Abstract
Activated sludge was fed phenol as the sole carbon source, and the phenol-loading rate was increased stepwise from 0.5 to 1.0 g liter−1 day−1 and then to 1.5 g liter−1 day−1. After the loading rate was increased to 1.5 g liter−1 day−1, nonflocculating bacteria outgrew the sludge, and the activated-sludge process broke down within 1 week. The bacterial population structure of the activated sludge was analyzed by temperature gradient gel electrophoresis (TGGE) of PCR-amplified 16S ribosomal DNA (rDNA) fragments. We found that the population diversity decreased as the phenol-loading rate increased and that two populations (designated populations R6 and R10) predominated in the sludge during the last several days before breakdown. The R6 population was present under the low-phenol-loading-rate conditions, while the R10 population was present only after the loading rate was increased to 1.5 g liter−1 day−1. A total of 41 bacterial strains with different repetitive extragenic palindromic sequence PCR patterns were isolated from the activated sludge under different phenol-loading conditions, and the 16S rDNA and gyrB fragments of these strains were PCR amplified and sequenced. Some bacterial isolates could be associated with major TGGE bands by comparing the 16S rDNA sequences. All of the bacterial strains affiliated with the R6 population had almost identical 16S rDNA sequences, while the gyrB phylogenetic analysis divided these strains into two physiologically divergent groups; both of these groups of strains could grow on phenol, while one group (designated the R6F group) flocculated in laboratory media and the other group (the R6T group) did not. A competitive PCR analysis in which specific gyrB sequences were used as the primers showed that a population shift from R6F to R6T occurred following the increase in the phenol-loading rate to 1.5 g liter−1 day−1. The R10 population corresponded to nonflocculating phenol-degrading bacteria. Our results suggest that an outbreak of nonflocculating catabolic populations caused the breakdown of the activated-sludge process. This study also demonstrated the usefulness of gyrB-targeted fine population analyses in microbial ecology.
One of the practical aspects of microbial ecology is its contribution to improvements in the design and operation of environment protection processes, including wastewater treatment and in situ bioremediation. One important step in such studies is to understand the diversity and abundance of the microbial populations that occur in these processes. In the 1990s, molecular approaches, especially those known as the rRNA phylogenetic framework (20, 21), have been proven to be useful for analyzing naturally occurring microbial populations. These approaches have been used to analyze microbial community structures in wastewater treatment processes (27, 29) and in situ bioremediation sites (7). Another important step is to identify functionally important populations in the microbial community (4, 35). In our previous study, a functionally dominant phenol-degrading population in activated sludge was identified and characterized by combining data obtained by direct molecular detection of populations in the sludge and data obtained by genetic and physiological analyses of isolated bacteria (35). One goal of such studies is to develop methods which allow for artificial control of microbial communities in the processes. We think that some clues to attaining this objective may be obtained by carefully analyzing the dynamics of major populations in the processes under different operational conditions.
We have studied phenol-digesting activated sludge (31–33, 35) for the following reasons. First, phenolic compounds are known to be major pollutants in wastewater from industrial plants, such as oil refineries, petrochemical plants, coking plants, and phenol resin industry plants (1a, 22, 32). Although these compounds have been treated for many years by using activated-sludge processes, more reliable processes are desired; this is especially relevant when there is a fluctuation in the loading rate of the compounds. Second, phenol-digesting activated sludge is considered an appropriate model for studying how to manage and control a microbial community. To date, a number of phenol-degrading bacteria have been isolated from activated sludge (26, 30, 35), and there is a great deal of information concerning the physiology and genetics of these organisms (14, 26, 35); this background information may be useful for understanding the physiological and molecular events that occur in phenol-digesting activated sludge. In addition, there have been many engineering studies of phenol-digesting activated sludge (2, 19, 23).
This study was conducted to investigate the changes in the major bacterial populations in phenol-digesting activated sludge in response to increases in the phenol-loading rate. Previous reports have suggested that population shifts in phenol-digesting activated sludge at a high phenol-loading rate are one of the possible causes of the breakdown of activated-sludge processes (23, 32). Okaygun and Akgerman (19) observed a shift in the microbial species at different influent phenol concentrations in a continuously stirred tank reactor; these authors used conventional plating to detect bacteria. In this study, using PCR-based direct-detection methods, we analyzed population shifts in phenol-digesting activated sludge following increases in the phenol-loading rate. Based on the results obtained, possible measures to enhance the phenol treatability of activated sludge are proposed.
MATERIALS AND METHODS
Operation of a laboratory activated-sludge process.
Samples of activated-sludge mixed liquor were obtained from the return sludge line of a municipal sewage treatment plant (Ohdaira, Kamaishi, Iwate, Japan) in March and June 1998. The activated sludge was acclimated to phenol as the sole carbon source in a laboratory activated-sludge unit composed of a 3-liter aeration tank and a 2-liter settling tank, as described previously (33). The inorganic ingredients in the feed were (per liter) 3.75 g of (NH4)2HPO4, 2.5 g of NH4H2PO4, 1.0 g of MgSO4 · 7H2O, 0.5 g of KCl, 0.1 g of NaCl, and 2 ml of trace element solution B (6); the pH of the feed was 6.8 to 7.0. The concentrations of the inorganic ingredients in the feed were kept constant under different phenol-loading conditions. The inorganic compounds were considered to be sufficiently supplied even at a feed phenol concentration of 5.0 g per liter (9). The phenol-loading rate was increased stepwise by increasing the phenol concentration in the feed. The dilution rate was kept constant at 6 liters per day, so the hydraulic residence time was 0.5 day. The concentration of mixed-liquor suspended solids (MLSS) in the aeration tank was kept between 1,800 and 2,500 mg per liter by continuously discarding the excess sludge from the aeration tank. Air was continuously supplied at a rate of 2 liters per min, and the temperature was maintained at approximately 25°C. The dissolved oxygen concentration was maintained at more than 5 mg per liter. The total direct count of bacteria in the activated sludge was determined by a fluorescent-microscopy method after staining with 4′,6-diamidino-2-phenylindole (DAPI) (33). The phenol concentration in the aeration tank was measured by a colorimetric assay performed with Phenol Test Wako (Wako Pure Chemicals) (33). The total organic carbon (TOC) concentration in the aeration tank was determined with a model TOC-5000 TOC meter (Shimadzu) (33). The phenol-oxygenating activity was measured at a phenol concentration of 10 μM with a Clark type oxygen electrode as described previously (32). One unit of activity was equivalent to 1 μmol of oxygen consumed per min, while the specific activity was defined as the amount of activity per gram of MLSS.
