Abstract
Background
Long non-coding RNA P73 antisense RNA 1 T (non-protein coding), also known as Lnc RNA TP73-AS1, is dysregulated in various tumors but the correlation between its expression and clinicopathological parameters and/or prognoses in cancer patients is inconclusive. Here, we performed a meta-analysis to evaluate the prognostic value of Lnc RNA TP73-AS1 for malignancies.
Methods
We systematically searched four online databases including PubMed, the Web of Science, Embase, and the Cochrane Library for eligible articles published up to June 29/2020. Odds ratios (ORs) and Pooled hazard ratios (HRs) with 95% confidence intervals (95% CIs) were used to assess the association of TP73-AS1 expression with prognostic and clinicopathological parameters. We further validated TP73-AS1 expression in various malignancies and its potential prognostic value using the GEPIA online database. We predicted potential biological processes and relevant signal mechanisms through the public databases.
Results
A total of 26 studies examining 14 cancers were analyzed to evaluate the relationship between TP73-AS1 expression, clinicopathological features and prognostic indicators. The results indicated that TP73-AS1 expression markedly correlates with TNM stage (OR = 3.27,95% CI:2.43–4.39, P < 0.00001), tumor size (OR = 3.00, 95%CI:2.08–4.35, P < 0.00001), lymph node metastasis (OR = 2.77, 95%CI:1.42–5.38,P < 0.00001) and distant metastasis (OR = 4.50,95%CI:2.
62–7.73,P < 0.00001). No correlation with age (OR = 1.12,95%CI:0.77–1.64, P > 0.05), gender (OR = 1.08, 95%CI:0.84–1.38, P > 0.05) or differentiation (OR = 1.39, 95%CI:0.71–2.70, P = 0.340) was observed. TP73-AS1 overexpression was a biomarker of poor Overall survival(OS)(HR = 1.85,95%CI:1.53–2.22, P < 0.00001) and Disease-Free-Survival (DFS) (HR = 1.57,95%CI:1.03–2.42, P < 0.05). Dysregulated TP73-AS1 expression and its prognostic value in various cancers was validated based on The Cancer Genome Atlas (TCGA). Further biological function predictions indicated that TP73-AS1 was involved in pro-oncogenic signaling.
Conclusions
The upregulation of Lnc RNA TP73-AS1 was related to detrimental clinicopathological parameters and can be considered an indicator of poor prognosis for cancer malignancies.
Keywords: Lnc RNA TP73-AS1, Clinicopathological parameters, Prognosis, malignancies, Meta-analysis, Bioinformatics
Introduction
Cancer is a global health problem with increasing morbidity and mortality with an additional economic burden to patients worldwide. According to recent cancer statistics, 1,762,450 new cancer cases and 606,880 cancer-related deaths occurred in the United States in 2019 [1]. Medical advances have allowed for the standardization of tumor treatments that can delay disease progress. However, patients with advanced cancers have a poor prognosis and quality of life. New cancer markers to enable early diagnosis and prognosis predictions are therefore urgently required.
Long non-coding RNAs (lncRNAs) were originally regarded as transcriptional noise and are defined as RNA transcripts greater than 200 nucleotides that do not encode proteins [2]. LncRNAs have been found to play crucial roles in modulating diverse aspects of tumorigenesis, including cell growth, invasion, metastasis and survival [3, 4]. Overexpression of the lncRNA TUC338 correlates with poor OS in prostate carcinoma [5]. Furthermore, the upregulation of LINC00520 is associated with a poor prognosis in patients with NPC [6]. LncRNAs therefore participate in tumorigenesis and can be used to predict the prognosis of malignancies.
TP73-AS1, located at region lp36.32, is a newly discovered lncRNA, also termed PDAM or KIAA0495 [7]. LncRNA TP73-AS1 expression is dysregulated in various human malignancies, including hepatocellular carcinoma, osteosarcoma and gastric cancer. Furthermore, previous studies indicate a correlation between abnormal TP73-AS1 expression and clinicopathological features and prognoses. However, owing to the small number of clinical patients studied, the significance of these associations is poorly characterized. Larger sample sizes are required.
In this study, we evaluated the utility of lncRNA TP73-AS1 as a prognostic cancer biomarker. A meta-analysis of all correlative studies of lncRNA TP73-AS1 and cancer was performed to assess the association between TP73-AS1 expression and clinicopathological characteristics including age, gender, TNM stage, tumor size, lymph node metastasis, distant metastasis, and differentiation. Our ultimate aim was to assess the value of lncRNA TP73-AS1 as an indicator of survival for different malignancies.
Methods and materials
Search strategy
We searched PubMed, Web of Science, Embase, and the Cochrane Library using the search terms “PDAM” OR “TP73-AS1” OR “KIAA0495” OR “PDAM lncRNA” OR “TP73-AS1 lncRNA” OR “KIAA0495 lncRNA” OR “lncRNA KIAA0495” OR “lncRNA PDAM” OR “lncRNA TP73-AS1” OR “long noncoding RNA TP73-AS1” and “Neoplasia” OR “Neoplasia’s” OR “Neoplasm” OR “Tumors” OR “Tumor” OR “Cancer” OR “Cancers” OR “Malignancy” OR “Malignancies” OR “Malignant Neoplasms” OR “Malignant Neoplasm” OR “Neoplasm, Malignant” OR “Neoplasms, Malignant” OR “Benign Neoplasms” OR “Neoplasms, Benign” OR “Benign Neoplasm” OR “Neoplasm, Benign” as keywords to identify all relevant publications. The search included manuscripts published up until June 29th 2020. Only articles written in English were included.
Inclusion and exclusion criteria
Inclusion criteria were as follows: (1) original primary research articles; (2) elucidated associations between TP73-AS1 expression and clinicopathological characterization or prognosis of various solid human malignancies; (3) division of patients into two groups according to TP73-AS1 expression from tissue sample analysis; (4) expression detectable by qRT-PCR; (5) reported data or Kaplan–Meier curves available for calculation of the hazard ratios (HR) (95% CI) for survival.
Exclusion criteria were as follows: (1) non-primary research articles, including case reports, reviews, conference abstracts, or letters; (2) duplicate studies; (3) studies lacking sufficient data or p-values to calculate the HR (95% CI); (4) nonhuman studies.
Data extraction and quality assessments
Studies were selected by two reviewers based on the inclusion and exclusion criteria. Disagreements were resolved by a third reviewer. The following information was collected from each study: author last name, publication year, country, sample type and detection method, criteria for patient categorization based on TP73-AS1 expression level, the hazard ratio (HR), corresponding 95% confident interval (CI) of overall survival (OS), disease free survival (DFS) and follow-up time. The HR (95% CI) of survival was obtained directly from the study direct or calculated using Engauge Digitizer 4.1 software if Kaplan–Meier curves were available.
The Newcastle–Ottawa quality assessment scale (NOS) that considers selection, outcome, and comparability, was used to assess the quality of the studies for inclusion. Manuscripts were given NOS scores from 0 to 9, with scores ≥ 6 showing the potential to compromise the study.
Statistical analysis
Pooled odds ratios (ORs) with 95% confidence intervals (CIs) were used to assess potential associations between of TP73-AS1 expression and clinicopathological features. Pooled HR values (95% CI) were calculated to identify TP73-AS1 as a prognostic biomarker in tumors. I2 statistics and chi-square Q tests were used to evaluate heterogeneity amongst eligible studies. A Chi-squared test of P < 0.10 or I2 > 50% indicated heterogeneity between the studies. A fixed model was used to integrate the results at an I2 ≤ 50%. Otherwise, the random-effects model was selected. Sensitivity analysis was performed to investigate the stability of the results. Publication bias was evaluated through a Begg’s assessments. STATA software version 12.0 was used for all statistical analysis. Significant differences were defined as P-values < 0.05.
TP73-AS1 expression for prognostic predictions
We compared the expression of TP73-AS1 in diverse tumor tissues with normal tissue, and validated its prognostic role using Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancer-pku.cn/), an online database from The Cancer Genome Atlas (TCGA) (https://tcga-data.nci.nih.gov). Targetscan (http://www.targetscan.org/vert_72/), mirdb (http://mirdb.org/), mirtarbase (http://mirtarbase.mbc.nctu.edu.tw/php/index.php), mircode (http://www.mircode.org/) were used to identify TP73-AS1 relevant ceRNAs. Cytoscape software was used to construct visualized lncRNA-miRNA-mRNA networks. TP73-AS1 biological functions and cancer-related pathways were predicted according to the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) via R package clusterprofiler.
