Abstract
Background
Coxiella burnetii (Cb) is the causative agent of the zoonotic disease Q fever which is distributed worldwide. Molecular typing of Cb strains is essential to find out the infectious source and prevent Q fever outbreaks, but there has been a lack of typing data for Cb strains in China. The aim of this study was to investigate the genotypes of Cb strains in wild rats in Yunnan Province, China.
Results
Eighty-six wild rats (Rattus flavipectus) were collected in Yunnan Province and 8 of the 86 liver samples from the wild rats were positive in Cb-specific quantitative PCR (qPCR). The Cb strains from the 8 rats were then typed into 3 genotypes using 10-spacer multispacer sequence typing (MST), and 2 of the 3 genotypes were recognized as novel ones. Moreover, the Cb strains in the wild rats were all identified as genotype 1 using 6-loci multilocus variable number of tandem repeat analysis (MLVA).
Conclusions
This is the first report of genotypic diversity of Cb strains from wild rats in China. Further studies are needed to explore the presence of more genotypes and to associate the genotypes circulating in the wildlife-livestock interaction with those causing human disease to further expand on the epidemiological aspects of the pathogen.
Keywords: Rattus flavipectus, Coxiella burnetii, Genotype, MST, MLVA, China
Background
Coxiella burnetii (Cb), an intracellular Gram-negative bacterium, is the causative agent of Q fever, a disease distributed worldwide except in New Zealand [1]. Cb is mainly found in livestock, such as goats, sheep and cattle, where chronic infection is associated with late abortion, stillbirth, and weak offspring [2]. Other reservoirs of Cb include birds, wild rodents and arthropods [3, 4]. Although ticks can transmit Cb in experimental systems, direct transmission of Cb to humans through ticks has never been properly documented [5, 6].
C. burnetii infection exhibits various acute and chronic clinical manifestations in humans. Acute Q fever typically presents as a flu-like illness with a high fever, headache, myalgia and malaise, and can develop into chronic Q fever which presents as endocarditis, hepatitis, and/or osteomyelitis [7, 8]. Human acute Cb infections often occur after inhalation of Cb-contaminated aerosols from infected animals, thus occupations that require close contact with livestock have a higher risk of acquiring Q fever [7, 9, 10].
The importance of Q fever has been increasing since the outbreak in the Netherlands from 2007 to 2010, where more than 4000 people became ill [11]. To prevent infection in humans, it is critical to determine the source of infection for each case including patients with Q fever exposed to wildlife. For this purpose, several genotyping methods have been described including sequencing analysis of individual genes like 16S, 23S [12], plasmid-based typing [13, 14], restriction fragment length polymorphism and pulsed-field gel electrophoresis (RFLP-PFGE) analysis [15], single nucleotide polymorphism (SNP) [16], multispacer sequence typing (MST) [17, 18], and multilocus variable number tandem repeats analysis (MLVA) [19]. Of the mentioned methods, MST and MLVA can be applied directly to DNA samples without previous cultivation of the strain, and have proven to be reliable and reproducible.
In China, Cb has been detected in wildlife, but few MST or MLVA genotypes of Cb strains have been reported [20, 21]. This study aimed to genotype Cb strains detected in wild rats in Yunnan Province. This study is the first report of MLVA and MST genotypes of Cb strains in wild rats in China, which provides further insight into the epidemiology and evolution of Q fever.
Results
Rodent species identification
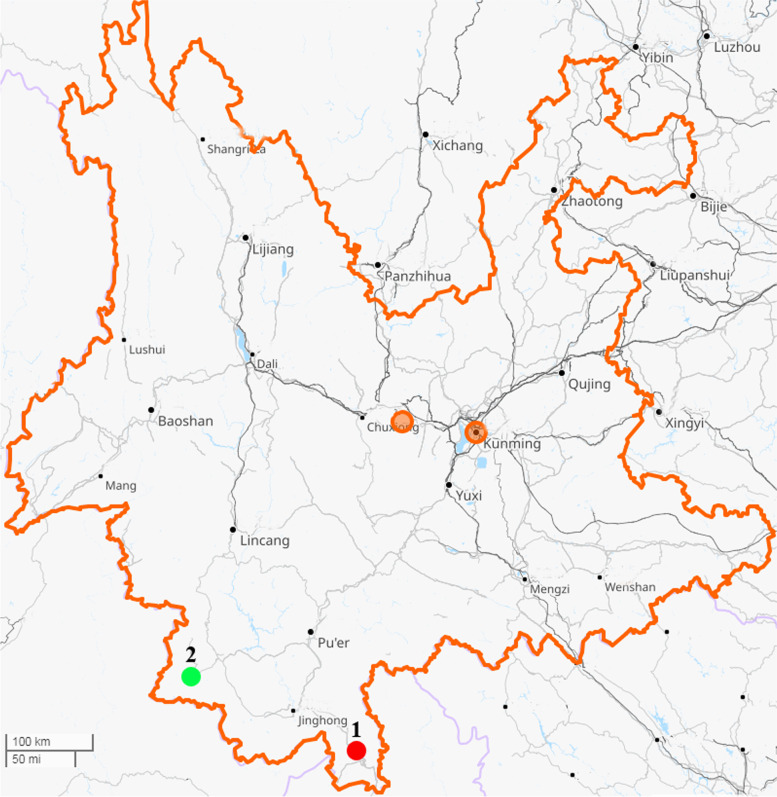
A total of 86 wild rats were collected from wild fields in Mengla County (n = 33) and Menglian County (n = 53) in Yunnan Province in August 2020 (Fig. 1). All rats were identified as Rattus flavipectus based on morphological identifications and confirmed by species-specific PCR and sequencing assays.
