2958.1991.tb00788.x. [DOI] [PubMed] [Google Scholar]
- 42.Shea J E, Hensel M, Gleeson C, Holden D W. Identification of a virulence locus encoding a second type III secretion system in Salmonella typhimurium. Proc Natl Acad Sci USA. 1996;93:2593–2597. doi: 10.1073/pnas.93.6.2593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Stephen J, Wallis T S, Starkey W G, Candy D C A, Osborne M P, Haddon S. Microbial toxins and diarrhoeal disease. London, United Kingdom: Pitman; 1985. Salmonellosis: in retrospect and prospect; pp. 175–192. [PubMed] [Google Scholar]
- 44.Tsolis R, Bäumler A J, Stojiljkovic I, Heffron F. Fur regulon of Salmonella typhimurium: identification of new iron-regulated genes. J Bacteriol. 1995;177:4628–4637. doi: 10.1128/jb.177.16.4628-4637.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tsolis R M, Bäumler A J, Heffron F, Stojiljkovic I. Contribution of TonB- and Feo-mediated iron uptake to growth of Salmonella typhimurium in the mouse. Infect Immun. 1996;64:4549–4556. doi: 10.1128/iai.64.11.4549-4556.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wagegg W, Braun V. Ferric citrate transport in Escherichia coli requires outer membrane receptor protein FecA. J Bacteriol. 1981;145:156–163. doi: 10.1128/jb.145.1.156-163.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Wallis T S. Interactions of bacteria and their products in whole animal systems. Methods Microbiol. 1998;27:103–112. [Google Scholar]
- 48.Young I G. Preparation of enterochelin from Escherichia coli. Prep Biochem. 1976;6:123–131. doi: 10.1080/00327487608061607. [DOI] [PubMed] [Google Scholar]