Results
Search results
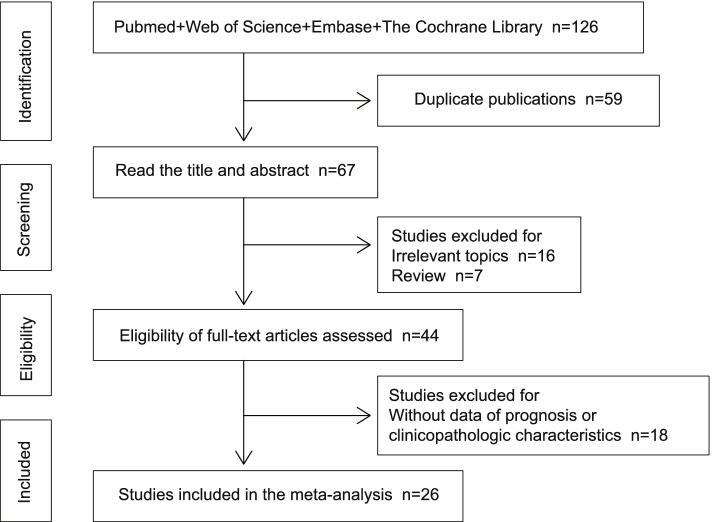
A total of 126 relevant publications were acquired after searching PubMed, Web of Science, Embase, and the Cochrane Library. Following the removal of duplicate publications, 67 studies were considered. Titles and abstracts were reviewed, and 16 non-relevant papers and 7 review articles were excluded. Further screening of the full texts of the 44 remaining articles led to the elimination of 18 reports due to a lack of study data regarding prognoses or clinicopathologic characteristics. Finally, 26 studies [8–33] from 2017 to 2020 met the criteria for the meta-analysis. Figure 1 describes the selection process for the included publications.
Fig. 1.
Screening process of the included studies
Included studies
The characteristics of the 26 included articles are summarized in Table 1. A total of 1770 patients with sample sizes ranging from 36 to 132 were evaluated and divided into high and low TP73-AS1 expression groups according to the mean cut-off value. All studies were performed in China and published from 2017 to 2020. The studies examined a wide variety of cancers, including hepatocellular carcinoma (2 studies), osteosarcoma (2 studies), gastric cancer (5 studies), ovarian cancer (2 studies), clear cell renal cell carcinoma (1 study), bladder cancer (1 study), breast cancer (3 studies), cholangiocarcinoma (1 study), lung cancer (3 studies), brain glioma (1 study), pancreatic cancer (1 study), colorectal cancer (2 studies),cervical cancer (1 study) and retinoblastoma (1 study). The expression of TP73-AS1 in tissues was through qRT-PCR analysis. OS data were reported in 19 studies, with 3 studies reporting DFS. The NOS assessment scores of the included studies ranged from 6 to 8.
Table 1.
Main characteristics of the studies included in the meta-analysis
| Author | Year | Country | Cancer | Sample | Method | patients number | OS | DFS |
Follow up (months) |
Data extraction method | NOS | Ref | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| high expression | low expression | total | HR (95% CL) | P | HR (95% CL) | P | ||||||||||
| Li | 2017 | China | hepatocellular carcinoma | tissue | qRT-PCR | 42 | 42 | 84 | 2.25(1.14–4.43) | 0.019 | NM | NM | 25 | reported | 8 | [8] |
| Chen | 2018 | China | osteosarcoma | tissue | qRT-PCR | 66 | 66 | 132 | 1.90(1.15–3.13) | 0.012 | NM | NM | 72 | reported | 8 | [27] |
| Ding | 2018 | China | gastric cancer | tissue | qRT-PCR | 38 | 34 | 72 | 1.19(0.47–3.03) | 0.710 | NM | NM | 60 | K-M | 7 | [11] |
| Li | 2018 | China | ovarian cancer | tissue | qRT-PCR | 36 | 26 | 62 | 2.16(1.05–4.46) | 0.036 | NM | NM | 60 | K-M | 7 | [28] |
| Liu | 2018 | China | ccRCC | tissue | qRT-PCR | 24 | 16 | 40 | 1.00(0.10–9.97) | 0.999 | 1.20(0.21–6.81) | 0.840 | 50 | K-M | 6 | [15] |
| Peng | 2018 | China | gastric cancer | tissue | qRT-PCR | 27 | 31 | 58 | 2.49(1.06–5.83) | 0.036 | NM | NM | 60 | K-M | 7 | [12] |
| Tuo | 2018 | China | bladder cancer | tissue | qRT-PCR | 64 | 64 | 128 | 0.79(0.24–2.63) | 0.706 | 0.83(0.37–1.84) | 0.639 | 60 | K-M | 7 | [16] |
| Wang | 2018 | china | ovarian cancer | tissue | qRT-PCR | 30 | 30 | 60 | NM | NM | NM | NM | NM | NM | 6 | [17] |
| Wang | 2018 | China | gastric cancer | tissue | qRT-PCR | 30 | 34 | 64 | 1.82(1.09–3.04) | 0.022 | 2.14(1.26–3.63) | 0.005 | 70 | K-M | 7 | [13] |
| Yang | 2018 | China | osteosarcoma | tissue | qRT-PCR | 23 | 23 | 46 | 1.92(0.71–5.18) | 0.199 | NM | NM | 50 | K-M | 6 | [10] |
| Yao | 2018 | China | breast cancer | tissue | qRT-PCR | 18 | 18 | 36 | 3.34(1.03–10.82) | 0.045 | NM | NM | 50 | reported | 8 | [18] |
| Yao | 2018 | China | cholangiocarcinoma | tissue | qRT-PCR | 41 | 34 | 75 | NM | NM | NM | NM | NM | NM | 6 | [29] |
| Zhang | 2018 | China | NSCLC | tissue | qRT-PCR | 22 | 23 | 45 | 1.76(0.65–4.74) | 0.267 | NM | NM | 60 | K-M | 6 | [21] |
| Zhang | 2018 | China | brain glioma | tissue | qRT-PCR | 24 | 23 | 47 | 2.46(1.13–5.35) | 0.023 | NM | NM | 40 | reported | 8 | [24] |
| Zhang | 2018 | China | gastric cancer | tissue | qRT-PCR | 41 | 35 | 76 | 1.42(0.72–2.81) | 0.310 | NM | NM | 60 | K-M | 6 | [14] |
| Zou | 2018 | China | breast cancer | tissue | qRT-PCR | 43 | 43 | 86 | NM | NM | NM | NM | NM | NM | 6 | [19] |
| Cui | 2019 | China | pancreatic cancer | tissue | qRT-PCR | 45 | 32 | 77 | 2.14(1.18–3.87) | 0.012 | NM | NM | 50 | K-M | 7 | [25] |
| Jia | 2019 | China | colorectal cancer | tissue | qRT-PCR | 30 | 31 | 61 | NM | NM | NM | NM | NM | NM | 8 | [26] |
| Liu | 2019 | China | lung adenocarcinoma | tissue | qRT-PCR | 37 | 43 | 80 | 1.60(0.45–5.71) | 0.467 | NM | NM | 60 | K-M | 6 | [23] |
| Ma | 2019 | China | hepatocellular carcinoma | tissue | qRT-PCR | 30 | 30 | 60 | NM | NM | NM | NM | NM | NM | 6 | [9] |