Fig. 1.
Map of the sampling sites in Yunnan Province, China. Rats were collected in Mengla County (sampling site 1) and Menglian County (sampling site 2) in Yunnan Province
C. burnetii detection
The presence of C. burnetii in the wild rats was first tested on their liver samples by nested PCR. Eight of 53 liver samples (15.09%, 8/53) collected in Menglian County were positive in PCR targeting Cb 16S rRNA, and the sequences obtained showed 99.17–99.44% nucleotide sequence identity to those of the known Cb strains in 16S rRNA comparison. No rats collected in Mengla County were positively detected in PCR.
The liver samples positively detected by Cb-specific PCR were further analyzed by qPCR to assess the Cb loads in the wild rats. As a result, all 8 liver samples showed a Cb concentration ranging from 4.40 × 104 to 2.53 × 106 copies/gram (Table 1).
Table 1.
Summary of qPCR of the rat samples used in MST and MLVA genotyping
| Sample ID | Host | Sample source | Ct in qPCR | Quantification in qPCR (Copies/gram) |
|---|---|---|---|---|
| 4 | Rattus flavipectus | liver | 34.15 | 1.45 × 105 |
| 5 | Rattus flavipectus | liver | 34.52 | 1.18 × 105 |
| 12 | Rattus flavipectus | liver | 34.35 | 1.34 × 105 |
| 16 | Rattus flavipectus | liver | 33.73 | 1.85 × 105 |
| 20 | Rattus flavipectus | liver | 34.28 | 1.35 × 105 |
| 21 | Rattus flavipectus | liver | 36.21 | 4.40 × 104 |
| 22 | Rattus flavipectus | liver | 35.72 | 5.85 × 104 |
| 44 | Rattus flavipectus | liver | 29.07 | 2.53 × 106 |
MST genotyping
Seven of the 8 liver samples positive for Cb were successfully amplified for all 10 spacers in MST genotyping. For samples 4 and 44, the allele codes found in the present study for loci Cox2-Cox5-Cox18-Cox20-Cox22-Cox37-Cox51-Cox56-Cox57-Cox61 were 3–8-5–3-4–1-6–7-6–5 and belonged to MST genotype 16. For sample 12, Cox61 was unable to be sequenced with good resolution, possibly due to poor quantity of DNA.
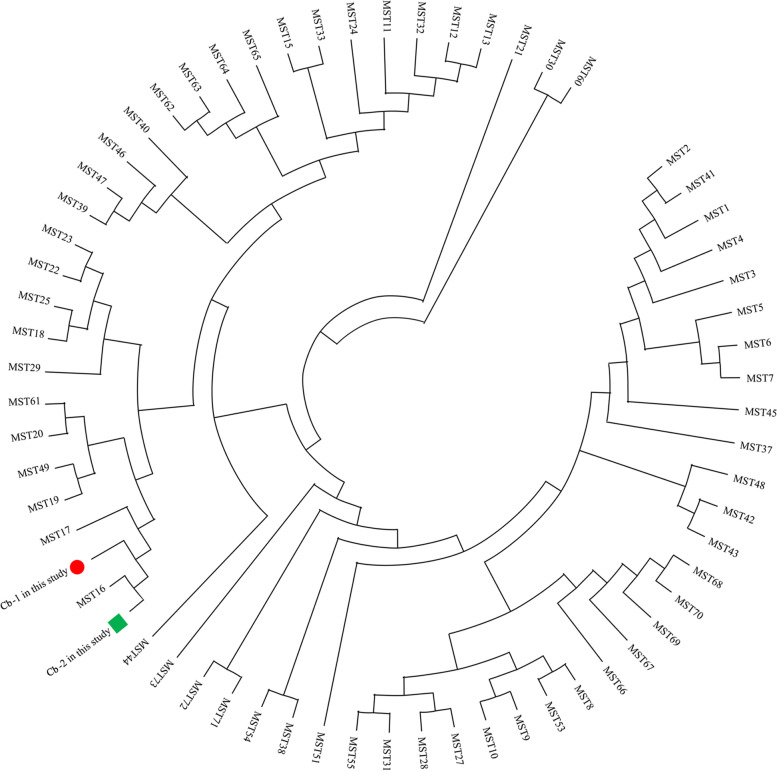
For samples 5, 16, 20, 21 and 22, spacer sequences of Cox 20 and/or Cox37 were diverse from those described in the MST database (Table 2). For Cox 20, mutations T/A at position 504 and G/A at position 513 of allele 3 were present. For Cox 37, a deletion of A at position 42 and a mutation T/G at position 33 of allele 1 were present. These data suggested the presence of new alleles, implying the existence of novel MST genotypes. A phylogenetic tree placed the MST genotypes of these strains (samples 5, 16, 20, 21 and 22) close to MST genotype 16 (Fig. 2).