DNA extraction from activated sludge.
DNA was extracted from 5 ml of a mixed liquor sample obtained from the aeration tank of the laboratory unit as described previously (33). The quantity and quality of the extracted DNA were checked by measuring the UV absorption spectrum of the DNA solution (25), and the DNA was dissolved in TE buffer (25) at a concentration of 100 μg per ml.
TGGE.
PCR primers P2 and P3 (containing a 40-bp GC clamp) (18) were used to amplify variable region V3 of bacterial 16S ribosomal DNA (rDNA) (corresponding to positions 341 to 534 in the Escherichia coli sequence). Amplification was performed with a Progene thermal cycler (Techne) as described previously (35). A temperature gradient gel electrophoresis (TGGE) system (Taitec) was used as described previously (35). The PCR products were electrophoresed in 10% (wt/vol) polyacrylamide gels at 250 V for 3.5 h by using a linear temperature gradient ranging from 45 to 60°C. After electrophoresis, the gel was stained with SYBR Green I (FMC Bioproducts) for 30 min. The nucleotide sequences of TGGE bands were determined as described previously (35).
Isolation of bacteria from activated sludge.
Bacteria were isolated from the activated sludge by direct plating on agar plates containing dCGY medium (referred to below as dCGY plates) (35). Mixed liquor from the phenol-digesting activated sludge (5 ml) obtained from the aeration tank of the laboratory unit was mixed with 0.5 ml of 50 mM sodium tripolyphosphate. In order to deflocculate the activated sludge, the mixture was treated in a blender (Wheaton Instruments) for 2 min. The resulting cell suspension was appropriately diluted with sterile MP medium (30) containing 5 mM sodium tripolyphosphate and then spread onto dCGY plates. The plates were incubated at 25°C for 7 days. All of the colonies that appeared on one plate were picked and grown in 5 ml of dCGY medium, and the dCGY medium cultures were restreaked onto dCGY plates. This purification procedure was repeated several times.
The purified colonies were subjected to repetitive extragenic palindromic sequence PCR (rep-PCR) to identify identical strains, as described previously (35). The rep-PCR analysis was repeated several times to determine the reproducibility of the method.
Sequencing of 16S rDNA of isolated bacteria.
A small amount of bacterial cells picked from a colony that developed on a dCGY plate was subjected to PCR in order to amplify an almost full-length 16S rDNA fragment. The nucleotide sequences of the primers used were 5′-AGAGTTTGATCCTGGCTCAG-3′ (E. coli 16S rDNA positions 8 to 27 [5]) and 5′-CAKAAAGGAGGTGATCC-3′ (E. coli 16S rDNA positions 1529 to 1546 [5]). Amplification was performed with a Progene thermal cycler (Techne) by using a 50-μl mixture containing 1.25 U of Taq DNA polymerase (Amplitaq Gold; Perkin-Elmer), 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, 0.001% (wt/vol) gelatin, each deoxynucleoside triphosphate at a concentration of 200 μM, 50 pmol of each primer, and a small amount of bacterial cells. The PCR conditions used were as follows: 10 min of activation of the polymerase at 94°C, followed by 40 cycles consisting of 1 min at 94°C, 1 min at 50°C, and 2 min at 72°C, and finally 10 min of extension at 72°C. The PCR products were electrophoresed through a 1.5% (wt/vol) agarose gel with TBE buffer (25) and then purified with a QIAquick gel extraction kit (QIAGEN). The extracted DNA was quantified by measuring the absorbance at 260 and 320 nm (23). The nucleotide sequences of the PCR products were then determined by using a Dye terminator cycle DNA sequencing kit (Perkin-Elmer) as described by Edwards et al. (10). The products of the sequencing reactions were analyzed with a model 377 DNA sequencer (Perkin-Elmer).
Nucleotide sequences of variable region V3 of 16S rDNA were determined as described previously (35).
Sequencing of gyrB genes of isolated bacteria.
A small amount of bacterial cells picked from a colony that developed on a dCGY plate was subjected to PCR in order to amplify a partial fragment of the gyrB gene. The primers used were GY-21 and UP-11 (16). Amplification was performed with a Progene thermal cycler by using a 50-μl mixture whose composition was the same as that of the mixture used for 16S rDNA amplification. The standard PCR conditions used were as follows: 10 min of activation of the polymerase at 94°C, followed by 40 cycles consisting of 1 min at 94°C, 1 min at 55°C, and 2 min at 72°C, and finally 10 min of extension at 72°C. The PCR products were electrophoresed and purified as described above. The nucleotide sequences of the PCR products were then determined by using Dye terminator cycle DNA sequencing kit with primer GY-21 or UP-11. The products of the sequencing reactions were analyzed with a model 377 DNA sequencer.
Nucleotide sequence analysis.
Database searches were conducted by using the BLAST program (15) with the GenBank database (for 16S rDNA) and the ICB database (for gyrB) (16). The sequences determined in this study and those retrieved from the databases were aligned by using ClustalW, version 1.7 (28). Alignments were refined by visual inspection. Neighbor-joining trees (24) were constructed by using njplot software in ClustalW, version 1.7. Nucleotide positions at which any sequence had a gap or an ambiguous base were not included in the calculations.