| Zhang | 2019 | China | cervical cancer | tissue | qRT-PCR | NM | NM | 56 | 2.37(0.75–7.50) | 0.142 | NM | NM | 60 | K-M | 7 | [20] |
| Zhu | 2019 | China | NSCLC | tissue | qRT-PCR | 33 | 39 | 72 | 0.88(0.28–2.76) | 0.825 | NM | NM | 60 | K-M | 8 | [22] |
| Li | 2019 | China | colorectal cancer | tissue | qRT-PCR | 33 | 37 | 70 | 1.31(0.59–2.92) | 0.510 | NM | NM | 60 | K-M | 7 | [32] |
| Liu | 2020 | China | gastric cancer | tissue | qRT-PCR | 34 | 34 | 68 | NM | NM | NM | NM | NM | NM | 6 | [30] |
| Liu | 2020 | China | breast cancer | tissue | qRT-PCR | 25 | 20 | 45 | NM | NM | NM | NM | NM | NM | 6 | [31] |
| Wang | 2020 | China | retinoblastoma | tissue | qRT-PCR | 37 | 33 | 70 | 2.72(0.85–8.68) | 0.090 | NM | NM | 60 | K-M | 7 | [33] |
Abbreviations: OS overall survival; DFS disease-free survival; HR hazard ratio; CI confidence interval; qRT-PCR quantitative reverse transcription polymerase chain reaction; NM not mentioned; K-M Kaplan–Meier plot; ccRCC Clear Cell Renal Cell Carcinoma; NSCLC non-small cell lung cancer; NOS Newcastle–Ottawa Scale, Ref reference
Correlation between TP73-AS1 and clinicopathologic parameters
Relationship between TP73-AS1 expression and age
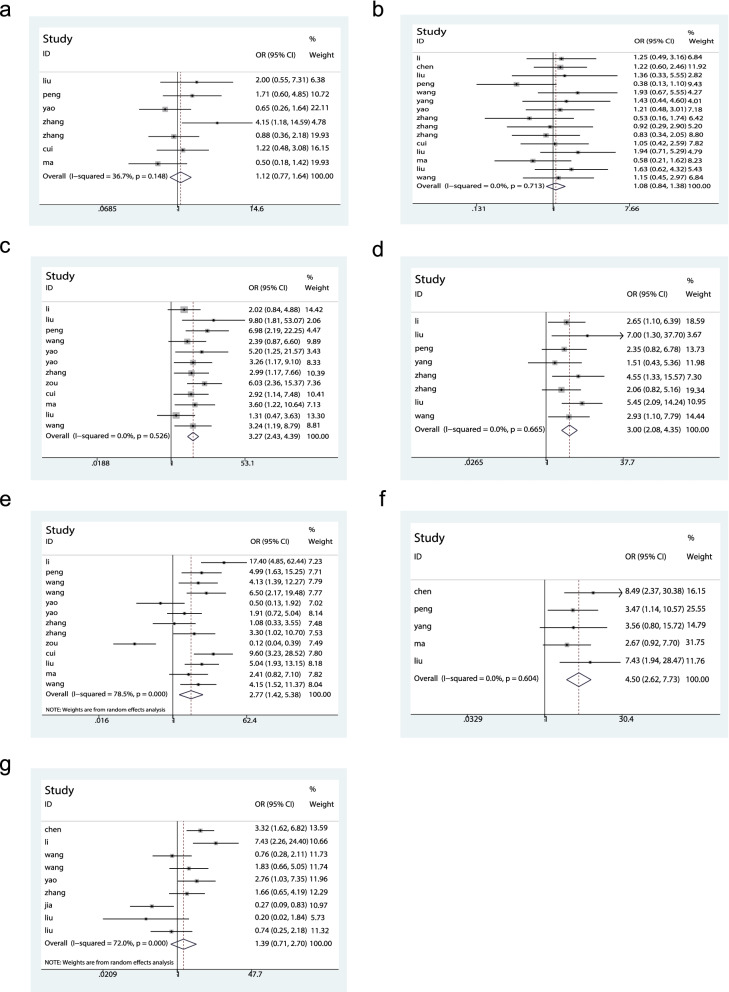
As shown in Fig. 2a, seven studies explored the correlation between TP73-AS1 expression and age. No noticeable heterogeneity was observed amongst the studies (I2 = 36.7%, P = 0.148) so a fixed model was employed for analysis. No significant correlation was identified between TP73-AS1 overexpression and patient age (OR = 1.12, 95% CI 0.77–1.64, P > 0.05).
Fig. 2.
Forest plots assessing the correlation between TP73-AS1 expression and clinicopathological parameters. a Age. b gender. c TNM stage. d tumor size. e lymph node metastasis. f distant metastasis and g differentiation
Relationship of TP73-AS1 expression with gender
Thirteen studies investigated potential associations between TP73-AS1 expression and gender. Upon assessment with the fixed model, no heterogeneity was observed amongst the studies (I2 = 0%, P = 0.713). As shown in Fig. 2b, expression of TP73-AS1 did not correlate with patient gender (OR = 1.08, 95% CI 0.84–1.38, P > 0.05).
Relationship of TP73-AS1 with TNM stage
TNM stage and TP73-AS1 expression were reported for 794 patients across twelve studies. The pooled results showed an OR = 3.27 (95% CI: 2.43–4.39, P < 0.00001) with no notable heterogeneity (I2 = 0%, P = 0.526). The data were therefore analyzed using a fixed model (Fig. 2c). The upregulation of TP73-AS1 was significantly associated with advanced TNM stage.
Relationship of TP73-AS1 expression with tumor size
The relationship between tumor size and TP73-AS1 expression was evaluated for 8 studies of 501 patients. Forest plots indicated no evident of heterogeneity amongst the studies (I2 = 0%, P = 0.665). Subsequent analysis indicated that the overexpression of TP73-AS1 correlated with a tumor size ≧ 5 cm (OR = 3.00, 95% CI: 2.08–4.35, P < 0.00001) (Fig. 2(d)).
Relationship between TP73-AS1 expression and lymph node metastasis
Data for total OR and 95% CI of LNM were collected from 13 studies. Data analysis yielded a pooled OR of 2.77 (95% CI 1.42–5.38, P < 0.00001) using a random model, owing to significant heterogeneity (I2 = 78.5%, P ≤ 0.001) (Fig. 2e). We concluded that TP73-AS1 overexpression was associated with lymph node metastasis of cancer.
Relationship between TP73-AS1 expression and distant metastasis
Five studies involving 341 patients were analyzed to evaluate the correlation between TP73-AS1 expression and distant metastasis. No significant heterogeneity was detected amongst the studies(I2 = 0%, P = 0.604, fixed-model). Model results indicated that TP73-AS1 upregulation was related to distant metastasis (OR = 4.50, 95% CI: 2.62–7.73, P < 0.00001), (Fig. 2f).
Relationship between TP73-AS1 expression and differentiation
As shown in Fig. 2g, eight studies were used to evaluate the correlation between TP73-AS1 expression and histological tumor differentiation. The random model was performed owing to heterogeneity amongst the studies (I2 = 72.0%, P ≤ 0.001) with a pooled OR = 1.39 (95% CI: 0.71–2.70, P = 0.340). No marked differences were detected in differentiation status between the two groups.
Association between TP73-AS1 and prognostic indicators
Relationship of TP73-AS1 expression with OS
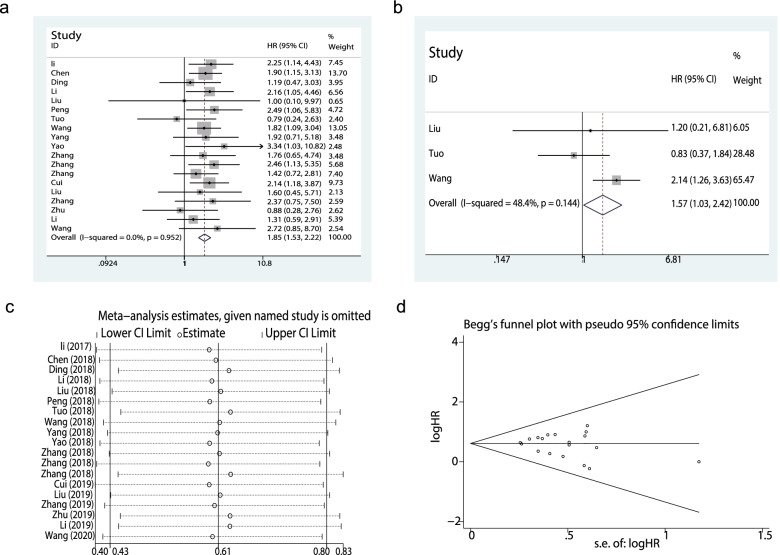
A total of 19 studies with data from 1315 patients were used to determine the utility of TP73-AS1 as a prognostic biomarker of cancer based on OS data. Pooled HR = 1.85 (95% CI: 1.53–2.22, P < 0.00001) (Fig. 3a). A fixed-effects model was used to estimate the HR of the studies, that showed no apparent heterogeneity (I2 = 0%, P = 0.952). These data indicated that the upregulation of TP73-AS1 was associated with a poorer OS amongst multiple types of malignancies.