Table 2.
MST genotypes of C. burnetii detected in rats in the present study
| MST Loci | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample ID | Cox 2 | Cox 5 | Cox 18 | Cox 20 | Cox 22 | Cox 37 | Cox 51 | Cox 56 | Cox 57 | Cox 61 | MST type |
| 4 | 3 | 8 | 5 | 3 | 4 | 1 | 6 | 7 | 6 | 5 | 16 |
| 5 | 3 | 8 | 5 | 3 | 4 | Novel | 6 | 7 | 6 | 5 | Novel |
| 12 | 3 | 8 | 5 | 3 | 4 | 1 | 6 | 7 | 6 | N/A | - |
| 16 | 3 | 8 | 5 | 3 | 4 | Novel | 6 | 7 | 6 | 5 | Novel |
| 20 | 3 | 8 | 5 | 3 | 4 | Novel | 6 | 7 | 6 | 5 | Novel |
| 21 | 3 | 8 | 5 | 3 | 4 | Novel | 6 | 7 | 6 | 5 | Novel |
| 22 | 3 | 8 | 5 | Novel | 4 | Novel | 6 | 7 | 6 | 5 | Novel |
| 44 | 3 | 8 | 5 | 3 | 4 | 1 | 6 | 7 | 6 | 5 | 16 |
N/A Negative assembly
Fig. 2.
Phylogenetic tree of Coxiella burnetii MST genotypes from this study with known genotypes. The obtained MST genotypes identified in this study and data of known MST genotypes based on the MST database (https://ifr48.timone.univ-mrs.fr/mst/coxiella_burnetii/) were used. Sequences were aligned using the MEGA X (Version 10.2.5) software package. Phylogenetic analysis was performed using the unweighted pair group method with arithmetic mean (UPGMA) method. Cb-1: genotype of samples 5, 16, 20, 21; Cb-2: genotype of sample 22
MLVA genotyping
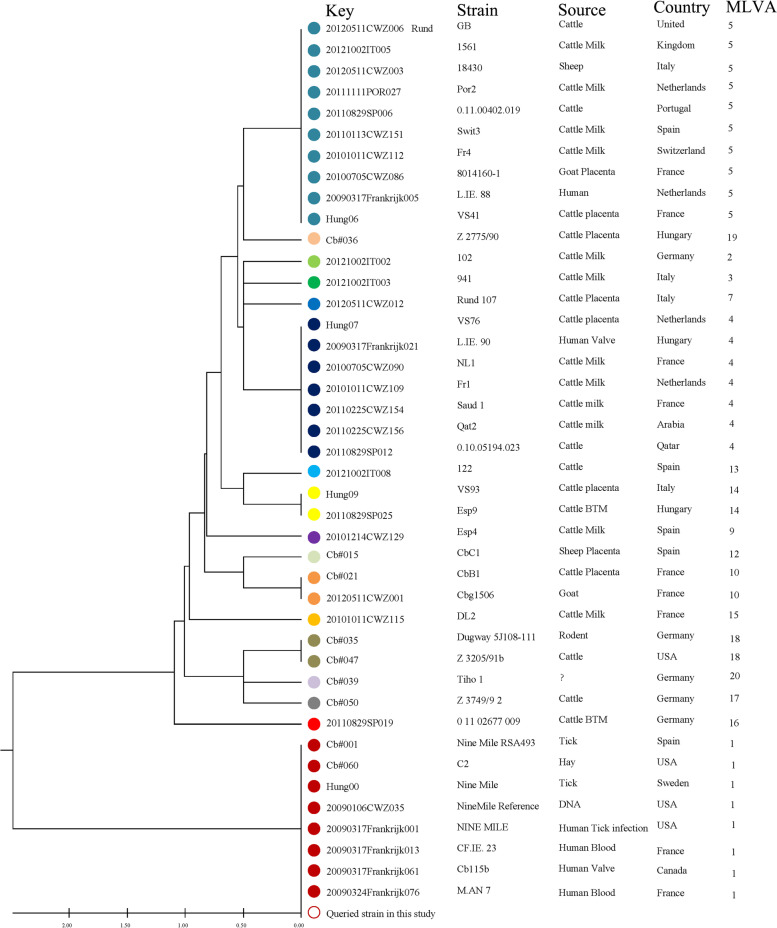
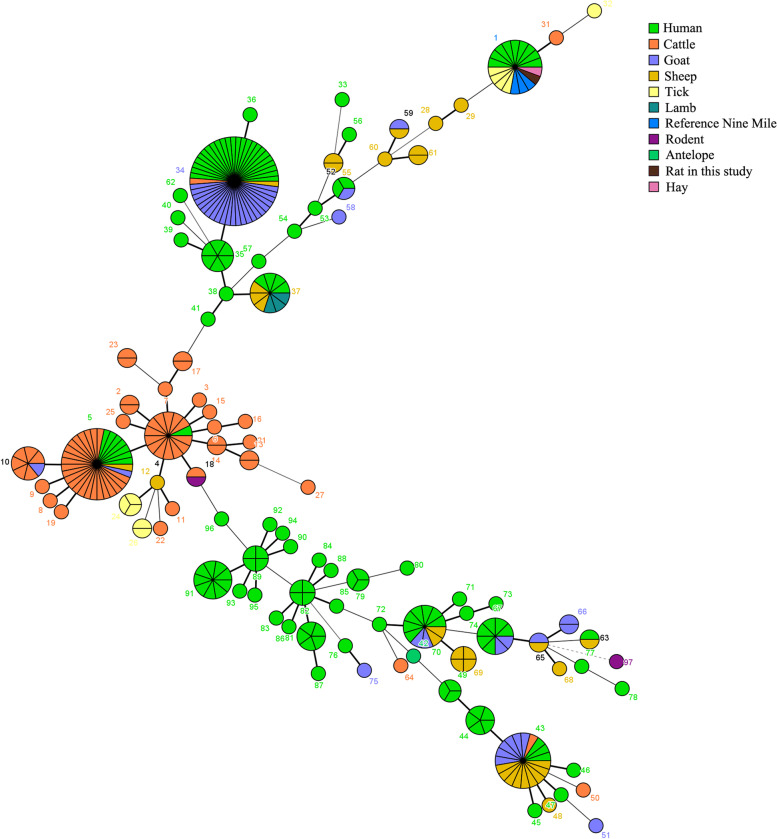
The strains in the 8 liver samples positive for Cb were characterized by the MLVA analysis. The allele codes found in the present study were 9–27-4–6-9–5 for loci ms23-ms24-ms27-ms28-ms33-ms34, and belonged to MLVA genotype 1, suggesting that all Cb strains identified in this study belong to the same genotype. MLVA genotype 1 has already been detected in ticks in the USA, and identified in Q fever patients in the USA, Canada, and France (Fig. 