Physiological characterization of the isolated bacteria.
Utilization of phenol and sugars as growth substrates by the isolated bacteria was examined as follows. Ten milliliters of MP medium was supplemented with phenol or sugar as the sole carbon source at an initial concentration of 50 mg per liter and was inoculated with a small amount of bacterial cells picked from a single colony on a dCGY plate. The preparation was then shaken reciprocally at 60 rpm, and the growth of bacteria was monitored by automatically measuring the optical density at 660 nm with a model TN-2612 Bio-photometer (Advantec) for 5 days. After this, floc formation in the culture was checked by visual inspection, and the substrate concentration was measured either by the colorimetric assay described above (for phenol) or by the phenol-sulfuric acid method (for sugars) (8).
cPCR.
The primers used for competitive PCR (cPCR) were designed by comparing the gyrB sequences of strains shown in Fig. 1. PCR primers R6F-F (5′-CAGGAGATTTTCAAAGAGAACTT-3′) and gyC4R (5′-CCTTCGCGCATGTCGT-3′) were used to specifically detect the R6F population (see below), PCR primers R6T-F (5′-ACCGAAATCTTCAAGGAAAACCA-3′) and gyC4R were used to specifically detect the R6T population, and PCR primers R10-F (5′-GCACGCGGCGCG-3′) and gyC4R were used to specifically detect the R10 population. Competitor fragments were produced by using a competitive DNA construction kit (Takara Shuzo Ltd.). The sizes of the target fragments and the competitors were 468 and 539 bp for the R6F population, 468 and 539 bp for the R6T population, and 586 and 528 bp for the R10 population, respectively. Amplification was performed with a Progene thermal cycler (Techne) by using a 50-μl mixture containing 1.25 U of Taq DNA polymerase, 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, 0.001% (wt/vol) gelatin, each deoxynucleoside triphosphate at a concentration of 200 μM, 50 pmol of each primer, 50 ng of activated-sludge DNA, and an appropriate amount of a competitor. The PCR conditions used were as follows: 10 min of activation of the polymerase at 94°C, followed by 35 cycles consisting of 1 min at 94°C, 1 min at 63°C (for the R6F and R6T populations) or 65°C (for the R10 population), and 2 min at 72°C, and finally 10 min of extension at 72°C. Two microliters of the PCR product was electrophoresed through a 1.5% (wt/vol) agarose gel with TBE buffer, and the gel was photographed after it was stained with SYBR Green I. The band intensity was quantified with image processing software (NIH IMAGE, version 1.60; National Institutes of Health), and then the copy number of a target sequence in the PCR mixture was determined by comparing the band intensities of the target fragment and the competitor. The number of bacterial cells was considered to be equal to the copy number of the gyrB sequence (3).
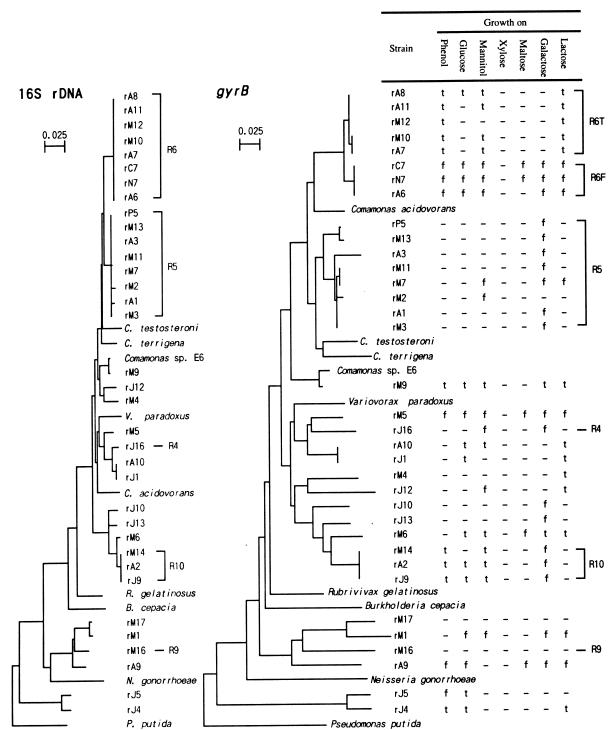
FIG. 1.
Phylogenetic trees based on the nucleotide sequences of 16S rDNA and gyrB genes, showing the relationships among the bacteria isolated from the phenol-digesting activated sludge. Pseudomonas putida JCM 6156 was used as the outgroup. The 16S rDNA sequences of previously described strains were retrieved from GenBank, and the accession numbers were as follows: Comamonas testosteroni, D87101; Comamonas terigena, AF078772; Comamonas acidovorans, AB020186; Comamonas sp. strain E6, AB008429; Variovorax paradoxus, D30793; Rubrivivax gelatinosus, D16213; Burkholderia cepacia, L28675; Neisseria gonorrhoeae, X07714; and P. putida, D37924. The gyrB sequences were retrieved from the ICB database (16); the accession numbers of these sequences were gy00024, gy10051, gy00022, gy10391, gy10066, gy10003, gy10339, gy00036, and gy10157, respectively. Bars = 0.025 substitution per amino acid site. f, floc formed; t, turbid; —, not grown.
Nucleotide sequence accession numbers.
The nucleotide sequences determined in this study have been deposited in the GSDB, DDBJ, EMBL, and NCBI nucleotide sequence databases under accession no. AB021320 to AB021362, AB021427 to AB021458, and AB021661.
RESULTS
Phenol digestion by activated sludge.