Fig. 3.
Forest plots assessing. a the correlation between TP73-AS1 expression and overall survival (OS). b TP73-AS1 expression and disease-free survival (DFS). c sensitivity analysis for OS; and. d Begg’s assessments of OS
Relationship of TP73-AS1 expression with DFS
Only three studies reported DFS data that could be used to assess the prognostic effects of TP73-AS1. Our analysis suggested that TP73-AS1 overexpression was associated with DFS (pooled HR: 1.57, 95% CI: 1.03–2.42, P < 0.05), (Fig. 3b). No obvious heterogeneity was observed amongst the studies (I2 = 48.4%, P = 0.144).
Sensitivity analysis and publication bias
A sensitivity analysis was used to evaluate the robustness of the pooled results. The data were deemed reliable without the removal of any studies (Fig. 3c). No publication bias was observed through Begg’s tests (Fig. 3d, P = 0.368).
Subgroup analysis of OS
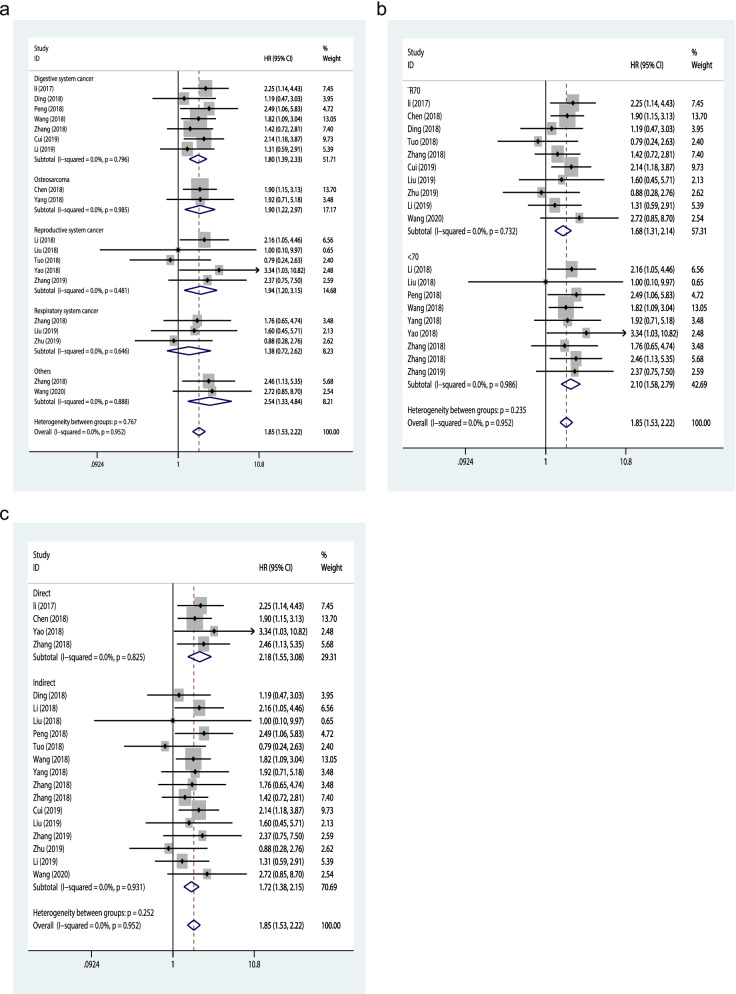
Subgroup analysis was performed in terms of cancer types, sample sizes and data extraction methods to further analyze the predictive value of TP73-AS1 (Fig. 4a–c, Table 2). As for different type of cancers, promoted TP73-AS1 expression level was closely associated with worse OS of the patients with digestive system cancers (HR = 1.80, 95%CI:1.39–2.33, P = 0.000), osteosarcomas (HR = 1.90, 95% CI:1.22–2.97, P = 0.005), respiratory system cancers (HR = 1.94, 95% CI:1.20–3.15, P = 0.007) and other cancers (HR = 2.54, 95% CI:1.33–4.84, P = 0.005), except for reproductive system cancers (HR = 1.38, 95% CI:0.72–2.62, P = 0.334). In the stratified analysis by sample sizes, we found that TP73-AS1 upregulation significantly related to unfavorable OS in the studies with sample size ≧70 (HR = 1.68, 95% CI:1.31–2.14, P = 0.000), as well as those with sample size < 70 (HR = 2.10, 95% CI:1.58–2.79, P = 0.000). When the studies were categorized according to different data extraction methods, the subgroup analysis demonstrated that the prognostic value of TP73-AS1 on the OS was not influenced by data extraction methods, that is, the HR provided in the research (HR = 2.18, 95% CI:1.55–3.08, P = 0.000) or extracted from the K-M curves (HR = 1.72, 95% CI:1.38–2.15, P = 0.000). No evident heterogeneity was found within the subgroups.
Fig. 4.
Forest plots for the subgroup analysis of OS. a cancer type; b sample size; c extracted method
Table 2.
Subgroup meta-analysis of the association between TP73-AS1 expression and OS
| Subgroup | Studies | HR | 95%CI | P value | Model | Heterogeneity | |
|---|---|---|---|---|---|---|---|
| I2 | P Value | ||||||
|
Cancer type Digestive system cancer |
7 | 1.80 | 1.39–2.33 | 0.000 | Fixed | 0% | 0.796 |
| Osteosarcoma | 2 | 1.90 | 1.22–2.97 | 0.005 | Fixed | 0% | 0.985 |
| Respiratory system cancer | 3 | 1.94 | 1.20–3.15 | 0.007 | Fixed | 0% | 0.481 |
| Reproductive system cancer | 5 | 1.38 | 0.72–2.62 | 0.334 | Fixed | 0% | 0.646 |
| Others | 2 | 2.54 | 1.33–4.84 | 0.005 | Fixed | 0% | 0.888 |
|
Sample size ≧70 |
10 | 1.68 | 1.31–2.14 | 0.000 | Fixed | 0% | 0.732 |
| < 70 | 9 | 2.10 | 1.58–2.79 | 0.000 | Fixed | 0% | 0.986 |
| Extracted method | |||||||
| Direct | 4 | 2.18 | 1.55–3.08 | 0.000 | Fixed | 0% | 0.825 |
| Indirect | 15 | 1.72 | 1.38–2.15 | 0.000 | Fixed | 0% | 0.931 |
OS overall survival; HR hazard ratio; 95% CI 95% confidence interval
Verification of TP73-AS1 expression and its prognostic value based on the TCGA
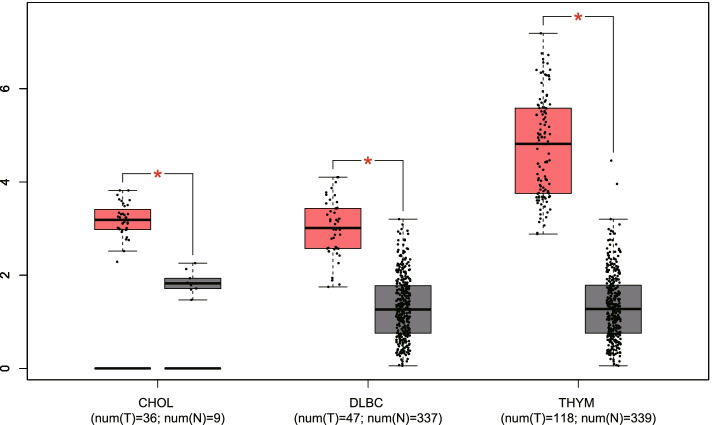
To validate the expression of TP73-AS1 in diverse cancers, we used the GEPIA online tool for gene analysis. As shown in Fig. 5, the expression of TP73-AS1 was dramatically upregulated in three malignancies including cholangiocarcinoma, lymphoid neoplasm diffuse large B-cell lymphoma and thymoma (|Log2fold change (FC)| cutoff > 1 and P < 0.01). In addition, log-rank analysis and Kaplan–Meier curves were used to verify the association between TP73-AS1 expression and the prognostic index of patients with different malignancies. Similar to our meta-analysis, the overexpression of TP73-AS1 correlated to a poorer OS in adrenocortical carcinoma (ACC) and low grade glioma (LGG) (log-rank P < 0.05) (Fig. 6a–b). Moreover, the upregulation of TP73-AS1 was associated with a poorer DFS in adrenocortical carcinoma (ACC), low grade glioma (LGG), colon adenocarcinoma (COAD), prostate adenocarcinoma (PRAD) and stomach adenocarcinoma (STAD) (log-rank P < 0.05) (Fig. 6c-g). These results indicated that TP73-AS1 serves as a novel prognostic biomarker for cancer malignancies.