3). A minimum spanning tree based on host origin of the MLVA analysis showed that the Cb strains detected in the present study were clustered with the previously described genotypes found primarily in ticks or Q–fever patients in different regions of the globe (Fig. 4).
Fig. 3.
UPGMA cluster analysis of Coxiella burnetii MLVA-6 genotypes. All data of selected samples are available in the MLVA-6 database (http://mlva.i2bc.paris-saclay.fr/mlvav4/genotyping/). Strain, source, geographical origin, and MLVA-6 type are indicated. The same genotype was coded with the same color, and the hollow dot indicates the genotype obtained in the present study
Fig. 4.
Minimum spanning tree of Coxiella burnetii strains based on the results of MLVA-6 analysis. All data of selected samples are available in the MLVA-6 database (http://mlva.i2bc.paris-saclay.fr/mlvav4/genotyping/). The minimum spanning tree provides information on the proportion of hosts of each identified genotype (see color index). Each circle represents a unique genotype, and the size of the pie charts represents the number of isolates of the corresponding genotype
Discussion
In China, Q fever was initially reported in 1950 and the Cb strain was first isolated from a chronic Q fever patient in Chongqing city in 1962 [22]. There have been dozens of studies focused on Q fever and the Cb strains detected and/or isolated from patients [22, 23], mammals [24], wild animals [20] and ticks [25]. However, very little information is available regarding the diversity of Cb strains in China. This study describes the genetic diversity of Cb strains detected in wild rats in Yunnan Province of China using MST and MLVA genotyping, which may be great value in tracking the disease [18, 19] and deciphering its potential zoonotic role in Yunnan.