Our previous study showed that the population structure in activated sludge became stable 10 days after phenol loading was started at a rate of 0.4 g per liter per day, and after that the activated sludge stably digested phenol (35). We repeated the same type of experiment with the Ohdaira activated sludge. We observed that the sludge could cope with phenol added at a rate of 0.5 g per liter per day and that the bacterial population structure in the sludge, as determined by TGGE, became stable 7 days after the phenol addition began (data not shown). In the present study, the initial phenol load added to the activated-sludge unit was 0.5 g per liter per day, and the phenol load was then increased to 1.0 g per liter per day. As shown in Fig. 2a, the activated sludge almost completely digested the phenol (the phenol concentrations were less than 1 mg per liter) for 21 days when the loading rate was 1.0 g per liter. Then, the loading rate was increased to 1.5 g per liter per day; phenol started to leak from the activated-sludge unit 7 days after the loading rate was increased. At this time, nonflocculating microorganisms became dominant, so the effluent water from the unit became turbid. Several days later, all of the flocs were washed out, and the phenol concentration in the aeration tank was approximately 750 mg per liter (i.e., the concentration in the feed). These observations indicate that the activated sludge could not cope with phenol at a loading rate of 1.5 g per liter per day.
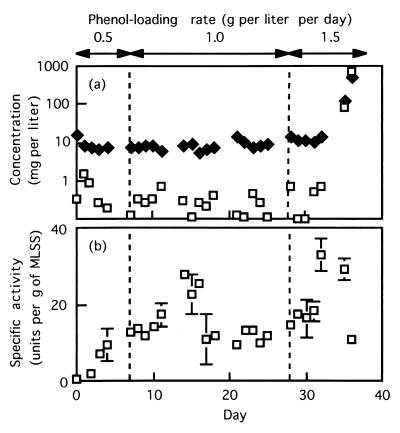
FIG. 2.
(a) Changes in the phenol concentration (□) and TOC concentration (⧫) in response to stepwise increases in the phenol-loading rate. (b) Changes in the specific phenol-oxygenating activity of activated sludge in response to stepwise increases in the phenol-loading rate.
Figure 2a also shows the TOC values during the experiment; the TOC value was approximately 10 mg per liter except on days 35 and 36, indicating that the quality of the effluent was good before the leakage of phenol. The phenol-oxygenating activity pattern is shown in Fig. 2b. The activity was between 10 and 15 U per g of MLSS except for days 14 to 17 and 32 to 35. High levels of activity were detected several days after the phenol-loading rate was increased.
TGGE.
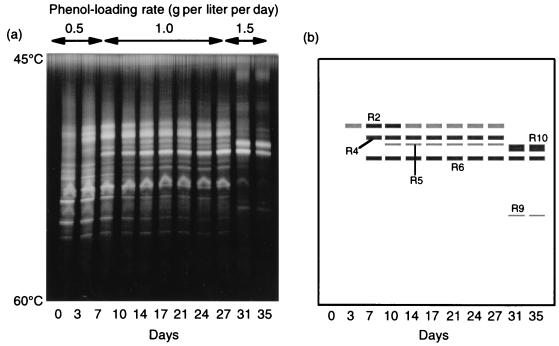
TGGE of partial 16S rDNA fragments that were PCR amplified from activated-sludge DNA was conducted to analyze changes in bacterial populations following the increases in the phenol-loading rate (Fig. 3). We found that the population diversity decreased after the loading rate was increased and that two populations (designated populations R6 and R10) eventually became dominant. The R6 band was detected from day 7 on, while the R10 band appeared after the loading rate was increased to 1.5 g per liter per day. Major bands that appeared after the phenol loading was begun were excised and sequenced (Table 1). Several bands at the same temperature position were excised from profiles on different days and were sequenced to confirm their identities. We found that the sequence of the R6 band was identical to the sequence of the dominant phenol-degrading population identified in our previous study (35). The database search suggested that the sequences were all affiliated with the beta subclass of the class Proteobacteria.
FIG. 3.
(a) TGGE profiles of the partial 16S rDNA fragments, showing shifts in major bacterial populations in phenol-digesting activated sludge in response to stepwise increases in the phenol-loading rate. (b) Drawing of the TGGE gel from panel a, showing the bands excised for sequence analysis.
TABLE 1.
Sequence analysis of major TGGE bands occurring after phenol loading
| TGGE band | Length (bases) | Phylogenetically related organism (accession no.) | % Identity |
|---|---|---|---|
| R2 | 194 | Aquaspirillum psychrophilum LMG 5408 (AF078755) | 97 |
| R4 | 194 | Comamonas sp. strain R7 (AJ002810) | 95 |
| R5 | 194 | Aquaspirillum delicatum LMG 4328 (AF078756) | 98 |
| R6 | 194 | Aquaspirillum delicatum LMG 4328 (AF078756) | 97 |
| R9 | 194 | Alcaligenes defragrans 65Phen (AJ005450) | 93 |
| R10 | 194 | Variovorax paradoxus E4 (AB008000) | 97 |
Isolation of bacteria.
In order to further characterize bacterial populations detected by the TGGE analysis, we attempted to isolate bacteria corresponding to the major TGGE bands (Table 2). Bacteria were isolated by the direct-plating method from the activated sludge when the phenol-loading rate was either 0.5, 1.0, or 1.5 g per liter per day. Bacteria isolated from each activated-sludge sample were examined by rep-PCR (data not shown), and then 16S rDNA sequences of strains that produced different rep-PCR patterns were determined (Table 2). In addition, the gyrB sequences of isolated strains belonging to the beta subclass of the class Proteobacteria were also determined (Table 2). As shown in Table 2, strains that produced identical rep-PCR patterns were obtained from the same and different activated-sludge samples. When the number of colonies picked and the number of rep-PCR patterns obtained were compared, it was clear that the genetic diversity of colonies was smaller in activated-sludge samples subjected to higher phenol-loading rates. Table 2 also shows that many strains had 16S rDNA sequences identical to the sequences in the TGGE bands, especially bands R5 and R6. However, no strain had a 16S rDNA sequence identical to the sequence of band R2.
TABLE 2.