Fig. 5.
TP73-AS1 expression in three types of cancer vs. normal tissue. “*”丨Log2FC丨 > 1 and P < 0.01. Abbreviations: CHOL: Cholangiocarcinoma; DLBC: Diffuse Large B-cell Lymphoma; THYM: Thymoma
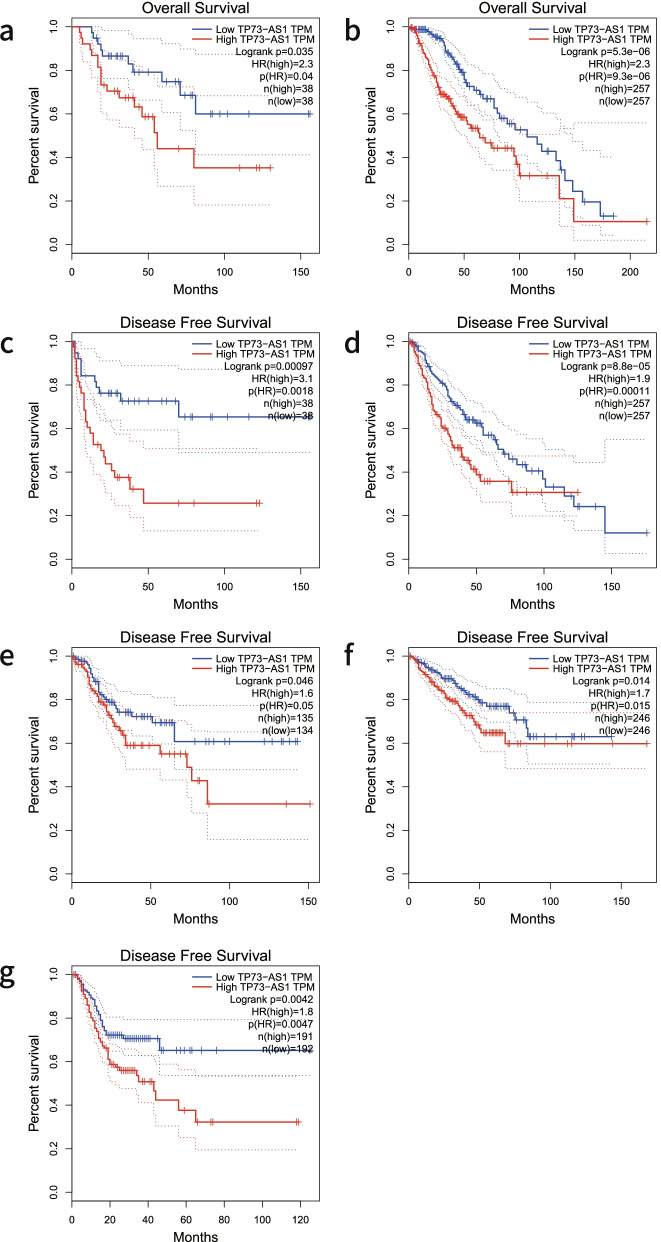
Fig. 6.
Verification of the prognostic value of TP73-AS1 in the TCGA database. a OS plots of TP73-AS1 in ACC. b OS plots of TP73-AS1 in LGG. c DFS plots of TP73-AS1 in ACC. d DFS plots of TP73-AS1 in LGG. e DFS plots of TP73-AS1 in COAD. f DFS plots of TP73-AS1 in PRAD. g DFS plots of TP73-AS1 in STAD. Abbreviations: TCGA: The Cancer Genome Atlas; ACC: Adenocarcinoma; Carcinoma; LGG: Brain Lower Grade Glioma; COAD: Colon Adenocarcinoma; PRAD: Prostate Adenocarcinoma; STAD: Stomach Adenocarcinoma
Prediction of TP73-AS1 function
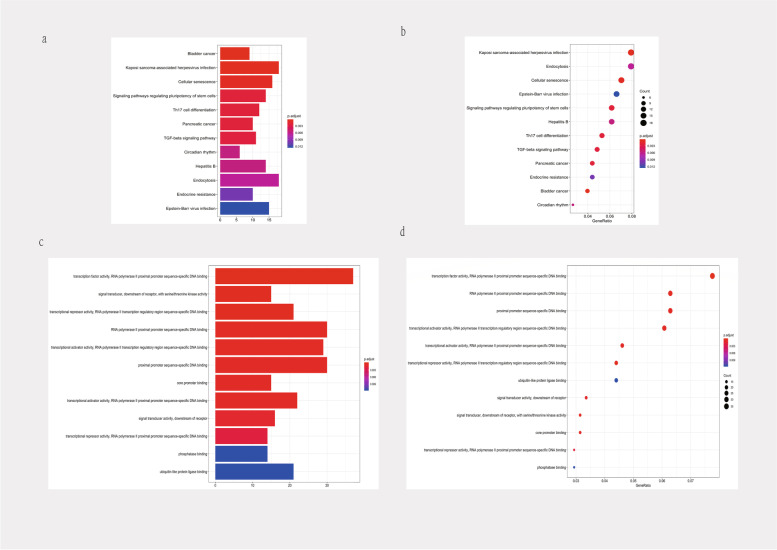
We predicted potential biological functions and the molecular mechanisms of TP73-AS1 in cells using public databases. We first explored ceRNA modulations for TP73-AS1 using targetscan, mirdb, mirtarbase, and mircode databases. TP73-AS1-miRNA-mRNA networks containing 8 miRNAs and 448 mRNA were constructed using cytoscape (Fig. 7). Figure 8 shows the top 12 KEGG and Go pathways. TP73-AS1 was predicted to participate in tumor signaling, including Kaposi sarcoma-associated herpesvirus infection, signaling pathways regulating the pluripotency of stem cells and TGF-beta signaling (Fig. 8a-b). GO functional enrichment analysis indicated that the molecular functions of TP73-AS1 included RNA polymerase II proximal promoter sequence-specific DNA binding, proximal promoter sequence-specific DNA binding, and core promoter binding (Fig. 8c-d).
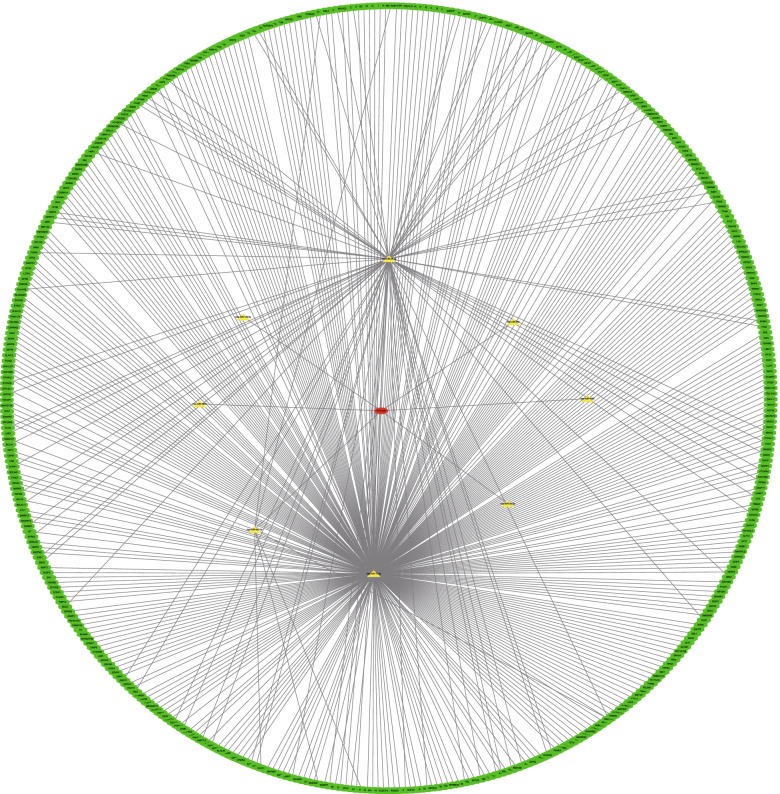
Fig.7.