To date, 74 MST genotypes of Cb have been recorded worldwide in the Coxiella MST database (https://ifr48.timone.univ-mrs.fr/mst/coxiella_burnetii/). In the present study, 2 samples (4 and 44) were recognized as MST genotype 16. Interestingly, the allele codes of Cox20 and/or Cox 37 from samples 5, 16, 20, 21 and 22 were designated as novel ones since no correspondence was found among the loci repeats profiles detected herein and those currently recorded in the Coxiella MST database, suggesting novel Cb genotypes circulating in China. Up to now the information regarding Cb diversity in China has been found only for strains from hedgehogs samples [20], describing two novel MST genotypes that are different from those typed in the present study.
Using phylogenetic analysis, the two novel MST genotypes in the present study were placed in a clade with MST genotypes 16 and 17, closer to genotype 16. MST genotype 16 is most often associated with Q fever patients and is occasionally recorded in cows or ticks, and it has a widespread distribution in the USA, Japan, Europe (Romania, France, Italy, Germany, Slovakia, Poland), and Central Africa [26]. MST genotype 17 has only been detected in Q fever patients in France according to the MST database. The presence of novel MST genotypes in the present study reinforces this cartographic feature of the genetic diversity based on MST typing of Cb strains.
Different genotyping methods of Cb strains seem to agree with each other [27, 28]. In addition to MST genotyping, MLVA genotyping was performed in the present study. The strains from the rats in Yunnan were recognized as genotype 1 using 6 MLVA loci (ms21, ms22, ms23, ms28, ms33, and ms34) (Fig. 3), and this genotype was mainly found in strains both from Q fever patients and ticks according to the database of MLVABank (http://mlva.u-psud.fr/mlvav4/genotyping/).
A minimum spanning tree allowing the observation of possible host-adapted lineages was drawn (Fig. 4). The obtained strains recognized as MLVA genotype 1 in the present study were clustered mainly with the strains from Q fever patients in France, Canada and the USA, suggesting that these strains from wild rats in Yunnan Province were linked in proximity to the Cb strains from ticks and patients in different regions of the world. This result seems to agree with the result of the MST genotype described above. In our previous study, Cb strains from ticks in Yunnan Province were also typed as MLVA genotype 1 [21], suggesting a wider distribution of this genotype in wildlife in this area.
Although novel MST genotypes of Cb in wild rats were identified in this study, it still has some limitations. The samples were collected only from rats from two locations in Yunnan Province and the number of tested samples was limited. More samples should be collected for analysis in future studies. Moreover, since cases of Q fever have been reported in Yunnan Province [22], there are no Cb genotyping data from patients with Q fever. More studies on the genotyping and pathogenesis of the strains isolated from patients, livestock, wild animals and ticks are necessary to better understand their roles in the epidemiology of Q fever in the province.
Conclusions
Two novel MST types were identified in wild rats collected from Yunnan Province, and the Cb genotypic diversity in wild rats was reported for the first time in China. Further studies are needed to explore the presence of more genotypes and to associate the genotypes circulating in the wildlife-livestock interaction with those causing human disease to further expand on the epidemiological aspects of the pathogen.
Methods
Ethics
All animal care and experimental procedures were in accordance with institutional policies to ensure the highest level of animal health and well-being and were approved by the Institutional Animal Care and Use Committee (IACUC) of the Academy of Military Medical Science (AMMS, Beijing, China) (approval number: IACUC-DWZX-2021–066).
Sample collection
An investigation was conducted in August 2020, and rats were collected from wild fields in Mengla County and Menglian County in Yunnan Province, China. Rat species were identified based on morphological characterization and by molecular biology methods based on the sequences of the species-specific mitochondrial cytochrome c oxidase I (COI) gene as previously described [29]. Rats were anesthetized with pentobarbital sodium (100 mg/kg body weight) and euthanized via cervical dislocation to collect the liver.
DNA extraction
Livers from all the mice were individually homogenized in 300 μL of phosphate buffered saline (PBS), and then the homogenate was subjected to DNA extraction using a QIAamp® Fast DNA Tissue Kit (Qiagen, Dusseldorf, Germany) following the procedure described before [21]. The extracted genomic DNA was stored at -20 °C.
Polymerase Chain Reaction (PCR)
Nested PCR targeting the 16S rRNA gene of Cb was used as described previously [30]. PCR amplifications were carried out in a 50 μL reaction mixture containing 1 × PrimeSTAR® HS (Premix) (TaKaRa, Beijing, China). The PCR products were electrophoresed on a 1.5% agarose gel and visualized under UV light. The positive amplicons were sequenced by TSINGKE Biological Technology (Beijing, China).
Samples positive for Cb were then tested by a quantitative PCR (qPCR) targeting the com1gene of Cb as described previously [31].