Isolates obtained
| Isolate | No. of colonies/total no. of colonies | Identical rep-PCR pattern(s) | 16S rDNA fragment
|
gyrB fragment
|
|||
|---|---|---|---|---|---|---|---|
| Length of sequenced part (bases)a | Phylogenetically related organism (% identity) | Identical TGGE band | Length of sequenced part (bases)a | Phylogenetically related organism (% identity)b | |||
| Isolates obtained from activated sludge at 0.5 g per liter per day (day 7) | |||||||
| rJ1 | 1/19 | rA10 | 1,464 | Variovorax paradoxus (95) | 651 | Variovorax paradoxus (90) | |
| rJ2 | 1/19 | NA | NA | ||||
| rJ3 | 1/19 | 1,464 | Moraxella osloensis (99) | — | |||
| rJ4 | 2/19 | 1,419 | Nevskia ramosa (93) | 831 | NHS | ||
| rJ5 | 2/19 | 1,467 | Nevskia ramosa (92) | 832 | NHS | ||
| rJ6 | 1/19 | 1,443 | Aureobacterium testaceum (97) | — | |||
| rJ7 | 1/19 | 1,452 | Sporichthya polymorpha (93) | — | |||
| rJ8 | 1/19 | 1,430 | Kocuria erythromyxa (97) | — | |||
| rJ9 | 1/19 | rM14, rA2 | 1,458 | Hydrogenophaga palleronii (92) | R10 | 971 | Variovorax paradoxus (81) |
| rJ10 | 1/19 | 1,454 | Hydrogenophaga palleronii (93) | 661 | Variovorax paradoxus (83) | ||
| rJ11 | 1/19 | 1,473 | Pseudomonas citronellolis (98) | — | |||
| rJ12 | 1/19 | 1,470 | Comamonas sp. strain E6 (97) | 910 | Variovorax paradoxus (80) | ||
| rJ13 | 1/19 | 1,480 | Hydrogenophaga palleronii (93) | 970 | Variovorax paradoxus (82) | ||
| rJ14 | 1/19 | 1,448 | Sporichthya polymorpha (93) | — | |||
| rJ15 | 1/19 | 1,487 | Xanthomonas campestris (94) | — | |||
| rJ16 | 1/19 | 1,461 | Variovorax paradoxus (96) | R4 | 970 | Variovorax paradoxus (88) | |
| rJ17 | 1/19 | 1,468 | Pseudomonas citronellolis (98) | — | |||
| Isolates obtained from activated sludge at 1.0 g per liter per day (day 15) | |||||||
| rM1 | 4/27 | 1,461 | Alcaligenes sp. strain R4 (94) | 903 | NHS | ||
| rM2 | 3/27 | 1,424 | Comamonas testosteroni (96) | R5 | 966 | Comamonas testosteroni (88) | |
| rM3 | 1/27 | 1,470 | Comamonas testosteroni (96) | R5 | 953 | Comamonas testosteroni (89) | |
| rM4 | 1/27 | 1,446 | Comamonas sp. strain E6 (97) | 970 | Variovorax paradoxus (83) | ||
| rM5 | 1/27 | 1,474 | Variovorax paradoxus (95) | 966 | Variovorax paradoxus (86) | ||
| rM6 | 3/27 | 1,466 | Hydrogenophaga palleronii (92) | 891 | Variovorax paradoxus (81) | ||
| rM7 | 1/27 | 1,464 | Comamonas testosteroni (96) | R5 | 891 | Comamonas testosteroni (88) | |
| rM8 | 2/27 | 194 | Xanthomonas campestris (98) | — | |||
| rM9 | 1/27 | 1,444 | Comamonas sp. strain E6 (98) | 917 | Comamonas sp. strain E6 (98) | ||
| rM10 | 2/27 | rA7 | 1,457 | Comamonas testosteroni (96) | R6 | 898 | Comamonas acidovorans (89) |
| rM11 | 2/27 | 1,456 | Comamonas testosteroni (96) | R5 | 913 | Comamonas testosteroni (88) | |
| rM12 | 1/27 | 1,448 | Comamonas testosteroni (96) | R6 | 901 | Comamonas acidovorans (90) | |
| rM13 | 1/27 | 1,453 | Comamonas testosteroni (96) | R5 | 905 | Comamonas testosteroni (88) | |
| rM14 | 1/27 | rJ9, rA2 | 1,474 | Hydrogenophaga palleronii (92) | R10 | 966 | Variovorax paradoxus (81) |
| rM15 | 1/27 | 189 | Flavobacterium heparinum (97) | — | |||
| rM16 | 1/27 | 1,470 | Alcaligenes sp. strain L6 (96) | R9 | 833 | NHS | |
| rM17 | 1/27 | 1,494 | Alcaligenes sp. strain R4 (94) | 899 | NHS | ||
| Isolates obtained from activated sludge at 1.5 g per liter per day (day 35) | |||||||
| rA1 | 1/22 | 1,475 | Comamonas testosteroni (96) | R5 | 959 | Comamonas testosteroni (89) | |
| rA2 | 9/22 | rJ9, rM14 | 1,499 | Hydrogenophaga palleronii (92) | R10 | 970 | Variovorax paradoxus (81) |
| rA3 | 2/22 | 1,445 | Comamonas testosteroni (96) | R5 | 970 | Comamonas testosteroni (86) | |
| rA4 | 2/22 | 189 | Sphingobacteium mizutae (93) | — | |||
| rA5 | 1/22 | NA | NA | ||||
| rA6 | 1/22 | 1,463 | Comamonas testosteroni (96) | R6 | 904 | Comamonas acidovorans (89) | |
| rA7 | 2/22 | rM10 | 1,447 | Comamonas testosteroni (96) | R6 | 886 | Comamonas acidovorans (89) |
| rA8 | 1/22 | 1,442 | Comamonas testosteroni (96) | R6 | 910 | Comamonas acidovorans (90) | |
| rA9 | 1/22 | 1,471 | Alcaligenes defragrans (98) | 846 | NHS | ||
| rA10 | 1/22 | rJ1 | 1,422 | Variovorax paradoxus (95) | 904 | Variovorax paradoxus (88) | |
| rA11 | 1/22 | 1,456 | Comamonas testosteroni (96) | R6 | 907 | Comamonas acidovorans (90) | |
NA, a DNA fragment could not be amplified; —, not analyzed.