Establishment of TP73-AS1-mediated ceRNA net. TP73-AS1-mediated ceRNA networks including 8 miRNAs and 448 mRNAs. Green octagons represent mRNAs; yellow triangles represent miRNAs; red ovals represent TP73-AS1
Fig. 8.
KEGG and GO term enrichment for TP73-AS1. a Barplots of KEGG molecular mechanisms. b dotplots of KEGG molecular mechanisms. c barplots of GO enrichment. d dotplots of GO enrichment
A flow diagram of the meta-analysis and bioinformatics analysis
In our paper, we first searched and selected studies met inclusion criteria and then extracted data from the papers.Thirdly, we analyzed the correlation between TP73-AS1 and clinicopathology of patients as well as prognostic indicators.Finally, we verificated TP73-AS1 expression and its prognostic value based on the TCGA and predicted its functions.The whole flow diagram was showed in Fig. 9.
Fig. 9.
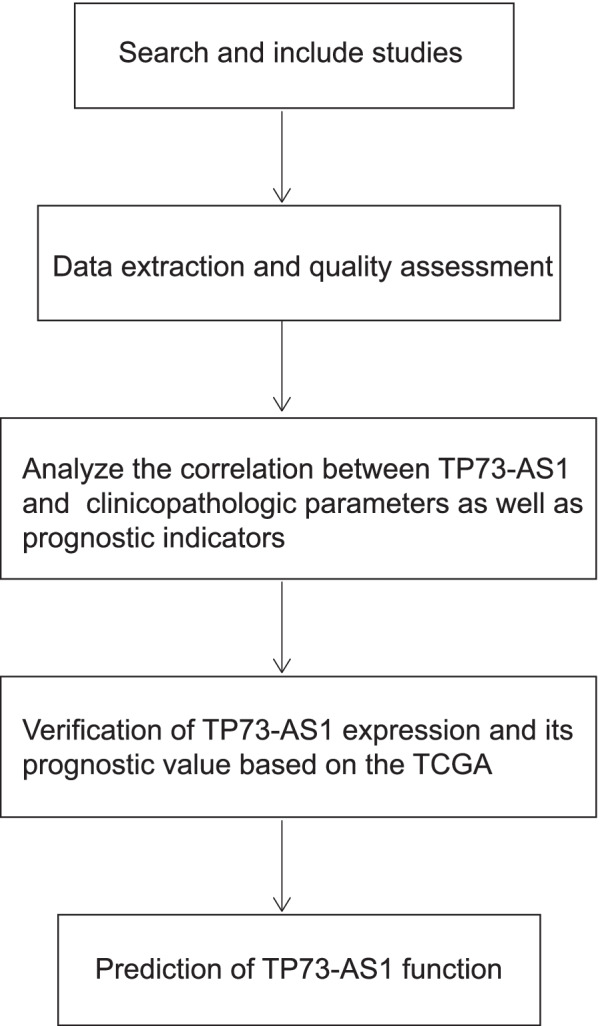
A flow diagram of the meta-analysis and bioinformatics analysis
Discussion
LncRNA TP73-AS1 is dysregulated in many tumor types. A meta-analysis was performed to examine the association between TP73-AS1 expression, clinicopathological features and prognostic values in patients. A total of twenty six studies examining 1770 patients and 14 cancers were included. The results suggested that the upregulation of TP73-AS1 significantly correlates with TNM stage, tumor size, lymph node metastasis and distant metastasis but not with the age, gender or tumor differentiation (Table 3). Furthermore, we found that TP73-AS1 overexpression is associated with poor OS and DFS, indicating that TP73-AS1 is an effective biomarker for the diagnoses and prognoses of malignancies. To further strengthen our conclusions, all finding were validated using the GEPIA database. TP73-AS1 was found to be overexpressed in CHOL, DLBC and THYM. Moreover, the OS of patients was lower in patients with high TP73-AS1 expression in ACC and LGG. Similar results were observed for the DFS of patients with ACC, LGG, COAD, PRAD and STAD.
Table 3.
Results of the association between TP73-AS1 and clinicopathological parameters
| Outcome | Studies | OR | 95%CI | P value | Model | Heterogeneity | |
|---|---|---|---|---|---|---|---|
| I2 | P Value | ||||||
| Age (< 60 vs ≥ 60) | 7 | 1.12 | 0.77–1.64 | 0.545 | Fixed | 36.7% | 0.148 |
| Gender (male vs female) | 15 | 1.08 | 0.84–1.38 | 0.560 | Fixed | 0% | 0.713 |
| TNM stage (III-IV vs I-II) | 12 | 3.27 | 2.43–4.39 | < 0.00001 | Fixed | 0% | 0.526 |
| tumor size (≥ 5 cm vs < 5 cm) | 8 | 3.00 | 2.08–4.35 | < 0.00001 | Fixed | 0% | 0.665 |
| Lymph node metastasis (positive vs negative) | 13 | 2.77 | 1.42–5.38 | 0.003 | Random | 78.5% | ≤ 0.001 |
| distant metastasis (yes vs no) | 5 | 4.50 | 2.62–7.73 | < 0.00001 | Fixed | 0% | 0.604 |
| Differentiation (poor vs well) | 9 | 1.39 | 0.71–2.70 | 0.340 | Random | 72.0% | ≤ 0.001 |
Previous studies investigated the molecular mechanisms of TP73-AS1 in diverse tumors, regarding cell proliferation, invasion, metastasis, and apoptosis. In hepatocellular carcinoma, TP73-AS1 promotes cell growth via a TP73-AS1/miR-200a/HMGB1/RAGE signaling axis and accelerates malignant progression through regulating microRNA-103 [8, 9]. In osteosarcoma, Yang et al. [10] found that TP73-AS1 silencing suppressed proliferation and invasion which when combined with miR-142 could avoid Rac1 binding. In gastric cancer, the downregulation of TP73-AS1 suppressed cell proliferation, invasion and migration via miR-194-5p/SDAD1, HMGB1/RAGE, WNT/β catenin or EMT/Bcl-2/caspase-3 signaling pathways but Liu et al. found that miR-223-5p may target TP73-AS1 to promote the invasion and migration of gastric cancer patients [11–14, 30]. In clear cell renal cell carcinoma, TP73-AS1 silencing suppressed cell proliferation and promoted apoptosis in vitro through the upregulation of KISS1 expression and the activation of PI3K/Akt/mTOR signaling [15]. In bladder cancer, Tuo et al. [16] showed that TP73-AS1 suppressed EMT. In the female reproductive system, Wang et al. [17] demonstrated that TP73‐AS1 facilitated cell proliferation and the metastasis of ovarian cancer through its regulation of MMP2 and MMP9 expression. In addition, TP73-AS1 boosted cell proliferation, invasion, and migration, and functioned as a ceRNA with miR-200a to prevent binding of TFAM, dampening the TP73-AS1/miR-200a/ZEB1 and TP73-AS1/miRNA-125a-3p/ metadherin axis in breast cancer [18, 19, 31]. Zhang et al. [20] also reported that the upregulation of TP73-AS1 promotes the tumorigenesis of cervical cancer by promoting CCND2 through the suppression of miR-607 expression. In the respiratory system, TP73-AS1 promotes tumor growth and cell cycle progression of NSCLC via a pathway involving TP73-AS1/miR-449a/EZH2 or through the regulation of miR-21 which controls the progression of lung adenocarcinoma through the PI3K/AKT axis [21–23]. We further explored the mechanisms of TP73-AS1 expression in the glioma, pancreatic cancer, colorectal cancer and retinoblastoma, and similar results were observed [24–26, 32, 33]. These studies demonstrated that TP73-AS1 can play an oncogenic role in addition to its role as a tumor suppressor, dependent on the cancer type. The molecular mechanisms governing the effects of TP73-AS1 in various malignancies are summarized in Table 4.
Table 4.