Multispacer Sequence Typing (MST)
MST was performed in PCR targeting 10 spacers that exhibited the highest variability, including Cox2, Cox5, Cox18, Cox20, Cox22, Cox37, Cox51, Cox56, Cox57 and Cox61 [2, 18]. Consensus sequences of each Cox spacer were blasted and compared with the sequences in the web-based MST database (https://ifr48.timone.univ-mrs.fr/mst/coxiella_burnetii/). Then the combination of alleles coded of all loci was used to assign the MST genotype. The MST genotypes identified in the present study were compared to genotypes included in the MST database and a phylogenetic analysis was performed using the MEGA X software.
Multilocus Variable Number Tandem Repeats Analysis (MLVA)
MLVA was performed in PCR targeting six highly variable loci, including ms23, ms24, ms27, ms28, ms33, and ms34 [19]. The forward and reverse primer sequences and PCR conditions were applied as described previously [32–34]. The Nine Mile strain of Cb (RSA493) which was considered 9–27-4–6-9–5 for loci ms23-ms24-ms27-ms28-ms33-ms34 was used as the reference. The MLVA pattern of the strains in the present study was identified using the MLVABank database (http://mlva.u-psud.fr/mlvav4/genotyping/). Clustering of the obtained MLVA profiles was performed with Bionumerics v.7.6 software (Applied Maths, Belgium).
Acknowledgements
Not applicable.
Abbreviations
- Cb
Coxiella burnetii
- RFLP-PFGE
Restriction fragment length polymorphism and pulsed-field gel electrophoresis
- SNP
Single nucleotide polymorphism
- MST
Multispacer sequence typing
- MLVA
Multilocus variable number tandem repeats analysis
- COI
Mitochondrial cytochrome c oxidase I
- PBS
Phosphate buffer saline
- PCR
Polymerase Chain Reaction
- qPCR
Quantitative PCR
Authors’ contributions
Jun Jiao and Qinghong Yuan: conceived and designed the study. Mengjiao Fu, Peisheng He, and Xuan OuYang: samples collection, rat species identification and laboratory work. Jun Jiao and Yonghui Yu: data curation. Jun Jiao and Xiaolu Xiong: writing—original draft. Bohai Wen and Dongsheng Zhou: writing—review & editing. The author(s) read and approved the final manuscript.
Funding
This work was supported by the National Key Research and Development Program of China [grant number 2019YFC1200500] and the National Natural Science Foundation of China [grant numbers 32000139 and 31970178].
Availability of data and materials
All data generated or analyzed during current study are available in the GenBank (https://www.ncbi.nlm.nih.gov/WebSub/?form=history&tool=genbank, BankIt2545710).
Declarations
Ethics approval and consent to participate
All animal care and experimental procedures were in accordance with institutional policies for animal health and well-being and were approved by the Institutional Animal Care and Use Committee (IACUC) of the Academy of Military Medical Science (AMMS, Beijing, China) (approval number: IACUC-DWZX-2021–066). All methods were performed in accordance with the ARRIVE guidelines (https://arriveguidelines.org).
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Mengjiao Fu and Peisheng He contributed equally to this work.
Contributor Information
Mengjiao Fu, Email: rainshenmin@163.com.
Peisheng He, Email: 672223510@qq.com.
Xuan OuYang, Email: ouyangxuan626@163.com.
Yonghui Yu, Email: yuyhjlu@163.com.
Bohai Wen, Email: bohaiwen@sohu.com.
Dongsheng Zhou, Email: dongshengzhou1977@gmail.com.
Xiaolu Xiong, Email: xiongxiaolu624@sohu.com.
Qinghong Yuan, Email: ynyuanqh@163.com.
Jun Jiao, Email: jiaojun51920@sina.com.