NHS, no homologous sequence (>80%) was found in the ICB database.
Phylogeny of isolates.
To determine the phylogenetic relationships among the isolated bacteria neighbor-joining trees were constructed by using the sequences of almost full-length 16S rDNA and partial gyrB fragments (Fig. 1). gyrB encodes the subunit B protein of DNA gyrase (topoisomerase type II) (17), and it has been suggested that this gene is useful for phylogenetic analysis of bacteria (12, 36). Strains rC7, rN7, and rP5 were isolated from phenol-acclimated Ohdaira activated sludge in our previous study (35), and their 16S rDNA and gyrB sequences were determined in this study. The data for these strains were also included in the trees. We found that the topologies of the 16S rDNA and gyrB trees were similar except for Comamonas acidovorans and strains rM4 and rJ12. Strains affiliated with the R6 population formed a monophyletic cluster on the 16S rDNA tree, while these strains clearly occurred in two distinct groups on the gyrB tree. Similarly, strains affiliated with the R5 population occurred in several groups on the gyrB tree but not on the 16S rDNA tree. The trees suggest that most of the isolates are affiliated with the beta subclass of the class Proteobacteria.
Physiological characterization of isolates.
Figure 1 also shows some growth characteristics of the isolated bacteria. The strains affiliated with the R6 and R10 populations could grow on phenol (at a concentration of 50 mg per liter), while the strains affiliated with the R4, R5, and R9 populations could not. The strains affiliated with the R6 population were divided into the following two groups on the basis of their growth characteristics on phenol: floc-forming strains and non-floc-forming strains. Interestingly, these groups are consistent with the phylogenetic groups based on the gyrB sequences; accordingly, these groups were designated the R6F and R6T populations. Figure 1 also shows that the strains affiliated with the R10 population did not form flocs when they were grown on phenol. Sugar utilization by the isolated bacteria was also examined (Fig. 1). Figure 1 shows that there were several differences between the R6F and R6T strains; most noticeably, the R6F strains formed flocs when they were grown on sugars, whereas the R6T strains did not. We found that most of the bacterial strains formed flocs when they were grown on galactose. The strains in the R10 population formed flocs only in the presence of galactose.
cPCR.
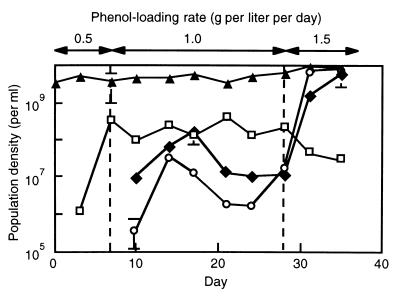
cPCR analyses to quantify the R6F, R6T, and R10 populations were conducted in order to examine (i) the quantitativeness of the TGGE analysis and (ii) differences in the dynamics of the R6F and R6T populations. The specificity of the gyrB-targeted PCR for each of the three populations was confirmed by analyzing the isolated strains (data not shown). Figure 4 shows the population dynamics determined by the cPCR. Although the R6 band was always present in the TGGE gel from day 7 on (Fig. 3), a shift from the R6F population to the R6T population was observed by using the gyrB-targeted cPCR; the density of the R6T population dramatically increased after the phenol-loading rate was increased to 1.5 g per liter per day. The cPCR also confirmed that the R10 population became dominant after the phenol-loading rate was increased to 1.5 g per liter per day (just before the breakdown of the activated-sludge process). The results of the TGGE and cPCR analyses were consistent, suggesting that the major bands which appeared on the TGGE gel (Fig. 3) corresponded to major populations in the activated sludge.
FIG. 4.
Dynamics of the R6F (□), R6T (⧫), and R10 (○) populations in the phenol-digesting activated sludge, as determined by the gyrB-targeted cPCR. The total direct count (▴) is also shown. Each datum point is the mean based on two or three determinations, and each error bar indicates the standard error.
DISCUSSION
We previously described a kinetic analysis of the phenol-oxygenating activity of phenol-acclimated Ohdaira activated sludge (35). When the MLSS concentration is 2,200 mg per liter (the average value in this study), the Vmax (11 U per g of dry cells) of the activated sludge corresponds to a phenol degradation rate of 1.7 g per liter per day (30). The phenol-oxygenating activities of the activated sludge determined in this study at phenol-loading rates of 0.5 and 1.0 g per liter per day (Fig. 2b) were equal to or somewhat greater than the activity found in the previous study. Therefore, we expected that the activated sludge might cope with phenol at a phenol-loading rate of 1.5 g per liter per day or higher. Unexpectedly, however, the activated sludge could not cope with phenol at this loading rate (Fig. 2a). We thought that the breakdown resulted from population shifts in the activated sludge after the increase in the loading rate because (i) phenol was almost completely digested for the first several days after the loading rate was increased to 1.5 g per liter per day and (ii) deflocculation preceded the breakdown. Actually, population shifts before the breakdown were observed in the TGGE (Fig. 3) and cPCR (Fig. 4) analyses. By combining the data on the population shifts and the data on the physiological characteristics of the corresponding isolates (Fig. 1), we concluded that the outbreak of nonflocculating phenol-degrading bacteria caused the breakdown of the activated-sludge process.
To maintain the MLSS concentration at a constant level, the sludge residence time (SRT) was shortened in response to the increase in the phenol-loading rate. Several investigators have suggested that the species compositions of heterogeneous cultures vary with changes in the dilution rate even when other environmental factors remain constant (19). The decrease in the SRT could have been a direct cause which provoked the population shifts, although the short SRT may have had an indirect effect on the bacterial population structure (i.e., a change in the protozoan grazing pressure) (1). With the short SRT, many of slowly growing protozoans which grazed on free-living bacteria could have been washed out, resulting in the increases in the population densities of the nonflocculating bacteria.