Summary of TP73-AS1 with their related signaling pathways
| Study | Cancer | Aberrant expression | Biological functions | Related signaling pathways |
|---|---|---|---|---|
| Li 2017 [8] | hepatocellular carcinoma | upregulation | promote cell proliferation | miR-200a/HMGB1/RAGE |
| Chen 2018 [27] | osteosarcoma | upregulation | promote cell proliferation, migration and invasion | |
| Ding 2018 [11] | gastric cancer | upregulation | promote cell growth and metastasis | miR-194-5p/SDAD1 |
| Li 2018 [28] | ovarian cancer | upregulation | promote cell proliferation | |
| Liu 2018 [15] | clear cell renal cell carcinoma | upregulation | promote cell proliferation, inhibit cell apoptosis | KISS/EZH2, PI3K/Akt/mTOR |
| Peng 2018 [12] | gastric cancer | upregulation | promote cell proliferation | HMGB1/RAGE |
| Tuo 2018 [16] | bladder cancer | downregulation | inhibit cell growth, cell cycle, migration and invasion, induce cell apoptosis | EMT |
| Wang 2018 [17] | ovarian cancer | upregulation | promoted cell proliferation, invasion, and migration | MMP2, MMP9 |
| Wang 2018 | gastric cancer | upregulation | promote cell proliferation, invasion | WNT/β-catenin |
| Yang 2018 [10] | osteosarcoma | upregulation | promote cell proliferation, invasion | miR-142/Rac1 |
| Yao 2018 [18] | breast cancer | upregulation | promote cell proliferation | miR-200a/TFAM |
| Yao 2018 [6] | cholangiocarcinoma | upregulation | promote cell proliferation, migration, invasion, inhibit cell apoptosis | |
| Zhang 2018 [21] | non-small cell lung cancer | upregulation | promote cell proliferation, tumor growth and cycle progression | miR-449a/EZH2 |
| Zhang 2018 [24] | brain glioma | upregulation | promote cell proliferation and invasion | miR-142/HMGB1/RAGE |
| Zhang 2018 [14] | gastric cancer | upregulation | promote cell migration and invasion | EMT/Bcl-2/caspase-3 |
| Zou 2018 [19] | breast cancer | upregulation | promote cell invasion and migration | miR-200a/ZEB1 |
| Cui 2019 [25] | pancreatic cancer | upregulation | promote migration and invasion | miR-141-3p/BDH2 |
| Jia 2019 [26] | colorectal cancer | downregulation | inhibite cell growth, promote apoptosis | miR-103/ PTEN |
| Liu 2019 [23] | Lung adenocarcinoma | upregulation | promote cell proliferation, migration, invasion, inhibit apoptosis | PI3K/AKT |
| Ma 2019 [9] | hepatocellular carcinoma | upregulation | promote cell proliferation, inhibit apoptosis | miR-103 |
| Zhang 2019 [20] | cervical cancer | upregulation | promote cell proliferation, migration and invasion | miR-607/cyclin D2 |
| Zhu 2019 [22] | non-small cell lung cancer | upregulation | promote cell proliferation. migration and invasion | miR-21 |
| Li 2019 [32] | colorectal cancer | upregulation | promote cell migration and invasion | TGF-β1 |
| Liu 2020 [30] | gastric cancer | downregulation | inhibit cell invasion and migration | miR-223-5p |
| Liu 2020 [31] | breast cancer | upregulation | promote cell proliferation, migration and invasion, inhibite apoptosis | miRNA-125a-3p/metadherin |
| Wang 2020 [33] | retinoblastoma | upregulation | promote cell proliferation, metastasis and invasion | miRNA-874-3p / TFAP2B |
LncRNAs indirectly modulate the function and expression of downstream genes through ceRNAs. We therefore constructed a TP73-AS1-mediated competing endogenous RNA network to evaluate possible functional and signaling pathways of TP73-AS1 in tumors. As shown in the network, TP73-AS1 could bind to eight miRNAs targeting more than 400 mRNAs. Furthermore, the predicted KEGG pathways indicated that TP73-AS1 regulates the cell proliferation, migration, invasion and apoptosis of malignancies via Kaposi sarcoma-associated herpesvirus infection, signaling pathways regulating the pluripotency of stem cells and TGF-beta signaling.
Some limitations of the study should be noted. First, all included studies were from China, so the results may only be applicable to Chinese or Asian populations. Supplementary analysis using the GEPIA database was used to compensate for this racial limitation. Secondly, a larger sample size and different tumor types are required to confirm our overall conclusions due to the limited sample size of limited number of carcinomas assessed. Thirdly, HR and 95% CI values of 15 studies were calculated from Kaplan–Meier curves. These values were likely to be less accurate than those obtained from direct measurements. Finally, we only predicted the biological functions and molecular mechanisms of TP73-AS1 ceRNA networks and utilized KEGG and GO enrichment analysis. Other broader regulatory mechanisms should now be investigated. Taken together, further studies should focus on other types of cancer, including hematological tumors with larger sample sizes to explore other potential functions of TP73-AS1 and its role during prooncogenic signaling. This information may provide novel therapeutic strategies for cancer patients.
Despite these limitations, we demonstrate that TP73-AS1 overexpression correlates with advanced TNM stage, larger tumor sizes, lymph node metastasis and distant metastasis. In addition, TP73-AS1 overexpression is related to a poor prognosis, indicating its utility as a diagnostic and prognostic marker of diverse malignancies. Further studies in larger patient cohorts and an array of cancer types are now required to validate these findings.
Conclusions
The upregulation of LncRNA TP73-AS1 correlates with detrimental clinicopathological features and can be considered an indicator of poor prognosis for malignancies in a clinical setting.
Acknowledgements
Not applicable.
Authors’ contributions
WZ designed the concept of study. CZC and YL collected and analyzed the data, wrote the manuscript; YH arranged the tables and figures; JJW and YQF revised the paper. All authors reviewed and approved the final manuscript.
Funding
This work was supported by the Development and Reform Commission of Hunan Province (NO. (2019, 412) and WU JIEPING MEDICAL FOUNDATION NO.320.6750.18540.
Availability of data and materials
All data are included in our paper.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69(1):7–34. doi: 10.3322/caac.21551. [DOI] [PubMed] [Google Scholar]
- 2.Ulitsky I. Evolution to the rescue: using comparative genomics to understand long non-coding RNAs. Nat Rev Genet. 2016;17(10):601–614. doi: 10.1038/nrg.2016.85. [DOI] [PubMed] [Google Scholar]
- 3.Zhao D, Wang S, Chu X, Han D. LncRNA HIF2PUT inhibited osteosarcoma stem cells proliferation, migration and invasion by regulating HIF2 expression. Artificial cells, nanomedicine, and biotechnology. 2019;47(1):1342–1348. doi: 10.1080/21691401.2019.1596934. [DOI] [PubMed] [Google Scholar]
- 4.Sha QK, Chen L, Xi JZ, Song H. Long non-coding RNA LINC00858 promotes cells proliferation, migration and invasion by acting as a ceRNA of miR-22-3p in colorectal cancer. Artificial cells, nanomedicine, and biotechnology. 2019;47(1):1057–1066. doi: 10.1080/21691401.2018.1544143. [DOI] [PubMed] [Google Scholar]
- 5.Li G, Zhang Y, Mao J, Hu P, Chen Q, Ding W, Pu R. LncRNA TUC338 is overexpressed in prostate carcinoma and downregulates miR-466. Gene. 2019;707:224–230. doi: 10.1016/j.gene.2019.05.026. [DOI] [PubMed] [Google Scholar]