References
- 1.Mioni MSR, Sidi-Boumedine K, Morales Dalanezi F, Fernandes Joaquim S, Denadai R, Reis Teixeira WS, Bahia Labruna M, Megid J. New genotypes of Coxiella burnetii circulating in Brazil and Argentina. Pathogens. 2019;9(1):30. [DOI] [PMC free article] [PubMed]
- 2.Di Domenico M, Curini V, De Massis F, Di Provvido A, Scacchia M, Camma C. Coxiella burnetii in central Italy: novel genotypes are circulating in cattle and goats. Vector Borne Zoonotic Dis. 2014;14(10):710–5. [DOI] [PMC free article] [PubMed]
- 3.Tokarevich NK, Panferova YA, Freylikhman OA, Blinova OV, Medvedev SG, Mironov SV, Grigoryeva LA, Tretyakov KA, Dimova T, Zaharieva MM, et al. Coxiella burnetii in ticks and wild birds. Ticks Tick Borne Dis. 2019;10(2):377–85. [DOI] [PubMed]
- 4.Barandika JF, Hurtado A, Garcia-Esteban C, Gil H, Escudero R, Barral M, Jado I, Juste RA, Anda P, Garcia-Perez AL. Tick-borne zoonotic bacteria in wild and domestic small mammals in northern Spain. Appl Environ Microbiol. 2007;73(19):6166–6171. doi: 10.1128/AEM.00590-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Duron O, Sidi-Boumedine K, Rousset E, Moutailler S, Jourdain E. the importance of ticks in q fever transmission: what has (and has not) been demonstrated? Trends Parasitol. 2015;31(11):536–552. doi: 10.1016/j.pt.2015.06.014. [DOI] [PubMed] [Google Scholar]
- 6.Pacheco RC, Echaide IE, Alves RN, Beletti ME, Nava S, Labruna MB. Coxiella burnetii in ticks, Argentina. Emerg Infect Dis. 2013;19(2):344–6. [DOI] [PMC free article] [PubMed]
- 7.Maurin M, Raoult D. Q fever. Clin Microbiol Rev. 1999;12(4):518–553. doi: 10.1128/CMR.12.4.518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hu X, Yu Y, Feng J, Fu M, Dai L, Lu Z, Luo W, Wang J, Zhou D, Xiong X, et al. Pathologic changes and immune responses against Coxiella burnetii in mice following infection via non-invasive intratracheal inoculation. PLoS One. 2019;14(12):e0225671. [DOI] [PMC free article] [PubMed]
- 9.Raoult D, Marrie T, Mege J. Natural history and pathophysiology of Q fever. Lancet Infect Dis. 2005;5(4):219–226. doi: 10.1016/S1473-3099(05)70052-9. [DOI] [PubMed] [Google Scholar]
- 10.Kwak W, Chu H, Hwang S, Park JH, Hwang KJ, Gwack J, Choi YS, Youn SK, Park MY. Epidemiological characteristics of serologically confirmed q Fever cases in South Korea, 2006–2011. Osong Public Health Res Perspect. 2013;4(1):34–38. doi: 10.1016/j.phrp.2012.12.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Roest HI, Tilburg JJ, van der Hoek W, Vellema P, van Zijderveld FG, Klaassen CH, Raoult D. The Q fever epidemic in The Netherlands: history, onset, response and reflection. Epidemiol Infect. 2011;139(1):1–12. doi: 10.1017/S0950268810002268. [DOI] [PubMed] [Google Scholar]
- 12.Stein A, Kruszewska D, Gouvernet J, Raoult D. Study of the 16S–23S ribosomal DNA internal spacer of Coxiella burnetii. Eur J Epidemiol. 1997;13(4):471–5. [DOI] [PubMed]
- 13.Willems H, Thiele D, Krauss H. Plasmid based differentiation and detection of Coxiella burnetii in clinical samples. Eur J Epidemiol. 1993;9(4):411–8. [DOI] [PubMed]
- 14.Jager C, Lautenschlager S, Willems H, Baljer G. Coxiella burnetii plasmid types QpDG and QpH1 are closely related and likely identical. Vet Microbiol. 2002;89(2–3):161–6. [DOI] [PubMed]
- 15.Jager C, Willems H, Thiele D, Baljer G. Molecular characterization of Coxiella burnetii isolates. Epidemiol Infect. 1998;120(2):157–64. [DOI] [PMC free article] [PubMed]
- 16.Huijsmans CJ, Schellekens JJ, Wever PC, Toman R, Savelkoul PH, Janse I, Hermans MH. Single-nucleotide-polymorphism genotyping of Coxiella burnetii during a Q fever outbreak in The Netherlands. Appl Environ Microbiol. 2011;77(6):2051–7. [DOI] [PMC free article] [PubMed]
- 17.Enright MC, Spratt BG. Multilocus sequence typing. Trends Microbiol. 1999;7(12):482–487. doi: 10.1016/S0966-842X(99)01609-1. [DOI] [PubMed] [Google Scholar]
- 18.Glazunova O, Roux V, Freylikman O, Sekeyova Z, Fournous G, Tyczka J, Tokarevich N, Kovacava E, Marrie TJ, Raoult D. Coxiella burnetii genotyping. Emerg Infect Dis. 2005;11(8):1211–7. [DOI] [PMC free article] [PubMed]