TGGE (13, 34) and denaturing gradient gel electrophoresis (11, 13, 18) coupled with PCR amplification of heterogeneous 16S rDNA fragments have been widely used to detect natural microbial populations, although attention should be given to the quantitative interpretation of the results of these analyses. For this reason, in our previous study a functionally dominant phenol-degrading population in activated sludge was assessed by the following three approaches: a TGGE analysis of 16S rDNA fragments to detect bacterial populations, a TGGE analysis of phenol hydroxylase gene fragments to detect enzyme populations, and a kinetic analysis of phenol-oxygenating activities expressed by bacteria (35). In this study, we used cPCR for quantitative backup of the TGGE results. As demonstrated in this study, a TGGE (denaturing gradient gel electrophoresis) profile would be a useful tool in microbial ecology, if this convenient method is combined with another quantitative method.
The gyrB-targeted phylogenetic analysis (Fig. 1) showed that one of the dominant phenol-degrading populations (the R6 band on the TGGE gel) was composed of two physiologically and genetically different groups of bacteria (the R6F and R6T populations). It is interesting that the gyrB-based grouping of the R6 bacteria was consistent with the grouping based on physiology. The almost full-length 16S rDNA sequences of the bacterial strains in the R6 group were identical except for one nucleotide in the sequence of strain rA7, so 16S rRNA-targeted population analyses, for instance fluorescence in situ hybridization (29), could not be used to detect the R6F and R6T populations in the sludge. Instead, cPCR of the gyrB genes was used for this purpose. This study thus demonstrated the usefulness of gyrB-targeted fine population analyses in microbial ecology, as demonstrated in our previous study (33). The ICB database (16) at our institute, which is accessible through the Internet, should facilitate the design of gyrB-targeted PCR primers to be used in such studies.
The results of the present study suggest that if the growth of nonflocculating phenol-degrading populations (i.e., the R6T and R10 populations) could be suppressed and if the flocculating phenol-degrading population (i.e., the R6F population) could be sustained in the activated sludge, the activity of the activated sludge process would be high enough to cope with phenol even under the high-phenol-loading-rate condition. To achieve this, the following two methods are conceivable: selective biostimulation of the flocculating population (e.g., by adding preferential growth substrates for the flocculating population) and bioaugmentation with flocculating phenol-degrading bacteria. These methods are currently being examined in our laboratory.
ACKNOWLEDGMENTS
We thank Ikuko Hiramatsu for technical assistance, Robert Kanaly for assistance in preparation of the manuscript, and Mitsuhiro Konno (Ohdaira Wastewater Treatment Plant, Kamaishi, Iwate, Japan) for kind help in sampling activated-sludge mixed liquor.
This work was supported by New Energy and Industrial Technology Development Organization (NEDO).
REFERENCES
- 1.Amann R, Lemmer H, Wagner M. Monitoring the community structure of wastewater treatment plants: a comparison of old and new techniques. FEMS Microbiol Ecol. 1998;25:205–216. [Google Scholar]
- 1a.American Petroleum Institute. Manual on the disposal of refinery wastes. Volume on liquid waste. Washington, D.C: American Petroleum Institute; 1969. [Google Scholar]
- 2.Beltrame P, Beltrame P L, Carniti P, Pitea D. Kinetics of phenol degradation by activated sludge: value of measurements in a batch reactor. Water Res. 1979;13:1305–1309. [Google Scholar]
- 3.Blattner F R, Plunkett G, 3rd, Bloch C A, Perna N T, Burland V, Riley M, Collado-Vides J, Glasner J D, Rode C K, Mayhew G F, Gregor J, Davis N W, Kirkpatrick H A, Goeden M A, Rose D J, Mau B, Shao Y. The complete genome sequence of Escherichia coli K-12. Science. 1997;277:1453–1474. doi: 10.1126/science.277.5331.1453. [DOI] [PubMed] [Google Scholar]
- 4.Bond P L, Hugenholtz P, Keller J, Blackall L L. Bacterial community structures of phosphate-removing and non-phosphate-removing activated sludge from sequencing batch reactors. Appl Environ Microbiol. 1995;61:1910–1916. doi: 10.1128/aem.61.5.1910-1916.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Brosius J, Dull T J, Sleeter D D, Noller H F. Gene organization and primary structure of a ribosomal RNA operon from Escherichia coli. J Mol Biol. 1981;148:107–127. doi: 10.1016/0022-2836(81)90508-8. [DOI] [PubMed] [Google Scholar]
- 6.Cote R J, Gherna R L. Nutrition and media. In: Gerhardt P, Murray R G E, Wood W A, Krieg N R, editors. Methods for general and molecular bacteriology. Washington, D. C: American Society for Microbiology; 1994. pp. 155–178. [Google Scholar]
- 7.Dojka M A, Hugenholtz P, Haack S K, Pace N R. Microbial diversity in a hydrocarbon- and chlorinated-solvent-contaminated aquifer undergoing intrinsic bioremediation. Appl Environ Microbiol. 1998;64:3869–3877. doi: 10.1128/aem.64.10.3869-3877.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dubois M, Gilles K A, Hamilton J K, Rebers P A, Smith F. Colorimetric method for determination of sugars and related substances. Anal Chem. 1956;28:350–356. [Google Scholar]
- 9.Eckenfelder W W, Musterman J L. Activated-sludge treatment of industrial wastewater. Lancaster, Pa: Technomic; 1995. [Google Scholar]