- 6.Xie T, Pi G, Yang B, Ren H, Yu J, Ren Q, Zhou X, Hu D, Zhang H, Zhang H, et al. Long non-coding RNA 520 is a negative prognostic biomarker and exhibits pro-oncogenic function in nasopharyngeal carcinoma carcinogenesis through regulation of miR-26b-3p/USP39 axis. Gene. 2019;707:44–52. doi: 10.1016/j.gene.2019.02.093. [DOI] [PubMed] [Google Scholar]
- 7.Wong KY, Li Z, Zhang X, Leung GK, Chan GC, Chim CS. Epigenetic silencing of a long non-coding RNA KIAA0495 in multiple myeloma. Mol Cancer. 2015;14:175. doi: 10.1186/s12943-015-0444-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Li S, Huang Y, Huang Y, Fu Y, Tang D, Kang R, Zhou R, Fan XG. The long non-coding RNA TP73-AS1 modulates HCC cell proliferation through miR-200a-dependent HMGB1/RAGE regulation. Journal of experimental & clinical cancer research : CR. 2017;36(1):51. doi: 10.1186/s13046-017-0519-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.C-X Ma W-CG L Tian: LncRNA TP73-AS1 Promotes Malignant Progression of Hepatoma by Regulating microRNA-103. Eur Rev Med Pharmacol Sci. 2019;23(11):4713–4722. doi: 10.26355/eurrev_201906_18052. [DOI] [PubMed] [Google Scholar]
- 10.Yang G, Song R, Wang L, Wu X: Knockdown of long non-coding RNA TP73-AS1 inhibits osteosarcoma cell proliferation and invasion through sponging miR-142. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie 2018, 103:1238–1245. [DOI] [PubMed]
- 11.Ding Z, Lan H, Xu R, Zhou X, Pan Y. LncRNA TP73-AS1 accelerates tumor progression in gastric cancer through regulating miR-194-5p/SDAD1 axis. Pathol Res Pract. 2018;214(12):1993–1999. doi: 10.1016/j.prp.2018.09.006. [DOI] [PubMed] [Google Scholar]
- 12.Peng J. si-TP73-AS1 suppressed proliferation and increased the chemotherapeutic response of GC cells to cisplatin. Oncol Lett. 2018;16(3):3706–3714. doi: 10.3892/ol.2018.9107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang Y, Xiao S, Wang B, Li Y, Chen Q. Knockdown of lncRNA TP73-AS1 inhibits gastric cancer cell proliferation and invasion via the WNT/beta-catenin signaling pathway. Oncol Lett. 2018;16(3):3248–3254. doi: 10.3892/ol.2018.9040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zhang W, Zhai Y, Wang W, Cao M, Ma C. Enhanced expression of lncRNA TP73-AS1 predicts unfavorable prognosis for gastric cancer and promotes cell migration and invasion by induction of EMT. Gene. 2018;678:377–383. doi: 10.1016/j.gene.2018.08.055. [DOI] [PubMed] [Google Scholar]
- 15.Liu G, Zhao X, Zhou J, Cheng X, Ye Z, Ji Z. LncRNA TP73-AS1 Promotes Cell Proliferation and Inhibits Cell Apoptosis in Clear Cell Renal Cell Carcinoma Through Repressing KISS1 Expression and Inactivation of PI3K/Akt/mTOR Signaling Pathway. Cellular physiology and biochemistry : international journal of experimental cellular physiology, biochemistry, and pharmacology. 2018;48(1):371–384. doi: 10.1159/000491767. [DOI] [PubMed] [Google Scholar]
- 16.Tuo Z, Zhang J, Xue W. LncRNA TP73-AS1 predicts the prognosis of bladder cancer patients and functions as a suppressor for bladder cancer by EMT pathway. Biochem Biophys Res Commun. 2018;499(4):875–881. doi: 10.1016/j.bbrc.2018.04.010. [DOI] [PubMed] [Google Scholar]
- 17.Wang X, Yang B, She Y, Ye Y. The lncRNA TP73-AS1 promotes ovarian cancer cell proliferation and metastasis via modulation of MMP2 and MMP9. J Cell Biochem. 2018;119(9):7790–7799. doi: 10.1002/jcb.27158. [DOI] [PubMed] [Google Scholar]
- 18.Yao J, Xu F, Zhang D, Yi W, Chen X, Chen G, Zhou E. TP73-AS1 promotes breast cancer cell proliferation through miR-200a-mediated TFAM inhibition. J Cell Biochem. 2018;119(1):680–690. doi: 10.1002/jcb.26231. [DOI] [PubMed] [Google Scholar]
- 19.Zou Q, Zhou E, Xu F, Zhang D, Yi W, Yao J. A TP73-AS1/miR-200a/ZEB1 regulating loop promotes breast cancer cell invasion and migration. J Cell Biochem. 2018;119(2):2189–2199. doi: 10.1002/jcb.26380. [DOI] [PubMed] [Google Scholar]
- 20.Zhang H, Xue B, Wang S, Li X, Fan T. Long non‑coding RNA TP73 antisense RNA 1 facilitates the proliferation and migration of cervical cancer cells via regulating microRNA‑607/cyclin D2. Mol Med Rep. 2019;20(4):3371–8. [DOI] [PubMed]
- 21.Zhang L, Fang F, He X. Long noncoding RNA TP73-AS1 promotes non-small cell lung cancer progression by competitively sponging miR-449a/EZH2. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie. 2018;104:705–11. [DOI] [PubMed]
- 22.Zhu D, Zhou J, Liu Y, Du L, Zheng Z, Qian X. LncRNA TP73-AS1 is upregulated in non-small cell lung cancer and predicts poor survival. Gene. 2019;710:98–102. doi: 10.1016/j.gene.2019.05.044. [DOI] [PubMed] [Google Scholar]
- 23.Liu C, Ren L, Deng J, Wang S. LncRNA TP73-AS1 promoted the progression of lung adenocarcinoma via PI3K/AKT pathway. Biosci Rep. 2019;39(1):BSR20180999. [DOI] [PMC free article] [PubMed]
- 24.Zhang R, Jin H, Lou F. The Long Non-Coding RNA TP73-AS1 Interacted With miR-142 to Modulate Brain Glioma Growth Through HMGB1/RAGE Pathway. J Cell Biochem. 2018;119(4):3007–3016. doi: 10.1002/jcb.26021. [DOI] [PubMed] [Google Scholar]
- 25.Cui XP, Wang CX, Wang ZY, Li J, Tan YW, Gu ST, Qin CK. LncRNA TP73-AS1 sponges miR-141–3p to promote the migration and invasion of pancreatic cancer cells through the up-regulation of BDH2. Biosci Rep. 2019;39(3):BSR20181937. [DOI] [PMC free article] [PubMed]
- 26.Jia Z, Peng J, Yang Z, Chen J, Liu L, Luo D, He P. Long non-coding RNA TP73AS1 promotes colorectal cancer proliferation by acting as a ceRNA for miR103 to regulate PTEN expression. Gene. 2019;685:222–229. doi: 10.1016/j.gene.2018.11.072. [DOI] [PubMed] [Google Scholar]
- 27.Chen X, Zhou Y, Liu S, Zhang D, Yang X, Zhou Q, Song Y, Liu Y. LncRNA TP73-AS1 predicts poor prognosis and functions as oncogenic lncRNA in osteosarcoma. J Cell Biochem. 2018;120(2):2569–75. [DOI] [PubMed]
- 28.Li X, Wang X, Mao L, Zhao S, Wei H. LncRNA TP73AS1 predicts poor prognosis and promotes cell proliferation in ovarian cancer via cell cycle and apoptosis regulation. Mol Med Rep. 2018;18(1):516–522. doi: 10.3892/mmr.2018.8951. [DOI] [PubMed] [Google Scholar]
- 29.Yao Y, Sun Y, Jiang Y, Qu L, Xu Y: Enhanced expression of lncRNA TP73-AS1 predicts adverse phenotypes for cholangiocarcinoma and exerts oncogenic properties in vitro and in vivo. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie 2018, 106:260–266. [DOI] [PubMed]
- 30.Liu J, Wang H, Liao X. MicroRNA-223-5p targets long non-coding RNA TP73 antisense RNA1 to promote the invasion of gastric cancer. Hum Cell. 2020;33(3):676–682. doi: 10.1007/s13577-020-00349-3. [DOI] [PubMed] [Google Scholar]
- 31.Liu Y, Wei G, Ma Q, Han Y. Knockdown of long noncoding RNA TP73-AS1 suppresses the malignant progression of breast cancer cells in vitro through targeting miRNA-125a-3p/metadherin axis. Thorac Cancer. 2020;11(2):394–407. doi: 10.1111/1759-7714.13283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Li M, Jin Y, Li Y. LncRNA TP73-AS1 Activates TGF-beta1 to Promote the Migration and Invasion of Colorectal Cancer Cell. Cancer Manag Res. 2019;11:10523–10529. doi: 10.2147/CMAR.S228490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang L, Wang C, Wu T, Sun F. Long non-coding RNA TP73-AS1 promotes TFAP2B-mediated proliferation, metastasis and invasion in retinoblastoma via decoying of miRNA-874-3p. J Cell Commun Signal. 2020;14(2):193–205. doi: 10.1007/s12079-020-00550-x. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data are included in our paper.