- 19.Arricau-Bouvery N, Hauck Y, Bejaoui A, Frangoulidis D, Bodier CC, Souriau A, Meyer H, Neubauer H, Rodolakis A, Vergnaud G. Molecular characterization of Coxiella burnetii isolates by infrequent restriction site-PCR and MLVA typing. BMC Microbiol. 2006;6:38. [DOI] [PMC free article] [PubMed]
- 20.Gong XQ, Xiao X, Liu JW, Han HJ, Qin XR, Lei SC, Yu XJ. Occurrence and Genotyping of Coxiella burnetii in Hedgehogs in China. Vector Borne Zoonotic Dis. 2020;20(8):580–5. [DOI] [PubMed]
- 21.Jiao J, Zhang J, He P, OuYang X, Yu Y, Wen B, Sun Y, Yuan Q, Xiong X. Identification of Tick-Borne Pathogens and Genotyping of Coxiella burnetii in Rhipicephalus microplus in Yunnan Province. China Front Microbiol. 2021;12:736484. [DOI] [PMC free article] [PubMed]
- 22.El-Mahallawy HS, Lu G, Kelly P, Xu D, Li Y, Fan W, Wang C. Q fever in China: a systematic review, 1989–2013. Epidemiol Infect. 2015;143(4):673–681. doi: 10.1017/S0950268814002593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Huang M, Ma J, Jiao J, Li C, Chen L, Zhu Z, Ruan F, Xing L, Zheng X, Fu M, et al. The epidemic of Q fever in 2018 to 2019 in Zhuhai city of China determined by metagenomic next-generation sequencing. PLoS Negl Trop Dis. 2021;15(7):e0009520. doi: 10.1371/journal.pntd.0009520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Li K, Luo H, Shahzad M. Epidemiology of Q fever in goats in Hubei province of China. Trop Anim Health Prod. 2018;50(6):1395–1398. doi: 10.1007/s11250-018-1561-3. [DOI] [PubMed] [Google Scholar]
- 25.Batu N, Wang Y, Liu Z, Huang T, Bao W, He H, Geri L. Molecular epidemiology of Rickettsia sp. and Coxiella burnetii collected from Hyalomma asiaticum in Bactrian camels (Camelus bactrianus) in inner Mongolia of China. Ticks Tick Borne Dis. 2020;11(6):101548. [DOI] [PubMed]
- 26.Szymanska-Czerwinska M, Jodelko A, Zareba-Marchewka K, Niemczuk K. Shedding and genetic diversity of Coxiella burnetii in Polish dairy cattle. PLoS One. 2019;14(1):e0210244. [DOI] [PMC free article] [PubMed]
- 27.Astobiza I, Tilburg JJ, Pinero A, Hurtado A, Garcia-Perez AL, Nabuurs-Franssen MH, Klaassen CH. Genotyping of Coxiella burnetii from domestic ruminants in northern Spain. BMC Vet Res. 2012;8:241. [DOI] [PMC free article] [PubMed]
- 28.Santos AS, Tilburg JJ, Botelho A, Barahona MJ, Nuncio MS, Nabuurs-Franssen MH, Klaassen CH. Genotypic diversity of clinical Coxiella burnetii isolates from Portugal based on MST and MLVA typing. Int J Med Microbiol. 2012;302(6):253–6. [DOI] [PubMed]
- 29.Jarquin-Diaz VH, Balard A, Macova A, Jost J, von RothSzepesbela T, Berktold K, Tank S, Kvicerova J, Heitlinger E. Generalist Eimeria species in rodents: Multilocus analyses indicate inadequate resolution of established markers. Ecol Evol. 2020;10(3):1378–1389. doi: 10.1002/ece3.5992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Duron O, Noel V, McCoy KD, Bonazzi M, Sidi-Boumedine K, Morel O, Vavre F, Zenner L, Jourdain E, Durand P, et al. The recent evolution of a maternally-inherited endosymbiont of ticks led to the emergence of the Q fever pathogen, Coxiella burnetii. PLoS Pathog. 2015;11(5):e1004892. [DOI] [PMC free article] [PubMed]
- 31.Lockhart MG, Graves SR, Banazis MJ, Fenwick SG, Stenos J. A comparison of methods for extracting DNA from Coxiella burnetii as measured by a duplex qPCR assay. Lett Appl Microbiol. 2011;52(5):514–20. [DOI] [PubMed]
- 32.Klaassen CH, Nabuurs-Franssen MH, Tilburg JJ, Hamans MA, Horrevorts AM. Multigenotype Q fever outbreak, the Netherlands. Emerg Infect Dis. 2009;15(4):613–614. doi: 10.3201/eid1504.081612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tilburg JJ, Rossen JW, van Hannen EJ, Melchers WJ, Hermans MH, van de Bovenkamp J, Roest HJ, de Bruin A, Nabuurs-Franssen MH, Horrevorts AM, et al. Genotypic diversity of Coxiella burnetii in the 2007–2010 Q fever outbreak episodes in The Netherlands. J Clin Microbiol. 2012;50(3):1076–8. [DOI] [PMC free article] [PubMed]
- 34.Gonzalez-Barrio D, Hagen F, Tilburg JJ, Ruiz-Fons F. Coxiella burnetii genotypes in Iberian Wildlife. Microb Ecol. 2016;72(4):890–7. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data generated or analyzed during current study are available in the GenBank (https://www.ncbi.nlm.nih.gov/WebSub/?form=history&tool=genbank, BankIt2545710).