- 10.Edwards U, Rogall T, Blöcker H, Emde M, Böttger E C. Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res. 1989;17:7843–7853. doi: 10.1093/nar/17.19.7843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ferris M J, Ward D M. Seasonal distributions of dominant 16S rRNA-defined populations in a hot spring microbial mat examined by denaturing gradient gel electrophoresis. Appl Environ Microbiol. 1997;63:1375–1381. doi: 10.1128/aem.63.4.1375-1381.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Harayama S, Yamamoto S. Phylogenetic identification of Pseudomonas strains based on a comparison of gyrB and rpoD sequences. In: Nakazawa T, Furukawa K, Haas D, Silver S, editors. Molecular biology of pseudomonads. Washington, D.C: American Society for Microbiology; 1996. pp. 250–258. [Google Scholar]
- 13.Heuer H, Krsek M, Baker P, Smalla K, Wellington E M H. Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel electrophoretic separation in denaturing gradients. Appl Environ Microbiol. 1997;63:3233–3241. doi: 10.1128/aem.63.8.3233-3241.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hino H, Watanabe K, Takahashi N. Phenol hydroxylase cloned from Ralstonia eutropha strain E2 exhibits novel kinetic properties. Microbiology. 1998;144:1765–1772. doi: 10.1099/00221287-144-7-1765. [DOI] [PubMed] [Google Scholar]
- 15.Karlin S, Altschul S F. Methods for assessing the statistical significance of molecular sequence features by using general scoring schemes. Proc Natl Acad Sci USA. 1990;87:2264–68. doi: 10.1073/pnas.87.6.2264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Marine Biotechnology Institute. ICB database. Iwate, Japan: Marine Biotechnology Institute; 1997. [Google Scholar]
- 17.McMacken R, Silver L, Geogopoulos C. DNA replication. In: Neidhardt F C, Ingraham J L, Low K B, Magasanik B, Schaechter M, Umbarger H E, editors. Escherichia coli and Salmonella typhimurium: cellular and molecular biology. Washington, D.C: American Society for Microbiology; 1987. pp. 578–580. [Google Scholar]
- 18.Muyzer G, de Waal E C, Uitterlinden A G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol. 1993;59:695–700. doi: 10.1128/aem.59.3.695-700.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Okaygun M S, Akgerman A. Microbial dynamics in a continuously stirred tank reactor with 100% cell recycle. Water Environ Res. 1992;64:811–816. [Google Scholar]
- 20.Olsen G J, Lane D L, Giovannoni S J, Pace N R. Microbial ecology and evolution: a ribosomal RNA approach. Annu Rev Microbiol. 1986;40:337–365. doi: 10.1146/annurev.mi.40.100186.002005. [DOI] [PubMed] [Google Scholar]
- 21.Pace N R, Stahl D A, Lane D L, Olsen G J. The analysis of natural microbial populations by rRNA sequences. Adv Microb Ecol. 1986;9:1–55. [Google Scholar]
- 22.Rebhun M, Galil N. Inhibition by hazardous compounds in an integrated oil refinery. J Water Pollut Control Fed. 1988;60:1953–1959. [Google Scholar]
- 23.Rozich A F, Gaudy A F., Jr Response of phenol-acclimated activated sludge process to quantitative shock loading. J Water Pollut Control Fed. 1985;57:795–804. [Google Scholar]
- 24.Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–425. doi: 10.1093/oxfordjournals.molbev.a040454. [DOI] [PubMed] [Google Scholar]
- 25.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1989. [Google Scholar]
- 26.Shingler V. Molecular and regulatory check points in phenol degradation by Pseudomonas sp. CF600. In: Nakazawa T, Furukawa K, Haas D, Silver S, editors. Molecular biology of pseudomonads. Washington, D.C: American Society for Microbiology; 1996. pp. 153–164. [Google Scholar]
- 27.Snaidr J, Amann R, Huber I, Ludwig W, Schleifer K H. Phylogenetic analysis and in situ identification of bacteria in activated sludge. Appl Environ Microbiol. 1997;63:2884–2896. doi: 10.1128/aem.63.7.2884-2896.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Thompson J D, Higgins D G, Gibson T J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wagner M, Amann R, Lemme H, Schleife K. Probing activated sludge with oligonucleotide specific for proteobacteria: inadequacy of culture-dependent methods for describing microbial community structure. Appl Environ Microbiol. 1993;59:1520–1525. doi: 10.1128/aem.59.5.1520-1525.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Watanabe K, Hino S, Onodera K, Kajie S, Takahashi N. Diversity in kinetics of bacterial phenol-oxygenating activity. J Ferment Bioeng. 1996;81:562–565. [Google Scholar]
- 31.Watanabe K, Hino S. Identification of a functionally important population in phenol-digesting activated sludge with antisera raised against isolated bacterial strains. Appl Environ Microbiol. 1996;62:3901–3904. doi: 10.1128/aem.62.10.3901-3904.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Watanabe K, Hino S, Takahashi N. Responses of activated sludge to an increase in phenol loading. J Ferment Bioeng. 1996;82:522–524. [Google Scholar]
- 33.Watanabe K, Yamamoto S, Hino S, Harayama S. Population dynamics of phenol-degrading bacteria in activated sludge determined by gyrB-targeted quantitative PCR. Appl Environ Microbiol. 1998;64:1203–1209. doi: 10.1128/aem.64.4.1203-1209.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Watanabe K, Harayama S. Rapid estimation of population densities of uncultured bacteria in the environment. Microbes Environ. 1998;13:123–127. [Google Scholar]
- 35.Watanabe K, Teramoto M, Futamata H, Harayama S. Molecular detection, isolation, and physiological characterization of functionally dominant phenol-degrading bacteria in activated sludge. Appl Environ Microbiol. 1998;64:4396–4402. doi: 10.1128/aem.64.11.4396-4402.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yamamoto S, Harayama S. PCR amplification and direct sequencing of gyrB genes with universal primers and their application to the detection and taxonomic analysis of Pseudomonas putida strains. Appl Environ Microbiol. 1995;61:1104–1109. doi: 10.1128/aem.61.3.1104-1109.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]